Submitted:
04 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Peptide preparation
Peptide Irradiation
LC-MS/MS and data analysis
Hydrogen peroxide assay
3. Results
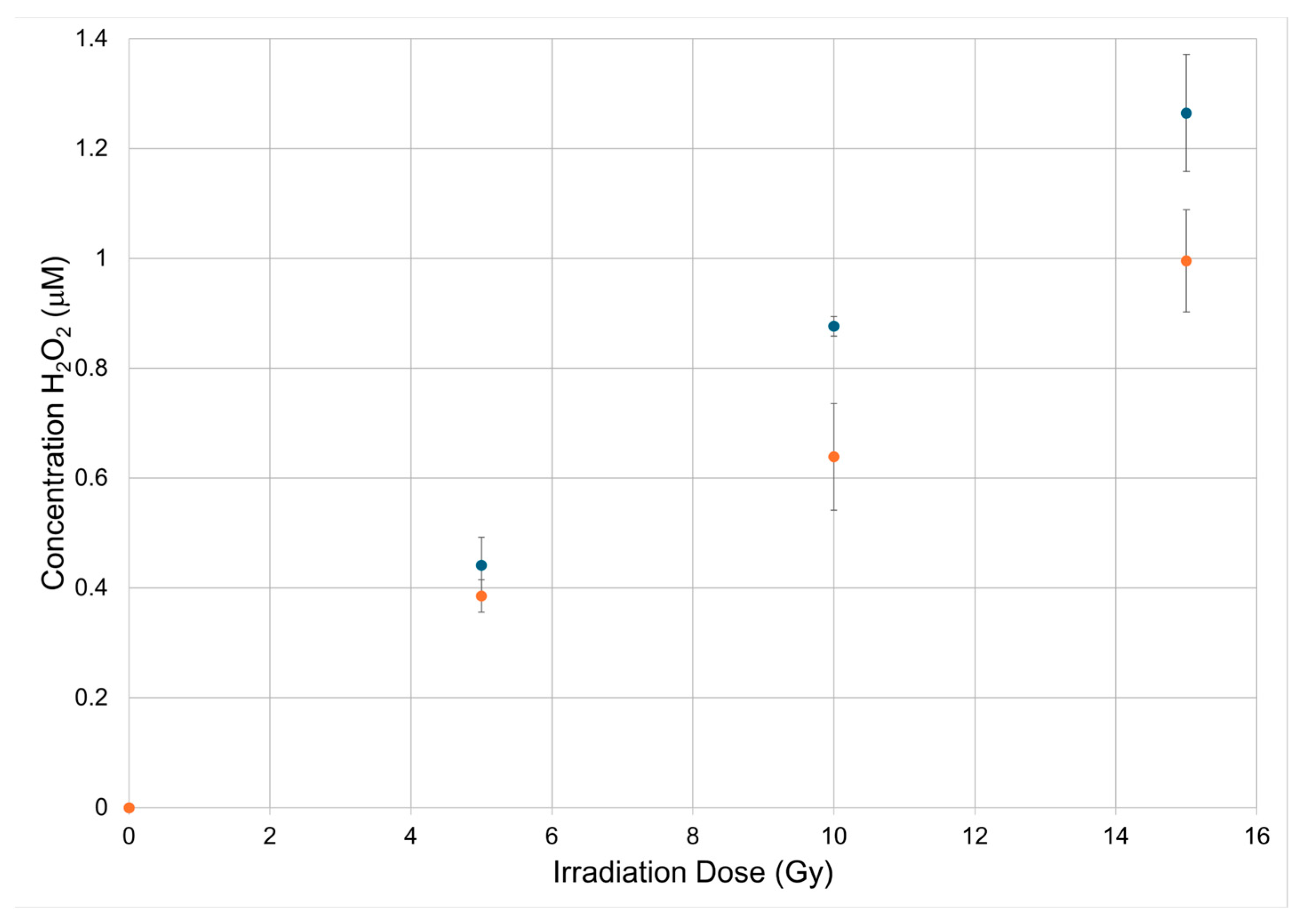
3.1. Measurement of Hydrogen Peroxide Due to Radiolysis
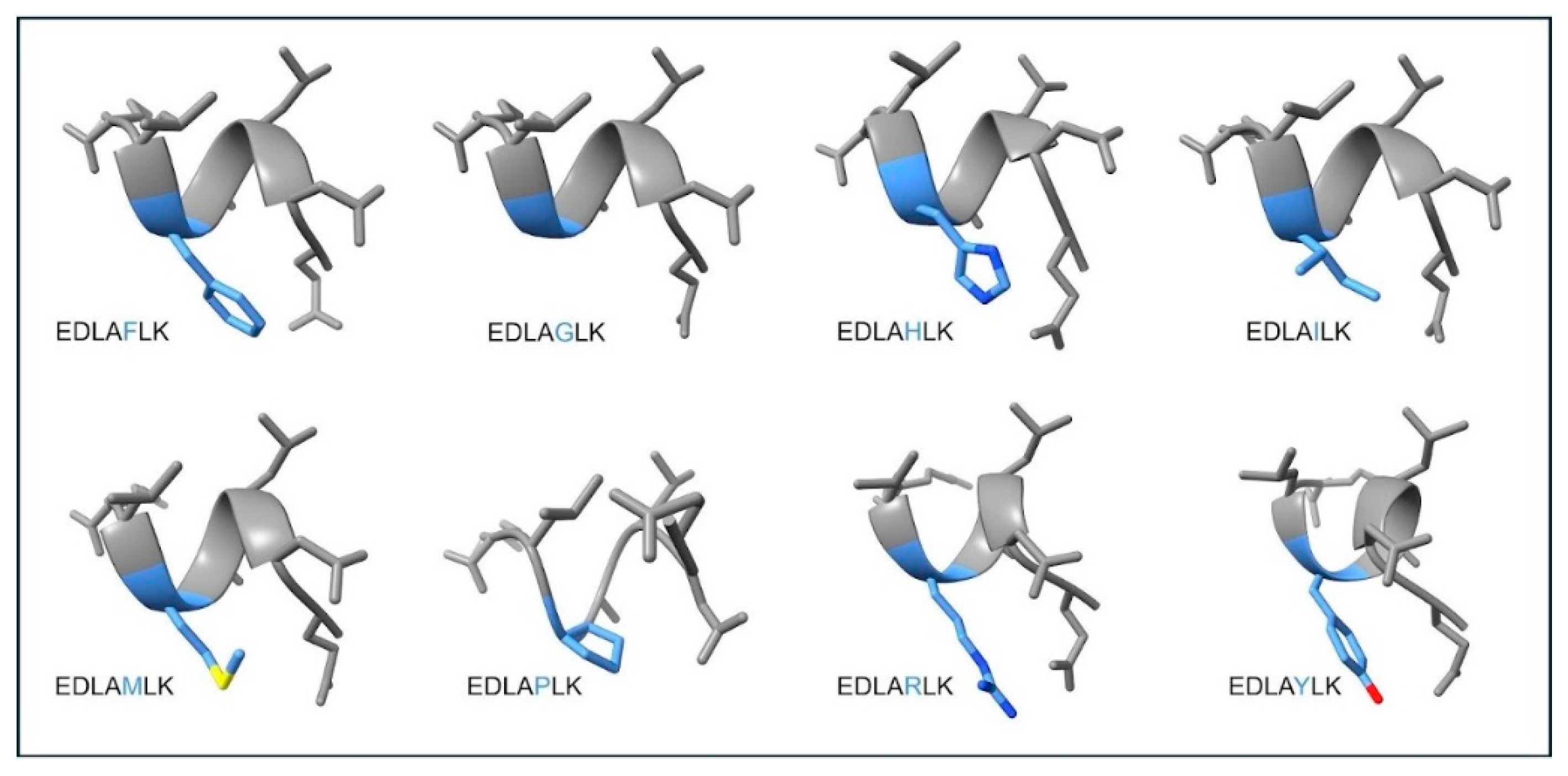
3.2. Peptide Design
3.3. Peptide Oxidation Under Aerated Conditions
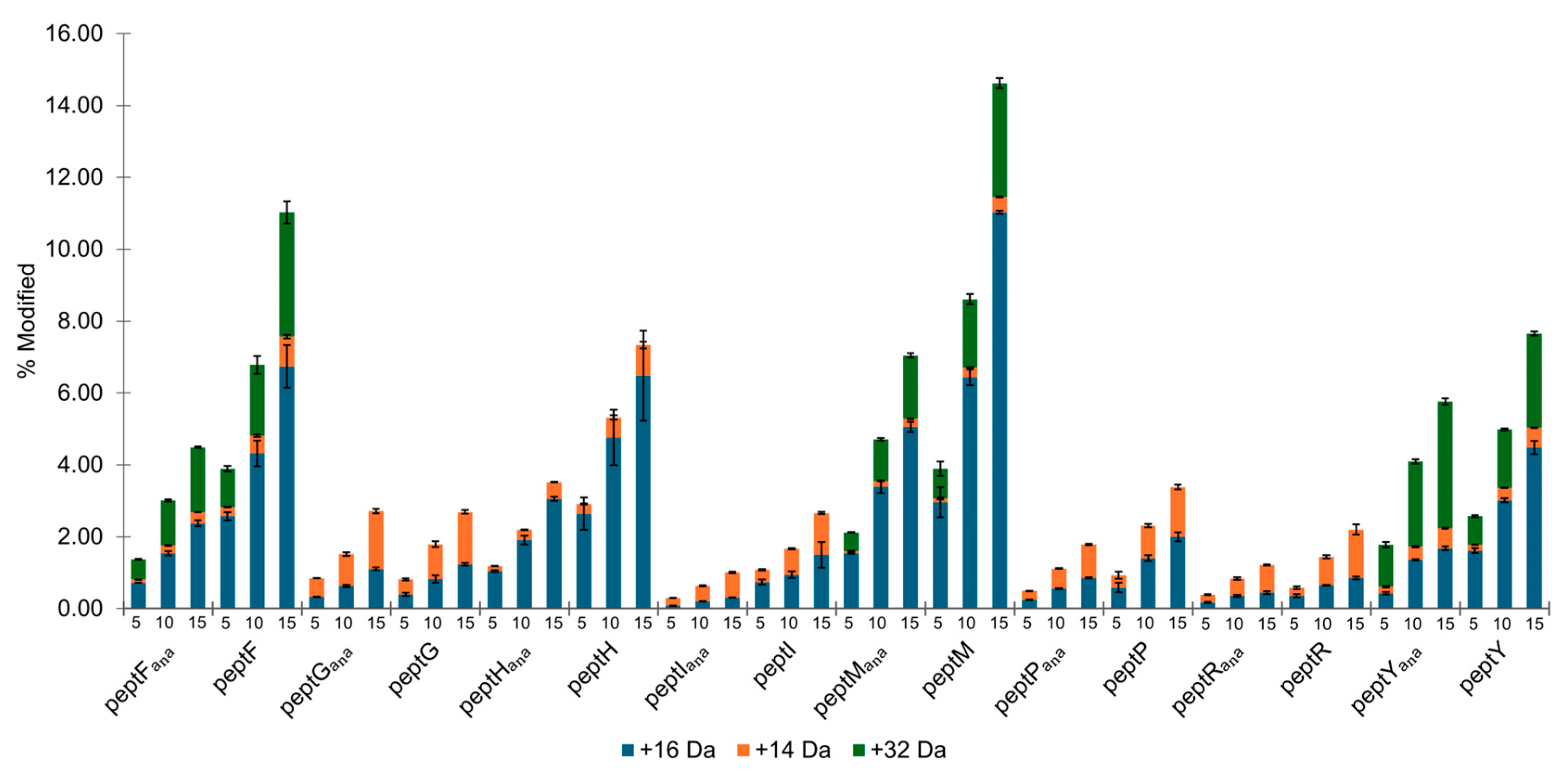
3.4. Peptide Oxidation Under Low Oxygen Conditions
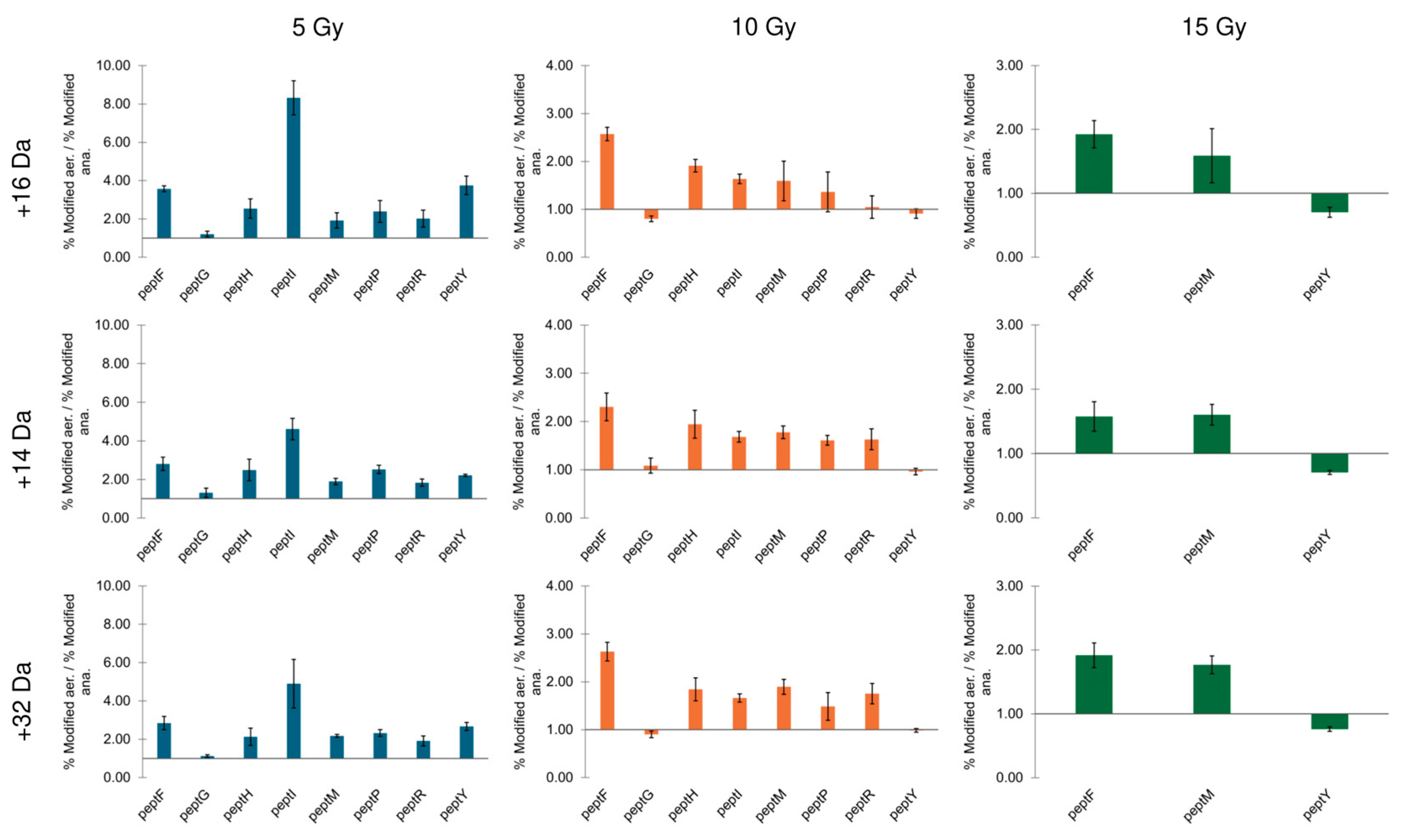
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| LCMS | Liquid Chromatography Mass Spectrometry |
| XFMS | X-ray Footprinting Mass Spectrometry |
References
- He, L.; McAndrew, R.; Barbu, R.; Gifford, G.; Halacoglu, C.; Drouin-Allaire, C.; Weber, L.; Kristensen, L. G.; Gupta, S.; Chen, Y.; et al. Structure and Interactions of HIV-1 gp41 CHR-NHR Reverse Hairpin Constructs Reveal Molecular Determinants of Antiviral Activity. J Mol Biol 2024, 436(16), 168650. [Google Scholar] [CrossRef]
- Dominguez-Martin, M. A.; Hammel, M.; Gupta, S.; Lechno-Yossef, S.; Sutter, M.; Rosenberg, D. J.; Chen, Y.; Petzold, C. J.; Ralston, C. Y.; Polivka, T.; Kerfeld, C. A. Structural analysis of a new carotenoid-binding protein: the C-terminal domain homolog of the OCP. Sci Rep 2020, 10(1), 15564. [Google Scholar] [CrossRef]
- Wheatley, P.; Gupta, S.; Pandini, A.; Chen, Y.; Petzold, C. J.; Ralston, C. Y.; Blair, D. F.; Khan, S. Allosteric Priming of E. coli CheY by the Flagellar Motor Protein FliM. Biophys J 2020, 119(6), 1108–1122. [Google Scholar] [CrossRef]
- Kristensen, L. G.; Gupta, S.; Chen, Y.; Petzold, C. J.; Ralston, C. Y. Residue-Specific Epitope Mapping of the PD-1/Nivolumab Interaction Using X-ray Footprinting Mass Spectrometry. Antibodies 2024, 13(3). [Google Scholar] [CrossRef]
- Sommer, M.; Sutter, M.; Gupta, S.; Kirst, H.; Turmo, A.; Lechno-Yossef, S.; Burton, R. L.; Saechao, C.; Sloan, N. B.; Cheng, X.; et al. Heterohexamers Formed by CcmK3 and CcmK4 Increase the Complexity of Beta Carboxysome Shells. Plant Physiol 2019, 179(1), 156–167. [Google Scholar] [CrossRef]
- Sutter, M.; Utschig, L. M.; Niklas, J.; Paul, S.; Kahan, D. N.; Gupta, S.; Poluektov, O. G.; Ferlez, B. H.; Tefft, N. M.; TerAvest, M. A.; et al. Electrochemical cofactor recycling of bacterial microcompartments. Proc Natl Acad Sci U S A 2024, 121(49), e2414220121. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Bavro, V. N.; D’Mello, R.; Tucker, S. J.; Venien-Bryan, C.; Chance, M. R. Conformational changes during the gating of a potassium channel revealed by structural mass spectrometry. Structure 2010, 18(7), 839–846. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S. Using X-ray Footprinting and Mass Spectrometry to Study the Structure and Function of Membrane Proteins. Protein Pept Lett 2019, 26(1), 44–54. [Google Scholar] [CrossRef] [PubMed]
- Watson, C.; Janik, I.; Zhuang, T.; Charvatova, O.; Woods, R. J.; Sharp, J. S. Pulsed electron beam water radiolysis for submicrosecond hydroxyl radical protein footprinting. Anal Chem 2009, 81(7), 2496–2505. [Google Scholar] [CrossRef]
- Hayes, J. J.; Kam, L.; Tullius, T. D. Footprinting protein-DNA complexes with gamma-rays. Methods Enzymol 1990, 186, 545–549. [Google Scholar] [CrossRef]
- Shcherbakova, I.; Mitra, S.; Beer, R. H.; Brenowitz, M. Fast Fenton footprinting: a laboratory-based method for the time-resolved analysis of DNA, RNA and proteins. Nucleic Acids Res 2006, 34(6), 48–49. [Google Scholar] [CrossRef] [PubMed]
- Sharp, J. S.; Becker, J. M.; Hettich, R. L. Analysis of protein solvent accessible surfaces by photochemical oxidation and mass spectrometry. Anal Chem 2004, 76(3), 672–683. [Google Scholar] [CrossRef] [PubMed]
- Xu, G.; Chance, M. R. Hydroxyl radical-mediated modification of proteins as probes for structural proteomics. Chem Rev 2007, 107(8), 3514–3543. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Celestre, R.; Petzold, C. J.; Chance, M. R.; Ralston, C. Development of a microsecond X-ray protein footprinting facility at the Advanced Light Source. J Synchrotron Radiat 2014, 21 Pt 4, 690–699. [Google Scholar] [CrossRef]
- Asuru, A.; Farquhar, E. R.; Sullivan, M.; Abel, D.; Toomey, J.; Chance, M. R.; Bohon, J. The XFP (17-BM) beamline for X-ray footprinting at NSLS-II. J Synchrotron Radiat 2019, 26 Pt 4, 1388–1399. [Google Scholar] [CrossRef]
- Kusumoto, T.; Danvin, A.; Mamiya, T.; Arnone, A.; Chefson, S.; Galindo, C.; Peaupardin, P.; Raffy, Q.; Kamiguchi, N.; Amano, D.; et al. Dose Rate Effects on Hydrated Electrons, Hydrogen Peroxide, and a OH Radical Molecular Probe Under Clinical Energy Protons. Radiat Res 2024, 201(4), 287–293. [Google Scholar] [CrossRef]
- Sorensen, B. S.; Overgaard, J.; Bassler, N. In vitro RBE-LET dependence for multiple particle types. Acta Oncol 2011, 50(6), 757–762. [Google Scholar] [CrossRef]
- Dean, R. T.; Fu, S.; Stocker, R.; Davies, M. J. Biochemistry and pathology of radical-mediated protein oxidation. Biochem J 1997, 324, 1–18. [Google Scholar] [CrossRef]
- Davies, M. J. Protein oxidation and peroxidation. Biochem J 2016, 473(7), 805–825. [Google Scholar] [CrossRef]
- Morgan, P. E.; Pattison, D. I.; Davies, M. J. Quantification of hydroxyl radical-derived oxidation products in peptides containing glycine, alanine, valine, and proline. Free Radic Biol Med 2012, 52(2), 328–339. [Google Scholar] [CrossRef]
- Sharp, J. S.; Tomer, K. B. Effects of anion proximity in peptide primary sequence on the rate and mechanism of leucine oxidation. Anal Chem 2006, 78(14), 4885–4893. [Google Scholar] [CrossRef]
- Galano, A.; Cruz-Torres, A. OH radical reactions with phenylalanine in free and peptide forms. Org Biomol Chem 2008, 6(4), 732–738. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S.; Inman, J. L.; Chant, J.; Obst-Huebl, L.; Nakamura, K.; Costello, S. M.; Marqusee, S.; Mao, J. H.; Kunz, L.; Paisley, R.; et al. A Novel Platform for Evaluating Dose Rate Effects on Oxidative Damage to Peptides: Toward a High-Throughput Method to Characterize the Mechanisms Underlying the FLASH Effect. Radiat Res 2023, 200(6), 523–530. [Google Scholar] [CrossRef]
- Kristensen, L. G.; Holton, J. M.; Rad, B.; Chen, Y.; Petzold, C. J.; Gupta, S.; Ralston, C. Y. Hydroxyl radical mediated damage of proteins in low oxygen solution investigated using X-ray footprinting mass spectrometry. J Synchrotron Radiat 2021, 28, 1333–1342. [Google Scholar] [CrossRef] [PubMed]
- Boyd, A.; Carver, M.; Dixon, R. Computed and Experimental Product Concentrations in the Radiolysis of Water. Radiat. Phys. Chem. 1980, 15, 8. [Google Scholar] [CrossRef]
- Garrison, W. M. Reaction Mechanisms in the Radiolysis of Peptides, Polypeptides, and Proteins. Chem Rev 1986, 87, 18. [Google Scholar]
- Megroureche, J.; Bekerat, H.; Bian, J.; Bui, A.; Sankey, J.; Childress, L.; Enger, S. A. Development of a hydrated electron dosimeter for radiotherapy applications: A proof of concept. Med Phys 2023, 50(11), 7245–7251. [Google Scholar] [CrossRef]
- Murphy, M. P.; Bayir, H.; Belousov, V.; Chang, C. J.; Davies, K. J. A.; Davies, M. J.; Dick, T. P.; Finkel, T.; Forman, H. J.; Janssen-Heininger, Y.; et al. Guidelines for measuring reactive oxygen species and oxidative damage in cells and in vivo. Nat Metab 2022, 4(6), 651–662. [Google Scholar] [CrossRef]
- Zhang, T.; Stengl, C.; Derksen, L.; Palskis, K.; Koritsidis, K.; Zink, K.; Adeberg, S.; Major, G.; Weishaar, D.; Theiss, U.; et al. Analysis of hydrogen peroxide production in pure water: Ultrahigh versus conventional dose-rate irradiation and mechanistic insights. Med Phys 2024, 51(10), 7439–7452. [Google Scholar] [CrossRef]
- Thomas, W.; Sunnerberg, J.; Reed, M.; Gladstone, D. J.; Zhang, R.; Harms, J.; Swartz, H. M.; Pogue, B. W. Proton and Electron Ultrahigh-Dose-Rate Isodose Irradiations Produce Differences in Reactive Oxygen Species Yields. Int J Radiat Oncol Biol Phys 2024, 118(1), 262–267. [Google Scholar] [CrossRef]
- Blain, G.; Vandenborre, J.; Villoing, D.; Fiegel, V.; Fois, G. R.; Haddad, F.; Koumeir, C.; Maigne, L.; Metivier, V.; Poirier, F.; et al. Proton Irradiations at Ultra-High Dose Rate vs. Conventional Dose Rate: Strong Impact on Hydrogen Peroxide Yield. Radiat Res 2022, 198(3), 318–324. [Google Scholar] [CrossRef]
- Kacem, H.; Psoroulas, S.; Boivin, G.; Folkerts, M.; Grilj, V.; Lomax, T.; Martinotti, A.; Meer, D.; Ollivier, J.; Petit, B.; et al. Comparing radiolytic production of H(2)O(2) and development of Zebrafish embryos after ultra high dose rate exposure with electron and transmission proton beams. Radiother Oncol 2022, 175, 197–202. [Google Scholar] [CrossRef]
- D-Kondo, J. N.; Garcia-Garcia, O. R.; LaVerne, J. A.; Faddegon, B.; Schuemann, J.; Shin, W. G.; Ramos-Mendez, J. An integrated Monte Carlo track-structure simulation framework for modeling inter and intra-track effects on homogenous chemistry. Phys Med Biol 2023, 68(12). [Google Scholar] [CrossRef] [PubMed]
- Derksen, L.; Flatten, V.; Engenhart-Cabillic, R.; Zink, K.; Baumann, K. S. A method to implement inter-track interactions in Monte Carlo simulations with TOPAS-nBio and their influence on simulated radical yields following water radiolysis. Phys Med Biol 2023, 68(13). [Google Scholar] [CrossRef]
- Abolfath, R.; Grosshans, D.; Mohan, R. Oxygen depletion in FLASH ultra-high-dose-rate radiotherapy: A molecular dynamics simulation. Med Phys 2020, 47(12), 6551–6561. [Google Scholar] [CrossRef]
- Lai, Y.; Jia, X.; Chi, Y. Modeling the effect of oxygen on the chemical stage of water radiolysis using GPU-based microscopic Monte Carlo simulations, with an application in FLASH radiotherapy. Phys Med Biol 2021, 66(2), 025004. [Google Scholar] [CrossRef]
- Lamiable, A.; Thevenet, P.; Rey, J.; Vavrusa, M.; Derreumaux, P.; Tuffery, P. PEP-FOLD3: faster de novo structure prediction for linear peptides in solution and in complex. Nucleic Acids Res 2016, 44(W1), W449–454. [Google Scholar] [CrossRef] [PubMed]
- Vogt, W. Oxidation of methionyl residues in proteins: tools, targets, and reversal. Free Radic Biol Med 1995, 18(1), 93–105. [Google Scholar] [CrossRef] [PubMed]
- Wecksler, A. T.; Wang, L.; Bernstein, L. J.; Huang, R. Y. C.; Gupta, S.; Kristensen, L. G.; Ralston, C. Y.; Sobott, F.; Gross, M. L.; Jones, L. M.; et al. Recommendations and Considerations for Hydroxyl Radical Protein Footprinting - Mass Spectrometry. Nature Methods 2026, in press. [Google Scholar]
- Liu, X. R.; Zhang, M. M.; Zhang, B.; Rempel, D. L.; Gross, M. L. Hydroxyl-Radical Reaction Pathways for the Fast Photochemical Oxidation of Proteins Platform As Revealed by (18)O Isotopic Labeling. Anal Chem 2019, 91(14), 9238–9245. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




