1. Introduction
Colorectal cancer (CRC), the third most common cancer worldwide, is a global health problem resulting in approximately 1.9 million new cases and 930,000 deaths according to 2020 data [
1]. Current demographic projections predict this burden will reach 3.2 million new cases by 2040 [
1]. While current therapeutic approaches primarily target actively proliferating cells, the development of chemoresistance and systemic toxicity necessitate the development of more specific and effective synthetic agents [
2].
Benzimidazole, a key pharmacophore with an aromatic heterocyclic core, finds widespread use in drug research [
3]. This framework has served as the basis for numerous synthetic derivatives exhibiting a variety of biological activities, including antibacterial, antiviral, antifungal, and antiproliferative effects [
4]. Benzimidazole derivatives, in particular, have attracted attention as target-based chemical frameworks in cancer treatment and have been studied by researchers in various cancer cell lines [
5,
6]. Benzimidazole derivatives exhibit potent anticancer effects by inhibiting cancer cell proliferation, inducing apoptosis, and modulating critical cellular signaling pathways [
7,
8]. These molecules possess multifaceted mechanisms for cell cycle arrest through DNA binding, disruption of microtubule dynamics, and specific enzyme inhibition [
9]. In addition to these comprehensive effects of benzimidazole derivatives, the inhibitory roles of benzimidazolium salts, particularly on the STAT3 signaling pathway, have been demonstrated [
10].
Autophagy is a bidirectional mechanism in CRC that plays both suppressive and supportive roles in tumor development by regulating stress adaptation [
11]. In CRC, the autophagy process is a central survival mechanism tightly controlled by the interaction between STAT3 and mTOR signaling pathways. The fact that STAT3 inhibition triggers autophagy by suppressing mTOR activity makes this axis a critical molecular target in terms of tumor progression and treatment response [
12]. The lethal autophagy mechanism, as it is called, can transform this process into a death mechanism when cellular stress exceeds a certain threshold [
13]. This phenomenon triggers pro-apoptotic signals from proteins such as Beclin-1 and Atg5, cleaved by caspases, leading the cell into an irreversible process of destruction. Consequently, the synergistic interaction between this initially protective mechanism and apoptosis presents a strategic therapeutic target that overcomes the chemoresistance shields of cancer cells and drives them into programmed cell death [
14].
Further molecular and mechanistic studies supporting the anticancer effects of benzimidazolium salts are of great importance in determining the place of these compounds in the rational drug development process. This work begins with benzimidazoline salts synthesized by Akkoç, which were initially evaluated for efficacy by MTT-based cytotoxicity experiments [
15]. Our preliminary experimental observations revealed that these compounds not only reduce cell viability but also cause significant changes in cell morphology. In particular, the formation of autophagosome-like vacuolar structures and a significant decrease in the adhesive properties of cells after drug application were observed. This series, exhibiting low IC₅₀ values and morphologically diverse effects, suggests that mechanisms beyond classical apoptotic cell death may be at play. Therefore, a more detailed evaluation of these compounds in different cancer cell types and elucidation of their mechanisms of action via apoptosis-autophagy-related molecular pathways constitute the main objective of the present study.
2. Materials and Methods
2.1. Synthesis of Benzimidazole-Based Compounds
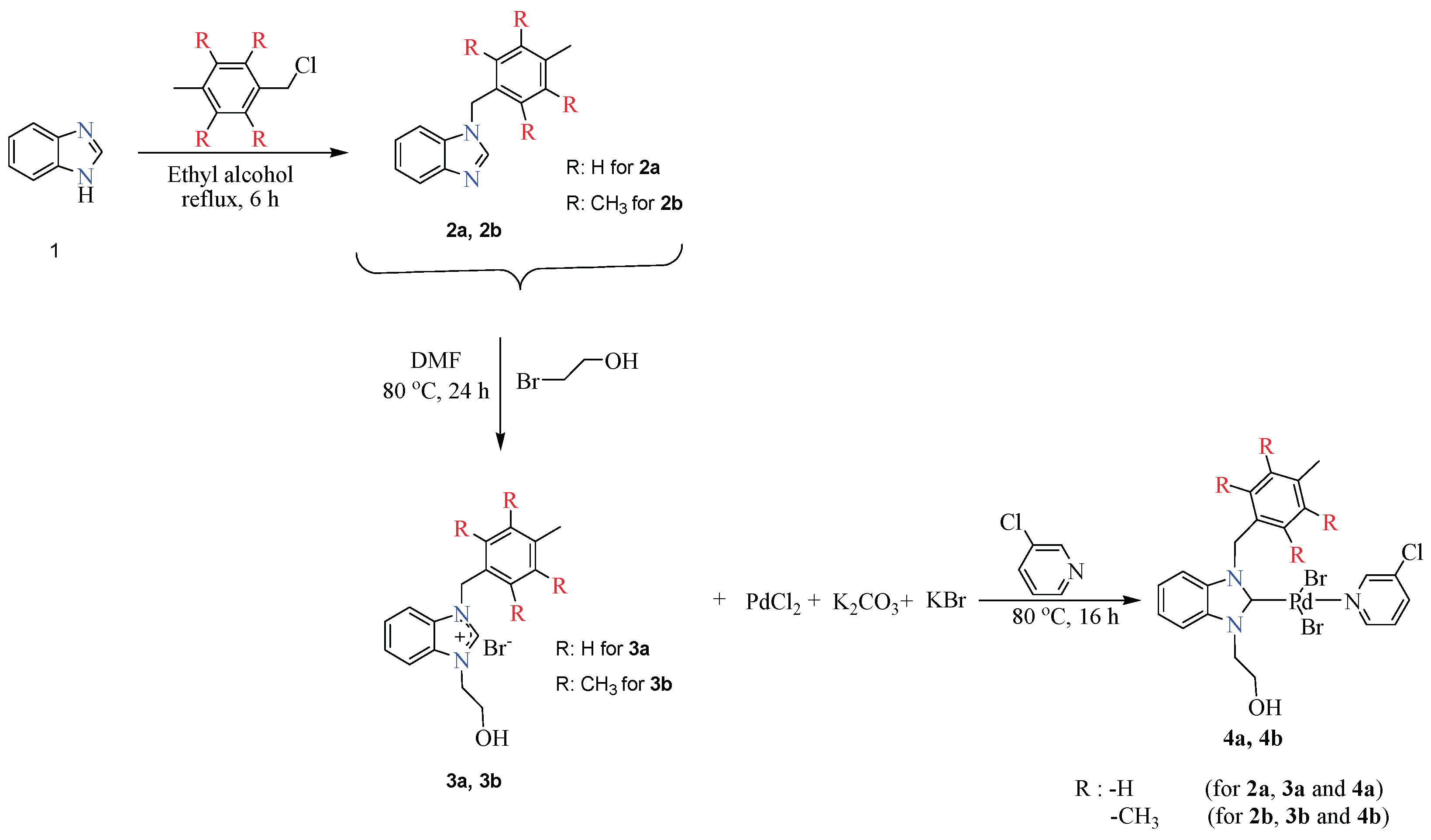
The 2-hydroxyethyl substituted benzimidazolium salts evaluated in this study and the palladium complexes derived from these salts were successfully synthesized in a previous study by our research group (
Scheme 1) [
15]. The synthesis processes, structural characterizations, and fundamental physical properties of these compounds are reported in detail in the relevant study. The relevant publication comprehensively presents the structural characterization (¹H NMR, ¹³C NMR, FT-IR, elemental analysis, etc.), physical properties, and fundamental chemical data of the synthesized benzimidazolium salts and palladium complexes.
In our previous study, the antiproliferative activities of 2-hydroxyethyl substituted benzimidazolium salts, which are 1-(2-hydroxyethyl)-3-(4-methylbenzyl)-1H-benzo[d]imidazol-3-ium bromide (3a), 1-(2-hydroxyethyl)-3-(2,3,4,5,6-pentamethylbenzyl)-1H-benzo[d]imidazol-3-ium bromide (3b), and their palladium complexes, which are dibromo-[1-(2-hydroxyethyl)-3-(4-methylbenzyl)benzimidazol-2-ylidene]-N-(3-chloropyridine) palladium (II) complex (4a), dibromo-[1-(2-hydroxyethyl)-3-(2,3,4,5,6-pentamethylbenzyl)benzimidazol-2-ylidene]-N-(3-chloropyridine) palladium (II) complex (4b), were evaluated in four different cancer cell lines (HeLa, MDA-MB-231, HepG2, and DLD-1) using the MTT assay method. The results obtained revealed that the organic benzimidazolium salts possess significantly higher cytotoxic potential compared to their metal complexes. Specifically, compound 3b exhibited measurable cytotoxic activity with an IC₅₀ value of 80.64 ± 1.25 µM in the HeLa cell line, while the palladium complexes (4a and 4b) did not show any antiproliferative effect in the same cell line. Similarly, compounds 3a and 3b showed moderate cytotoxic activity in HepG2 cells, while the metal complexes were also found to be ineffective against this cell line.
The results obtained in MDA-MB-231 and DLD-1 cell lines more clearly demonstrated the potent and selective antiproliferative effects of organic compounds. In MDA-MB-231 breast cancer cells, compound
3b in particular showed the highest cytotoxic activity with an IC₅₀ value of 7.59 ± 0.68 µM and was found to be more effective than cisplatin, which was used as a positive control. In addition, compounds
3a and
3b were found to have a dose-dependent effect against breast cancer cells. In the DLD-1 CRC cell line, compound
3a showed a remarkable antiproliferative effect with an IC₅₀ value of 9.19 ± 1.51 µM, which was very close to the IC₅₀ value of cisplatin. In contrast, compounds
4a and
4b did not show cytotoxic activity in either cell line. These results indicate that the 2-hydroxyethyl substituted benzimidazolium salts synthesized in our previous study [
15] are promising anticancer candidates, particularly MDA-MB-231 and DLD-1 cancer cells, and provide a strong basis for further biological studies. Therefore, mecanistic experiments were conducted in this study.
2.2. Cell Culture
Human breast cancer cell line MDA-MB-231, mouse breast cancer cell line 4T1, DLD-1 and HT-29 human CRC cell lines were obtained from the American Type Culture Collection (ATCC). MDA-MB-231, 4T1, DLD-1 and HT-29 cells were cultured in a single layer in RPMI 1640 or Dulbecco's Modified Eagle Medium (DMEM, Gibco) supplemented with 10% fetal bovine serum (FBS, Gibco), 4.5 g/L L-glutamine, 1 mM sodium pyruvate, 100 U/mL penicillin-streptomycin and 1% amphotericin B. Cultures were maintained at 37°C in a humidified incubator containing 5% CO₂.
2.3. MTT Cell Viability Test
The cytotoxic effect of compounds 3a, 3b, 4a and 4b on MDA-MB-231, 4T1, DLL-1, and HT-29 cells was evaluated using the MTT [3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide] test. Cells were seeded into 96-well plates at a cell density of 6×10³ per well and incubated overnight. They were then treated with various concentrations of the compound (1–160 µM) for 48 hours. After treatment, 10 µL of MTT (5 mg/ml) solution was added to each well and incubated at 37 °C for 4 hours, and 50 µL of DMSO was added, and absorbance was measured at 570 nm using a microplate reader. Cell viability was expressed as a percentage compared to an untreated control group, which was considered to be 100%.
2.4. Immunofluorescence (IF) Staining
HT-29 and DLD-1 colorectal cancer cells were seeded onto sterile glass coverslips placed in 6-well culture plates and subsequently subjected to 3a treatment under the specified experimental conditions. Upon completion of the incubation period, cells were fixed in 4% paraformaldehyde for 15 min at room temperature and rinsed thoroughly with phosphate-buffered saline (PBS). Cellular permeabilization was achieved using Triton X-100 for 10 min at room temperature, followed by blocking of non-specific antibody binding with 5% bovine serum albumin (BSA). Immunostaining was performed by incubating the cells overnight at 4 °C in a humidified chamber with primary antibodies targeting LC3 (bs-8878R, Bioss) and mTOR (bs-1992R, Bioss), each diluted 1:200 in blocking buffer. After extensive washing with PBS, cells were incubated with a FITC-conjugated secondary antibody for 1 h at room temperature. Nuclear counterstaining and mounting were carried out using a DAPI-containing mounting medium (Sigma). Fluorescence images were acquired using an Olympus BX53 fluorescence microscope equipped with a DP74 camera system. Quantitative image analysis was conducted using ImageJ software. Mean fluorescence intensity values were determined from 30 cells per condition, selected from randomly chosen microscopic fields
2.5. Annexin V Apoptotic Analysis
To evaluate the apoptotic effects of compound 3a, DLD-1 and HT-29 cells were treated for 48 hours and then stained with Annexin V-APC and Propidium Iodide (PI). After staining, they were analyzed by flow cytometry to distinguish between viable and apoptotic cell populations. Apoptotic percentages were measured for each group and compared with controls.
2.6. Cell Migration Experiment
A wound healing experiment was performed to evaluate the effects of compound
3a on cell migration. Details of the method are given in our previous study [
16]. Briefly, HT-29 CRC cells were seeded into 6-well plates at a density of 10*105 cells per well. After the cells reached approximately 70-80% density, at the end of 48 hours of incubation, a wound was created on the culture surface using a sterile 100 µL pipette tip. Wound widths were visualized at 0 and 48 hours and then percentages were determined using ImageJ software.
2.7. Cell Adhesion Experiment
HT-29 CRC cells were treated with compound
3a for 24 hours. Control group cells received no treatment. Then, the cell adhesion experiment was performed according to the literature [
17]. Microplate wells were coated with 1% Poly-D-lysine and blocked with heat-denatured bovine serum albumin (BSA) to prevent non-specific binding. After fixing the cells with methanol, they were stained with 0.1% crystal violet, and the amount of bound cells was visualized by removing excess dye. In the final step, the dye bound to the cells was dissolved in 10% acetic acid and analyzed spectrophotometrically at a wavelength of 570 nm in a microplate reader.
2.8. Gene Expression Analysis
HT-29 cells were incubated with compound 3a at an IC50 (3.7 µM) concentration for 48 hours, followed by total RNA isolation using Trizol reagent (Invitrogen, USA) according to the manufacturer's instructions. cDNA synthesis was performed from the isolated RNA samples via reverse transcriptase enzyme; then, the expression levels of apoptosis-related (BCL-2, BAX, and Caspase-3), autophagy-related genes (Mtor and BECN1), adhesion-related (VIM and ITGβ1), AKT, and Stat3 genes were analyzed using quantitative real-time PCR (qPCR). Gene expression data were normalized relative to the beta-actin gene used as an internal control, and relative mRNA changes were calculated using the 2-∆∆Ct method. For statistical comparisons, the control group expression level was fixed at 1, and fold changes of 3a application on target genes were determined relative to this reference value.
2.9. Statistical Analysis
Statistical analyses were performed using GraphPad Prism software. Student t-test was applied for pairs of samples. For data with more than two samples, one-way ANOVA followed by Tukey's multiple comparison test was applied, IC50 values and confidence intervals were obtained from nonlinear regression analyses. A p-value < 0.05 was considered statistically significant.
4. Discussion
CRC, has become a global health crisis, presents a difficult barrier to overcome in current treatment protocols, particularly due to chemoresistance mechanisms developing in metastatic processes [
18]. Despite the widespread application of conventional cytotoxic agents, the ability of cancer cells to evade apoptosis through genomic instability and disrupt DNA repair mechanisms significantly limits the clinical efficacy of existing regimens. To overcome these treatment bottlenecks and control resistant cell populations, the discovery of innovative synthetic compounds that simultaneously target cellular survival and adhesion dynamics is an indispensable priority for modern oncology research [
18,
19]. Benzimidazole salts stand out for their superior solubility and stability, which are critical for overcoming the bioavailability hurdles of neutral compounds. Their unique ionic profile enables more precise interaction with cellular targets, effectively dismantling oncogenic signaling to drive resistant cancer cells toward programmed death [
5,
20].
Benzimidazole derivative drugs have been proven to have a highly effective anticancer effect in many cancers, particularly colon cancer [
5,
6,
7,
21]. Our previous study showed that organic benzimidazolium salts (
3a and 3b) have a much higher cytotoxic potential compared to metal complexes (4a and 4b) and exhibit antiproliferative activity comparable to cisplatin, particularly in DLD-1 and MDA-MB-231 cell lines
15. In this study, the effects of these four compounds on cell proliferation were investigated in aggressive mouse breast cancer cells (4T1), aggressive human breast cancer cells (MDA-MB231), and aggressive HT-29 and DLD-1 cells. Similar to the previous study,
3a was the most effective compound in colon cancer, while
3a and 3b showed similar levels of anticancer activity in breast cancer. 4a and 4b did not show any anticancer activity, as in the previous study. In this case, the properties of benzimidazole suggest that the 2-hydroxyethyl substituent interacts more effectively with biological macromolecules in its free form, but complexation with palladium significantly reduces its anticancer activity by masking or halting the molecule's active pharmacological sites.
Autophagy functions as a protective mechanism in the early stages of cancer, preventing tumor formation by maintaining genomic stability, while in advanced stages, its over-induction becomes a strategic therapeutic weapon, triggering programmed cell death by creating a metabolic impasse in cancer cells [
22]. LC3, a specific marker of autophagosome formation, is widely used in autophagy measurements [
23]. One of the striking findings of this study is that
3a significantly induces autophagy. Immunofluorescence analyses show an increase in LC3 signaling and suppression of mTOR activity, indicating that
3a activates the autophagic flow. Autophagy can play both tumor-suppressing and tumor-promoting roles in CRC, depending on the context. Particularly in advanced cancers, autophagy can support cell survival under metabolic stress, while excessive or dysregulated autophagy can lead to cell death [
24]. In this study, the observation of autophagy induction along with apoptosis suggests that
3a drives cells not towards protective autophagy, but towards a process described as "lethal autophagy." Numerous protein molecules involved in tumor pathogenesis have been found to influence tumor formation by regulating autophagy and apoptosis; most of these are via the autophagy protein LC3, beclin-1, the apoptosis protein family Bcl-2, BAX, or the interaction between the two
[25]. Autophagy and apoptosis are in constant interaction via common molecular bridges such as the BH3 domain and p53 to maintain cellular homeostasis and make decisions about survival or death under stressful conditions [
26]. Apoptosis analyses showed that compound
3a caused significant apoptotic cell death in both HT-29 and DLD-1 cells. This finding, confirmed by Annexin V/PI assays, is also supported by gene expression results. The decrease in anti-apoptotic BCL-2 expression, accompanied by an increase in pro-apoptotic BAX and caspase-3 expression, suggests that
3a activates the intrinsic apoptotic pathway. Given that apoptotic mechanisms are frequently suppressed in CRC, contributing to chemotherapy resistance, the ability of
3a to restore apoptotic balance is of therapeutic importance. It is thought that compound
3a causes cell apoptosis by triggering lethal autophagy.
The AKT/mTOR/STAT3 signaling pathways are key regulators of proliferation, survival, migration, and chemotherapy resistance in CRC cells [
27]. Gene expression analyses revealed that compound
3a significantly suppressed AKT, mTOR, and STAT3 expression. The literature reports that constitutive activation of STAT3 is strongly associated with CRC progression, poor prognosis, and metastasis. STAT3 can also suppress autophagy by activating the mTOR pathway [
28]. Therefore, the inhibition of the STAT3 and mTOR axis by compound
3a is mechanistically consistent with the activation of autophagy and the attenuation of cellular survival signals.
Functional analyses demonstrate that molecular findings directly reflect cellular behavior. In wound-healing, cell adhesion, and colony formation experiments,
3a was found to significantly inhibit the migration capacity, surface adhesion ability, and clonal growth of HT-29 cells. These findings are consistent with the decrease in Vimentin (VIM) and integrin β1 (ITGB1) expression observed in gene expression analyses. Integrins and EMT-related proteins play a critical role in CRC metastasis, and suppression of these proteins significantly reduces the invasive potential of tumor cells
[29,30].
The literature increasingly features investigations on the anticancer potential of benzimidazole derivatives, however most of these molecules lack rigorous mechanistic evaluations
[8]. This study is important because it shows that
3a not only suppresses proliferation but also reverses the autophagy-apoptosis balance against the tumor by targeting the AKT/mTOR/STAT3 axis. Especially in a heterogeneous and treatment-resistant tumor type like CRC, such multi-target agents hold promise for next-generation therapeutic strategies.
5. Conclusions
Although this study has some limitations, the findings clearly demonstrate that compound 3a exhibits a multifaceted anticancer effect in colon cancer cells. While the study was limited to in vitro models, the simultaneous induction of autophagy and apoptosis by 3a, and the association of these processes with the suppression of AKT/mTOR/STAT3 signaling pathways, provides strong clues about the compound's mechanism of action. Furthermore, the effective inhibition of migration, colony formation, and cell adhesion suggests that 3a can also target cellular characteristics associated with metastasis.
In conclusion, the benzimidazole derivative compound 3a emerges as a promising anticancer candidate that suppresses proliferation, triggers autophagy-mediated apoptotic cell death, and reduces metastatic potential in colon cancer cells. These findings strongly support the need for further evaluation of 3a's therapeutic potential through advanced mechanistic approaches and in vivo models.
Author Contributions
Conceptualization, M.Ç..; methodology, M.Ç. and E.A.; software, M.Ç. and E.A.; validation, M.Ç., E.A., S.A., H.Ö., H.A. and S.A.; formal analysis, M.Ç. and E.A.; investigation, M.Ç. and E.A.; resources, M.Ç.; data curation, M.Ç. and E.A.; writing—original draft preparation, M.Ç., E.A. and S.A.; writing—review and editing, M.Ç. and E.A.; visualization, M.Ç. and E.A.; supervision, M.Ç. ; project administration, M.Ç.; funding acquisition, M.Ç. . All authors have read and agreed to the published version of the manuscript.
Scheme 1.
Synthesis of compounds 2a-b, 3a-b and 4a-b.
Scheme 1.
Synthesis of compounds 2a-b, 3a-b and 4a-b.
Figure 1.
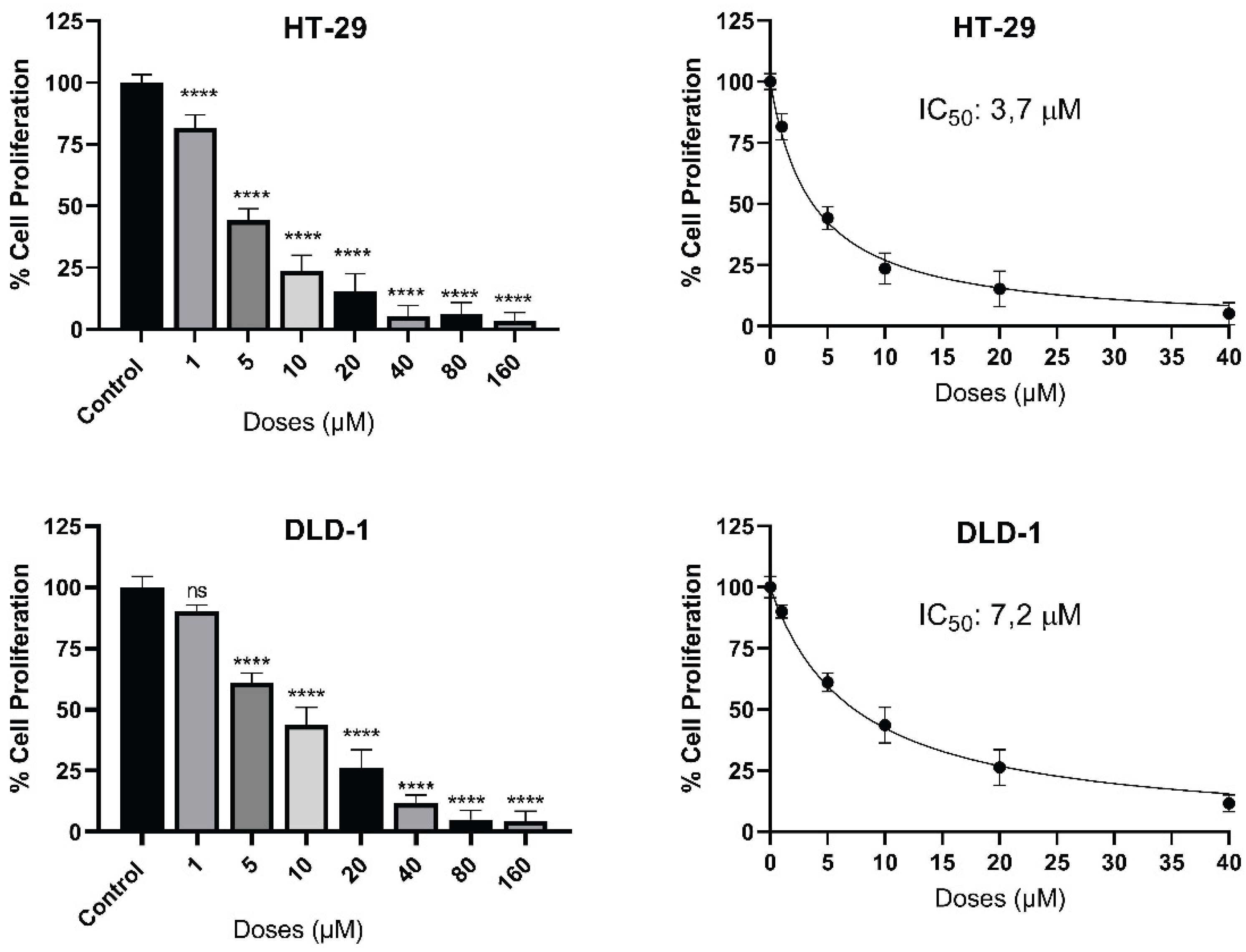
Cytotoxic profile and dose-response curves of compound 3a in colorectal cancer cells. The bar charts and non-linear regression curves illustrate a significant, dose-dependent inhibition of cell proliferation in both HT-29 and DLD-1 cell lines after treatment. While compound 3a exhibited high potency across both lines, HT-29 cells showed greater sensitivity with an IC50 of 3.7 µM, compared to 7.2 µM in DLD-1 cells. Statistically significant differences compared to the control group are indicated as ns: non signifant and **** p < 0.0001.
Figure 1.
Cytotoxic profile and dose-response curves of compound 3a in colorectal cancer cells. The bar charts and non-linear regression curves illustrate a significant, dose-dependent inhibition of cell proliferation in both HT-29 and DLD-1 cell lines after treatment. While compound 3a exhibited high potency across both lines, HT-29 cells showed greater sensitivity with an IC50 of 3.7 µM, compared to 7.2 µM in DLD-1 cells. Statistically significant differences compared to the control group are indicated as ns: non signifant and **** p < 0.0001.
Figure 2.
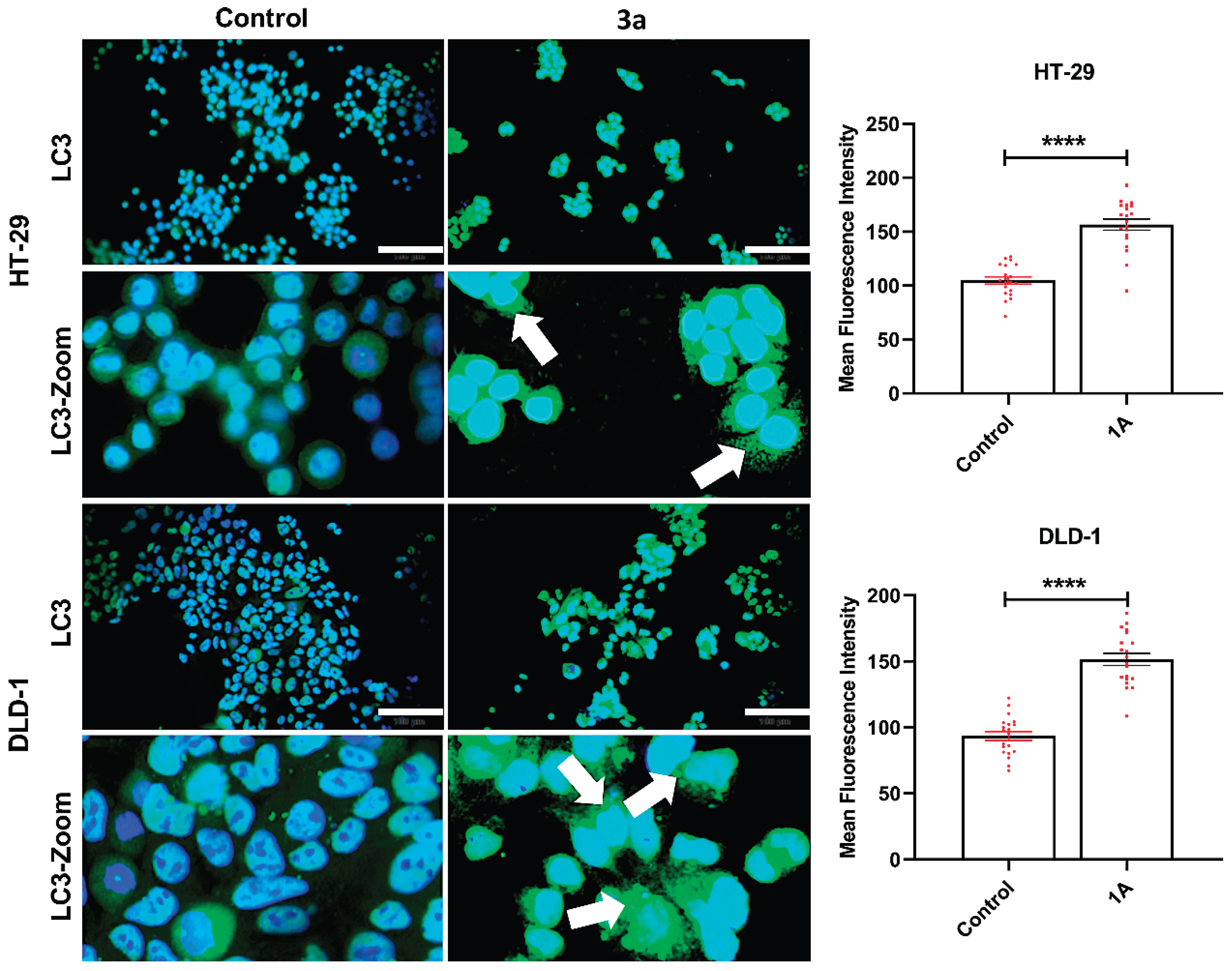
3a treatment enhances LC3 expression and induces punctate LC3 accumulation in colorectal cancer cells. Representative immunofluorescence images of LC3 staining in HT-29 and DLD-1 colorectal cancer cells under control conditions and following 3a treatment. Low-magnification images and corresponding high-magnification (zoom) views illustrate LC3 localization patterns. Quantitative fluorescence analysis revealed a significant increase in LC3 mean fluorescence intensity in both HT-29 and DLD-1 cells upon 3a exposure compared to controls (****p < 0.0001). Notably, 3a-treated cells exhibited prominent punctate LC3-positive structures within the cytoplasm (indicated by arrows), in contrast to the predominantly diffuse LC3 distribution observed in control cells. Nuclei were counterstained with DAPI. Scale bar: 100 µm.
Figure 2.
3a treatment enhances LC3 expression and induces punctate LC3 accumulation in colorectal cancer cells. Representative immunofluorescence images of LC3 staining in HT-29 and DLD-1 colorectal cancer cells under control conditions and following 3a treatment. Low-magnification images and corresponding high-magnification (zoom) views illustrate LC3 localization patterns. Quantitative fluorescence analysis revealed a significant increase in LC3 mean fluorescence intensity in both HT-29 and DLD-1 cells upon 3a exposure compared to controls (****p < 0.0001). Notably, 3a-treated cells exhibited prominent punctate LC3-positive structures within the cytoplasm (indicated by arrows), in contrast to the predominantly diffuse LC3 distribution observed in control cells. Nuclei were counterstained with DAPI. Scale bar: 100 µm.
Figure 3.
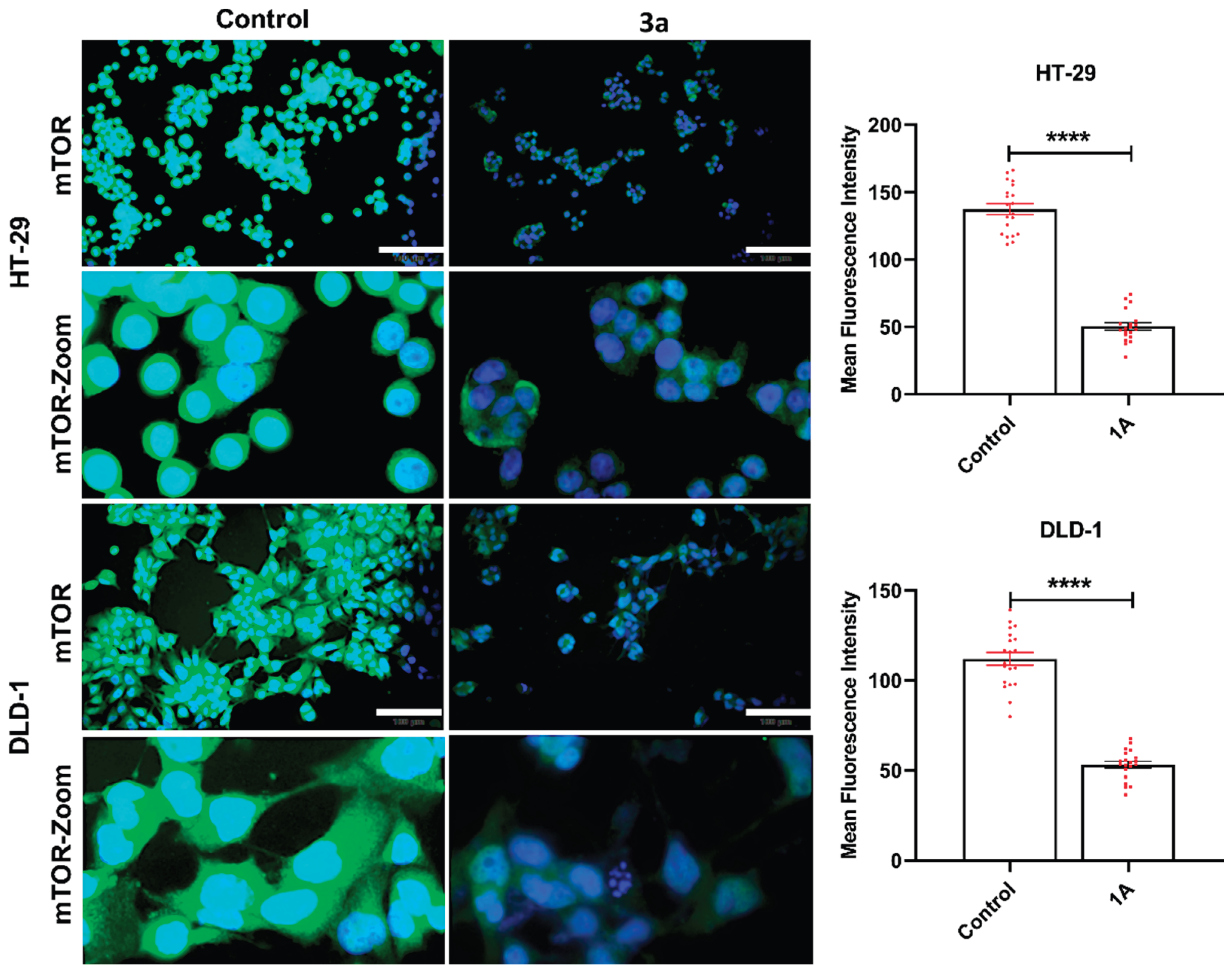
3a treatment reduces mTOR fluorescence intensity in colorectal cancer cells. Representative immunofluorescence images depicting mTOR expression in HT-29 and DLD-1 cells under control conditions and following 3a treatment, shown at low magnification and corresponding zoomed-in views. Quantitative analysis demonstrated a significant reduction in mTOR mean fluorescence intensity in both colorectal cancer cell lines upon 3a treatment compared to control cells (****p < 0.0001). The decrease in mTOR signal intensity was consistently observed across independent experiments. Nuclei were counterstained with DAPI. Scale bar: 100 µm.
Figure 3.
3a treatment reduces mTOR fluorescence intensity in colorectal cancer cells. Representative immunofluorescence images depicting mTOR expression in HT-29 and DLD-1 cells under control conditions and following 3a treatment, shown at low magnification and corresponding zoomed-in views. Quantitative analysis demonstrated a significant reduction in mTOR mean fluorescence intensity in both colorectal cancer cell lines upon 3a treatment compared to control cells (****p < 0.0001). The decrease in mTOR signal intensity was consistently observed across independent experiments. Nuclei were counterstained with DAPI. Scale bar: 100 µm.
Figure 4.
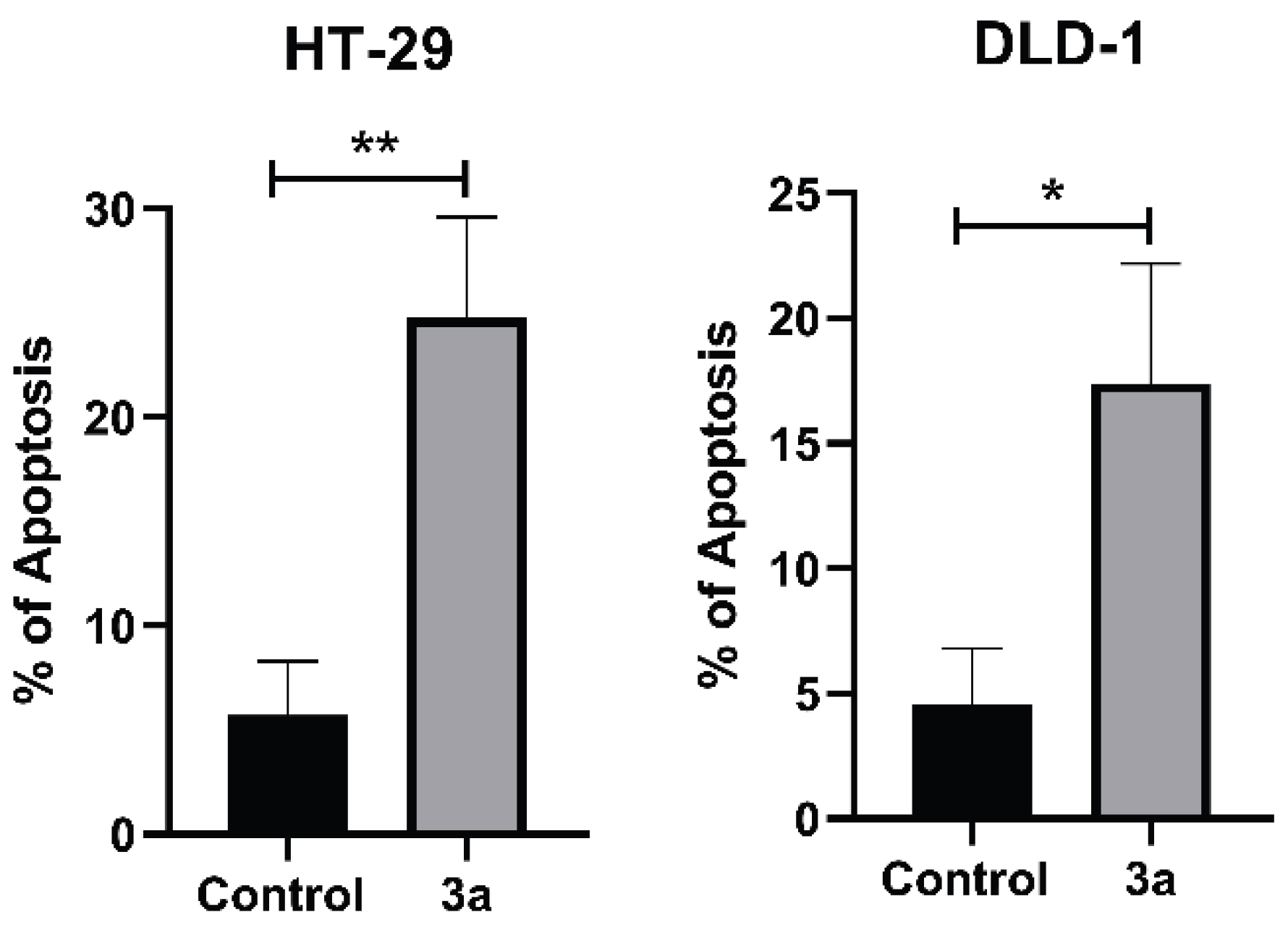
The effect of compound 3a on apoptosis induction colorectal cancer cells. HT-29 and DLD-1 colorectal cancer cells were treated with compound 3a at the respective IC50 concentrations for 48 hours. The percentage of apoptotic cells was determined by flow cytometry using Annexin V-FITC/PI double staining. Data are presented as mean ± SD of three independent experiments. Statistically significant differences compared to the control group are indicated as *p < 0.05 and **p < 0.01.
Figure 4.
The effect of compound 3a on apoptosis induction colorectal cancer cells. HT-29 and DLD-1 colorectal cancer cells were treated with compound 3a at the respective IC50 concentrations for 48 hours. The percentage of apoptotic cells was determined by flow cytometry using Annexin V-FITC/PI double staining. Data are presented as mean ± SD of three independent experiments. Statistically significant differences compared to the control group are indicated as *p < 0.05 and **p < 0.01.
Figure 5.
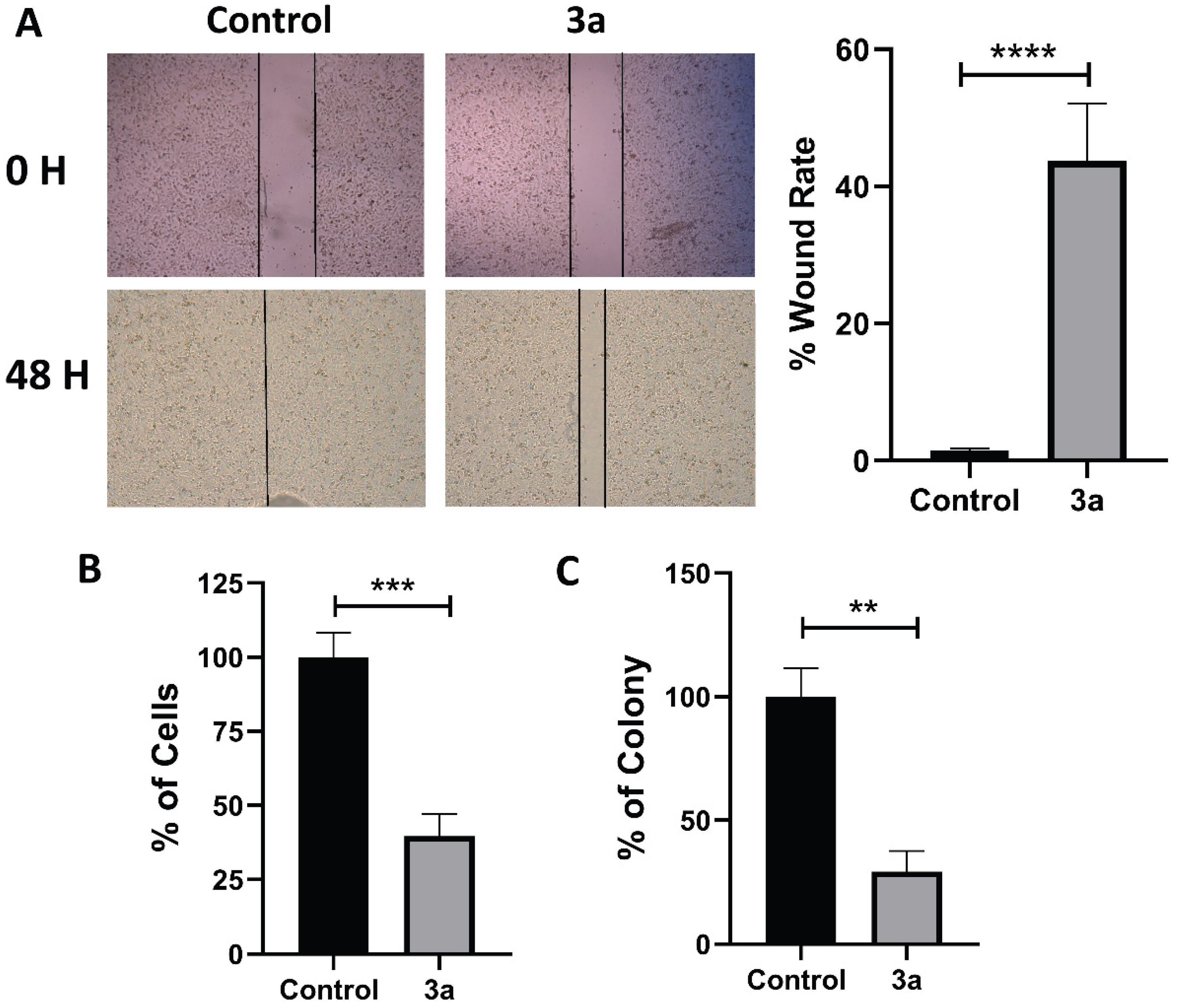
Treatment of CRC cells with 3a decreased migratory, colonization, and adhesion characteristics. A) Comparative analysis of the inhibitory effects of compound 3a on the cell migration ability of HT-29 cells revealed that wound closure was not completed in the treatment group compared to the control group. This finding supports the strong potential anti-metastatic effect of 3a. In the study, wound widths were analyzed using Image J. B) A statistically significant decrease in the adhesion rate was observed between the control and treatment groups of HT-29 CRC cells treated with compound 3a. C)It is noteworthy that the treatment group showed a significant decrease in colony count in HT-29 cells compared to the control group. Data are presented as mean ± SD of three independent experiments. Statistically significant differences compared to the control group are indicated as ** p < 0.01, *** p < 0.001 and **** p < 0.0001.
Figure 5.
Treatment of CRC cells with 3a decreased migratory, colonization, and adhesion characteristics. A) Comparative analysis of the inhibitory effects of compound 3a on the cell migration ability of HT-29 cells revealed that wound closure was not completed in the treatment group compared to the control group. This finding supports the strong potential anti-metastatic effect of 3a. In the study, wound widths were analyzed using Image J. B) A statistically significant decrease in the adhesion rate was observed between the control and treatment groups of HT-29 CRC cells treated with compound 3a. C)It is noteworthy that the treatment group showed a significant decrease in colony count in HT-29 cells compared to the control group. Data are presented as mean ± SD of three independent experiments. Statistically significant differences compared to the control group are indicated as ** p < 0.01, *** p < 0.001 and **** p < 0.0001.
Figure 6.
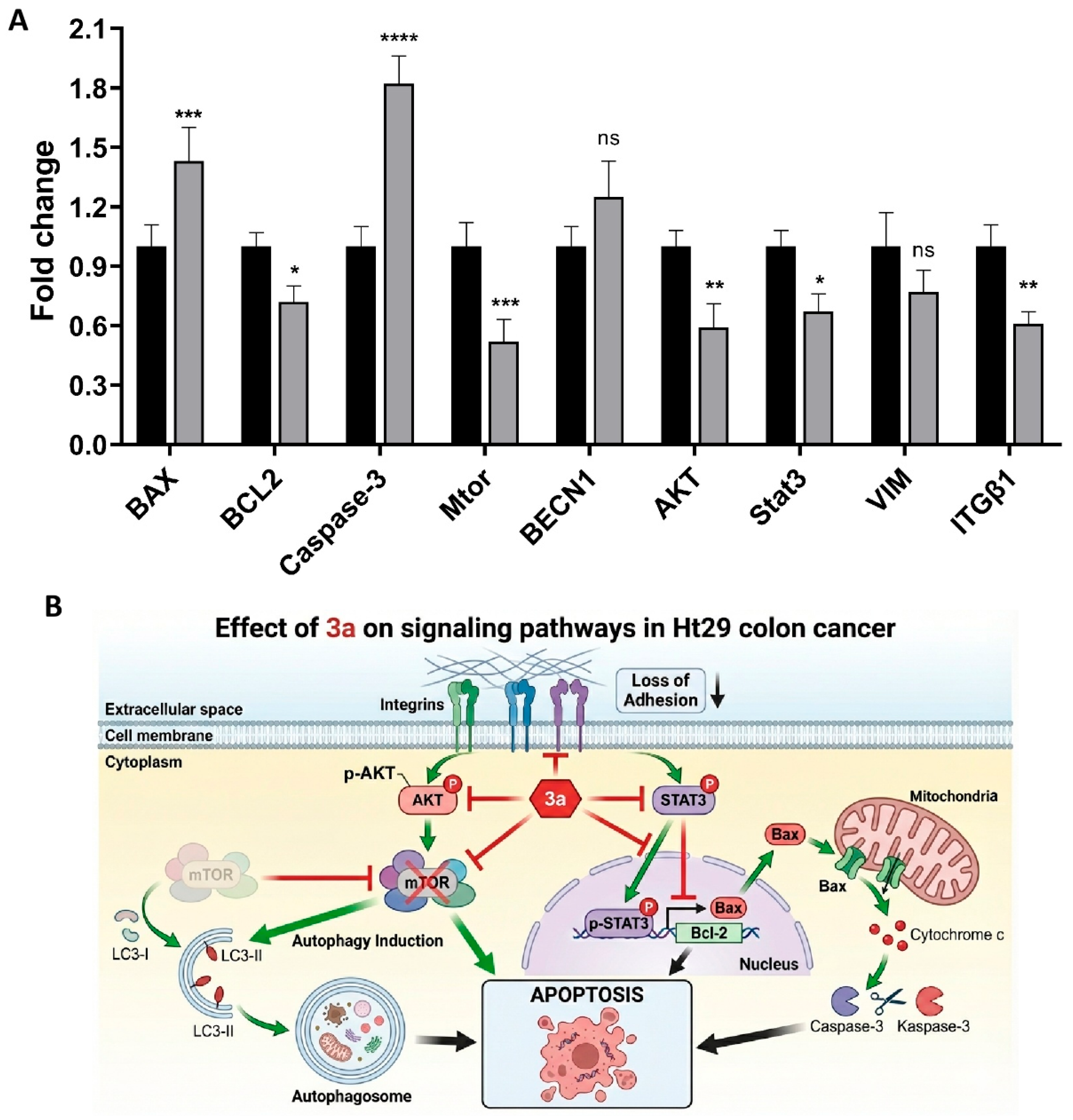
Gene expression profile of HT-29 cells treated with compound 3a. A) mRNA expression levels of key genes involved in apoptosis (BAX, BCL2, Caspase-3), autophagy (mtor, BECN1), survival signaling (AKT, Stat3), and adhesion (VIM, ITGβ1) were analyzed by RT-qPCR after 48 hours of treatment with compound 3a. GAPDH was used as the housekeeping gene. Fold change values are shown with the control group fixed at 1. Significant downregulations were observed in the AKT/mTOR/STAT3 axis and adhesion markers, while pro-apoptotic markers were significantly upregulated.B) Schematic representation of altered gene expression in HT-29. Drawn by artificial intelligence based on information from the literature, then revised and checked before being added.
Figure 6.
Gene expression profile of HT-29 cells treated with compound 3a. A) mRNA expression levels of key genes involved in apoptosis (BAX, BCL2, Caspase-3), autophagy (mtor, BECN1), survival signaling (AKT, Stat3), and adhesion (VIM, ITGβ1) were analyzed by RT-qPCR after 48 hours of treatment with compound 3a. GAPDH was used as the housekeeping gene. Fold change values are shown with the control group fixed at 1. Significant downregulations were observed in the AKT/mTOR/STAT3 axis and adhesion markers, while pro-apoptotic markers were significantly upregulated.B) Schematic representation of altered gene expression in HT-29. Drawn by artificial intelligence based on information from the literature, then revised and checked before being added.
Table 1.
IC50 values of compounds at 72 hours in 4 different cancer cell(Breast and Colon) types.
Table 1.
IC50 values of compounds at 72 hours in 4 different cancer cell(Breast and Colon) types.
| IC50 (µM) |
|---|
| |
MDA-MB-231 |
4T1 |
HT-29 |
DLD-1 |
| 3a |
8,2 |
6,4 |
3,7 |
7,2 |
| 3b |
8,7 |
10,2 |
28,6 |
43,4 |
| 4a |
100< |
100< |
100< |
100< |
| 4b |
100< |
100< |
100< |
100< |