Submitted:
29 January 2026
Posted:
30 January 2026
You are already at the latest version
Abstract
Keywords:
Introduction
1. Intestinal Disease Models
1.1. Dextran Sulfate Sodium (DSS)-Induced Models (Colitis)
1.2. Intestinal Viral Infections
1.3. Regulation by Commensal Microbiota–Derived Metabolites Under Homeostatic Conditions
1.4. Bacterial and Fungal Pathogens
2. Systemic Infection Models
2.1. Listeria Monocytogenes, Salmonella Typhimurium, and Escherichia coli (Systemic Infection Models)
2.2. Salmonella Typhimurium (Systemic Dissemination Following Intestinal Infection)
2.3. Listeria Monocytogenes and Staphylococcus aureus (In Vitro Macrophage Infection and Systemic Infection Models)
2.4. Sepsis Model
2.5. Sepsis Model (Klebsiella Pneumoniae–Induced)
2.6. Sepsis Model (Platelet-Specific Nlrp6 Deletion)
2.7. Sepsis Model (Effect of Gut Microbiota on Sepsis)
3. Lung Infection and Lung Cancer Models
3.1. Staphylococcus aureus Pneumonia
3.2. Streptococcus pneumoniae Infection
3.3. Birch Pollen Exposure and Helminth Infection Models
4. Hepatitis and Hepatocellular Carcinoma Models
4.1. Non-Alcoholic Steatohepatitis (NASH)
4.2. Alcoholic Hepatitis (AH) / Alcoholic Liver Disease (ALD)
4.3. Liver Injury Model After Allogeneic Hematopoietic Stem Cell Transplantation
4.4. Liver Cancer Models
4.4.1. Impact of Beneficial Gut Microbiota on Liver Cancer Progression
4.4.2. Chemically Induced and Transplantation HCC Models Using Conventional and Conditional Nlrp6 KO Mice
4.4.3. Subcutaneous HCC Transplantation Model with Intestinal Fungal Overgrowth
4.4.4. Colorectal Cancer Liver Metastasis Model
5. Gastric Cancer
5.1. Clinical Relevance and Functional Validation in Gastric Cancer Cell Lines
5.2. Regulation of NLRP6 Expression by Helicobacter Pylori
5.3. GRP78 as a Downstream Effector of NLRP6 Tumor-Suppressive Function
5.4. Epigenetic Suppression of NLRP6 by the OIP5-AS1/EZH2 Axis
6. Acute Injury Models
6.1. Acute Kidney Injury (AKI) Model
6.2. Acute Neural Injury (Sciatic Nerve Injury Model)
6.3. Intracerebral Hemorrhage (ICH) Model
6.4. Neural Stem Cells and Stress Resilience
6.5. miRNA-Mediated Regulation of NLRP6 in Astrocytes
6.6. Glioma and Glioblastoma
Concluding Remarks
Author Contributions
Funding
Conflicts of Interest
References
- Zheng, D.; Kern, L.; Elinav, E. The NLRP6 inflammasome. Immunology 2021, 162, 281–289. [Google Scholar] [CrossRef]
- Ghimire, L.; Paudel, S.; Jin, L.; Jeyaseelan, S. The NLRP6 inflammasome in health and disease. Mucosal Immunol 2020, 13, 388–398. [Google Scholar] [CrossRef]
- Sundaram, B.; Tweedell, R.E.; Prasanth Kumar, S.; Kanneganti, T.D. The NLR family of innate immune and cell death sensors. Immunity 2024, 57, 674–699. [Google Scholar] [CrossRef]
- Li, R.; Zan, Y.; Wang, D.; Chen, X.; Wang, A.; Tan, H.; Zhang, G.; Ding, S.; Shen, C.; Wu, H.; et al. A mouse model to distinguish NLRP6-mediated inflammasome-dependent and -independent functions. Proc Natl Acad Sci U S A 2024, 121, e2321419121. [Google Scholar] [CrossRef]
- Levy, M.; Thaiss, C.A.; Zeevi, D.; Dohnalova, L.; Zilberman-Schapira, G.; Mahdi, J.A.; David, E.; Savidor, A.; Korem, T.; Herzig, Y.; et al. Microbiota-Modulated Metabolites Shape the Intestinal Microenvironment by Regulating NLRP6 Inflammasome Signaling. Cell 2015, 163, 1428–1443. [Google Scholar] [CrossRef] [PubMed]
- Elinav, E.; Strowig, T.; Kau, A.L.; Henao-Mejia, J.; Thaiss, C.A.; Booth, C.J.; Peaper, D.R.; Bertin, J.; Eisenbarth, S.C.; Gordon, J.I.; et al. NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 2011, 145, 745–757. [Google Scholar] [CrossRef]
- Henao-Mejia, J.; Elinav, E.; Jin, C.; Hao, L.; Mehal, W.Z.; Strowig, T.; Thaiss, C.A.; Kau, A.L.; Eisenbarth, S.C.; Jurczak, M.J.; et al. Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 2012, 482, 179–185. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.Y.; Liu, M.; Wang, F.; Bertin, J.; Nunez, G. A functional role for Nlrp6 in intestinal inflammation and tumorigenesis. J Immunol 2011, 186, 7187–7194. [Google Scholar] [CrossRef] [PubMed]
- Normand, S.; Delanoye-Crespin, A.; Bressenot, A.; Huot, L.; Grandjean, T.; Peyrin-Biroulet, L.; Lemoine, Y.; Hot, D.; Chamaillard, M. Nod-like receptor pyrin domain-containing protein 6 (NLRP6) controls epithelial self-renewal and colorectal carcinogenesis upon injury. Proc Natl Acad Sci U S A 2011, 108, 9601–9606. [Google Scholar] [CrossRef]
- Wlodarska, M.; Thaiss, C.A.; Nowarski, R.; Henao-Mejia, J.; Zhang, J.P.; Brown, E.M.; Frankel, G.; Levy, M.; Katz, M.N.; Philbrick, W.M.; et al. NLRP6 inflammasome orchestrates the colonic host-microbial interface by regulating goblet cell mucus secretion. Cell 2014, 156, 1045–1059. [Google Scholar] [CrossRef]
- Ghimire, L.; Paudel, S.; Jin, L.; Baral, P.; Cai, S.; Jeyaseelan, S. NLRP6 negatively regulates pulmonary host defense in Gram-positive bacterial infection through modulating neutrophil recruitment and function. PLoS Pathog 2018, 14, e1007308. [Google Scholar] [CrossRef]
- Li, S.; Fu, Y.; Jia, X.; Liu, Z.; Qian, Z.; Zha, H.; Lei, G.; Yu, L.; Zhang, X.; Zhang, T.; et al. NLRP6 deficiency enhances macrophage-mediated phagocytosis via E-Syt1 to inhibit hepatocellular carcinoma progression. Gut 2025, 74, 1883–1895. [Google Scholar] [CrossRef]
- Valino-Rivas, L.; Cuarental, L.; Nunez, G.; Sanz, A.B.; Ortiz, A.; Sanchez-Nino, M.D. Loss of NLRP6 expression increases the severity of acute kidney injury. Nephrol Dial Transplant 2020, 35, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.F.; Li, Z.G.; Zhang, Y.; Ju, X.H.; Liu, X.W.; Zhou, A.M.; Chen, J. NLRP6 Inflammasome Ameliorates Brain Injury after Intracerebral Hemorrhage. Front Cell Neurosci 2017, 11, 206. [Google Scholar] [CrossRef]
- Anand, P.K.; Malireddi, R.K.; Lukens, J.R.; Vogel, P.; Bertin, J.; Lamkanfi, M.; Kanneganti, T.D. NLRP6 negatively regulates innate immunity and host defence against bacterial pathogens. Nature 2012, 488, 389–393. [Google Scholar] [CrossRef]
- Chenuet, P.; Marquant, Q.; Fauconnier, L.; Youness, A.; Mellier, M.; Marchiol, T.; Rouxel, N.; Messaoud-Nacer, Y.; Maillet, I.; Ledru, A.; et al. NLRP6 negatively regulates type 2 immune responses in mice. Allergy 2022, 77, 3320–3336. [Google Scholar] [CrossRef]
- Wang, P.; Zhu, S.; Yang, L.; Cui, S.; Pan, W.; Jackson, R.; Zheng, Y.; Rongvaux, A.; Sun, Q.; Yang, G.; et al. Nlrp6 regulates intestinal antiviral innate immunity. Science 2015, 350, 826–830. [Google Scholar] [CrossRef]
- Hara, H.; Seregin, S.S.; Yang, D.; Fukase, K.; Chamaillard, M.; Alnemri, E.S.; Inohara, N.; Chen, G.Y.; Nunez, G. The NLRP6 Inflammasome Recognizes Lipoteichoic Acid and Regulates Gram-Positive Pathogen Infection. Cell 2018, 175, 1651–1664 e1614. [Google Scholar] [CrossRef]
- Shen, C.; Li, R.; Negro, R.; Cheng, J.; Vora, S.M.; Fu, T.M.; Wang, A.; He, K.; Andreeva, L.; Gao, P.; et al. Phase separation drives RNA virus-induced activation of the NLRP6 inflammasome. Cell 2021, 184, 5759–5774 e5720. [Google Scholar] [CrossRef]
- Hu, B.; Elinav, E.; Huber, S.; Strowig, T.; Hao, L.; Hafemann, A.; Jin, C.; Wunderlich, C.; Wunderlich, T.; Eisenbarth, S.C.; et al. Microbiota-induced activation of epithelial IL-6 signaling links inflammasome-driven inflammation with transmissible cancer. Proc Natl Acad Sci U S A 2013, 110, 9862–9867. [Google Scholar] [CrossRef]
- Gremel, G.; Wanders, A.; Cedernaes, J.; Fagerberg, L.; Hallstrom, B.; Edlund, K.; Sjostedt, E.; Uhlen, M.; Ponten, F. The human gastrointestinal tract-specific transcriptome and proteome as defined by RNA sequencing and antibody-based profiling. J Gastroenterol 2015, 50, 46–57. [Google Scholar] [CrossRef]
- Lu, Y.; Chen, J.; Zheng, J.; Hu, G.; Wang, J.; Huang, C.; Lou, L.; Wang, X.; Zeng, Y. Mucosal adherent bacterial dysbiosis in patients with colorectal adenomas. Sci Rep 2016, 6, 26337. [Google Scholar] [CrossRef]
- Seregin, S.S.; Golovchenko, N.; Schaf, B.; Chen, J.; Eaton, K.A.; Chen, G.Y. NLRP6 function in inflammatory monocytes reduces susceptibility to chemically induced intestinal injury. Mucosal Immunol 2017, 10, 434–445. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, K.; Yoshimoto, T.; Tsutsui, H.; Okamura, H. Interleukin-18 regulates both Th1 and Th2 responses. Annu Rev Immunol 2001, 19, 423–474. [Google Scholar] [CrossRef]
- Dai, S.M.; Matsuno, H.; Nakamura, H.; Nishioka, K.; Yudoh, K. Interleukin-18 enhances monocyte tumor necrosis factor alpha and interleukin-1beta production induced by direct contact with T lymphocytes: implications in rheumatoid arthritis. Arthritis Rheum 2004, 50, 432–443. [Google Scholar] [CrossRef] [PubMed]
- Naito, Y.; Takagi, T.; Handa, O.; Ishikawa, T.; Nakagawa, S.; Yamaguchi, T.; Yoshida, N.; Minami, M.; Kita, M.; Imanishi, J.; et al. Enhanced intestinal inflammation induced by dextran sulfate sodium in tumor necrosis factor-alpha deficient mice. J Gastroenterol Hepatol 2003, 18, 560–569. [Google Scholar] [CrossRef]
- Noti, M.; Corazza, N.; Mueller, C.; Berger, B.; Brunner, T. TNF suppresses acute intestinal inflammation by inducing local glucocorticoid synthesis. J Exp Med 2010, 207, 1057–1066. [Google Scholar] [CrossRef]
- Schneider, K.M.; Mohs, A.; Gui, W.; Galvez, E.J.C.; Candels, L.S.; Hoenicke, L.; Muthukumarasamy, U.; Holland, C.H.; Elfers, C.; Kilic, K.; et al. Imbalanced gut microbiota fuels hepatocellular carcinoma development by shaping the hepatic inflammatory microenvironment. Nat Commun 2022, 13, 3964. [Google Scholar] [CrossRef]
- Wang, J.; Wang, L.; Lu, W.; Farhataziz, N.; Gonzalez, A.; Xing, J.; Zhang, Z. TRIM29 controls enteric RNA virus-induced intestinal inflammation by targeting NLRP6 and NLRP9b signaling pathways. Mucosal Immunol 2025, 18, 135–150. [Google Scholar] [CrossRef] [PubMed]
- Winsor, N.J.; Bayer, G.; Singh, O.; Chan, J.K.; Li, L.Y.; Lieng, B.Y.; Foerster, E.; Popovic, A.; Tsankov, B.K.; Maughan, H.; et al. Cross-kingdom-mediated detection of intestinal protozoa through NLRP6. Cell Host Microbe 2025, 33, 388–407 e389. [Google Scholar] [CrossRef]
- Mukherjee, S.; Kumar, R.; Tsakem Lenou, E.; Basrur, V.; Kontoyiannis, D.L.; Ioakeimidis, F.; Mosialos, G.; Theiss, A.L.; Flavell, R.A.; Venuprasad, K. Deubiquitination of NLRP6 inflammasome by Cyld critically regulates intestinal inflammation. Nat Immunol 2020, 21, 626–635. [Google Scholar] [CrossRef]
- Mao, X.; Qiu, X.; Jiao, C.; Lu, M.; Zhao, X.; Li, X.; Li, J.; Ma, J.; Zhang, H. Candida albicans SC5314 inhibits NLRP3/NLRP6 inflammasome expression and dampens human intestinal barrier activity in Caco-2 cell monolayer model. Cytokine 2020, 126, 154882. [Google Scholar] [CrossRef]
- Deng, Q.; Yang, S.; Huang, K.; Zhu, Y.; Sun, L.; Cao, Y.; Dong, K.; Li, Y.; Wu, S.; Huang, R. NLRP6 induces RIP1 kinase-dependent necroptosis via TAK1-mediated p38(MAPK)/MK2 phosphorylation in S. typhimurium infection. iScience 2024, 27, 109339. [Google Scholar] [CrossRef]
- Sun, L.; Huang, K.; Deng, Q.; Zhu, Y.; Cao, Y.; Dong, K.; Yang, S.; Li, Y.; Wu, S.; Huang, R. REV-ERBalpha negatively regulates NLRP6 transcription and reduces the severity of Salmonella infection in mice. Heliyon 2024, 10, e28432. [Google Scholar] [CrossRef]
- Pourcet, B.; Zecchin, M.; Ferri, L.; Beauchamp, J.; Sitaula, S.; Billon, C.; Delhaye, S.; Vanhoutte, J.; Mayeuf-Louchart, A.; Thorel, Q.; et al. Nuclear Receptor Subfamily 1 Group D Member 1 Regulates Circadian Activity of NLRP3 Inflammasome to Reduce the Severity of Fulminant Hepatitis in Mice. Gastroenterology 2018, 154, 1449–1464 e1420. [Google Scholar] [CrossRef]
- Lam, M.T.; Cho, H.; Lesch, H.P.; Gosselin, D.; Heinz, S.; Tanaka-Oishi, Y.; Benner, C.; Kaikkonen, M.U.; Kim, A.S.; Kosaka, M.; et al. Rev-Erbs repress macrophage gene expression by inhibiting enhancer-directed transcription. Nature 2013, 498, 511–515. [Google Scholar] [CrossRef]
- Ghimire, L.; Paudel, S.; Le, J.; Jin, L.; Cai, S.; Bhattarai, D.; Jeyaseelan, S. NLRP6 negatively regulates host defense against polymicrobial sepsis. Front Immunol 2024, 15, 1248907. [Google Scholar] [CrossRef] [PubMed]
- Radulovic, K.; Ayata, C.K.; Mak'Anyengo, R.; Lechner, K.; Wuggenig, P.; Kaya, B.; Hruz, P.; Gomez de Aguero, M.; Broz, P.; Weigmann, B.; et al. NLRP6 Deficiency in CD4 T Cells Decreases T Cell Survival Associated with Increased Cell Death. J Immunol 2019, 203, 544–556. [Google Scholar] [CrossRef]
- Cai, S.; Paudel, S.; Jin, L.; Ghimire, L.; Taylor, C.M.; Wakamatsu, N.; Bhattarai, D.; Jeyaseelan, S. NLRP6 modulates neutrophil homeostasis in bacterial pneumonia-derived sepsis. Mucosal Immunol 2021, 14, 574–584. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Chen, S.; Gui, X.; Li, Y.; Sun, Y.; Zhu, H.; Dai, Y.; Zhang, J.; Li, X.; Ju, W.; et al. Platelet NLRP6 protects against microvascular thrombosis in sepsis. Blood 2025, 146, 382–395. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Huang, Y.; Ye, Z.; Chen, W.; Zhang, N.; Wen, Z.; Ge, C. Short-chain fatty acids attenuate sepsis-induced gut dysbiosis and hippocampal neuroinflammation via NLRP6 inflammasome activation in mice. Int J Surg 2026, 112, 570–582. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Wu, X.; Peng, L.; Chen, T.; Huang, Q.; Wang, Y.; Ye, C.; Peng, Y.; Hu, D.; Fang, R. The Critical Role of NLRP6 Inflammasome in Streptococcus pneumoniae Infection In Vitro and In Vivo. Int J Mol Sci 2021, 22. [Google Scholar] [CrossRef] [PubMed]
- Forner, A.; Reig, M.; Bruix, J. Hepatocellular carcinoma. Lancet 2018, 391, 1301–1314. [Google Scholar] [CrossRef] [PubMed]
- Luedde, T.; Kaplowitz, N.; Schwabe, R.F. Cell death and cell death responses in liver disease: mechanisms and clinical relevance. Gastroenterology 2014, 147, 765–783 e764. [Google Scholar] [CrossRef]
- Macpherson, A.J.; Heikenwalder, M.; Ganal-Vonarburg, S.C. The Liver at the Nexus of Host-Microbial Interactions. Cell Host Microbe 2016, 20, 561–571. [Google Scholar] [CrossRef]
- Huang, C.; Liu, Q.; Tang, Q.; Jing, X.; Wu, T.; Zhang, J.; Zhang, G.; Zhou, J.; Zhang, Z.; Zhao, Y.; et al. Hepatocyte-specific deletion of Nlrp6 in mice exacerbates the development of non-alcoholic steatohepatitis. Free Radic Biol Med 2021, 169, 110–121. [Google Scholar] [CrossRef]
- Ji, X.; Li, L.; Lu, P.; Li, X.; Tian, D.; Liu, M. NLRP6 exerts a protective role via NF-kB with involvement of CCL20 in a mouse model of alcoholic hepatitis. Biochem Biophys Res Commun 2020, 528, 485–492. [Google Scholar] [CrossRef]
- Mainz, R.E.; Albers, S.; Haque, M.; Sonntag, R.; Treichel, N.S.; Clavel, T.; Latz, E.; Schneider, K.M.; Trautwein, C.; Otto, T. NLRP6 Inflammasome Modulates Disease Progression in a Chronic-Plus-Binge Mouse Model of Alcoholic Liver Disease. Cells 2022, 11. [Google Scholar] [CrossRef]
- Li, M.; Chen, Y.; Shi, J.; Ju, W.; Qi, K.; Fu, C.; Li, Z.; Zhang, X.; Qiao, J.; Xu, K.; et al. NLRP6 deficiency aggravates liver injury after allogeneic hematopoietic stem cell transplantation. Int Immunopharmacol 2019, 74, 105740. [Google Scholar] [CrossRef]
- Liu, Z.; Li, Y.; Li, C.; Lei, G.; Zhou, L.; Chen, X.; Jia, X.; Lu, Y. Intestinal Candida albicans Promotes Hepatocarcinogenesis by Up-Regulating NLRP6. Front Microbiol 2022, 13, 812771. [Google Scholar] [CrossRef]
- Chang, L.; Xu, L.; Tian, Y.; Liu, Z.; Song, M.; Li, S.; Zhang, X.; Chen, Y.; Hao, Q.; Lu, Y.; et al. NLRP6 deficiency suppresses colorectal cancer liver metastasis growth by modulating M-MDSC-induced immunosuppressive microenvironment. Biochim Biophys Acta Mol Basis Dis 2024, 1870, 167035. [Google Scholar] [CrossRef] [PubMed]
- Lucke, J.; Zhang, T.; Zazara, D.E.; Seeger, P.; Izbicki, J.R.; Hackert, T.; Huber, S.; Giannou, A.D. Protocol for generating lung and liver metastasis in mice using models that bypass intravasation. STAR Protoc 2024, 5, 102696. [Google Scholar] [CrossRef] [PubMed]
- Rao, X.; Zhou, X.; Wang, G.; Jie, X.; Xing, B.; Xu, Y.; Chen, Y.; Li, J.; Zhu, K.; Wu, Z.; et al. NLRP6 is required for cancer-derived exosome-modified macrophage M2 polarization and promotes metastasis in small cell lung cancer. Cell Death Dis 2022, 13, 891. [Google Scholar] [CrossRef]
- Kempster, S.L.; Belteki, G.; Forhead, A.J.; Fowden, A.L.; Catalano, R.D.; Lam, B.Y.; McFarlane, I.; Charnock-Jones, D.S.; Smith, G.C. Developmental control of the Nlrp6 inflammasome and a substrate, IL-18, in mammalian intestine. Am J Physiol Gastrointest Liver Physiol 2011, 300, G253–263. [Google Scholar] [CrossRef]
- Wang, H.; Xu, G.; Huang, Z.; Li, W.; Cai, H.; Zhang, Y.; Xiong, D.; Liu, G.; Wang, S.; Xue, Z.; et al. LRP6 targeting suppresses gastric tumorigenesis via P14(ARF)-Mdm2-P53-dependent cellular senescence. Oncotarget 2017, 8, 111597–111607. [Google Scholar] [CrossRef]
- Wang, Q.; Wang, C.; Chen, J. NLRP6, decreased in gastric cancer, suppresses tumorigenicity of gastric cancer cells. Cancer Manag Res 2018, 10, 6431–6444. [Google Scholar] [CrossRef] [PubMed]
- Tabassam, F.H.; Graham, D.Y.; Yamaoka, Y. Helicobacter pylori-associated regulation of forkhead transcription factors FoxO1/3a in human gastric cells. Helicobacter 2012, 17, 193–202. [Google Scholar] [CrossRef]
- Wang, X.; Wu, X.; Wang, Q.; Zhang, Y.; Wang, C.; Chen, J. NLRP6 suppresses gastric cancer growth via GRP78 ubiquitination. Exp Cell Res 2020, 395, 112177. [Google Scholar] [CrossRef]
- Lin, M.; Mo, Y.; Li, C.M.; Liu, Y.Z.; Feng, X.P. GRP78 as a potential therapeutic target in cancer treatment: an updated review of its role in chemoradiotherapy resistance of cancer cells. Med Oncol 2025, 42, 49. [Google Scholar] [CrossRef]
- Hernandez, I.; Cohen, M. Linking cell-surface GRP78 to cancer: From basic research to clinical value of GRP78 antibodies. Cancer Lett 2022, 524, 1–14. [Google Scholar] [CrossRef]
- Bai, Y.; Li, S. Long noncoding RNA OIP5-AS1 aggravates cell proliferation, migration in gastric cancer by epigenetically silencing NLRP6 expression via binding EZH2. J Cell Biochem 2020, 121, 353–362. [Google Scholar] [CrossRef]
- Ghafouri-Fard, S.; Dashti, S.; Farsi, M.; Hussen, B.M.; Taheri, M. A review on the role of oncogenic lncRNA OIP5-AS1 in human malignancies. Biomed Pharmacother 2021, 137, 111366. [Google Scholar] [CrossRef]
- Izquierdo, M.C.; Sanz, A.B.; Mezzano, S.; Blanco, J.; Carrasco, S.; Sanchez-Nino, M.D.; Benito-Martin, A.; Ruiz-Ortega, M.; Egido, J.; Ortiz, A. TWEAK (tumor necrosis factor-like weak inducer of apoptosis) activates CXCL16 expression during renal tubulointerstitial inflammation. Kidney Int 2012, 81, 1098–1107. [Google Scholar] [CrossRef]
- Ydens, E.; Demon, D.; Lornet, G.; De Winter, V.; Timmerman, V.; Lamkanfi, M.; Janssens, S. Nlrp6 promotes recovery after peripheral nerve injury independently of inflammasomes. J Neuroinflammation 2015, 12, 143. [Google Scholar] [CrossRef] [PubMed]
- Meng, C.; Zhang, J.; Zhang, L.; Wang, Y.; Li, Z.; Zhao, J. Effects of NLRP6 in Cerebral Ischemia/Reperfusion (I/R) Injury in Rats. J Mol Neurosci 2019, 69, 411–418. [Google Scholar] [CrossRef]
- Xiao, H.; Chen, H.; Jiang, R.; Zhang, L.; Wang, L.; Gan, H.; Jiang, N.; Zhao, J.; Zhai, X.; Liang, P. NLRP6 contributes to inflammation and brain injury following intracerebral haemorrhage by activating autophagy. J Mol Med (Berl) 2020, 98, 1319–1331. [Google Scholar] [CrossRef]
- Huang, X.; Tan, J.; Ji, Y.; Luo, J.; Zhao, Y.; Zhao, J. BRCC3 mediates inflammation and pyroptosis in cerebral ischemia/reperfusion injury by activating the NLRP6 inflammasome. CNS Neurosci Ther 2024, 30, e14697. [Google Scholar] [CrossRef]
- Tang, C.; Wang, Q.; Shen, J.; Wang, C.; Ding, H.; Wen, S.; Yang, F.; Jiao, R.; Wu, X.; Li, J.; et al. Neuron stem cell NLRP6 sustains hippocampal neurogenesis to resist stress-induced depression. Acta Pharm Sin B 2023, 13, 2017–2038. [Google Scholar] [CrossRef]
- Du, Y.; He, C.; An, Y.; Huang, Y.; Zhang, H.; Fu, W.; Wang, M.; Shan, Z.; Xie, J.; Yang, Y.; et al. The Role of Short Chain Fatty Acids in Inflammation and Body Health. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef] [PubMed]
- Oladapo, A.; Kannan, M.; Deshetty, U.M.; Singh, S.; Buch, S.; Periyasamy, P. Methamphetamine-mediated astrocytic pyroptosis and neuroinflammation involves miR-152-NLRP6 inflammasome signaling axis. Redox Biol 2025, 80, 103517. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Kannan, M.; Oladapo, A.; Deshetty, U.M.; Ray, S.; Buch, S.; Periyasamy, P. Ethanol modulates astrocyte activation and neuroinflammation via miR-339/NLRP6 inflammasome signaling. Free Radic Biol Med 2025, 226, 1–12. [Google Scholar] [CrossRef]
- Zhi, F.; Li, B.; Zhang, C.; Xia, F.; Wang, R.; Xie, W.; Cai, S.; Zhang, D.; Kong, R.; Hu, Y.; et al. NLRP6 potentiates PI3K/AKT signalling by promoting autophagic degradation of p85alpha to drive tumorigenesis. Nat Commun 2023, 14, 6069. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Cao, F.; Xiong, Y.; Zhou, H. SP1 transcriptionally activates NLRP6 inflammasome and induces immune evasion and radioresistance in glioma cells. Int Immunopharmacol 2021, 98, 107858. [Google Scholar] [CrossRef] [PubMed]
- Leng, F.; Yin, H.; Qin, S.; Zhang, K.; Guan, Y.; Fang, R.; Wang, H.; Li, G.; Jiang, Z.; Sun, F.; et al. NLRP6 self-assembles into a linear molecular platform following LPS binding and ATP stimulation. Sci Rep 2020, 10, 198. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Xie, P.; Wang, J.; Yang, F.; Li, S.; Jiang, H.; Wu, X.; Zhou, F.; Li, J. Nlrp6 protects from corticosterone-induced NSPC ferroptosis by modulating RIG-1/MAVS-mediated mitophagy. Redox Biol 2024, 73, 103196. [Google Scholar] [CrossRef]
- Wang, D.; Chen, X.; Sui, K.; Wang, A.; Li, R.; Zhou, W.; Zhu, X.; Hua, Y.; Yuan, S.; Zhou, R.; et al. UBE2O-mediated monoubiquitination licenses NLRP6 inflammasome activation in the intestine. Cell Host Microbe 2026, 34, 86–102 e108. [Google Scholar] [CrossRef]
- Premadasa, L.S.; McDew-White, M.; Romero, L.; Gondo, B.; Drawec, J.A.; Ling, B.; Okeoma, C.M.; Mohan, M. Epigenetic modulation of the NLRP6 inflammasome sensor as a therapeutic modality to reduce necroptosis-driven gastrointestinal mucosal dysfunction in HIV/SIV infection. Cell Commun Signal 2025, 23, 199. [Google Scholar] [CrossRef]
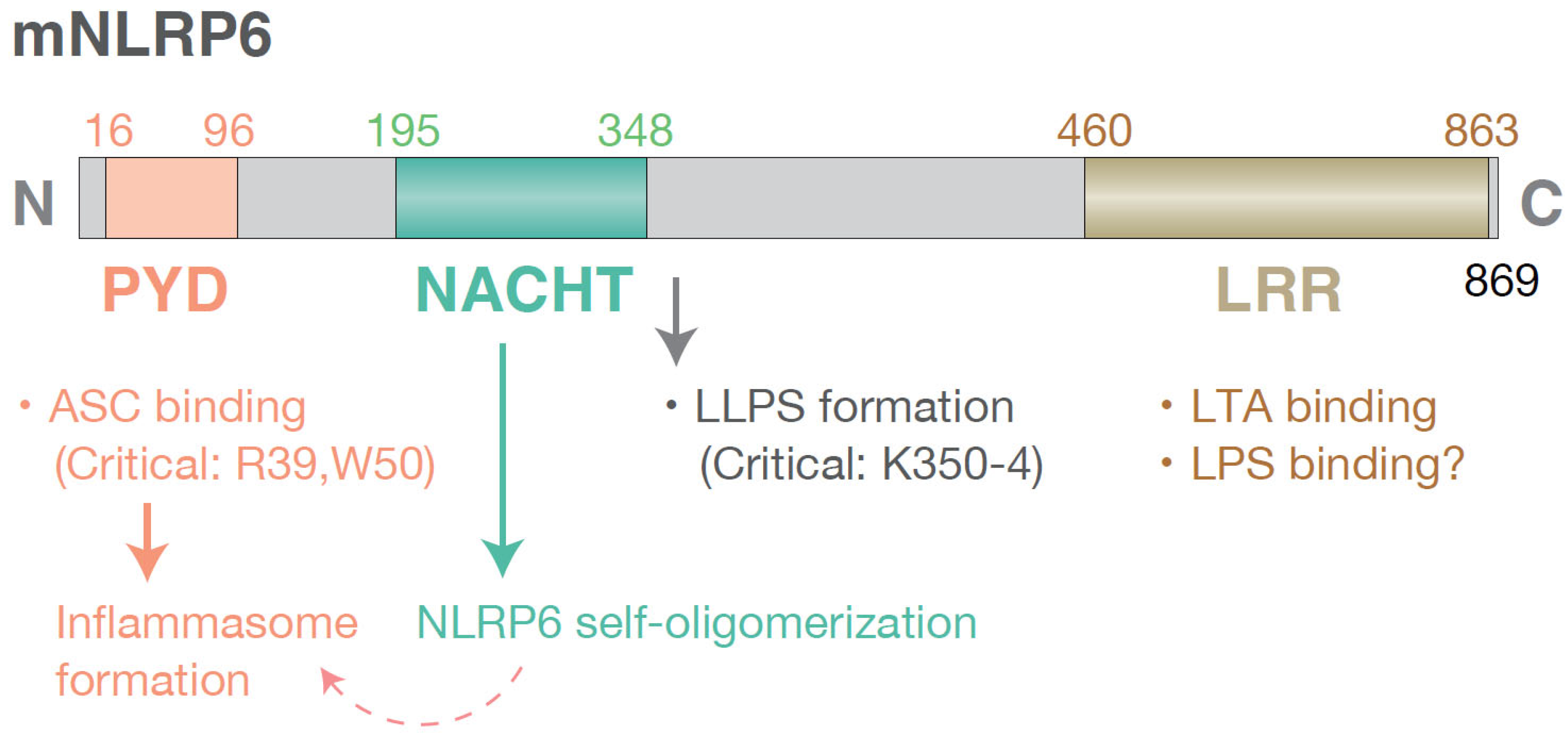
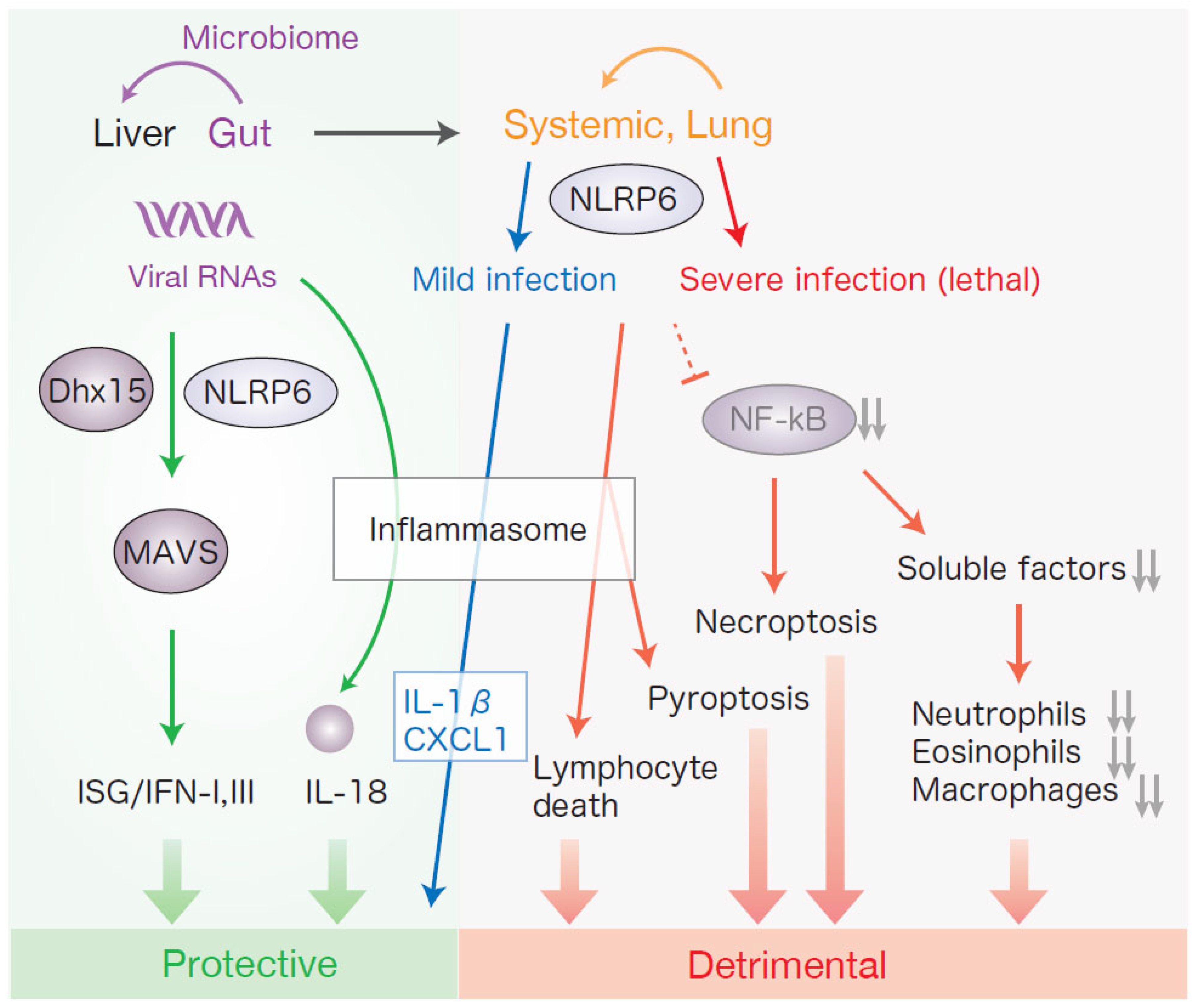
| Upstream regulators | Mechanisms | Locations |
| Taurine, Histamine, Spermine | Induction (taurine) or reduction (spermine and histamine) of NLRP6 inflammasome assembly | Colon [5] |
| Sphingolipids | Induction of NLRP6 inflammasome assembly | Colon [30] |
| ssRNA(EMCV) | MAVS-mediated IFN/ISG induction (together with Dhx15) | Intestinal tract [17] |
| dsRNA | Induction of LLPS of NLRP6 | MEF [19] |
| Lipoteichoic acid (LTA) | Recruitment of caspase-1 and caspase-11 to the NLRP6 inflammasome | BMDM [18] |
| α-hemolysin (hla) | Activation of the NLRP6 inflammasome | BMDM [11] |
| PPAR-γ | Transcriptional upregulation of NLRP6 | Human colon cancer cell line (Caco-2) [54] |
| Short-chain fatty acid (SCFA) | Transcriptional upregulation of Nlrp6 | Mouse neural stem cell line (NE-4C) [75] |
| SP1 | Transcriptional upregulation of NLRP6 | Human glioblastoma cell line (U87) [73] |
| REV-ERBα | Transcriptional repression of Nlrp6 | Intestinal epithelial cells [34] |
| Polyunsaturated fat (PUFA) diet, Methionine-choline deficient (MCD) diet | Transcriptional repression of Nlrp6 | Liver [46] |
| H. pylori/FOXO3 | Transcriptional repression of NLRP6 | Human gastric cancer cell lines (BGC-823, HGC-27) [56] |
| miR-152, miR-339 | Transcriptional repression of Nlrp6 (NLRP6) | Mouse primary astrocytes [56], Primary human astrocytes [71] |
| OIP5-AS1 (lncRNA)/ EZH2 | Increasing the H3K27me3 enrichment of NLRP6 promoter to represses NLRP6 expression | Human gastric cancer cell lines (AGS, SGC7901) [61] |
| Cyld (deubiquitinase) | Deubiquitination inhibited the NLRP6–ASC inflammasome complex | Colon [31] |
| BRCC3 (deubiquitinase) | Reduction of NLRP6 ubiquitination for NLRP6 stabilization | Mouse Hippocampal Nneuron Cell Line (HT22), HEK293T cells [67] |
| UBE2O* (ubiquitin conjugating enzyme) | NLRP6 monoubiquitination promotes oligomerization and cytoplasmic localization. | Intestinal epithelial cells [76] |
| TRIM29 (E3 ubiquitin ligase) | Promotes ubiquitination (degradation-inducing) of NLRP6 | Mouse intestinal organoids [29] |
| Exosomes from small cell lung cancer (SCLC) | Induction of NLRP6 expression | BMDM [53] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




