Submitted:
30 January 2026
Posted:
02 February 2026
You are already at the latest version
Abstract
Keywords:
I. Introduction
II. Mitochondrial Function and Aging
III. Mitochondrial Drivers of Aging
Mitochondrial Driver of Aging Nº1: ATP Crisis
Mitochondrial Driver of Aging N°2: ETC-SCs Assembly Defects
Mitochondrial Driver of Aging N°3: ROS
Mitochondrial Driver of Aging N°4: mtDNA Damage
Mitochondrial Driver of Aging N°5: mtDNA Mutations
Mitochondrial Driver of Aging N°6: mtDNA Heteroplasmy
Mitochondrial Driver of Aging N°7: mtDNA Repair Mechanisms
Mitochondrial Driver of Aging N°8: mtDNA Copy Number Decrease
Mitochondrial Driver of Aging N°9 & 10: Mito Dynamics Deficiency
Mitochondrial Driver of Aging N°11: Mito-Biogenesis Deficiency
Mitochondrial Driver of Aging N°12: Mitophagy Deficiency
IV. Discussion
The Vicious Cycle of Mitophagy, Biogenesis, ROS, and Mutations
V. Acknowledgments and Conflicts of Interest
References
- D. N. J. De Grey and Michael. Rae, “Ending aging: the rejuvenation breakthroughs that could reverse human aging in our lifetime,” p. 433, 2013.
- J. P. De Magalhães, M. Stevens, and D. Thornton, “The Business of Anti-Aging Science,” Trends Biotechnol., vol. 35, no. 11, pp. 1062–1073, Nov. 2017. [CrossRef]
- M. A. Lee R, “Cost of Aging -- Finance & Development.” Accessed: Nov. 14, 2025. [Online]. Available: https://www.imf.org/external/pubs/ft/fandd/2017/03/lee.htm.
- S. Ogura and M. M. Jakovljevic, “Editorial: Global population aging - health care, social and economic consequences,” Front. Public Health, vol. 6, no. NOV, p. 430033, Nov. 2018. [CrossRef]
- W. James, J. Cossman, and J. K. Wolf, “Persistence of death in the United States: The remarkably different mortality patterns between America’s Heartland and Dixieland,” Demogr. Res., vol. 39, no. 1, pp. 897–910, Oct. 2018. [CrossRef]
- European Union Stats, “Ageing Europe-statistics on population developments Statistics Explained,” 2020, Accessed: Nov. 14, 2025. [Online]. Available: https://ec.europa.eu/eurostat/statisticsexplained/.
- W. Lutz, W. Sanderson, and S. Scherbov, “The coming acceleration of global population ageing,” Nature 2007 451:7179, vol. 451, no. 7179, pp. 716–719, Jan. 2008. [CrossRef]
- Z. Ismail, W. I. W. Ahmad, S. H. Hamjah, and I. K. Astina, “The Impact of Population Ageing: A Review,” Iran. J. Public Health, vol. 50, no. 12, pp. 2451–2460, 2021. [CrossRef]
- United Nations, “World Population Prospects 2022 World Population Prospects 2022 Summary of Results,” 2022.
- S. jin Li et al., “Population aging and trends of pulmonary tuberculosis incidence in the elderly,” BMC Infect. Dis., vol. 21, no. 1, Dec. 2021. [CrossRef]
- L. Wu, Z. Huang, and Z. Pan, “The spatiality and driving forces of population ageing in China,” PLoS One, vol. 16, no. 1, p. e0243559, Jan. 2021. [CrossRef]
- N. Glasgow and E. H. Berry, “Rural aging in 21st century America,” Rural Aging in 21st Century America, pp. 1–384, Jan. 2013. [CrossRef]
- G. Marois, A. Bélanger, and W. Lutz, “Population aging, migration, and productivity in Europe,” Proc. Natl. Acad. Sci. U. S. A., vol. 117, no. 14, pp. 7690–7695, Apr. 2020. [CrossRef]
- Z. Li et al., “Aging and age-related diseases: from mechanisms to therapeutic strategies,” Biogerontology 2021 22:2, vol. 22, no. 2, pp. 165–187, Jan. 2021. [CrossRef]
- Y. Cai et al., “The landscape of aging,” Sci. China Life Sci., vol. 65, no. 12, pp. 2354–2454, Dec. 2022. [CrossRef]
- J. P. Da Costa, R. Vitorino, G. M. Silva, C. Vogel, A. C. Duarte, and T. Rocha-Santos, “A synopsis on aging-Theories, mechanisms and future prospects,” Ageing Res. Rev., vol. 29, pp. 90–112, Aug. 2016. [CrossRef]
- S. Tabibzadeh, “Cell-centric hypotheses of aging,” Front. Biosci. (Landmark Ed)., vol. 26, no. 1, pp. 1–49, Jan. 2021. [CrossRef]
- D. Gems, “The hyperfunction theory: an emerging paradigm for the biology of aging,” Ageing Res. Rev., vol. 74, p. 101557, Feb. 2022. [CrossRef]
- A. A. Johnson, M. N. Shokhirev, and B. Shoshitaishvili, “Revamping the evolutionary theories of aging,” Ageing Res. Rev., vol. 55, Nov. 2019. [CrossRef]
- M. I. Brengdahl, C. M. Kimber, V. N. Shenoi, M. Dumea, A. Mital, and U. Friberg, “Age-specific effects of deletions: implications for aging theories,” Evolution, vol. 77, no. 1, pp. 254–263, Jan. 2023. [CrossRef]
- P. R. Rich, “A perspective on Peter Mitchell and the chemiosmotic theory,” J. Bioenerg. Biomembr., vol. 40, no. 5, pp. 407–410, Oct. 2008. [CrossRef]
- López-Otín, M. A. Blasco, L. Partridge, M. Serrano, and G. Kroemer, “Hallmarks of aging: An expanding universe,” Cell, vol. 186, no. 2, pp. 243–278, Jan. 2023. [CrossRef]
- S. K. Bhutia, P. P. Naik, D. P. Panigrahi, C. S. Bhol, and K. K. Mahapatra, “Mitophagy, Diseases, and Aging,” Models, Molecules and Mechanisms in Biogerontology, pp. 177–191, 2019. [CrossRef]
- A. Dillin et al., “Rates of Behavior and Aging Specified by Mitochondrial Function During Development,” Science (1979)., vol. 298, no. 5602, pp. 2398–2401, Dec. 2002. [CrossRef]
- C. Wallace, “Mitochondria and Cancer: Warburg Addressed,” Cold Spring Harb. Symp. Quant. Biol., vol. 70, pp. 363–374, Jan. 2005. [CrossRef]
- C. Wallace, “A mitochondrial paradigm of metabolic and degenerative diseases, aging, and cancer: A dawn for evolutionary medicine,” Annu. Rev. Genet., vol. 39, no. Volume 39, 2005, pp. 359–407, Dec. 2005. [CrossRef]
- S. Bartman, G. Coppotelli, and J. M. Ross, “Mitochondrial Dysfunction: A Key Player in Brain Aging and Diseases,” Curr. Issues Mol. Biol., vol. 46, no. 3, pp. 1987–2026, Mar. 2024. [CrossRef]
- M. Picard and O. S. Shirihai, “Mitochondrial signal transduction,” Cell Metab., vol. 34, no. 11, pp. 1620–1653, Nov. 2022. [CrossRef]
- S. van der Rijt, M. Molenaars, R. L. McIntyre, G. E. Janssens, and R. H. Houtkooper, “Integrating the Hallmarks of Aging Throughout the Tree of Life: A Focus on Mitochondrial Dysfunction,” Front. Cell Dev. Biol., vol. 8, Nov. 2020. [CrossRef]
- B. Kothe, S. Klein, and S. N. Petrosky, “Urolithin A as a Potential Agent for Prevention of Age-Related Disease: A Scoping Review,” Cureus, vol. 15, no. 7, Jul. 2023. [CrossRef]
- S. B. Poudel et al., “Effects of GH/IGF on the Aging Mitochondria,” Cells, vol. 9, no. 6, Jun. 2020. [CrossRef]
- Maldonado, S. Morales-Pison, F. Urbina, and A. Solari, “Aging Hallmarks and the Role of Oxidative Stress,” Antioxidants 2023, Vol. 12, Page 651, vol. 12, no. 3, p. 651, Mar. 2023. [CrossRef]
- A. De Gaetano, L. Gibellini, G. Zanini, M. Nasi, A. Cossarizza, and M. Pinti, “Mitophagy and Oxidative Stress: The Role of Aging,” Antioxidants, vol. 10, no. 5, p. 794, May 2021. [CrossRef]
- M. C. Sanchez, S. Lancel, E. Boulanger, and R. Neviere, “Targeting Oxidative Stress and Mitochondrial Dysfunction in the Treatment of Impaired Wound Healing: A Systematic Review,” Antioxidants (Basel)., vol. 7, no. 8, Aug. 2018. [CrossRef]
- Y. Li, L. Berliocchi, Z. Li, and L. J. Rasmussen, “Interactions between mitochondrial dysfunction and other hallmarks of aging: Paving a path toward interventions that promote healthy old age,” Aging Cell, vol. 23, no. 1, Jan. 2024. [CrossRef]
- J. I. Jiménez-Loygorri et al., “Mitophagy curtails cytosolic mtDNA-dependent activation of cGAS/STING inflammation during aging,” Nat. Commun., vol. 15, no. 1, Dec. 2024. [CrossRef]
- J. Tian et al., “Hippocampal transcriptome-wide association study and pathway analysis of mitochondrial solute carriers in Alzheimer’s disease,” Translational Psychiatry 2024 14:1, vol. 14, no. 1, pp. 250-, Jun. 2024. [CrossRef]
- S. Xie, S. C. Xu, W. Deng, and Q. Tang, “Metabolic landscape in cardiac aging: insights into molecular biology and therapeutic implications,” Signal Transduct. Target. Ther., vol. 8, no. 1, Dec. 2023. [CrossRef]
- L. Holper, D. Ben-Shachar, and J. Mann, “Multivariate meta-analyses of mitochondrial complex I and IV in major depressive disorder, bipolar disorder, schizophrenia, Alzheimer disease, and Parkinson disease,” Neuropsychopharmacology, vol. 44, no. 5, pp. 837–849, Apr. 2019. [CrossRef]
- S. Chen et al., “TOMM40 genetic variants associated with healthy aging and longevity: a systematic review,” BMC Geriatrics 2022 22:1, vol. 22, no. 1, pp. 667-, Aug. 2022. [CrossRef]
- J. I. Escrig-Larena, S. Delgado-Pulido, and M. Mittelbrunn, “Mitochondria during T cell aging,” Semin. Immunol., vol. 69, p. 101808, Sep. 2023. [CrossRef]
- C. C. S. Chini, H. S. Cordeiro, N. L. K. Tran, and E. N. Chini, “NAD metabolism: Role in senescence regulation and aging,” Aging Cell, vol. 23, no. 1, Jan. 2024. [CrossRef]
- X. Yang, R. Zhang, K. Nakahira, and Z. Gu, “Mitochondrial DNA Mutation, Diseases, and Nutrient-Regulated Mitophagy,” Annu. Rev. Nutr., vol. 39, no. Volume 39, 2019, pp. 201–226, Aug. 2019. [CrossRef]
- A. Suomalainen and J. Nunnari, “Mitochondria at the crossroads of health and disease,” Cell, vol. 187, no. 11, pp. 2601–2627, May 2024. [CrossRef]
- D. A. Harris and A. M. Das, “Control of mitochondrial ATP synthesis in the heart,” Biochem. J., vol. 280 (Pt 3), no. Pt 3, pp. 561–573, 1991. [CrossRef]
- D. J. Pagliarini and J. Rutter, “Hallmarks of a new era in mitochondrial biochemistry,” Genes Dev., vol. 27, no. 24, pp. 2615–2627, Dec. 2013. [CrossRef]
- R. Kerr, S. Jabbari, and I. G. Johnston, “Intracellular Energy Variability Modulates Cellular Decision-Making Capacity,” Scientific Reports 2019 9:1, vol. 9, no. 1, pp. 20196-, Dec. 2019. [CrossRef]
- Fernandez-Vizarra and M. Zeviani, “Mitochondrial disorders of the OXPHOS system,” FEBS Lett., vol. 595, no. 8, pp. 1062–1106, Apr. 2021. [CrossRef]
- J. A. Amorim, G. Coppotelli, A. P. Rolo, C. M. Palmeira, J. M. Ross, and D. A. Sinclair, “Mitochondrial and metabolic dysfunction in ageing and age-related diseases,” Nat. Rev. Endocrinol., vol. 18, no. 4, pp. 243–258, Apr. 2022. [CrossRef]
- M. Patron, H. G. Sprenger, and T. Langer, “m-AAA proteases, mitochondrial calcium homeostasis and neurodegeneration,” Cell Res., vol. 28, no. 3, pp. 296–306, Mar. 2018. [CrossRef]
- R. Rizzuto et al., “Close contacts with the endoplasmic reticulum as determinants of mitochondrial Ca2+ responses,” Science, vol. 280, no. 5370, pp. 1763–1766, Jun. 1998. [CrossRef]
- R. H. Houtkooper et al., “Mitonuclear protein imbalance as a conserved longevity mechanism,” Nature, vol. 497, no. 7450, pp. 451–457, 2013. [CrossRef]
- J. C. Goldstein, N. J. Waterhouse, P. Juin, G. I. Evan, and D. R. Green, “The coordinate release of cytochrome c during apoptosis is rapid, complete and kinetically invariant,” Nat. Cell Biol., vol. 2, no. 3, pp. 156–162, 2000. [CrossRef]
- X. Liu, C. N. Kim, J. Yang, R. Jemmerson, and X. Wang, “Induction of apoptotic program in cell-free extracts: requirement for dATP and cytochrome c,” Cell, vol. 86, no. 1, pp. 147–157, Jul. 1996. [CrossRef]
- M. Feric, T. G. Demarest, J. Tian, D. L. Croteau, V. A. Bohr, and T. Misteli, “Self-assembly of multi-component mitochondrial nucleoids via phase separation,” EMBO J., vol. 40, no. 6, Mar. 2021. [CrossRef]
- A. S. Monzel, M. Levin, and M. Picard, “The energetics of cellular life transitions,” Life metabolism, vol. 3, no. 3, Jun. 2024. [CrossRef]
- R. Lill, R. Lill, and S. A. Freibert, “Mechanisms of Mitochondrial Iron-Sulfur Protein Biogenesis,” Annu. Rev. Biochem., vol. 89, pp. 471–499, Jun. 2020. [CrossRef]
- D. HARMAN, “Aging: a theory based on free radical and radiation chemistry,” J. Gerontol., vol. 11, no. 3, pp. 298–300, 1956. [CrossRef]
- W. Yang and S. Hekimi, “A Mitochondrial Superoxide Signal Triggers Increased Longevity in Caenorhabditis elegans,” PLoS Biol., vol. 8, no. 12, p. e1000556, Dec. 2010. [CrossRef]
- C. Fang, X. Wei, and Y. Wei, “Mitochondrial DNA in the regulation of innate immune responses,” Protein Cell, vol. 7, no. 1, pp. 11–16, Jan. 2016. [CrossRef]
- A. Fairbrother-Browne et al., “Mitochondrial-nuclear cross-talk in the human brain is modulated by cell type and perturbed in neurodegenerative disease,” Commun. Biol., vol. 4, no. 1, Dec. 2021. [CrossRef]
- R. J. Youle and D. P. Narendra, “Mechanisms of mitophagy,” Nat. Rev. Mol. Cell Biol., vol. 12, no. 1, pp. 9–14, Jan. 2011. [CrossRef]
- B. M. Baker, A. M. Nargund, T. Sun, and C. M. Haynes, “Protective coupling of mitochondrial function and protein synthesis via the eIF2α kinase GCN-2,” PLoS Genet., vol. 8, no. 6, Jun. 2012. [CrossRef]
- J. Yun and T. Finkel, “Mitohormesis,” Cell Metab., vol. 19, no. 5, pp. 757–766, May 2014. [CrossRef]
- K. Palikaras, E. Lionaki, and N. Tavernarakis, “Coordination of mitophagy and mitochondrial biogenesis during ageing in C. elegans,” Nature, vol. 521, no. 7553, pp. 525–528, May 2015. [CrossRef]
- E. Owusu-Ansah, W. Song, and N. Perrimon, “XMuscle mitohormesis promotes longevity via systemic repression of insulin signaling,” Cell, vol. 155, no. 3, p. 699, Oct. 2013. [CrossRef]
- Q. Zhao, J. Wang, I. V. Levichkin, S. Stasinopoulos, M. T. Ryan, and N. J. Hoogenraad, “A mitochondrial specific stress response in mammalian cells,” EMBO J., vol. 21, no. 17, pp. 4411–4419, Sep. 2002. [CrossRef]
- R. D. Martinus et al., “Selective induction of mitochondrial chaperones in response to loss of the mitochondrial genome,” Eur. J. Biochem., vol. 240, no. 1, pp. 98–103, Jan. 1996. [CrossRef]
- K. M. Berendzen et al., “Neuroendocrine Coordination of Mitochondrial Stress Signaling and Proteostasis,” Cell, vol. 166, no. 6, pp. 1553-1563.e10, Sep. 2016. [CrossRef]
- N. S. Chandel, “Mitochondria as signaling organelles,” BMC Biology 2014 12:1, vol. 12, no. 1, pp. 34-, May 2014. [CrossRef]
- C. Lee et al., “The mitochondrial-derived peptide MOTS-c promotes metabolic homeostasis and reduces obesity and insulin resistance,” Cell Metab., vol. 21, no. 3, pp. 443–454, Mar. 2015. [CrossRef]
- Y. Hashimoto, S. Ookuma, and E. Nishida, “Lifespan extension by suppression of autophagy genes in Caenorhabditis elegans,” Genes Cells, vol. 14, no. 6, pp. 717–726, 2009. [CrossRef]
- R. C. Scarpulla, R. B. Vega, and D. P. Kelly, “Transcriptional integration of mitochondrial biogenesis,” Trends in Endocrinology and Metabolism, vol. 23, no. 9, pp. 459–466, Sep. 2012. [CrossRef]
- E. Nisoli et al., “Calorie restriction promotes mitochondrial biogenesis by inducing the expression of eNOS,” Science, vol. 310, no. 5746, pp. 314–317, Oct. 2005. [CrossRef]
- A. M. Pickrell and R. J. Youle, “The roles of PINK1, Parkin, and mitochondrial fidelity in parkinson’s disease,” Neuron, vol. 85, no. 2, pp. 257–273, Jan. 2015. [CrossRef]
- B. Cannon and J. Nedergaard, “Brown adipose tissue: function and physiological significance,” Physiol. Rev., vol. 84, no. 1, pp. 277–359, Jan. 2004. [CrossRef]
- S. H. Du et al., “Nicotinamide mononucleotide ameliorates acute lung injury by inducing mitonuclear protein imbalance and activating the UPRmt,” Exp. Biol. Med., vol. 247, no. 14, pp. 1264–1276, Jul. 2022. [CrossRef]
- R. Liang, L. Zhu, Y. Huang, J. Chen, and Q. Tang, “Mitochondria: fundamental characteristics, challenges, and impact on aging,” Biogerontology, vol. 25, no. 6, pp. 923–941, Nov. 2024. [CrossRef]
- Tirichen, H. Yaigoub, W. Xu, C. Wu, R. Li, and Y. Li, “Mitochondrial Reactive Oxygen Species and Their Contribution in Chronic Kidney Disease Progression Through Oxidative Stress,” Front. Physiol., vol. 12, Apr. 2021. [CrossRef]
- L. Liu, X. Liao, H. Wu, Y. Li, Y. Zhu, and Q. Chen, “Mitophagy and Its Contribution to Metabolic and Aging-Associated Disorders,” Antioxid. Redox Signal., vol. 32, no. 12, pp. 906–927, Apr. 2020. [CrossRef]
- S. Javadov, S. Jang, X. R. Chapa-Dubocq, Z. Khuchua, and A. K. Camara, “Mitochondrial respiratory supercomplexes in mammalian cells: structural versus functional role,” J. Mol. Med. (Berl)., vol. 99, no. 1, pp. 57–73, Jan. 2021. [CrossRef]
- V. Larosa and C. Remacle, “Insights into the respiratory chain and oxidative stress,” Biosci. Rep., vol. 38, no. 5, Oct. 2018. [CrossRef]
- T. Lobo-Jarne and C. Ugalde, “Respiratory chain supercomplexes: Structures, function and biogenesis,” Semin. Cell Dev. Biol., vol. 76, pp. 179–190, Apr. 2018. [CrossRef]
- J. A. Letts, K. Fiedorczuk, and L. A. Sazanov, “The architecture of respiratory supercomplexes,” Nature 2016 537:7622, vol. 537, no. 7622, pp. 644–648, Sep. 2016. [CrossRef]
- S. Guan, L. Zhao, and R. Peng, “Mitochondrial Respiratory Chain Supercomplexes: From Structure to Function,” Int. J. Mol. Sci., vol. 23, no. 22, Nov. 2022. [CrossRef]
- A. Toth et al., “Kinetic coupling of the respiratory chain with ATP synthase, but not proton gradients, drives ATP production in cristae membranes,” Proc. Natl. Acad. Sci. U. S. A., vol. 117, no. 5, pp. 2412–2421, Feb. 2020. [CrossRef]
- L. D. Zorova et al., “Mitochondrial membrane potential,” Anal. Biochem., vol. 552, pp. 50–59, Jul. 2018. [CrossRef]
- Martínez-Reyes et al., “TCA cycle and mitochondrial membrane potential are necessary for diverse biological functions,” Mol. Cell, vol. 61, no. 2, p. 199, Jan. 2015. [CrossRef]
- Mengel-From, M. Thinggaard, C. Dalgård, K. O. Kyvik, K. Christensen, and L. Christiansen, “Mitochondrial DNA copy number in peripheral blood cells declines with age and is associated with general health among elderly,” Hum. Genet., vol. 133, no. 9, pp. 1149–1159, Sep. 2014. [CrossRef]
- G. Sturm et al., “OxPhos defects cause hypermetabolism and reduce lifespan in cells and in patients with mitochondrial diseases,” Commun. Biol., vol. 6, no. 1, Dec. 2023. [CrossRef]
- X. Zheng et al., “Metabolic reprogramming during neuronal differentiation from aerobic glycolysis to neuronal oxidative phosphorylation,” Elife, vol. 5, no. JUN2016, p. e13374, Jun. 2016. [CrossRef]
- E. S. Chocron, E. Munkácsy, and A. M. Pickering, “Cause or casualty: The role of mitochondrial DNA in aging and age-associated disease,” Biochim. Biophys. Acta Mol. Basis Dis., vol. 1865, no. 2, pp. 285–297, Feb. 2019. [CrossRef]
- U. Basu, A. M. Bostwick, K. Das, K. E. Dittenhafer-Reed, and S. S. Patel, “Structure, mechanism, and regulation of mitochondrial DNA transcription initiation,” Journal of Biological Chemistry, vol. 295, no. 52, pp. 18406–18425, Dec. 2020. [CrossRef]
- W. Wei et al., “Nuclear-embedded mitochondrial DNA sequences in 66,083 human genomes,” Nature, vol. 611, no. 7934, pp. 105–114, Nov. 2022. [CrossRef]
- G. De Benedittis et al., “Alteration of Mitochondrial DNA Copy Number and Increased Expression Levels of Mitochondrial Dynamics-Related Genes in Sjögren’s Syndrome,” Biomedicines, vol. 10, no. 11, Nov. 2022. [CrossRef]
- E. S. Ramos et al., “Mitochondrial fusion is required for regulation of mitochondrial DNA replication,” PLoS Genet., vol. 15, no. 6, Jun. 2019. [CrossRef]
- P. Katajisto et al., “Stem cells. Asymmetric apportioning of aged mitochondria between daughter cells is required for stemness,” Science, vol. 348, no. 6232, pp. 340–343, Apr. 2015. [CrossRef]
- D. C. Wallace, “A mitochondrial bioenergetic etiology of disease,” J. Clin. Invest., vol. 123, no. 4, pp. 1405–1412, Apr. 2013. [CrossRef]
- R. Gupta et al., “Nuclear genetic control of mtDNA copy number and heteroplasmy in humans,” Nature, vol. 620, no. 7975, pp. 839–848, Aug. 2023. [CrossRef]
- N. A. Bonekamp et al., “High levels of TFAM repress mammalian mitochondrial DNA transcription in vivo,” Life Sci. Alliance, vol. 4, no. 11, 2021. [CrossRef]
- S. M. A. Radzak, S. Z. N. M. Khair, F. Ahmad, A. Patar, Z. Idris, and A. A. M. Yusoff, “Insights regarding mitochondrial DNA copy number alterations in human cancer (Review),” Int. J. Mol. Med., vol. 50, no. 2, Aug. 2022. [CrossRef]
- B. Peter and M. Falkenberg, “TWINKLE and Other Human Mitochondrial DNA Helicases: Structure, Function and Disease,” Genes (Basel)., vol. 11, no. 4, Apr. 2020. [CrossRef]
- A. Picca et al., “The contribution of mitochondrial DNA alterations to aging, cancer, and neurodegeneration,” Exp. Gerontol., vol. 178, Jul. 2023. [CrossRef]
- Picca et al., “Circulating Mitochondrial DNA and Inter-Organelle Contact Sites in Aging and Associated Conditions,” Cells, vol. 11, no. 4, Feb. 2022. [CrossRef]
- N. N. Wu, Y. Zhang, and J. Ren, “Mitophagy, Mitochondrial Dynamics, and Homeostasis in Cardiovascular Aging,” Oxid. Med. Cell. Longev., vol. 2019, 2019. [CrossRef]
- X. Hong et al., “Mitochondrial dynamics maintain muscle stem cell regenerative competence throughout adult life by regulating metabolism and mitophagy,” Cell Stem Cell, vol. 29, no. 9, pp. 1298-1314.e10, Sep. 2022. [CrossRef]
- Zhang et al., “The role of mitochondrial quality surveillance in skin aging: Focus on mitochondrial dynamics, biogenesis and mitophagy,” Ageing Res. Rev., vol. 87, Jun. 2023. [CrossRef]
- G. Chen, G. Kroemer, and O. Kepp, “Mitophagy: An Emerging Role in Aging and Age-Associated Diseases,” Front. Cell Dev. Biol., vol. 8, Mar. 2020. [CrossRef]
- W. N. Xu et al., “Mitochondrial NDUFA4L2 attenuates the apoptosis of nucleus pulposus cells induced by oxidative stress via the inhibition of mitophagy,” Exp. Mol. Med., vol. 51, no. 11, Nov. 2019. [CrossRef]
- L. Kang, S. Liu, J. Li, Y. Tian, Y. Xue, and X. Liu, “Parkin and Nrf2 prevent oxidative stress-induced apoptosis in intervertebral endplate chondrocytes via inducing mitophagy and anti-oxidant defenses,” Life Sci., vol. 243, Feb. 2020. [CrossRef]
- L. Xie et al., “CircERCC2 ameliorated intervertebral disc degeneration by regulating mitophagy and apoptosis through miR-182-5p/SIRT1 axis,” Cell Death Dis., vol. 10, no. 10, Oct. 2019. [CrossRef]
- Huang et al., “Compression-induced senescence of nucleus pulposus cells by promoting mitophagy activation via the PINK1/PARKIN pathway,” J. Cell. Mol. Med., vol. 24, no. 10, pp. 5850–5864, May 2020. [CrossRef]
- Liu, J. Wang, and Y. Zhou, “Upregulation of BNIP3 and translocation to mitochondria in nutrition deprivation induced apoptosis in nucleus pulposus cells,” Joint Bone Spine, vol. 79, no. 2, pp. 186–191, Mar. 2012. [CrossRef]
- Liu, C. Yuan, L. Pu, and J. Wang, “Nutrient deprivation induces apoptosis of nucleus pulposus cells via activation of the BNIP3/AIF signalling pathway,” Mol. Med. Rep., vol. 16, no. 5, pp. 7253–7260, Nov. 2017. [CrossRef]
- Z. He, L. Pu, C. Yuan, M. Jia, and J. Wang, “Nutrition deficiency promotes apoptosis of cartilage endplate stem cells in a caspase-independent manner partially through upregulating BNIP3,” Acta Biochim. Biophys. Sin. (Shanghai)., vol. 49, no. 1, pp. 25–32, Jan. 2017. [CrossRef]
- H. J. Shin et al., “Pink1-Mediated Chondrocytic Mitophagy Contributes to Cartilage Degeneration in Osteoarthritis,” J. Clin. Med., vol. 8, no. 11, Nov. 2019. [CrossRef]
- Sun, X. Jing, J. Guo, X. Yao, and F. Guo, “Mitophagy in degenerative joint diseases,” Autophagy, vol. 17, no. 9, pp. 2082–2092, 2021. [CrossRef]
- Babbar, S. Basu, B. Yang, D. L. Croteau, and V. A. Bohr, “Mitophagy and DNA damage signaling in human aging,” Mech. Ageing Dev., vol. 186, Mar. 2020. [CrossRef]
- A. Diot, K. Morten, and J. Poulton, “Mitophagy plays a central role in mitochondrial ageing,” Mamm. Genome, vol. 27, no. 7–8, pp. 381–395, Aug. 2016. [CrossRef]
- Y. Chen et al., “Mfn2 is involved in intervertebral disc degeneration through autophagy modulation,” Osteoarthritis Cartilage, vol. 28, no. 3, pp. 363–374, Mar. 2020. [CrossRef]
- K. Palikaras, I. Daskalaki, M. Markaki, and N. Tavernarakis, “Mitophagy and age-related pathologies: Development of new therapeutics by targeting mitochondrial turnover,” Pharmacol. Ther., vol. 178, pp. 157–174, Oct. 2017. [CrossRef]
- Bakula and M. Scheibye-Knudsen, “MitophAging: Mitophagy in Aging and Disease,” Front. Cell Dev. Biol., vol. 8, Apr. 2020. [CrossRef]
- T. A. Banarase et al., “Mitophagy regulation in aging and neurodegenerative disease,” Biophys. Rev., vol. 15, no. 2, pp. 239–255, Apr. 2023. [CrossRef]
- J. Faitg, D. D’Amico, C. Rinsch, and A. Singh, “Mitophagy Activation by Urolithin A to Target Muscle Aging,” Calcif. Tissue Int., vol. 114, no. 1, pp. 53–59, Jan. 2024. [CrossRef]
- J. Guo and W. C. Chiang, “Mitophagy in aging and longevity,” IUBMB Life, vol. 74, no. 4, pp. 296–316, Apr. 2022. [CrossRef]
- Tran and P. H. Reddy, “Defective Autophagy and Mitophagy in Aging and Alzheimer’s Disease,” Front. Neurosci., vol. 14, Jan. 2021. [CrossRef]
- T. J. LaRocca, R. A. Gioscia-Ryan, C. M. Hearon, and D. R. Seals, “The autophagy enhancer spermidine reverses arterial aging,” Mech. Ageing Dev., vol. 134, no. 7–8, pp. 314–320, Jul. 2013. [CrossRef]
- C. G. McCarthy et al., “Reconstitution of autophagy ameliorates vascular function and arterial stiffening in spontaneously hypertensive rats,” Am. J. Physiol. Heart Circ. Physiol., vol. 317, no. 5, pp. H1013–H1027, 2019. [CrossRef]
- Z. J. Schreckenberger, C. F. Wenceslau, B. Joe, and C. G. McCarthy, “Mitophagy in Hypertension-Associated Premature Vascular Aging,” Am. J. Hypertens., vol. 33, no. 9, pp. 804–812, Sep. 2020. [CrossRef]
- A. Rappe and T. G. McWilliams, “Mitophagy in the aging nervous system,” Front. Cell Dev. Biol., vol. 10, Oct. 2022. [CrossRef]
- B. Lombard, K. F. Chua, R. Mostoslavsky, S. Franco, M. Gostissa, and F. W. Alt, “DNA repair, genome stability, and aging,” Cell, vol. 120, no. 4, pp. 497–512, Feb. 2005. [CrossRef]
- C. McMahon et al., “TRIBE: Hijacking an RNA-Editing Enzyme to Identify Cell-Specific Targets of RNA-Binding Proteins,” Cell, vol. 165, no. 3, pp. 742–753, Apr. 2016. [CrossRef]
- L. Hayflick and P. S. Moorhead, “The serial cultivation of human diploid cell strains,” Exp. Cell Res., vol. 25, no. 3, pp. 585–621, 1961. [CrossRef]
- M. Zhou et al., “Acupoint catgut embedding improves senescence in a rat model of ageing by regulating mitophagy via the PINK1 pathway,” J. Cell. Mol. Med., vol. 25, no. 8, pp. 3816–3828, Apr. 2021. [CrossRef]
- M. Zatyka, S. Sarkar, and T. Barrett, “Autophagy in Rare (NonLysosomal) Neurodegenerative Diseases,” J. Mol. Biol., vol. 432, no. 8, pp. 2735–2753, Apr. 2020. [CrossRef]
- S. Salvioli et al., “Genes, ageing and longevity in humans: problems, advantages and perspectives,” Free Radic. Res., vol. 40, no. 12, pp. 1303–1323, Dec. 2006. [CrossRef]
- J. Zhang and P. A. Ney, “Reticulocyte mitophagy: monitoring mitochondrial clearance in a mammalian model,” Autophagy, vol. 6, no. 3, pp. 405–408, Apr. 2010. [CrossRef]
- T. Ito et al., “Regulation of myeloid leukaemia by the cell-fate determinant Musashi,” Nature, vol. 466, no. 7307, pp. 765–768, Aug. 2010. [CrossRef]
- W. J. Liang and Å. B. Gustafsson, “The Aging Heart: Mitophagy at the Center of Rejuvenation,” Front. Cardiovasc. Med., vol. 7, Feb. 2020. [CrossRef]
- S. A. Detmer and D. C. Chan, “Functions and dysfunctions of mitochondrial dynamics,” Nat. Rev. Mol. Cell Biol., vol. 8, no. 11, pp. 870–879, Nov. 2007. [CrossRef]
- R. Bernardi and P. P. Pandolfi, “Structure, dynamics and functions of promyelocytic leukaemia nuclear bodies,” Nat. Rev. Mol. Cell Biol., vol. 8, no. 12, pp. 1006–1016, Dec. 2007. [CrossRef]
- Yetkin-Arik et al., “The role of glycolysis and mitochondrial respiration in the formation and functioning of endothelial tip cells during angiogenesis,” Sci. Rep., vol. 9, no. 1, Dec. 2019. [CrossRef]
- Gauba, L. Guo, and H. Du, “Cyclophilin D Promotes Brain Mitochondrial F1FO ATP Synthase Dysfunction in Aging Mice,” J. Alzheimers Dis., vol. 55, no. 4, pp. 1351–1362, 2017. [CrossRef]
- G. Nicholls, “Oxidative stress and energy crises in neuronal dysfunction,” Ann. N. Y. Acad. Sci., vol. 1147, pp. 53–60, 2008. [CrossRef]
- G. Nicholls, “Spare respiratory capacity, oxidative stress and excitotoxicity,” Biochem. Soc. Trans., vol. 37, no. Pt 6, pp. 1385–1388, 2009. [CrossRef]
- C. Desler, T. L. Hansen, J. B. Frederiksen, M. L. Marcker, K. K. Singh, and L. Juel Rasmussen, “Is There a Link between Mitochondrial Reserve Respiratory Capacity and Aging?,” J. Aging Res., vol. 2012, 2012. [CrossRef]
- S. N. Chaudhari and E. T. Kipreos, “The Energy Maintenance Theory of Aging: Maintaining Energy Metabolism to Allow Longevity,” Bioessays, vol. 40, no. 8, Aug. 2018. [CrossRef]
- Y. Yuan, V. F. Cruzat, P. Newshome, J. Cheng, Y. Chen, and Y. Lu, “Regulation of SIRT1 in aging: Roles in mitochondrial function and biogenesis,” Mech. Ageing Dev., vol. 155, pp. 10–21, Apr. 2016. [CrossRef]
- C. Abboudi, “The Effects of Aging on Skeletal Muscle ATP Production,” The Science Journal of the Lander College of Arts and Sciences, vol. 12, no. 2, p. 10, Jan. 2019, Accessed: Nov. 15, 2025. [Online]. Available: https://touroscholar.touro.edu/sjlcas/vol12/iss2/10.
- Distefano and B. H. Goodpaster, “Effects of Exercise and Aging on Skeletal Muscle,” Cold Spring Harb. Perspect. Med., vol. 8, no. 3, Mar. 2018. [CrossRef]
- A. Hoshino et al., “The ADP/ATP translocase drives mitophagy independent of nucleotide exchange,” Nature, vol. 575, no. 7782, pp. 375–379, Nov. 2019. [CrossRef]
- Mikolajewicz, E. A. Zimmermann, B. M. Willie, and S. V. Komarova, “Mechanically stimulated ATP release from murine bone cells is regulated by a balance of injury and repair,” Elife, vol. 7, Oct. 2018. [CrossRef]
- V. Warnsmann, J. Meisterknecht, I. Wittig, and H. D. Osiewacz, “Aging of Podospora anserina Leads to Alterations of OXPHOS and the Induction of Non-Mitochondrial Salvage Pathways,” Cells, vol. 10, no. 12, Dec. 2021. [CrossRef]
- L. M. de Smalen et al., “Impaired age-associated mitochondrial translation is mitigated by exercise and PGC-1α,” Proc. Natl. Acad. Sci. U. S. A., vol. 120, no. 36, 2023. [CrossRef]
- Z. Vue et al., “3D reconstruction of murine mitochondria reveals changes in structure during aging linked to the MICOS complex,” Aging Cell, vol. 22, no. 12, Dec. 2023. [CrossRef]
- M. Frenzel, H. Rommelspacher, M. D. Sugawa, and N. A. Dencher, “Ageing alters the supramolecular architecture of OxPhos complexes in rat brain cortex,” Exp. Gerontol., vol. 45, no. 7–8, pp. 563–572, Aug. 2010. [CrossRef]
- Lopez-Fabuel, M. Resch-Beusher, M. Carabias-Carrasco, A. Almeida, and J. P. Bolaños, “Mitochondrial Complex I Activity is Conditioned by Supercomplex I-III2-IV Assembly in Brain Cells: Relevance for Parkinson’s Disease,” Neurochem. Res., vol. 42, no. 6, pp. 1676–1682, Jun. 2017. [CrossRef]
- A. Grünewald, K. R. Kumar, and C. M. Sue, “New insights into the complex role of mitochondria in Parkinson’s disease,” Prog. Neurobiol., vol. 177, pp. 73–93, Jun. 2019. [CrossRef]
- M. R. Cookson, “Parkinsonism due to mutations in PINK1, parkin, and DJ-1 and oxidative stress and mitochondrial pathways,” Cold Spring Harb. Perspect. Med., vol. 2, no. 9, 2012. [CrossRef]
- M. Kenney and J. P. Bennett, “Alzheimer’s Disease Frontal Cortex Mitochondria Show a Loss of Individual Respiratory Proteins but Preservation of Respiratory Supercomplexes,” Int. J. Alzheimers Dis., vol. 2019, 2019. [CrossRef]
- S. K. Sagwal and S. Bekeschus, “ROS Pleiotropy in Melanoma and Local Therapy with Physical Modalities,” Oxid. Med. Cell. Longev., vol. 2021, 2021. [CrossRef]
- Zhang, H. Yu, M. Q. Man, and L. Hu, “Aging in the dermis: Fibroblast senescence and its significance,” Aging Cell, vol. 23, no. 2, Feb. 2024. [CrossRef]
- Song et al., “FOXO-regulated OSER1 reduces oxidative stress and extends lifespan in multiple species,” Nat. Commun., vol. 15, no. 1, Dec. 2024. [CrossRef]
- M. Zhao et al., “Mitochondrial ROS promote mitochondrial dysfunction and inflammation in ischemic acute kidney injury by disrupting TFAM-mediated mtDNA maintenance,” Theranostics, vol. 11, no. 4, pp. 1845–1863, Jan. 2021. [CrossRef]
- A. Sen et al., “Mitochondrial membrane proteins and VPS35 orchestrate selective removal of mtDNA,” Nat. Commun., vol. 13, no. 1, Dec. 2022. [CrossRef]
- Yeo, C. Kang, M. C. Gomez-Cabrera, J. Vina, and L. L. Ji, “Intensified mitophagy in skeletal muscle with aging is downregulated by PGC-1alpha overexpression in vivo,” Free Radic. Biol. Med., vol. 130, pp. 361–368, Jan. 2019. [CrossRef]
- F. Fang et al., “Mitophagy inhibits amyloid-β and tau pathology and reverses cognitive deficits in models of Alzheimer’s disease,” Nat. Neurosci., vol. 22, no. 3, pp. 401–412, Mar. 2019. [CrossRef]
- N. Sun et al., “Measuring In Vivo Mitophagy,” Mol. Cell, vol. 60, no. 4, pp. 685–696, Nov. 2015. [CrossRef]
- E. Tresse et al., “Mitochondrial DNA damage triggers spread of Parkinson’s disease-like pathology,” Mol. Psychiatry, vol. 28, no. 11, pp. 4902–4914, Nov. 2023. [CrossRef]
- A. T. Erlich and D. A. Hood, “Mitophagy Regulation in Skeletal Muscle: Effect of Endurance Exercise and Age,” Journal of Science in Sport and Exercise 2019 1:3, vol. 1, no. 3, pp. 228–236, Nov. 2019. [CrossRef]
- M. Villanueva-Paz et al., “Parkin-mediated mitophagy and autophagy flux disruption in cellular models of MERRF syndrome,” Biochim. Biophys. Acta Mol. Basis Dis., vol. 1866, no. 6, Jun. 2020. [CrossRef]
- Xiong et al., “Modulation of miR-34a/SIRT1 signaling protects cochlear hair cells against oxidative stress and delays age-related hearing loss through coordinated regulation of mitophagy and mitochondrial biogenesis,” Neurobiol. Aging, vol. 79, pp. 30–42, Jul. 2019. [CrossRef]
- M. Vermulst et al., “Mitochondrial point mutations do not limit the natural lifespan of mice,” Nat. Genet., vol. 39, no. 4, pp. 540–543, Apr. 2007. [CrossRef]
- A. R. Lee et al., “Involvement of mitochondrial biogenesis during the differentiation of human periosteum-derived mesenchymal stem cells into adipocytes, chondrocytes and osteocytes,” Arch. Pharm. Res., vol. 42, no. 12, pp. 1052–1062, Dec. 2019. [CrossRef]
- Sarangarajan, S. Meera, R. Rukkumani, P. Sankar, and G. Anuradha, “Antioxidants: Friend or foe?,” Asian Pac. J. Trop. Med., vol. 10, no. 12, pp. 1111–1116, Dec. 2017. [CrossRef]
- A. Fontana and H. L. Gahlon, “Mechanisms of replication and repair in mitochondrial DNA deletion formation,” Nucleic Acids Res., vol. 48, no. 20, pp. 11244–11258, Nov. 2020. [CrossRef]
- Li, J. Slone, L. Fei, and T. Huang, “Mitochondrial DNA Variants and Common Diseases: A Mathematical Model for the Diversity of Age-Related mtDNA Mutations,” Cells, vol. 8, no. 6, 2019. [CrossRef]
- G. Shabalina et al., “Enhanced ROS Production in Mitochondria from Prematurely Aging mtDNA Mutator Mice,” Biochemistry (Mosc)., vol. 89, no. 2, pp. 279–298, Feb. 2024. [CrossRef]
- E. Vernucci et al., “Mitophagy and Oxidative Stress in Cancer and Aging: Focus on Sirtuins and Nanomaterials,” Oxid. Med. Cell. Longev., vol. 2019, 2019. [CrossRef]
- Adam Held, “An Introduction to Reactive Oxygen Species - Measurement of ROS in Cells,” 2012.
- G. Waris and H. Ahsan, “Reactive oxygen species: role in the development of cancer and various chronic conditions,” J. Carcinog., vol. 5, p. 14, May 2006. [CrossRef]
- H. A. L. Tuppen, E. L. Blakely, D. M. Turnbull, and R. W. Taylor, “Mitochondrial DNA mutations and human disease,” Biochim. Biophys. Acta Bioenerg., vol. 1797, no. 2, pp. 113–128, Feb. 2010. [CrossRef]
- “Understanding the Process of Aging: The Roles of Mitochondria: Free Radicals, and Antioxidants,” Understanding the Process of Aging, Jan. 1999. [CrossRef]
- N. Ghosh, A. Das, S. Chaffee, S. Roy, and C. K. Sen, “Reactive oxygen species, oxidative damage and cell death,” Immunity and Inflammation in Health and Disease: Emerging Roles of Nutraceuticals and Functional Foods in Immune Support, pp. 45–55, Jan. 2017. [CrossRef]
- A. Ayala, M. F. Muñoz, and S. Argüelles, “Lipid peroxidation: production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal,” Oxid. Med. Cell. Longev., vol. 2014, 2014. [CrossRef]
- B. K. Kennedy et al., “Geroscience: Linking aging to chronic disease,” Cell, vol. 159, no. 4, pp. 709–713, Nov. 2014. [CrossRef]
- C. Heidi Chial, “mtDNA and Mitochondrial Diseases | Learn Science at Scitable,” Nature Education, 2008, Accessed: Nov. 22, 2025. [Online]. Available: https://www.nature.com/scitable/topicpage/mtdna-and-mitochondrial-diseases-903/.
- W. Gray, “Mitochondrial evolution,” Cold Spring Harb. Perspect. Biol., vol. 4, no. 9, Sep. 2012. [CrossRef]
- Hahn and S. Zuryn, “The Cellular Mitochondrial Genome Landscape in Disease,” Trends Cell Biol., vol. 29, no. 3, pp. 227–240, Mar. 2019. [CrossRef]
- P. Burgstaller, I. G. Johnston, and J. Poulton, “Mitochondrial DNA disease and developmental implications for reproductive strategies,” Mol. Hum. Reprod., vol. 21, no. 1, p. 11, Jun. 2014. [CrossRef]
- Yu, “Generation, function and diagnostic value of mitochondrial DNA copy number alterations in human cancers,” Life Sci., vol. 89, no. 3–4, pp. 65–71, Jul. 2011. [CrossRef]
- A. Lodato et al., “Aging and neurodegeneration are associated with increased mutations in single human neurons,” Science, vol. 359, no. 6375, pp. 555–559, Feb. 2018. [CrossRef]
- A. Risques and S. R. Kennedy, “Aging and the rise of somatic cancer-associated mutations in normal tissues,” PLoS Genet., vol. 14, no. 1, Jan. 2018. [CrossRef]
- H. Ma et al., “Germline and somatic mtDNA mutations in mouse aging,” PLoS One, vol. 13, no. 7, p. e0201304, Jul. 2018. [CrossRef]
- A. M. Orogo et al., “Accumulation of Mitochondrial DNA Mutations Disrupts Cardiac Progenitor Cell Function and Reduces Survival,” J. Biol. Chem., vol. 290, no. 36, p. 22061, Sep. 2015. [CrossRef]
- J. Niemann et al., “An mtDNA mutation accelerates liver aging by interfering with the ROS response and mitochondrial life cycle,” Free Radic. Biol. Med., vol. 102, pp. 174–187, Jan. 2017. [CrossRef]
- A. Trifunovic et al., “Premature ageing in mice expressing defective mitochondrial DNA polymerase,” Nature, vol. 429, no. 6990, pp. 417–423, May 2004. [CrossRef]
- Yang et al., “NAD+ dependent UPRmt activation underlies intestinal aging caused by mitochondrial DNA mutations,” Nature Communications 2024 15:1, vol. 15, no. 1, pp. 546-, Jan. 2024. [CrossRef]
- Y. Quan, Y. Xin, G. Tian, J. Zhou, and X. Liu, “Mitochondrial ROS-Modulated mtDNA: A Potential Target for Cardiac Aging,” Oxid. Med. Cell. Longev., vol. 2020, no. 1, p. 9423593, Jan. 2020. [CrossRef]
- L. Samstag et al., “Deleterious mitochondrial DNA point mutations are overrepresented in Drosophila expressing a proofreading-defective DNA polymerase γ,” PLoS Genet., vol. 14, no. 11, Nov. 2018. [CrossRef]
- A. D. N. J. De Grey, “A proposed refinement of the mitochondrial free radical theory of aging,” Bioessays, vol. 19, no. 2, pp. 161–166, 1997. [CrossRef]
- G. S. Nido et al., “Ultradeep mapping of neuronal mitochondrial deletions in Parkinson’s disease,” Neurobiol. Aging, vol. 63, pp. 120–127, Mar. 2018. [CrossRef]
- A. Dabravolski, N. K. Sadykhov, A. G. Kartuesov, E. E. Borisov, V. N. Sukhorukov, and A. N. Orekhov, “The Role of Mitochondrial Abnormalities in Diabetic Cardiomyopathy,” Int. J. Mol. Sci., vol. 23, no. 14, Jul. 2022. [CrossRef]
- A. L. M. Smith et al., “Age-associated mitochondrial DNA mutations cause metabolic remodelling that contributes to accelerated intestinal tumorigenesis,” Nat. Cancer, vol. 1, no. 10, pp. 976–989, Oct. 2020. [CrossRef]
- J. Sturgis et al., “Modeling aging and retinal degeneration with mitochondrial DNA mutation burden,” bioRxiv, Dec. 2023. [CrossRef]
- E. R. Árnadóttir et al., “The rate and nature of mitochondrial DNA mutations in human pedigrees,” Cell, vol. 187, no. 15, pp. 3904-3918.e8, Jul. 2024. [CrossRef]
- J. R. Connell et al., “Pedigree derived mutation rate across the entire mitochondrial genome of the Norfolk Island population,” Sci. Rep., vol. 12, no. 1, Dec. 2022. [CrossRef]
- Sigurdardóttir, A. Helgason, J. R. Gulcher, K. Stefansson, and P. Donnelly, “The mutation rate in the human mtDNA control region,” Am. J. Hum. Genet., vol. 66, no. 5, pp. 1599–1609, 2000. [CrossRef]
- Santos et al., “Understanding Differences Between Phylogenetic and Pedigree-Derived mtDNA Mutation Rate: A Model Using Families from the Azores Islands (Portugal),” Mol. Biol. Evol., vol. 22, no. 6, pp. 1490–1505, Jun. 2005. [CrossRef]
- J. Wen et al., “Role of mismatch repair in aging,” Int. J. Biol. Sci., vol. 17, no. 14, p. 3923, 2021. [CrossRef]
- R. Filipovic and W. H. Koppenol, “The Haber-Weiss reaction – The latest revival,” Free Radic. Biol. Med., vol. 145, pp. 221–222, Dec. 2019. [CrossRef]
- L. Mao et al., “Estimation of the mtDNA mutation rate in aging mice by proteome analysis and mathematical modeling,” Exp. Gerontol., vol. 41, no. 1, pp. 11–24, Jan. 2006. [CrossRef]
- B. Nabholz, S. Glémin, and N. Galtier, “The erratic mitochondrial clock: variations of mutation rate, not population size, affect mtDNA diversity across birds and mammals,” BMC Evol. Biol., vol. 9, no. 1, 2009. [CrossRef]
- G. Lou, K. Palikaras, S. Lautrup, M. Scheibye-Knudsen, N. Tavernarakis, and E. F. Fang, “Mitophagy and Neuroprotection,” Trends Mol. Med., vol. 26, no. 1, pp. 8–20, Jan. 2020. [CrossRef]
- S. Schaack, E. K. H. Ho, and F. MacRae, “Disentangling the intertwined roles of mutation, selection and drift in the mitochondrial genome,” Philos. Trans. R. Soc. Lond. B Biol. Sci., vol. 375, no. 1790, 2020. [CrossRef]
- Lane, “Mitonuclear match: optimizing fitness and fertility over generations drives ageing within generations,” Bioessays, vol. 33, no. 11, pp. 860–869, Nov. 2011. [CrossRef]
- Lane, “The Problem with Mixing Mitochondria,” Cell, vol. 151, no. 2, pp. 246–248, Oct. 2012. [CrossRef]
- M. S. Sharpley et al., “Heteroplasmy of mouse mtDNA is genetically unstable and results in altered behavior and cognition,” Cell, vol. 151, no. 2, pp. 333–343, Oct. 2012. [CrossRef]
- J. B. Stewart and P. F. Chinnery, “The dynamics of mitochondrial DNA heteroplasmy: implications for human health and disease,” Nat. Rev. Genet., vol. 16, no. 9, pp. 530–542, Aug. 2015. [CrossRef]
- F. Ghiselli et al., “Natural Heteroplasmy and Mitochondrial Inheritance in Bivalve Molluscs,” Integr. Comp. Biol., vol. 59, no. 4, pp. 1016–1032, Oct. 2019. [CrossRef]
- L. N. Lakshmanan, Z. Yee, L. F. Ng, R. Gunawan, B. Halliwell, and J. Gruber, “Clonal expansion of mitochondrial DNA deletions is a private mechanism of aging in long-lived animals,” Aging Cell, vol. 17, no. 5, Oct. 2018. [CrossRef]
- A. E. Vincent et al., “Subcellular origin of mitochondrial DNA deletions in human skeletal muscle,” Ann. Neurol., vol. 84, no. 2, pp. 289–301, Aug. 2018. [CrossRef]
- M. Li, A. Schönberg, M. Schaefer, R. Schroeder, I. Nasidze, and M. Stoneking, “Detecting heteroplasmy from high-throughput sequencing of complete human mitochondrial DNA genomes,” Am. J. Hum. Genet., vol. 87, no. 2, pp. 237–249, Aug. 2010. [CrossRef]
- Nissanka and C. T. Moraes, “Mitochondrial DNA heteroplasmy in disease and targeted nuclease-based therapeutic approaches,” EMBO Rep., vol. 21, no. 3, Mar. 2020. [CrossRef]
- K. Kopinski et al., “Regulation of nuclear epigenome by mitochondrial DNA heteroplasmy,” Proc. Natl. Acad. Sci. U. S. A., vol. 116, no. 32, pp. 16028–16035, Aug. 2019. [CrossRef]
- J. A. Irwin et al., “Investigation of heteroplasmy in the human mitochondrial DNA control region: a synthesis of observations from more than 5000 global population samples,” J. Mol. Evol., vol. 68, no. 5, pp. 516–527, May 2009. [CrossRef]
- A. D. Jayaprakash et al., “Stable heteroplasmy at the single-cell level is facilitated by intercellular exchange of mtDNA,” Nucleic Acids Res., vol. 43, no. 4, p. 2177, Feb. 2015. [CrossRef]
- J. Naue et al., “Evidence for frequent and tissue-specific sequence heteroplasmy in human mitochondrial DNA,” Mitochondrion, vol. 20, no. 1, pp. 82–94, 2015. [CrossRef]
- Y. He et al., “Heteroplasmic mitochondrial DNA mutations in normal and tumour cells,” Nature, vol. 464, no. 7288, pp. 610–614, Mar. 2010. [CrossRef]
- C. Samuels et al., “Recurrent tissue-specific mtDNA mutations are common in humans,” PLoS Genet., vol. 9, no. 11, Nov. 2013. [CrossRef]
- H. Goto et al., “Dynamics of mitochondrial heteroplasmy in three families investigated via a repeatable re-sequencing study,” Genome Biol., vol. 12, no. 6, Jun. 2011. [CrossRef]
- L. A. Zinovkina, “Mechanisms of Mitochondrial DNA Repair in Mammals,” Biochemistry (Mosc)., vol. 83, no. 3, pp. 233–249, Mar. 2018. [CrossRef]
- Sharma and H. Sampath, “Mitochondrial DNA Integrity: Role in Health and Disease,” Cells, vol. 8, no. 2, Feb. 2019. [CrossRef]
- M. Ding et al., “Melatonin prevents Drp1-mediated mitochondrial fission in diabetic hearts through SIRT1-PGC1α pathway,” J. Pineal Res., vol. 65, no. 2, Sep. 2018. [CrossRef]
- C. Driver, K. D. S. Bamitale, A. Kazi, M. Olla, N. A. Nyane, and P. M. O. Owira, “Cardioprotective Effects of Metformin,” J. Cardiovasc. Pharmacol., vol. 72, no. 2, pp. 121–127, Aug. 2018. [CrossRef]
- G. B. Stefano and R. M. Kream, “Mitochondrial DNA heteroplasmy in human health and disease,” Biomed. Rep., vol. 4, no. 3, p. 259, Mar. 2016. [CrossRef]
- A. A. Elorza and J. P. Soffia, “mtDNA Heteroplasmy at the Core of Aging-Associated Heart Failure. An Integrative View of OXPHOS and Mitochondrial Life Cycle in Cardiac Mitochondrial Physiology,” Front. Cell Dev. Biol., vol. 9, Feb. 2021. [CrossRef]
- J. S. L. Berg Jeremy, Biochemistry, Fifth Edition. W.H. FREEMAN AND COMPANY, 2013.
- Kivrak, K. Yurt, A. Kaplan, I. Alkan, and G. Altun, “Effects of electromagnetic fields exposure on the antioxidant defense system,” J. Microsc. Ultrastruct., vol. 5, no. 4, p. 167, 2017. [CrossRef]
- N. Fuku et al., “The mitochondrial-derived peptide MOTS-c: a player in exceptional longevity?,” Aging Cell, vol. 14, no. 6, pp. 921–923, Dec. 2015. [CrossRef]
- H. Kobayashi and S. Imanaka, “Mitochondrial DNA Damage and Its Repair Mechanisms in Aging Oocytes,” Int. J. Mol. Sci., vol. 25, no. 23, Dec. 2024. [CrossRef]
- C. Tang, J. Cai, X. M. Yin, J. M. Weinberg, M. A. Venkatachalam, and Z. Dong, “Mitochondrial quality control in kidney injury and repair,” Nat. Rev. Nephrol., vol. 17, no. 5, pp. 299–318, May 2021. [CrossRef]
- A. Bohr and G. L. Dianov, “Oxidative DNA damage processing in nuclear and mitochondrial DNA,” Biochimie, vol. 81, no. 1–2, pp. 155–160, 1999. [CrossRef]
- N. B. Larsen, M. Rasmussen, and L. J. Rasmussen, “Nuclear and mitochondrial DNA repair: Similar pathways?,” Mitochondrion, vol. 5, no. 2, pp. 89–108, Apr. 2005. [CrossRef]
- A. Hiona and C. Leeuwenburgh, “The role of mitochondrial DNA mutations in aging and sarcopenia: Implications for the mitochondrial vicious cycle theory of aging,” Exp. Gerontol., vol. 43, no. 1, pp. 24–33, Jan. 2008. [CrossRef]
- Gredilla, I. Sánchez-Román, A. Gómez, M. López-Torres, and G. Barja, “Mitochondrial base excision repair positively correlates with longevity in the liver and heart of mammals,” Geroscience, vol. 42, no. 2, pp. 653–665, Apr. 2020. [CrossRef]
- Gredilla, L. Weissman, J. L. Yang, V. A. Bohr, and T. Stevnsner, “Mitochondrial base excision repair in mouse synaptosomes during normal aging and in a model of Alzheimer’s disease,” Neurobiol. Aging, vol. 33, no. 4, p. 694, Apr. 2010. [CrossRef]
- SenGupta et al., “Base excision repair causes age-dependent accumulation of single-stranded DNA breaks that contribute to Parkinson disease pathology,” Cell Rep., vol. 36, no. 10, Sep. 2021. [CrossRef]
- A. Chen et al., “Increased mitochondrial DNA damage and decreased base excision repair in the auditory cortex of D-galactose-induced aging rats,” Mol. Biol. Rep., vol. 38, no. 6, pp. 3635–3642, Aug. 2011. [CrossRef]
- Y. Zhong et al., “Mitochondrial transcription factor A overexpression and base excision repair deficiency in the inner ear of rats with D-galactose-induced aging,” FEBS J., vol. 278, no. 14, pp. 2500–2510, Jul. 2011. [CrossRef]
- S. K. Tadi, R. Sebastian, S. Dahal, R. K. Babu, B. Choudhary, and S. C. Raghavan, “Microhomology-mediated end joining is the principal mediator of double-strand break repair during mitochondrial DNA lesions,” Mol. Biol. Cell, vol. 27, no. 2, pp. 223–235, Jan. 2016. [CrossRef]
- K. Allkanjari and R. A. Baldock, “Beyond base excision repair: an evolving picture of mitochondrial DNA repair,” Biosci. Rep., vol. 41, no. 10, p. BSR20211320, Oct. 2021. [CrossRef]
- Di Carlo and C. Sorrentino, “Oxidative Stress and Age-Related Tumors,” Antioxidants (Basel)., vol. 13, no. 9, Sep. 2024. [CrossRef]
- P. Volobaev, S. S. Kunizheva, L. I. Uralsky, D. A. Kupriyanova, and E. I. Rogaev, “Quantifying human genome parameters in aging,” Vavilovskii Zhurnal Genet. Selektsii, vol. 27, no. 5, pp. 495–501, 2023. [CrossRef]
- L. Wu et al., “Increased p66Shc in the Inner Ear of D-Galactose-Induced Aging Mice with Accumulation of Mitochondrial DNA 3873-bp Deletion: p66Shc and mtDNA Damage in the Inner Ear during Aging,” PLoS One, vol. 7, no. 11, p. e50483, Nov. 2012. [CrossRef]
- C. A. Castellani, R. J. Longchamps, J. Sun, E. Guallar, and D. E. Arking, “Thinking outside the nucleus: Mitochondrial DNA copy number in health and disease,” Mitochondrion, vol. 53, pp. 214–223, Jul. 2020. [CrossRef]
- Zhang et al., “The distinct spatiotemporal distribution and effect of feed restriction on mtDNA copy number in broilers,” Scientific Reports 2020 10:1, vol. 10, no. 1, pp. 3240-, Feb. 2020. [CrossRef]
- B. Singh, T. R. Schoeb, P. Bajpai, A. Slominski, and K. K. Singh, “Reversing wrinkled skin and hair loss in mice by restoring mitochondrial function,” Cell Death & Disease 2018 9:7, vol. 9, no. 7, pp. 735-, Jul. 2018. [CrossRef]
- Z. Zhang et al., “Decrease of MtDNA copy number affects mitochondrial function and involves in the pathological consequences of ischaemic stroke,” J. Cell. Mol. Med., vol. 26, no. 15, pp. 4157–4168, Aug. 2022. [CrossRef]
- N. Sun, R. J. Youle, and T. Finkel, “The Mitochondrial Basis of Aging,” Mol. Cell, vol. 61, no. 5, pp. 654–666, Mar. 2016. [CrossRef]
- R. R. Thomas et al., “RhTFAM treatment stimulates mitochondrial oxidative metabolism and improves memory in aged mice,” Aging, vol. 4, no. 9, pp. 620–635, 2012. [CrossRef]
- D. T. Soltys et al., “Lower mitochondrial DNA content but not increased mutagenesis associates with decreased base excision repair activity in brains of AD subjects,” Neurobiol. Aging, vol. 73, pp. 161–170, Jan. 2019. [CrossRef]
- D. Edgar and A. Trifunovic, “The mtDNA mutator mouse: Dissecting mitochondrial involvement in aging,” Aging, vol. 1, no. 12, pp. 1028–1032, 2009. [CrossRef]
- N. Keshavan et al., “The natural history of infantile mitochondrial DNA depletion syndrome due to RRM2B deficiency,” Genetics in Medicine, vol. 22, no. 1, pp. 199–209, Jan. 2020. [CrossRef]
- C. Viscomi and M. Zeviani, “MtDNA-maintenance defects: syndromes and genes,” J. Inherit. Metab. Dis., vol. 40, no. 4, pp. 587–599, Jul. 2017. [CrossRef]
- Wang et al., “Ablation of Shank3 alleviates cardiac dysfunction in aging mice by promoting CaMKII activation and Parkin-mediated mitophagy,” Redox Biol., vol. 58, p. 102537, Dec. 2022. [CrossRef]
- R. Filograna, M. Mennuni, D. Alsina, and N. G. Larsson, “Mitochondrial DNA copy number in human disease: the more the better?,” FEBS Lett., vol. 595, no. 8, pp. 976–1002, Apr. 2021. [CrossRef]
- Mizuno et al., “Low mitochondrial DNA copy number in peripheral blood mononuclear cells is associated with future mortality risk: a long-term follow-up study from Japan,” J. Nutr. Health Aging, vol. 28, no. 1, p. 100013, Jan. 2024. [CrossRef]
- Sun and J. C. St John, “Modulation of mitochondrial DNA copy number in a model of glioblastoma induces changes to DNA methylation and gene expression of the nuclear genome in tumours,” Epigenetics Chromatin, vol. 11, no. 1, Sep. 2018. [CrossRef]
- K. Foote et al., “Restoring mitochondrial DNA copy number preserves mitochondrial function and delays vascular aging in mice,” Aging Cell, vol. 17, no. 4, Aug. 2018. [CrossRef]
- J. Ding et al., “Assessing Mitochondrial DNA Variation and Copy Number in Lymphocytes of ~2,000 Sardinians Using Tailored Sequencing Analysis Tools,” PLoS Genet., vol. 11, no. 7, Jul. 2015. [CrossRef]
- J. Knez et al., “Correlates of Peripheral Blood Mitochondrial DNA Content in a General Population,” Am. J. Epidemiol., vol. 183, no. 2, pp. 138–146, Jan. 2016. [CrossRef]
- R. Zhang, Y. Wang, K. Ye, M. Picard, and Z. Gu, “Independent impacts of aging on mitochondrial DNA quantity and quality in humans,” BMC Genomics 2017 18:1, vol. 18, no. 1, pp. 890-, Nov. 2017. [CrossRef]
- A. Herbst et al., “Skeletal muscle mitochondrial DNA copy number and mitochondrial DNA deletion mutation frequency as predictors of physical performance in older men and women,” Geroscience, vol. 43, no. 3, pp. 1253–1264, Jun. 2021. [CrossRef]
- X. Liu et al., “Association of mitochondrial DNA copy number with cardiometabolic diseases,” Cell Genomics, vol. 1, no. 1, Oct. 2021. [CrossRef]
- E. Coskun, M. F. Beal, and D. C. Wallace, “Alzheimer’s brains harbor somatic mtDNA control-region mutations that suppress mitochondrial transcription and replication,” Proc. Natl. Acad. Sci. U. S. A., vol. 101, no. 29, pp. 10726–10731, Jul. 2004. [CrossRef]
- B. Rodríguez-Santiago, J. Casademont, and V. Nunes, “Is mitochondrial DNA depletion involved in Alzheimer’s disease?,” Eur. J. Hum. Genet., vol. 9, no. 4, pp. 279–285, 2001. [CrossRef]
- C. Dölle et al., “Defective mitochondrial DNA homeostasis in the substantia nigra in Parkinson disease,” Nat. Commun., vol. 7, Nov. 2016. [CrossRef]
- A. Pyle, H. Anugrha, M. Kurzawa-Akanbi, A. Yarnall, D. Burn, and G. Hudson, “Reduced mitochondrial DNA copy number is a biomarker of Parkinson’s disease,” Neurobiol. Aging, vol. 38, pp. 216.e7-216.e10, Jul. 2015. [CrossRef]
- X. bin Wang, N. hua Cui, S. Zhang, Z. jin Liu, J. fen Ma, and L. Ming, “Leukocyte telomere length, mitochondrial DNA copy number, and coronary artery disease risk and severity: A two-stage case-control study of 3064 Chinese subjects,” Atherosclerosis, vol. 284, pp. 165–172, May 2019. [CrossRef]
- Zhang et al., “Association between mitochondrial DNA copy number and sudden cardiac death: findings from the Atherosclerosis Risk in Communities study (ARIC),” Eur. Heart J., vol. 38, no. 46, pp. 3443–3448, Dec. 2017. [CrossRef]
- F. N. Ashar et al., “Association of Mitochondrial DNA Copy Number With Cardiovascular Disease,” JAMA Cardiol., vol. 2, no. 11, pp. 1247–1255, Nov. 2017. [CrossRef]
- M. Ikeda et al., “Overexpression of TFAM or twinkle increases mtDNA copy number and facilitates cardioprotection associated with limited mitochondrial oxidative stress,” PLoS One, vol. 10, no. 3, Mar. 2015. [CrossRef]
- Filograna et al., “Modulation of mtDNA copy number ameliorates the pathological consequences of a heteroplasmic mtDNA mutation in the mouse,” Sci. Adv., vol. 5, no. 4, 2019. [CrossRef]
- K. Li et al., “Leukocyte telomere length and mitochondrial DNA copy number associate with endothelial function in aging-related cardiovascular disease,” Front. Cardiovasc. Med., vol. 10, 2023. [CrossRef]
- B. Pillai et al., “Sirt3 protects mitochondrial DNA damage and blocks the development of doxorubicin-induced cardiomyopathy in mice,” Am. J. Physiol. Heart Circ. Physiol., vol. 310, no. 8, pp. H962–H972, Apr. 2016. [CrossRef]
- J. Marín-García, “Mitochondrial DNA repair: a novel therapeutic target for heart failure,” Heart Fail. Rev., vol. 21, no. 5, pp. 475–487, Sep. 2016. [CrossRef]
- A. Hahn et al., “Misregulation of mitochondrial 6mA promotes the propagation of mutant mtDNA and causes aging in C. elegans,” Cell Metab., vol. 36, no. 12, pp. 2528-2541.e11, Dec. 2024. [CrossRef]
- Long et al., “Maintaining mitochondrial DNA copy number mitigates ROS-induced oocyte decline and female reproductive aging,” Commun. Biol., vol. 7, no. 1, Dec. 2024. [CrossRef]
- Sreedhar, L. Aguilera-Aguirre, and K. K. Singh, “Mitochondria in skin health, aging, and disease,” Cell Death Dis., vol. 11, no. 6, Jun. 2020. [CrossRef]
- Sherkhane, G. Chayanika, A. Sood, D. K. Khatri, and S. B. Singh, “Mitochondrial remodelling-a vicious cycle in diabetic complications,” Mol. Biol. Rep., vol. 48, no. 5, pp. 4721–4731, May 2021. [CrossRef]
- Sebastián et al., “Mfn2 deficiency links age-related sarcopenia and impaired autophagy to activation of an adaptive mitophagy pathway,” EMBO J., vol. 35, no. 15, pp. 1677–1693, Aug. 2016. [CrossRef]
- Iqbal, O. Ostojic, K. Singh, A. M. Joseph, and D. A. Hood, “Expression of mitochondrial fission and fusion regulatory proteins in skeletal muscle during chronic use and disuse,” Muscle Nerve, vol. 48, no. 6, pp. 963–970, 2013. [CrossRef]
- L. Zhao et al., “Evidence for association of mitochondrial metabolism alteration with lipid accumulation in aging rats,” Exp. Gerontol., vol. 56, pp. 3–12, 2014. [CrossRef]
- Tezze et al., “Age-Associated Loss of OPA1 in Muscle Impacts Muscle Mass, Metabolic Homeostasis, Systemic Inflammation, and Epithelial Senescence,” Cell Metab., vol. 25, no. 6, pp. 1374-1389.e6, Jun. 2017. [CrossRef]
- A. M. Joseph et al., “The impact of aging on mitochondrial function and biogenesis pathways in skeletal muscle of sedentary high- and low-functioning elderly individuals,” Aging Cell, vol. 11, no. 5, pp. 801–809, Oct. 2012. [CrossRef]
- J. Faitg, J. P. Leduc-Gaudet, O. Reynaud, G. Ferland, P. Gaudreau, and G. Gouspillou, “Effects of Aging and Caloric Restriction on Fiber Type Composition, Mitochondrial Morphology and Dynamics in Rat Oxidative and Glycolytic Muscles,” Front. Physiol., vol. 10, no. APR, 2019. [CrossRef]
- J. P. Leduc-Gaudet et al., “Mitochondrial morphology is altered in atrophied skeletal muscle of aged mice,” Oncotarget, vol. 6, no. 20, pp. 17923–17937, 2015. [CrossRef]
- Favaro et al., “DRP1-mediated mitochondrial shape controls calcium homeostasis and muscle mass,” Nature Communications 2019 10:1, vol. 10, no. 1, pp. 2576-, Jun. 2019. [CrossRef]
- M. Dulac et al., “Drp1 knockdown induces severe muscle atrophy and remodelling, mitochondrial dysfunction, autophagy impairment and denervation,” J. Physiol., vol. 598, no. 17, pp. 3691–3710, Sep. 2020. [CrossRef]
- M. Dulac et al., “Regulation of muscle and mitochondrial health by the mitochondrial fission protein Drp1 in aged mice,” J. Physiol., vol. 599, no. 17, pp. 4045–4063, Sep. 2021. [CrossRef]
- A. M. Joseph et al., “Dysregulation of mitochondrial quality control processes contribute to sarcopenia in a mouse model of premature aging,” PLoS One, vol. 8, no. 7, Jul. 2013. [CrossRef]
- L. García-Prat et al., “Autophagy maintains stemness by preventing senescence,” Nature, vol. 529, no. 7584, pp. 37–42, Jan. 2016. [CrossRef]
- C. A. McMullen, A. L. Ferry, J. L. Gamboa, F. H. Andrade, and E. E. Dupont-Versteegden, “Age-related changes of cell death pathways in rat extraocular muscle,” Exp. Gerontol., vol. 44, no. 6–7, pp. 420–425, Jun. 2009. [CrossRef]
- M. Gaugler, A. Brown, E. Merrell, M. DiSanto-Rose, J. A. Rathmacher, and T. H. Reynolds IV, “PKB signaling and atrogene expression in skeletal muscle of aged mice,” J. Appl. Physiol. (1985)., vol. 111, no. 1, pp. 192–199, Jul. 2011. [CrossRef]
- M. J. Drummond et al., “Downregulation of E3 ubiquitin ligases and mitophagy-related genes in skeletal muscle of physically inactive, frail older women: a cross-sectional comparison,” J. Gerontol. A Biol. Sci. Med. Sci., vol. 69, no. 8, pp. 1040–1048, 2014. [CrossRef]
- W. Russ, A. M. Wills, I. M. Boyd, and J. Krause, “Weakness, SR function and stress in gastrocnemius muscles of aged male rats,” Exp. Gerontol., vol. 50, no. 1, pp. 40–44, Feb. 2014. [CrossRef]
- P. Leduc-Gaudet, O. Reynaud, S. N. Hussain, and G. Gouspillou, “Parkin overexpression protects from ageing-related loss of muscle mass and strength,” J. Physiol., vol. 597, no. 7, pp. 1975–1991, Apr. 2019. [CrossRef]
- Gouspillou et al., “Increased sensitivity to mitochondrial permeability transition and myonuclear translocation of endonuclease G in atrophied muscle of physically active older humans,” FASEB J., vol. 28, no. 4, pp. 1621–1633, 2014. [CrossRef]
- A. R. Konopka, M. K. Suer, C. A. Wolff, and M. P. Harber, “Markers of human skeletal muscle mitochondrial biogenesis and quality control: effects of age and aerobic exercise training,” J. Gerontol. A Biol. Sci. Med. Sci., vol. 69, no. 4, pp. 371–378, Apr. 2014. [CrossRef]
- D. Mellem et al., “Fragmentation of the mitochondrial network in skin in vivo,” PLoS One, vol. 12, no. 6, Jun. 2017. [CrossRef]
- D. Gupta, S. Archoo, S. H. Naikoo, and S. T. Abdullah, “Rosmarinic Acid: A Naturally Occurring Plant Based Agent Prevents Impaired Mitochondrial Dynamics and Apoptosis in Ultraviolet-B-Irradiated Human Skin Cells,” Photochem. Photobiol., vol. 98, no. 4, pp. 925–934, Jul. 2021. [CrossRef]
- R. B. Hamanaka et al., “Mitochondrial reactive oxygen species promote epidermal differentiation and hair follicle development,” Sci. Signal., vol. 6, no. 261, Feb. 2013. [CrossRef]
- Miwa, S. Kashyap, E. Chini, and T. von Zglinicki, “Mitochondrial dysfunction in cell senescence and aging,” J. Clin. Invest., vol. 132, no. 13, Jul. 2022. [CrossRef]
- E. Oblong et al., “Metabolic dysfunction in human skin: Restoration of mitochondrial integrity and metabolic output by nicotinamide (niacinamide) in primary dermal fibroblasts from older aged donors,” Aging Cell, vol. 19, no. 10, p. e13248, Oct. 2020. [CrossRef]
- Rorteau et al., “Maintenance of Chronological Aging Features in Culture of Normal Human Dermal Fibroblasts from Old Donors,” Cells, vol. 11, no. 5, Mar. 2022. [CrossRef]
- A. Bowman and M. A. Birch-Machin, “Age-Dependent Decrease of Mitochondrial Complex II Activity in Human Skin Fibroblasts,” Journal of Investigative Dermatology, vol. 136, no. 5, pp. 912–919, 2016. [CrossRef]
- D. F. Dai, T. Chen, S. C. Johnson, H. Szeto, and P. S. Rabinovitch, “Cardiac aging: from molecular mechanisms to significance in human health and disease,” Antioxid. Redox Signal., vol. 16, no. 12, pp. 1492–1536, Jun. 2012. [CrossRef]
- Ljubicic, K. J. Menzies, and D. A. Hood, “Mitochondrial dysfunction is associated with a pro-apoptotic cellular environment in senescent cardiac muscle,” Mech. Ageing Dev., vol. 131, no. 2, pp. 79–88, Feb. 2010. [CrossRef]
- W. Dorn, “Parkin-dependent mitophagy in the heart,” J. Mol. Cell. Cardiol., vol. 95, pp. 42–49, Jun. 2016. [CrossRef]
- J. Gollmer, A. Zirlik, and H. Bugger, “Mitochondrial Mechanisms in Diabetic Cardiomyopathy,” Diabetes Metab. J., vol. 44, no. 1, pp. 33–53, Feb. 2020. [CrossRef]
- T. Breitzig, M. D. Alleyn, R. F. Lockey, and N. Kolliputi, “A mitochondrial delicacy: dynamin-related protein 1 and mitochondrial dynamics,” Am. J. Physiol. Cell Physiol., vol. 315, no. 1, pp. C80–C90, Jul. 2018. [CrossRef]
- Y. Y. Park, S. Lee, M. Karbowski, A. Neutzner, R. J. Youle, and H. Cho, “Loss of MARCH5 mitochondrial E3 ubiquitin ligase induces cellular senescence through dynamin-related protein 1 and mitofusin 1,” J. Cell Sci., vol. 123, no. Pt 4, pp. 619–626, Feb. 2010. [CrossRef]
- T. J. Cahill et al., “Resistance of Dynamin-related Protein 1 Oligomers to Disassembly Impairs Mitophagy, Resulting in Myocardial Inflammation and Heart Failure,” Journal of Biological Chemistry, vol. 290, no. 43, pp. 25907–25919, Oct. 2015. [CrossRef]
- T. T. Lee et al., “Loss of Fis1 impairs proteostasis during skeletal muscle aging in Drosophila,” Aging Cell, vol. 20, no. 6, p. e13379, Jun. 2021. [CrossRef]
- G. Rontoyanni, O. N. Lopez, G. T. Fankhauser, Z. F. Cheema, B. B. Rasmussen, and C. Porter, “Mitochondrial Bioenergetics in the Metabolic Myopathy Accompanying Peripheral Artery Disease,” Front. Physiol., vol. 8, no. MAR, Mar. 2017. [CrossRef]
- He, Y. Ma, R. Wang, J. Zhang, L. Jing, and P. A. Li, “Deletion of Mitochondrial Uncoupling Protein 2 Exacerbates Mitochondrial Damage in Mice Subjected to Cerebral Ischemia and Reperfusion Injury under both Normo- and Hyperglycemic Conditions,” Int. J. Biol. Sci., vol. 16, no. 15, pp. 2788–2802, 2020. [CrossRef]
- P. Semadhi, D. Mulyaty, E. Halimah, and J. Levita, “Healthy mitochondrial DNA in balanced mitochondrial dynamics: A potential marker for neuro-aging prediction (Review),” Biomed. Rep., vol. 19, no. 3, Sep. 2023. [CrossRef]
- E. Zukowski et al., “STAT3 modulates CD4+ T mitochondrial dynamics and function in aging,” Aging Cell, vol. 22, no. 11, Nov. 2023. [CrossRef]
- J. C. Campos et al., “Exercise preserves physical fitness during aging through AMPK and mitochondrial dynamics,” Proc. Natl. Acad. Sci. U. S. A., vol. 120, no. 2, Jan. 2023. [CrossRef]
- T. Yu et al., “Premature aging is associated with higher levels of 8-oxoguanine and increased DNA damage in the Polg mutator mouse,” Aging Cell, vol. 21, no. 9, Sep. 2022. [CrossRef]
- J. F. Halling et al., “PGC-1α regulates mitochondrial properties beyond biogenesis with aging and exercise training,” Am. J. Physiol. Endocrinol. Metab., vol. 317, no. 3, pp. E513–E525, Sep. 2019. [CrossRef]
- J. Wang et al., “Spermidine alleviates cardiac aging by improving mitochondrial biogenesis and function,” Aging, vol. 12, no. 1, pp. 650–671, Jan. 2020. [CrossRef]
- L. D. Popov, “Mitochondrial biogenesis: An update,” J. Cell. Mol. Med., vol. 24, no. 9, pp. 4892–4899, May 2020. [CrossRef]
- Li and Z. Cai, “SIRT3 regulates mitochondrial biogenesis in aging-related diseases,” J. Biomed. Res., vol. 37, no. 2, pp. 77–88, 2022. [CrossRef]
- M. H. Muhammad and M. M. Allam, “Resveratrol and/or exercise training counteract aging-associated decline of physical endurance in aged mice; targeting mitochondrial biogenesis and function,” Journal of Physiological Sciences, vol. 68, no. 5, pp. 681–688, Sep. 2018. [CrossRef]
- Y. Kim, M. Triolo, and D. A. Hood, “Impact of Aging and Exercise on Mitochondrial Quality Control in Skeletal Muscle,” Oxid. Med. Cell. Longev., vol. 2017, 2017. [CrossRef]
- A. Li, X. Hou, and S. Hao, “Mitochondrial biogenesis in neurodegeneration,” J. Neurosci. Res., vol. 95, no. 10, pp. 2025–2029, Oct. 2017. [CrossRef]
- C. Ploumi, I. Daskalaki, and N. Tavernarakis, “Mitochondrial biogenesis and clearance: a balancing act,” FEBS J., vol. 284, no. 2, pp. 183–195, Jan. 2017. [CrossRef]
- Tian, W. He, R. Yang, and Y. Liu, “Dl-3-n-butylphthalide protects the heart against ischemic injury and H9c2 cardiomyoblasts against oxidative stress: involvement of mitochondrial function and biogenesis,” J. Biomed. Sci., vol. 24, no. 1, 2017. [CrossRef]
- L. Tao et al., “Exercise Training Protects Against Acute Myocardial Infarction via Improving Myocardial Energy Metabolism and Mitochondrial Biogenesis,” Cell. Physiol. Biochem., vol. 37, no. 1, pp. 162–175, Aug. 2015. [CrossRef]
- L. Xu et al., “GLIS1 alleviates cell senescence and renal fibrosis through PGC1-α mediated mitochondrial quality control in kidney aging,” Free Radic. Biol. Med., vol. 209, no. Pt 1, pp. 171–184, Nov. 2023. [CrossRef]
- Wang et al., “Pentoxifylline Enhances Antioxidative Capability and Promotes Mitochondrial Biogenesis in D-Galactose-Induced Aging Mice by Increasing Nrf2 and PGC-1 α through the cAMP-CREB Pathway,” Oxid. Med. Cell. Longev., vol. 2021, 2021. [CrossRef]
- Liu, C. E. Chang, A. C. Wooldredge, B. Fong, B. K. Kennedy, and C. Zhou, “Tom70-based transcriptional regulation of mitochondrial biogenesis and aging,” Elife, vol. 11, Mar. 2022. [CrossRef]
- D’Aquila, D. Bellizzi, and G. Passarino, “Mitochondria in health, aging and diseases: the epigenetic perspective,” Biogerontology, vol. 16, no. 5, pp. 569–585, Oct. 2015. [CrossRef]
- M. Parodi-Rullán, X. R. Chapa-Dubocq, and S. Javadov, “Acetylation of Mitochondrial Proteins in the Heart: The Role of SIRT3,” Front. Physiol., vol. 9, no. AUG, Aug. 2018. [CrossRef]
- L. Liu, Y. Li, G. Chen, and Q. Chen, “Crosstalk between mitochondrial biogenesis and mitophagy to maintain mitochondrial homeostasis,” J. Biomed. Sci., vol. 30, no. 1, Dec. 2023. [CrossRef]
- K. Palikaras, E. Lionaki, and N. Tavernarakis, “Mechanisms of mitophagy in cellular homeostasis, physiology and pathology,” Nat. Cell Biol., vol. 20, no. 9, pp. 1013–1022, Sep. 2018. [CrossRef]
- M. Cui et al., “HKDC1, a target of TFEB, is essential to maintain both mitochondrial and lysosomal homeostasis, preventing cellular senescence,” Proc. Natl. Acad. Sci. U. S. A., vol. 121, no. 2, 2024. [CrossRef]
- M. Triolo, D. Bhattacharya, and D. A. Hood, “Denervation induces mitochondrial decline and exacerbates lysosome dysfunction in middle-aged mice,” Aging, vol. 14, no. 22, pp. 8900–8913, 2022. [CrossRef]
- N. M. Christensen, S. Ringholm, B. T. Buch, A. Gudiksen, J. F. Halling, and H. Pilegaard, “Muscle PGC-1α modulates hepatic mitophagy regulation during aging,” Exp. Gerontol., vol. 172, Feb. 2023. [CrossRef]
- E. Marzetti, R. Calvani, H. J. Coelho-Junior, and A. Picca, “Mitochondrial pathways and sarcopenia in the geroscience era,” Journal of Nutrition, Health and Aging, vol. 28, no. 12, Dec. 2024. [CrossRef]
- A. Rappe et al., “Longitudinal autophagy profiling of the mammalian brain reveals sustained mitophagy throughout healthy aging,” EMBO J., vol. 43, no. 23, pp. 6199–6231, Dec. 2024. [CrossRef]
- E. T. Schmid, J. H. Pyo, and D. W. Walker, “Neuronal induction of BNIP3-mediated mitophagy slows systemic aging in Drosophila,” Nat. Aging, vol. 2, no. 6, pp. 494–507, Jun. 2022. [CrossRef]
- Luo, R. Zhang, J. Krigman, A. McAdams, S. Ozgen, and N. Sun, “A Healthy Heart and a Healthy Brain: Looking at Mitophagy,” Front. Cell Dev. Biol., vol. 8, May 2020. [CrossRef]
- P. Leduc-Gaudet, S. N. A. Hussain, E. Barreiro, and G. Gouspillou, “Mitochondrial Dynamics and Mitophagy in Skeletal Muscle Health and Aging,” Int. J. Mol. Sci., vol. 22, no. 15, Aug. 2021. [CrossRef]
- P. Rai et al., “IRGM1 links mitochondrial quality control to autoimmunity,” Nature Immunology 2021 22:3, vol. 22, no. 3, pp. 312–321, Jan. 2021. [CrossRef]
- Hu et al., “Stabilization of HIF-1α alleviates osteoarthritis via enhancing mitophagy,” Cell Death & Disease 2020 11:6, vol. 11, no. 6, pp. 481-, Jun. 2020. [CrossRef]
- N. Oleinik et al., “Alterations of lipid-mediated mitophagy result in aging-dependent sensorimotor defects,” Aging Cell, vol. 22, no. 10, Oct. 2023. [CrossRef]
- Mito et al., “Mosaic dysfunction of mitophagy in mitochondrial muscle disease,” Cell Metab., vol. 34, no. 2, pp. 197-208.e5, Feb. 2022. [CrossRef]
- A. L. Hughes and D. E. Gottschling, “An early age increase in vacuolar pH limits mitochondrial function and lifespan in yeast,” Nature 2012 492:7428, vol. 492, no. 7428, pp. 261–265, Nov. 2012. [CrossRef]
- Shirakabe et al., “Drp1-Dependent Mitochondrial Autophagy Plays a Protective Role Against Pressure Overload-Induced Mitochondrial Dysfunction and Heart Failure,” Circulation, vol. 133, no. 13, pp. 1249–1263, Mar. 2016. [CrossRef]
- M. Schneider et al., “Age-Related Deterioration of Mitochondrial Function in the Intestine,” Oxid. Med. Cell. Longev., vol. 2020, p. 4898217, 2020. [CrossRef]
- M. Golpich, E. Amini, Z. Mohamed, R. Azman Ali, N. Mohamed Ibrahim, and A. Ahmadiani, “Mitochondrial Dysfunction and Biogenesis in Neurodegenerative diseases: Pathogenesis and Treatment,” CNS Neurosci. Ther., vol. 23, no. 1, pp. 5–22, Jan. 2017. [CrossRef]
- S. Rizza et al., “S-nitrosylation drives cell senescence and aging in mammals by controlling mitochondrial dynamics and mitophagy,” Proc. Natl. Acad. Sci. U. S. A., vol. 115, no. 15, pp. E3388–E3397, 2018. [CrossRef]
- S. Lautrup, D. A. Sinclair, M. P. Mattson, and E. F. Fang, “NAD+ in Brain Aging and Neurodegenerative Disorders,” Cell Metab., vol. 30, no. 4, pp. 630–655, Oct. 2019. [CrossRef]
- E. F. Fang et al., “Tomatidine enhances lifespan and healthspan in C. elegans through mitophagy induction via the SKN-1/Nrf2 pathway,” Scientific Reports 2017 7:1, vol. 7, no. 1, pp. 46208-, Apr. 2017. [CrossRef]
- D. Ryu et al., “Urolithin A induces mitophagy and prolongs lifespan in C. elegans and increases muscle function in rodents,” Nat. Med., vol. 22, no. 8, pp. 879–888, Aug. 2016. [CrossRef]
- E. M. Fivenson et al., “Mitophagy in neurodegeneration and aging,” Neurochem. Int., vol. 109, pp. 202–209, Oct. 2017. [CrossRef]
- M. Castellazzi et al., “Autophagy and mitophagy biomarkers are reduced in sera of patients with Alzheimer’s disease and mild cognitive impairment,” Scientific Reports 2019 9:1, vol. 9, no. 1, pp. 20009-, Dec. 2019. [CrossRef]
- P. Martín-Maestro, R. Gargini, G. Perry, J. Avila, and V. García-Escudero, “PARK2 enhancement is able to compensate mitophagy alterations found in sporadic Alzheimer’s disease,” Hum. Mol. Genet., vol. 25, no. 4, pp. 792–806, Feb. 2016. [CrossRef]
- Götz, F. Chen, J. Van Dorpe, and R. M. Nitsch, “Formation of neurofibrillary tangles in P301l tau transgenic mice induced by Abeta 42 fibrils,” Science, vol. 293, no. 5534, pp. 1491–1495, Aug. 2001. [CrossRef]
- L. Schulz et al., “A new link to mitochondrial impairment in tauopathies,” Mol. Neurobiol., vol. 46, no. 1, pp. 205–216, 2012. [CrossRef]
- R. K. Lane, T. Hilsabeck, and S. L. Rea, “The role of mitochondrial dysfunction in age-related diseases,” Biochim. Biophys. Acta Bioenerg., vol. 1847, no. 11, pp. 1387–1400, Nov. 2015. [CrossRef]
- Wang and R. L. Davis, “Early Mitochondrial Fragmentation and Dysfunction in a Drosophila Model for Alzheimer’s Disease,” Mol. Neurobiol., vol. 58, no. 1, pp. 143–155, Jan. 2021. [CrossRef]
- Manczak, M. J. Calkins, and P. H. Reddy, “Impaired mitochondrial dynamics and abnormal interaction of amyloid beta with mitochondrial protein Drp1 in neurons from patients with Alzheimer’s disease: implications for neuronal damage,” Hum. Mol. Genet., vol. 20, no. 13, pp. 2495–2509, Jul. 2011. [CrossRef]
- Manczak and P. H. Reddy, “Abnormal interaction between the mitochondrial fission protein Drp1 and hyperphosphorylated tau in Alzheimer’s disease neurons: implications for mitochondrial dysfunction and neuronal damage,” Hum. Mol. Genet., vol. 21, no. 11, pp. 2538–2547, Jun. 2012. [CrossRef]
- Zhao et al., “Treadmill Exercise Attenuates Aβ-Induced Mitochondrial Dysfunction and Enhances Mitophagy Activity in APP/PS1 Transgenic Mice,” Neurochem. Res., vol. 45, no. 5, pp. 1202–1214, May 2020. [CrossRef]
- R. J. Youle and D. P. Narendra, “Mechanisms of mitophagy,” Nat. Rev. Mol. Cell Biol., vol. 12, no. 1, pp. 9–14, Jan. 2011. [CrossRef]
- S. M. S. Yakhine-Diop et al., “Impaired Mitophagy and Protein Acetylation Levels in Fibroblasts from Parkinson’s Disease Patients,” Mol. Neurobiol., vol. 56, no. 4, pp. 2466–2481, Apr. 2019. [CrossRef]
- Cai et al., “Overexpression of PGC-1α influences the mitochondrial unfolded protein response (mtUPR) induced by MPP+ in human SH-SY5Y neuroblastoma cells,” Sci. Rep., vol. 10, no. 1, Dec. 2020. [CrossRef]
- M. W. Lin, C. C. Lin, Y. H. Chen, H. Bin Yang, and S. Y. Hung, “Celastrol Inhibits Dopaminergic Neuronal Death of Parkinson’s Disease through Activating Mitophagy,” Antioxidants (Basel)., vol. 9, no. 1, Jan. 2019. [CrossRef]
- M. D. Umare et al., “Interweaving of reactive oxygen species and major neurological and psychiatric disorders,” Ann. Pharm. Fr., vol. 80, no. 4, pp. 409–425, Jul. 2022. [CrossRef]
- A. H. V. Schapira and M. Gegg, “Mitochondrial contribution to Parkinson’s disease pathogenesis,” Parkinsons Dis., vol. 2011, 2011. [CrossRef]
- M. J. Quinn, P. I. Moreira, A. F. Ambrósio, and C. H. Alves, “PINK1/PARKIN signalling in neurodegeneration and neuroinflammation,” Acta Neuropathol. Commun., vol. 8, no. 1, p. 189, Nov. 2020. [CrossRef]
- L. Ma, J. Zhu, Q. Gao, M. J. Rebecchi, Q. Wang, and L. Liu, “Restoring Pharmacologic Preconditioning in the Aging Heart: Role of Mitophagy/Autophagy,” J. Gerontol. A Biol. Sci. Med. Sci., vol. 72, no. 4, pp. 489–498, 2017. [CrossRef]
- J. Ren et al., “Akt2 ablation prolongs life span and improves myocardial contractile function with adaptive cardiac remodeling: role of Sirt1-mediated autophagy regulation,” Aging Cell, vol. 16, no. 5, pp. 976–987, Oct. 2017. [CrossRef]
- A. Aliper et al., “Towards natural mimetics of metformin and rapamycin,” Aging, vol. 9, no. 11, pp. 2245–2268, 2017. [CrossRef]
- J. Wen et al., “Role of mitophagy in the hallmarks of aging,” J. Biomed. Res., vol. 37, no. 1, p. 1, 2022. [CrossRef]
- Y. Guo et al., “Sirt3-mediated mitophagy regulates AGEs-induced BMSCs senescence and senile osteoporosis,” Redox Biol., vol. 41, May 2021. [CrossRef]
- J. S. Harrington, S. W. Ryter, M. Plataki, D. R. Price, and A. M. K. Choi, “Mitochondria in health, disease, and aging,” Physiol. Rev., vol. 103, no. 4, pp. 2349–2422, 2023. [CrossRef]
- N. S. Swerdlow and H. M. Wilkins, “Mitophagy and the Brain,” Int. J. Mol. Sci., vol. 21, no. 24, Dec. 2020. [CrossRef]
- M. F. Alexeyev, S. P. LeDoux, and G. L. Wilson, “Mitochondrial DNA and aging,” Clin. Sci. (Lond)., vol. 107, no. 4, pp. 355–364, Oct. 2004. [CrossRef]
- O. S. Ademowo, O. Oyebode, R. Edward, M. E. Conway, H. R. Griffiths, and I. H. K. Dias, “Effects of carotenoids on mitochondrial dysfunction,” Biochem. Soc. Trans., vol. 52, no. 1, p. 65, Feb. 2024. [CrossRef]
- H. Alizadeh Pahlavani, I. Laher, B. Knechtle, and H. Zouhal, “Exercise and mitochondrial mechanisms in patients with sarcopenia,” Front. Physiol., vol. 13, Dec. 2022. [CrossRef]
- Fularski et al., “Statins in Chronic Kidney Disease-Effects on Atherosclerosis and Cellular Senescence,” Cells, vol. 12, no. 13, Jul. 2023. [CrossRef]
- M. Ge, F. Fontanesi, S. Merscher, and A. Fornoni, “The Vicious Cycle of Renal Lipotoxicity and Mitochondrial Dysfunction,” Front. Physiol., vol. 11, Jul. 2020. [CrossRef]
- A. Paoli and G. Cerullo, “Investigating the Link between Ketogenic Diet, NAFLD, Mitochondria, and Oxidative Stress: A Narrative Review,” Antioxidants (Basel)., vol. 12, no. 5, May 2023. [CrossRef]
- Poerwoatmodjo, G. J. Schenk, J. J. G. Geurts, and A. Luchicchi, “Cysteine Proteases and Mitochondrial Instability: A Possible Vicious Cycle in MS Myelin?,” Front. Cell. Neurosci., vol. 14, Dec. 2020. [CrossRef]
- D. Nguyen et al., “A new vicious cycle involving glutamate excitotoxicity, oxidative stress and mitochondrial dynamics,” Cell Death Dis., vol. 2, no. 12, Dec. 2011. [CrossRef]
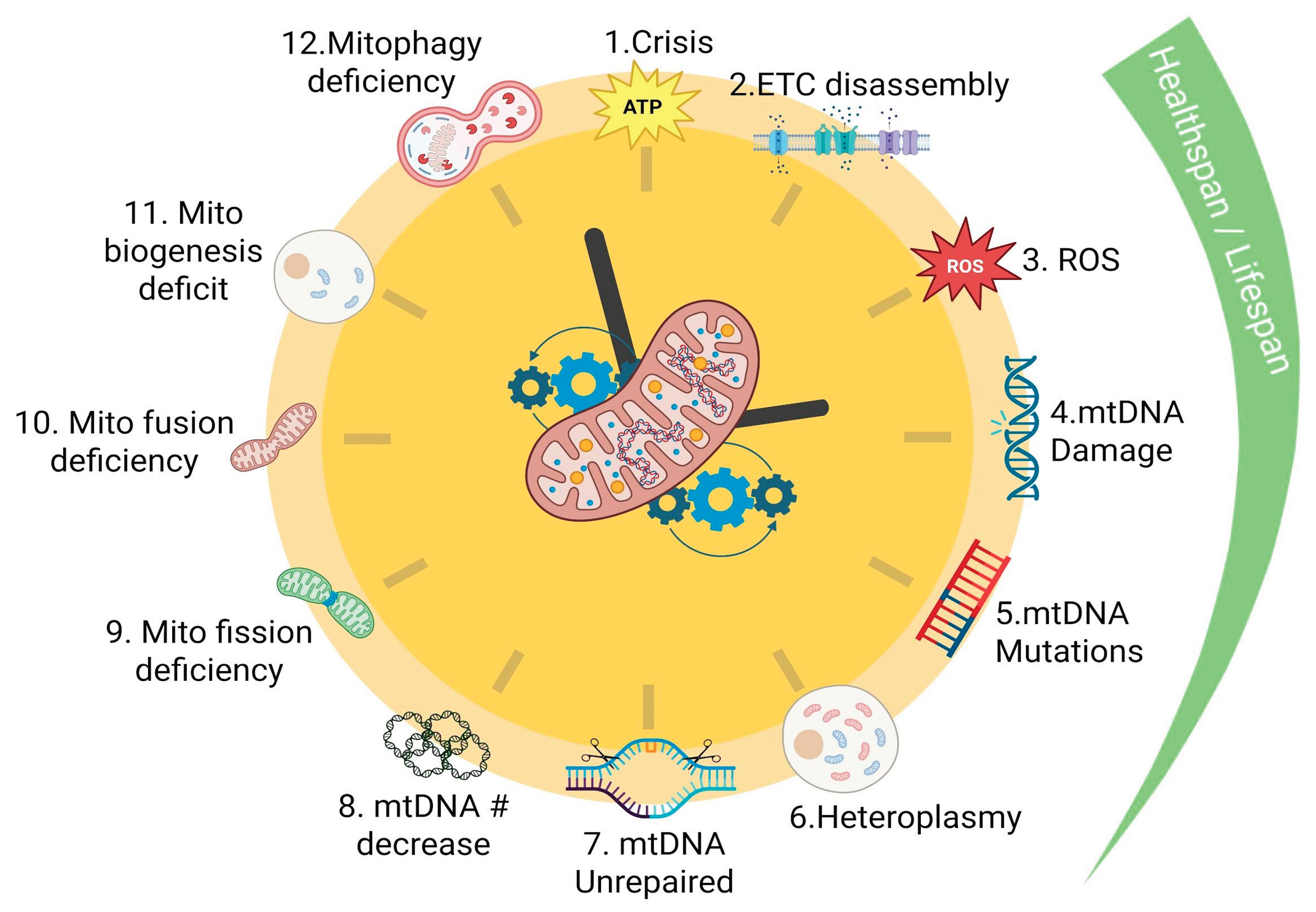
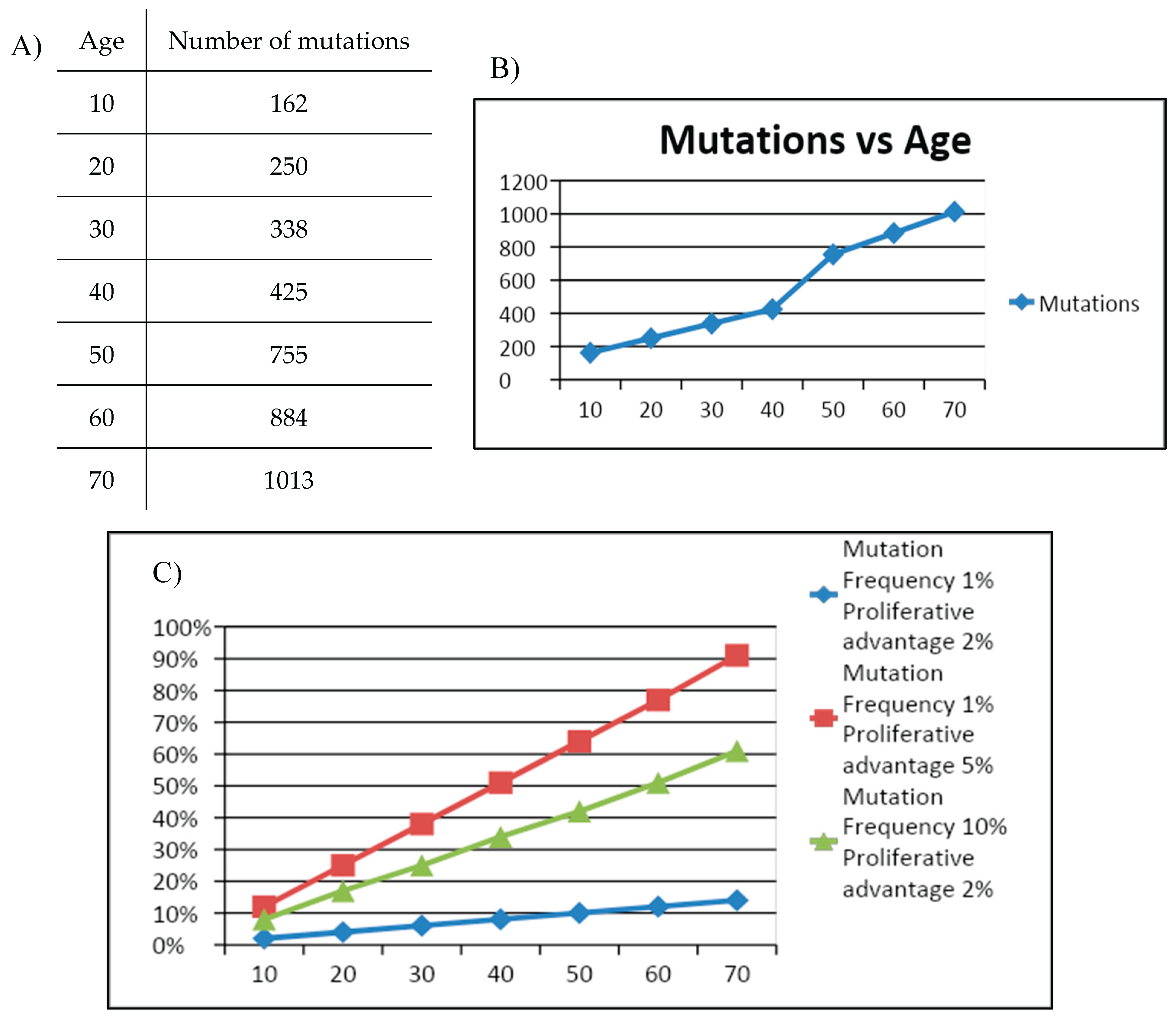
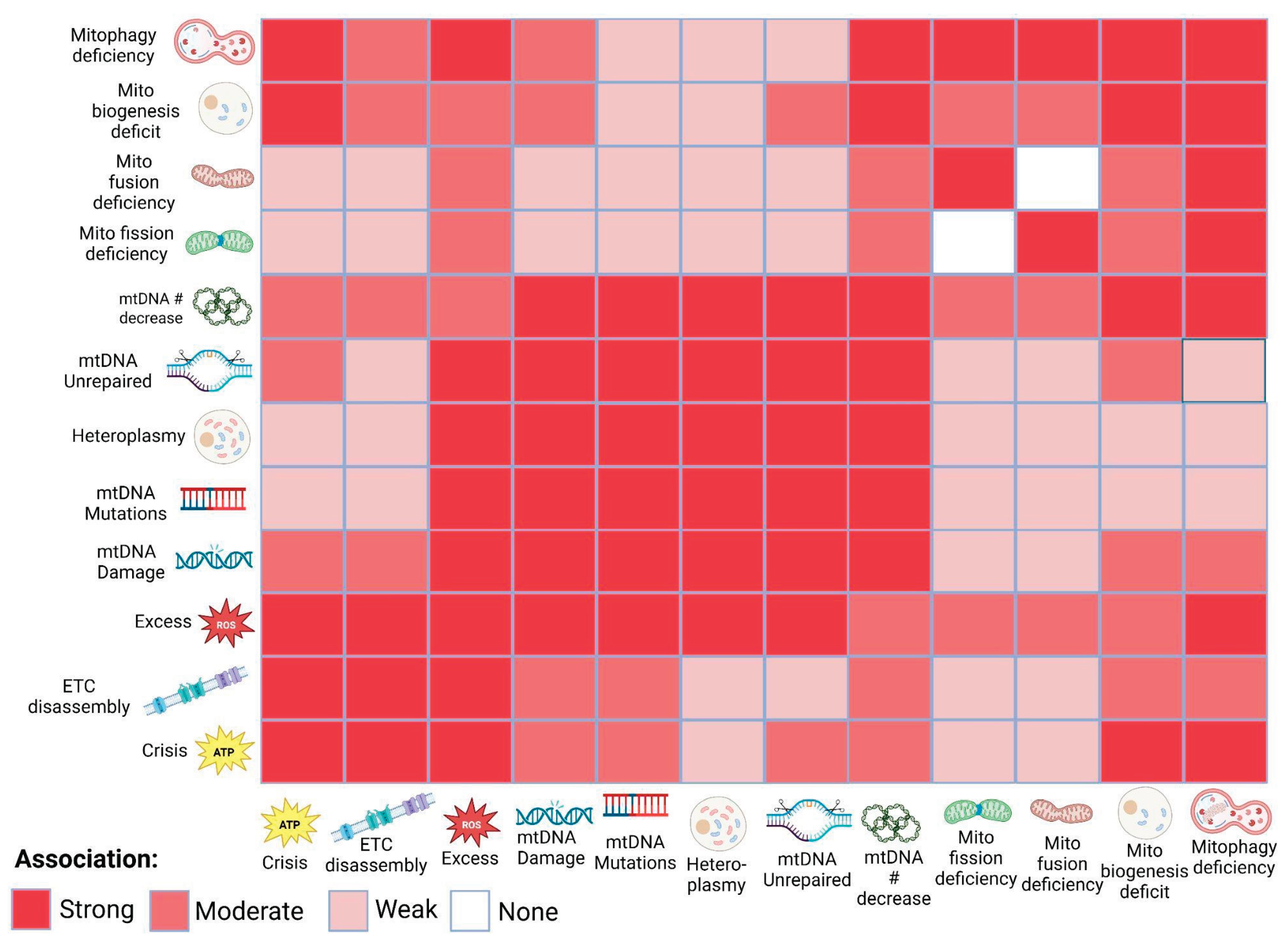
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



