Submitted:
03 February 2026
Posted:
05 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
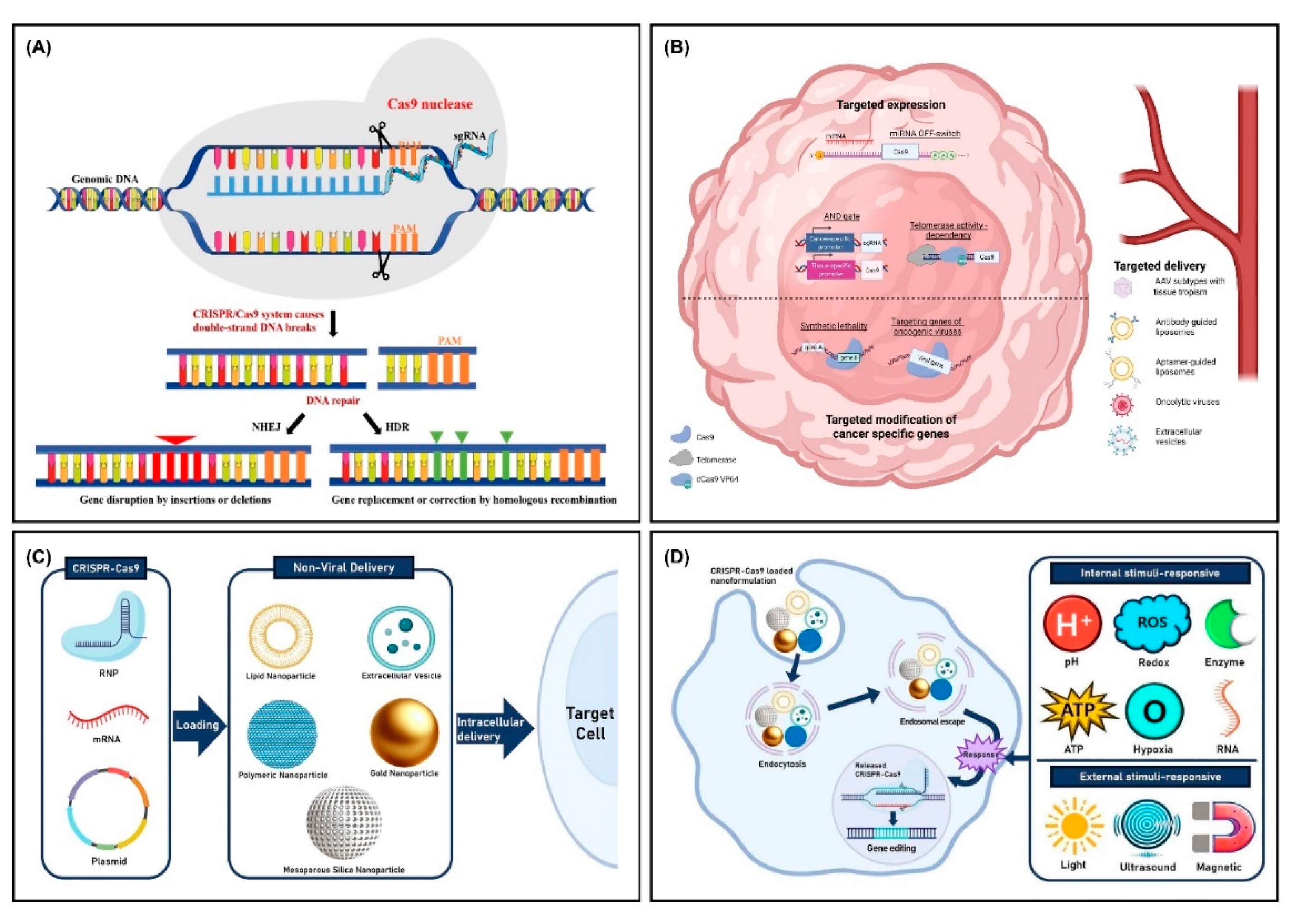
2. CRISPR-Cas9 Technology in Cancer Research
| Pathological Condition | Phase | Status | CRISPR-Engineered Therapeutic Agent | Other Therapies | Identifier |
|---|---|---|---|---|---|
| B-cell acute lymphoblastic leukemia | Phase 1 | Recruiting | Allogenic transplantation of engineered T cells—PBLTT52CAR19 | – | NCT04557436 |
| CD19+ leukemia and lymphoma | Phase 1 | Withdrawn | Allogenic transplantation of engineered T cells—PACE CART19 | – | NCT05037669 |
| Gastrointestinal cancers | Ph½ | Recruiting | Autologous transplantation of CISH-CRISPR TILs | Cyclophosphamide, fludarabine, aldesleukin | NCT04426669 |
| HIV + hematologic malignancies | N/A | Unknown | CRISPR/Cas9-modified CD34+ hematopoietic stem/progenitor cells (CCR5 knockout) | – | NCT03164135 |
| HPV-related malignant neoplasm | Phase 1 | Unknown | Local application of HPV16/18 E6/E7 CRISPR | – | NCT03057912 |
| EBV-associated malignancies | Ph½ | Recruiting | Autologous transplantation of PD-1 knockout EBV-CTL cells | Cyclophosphamide, fludarabine, interleukin-2 | NCT03044743 |
| Non-small cell lung cancer | Phase 1 | Completed | Autologous PD-1 knockout T cells | Cyclophosphamide | NCT02793856 |
| Renal cell carcinoma | Phase 1 | Withdrawn (no funding) | Autologous PD-1 knockout T cells | Cyclophosphamide, interleukin-2 | NCT02867332 |
| Prostate cancer | Phase 1 | Withdrawn (no funding) | Autologous PD-1 knockout T cells | Cyclophosphamide, interleukin-2 | NCT02867345 |
| Bladder cancer | Phase 1 | Withdrawn (no funding) | Autologous PD-1 knockout T cells | Cyclophosphamide, interleukin-2 | NCT02863913 |
| Hepatocellular carcinoma | Phase 1 | Recruiting | Autologous PD-1 knockout T cells | TACE | NCT04417764 |
| Esophageal cancer | Phase 1 | Completed | Autologous PD-1 knockout T cells | – | NCT03081715 |
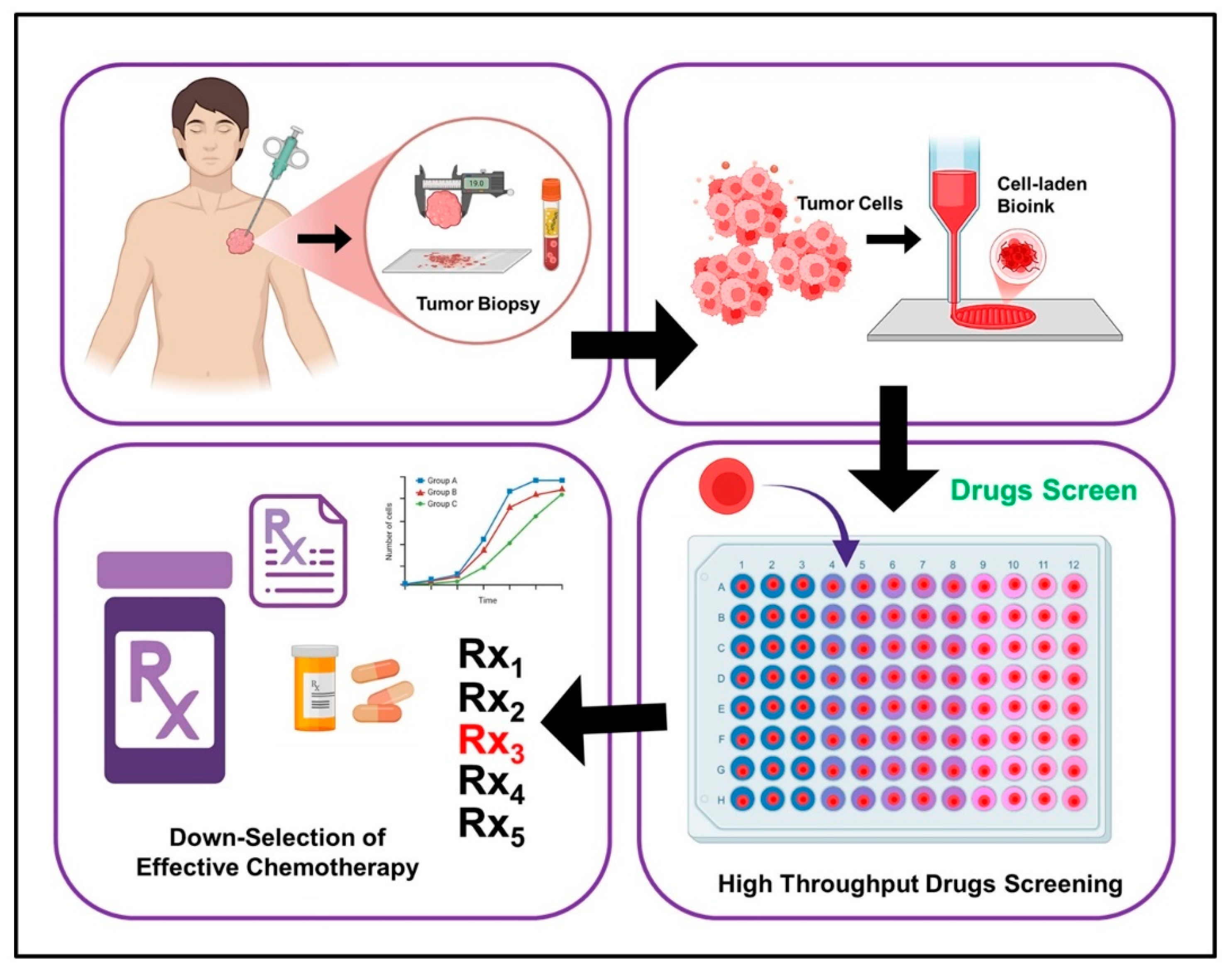
3.3D Bioprinting for Cancer Modeling
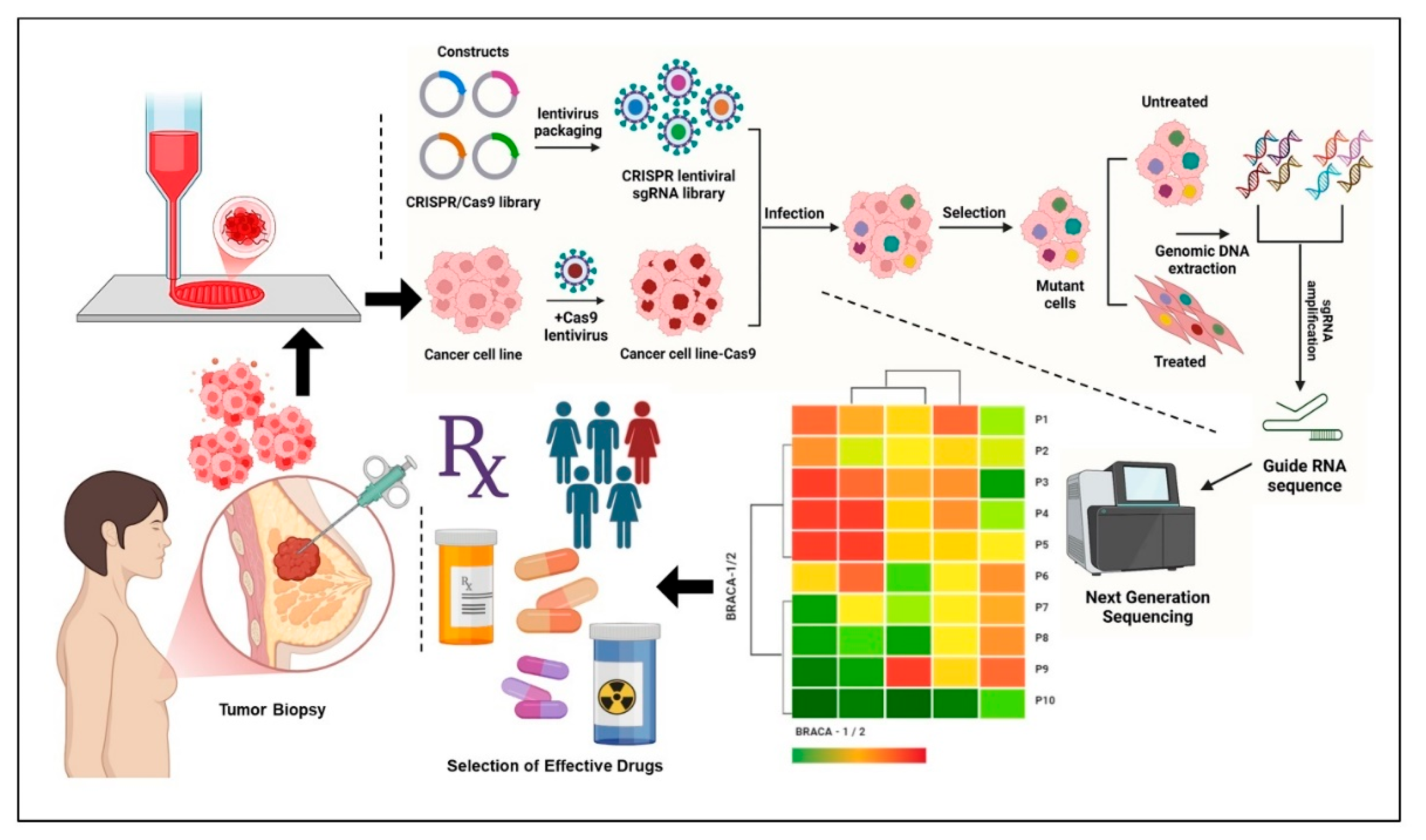
4. Integration of CRISPR-Cas9 with 3D Bioprinted Cancer Models
5. Modeling Tumor Progression and Metastasis by Integration of CRISPR-Cas9 with 3D Bioprinted Cancer Models
6. Integration of CRISPR-Cas9 with 3D Bioprinted Models for Evaluation of Treatment Response
7. Integration of CRISPR-Cas9 with 3D Bioprinting: Challenges and Limitations
8. Integration of CRISPR-Cas9 with 3D Bioprinted Cancer Models: Future Perspectives
9. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Walrath, J.C.; Hawes, J.J.; Van Dyke, T.; Reilly, K.M. Genetically engineered mouse models in cancer research. Adv Cancer Res. 2010, 106, 113–64. [Google Scholar]
- Xiao, W.; Jiang, W.; Chen, Z.; Huang, Y.; Mao, J.; Zheng, W.; et al. Advance in peptide-based drug development: delivery platforms, therapeutics and vaccines. Signal Transduction and Targeted Therapy 2025, 10, 74. [Google Scholar] [CrossRef]
- Selicharova, I.; Sanda, M.; Mladkova, J.; Ohri, S.S.; Vashishta, A.; Fusek, M.; et al. 2-DE analysis of breast cancer cell lines 1833 and 4175 with distinct metastatic organ-specific potentials: comparison with parental cell line MDA-MB-231. Oncol Rep. 2008, 19, 1237–44. [Google Scholar] [CrossRef]
- Ran, R.; Harrison, H.; Syamimi Ariffin, N.; Ayub, R.; Pegg, H.J.; Deng, W.; et al. A role for CBFβ in maintaining the metastatic phenotype of breast cancer cells. Oncogene 2020, 39, 2624–37. [Google Scholar] [CrossRef]
- Sajjad, H.; Imtiaz, S.; Noor, T.; Siddiqui, Y.H.; Sajjad, A.; Zia, M. Cancer models in preclinical research: A chronicle review of advancement in effective cancer research. Animal Model Exp Med. 2021, 4, 87–103. [Google Scholar] [CrossRef] [PubMed]
- Choi, S.Y.C.; Lin, D.; Gout, P.W.; Collins, C.C.; Xu, Y.; Wang, Y. Lessons from patient-derived xenografts for better in vitro modeling of human cancer. Advanced Drug Delivery Reviews 2014, 79-80, 222–37. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Koo, B.-K.; Knoblich, J.A. Human organoids: model systems for human biology and medicine. Nature Reviews Molecular Cell Biology 2020, 21, 571–84. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.C.; Liang, S.B.; Luo, M.; Wang, X.P. Intratumoral heterogeneity and drug resistance in cancer. Cancer Cell Int. 2025, 25, 103. [Google Scholar] [CrossRef]
- Yang, D.; Liu, J.; Qian, H.; Zhuang, Q. Cancer-associated fibroblasts: from basic science to anticancer therapy. Experimental & Molecular Medicine 2023, 55, 1322–32. [Google Scholar] [CrossRef]
- de Visser, K.E.; Joyce, J.A. The evolving tumor microenvironment: From cancer initiation to metastatic outgrowth. Cancer Cell. 2023, 41, 374–403. [Google Scholar] [CrossRef]
- Quail, D.F.; Joyce, J.A. Microenvironmental regulation of tumor progression and metastasis. Nat Med. 2013, 19, 1423–37. [Google Scholar] [CrossRef]
- Glabman, R.A.; Choyke, P.L.; Sato, N. Cancer-Associated Fibroblasts: Tumorigenicity and Targeting for Cancer Therapy. Cancers (Basel) 2022, 14(16). [Google Scholar] [CrossRef] [PubMed]
- Zhang, F.; Ma, Y.; Li, D.; Wei, J.; Chen, K.; Zhang, E.; et al. Cancer associated fibroblasts and metabolic reprogramming: unraveling the intricate crosstalk in tumor evolution. J Hematol Oncol. 2024, 17, 80. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhou, Y.; Wang, Y.; Yang, J.; Li, Z.; Liu, F.; et al. Overcoming cancer treatment resistance: Unraveling the role of cancer-associated fibroblasts. Journal of the National Cancer Center 2025, 5, 237–51. [Google Scholar] [CrossRef]
- Makesh, K.Y.; Navaneethan, A.; Ajay, M.; Munuswamy-Ramanujam, G.; Chinnasamy, A.; Gnanasampanthapandian, D.; et al. A Concise Review of Organoid Tissue Engineering: Regenerative Applications and Precision Medicine. Organoids 2025, 4, 16. [Google Scholar] [CrossRef]
- Feng, X.; Li, Z.; Liu, Y.; Chen, D.; Zhou, Z. CRISPR/Cas9 technology for advancements in cancer immunotherapy: from uncovering regulatory mechanisms to therapeutic applications. Exp Hematol Oncol. 2024, 13, 102. [Google Scholar] [CrossRef]
- Pasupuleti, V.; Vora, L.; Prasad, R.; Nandakumar, D.N.; Khatri, D.K. Glioblastoma preclinical models: Strengths and weaknesses. Biochimica et Biophysica Acta (BBA) - Reviews on Cancer 2024, 1879, 189059. [Google Scholar] [CrossRef]
- Dell, A.C.; Wagner, G.; Own, J.; Geibel, J.P. 3D Bioprinting Using Hydrogels: Cell Inks and Tissue Engineering Applications. Pharmaceutics 2022, 14(12). [Google Scholar] [CrossRef]
- Gogoi, D.; Kumar, M.; Singh, J. A comprehensive review on hydrogel-based bio-ink development for tissue engineering scaffolds using 3D printing. Annals of 3D Printed Medicine 2024, 15, 100159. [Google Scholar] [CrossRef]
- Tripathi, S.; Mandal, S.S.; Bauri, S.; Maiti, P. 3D bioprinting and its innovative approach for biomedical applications. MedComm (2020) 2023, 4, e194. [Google Scholar] [CrossRef]
- Fatimi, A.; Okoro, O.V.; Podstawczyk, D.; Siminska-Stanny, J.; Shavandi, A. Natural Hydrogel-Based Bio-Inks for 3D Bioprinting in Tissue Engineering: A Review. Gels 2022, 8, 179. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Zhu, T.; Cui, H.; Cui, H. Integrating 3D Bioprinting and Organoids to Better Recapitulate the Complexity of Cellular Microenvironments for Tissue Engineering. Adv Healthc Mater. 2025, 14, e2403762. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.H.; Liu, N.; Zhang, H.; Yin, Z.S.; Zha, Z.G. From cells to organs: progress and potential in cartilaginous organoids research. J Transl Med. 2023, 21, 926. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Guo, F.; Jin, Y.; Ma, Y. Applications of human organoids in the personalized treatment for digestive diseases. Signal Transduction and Targeted Therapy 2022, 7, 336. [Google Scholar] [CrossRef]
- Aljabali, A.A.A.; El-Tanani, M.; Tambuwala, M.M. Principles of CRISPR-Cas9 technology: Advancements in genome editing and emerging trends in drug delivery. Journal of Drug Delivery Science and Technology 2024, 92, 105338. [Google Scholar] [CrossRef]
- Balasubramanian, A.; Veluswami, K.; Rao, S.; Aggarwal, S.; Mani, S. Exploring Clustered Regularly Interspaced Short Palindromic Repeats-CRISPR-Associated Protein 9 (CRISPR-Cas9) as a Therapeutic Modality for Cancer: A Scoping Review. Cureus 2024, 16, e64324. [Google Scholar] [CrossRef]
- Shamjetsabam, N.D.; Rana, R.; Malik, P.; Ganguly, N.K. CRISPR/Cas9, an overview of recent developments and applications in cancer research. Int J Surg. 2024, 110, 6198–213. [Google Scholar] [CrossRef]
- Li, H.; Yang, Y.; Hong, W.; Huang, M.; Wu, M.; Zhao, X. Applications of genome editing technology in the targeted therapy of human diseases: mechanisms, advances and prospects. Signal Transduct Target Ther. 2020, 5, 1. [Google Scholar] [CrossRef]
- Schiermeyer, A.; Schneider, K.; Kirchhoff, J.; Schmelter, T.; Koch, N.; Jiang, K.; et al. Targeted insertion of large DNA sequences by homology-directed repair or non-homologous end joining in engineered tobacco BY-2 cells using designed zinc finger nucleases. Plant Direct 2019, 3, e00153. [Google Scholar] [CrossRef]
- Nie, D.; Guo, T.; Yue, M.; Li, W.; Zong, X.; Zhu, Y.; et al. Research Progress on Nanoparticles-Based CRISPR/Cas9 System for Targeted Therapy of Tumors. Biomolecules 2022, 12, 1239. [Google Scholar] [CrossRef]
- Balon, K.; Sheriff, A.; Jacków, J.; Łaczmański, Ł. Targeting Cancer with CRISPR/Cas9-Based Therapy. International Journal of Molecular Sciences 2022, 23, 573. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Rho, W.-Y.; Kim, Y.-H.; Chang, H.; Jun, B.-H. CRISPR-Cas9 Gene Therapy: Non-Viral Delivery and Stimuli-Responsive Nanoformulations. Molecules 2025, 30, 542. [Google Scholar] [CrossRef] [PubMed]
- Ran, F.A.; Hsu, P.D.; Lin, C.-Y.; Gootenberg, J.S.; Konermann, S.; Trevino, A.E.; et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell. 2013, 154, 1380–9. [Google Scholar] [CrossRef] [PubMed]
- Jacków, J.; Guo, Z.; Hansen, C.; Abaci, H.E.; Doucet, Y.S.; Shin, J.U.; et al. CRISPR/Cas9-based targeted genome editing for correction of recessive dystrophic epidermolysis bullosa using iPS cells. Proceedings of the National Academy of Sciences 2019, 116, 26846–52. [Google Scholar] [CrossRef]
- Kocher, T.; Wagner, R.N.; Klausegger, A.; Guttmann-Gruber, C.; Hainzl, S.; Bauer, J.W.; et al. Improved double-nicking strategies for COL7A1-editing by homologous recombination. Molecular Therapy Nucleic Acids 2019, 18, 496–507. [Google Scholar] [CrossRef]
- Zhang, M.; Eshraghian, E.A.; Al Jammal, O.; Zhang, Z.; Zhu, X. CRISPR technology: The engine that drives cancer therapy. Biomedicine & pharmacotherapy 2021, 133, 111007. [Google Scholar]
- Chen, Z.-H.; Yu, Y.P.; Zuo, Z.-H.; Nelson, J.B.; Michalopoulos, G.K.; Monga, S.; et al. Targeting genomic rearrangements in tumor cells through Cas9-mediated insertion of a suicide gene. Nature biotechnology 2017, 35, 543–50. [Google Scholar] [CrossRef]
- Kwon, D.Y.; Zhao, Y.-T.; Lamonica, J.M.; Zhou, Z. Locus-specific histone deacetylation using a synthetic CRISPR-Cas9-based HDAC. Nature communications 2017, 8, 15315. [Google Scholar] [CrossRef]
- Wang, J.; Balan, V.; Marincola, F. CRISPR technology for immuno-oncology applications. Methods in Enzymology 2020, 635, 251–66. [Google Scholar]
- Azangou-Khyavy, M.; Ghasemi, M.; Khanali, J.; Boroomand-Saboor, M.; Jamalkhah, M.; Soleimani, M.; et al. CRISPR/Cas: from tumor gene editing to T cell-based immunotherapy of cancer. Frontiers in Immunology 2020, 11, 2062. [Google Scholar] [CrossRef]
- Perincheri, S.; Hui, P. KRAS mutation testing in clinical practice. Expert review of molecular diagnostics 2015, 15, 375–84. [Google Scholar] [CrossRef] [PubMed]
- Cheung, A.H.-K.; Chow, C.; Zhang, J.; Zhou, Y.; Huang, T.; Ng, K.C.-K.; et al. Specific targeting of point mutations in EGFR L858R-positive lung cancer by CRISPR/Cas9. Laboratory Investigation 2018, 98, 968–76. [Google Scholar] [CrossRef] [PubMed]
- Weng, A.P.; Ferrando, A.A.; Lee, W.; Morris, I.V.J.P.; Silverman, L.B.; Sanchez-Irizarry, C.; et al. Activating mutations of NOTCH1 in human T cell acute lymphoblastic leukemia. Science 2004, 306, 269–71. [Google Scholar] [CrossRef] [PubMed]
- Misiorek, J.O.; Przybyszewska-Podstawka, A.; Kałafut, J.; Paziewska, B.; Rolle, K.; Rivero-Müller, A.; et al. Context matters: NOTCH signatures and pathway in cancer progression and metastasis. Cells 2021, 10, 94. [Google Scholar] [CrossRef]
- Komor, A.C.; Kim, Y.B.; Packer, M.S.; Zuris, J.A.; Liu, D.R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 2016, 533, 420–4. [Google Scholar] [CrossRef]
- Anzalone, A.V.; Koblan, L.W.; Liu, D.R. Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nature biotechnology 2020, 38, 824–44. [Google Scholar] [CrossRef]
- Liu, C.; Zhang, L.; Liu, H.; Cheng, K. Delivery strategies of the CRISPR-Cas9 gene-editing system for therapeutic applications. Journal of Controlled Release 2017, 266, 17–26. [Google Scholar] [CrossRef]
- Yip, B.H. Recent advances in CRISPR/Cas9 delivery strategies. Biomolecules 2020, 10, 839. [Google Scholar] [CrossRef]
- Wang, L.; Zheng, W.; Liu, S.; Li, B.; Jiang, X. Delivery of CRISPR/Cas9 by novel strategies for gene therapy. ChemBioChem 2019, 20, 634–43. [Google Scholar] [CrossRef]
- Liu, B.; Saber, A.; Haisma, H.J. CRISPR/Cas9, a powerful tool for identification of new targets for cancer treatment. Drug Discovery Today 2019, 24, 955–70. [Google Scholar] [CrossRef]
- Yao, S.; He, Z.; Chen, C. CRISPR/Cas9-Mediated Genome Editing of Epigenetic Factors for Cancer Therapy. Human Gene Therapy 2015, 26, 463–71. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Alphonse, M.; Liu, Q. Strategies for nonviral nanoparticle-based delivery of CRISPR/Cas9 therapeutics. WIREs Nanomedicine and Nanobiotechnology 2020, 12, e1609. [Google Scholar] [CrossRef] [PubMed]
- Xu, X.; Wan, T.; Xin, H.; Li, D.; Pan, H.; Wu, J.; et al. Delivery of CRISPR/Cas9 for therapeutic genome editing. The Journal of Gene Medicine 2019, 21, e3107. [Google Scholar] [CrossRef]
- Samareh Salavatipour, M.; Poursalehi, Z.; Hosseini Rouzbahani, N.; Mohammadyar, S.; Vasei, M. CRISPR-Cas9 in basic and translational aspects of cancer therapy. Bioimpacts 2024, 14, 30087. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Li, Z.; Liu, Y.; Chen, D.; Zhou, Z. CRISPR/Cas9 technology for advancements in cancer immunotherapy: from uncovering regulatory mechanisms to therapeutic applications. Experimental Hematology & Oncology 2024, 13, 102. [Google Scholar] [CrossRef]
- Allemailem, K.S.; Alsahli, M.A.; Almatroudi, A.; Alrumaihi, F.; Alkhaleefah, F.K.; Rahmani, A.H.; et al. Current updates of CRISPR/Cas9-mediated genome editing and targeting within tumor cells: an innovative strategy of cancer management. Cancer Commun (Lond) 2022, 42, 1257–87. [Google Scholar] [CrossRef]
- Li, T.; Li, S.; Kang, Y.; Zhou, J.; Yi, M. Harnessing the evolving CRISPR/Cas9 for precision oncology. J Transl Med. 2024, 22, 749. [Google Scholar] [CrossRef]
- Seijas, A.; Cora, D.; Novo, M.; Al-Soufi, W.; Sánchez, L.; Arana, Á.J. CRISPR/Cas9 Delivery Systems to Enhance Gene Editing Efficiency. International Journal of Molecular Sciences 2025, 26, 4420. [Google Scholar] [CrossRef]
- Laurent, M.; Geoffroy, M.; Pavani, G.; Guiraud, S. CRISPR-Based Gene Therapies: From Preclinical to Clinical Treatments. Cells 2024, 13(10). [Google Scholar] [CrossRef]
- Xu, Y.; Le, H.; Wu, Q.; Wang, N.; Gong, C. Advancements in CRISPR/Cas systems for disease treatment. Acta Pharmaceutica Sinica B 2025, 15, 2818–44. [Google Scholar] [CrossRef]
- Lei, T.; Wang, Y.; Zhang, Y.; Yang, Y.; Cao, J.; Huang, J.; et al. Leveraging CRISPR gene editing technology to optimize the efficacy, safety and accessibility of CAR T-cell therapy. Leukemia 2024, 38, 2517–43. [Google Scholar] [CrossRef] [PubMed]
- Wu, B.-X.; Wu, Z.; Hou, Y.-Y.; Fang, Z.-X.; Deng, Y.; Wu, H.-T.; et al. Application of three-dimensional (3D) bioprinting in anti-cancer therapy. Heliyon 2023, 9, e20475. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Jiao, J.; Yang, L.; Wang, Y.; Jiang, W.; Yu, T.; et al. Advanced tumor organoid bioprinting strategy for oncology research. Mater Today Bio 2024, 28, 101198. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Liu, J.; Xu, S.; Wang, J. 3D Bioprinting: An Important Tool for Tumor Microenvironment Research. Int J Nanomedicine 2023, 18, 8039–57. [Google Scholar] [CrossRef]
- Sun, D.; Gao, W.; Hu, H.; Zhou, S. Why 90% of clinical drug development fails and how to improve it? Acta Pharm Sin B 2022, 12, 3049–62. [Google Scholar] [CrossRef]
- Wang, H.; Brown, P.C.; Chow, E.C.Y.; Ewart, L.; Ferguson, S.S.; Fitzpatrick, S.; et al. 3D cell culture models: Drug pharmacokinetics, safety assessment, and regulatory consideration. Clin Transl Sci. 2021, 14, 1659–80. [Google Scholar] [CrossRef]
- Milner, E.; Ainsworth, M.; McDonough, M.; Stevens, B.; Buehrer, J.; Delzell, R.; et al. Emerging Three-Dimensional Hepatic Models in Relation to Traditional Two-Dimensional In Vitro Assays for Evaluating Drug Metabolism and Hepatoxicity. Medicine in Drug Discovery 2020, 8, 100060. [Google Scholar] [CrossRef]
- Kapałczyńska, M.; Kolenda, T.; Przybyła, W.; Zajączkowska, M.; Teresiak, A.; Filas, V.; et al. 2D and 3D cell cultures - a comparison of different types of cancer cell cultures. Arch Med Sci. 2018, 14, 910–9. [Google Scholar] [CrossRef]
- Close, D.A.; Johnston, P.A. Miniaturization and characterization of patient derived hepatocellular carcinoma tumor organoid cultures for cancer drug discovery applications. SLAS Discovery 2025, 30, 100201. [Google Scholar] [CrossRef]
- Zhao, Z.; Chen, X.; Dowbaj, A.M.; Sljukic, A.; Bratlie, K.; Lin, L.; et al. Organoids. Nature Reviews Methods Primers 2022, 2, 94. [Google Scholar] [CrossRef]
- Yang, H.; Sun, L.; Liu, M.; Mao, Y. Patient-derived organoids: a promising model for personalized cancer treatment. Gastroenterol Rep (Oxf) 2018, 6, 243–5. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Yuan, F.; Zuo, X.; Li, M. Breakthroughs and challenges of organoid models for assessing cancer immunotherapy: a cutting-edge tool for advancing personalised treatments. Cell Death Discov. 2025, 11, 222. [Google Scholar] [CrossRef] [PubMed]
- Gungor-Ozkerim, P.S.; Inci, I.; Zhang, Y.S.; Khademhosseini, A.; Dokmeci, M.R. Bioinks for 3D bioprinting: an overview. Biomater Sci. 2018, 6, 915–46. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.B.; Fazel Anvari-Yazdi, A.; Duan, X.; Zimmerling, A.; Gharraei, R.; Sharma, N.K.; et al. Biomaterials / bioinks and extrusion bioprinting. Bioactive Materials 2023, 28, 511–36. [Google Scholar] [CrossRef]
- Wang, H.; Yu, H.; Zhou, X.; Zhang, J.; Zhou, H.; Hao, H.; et al. An Overview of Extracellular Matrix-Based Bioinks for 3D Bioprinting. Front Bioeng Biotechnol. 2022, 10, 905438. [Google Scholar] [CrossRef]
- Johari, N.; Adabavazeh, Z.; Baino, F. PVA-based bioinks for 3D bioprinting: A comprehensive review of their applications in tissue engineering. Bioprinting 2025, 49, e00419. [Google Scholar] [CrossRef]
- Fang, L.; Liu, Y.; Qiu, J.; Wan, W. Bioprinting and its Use in Tumor-On-A-Chip Technology for Cancer Drug Screening: A Review. Int J Bioprint 2022, 8, 603. [Google Scholar] [CrossRef]
- Monteiro, M.V.; Zhang, Y.S.; Gaspar, V.M.; Mano, J.F. 3D-bioprinted cancer-on-a-chip: level-up organotypic in vitro models. Trends Biotechnol. 2022, 40, 432–47. [Google Scholar] [CrossRef]
- Shukla, A.K.; Yoon, S.; Oh, S.-O.; Lee, D.; Ahn, M.; Kim, B.S. Advancement in Cancer Vasculogenesis Modeling through 3D Bioprinting Technology. Biomimetics 2024, 9, 306. [Google Scholar] [CrossRef]
- Anari, E.; Anari, R.; Pazoki-Toroudi, H. A decade of scientific advancements and collaborations on CRISPR-Cas9 application in cancer research: a bibliometric review. Discov Oncol. 2025, 16, 1232. [Google Scholar] [CrossRef]
- Li, T.; Yang, Y.; Qi, H.; Cui, W.; Zhang, L.; Fu, X.; et al. CRISPR/Cas9 therapeutics: progress and prospects. Signal Transduction and Targeted Therapy 2023, 8, 36. [Google Scholar] [CrossRef] [PubMed]
- Chehelgerdi, M.; Chehelgerdi, M.; Khorramian-Ghahfarokhi, M.; Shafieizadeh, M.; Mahmoudi, E.; Eskandari, F.; et al. Comprehensive review of CRISPR-based gene editing: mechanisms, challenges, and applications in cancer therapy. Molecular Cancer 2024, 23, 9. [Google Scholar] [CrossRef] [PubMed]
- Shukla, A.K.; Gao, G.; Kim, B.S. Applications of 3D Bioprinting Technology in Induced Pluripotent Stem Cells-Based Tissue Engineering. Micromachines (Basel) 2022, 13(2). [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zhou, G.; Jiang, Q.; Li, L. In situ 3D bioprinting: The future of regenerative medicine. In Fundamental Research; 2025. [Google Scholar]
- Mallya, D.; Gadre, M.A.; Varadharajan, S.; Vasanthan, K.S. 3D bioprinting for the construction of drug testing models-development strategies and regulatory concerns. Front Bioeng Biotechnol. 2025, 13, 1457872. [Google Scholar] [CrossRef]
- Cordeiro, S.; Oliveira, B.B.; Valente, R.; Ferreira, D.; Luz, A.; Baptista, P.V.; et al. Breaking the mold: 3D cell cultures reshaping the future of cancer research. Front Cell Dev Biol. 2024, 12, 1507388. [Google Scholar] [CrossRef]
- Ahmed, T. Biomaterial-based in vitro 3D modeling of glioblastoma multiforme. Cancer Pathogenesis and Therapy 2023, 1, 177–94. [Google Scholar] [CrossRef]
- Yamanaka, Y. CRISPR/Cas9 Genome Editing as a Strategy to Study the Tumor Microenvironment in Transgenic Mice. Methods Mol Biol. 2016, 1458, 261–71. [Google Scholar]
- Begagić, E.; Bečulić, H.; Đuzić, N.; Džidić-Krivić, A.; Pugonja, R.; Muharemović, A.; et al. CRISPR/Cas9-Mediated Gene Therapy for Glioblastoma: A Scoping Review. Biomedicines 2024, 12(1). [Google Scholar] [CrossRef]
- Slika, H.; Karimov, Z.; Alimonti, P.; Abou-Mrad, T.; De Fazio, E.; Alomari, S.; et al. Preclinical Models and Technologies in Glioblastoma Research: Evolution, Current State, and Future Avenues. International Journal of Molecular Sciences 2023, 24, 16316. [Google Scholar] [CrossRef]
- Lu, P.; Ruan, D.; Huang, M.; Tian, M.; Zhu, K.; Gan, Z.; et al. Harnessing the potential of hydrogels for advanced therapeutic applications: current achievements and future directions. Signal Transduction and Targeted Therapy 2024, 9, 166. [Google Scholar] [CrossRef]
- Porter, R.J.; Murray, G.I.; McLean, M.H. Current concepts in tumour-derived organoids. British Journal of Cancer 2020, 123, 1209–18. [Google Scholar] [CrossRef] [PubMed]
- Dave, R.; Pandey, K.; Patel, R.; Gour, N.; Bhatia, D. Leveraging 3D cell culture and AI technologies for next-generation drug discovery. Cell Biomaterials 2025, 1(3). [Google Scholar] [CrossRef]
- Stefanoudakis, D. Integrating CRISPR Technology with Key Genetic Markers in Pancreatic Cancer: A New Frontier in Targeted Therapies. SynBio 2025, 3, 1. [Google Scholar] [CrossRef]
- Seager, R.J.; Hajal, C.; Spill, F.; Kamm, R.D.; Zaman, M.H. Dynamic interplay between tumour, stroma and immune system can drive or prevent tumour progression. Converg Sci Phys Oncol. 2017, 3. [Google Scholar] [CrossRef]
- Kozlova, N.; Grossman, J.E.; Iwanicki, M.P.; Muranen, T. The Interplay of the Extracellular Matrix and Stromal Cells as a Drug Target in Stroma-Rich Cancers. Trends Pharmacol Sci. 2020, 41, 183–98. [Google Scholar] [CrossRef]
- El-Tanani, M.; Rabbani, S.A.; Babiker, R.; Rangraze, I.; Kapre, S.; Palakurthi, S.S.; et al. Unraveling the tumor microenvironment: Insights into cancer metastasis and therapeutic strategies. Cancer Letters 2024, 591, 216894. [Google Scholar] [CrossRef]
- Castaneda, M.; den Hollander, P.; Kuburich, N.A.; Rosen, J.M.; Mani, S.A. Mechanisms of cancer metastasis. Seminars in Cancer Biology 2022, 87, 17–31. [Google Scholar] [CrossRef]
- Liu, Q.; Zhang, H.; Jiang, X.; Qian, C.; Liu, Z.; Luo, D. Factors involved in cancer metastasis: a better understanding to "seed and soil" hypothesis. Mol Cancer 2017, 16, 176. [Google Scholar] [CrossRef]
- Shi, X.; Wang, X.; Yao, W.; Shi, D.; Shao, X.; Lu, Z.; et al. Mechanism insights and therapeutic intervention of tumor metastasis: latest developments and perspectives. Signal Transduction and Targeted Therapy 2024, 9, 192. [Google Scholar] [CrossRef]
- Khawar, M.B.; Wang, Y.; Majeed, A.; Afzal, A.; Haneef, K.; Sun, H. Mini-organs with big impact: Organoids in liver cancer studies. Oncol Res. 2023, 31, 677–88. [Google Scholar] [CrossRef]
- Hu, Y.; Peng, Z.; Qiu, M.; Xue, L.; Ren, H.; Wu, X.; et al. Developing biotechnologies in organoids for liver cancer. Biomedical Technology 2025, 9, 100067. [Google Scholar] [CrossRef]
- Peng, Z.; Lv, X.; Sun, H.; Zhao, L.; Huang, S. 3D tumor cultures for drug resistance and screening development in clinical applications. Mol Cancer 2025, 24, 93. [Google Scholar] [CrossRef] [PubMed]
- Ishibashi, A.; Saga, K.; Hisatomi, Y.; Li, Y.; Kaneda, Y.; Nimura, K. A simple method using CRISPR-Cas9 to knock-out genes in murine cancerous cell lines. Sci Rep. 2020, 10, 22345. [Google Scholar] [CrossRef] [PubMed]
- Imtiaz, S.; Ferdous, U.T.; Nizela, A.; Hasan, A.; Shakoor, A.; Zia, A.W.; et al. Mechanistic study of cancer drug delivery: Current techniques, limitations, and future prospects. European Journal of Medicinal Chemistry 2025, 290, 117535. [Google Scholar] [CrossRef]
- Wang, X.-H.; Wang, W.-Y.; Sun, Z.-J. Immune organoid for cancer immunotherapy. Acta Pharmaceutica Sinica B 2025, 15, 3419–35. [Google Scholar] [CrossRef]
- Alves, E.; Taifour, S.; Dolcetti, R.; Chee, J.; Nowak, A.K.; Gaudieri, S.; et al. Reprogramming the anti-tumor immune response via CRISPR genetic and epigenetic editing. Mol Ther Methods Clin Dev. 2021, 21, 592–606. [Google Scholar] [CrossRef]
- Al-Kabani, A.; Huda, B.; Haddad, J.; Yousuf, M.; Bhurka, F.; Ajaz, F.; et al. Exploring Experimental Models of Colorectal Cancer: A Critical Appraisal from 2D Cell Systems to Organoids, Humanized Mouse Avatars, Organ-on-Chip, CRISPR Engineering, and AI-Driven Platforms—Challenges and Opportunities for Translational Precision Oncology. Cancers 2025, 17, 2163. [Google Scholar]
- Gnatowski, P.; Piłat, E.; Kucińska-Lipka, J.; Saeb, M.R.; Hamblin, M.R.; Mozafari, M. Recent advances in 3D bioprinted tumor models for personalized medicine. Transl Oncol. 2023, 37, 101750. [Google Scholar] [CrossRef]
- Ireson, C.R.; Alavijeh, M.S.; Palmer, A.M.; Fowler, E.R.; Jones, H.J. The role of mouse tumour models in the discovery and development of anticancer drugs. British Journal of Cancer 2019, 121, 101–8. [Google Scholar] [CrossRef]
- Zhang, W.; Ding, Y.; He, H.; Chen, K.; Zeng, Q.; Cao, X.; et al. Prospects and challenges of ovarian cancer organoids in chemotherapy research (Review). Oncol Lett. 2025, 29, 198. [Google Scholar] [CrossRef]
- Marei, H.E. Stem cell therapy: a revolutionary cure or a pandora's box. Stem Cell Res Ther. 2025, 16, 255. [Google Scholar] [CrossRef] [PubMed]
- Sadiq, I.Z.; Abubakar, F.S.; Katsayal, B.S.; Ibrahim, B.; Adamu, A.; Usman, M.A.; et al. Stem cells in regenerative medicine: Unlocking therapeutic potential through stem cell therapy, 3D bioprinting, gene editing, and drug discovery. Biomedical Engineering Advances 2025, 9, 100172. [Google Scholar] [CrossRef]
- Chehelgerdi, M.; Behdarvand Dehkordi, F.; Chehelgerdi, M.; Kabiri, H.; Salehian-Dehkordi, H.; Abdolvand, M.; et al. Exploring the promising potential of induced pluripotent stem cells in cancer research and therapy. Mol Cancer 2023, 22, 189. [Google Scholar] [CrossRef] [PubMed]
- Eskandar, K. 3D bioprinting for facial reconstruction: Advances and challenges. Regenesis Repair Rehabilitation 2025, 1, 16–23. [Google Scholar] [CrossRef]
- Gopalakrishnan, M.; Kannan, D.; Elumalai, K.; Karunakar, K.; Jayaraj, S.; Devaraji, M.; et al. Advanced 3D biomaterials and bioprinting strategies for in vitro modeling of neurodegenerative diseases. Biomedical Technology 2025, 11, 100089. [Google Scholar] [CrossRef]
- Hoang, V.T.; Nguyen, Q.T.; Phan, T.T.K.; Pham, T.H.; Dinh, N.T.H.; Anh, L.P.H.; et al. Tissue Engineering and Regenerative Medicine: Perspectives and Challenges. MedComm (2020) 2025, 6, e70192. [Google Scholar] [CrossRef]
- Ayanoğlu, F.B.; Elçin, A.E.; Elçin, Y.M. Bioethical issues in genome editing by CRISPR-Cas9 technology. Turk J Biol. 2020, 44, 110–20. [Google Scholar] [CrossRef]
- Farag, V.E.; Devey, E.A.; Leong, K.W. The Interface of Gene Editing with Regenerative Medicine. Engineering 2025, 46, 73–100. [Google Scholar] [CrossRef]
- Brokowski, C.; Adli, M. CRISPR Ethics: Moral Considerations for Applications of a Powerful Tool. J Mol Biol. 2019, 431, 88–101. [Google Scholar] [CrossRef]
- Jose, A.; Kulkarni, P.; Thilakan, J.; Munisamy, M.; Malhotra, A.G.; Singh, J.; et al. Integration of pan-omics technologies and three-dimensional in vitro tumor models: an approach toward drug discovery and precision medicine. Molecular Cancer 2024, 23, 50. [Google Scholar] [CrossRef]
- Qu, S.; Xu, R.; Yi, G.; Li, Z.; Zhang, H.; Qi, S.; et al. Patient-derived organoids in human cancer: a platform for fundamental research and precision medicine. Mol Biomed. 2024, 5, 6. [Google Scholar] [CrossRef]
- Lv, J.; Du, X.; Wang, M.; Su, J.; Wei, Y.; Xu, C. Construction of tumor organoids and their application to cancer research and therapy. Theranostics 2024, 14, 1101–25. [Google Scholar] [CrossRef] [PubMed]
- Khorsandi, D.; Yang, J.-W.; Foster, S.; Khosravi, S.; Hosseinzadeh Kouchehbaghi, N.; Zarei, F.; et al. Patient-Derived Organoids as Therapy Screening Platforms in Cancer Patients. Advanced Healthcare Materials 2024, 13, 2302331. [Google Scholar] [CrossRef] [PubMed]
- Parvin, N.; Joo, S.W.; Jung, J.H.; Mandal, T.K. Multimodal AI in Biomedicine: Pioneering the Future of Biomaterials, Diagnostics, and Personalized Healthcare. Nanomaterials 2025, 15, 895. [Google Scholar] [CrossRef] [PubMed]
- Imani, S.; Li, X.; Chen, K.; Maghsoudloo, M.; Jabbarzadeh Kaboli, P.; Hashemi, M.; et al. Computational biology and artificial intelligence in mRNA vaccine design for cancer immunotherapy. Front Cell Infect Microbiol. 2024, 14, 1501010. [Google Scholar] [CrossRef]
- Wang, H.; Li, X.; You, X.; Zhao, G. Harnessing the power of artificial intelligence for human living organoid research. Bioactive Materials 2024, 42, 140–64. [Google Scholar] [CrossRef]
- Ravichandran, M.; Maddalo, D. Applications of CRISPR-Cas9 for advancing precision medicine in oncology: from target discovery to disease modeling. Front Genet. 2023, 14, 1273994. [Google Scholar] [CrossRef]
- Li, T.; Li, S.; Kang, Y.; Zhou, J.; Yi, M. Harnessing the evolving CRISPR/Cas9 for precision oncology. Journal of Translational Medicine 2024, 22, 749. [Google Scholar] [CrossRef]
- Cao, R.; Fomina, A.; McGuigan, A.P. Tissue-engineered Cancer Models in Drug Screening. In Biomaterial Based Approaches to Study the Tumour; Winter, J.O., Rao, S., Eds.; The Royal Society of Chemistry: Microenvironment, 2022; p. 0. [Google Scholar]
| Strategy | Mechanism/Approach | Advatages |
|---|---|---|
| In silico sgRNA design | Bioinformatics tools to predict and avoid off-target sequences | Reduces the likelihood of off-target cleavage |
| High-fidelity Cas9 variants (e.g., eSpCas9) | Engineered Cas9 with reduced non-specific binding | Minimizes unintended genome editing |
| Paired Cas9 nickases | Two Cas9 nickases targeting opposite strands, creating DSB only at the intended site | Decreases off-target DSBs and indels |
| Truncated sgRNAs | Shortened guide RNAs to reduce off-target binding | Enhances specificity without sacrificing on-target efficiency |
| Genome-wide off-target screening | Techniques like GUIDE-seq, CIRCLE-seq, and Digenome-seq | Validates safety before clinical application |
| Cas9 Modification | Application |
|---|---|
| Cas9 nickase | Introduces a single-strand break in DNA; using a pair of nickases to target opposite strands enhances specificity and reduces off-target mutations. |
| deadCas9 (dCas9) | Lacks nuclease activity; functions as a programmable DNA-binding scaffold to recruit regulatory proteins or effector molecules. |
| dCas9 with KRAB domain | Inhibits gene expression by recruiting transcriptional repressors, silencing downstream gene activity. |
| dCas9 with VP64 | Activates transcription of downstream genes by recruiting transcriptional activators. |
| dCas9 with methylases/demethylases or histone deacetylases | Modulates epigenetic marks at target loci, enabling control of gene expression through epigenetic regulation. |
|
Cas9 nickase with adenine or cytosine deaminase (Base editors; BEs) |
Allows precise base transitions (e.g., A→G, C→T) without introducing DSBs, offering efficient and accurate gene correction. |
|
Cas9 nickase with a reverse transcription domain (Prime editors; PEs) |
Permits the insertion, deletion, or substitution of specific DNA sequences with high precision and without generating DSBs. |
| Delivery Type | Method | Advantages | Disadvantages |
|---|---|---|---|
| Capsule | Viral vector | - High delivery efficiency - Broad range of available viral systems | - Immunogenic response - Potential pathogenicity - Genome integration risk |
| AAV (Adeno-associated virus) | - Low immunogenicity - Non-integrating nature - Tissue-specific tropism via serotypes | - Low packaging capacity | |
| Lipid-based vector | - Customizable surface modifications for targeted delivery | - Low transfection efficiency - Possible cytotoxicity | |
| Cas9 Form | Plasmid DNA | - Stable and long-term expression - Easy to produce | - Slow onset of gene editing - Requires nuclear entry |
| mRNA | - Rapid onset of editing - Transient expression reduces off-target effects | - Unstable and rapidly degraded - Requires cold-chain storage | |
| Protein (Cas9 RNP complex) | - Immediate gene editing upon delivery - Lower risk of off-target effects due to short activity window | - Difficult intracellular delivery - Risk of bacterial contaminants |
| Consideration | Description |
|---|---|
| Germline Editing | Potential for heritable genetic modifications in bioprinted reproductive tissues |
| Human Enhancement | Ethical debate over non-therapeutic gene edits for performance or appearance |
| Chimeric Constructs | Concerns over creating human-animal hybrids with cognitive functions |
| Informed Consent and Data Privacy | Challenges in obtaining consent for the use of edited tissues and genomic data |
| Regulatory Oversight | Lack of clear, unified regulations for combined bioprinting and gene editing |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




