Submitted:
04 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
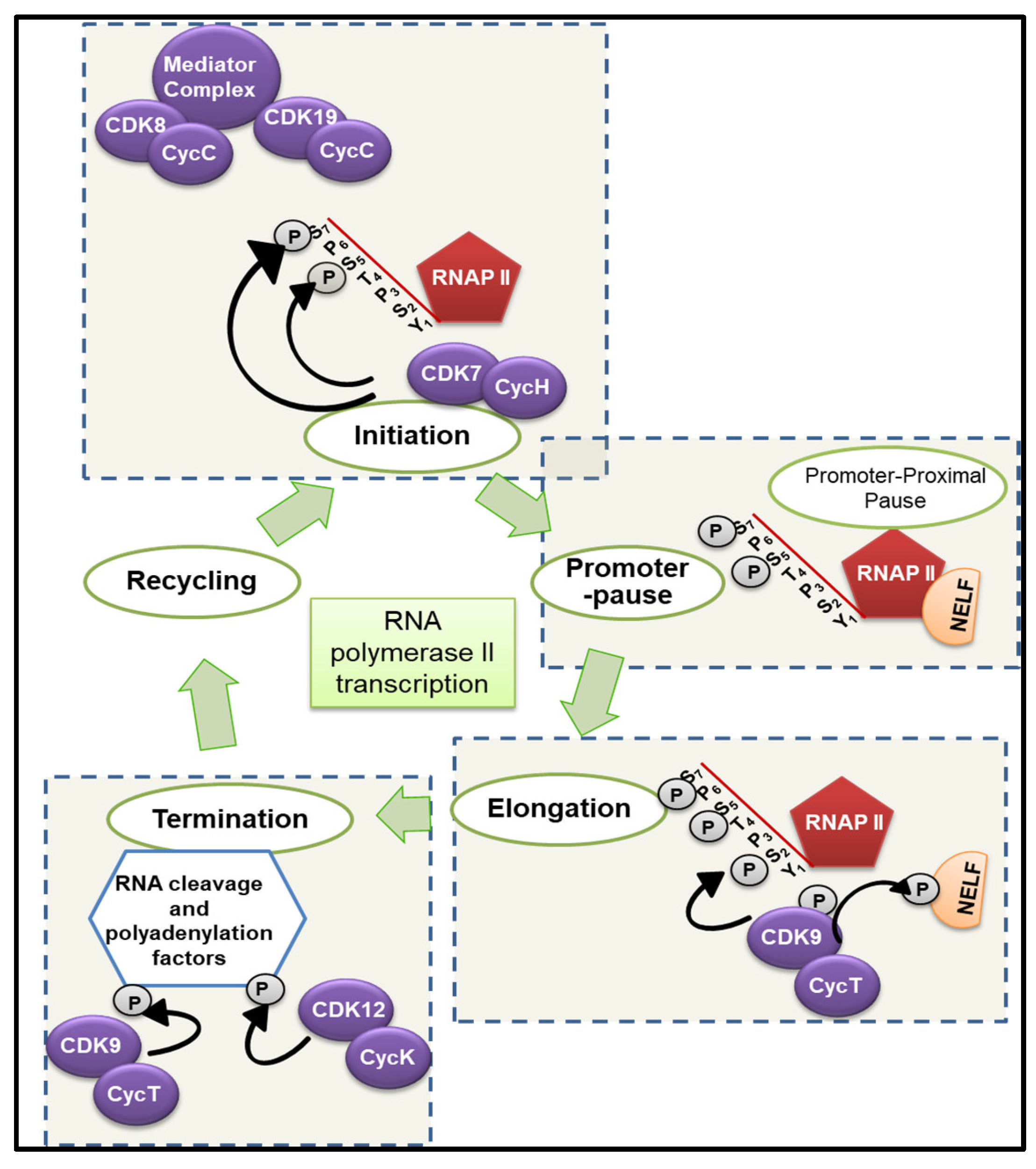
1.1. Transcription-Associated CDKs (tCDKs)
1.1.1. Cyclin-Dependent Kinase 7 (CDK7)
1.1.2. Cyclin-Dependent Kinase 8 (CDK8) and Cyclin-Dependent Kinase 19 (CDK19)
1.1.3. Cyclin-Dependent Kinase 9 (CDK9)
1.1.4. Cyclin-Dependent Kinase 10 (CDK10)
1.1.5. Cyclin-Dependent Kinase 11 (CDK11)
1.1.6. Cyclin-Dependent Kinase 12 (CDK12) and Cyclin-Dependent Kinase 13 (CDK13)
2. tCDKs in GI Malignancies
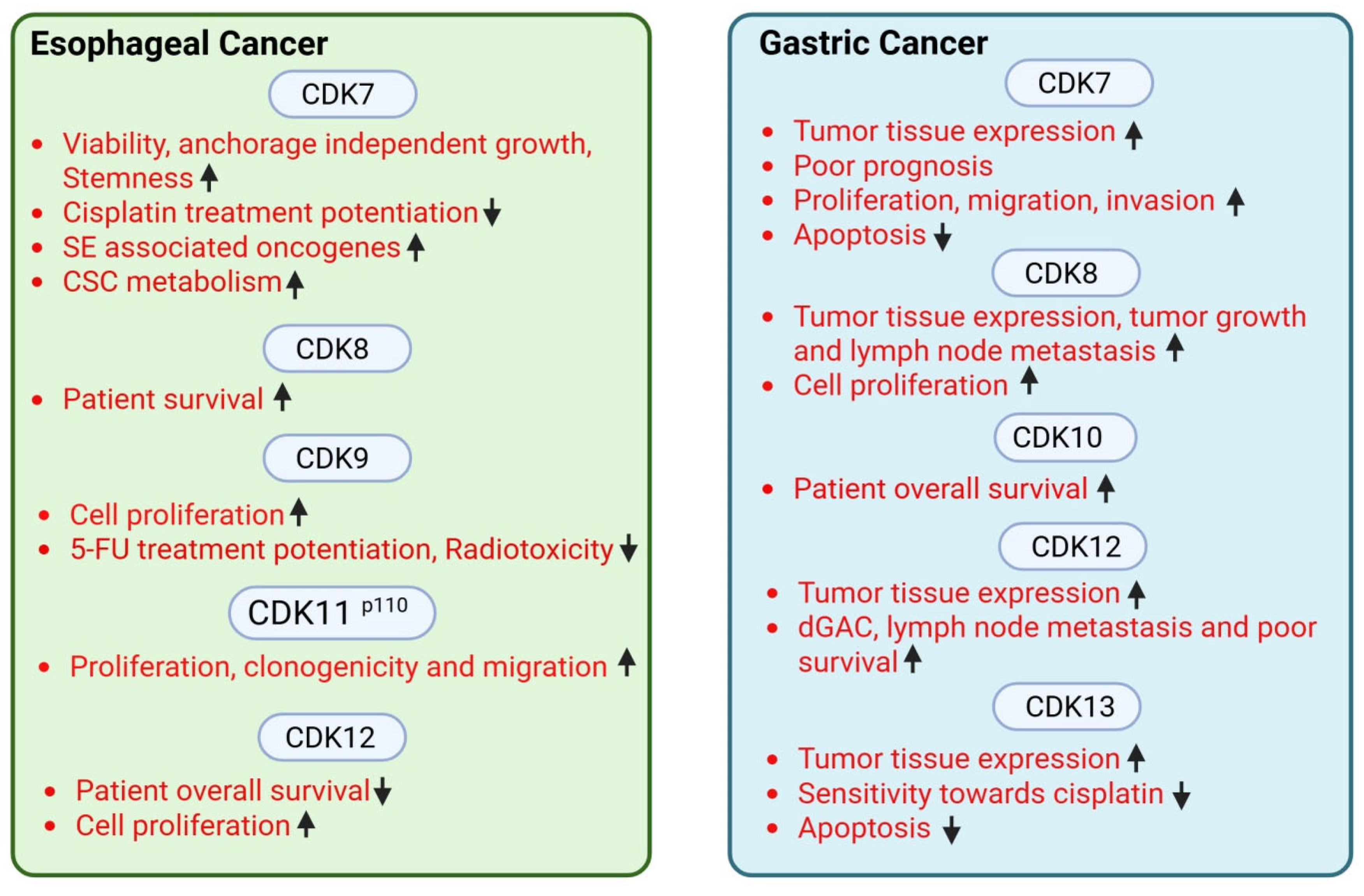
2.1. Esophageal Cancer
2.2. Gastric Cancer
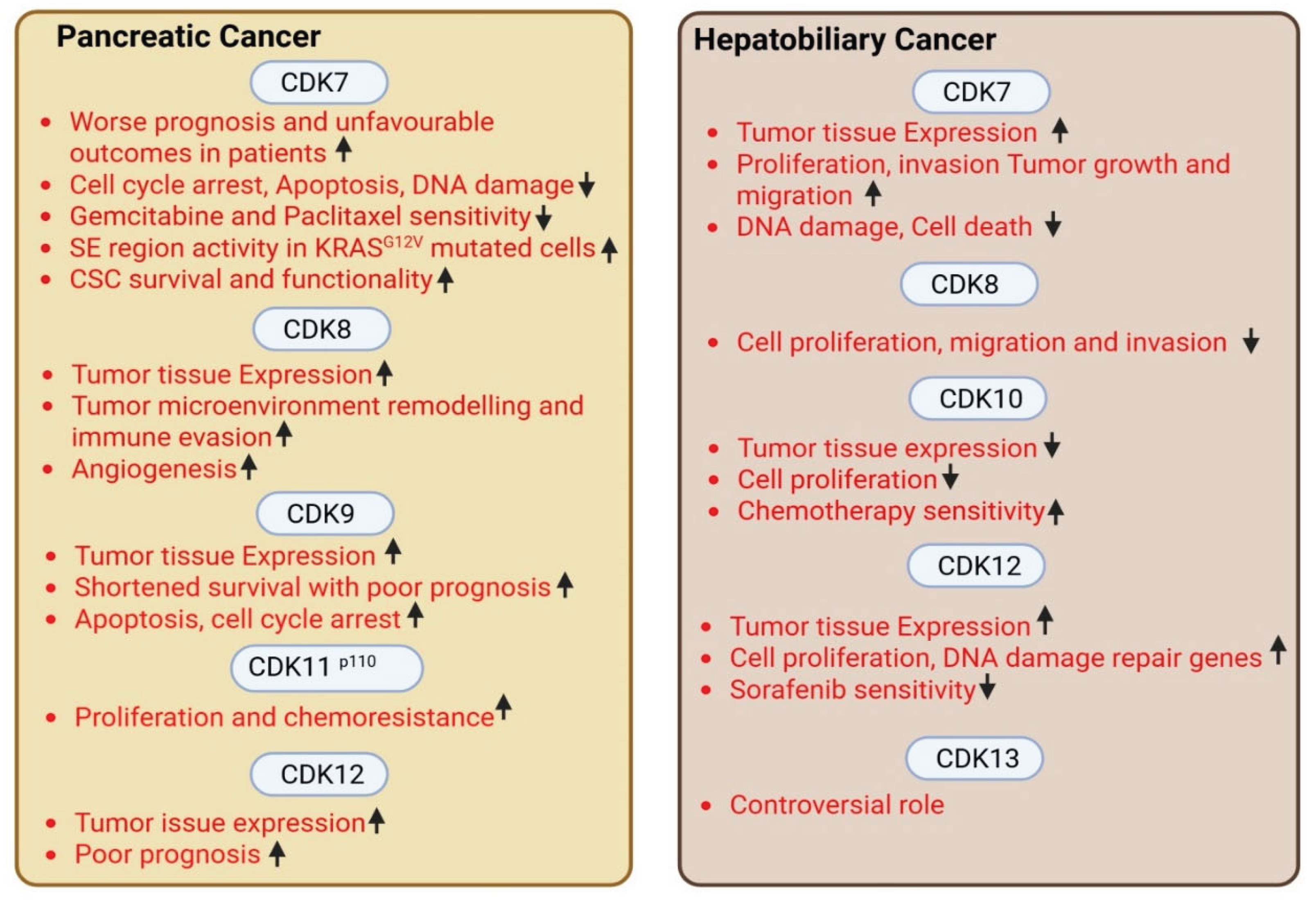
2.3. Pancreatic Cancer
2.4. Hepatobiliary Cancer
3. Key Challenges and Future Directions
| Trial identifier | tCDK targeted | tCDK inhibitor used | GI tumors targeted | Non-GI tumors targeted | Trial status |
|---|---|---|---|---|---|
| NCT04726332 [109] | CDK7 | XL102 | NA | Neoplasm Malignant, Epithelial Ovarian Cancer, Triple Negative Breast Cancer Hormone Receptor Positive (HR+) Breast Carcinoma, Metastatic Castration-resistant Prostate Cancer (CRPC) |
Terminated |
| NCT05394103 | CDK7 | Q901 | colorectal, Pancreatic cancer | advanced or metastatic ovarian, CRPC, HR+ HER2- breast, endometrial cancer, small-cell lung cancer | Recruiting |
| NCT04247126 | CDK7 | SY5609 | pancreatic ductal adenocarcinoma (PDAC) | HR positive, HER2-negative breast cancer | Completed |
| NCT03134638 | CDK7 | SY-1365 | NA | HR+ metastatic breast cancer, Ovarian cancer, advanced solid tumors of any histology | Terminated |
| NCT03363893 [110] | CDK7 | CT7001/Samuraciclib | NA | Triple Negative Breast Cancer (TNBC), CRPC, (HR+ve)/human epidermal growth factor-2 negative (HER2-ve) breast cancer | Completed |
| NCT03065010 | CDK8/ CDK19 |
BCD-115 | NA | ER(+) HER2(-) Local Advanced and Metastatic Breast Cancer | Completed |
| NCT06987058 | CDK8/ CDK19 |
RVU120/SEL120 |
NA | Advanced Solid Tumors, Acute Myeloid Leukaemia (AML), High-risk Myelodysplastic Syndrome |
Not yet recruiting |
| NCT06268574 | CDK8/ CDK19 |
RVU120/SEL120 |
NA | Acute Myeloid Leukemia (AML), High-risk Myelodysplastic Syndrome (MDS) |
Active, not recruiting |
| NCT06243458 | CDK8/ CDK19 |
RVU120/SEL120 |
NA | Low-risk Myelodysplastic Syndrome | Active, not recruiting |
| NCT06191263 | CDK8/ CDK19 |
RVU120/SEL120 | NA | Relapsed/refractory AML | Recruiting |
| NCT06532058 | CDK9 | QHRD107 | NA | Relapsed/Refractory Acute Myeloid Leukemia | Recruiting |
| NCT04588922 | CDK9 | SLS009 (formerly GFH009) | NA | Hematologic Malignancies | Recruiting |
| NCT00835419 | CDK9 | P276-00/Riviciclib | NA | Advanced-metastatic melanoma | Completed |
| NCT00408018 | CDK9 | P276-00/Riviciclib | NA | metastatic or unresectable malignancy | Terminated |
| NCT05168904 | CDK9 | Fadraciclib (Formerly CYC065) | NA | Leukemia, MDS | Suspended |
| NCT04983810 | CDK9 | Fadraciclib (Formerly CYC065) | Biliary tract cancer, HCC, colorectal cancer | advanced solid tumors or lymphoma, Endometrial or Ovarian cancer, Breast cancer | Recruiting |
| NCT04978779 | CDK9 | VIP152 | NA | High-risk Chronic Lymphocytic Leukemia | Terminated |
| NCT03263637 [111] | CDK9 | AZD4573 | NA | Leukemia, Lymphoma | Completed |
| NCT02745743 | CDK9 | BAY1251152 | NA | Hematological malignancy | Completed |
| NCT04718675 | CDK9 | KB-0742 | NA | High Grade Serous Ovarian Cancer | Terminated |
| NCT05159518 | CDK9 | PRT2527 | NA | Advanced/metastatic sarcomas displaying documented gene fusion, castrate resistant prostate cancer, hormone receptor positive HER2-negative breast cancer, Advanced/metastatic non-small cell lung cancer, and solid tumors displaying MYC amplification | Completed |
4. Conclusions
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Malumbres, M. Cyclin-dependent kinases. Genome Biol 2014, 15, 122. [Google Scholar] [CrossRef] [PubMed]
- Donovan, M.G.; Galbraith, M.D.; Espinosa, J.M. Multi-omics investigation reveals functional specialization of transcriptional cyclin dependent kinases in cancer biology. Sci Rep 2022, 12, 22505. [Google Scholar] [CrossRef]
- Galbraith, M.D.; Bender, H.; Espinosa, J.M. Therapeutic targeting of transcriptional cyclin-dependent kinases. Transcription 2019, 10, 118–136. [Google Scholar] [CrossRef]
- Zeng, M.; Kwiatkowski, N.P.; Zhang, T.; Nabet, B.; Xu, M.; Liang, Y.; Quan, C.; et al. Targeting MYC dependency in ovarian cancer through inhibition of CDK7 and CDK12/13. Elife 2018, 7. [Google Scholar] [CrossRef]
- Guo, Z.; Stiller, J.W. Comparative genomics of cyclin-dependent kinases suggest co-evolution of the RNAP II C-terminal domain and CTD-directed CDKs. BMC Genomics 2004, 5, 69. [Google Scholar] [CrossRef] [PubMed]
- Belew, M.D.; Chen, J.; Cheng, Z. Emerging roles of cyclin-dependent kinase 7 in health and diseases. Trends Mol Med 2025, 31, 138–151. [Google Scholar] [CrossRef]
- Ebmeier, C.C.; Erickson, B.; Allen, B.L.; Allen, M.A.; Kim, H.; Fong, N.; Jacobsen, J.R.; et al. Human TFIIH Kinase CDK7 Regulates Transcription-Associated Chromatin Modifications. Cell Rep 2017, 20, 1173–1186. [Google Scholar] [CrossRef]
- Yao, Y.; Ng, J.F.; Park, W.D.; Samur, M.; Morelli, E.; Encinas Mayoral, J.; Chyra, Z.; et al. CDK7 controls E2F- and MYC-driven proliferative and metabolic vulnerabilities in multiple myeloma. Blood 2023, 141, 2841–2852. [Google Scholar] [CrossRef]
- Piccolo, S. Linking cancer transcriptional addictions by CDK7 to YAP/TAZ. Genes Dev 2020, 34, 4–6. [Google Scholar] [CrossRef]
- Christensen, C.L.; Kwiatkowski, N.; Abraham, B.J.; Carretero, J.; Al-Shahrour, F.; Zhang, T.; Chipumuro, E.; et al. Targeting transcriptional addictions in small cell lung cancer with a covalent CDK7 inhibitor. Cancer Cell 2014, 26, 909–922. [Google Scholar] [CrossRef]
- Pellarin, I.; Dall’Acqua, A.; Favero, A.; Segatto, I.; Rossi, V.; Crestan, N.; Karimbayli, J.; et al. Cyclin-dependent protein kinases and cell cycle regulation in biology and disease. Signal Transduct Target Ther 2025, 10, 11. [Google Scholar] [CrossRef] [PubMed]
- Dannappel, M.V.; Sooraj, D.; Loh, J.J.; Firestein, R. Molecular and in vivo Functions of the CDK8 and CDK19 Kinase Modules. Front Cell Dev Biol 2018, 6, 171. [Google Scholar] [CrossRef] [PubMed]
- Prieto, S.; Dubra, G.; Camasses, A.; Aznar, A.B.; Begon-Pescia, C.; Simboeck, E.; Pirot, N.; et al. CDK8 and CDK19 act redundantly to control the CFTR pathway in the intestinal epithelium. EMBO Rep 2023, 24, e54261. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Ruiz, M.J.; Popoola, O.; Mitsopoulos, K.; Te-Poele, R.; Samant, R.S.; Box, G.; Court, W.; et al. Mediator Kinase Inhibitor Selectivity and Activity in Colorectal Cancer. ACS Chem Biol 2025, 20, 1792–1804. [Google Scholar] [CrossRef]
- Adler, A.S.; McCleland, M.L.; Truong, T.; Lau, S.; Modrusan, Z.; Soukup, T.M.; Roose-Girma, M.; et al. CDK8 maintains tumor dedifferentiation and embryonic stem cell pluripotency. Cancer Res 2012, 72, 2129–2139. [Google Scholar] [CrossRef]
- Firestein, R.; Bass, A.J.; Kim, S.Y.; Dunn, I.F.; Silver, S.J.; Guney, I.; Freed, E.; et al. CDK8 is a colorectal cancer oncogene that regulates beta-catenin activity. Nature 2008, 455, 547–551. [Google Scholar] [CrossRef]
- Becker, F.; Joerg, V.; Hupe, M.C.; Roth, D.; Krupar, R.; Lubczyk, V.; Kuefer, R.; et al. Increased mediator complex subunit CDK19 expression associates with aggressive prostate cancer. Int J Cancer 2020, 146, 577–588. [Google Scholar] [CrossRef]
- Serrao, A.; Jenkins, L.M.; Chumanevich, A.A.; Horst, B.; Liang, J.; Gatza, M.L.; Lee, N.Y.; et al. Mediator kinase CDK8/CDK19 drives YAP1-dependent BMP4-induced EMT in cancer. Oncogene 2018, 37, 4792–4808. [Google Scholar] [CrossRef]
- Parua, P.K.; Fisher, R.P. Dissecting the Pol II transcription cycle and derailing cancer with CDK inhibitors. Nat Chem Biol 2020, 16, 716–724. [Google Scholar] [CrossRef]
- Mo, C.; Wei, N.; Li, T.; Ahmed Bhat, M.; Mohammadi, M.; Kuang, C. CDK9 inhibitors for the treatment of solid tumors. Biochem Pharmacol 2024, 229, 116470. [Google Scholar] [CrossRef]
- Ranjan, A.; Pang, Y.; Butler, M.; Merchant, M.; Kim, O.; Yu, G.; Su, Y.T.; et al. Targeting CDK9 for the Treatment of Glioblastoma. Cancers (Basel) 2021, 13. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Zhao, Y.; Yiminniyaze, R.; Zhu, N.; Zhang, Y.; Wumaier, G.; Xia, J.; et al. CDK10 suppresses metastasis of lung adenocarcinoma through inhibition of the ETS2/c-Raf/p-MEK/p-ERK signaling loop. Mol Carcinog 2024, 63, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Duster, R.; Ji, Y.; Pan, K.T.; Urlaub, H.; Geyer, M. Functional characterization of the human Cdk10/Cyclin Q complex. Open Biol 2022, 12, 210381. [Google Scholar] [CrossRef]
- Hong, C.; Meng, Y.; Qiu, A.; Zhang, H.; Yang, L.; Hong, Y.; Huang, Y. Downregulated CDK10 promotes cancer progression and radioresistance in lung cancer through activating the JNK/c-Jun signaling pathway. BMB Rep 2024, 57, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Guo, L.; Dai, H.; Zheng, Z.; Yan, J.; Liu, J.; Zhang, S.; et al. RNF115 aggravates tumor progression through regulation of CDK10 degradation in thyroid carcinoma. Cell Biol Toxicol 2024, 40, 14. [Google Scholar] [CrossRef]
- Li, H.; You, Y.; Liu, J. Cyclin-dependent kinase 10 prevents glioma metastasis via modulation of Snail expression. Mol Med Rep 2018, 18, 1165–1170. [Google Scholar] [CrossRef]
- Bazzi, Z.A.; Tai, I.T. CDK10 in Gastrointestinal Cancers: Dual Roles as a Tumor Suppressor and Oncogene. Front Oncol 2021, 11, 655479. [Google Scholar] [CrossRef]
- Rajecky, M.; Gajduskova, P.; Manik, P.; Hluchy, M.; Hegedusova, E.; Krystofova, K.; Potesil, D.; et al. CDK7-CDK11 axis in spliceosome regulation and pre-mRNA splicing. Nucleic Acids Res 2025, 53. [Google Scholar] [CrossRef]
- Devlin, J.R.; Martin, B.; Bartonicek, N.; Ting, K.; Fan, Z.; Todorovski, I.; Bjelosevic, F.; et al. A CDK11-dependent RNA polymerase II pause-checkpoint precedes CDK9-mediated transition to transcriptional elongation. Mol Cell 2025, 85, 3256–3274.e3214. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhu, L.; Yang, K.; Yang, J.; Wang, D.; Zhang, W.; Zhang, J.; et al. CDK11 Mediates Autophagy to Promote Breast Cancer Cell Proliferation and Migration by Regulating BCL-2. FASEB J 2025, 39, e70997. [Google Scholar] [CrossRef]
- Zhang, Y.; Lu, J.; Qi, M.; Zhang, X.; Ou, D. CDK11 Promotes Paclitaxel Resistance in Cervical Cancer by Regulating LATS1-Mediated Hippo Signaling Pathway Through Phosphorylation of NF2. FASEB J 2025, 39, e71085. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Gao, Y.; Shen, J.; Yang, W.; Choy, E.; Mankin, H.; Hornicek, F.J.; et al. Cyclin-Dependent Kinase 11 (CDK11) Is Required for Ovarian Cancer Cell Growth In Vitro and In Vivo, and Its Inhibition Causes Apoptosis and Sensitizes Cells to Paclitaxel. Mol Cancer Ther 2016, 15, 1691–1701. [Google Scholar] [CrossRef]
- Blazek, D.; Kohoutek, J.; Bartholomeeusen, K.; Johansen, E.; Hulinkova, P.; Luo, Z.; Cimermancic, P.; et al. The Cyclin K/Cdk12 complex maintains genomic stability via regulation of expression of DNA damage response genes. Genes Dev 2011, 25, 2158–2172. [Google Scholar] [CrossRef]
- Dai, W.; Wu, J.; Peng, X.; Hou, W.; Huang, H.; Cheng, Q.; Liu, Z.; et al. CDK12 orchestrates super-enhancer-associated CCDC137 transcription to direct hepatic metastasis in colorectal cancer. Clin Transl Med 2022, 12, e1087. [Google Scholar] [CrossRef] [PubMed]
- Dai, W.; Xie, D.; Huang, H.; Li, J.; Guo, C.; Cao, F.; Yang, L.; et al. New insights into the dule roles CDK12 in human cancers: Mechanisms and interventions for cancer therapy. J Pharm Anal 2025, 15, 101173. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Wang, J.; Yi, Z.; Li, C.; Wang, H.; Zhang, J.; Wang, T.; et al. CDK12 inhibition enhances sensitivity of HER2+ breast cancers to HER2-tyrosine kinase inhibitor via suppressing PI3K/AKT. Eur J Cancer 2021, 145, 92–108. [Google Scholar] [CrossRef]
- Tellier, M.; Zaborowska, J.; Caizzi, L.; Mohammad, E.; Velychko, T.; Schwalb, B.; Ferrer-Vicens, I.; et al. CDK12 globally stimulates RNA polymerase II transcription elongation and carboxyl-terminal domain phosphorylation. Nucleic Acids Res 2020, 48, 7712–7727. [Google Scholar] [CrossRef]
- Cheng, L.; Zhou, S.; Zhou, S.; Shi, K.; Cheng, Y.; Cai, M.C.; Ye, K.; et al. Dual Inhibition of CDK12/CDK13 Targets Both Tumor and Immune Cells in Ovarian Cancer. Cancer Res 2022, 82, 3588–3602. [Google Scholar] [CrossRef]
- Bajrami, I.; Frankum, J.R.; Konde, A.; Miller, R.E.; Rehman, F.L.; Brough, R.; Campbell, J.; et al. Genome-wide profiling of genetic synthetic lethality identifies CDK12 as a novel determinant of PARP1/2 inhibitor sensitivity. Cancer Res 2014, 74, 287–297. [Google Scholar] [CrossRef]
- Tien, J.C.; Zhai, Y.; Wu, R.; Zhang, Y.; Chang, Y.; Cheng, Y.; Todd, A.J.; et al. Defining CDK12 as a tumor suppressor and therapeutic target in mouse models of tubo-ovarian high-grade serous carcinoma. Proc Natl Acad Sci U S A 2025, 122, e2426909122. [Google Scholar] [CrossRef]
- Tien, J.C.; Luo, J.; Chang, Y.; Zhang, Y.; Cheng, Y.; Wang, X.; Yang, J.; et al. CDK12 loss drives prostate cancer progression, transcription-replication conflicts, and synthetic lethality with paralog CDK13. Cell Rep Med 2024, 5, 101758. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, X.N.; Zhang, H.; Wen, J.K.; Gao, H.T.; Shi, B.; Wang, D.D.; et al. CDK13 promotes lipid deposition and prostate cancer progression by stimulating NSUN5-mediated m5C modification of ACC1 mRNA. Cell Death Differ 2023, 30, 2462–2476. [Google Scholar] [CrossRef]
- Insco, M.L.; Abraham, B.J.; Dubbury, S.J.; Kaltheuner, I.H.; Dust, S.; Wu, C.; Chen, K.Y.; et al. Oncogenic CDK13 mutations impede nuclear RNA surveillance. Science 2023, 380, eabn7625. [Google Scholar] [CrossRef]
- Lv, M.; Gong, Y.; Liu, X.; Wang, Y.; Wu, Q.; Chen, J.; Min, Q.; et al. CDK7-YAP-LDHD axis promotes D-lactate elimination and ferroptosis defense to support cancer stem cell-like properties. Signal Transduct Target Ther 2023, 8, 302. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.Y.; Lin, D.C.; Mayakonda, A.; Hazawa, M.; Ding, L.W.; Chien, W.W.; Xu, L.; et al. Targeting super-enhancer-associated oncogenes in oesophageal squamous cell carcinoma. Gut 2017, 66, 1358–1368. [Google Scholar] [CrossRef]
- Zeng, H.; Yang, H.; Song, Y.; Fang, D.; Chen, L.; Zhao, Z.; Wang, C.; et al. Transcriptional inhibition by CDK7/9 inhibitor SNS-032 suppresses tumor growth and metastasis in esophageal squamous cell carcinoma. Cell Death Dis 2021, 12, 1048. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.Y.; Peng, L.; Chen, Y.; Liao, L.D.; Chen, J.X.; Li, M.; Li, Y.Y.; et al. Characterization of super-enhancer-associated functional lncRNAs acting as ceRNAs in ESCC. Mol Oncol 2020, 14, 2203–2230. [Google Scholar] [CrossRef]
- Zhang, J.; Zhu, J.; Yang, L.; Guan, C.; Ni, R.; Wang, Y.; Ji, L.; et al. Interaction with CCNH/CDK7 facilitates CtBP2 promoting esophageal squamous cell carcinoma (ESCC) metastasis via upregulating epithelial-mesenchymal transition (EMT) progression. Tumour Biol 2015, 36, 6701–6714. [Google Scholar] [CrossRef] [PubMed]
- Roninson, I.B.; Gyorffy, B.; Mack, Z.T.; Shtil, A.A.; Shtutman, M.S.; Chen, M.; Broude, E.V. Identifying Cancers Impacted by CDK8/19. Cells 2019, 8. [Google Scholar] [CrossRef]
- Wang, B.Y.; Liu, Q.Y.; Cao, J.; Chen, J.W.; Liu, Z.S. Selective CDK7 inhibition with BS-181 suppresses cell proliferation and induces cell cycle arrest and apoptosis in gastric cancer. Drug Des Devel Ther 2016, 10, 1181–1189. [Google Scholar] [CrossRef]
- Song, X.Q.; Chen, B.B.; Jin, Y.M.; Wang, C.Y. DNMT1-mediated epigenetic suppression of FBXO32 expression promoting cyclin dependent kinase 9 (CDK9) survival and esophageal cancer cell growth. Cell Cycle 2024, 23, 262–278. [Google Scholar] [CrossRef]
- Tong, Z.; Mejia, A.; Veeranki, O.; Verma, A.; Correa, A.M.; Dokey, R.; Patel, V.; et al. Targeting CDK9 and MCL-1 by a new CDK9/p-TEFb inhibitor with and without 5-fluorouracil in esophageal adenocarcinoma. Ther Adv Med Oncol 2019, 11, 1758835919864850. [Google Scholar] [CrossRef]
- Veeranki, O.L.; Tong, Z.; Dokey, R.; Mejia, A.; Zhang, J.; Qiao, Y.; Singh, P.K.; et al. Targeting cyclin-dependent kinase 9 by a novel inhibitor enhances radiosensitization and identifies Axl as a novel downstream target in esophageal adenocarcinoma. Oncotarget 2019, 10, 4703–4718. [Google Scholar] [CrossRef] [PubMed]
- Du, Y.; Yan, D.; Yuan, Y.; Xu, J.; Wang, S.; Yang, Z.; Cheng, W.; et al. CDK11(p110) plays a critical role in the tumorigenicity of esophageal squamous cell carcinoma cells and is a potential drug target. Cell Cycle 2019, 18, 452–466. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Huang, S.; He, Y.; Zhang, Y.; Chen, S.; Huang, M.; He, F.; et al. Discovery of 4-(2-(methylamino)thiazol-5-yl)pyrimidin-2-amine derivatives as novel cyclin-dependent kinase 12 (CDK12) inhibitors for the treatment of esophageal squamous cell carcinoma. Bioorg Chem 2025, 158, 108302. [Google Scholar] [CrossRef]
- Wang, Q.; Li, M.; Zhang, X.; Huang, H.; Huang, J.; Ke, J.; Ding, H.; et al. Upregulation of CDK7 in gastric cancer cell promotes tumor cell proliferation and predicts poor prognosis. Exp Mol Pathol 2016, 100, 514–521. [Google Scholar] [CrossRef]
- Huang, J.R.; Qin, W.M.; Wang, K.; Fu, D.R.; Zhang, W.J.; Jiang, Q.W.; Yang, Y.; et al. Cyclin-dependent kinase 7 inhibitor THZ2 inhibits the growth of human gastric cancer in vitro and in vivo. Am J Transl Res 2018, 10, 3664–3676. [Google Scholar]
- Xu, T.; Xie, M.; Jing, X.; Jiang, H.; Wu, X.; Wang, X.; Shu, Y. Loss of miR-26b-5p promotes gastric cancer progression via miR-26b-5p-PDE4B/CDK8-STAT3 feedback loop. J Transl Med 2023, 21, 77. [Google Scholar] [CrossRef]
- Kim, M.Y.; Han, S.I.; Lim, S.C. Roles of cyclin-dependent kinase 8 and beta-catenin in the oncogenesis and progression of gastric adenocarcinoma. Int J Oncol 2011, 38, 1375–1383. [Google Scholar]
- Song, Y.Q.; Ma, X.H.; Ma, G.L.; Lin, B.; Liu, C.; Deng, Q.J.; Lv, W.P. MicroRNA-107 promotes proliferation of gastric cancer cells by targeting cyclin dependent kinase 8. Diagn Pathol 2014, 9, 164. [Google Scholar] [CrossRef]
- Zhao, B.W.; Chen, S.; Li, Y.F.; Xiang, J.; Zhou, Z.W.; Peng, J.S.; Chen, Y.B. Low Expression of CDK10 Correlates with Adverse Prognosis in Gastric Carcinoma. J Cancer 2017, 8, 2907–2914. [Google Scholar] [CrossRef]
- Wang, B.; Zhang, Y.; Qing, T.; Xing, K.; Li, J.; Zhen, T.; Zhu, S.; et al. Comprehensive analysis of metastatic gastric cancer tumour cells using single-cell RNA-seq. Sci Rep 2021, 11, 1141. [Google Scholar] [CrossRef]
- Gao, L.Z.; Wang, J.Q.; Chen, J.L.; Zhang, X.L.; Zhang, M.M.; Wang, S.L.; Zhao, C. CDK12 Promotes the Proliferation, Migration, and Angiogenesis of Gastric Carcinoma via Activating the PI3K/AKT/mTOR Signaling Pathway. Appl Biochem Biotechnol 2023, 195, 6913–6926. [Google Scholar] [CrossRef]
- Ji, J.; Zhou, C.; Wu, J.; Cai, Q.; Shi, M.; Zhang, H.; Yu, Y.; et al. Expression pattern of CDK12 protein in gastric cancer and its positive correlation with CD8(+) cell density and CCL12 expression. Int J Med Sci 2019, 16, 1142–1148. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Fan, H.; Li, T.; Sihong, L.; Qiao, S.; Bi, J. Low expression of CDK12 in gastric cancer is correlated with advanced stage and poor outcome. Pathol Res Pract 2020, 216, 152962. [Google Scholar] [CrossRef]
- Xin, L.; Zou, Y.H.; Liu, C.X.; Lu, H.; Fan, L.J.; Xu, H.S.; Zhou, Q.; et al. Methionine restriction promotes cisplatin sensitivity of gastric cancer resistant cells by down-regulating circ-CDK13 level. Exp Cell Res 2024, 443, 114315. [Google Scholar] [CrossRef] [PubMed]
- Wu, Z.; Wang, M.; Li, F.; Wang, F.; Jia, J.; Feng, Z.; Huo, X.; et al. CDK13-Mediated Cell Cycle Disorder Promotes Tumorigenesis of High HMGA2 Expression Gastric Cancer. Front Mol Biosci 2021, 8, 707295. [Google Scholar] [CrossRef] [PubMed]
- Zeng, S.; Lan, B.; Ren, X.; Zhang, S.; Schreyer, D.; Eckstein, M.; Yang, H.; et al. CDK7 inhibition augments response to multidrug chemotherapy in pancreatic cancer. J Exp Clin Cancer Res 2022, 41, 241. [Google Scholar] [CrossRef]
- Lu, P.; Geng, J.; Zhang, L.; Wang, Y.; Niu, N.; Fang, Y.; Liu, F.; et al. THZ1 reveals CDK7-dependent transcriptional addictions in pancreatic cancer. Oncogene 2019, 38, 3932–3945. [Google Scholar] [CrossRef]
- Huang, L.; Yang, H.; Chen, K.; Yuan, J.; Li, J.; Dai, G.; Gu, M.; et al. The suppressive efficacy of THZ1 depends on KRAS mutation subtype and is associated with super-enhancer activity and the PI3K/AKT/mTOR signalling in pancreatic ductal adenocarcinoma: A hypothesis-generating study. Clin Transl Med 2023, 13, e1500. [Google Scholar] [CrossRef]
- Huang, C.S.; You, X.; Dai, C.; Xu, Q.C.; Li, F.; Wang, L.; Huang, X.T.; et al. Targeting Super-Enhancers via Nanoparticle-Facilitated BRD4 and CDK7 Inhibitors Synergistically Suppresses Pancreatic Ductal Adenocarcinoma. Adv Sci (Weinh) 2020, 7, 1902926. [Google Scholar] [CrossRef]
- Valles-Marti, A.; de Goeij-de Haas, R.R.; Henneman, A.A.; Piersma, S.R.; Pham, T.V.; Knol, J.C.; Verheij, J.; et al. Kinase activities in pancreatic ductal adenocarcinoma with prognostic and therapeutic avenues. Mol Oncol 2024, 18, 2020–2041. [Google Scholar] [CrossRef]
- Kazi, A.; Chen, L.; Xiang, S.; Vangipurapu, R.; Yang, H.; Beato, F.; Fang, B.; et al. Global Phosphoproteomics Reveal CDK Suppression as a Vulnerability to KRas Addiction in Pancreatic Cancer. Clin Cancer Res 2021, 27, 4012–4024. [Google Scholar] [CrossRef] [PubMed]
- McAndrews, K.M.; Mahadevan, K.K.; Li, B.; Sockwell, A.M.; Morse, S.J.; Kelly, P.J.; Patel, S.I.; et al. CDK8 remodels the tumor microenvironment to resist the therapeutic efficacy of targeted KRAS (G12D) inhibition in pancreatic ductal adenocarcinoma. bioRxiv 2025. [Google Scholar]
- Xu, W.; Wang, Z.; Zhang, W.; Qian, K.; Li, H.; Kong, D.; Li, Y.; et al. Mutated K-ras activates CDK8 to stimulate the epithelial-to-mesenchymal transition in pancreatic cancer in part via the Wnt/beta-catenin signaling pathway. Cancer Lett 2015, 356, 613–627. [Google Scholar] [CrossRef]
- Kartha, N.; Gianopulos, J.E.; Schrank, Z.; Cavender, S.M.; Dobersch, S.; Kynnap, B.D.; Wallace-Povirk, A.; et al. Sirtuin 6 is required for the integrated stress response and resistance to inhibition of transcriptional cyclin-dependent kinases. Sci Transl Med 2023, 15, eabn9674. [Google Scholar] [CrossRef] [PubMed]
- Bian, Y.; Teper, Y.; Mathews Griner, L.A.; Aiken, T.J.; Shukla, V.; Guha, R.; Shinn, P.; et al. Target Deconvolution of a Multikinase Inhibitor with Antimetastatic Properties Identifies TAOK3 as a Key Contributor to a Cancer Stem Cell-Like Phenotype. Mol Cancer Ther 2019, 18, 2097–2110. [Google Scholar] [CrossRef]
- Wei, R.; Kong, L.; Xiao, Y.; Yuan, H.; Song, Y.; Wang, J.; Yu, H.; et al. CDK8 regulates the angiogenesis of pancreatic cancer cells in part via the CDK8-beta-catenin-KLF2 signal axis. Exp Cell Res 2018, 369, 304–315. [Google Scholar] [CrossRef]
- Kretz, A.L.; Schaum, M.; Richter, J.; Kitzig, E.F.; Engler, C.C.; Leithauser, F.; Henne-Bruns, D.; et al. CDK9 is a prognostic marker and therapeutic target in pancreatic cancer. Tumour Biol 2017, 39, 1010428317694304. [Google Scholar] [CrossRef]
- King, H.M.; Rana, S.; Kubica, S.P.; Mallareddy, J.R.; Kizhake, S.; Ezell, E.L.; Zahid, M.; et al. Aminopyrazole based CDK9 PROTAC sensitizes pancreatic cancer cells to venetoclax. Bioorg Med Chem Lett 2021, 43, 128061. [Google Scholar] [CrossRef]
- Xu, H.; Wu, D.; Xiao, M.; Lei, Y.; Lei, Y.; Yu, X.; Shi, S. PP2A complex disruptor SET prompts widespread hypertranscription of growth-essential genes in the pancreatic cancer cells. Sci Adv 2024, 10, eadk6633. [Google Scholar] [CrossRef]
- Ruff, J.P.; Kretz, A.L.; Kornmann, M.; Henne-Bruns, D.; Lemke, J.; Traub, B. The Novel, Orally Bioavailable CDK9 Inhibitor Atuveciclib Sensitises Pancreatic Cancer Cells to TRAIL-induced Cell Death. Anticancer Res 2021, 41, 5973–5985. [Google Scholar] [CrossRef]
- Moser, R.; Annis, J.; Nikolova, O.; Whatcott, C.; Gurley, K.; Mendez, E.; Moran-Jones, K.; et al. Pharmacologic Targeting of TFIIH Suppresses KRAS-Mutant Pancreatic Ductal Adenocarcinoma and Synergizes with TRAIL. Cancer Res 2022, 82, 3375–3393. [Google Scholar] [CrossRef]
- Ge, Y.; Lei, W.; Ma, Y.; Wang, Y.; Wei, B.; Chen, X.; Ru, G.; et al. Synergistic antitumor effects of CDK inhibitor SNS-032 and an oncolytic adenovirus co-expressing TRAIL and Smac in pancreatic cancer. Mol Med Rep 2017, 15, 3521–3528. [Google Scholar] [CrossRef]
- Xu, Z.; Zhang, B.; Liu, Z.; Gou, S. Design, synthesis and anticancer evaluation of selective 2,4-disubstituted pyrimidine CDK9 inhibitors. Eur J Med Chem 2022, 244, 114875. [Google Scholar] [CrossRef]
- Blake, D.R.; Vaseva, A.V.; Hodge, R.G.; Kline, M.P.; Gilbert, T.S.K.; Tyagi, V.; Huang, D.; et al. Application of a MYC degradation screen identifies sensitivity to CDK9 inhibitors in KRAS-mutant pancreatic cancer. Sci Signal 2019, 12. [Google Scholar] [CrossRef]
- Sui, Y.; Wang, T.; Mei, Y.; Zhu, Y.; Jiang, W.; Shen, J.; Yan, S.; et al. Targeting Super-Enhancer-Driven Transcriptional Dependencies Suppresses Aberrant Hedgehog Pathway Activation and Overcomes Smoothened Inhibitor Resistance. Cancer Res 2024, 84, 2690–2706. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Chen, J.; Zeng, T.; Huang, Q.; Chen, D.; Chen, H.; Chen, J.; et al. WZ35 inhibits gastric cancer cell metastasis by depleting glutathione to promote cellular metabolic remodeling. Cancer Lett 2023, 555, 216044. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.Q.; Li, Z.L.; Zhang, Q.; Yin, Q.; Zhang, Y.L.; Ni, W.J.; Jiang, L.Z.; et al. CDK12 is a potential biomarker for diagnosis, prognosis and immunomodulation in pan-cancer. Sci Rep 2024, 14, 6574. [Google Scholar] [CrossRef]
- Duan, J.; He, Y.; Fu, X.; Deng, Y.; Zheng, M.; Lu, D. CDK7 activated beta-catenin/TCF signaling in hepatocellular carcinoma. Exp Cell Res 2018, 370, 461–467. [Google Scholar] [CrossRef]
- Zhong, L.; Yang, S.; Jia, Y.; Lei, K. Inhibition of cyclin-dependent kinase 7 suppresses human hepatocellular carcinoma by inducing apoptosis. J Cell Biochem 2018, 119, 9742–9751. [Google Scholar] [CrossRef]
- Xie, G.; Zhu, A.; Gu, X. Converged DNA Damage Response Renders Human Hepatocellular Carcinoma Sensitive to CDK7 Inhibition. Cancers (Basel) 2022, 14. [Google Scholar] [CrossRef]
- Huang, C.H.; Lujambio, A.; Zuber, J.; Tschaharganeh, D.F.; Doran, M.G.; Evans, M.J.; Kitzing, T.; et al. CDK9-mediated transcription elongation is required for MYC addiction in hepatocellular carcinoma. Genes Dev 2014, 28, 1800–1814. [Google Scholar] [CrossRef]
- Tsang, F.H.; Law, C.T.; Tang, T.C.; Cheng, C.L.; Chin, D.W.; Tam, W.V.; Wei, L.; et al. Aberrant Super-Enhancer Landscape in Human Hepatocellular Carcinoma. Hepatology 2019, 69, 2502–2517. [Google Scholar] [CrossRef]
- Wang, C.; Jin, H.; Gao, D.; Wang, L.; Evers, B.; Xue, Z.; Jin, G.; et al. A CRISPR screen identifies CDK7 as a therapeutic target in hepatocellular carcinoma. Cell Res 2018, 28, 690–692. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Li, D.; Lu, L.; Song, D.; Li, P. LncRNA HEIH modulates the proliferation, migration, and invasion of hepatocellular carcinoma cells by regulating the miR-193a-5p/CDK8 axis. Transl Cancer Res 2024, 13, 423–436. [Google Scholar] [CrossRef]
- Umemoto, K.; Yamamoto, H.; Oikawa, R.; Takeda, H.; Doi, A.; Horie, Y.; Arai, H.; et al. The Molecular Landscape of Pancreatobiliary Cancers for Novel Targeted Therapies From Real-World Genomic Profiling. J Natl Cancer Inst 2022, 114, 1279–1286. [Google Scholar] [CrossRef]
- Wang, C.; Wang, H.; Lieftink, C.; du Chatinier, A.; Gao, D.; Jin, G.; Jin, H.; et al. CDK12 inhibition mediates DNA damage and is synergistic with sorafenib treatment in hepatocellular carcinoma. Gut 2020, 69, 727–736. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, Y.; Lu, L.; Lu, Y.; Tang, Q.; Pu, J. Insight into the molecular mechanism of LINC00152/miR-215/CDK13 axis in hepatocellular carcinoma progression. J Cell Biochem 2019, 120, 18816–18825. [Google Scholar] [CrossRef]
- Kim, H.E.; Kim, D.G.; Lee, K.J.; Son, J.G.; Song, M.Y.; Park, Y.M.; Kim, J.J.; et al. Frequent amplification of CENPF, GMNN and CDK13 genes in hepatocellular carcinomas. PLoS One 2012, 7, e43223. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Chen, G.; Cai, Z.; Li, Z.; Qiu, L.; Xu, H.; Yuan, Y.; et al. CDK13 RNA Over-Editing Mediated by ADAR1 Associates with Poor Prognosis of Hepatocellular Carcinoma Patients. Cell Physiol Biochem 2018, 47, 2602–2612. [Google Scholar] [CrossRef]
- Kolodziejczyk, A.; Sicinski, P. Targeting Transcriptional Cyclin-Dependent Kinases in Cancer. Mol Cancer Ther 2025, 24, 1497–1510. [Google Scholar] [CrossRef]
- Lai, C.F.; Cushing, V.I.; Olden, E.; Bevan, C.L.; Coombes, R.C.; Greber, B.J.; Buluwela, L.; et al. Resistance to CDK7 inhibitors directed by acquired mutation of a conserved residue in cancer cells. EMBO J 2025, 44, 5860–5889. [Google Scholar] [CrossRef] [PubMed]
- Marei, H.E.; Bedair, K.; Hasan, A.; Al-Mansoori, L.; Gaiba, A.; Morrione, A.; Cenciarelli, C.; et al. Targeting CDKs in cancer therapy: advances in PROTACs and molecular glues. NPJ Precis Oncol 2025, 9, 204. [Google Scholar] [CrossRef] [PubMed]
- Toure, M.A.; Motoyama, K.; Xiang, Y.; Urgiles, J.; Kabinger, F.; Koglin, A.S.; Iyer, R.S.; et al. Targeted degradation of CDK9 potently disrupts the MYC-regulated network. Cell Chem Biol 2025, 32, 542–555.e510. [Google Scholar] [CrossRef] [PubMed]
- Raina, K.; Forbes, C.D.; Stronk, R.; Rappi, J.P., Jr.; Eastman, K.J.; Zaware, N.; Yu, X.; et al. Regulated induced proximity targeting chimeras-RIPTACs-A heterobifunctional small molecule strategy for cancer selective therapies. Cell Chem Biol 2024, 31, 1490–1502.e1442. [Google Scholar] [CrossRef]
- Yalala, S.; Gondane, A.; Poulose, N.; Liang, J.; Mills, I.G.; Itkonen, H.M. CDK9 inhibition activates innate immune response through viral mimicry. FASEB J 2024, 38, e23628. [Google Scholar] [CrossRef]
- Kim, S.; Son, E.; Park, H.R.; Kim, M.; Yang, H.W. Dual targeting of CDK4/6 and CDK7 augments tumor response and antitumor immunity in breast cancer models. J Clin Invest 2025, 135. [Google Scholar] [CrossRef]
- Gaur, T.; Poddutoori, R.; Khare, L.; Bagal, B.; Rashmi, S.; Patkar, N.; Tembhare, P.; et al. Novel covalent CDK7 inhibitor potently induces apoptosis in acute myeloid leukemia and synergizes with Venetoclax. J Exp Clin Cancer Res 2023, 42, 186. [Google Scholar] [CrossRef]
- Coombes, R.C.; Howell, S.; Lord, S.R.; Kenny, L.; Mansi, J.; Mitri, Z.; Palmieri, C.; et al. Dose escalation and expansion cohorts in patients with advanced breast cancer in a Phase I study of the CDK7-inhibitor samuraciclib. Nat Commun 2023, 14, 4444. [Google Scholar] [CrossRef]
- Cidado, J.; Boiko, S.; Proia, T.; Ferguson, D.; Criscione, S.W.; San Martin, M.; Pop-Damkov, P.; et al. AZD4573 Is a Highly Selective CDK9 Inhibitor That Suppresses MCL-1 and Induces Apoptosis in Hematologic Cancer Cells. Clin Cancer Res 2020, 26, 922–934. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




