Submitted:
28 January 2026
Posted:
29 January 2026
You are already at the latest version
Abstract
Keywords:
Introduction
2.5. Bioinformatic Analysis
Materials and Methods
| Group | Cell Type | Treatment | Concentration |
| 1 | Human PBMC | Vehicle Control | — |
| 2 | Metadichol | 0.1pg/mL | |
| 3 | 1 pg/mL | ||
| 4 | 100 pg/mL | ||
| 5 | 1 ng/mL | ||
| 6 | 100 ng/mL |
| Test concentrations | ||||||
| RNA yield (ng/µl) | 0 | 0.1 pg/ ml | 1 pg/ ml | 100 pg/ ml | 1 ng/ ml | 100 ng/ ml |
| Human PBMC's | 425.2 | 410.4 | 380.9 | 412.8 | 438.6 | 446.2 |
| No | Gene | Primers | Amplicon size | Annealing temperature | |
| 1 | EIF5 | F | AATGGCTCCGTATCCAGCAGTG | 124 | 65 |
| R | GCTTCCTCAGTTGTATCTTCTCC | ||||
| 2 | PTPRO | F | GGTGTCTGTAGAGGATGTGACTG | 105 | 65 |
| R | AGTGATGAGGCGCTGTGGCTTT | ||||
| 3 | PTMS | F | AGAAACTGCCGAGGATGGAGAG | 131 | 67 |
| R | TGCCGTTTGGGATCCGCTTCAT | ||||
| 4 | B2M | F | CCACTGAAAAAGATGAGTATGCCT | 122 | 65 |
| R | CCAATCCAAATGCGGCATCTTCA | ||||
| 5 | HPRT1 | F | CATTATGCTGAGGATTTGGAAAGG | 128 | 65 |
| R | CTTGAGCACACAGAGGGCTACA | ||||
| 6 | RPLPO | F | TGGTCATCCAGCAGGTGTTCGA | 118 | 65 |
| R | ACAGACACTGGCAACATTGCGG | ||||
| 7 | HSPA9 | F | GCCTTGCTACGGCACATTGTGA | 131 | 65 |
| R | CTGCACAGATGAGGAGAGTTCAC | ||||
| 8 | PTMA | F | GGCTGACAATGAGGTAGACGAAG | 116 | 67 |
| R | GTAGCTGACTCAGCTTCCTCATC | ||||
| 9 | EIF1 | F | ACTGTCCAAGGGATCGCTGATG | 138 | 67 |
| R | TCTTGCGTTGGTCACCCTGTAG | ||||
| 10 | EIF4A2 | F | TACTGACTTGTTGGCTCGC | 109 | 62 |
| R | GACCCCCTCTGCCAATTCT | ||||
| 11 | EIF2S2 | F | GCAGGCTCAGAAAGAGACTACAC | 139 | 67 |
| R | GTTCCTACTCGGACGACTTGTG | ||||
| 12 | ACTB | F | CACCATTGGCAATGAGCGGTTC | 134 | 67 |
| R | AGGTCTTTGCGGATGTCCACGT | ||||
| 13 | TUBB | F | CTGGACCGCATCTCTGTGTACT | 116 | 67 |
| R | GCCAAAAGGACCTGAGCGAACA | ||||
| 14 | ATF4 | F | TTCTCCAGCGACAAGGCTAAGG | 116 | 67 |
| R | CTCCAACATCCAATCTGTCCCG | ||||
| 15 | HIF1A | F | TATGAGCCAGAAGAACTTTTAGGC | 144 | 67 |
| R | CACCTCTTTTGGCAAGCATCCTG | ||||
| 16 | FDFT1 | F | TGTGACCTCTGAACAGGAGTGG | 141 | 67 |
| R | GCCCATAGAGTTGGCACGTTCT | ||||
| 17 | CANX | F | GCTGGTTAGATGATGAGCCTGAG | 138 | 67 |
| R | ACACCACATCCAGGAGCTGACT | ||||
| 18 | HSPA5 | F | CTGTCCAGGCTGGTGTGCTCT | 142 | 67 |
| R | CTTGGTAGGCACCACTGTGTTC | ||||
| 19 | GAPDH | F | GTCTCCTCTGACTTCAACAGCG | 132 | 67 |
| R | ACCACCCTGTTGCTGTAGCCAA | ||||
| R | GGAATTGACAGTTGGGTCCAGG | ||||
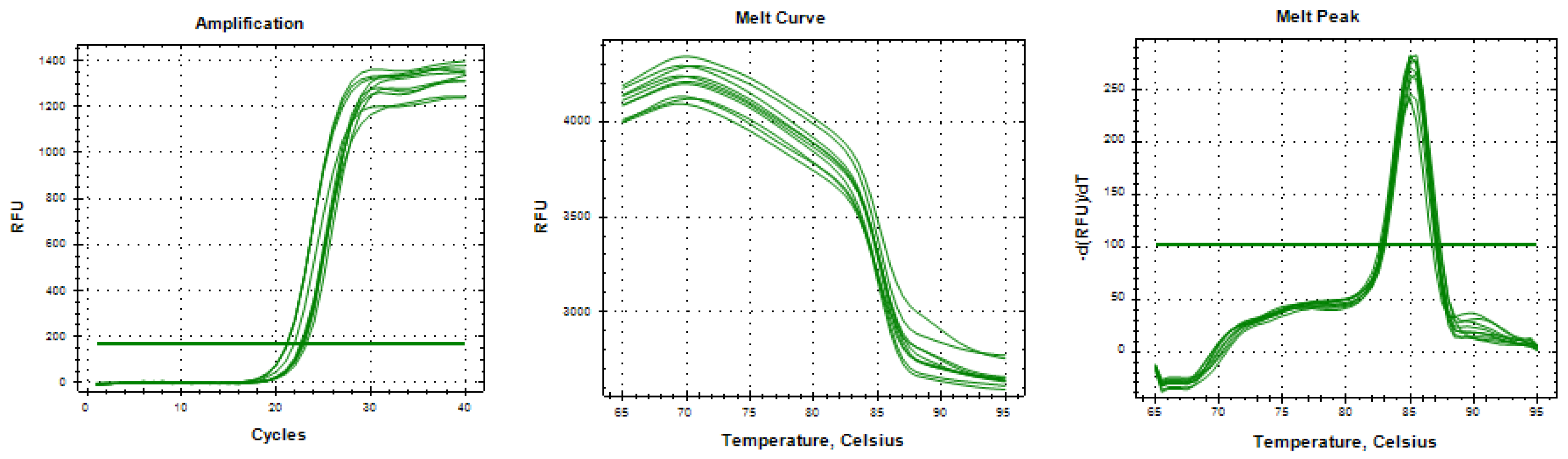
Results
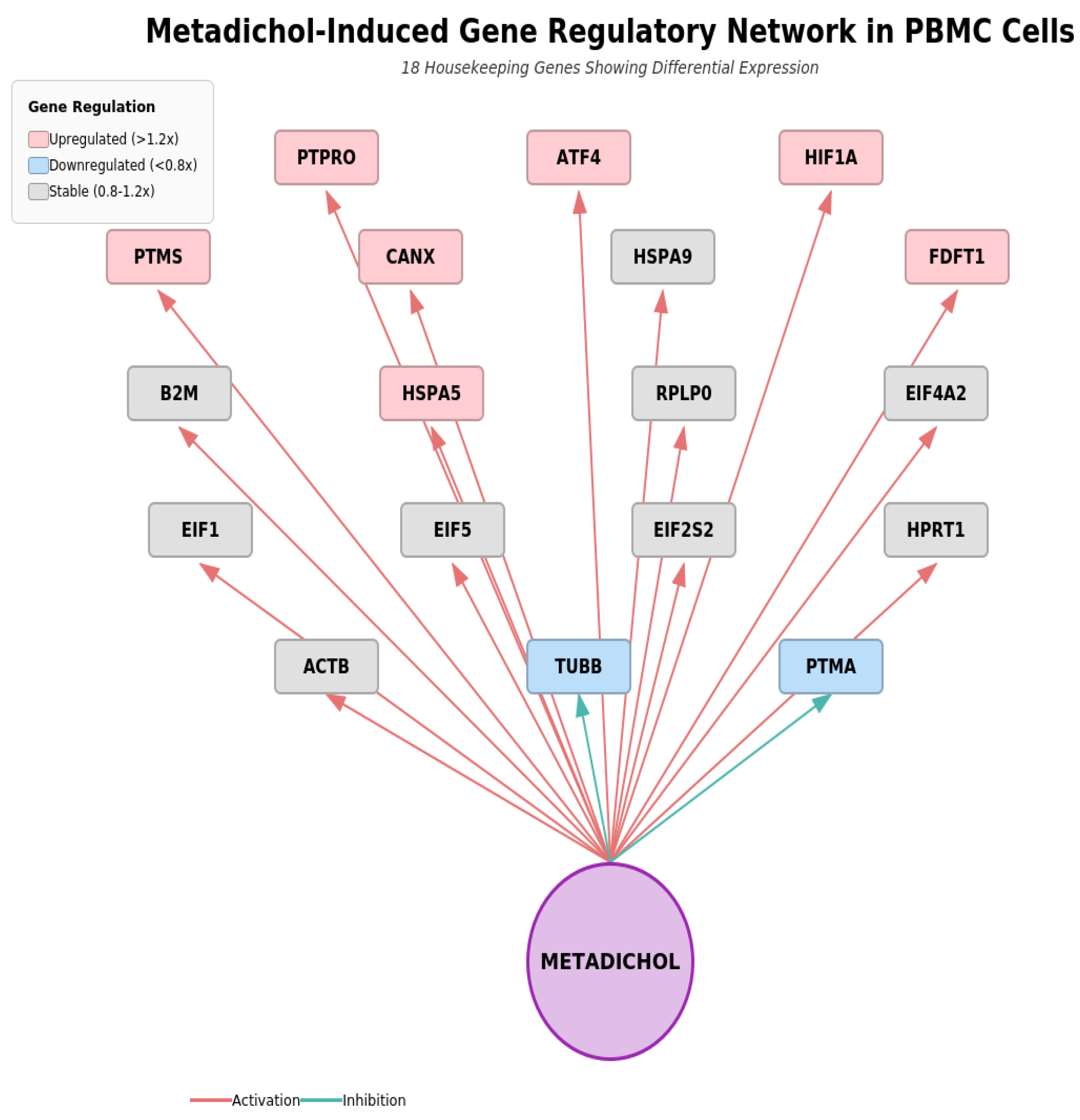
| Gene | Control | 0.1 pg/mL | 1 pg/mL | 100 pg/mL | 1 ng/mL | 100 ng/mL |
| ACTB | 1.00 | 1.09 | 1.04 | 0.94 | 1.00 | 0.98 |
| PTMA | 1.00 | 1.05 | 0.92 | 1.00 | 0.97 | 0.75* |
| ATF4 | 1.00 | 2.28** | 4.06* | 2.91** | 1.73** | 2.61* |
| PTPRO | 1.00 | 2.44* | 3.40* | 6.02*** | 0.47* | 2.98 |
| FDFT1 | 1.00 | 1.05 | 1.30 | 1.32 | 1.01 | 0.99 |
| PTMS | 1.00 | 1.52* | 1.16 | 1.91 | 0.87 | 0.60 |
| CANX | 1.00 | 1.54 | 1.08 | 1.56 | 1.08 | 0.96 |
| RPLP0 | 1.00 | 1.07 | 1.02 | 0.94 | 1.26 | 1.04 |
| B2M | 1.00 | 2.91 | 6.02 | 1.32 | 1.91 | 1.56 |
| HSPA9 | 1.00 | 1.23 | 1.31* | 1.00 | 0.96 | 0.83 |
| EIF4A2 | 1.00 | 1.19 | 0.95 | 0.97 | 0.84 | 1.26 |
| TUBB | 1.00 | 0.95 | 0.92 | 0.86 | 1.12 | 0.91 |
| HPRT1 | 1.00 | 0.87 | 1.13 | 0.86 | 1.03 | 1.20 |
| EIF1 | 1.00 | 1.02 | 0.93 | 1.12 | 1.01 | 0.85* |
| EIF5 | 1.00 | 0.89* | 1.08 | 0.95 | 1.17 | 1.02 |
| HIF1A | 1.00 | 1.61 | 0.45* | 1.41 | 0.72 | 1.01 |
| EIF2S2 | 1.00 | 1.03 | 1.04 | 0.89 | 1.18 | 1.08 |
| HSPA5 | 1.00 | 0.98 | 0.90 | 0.87 | 1.03 | 1.33* |
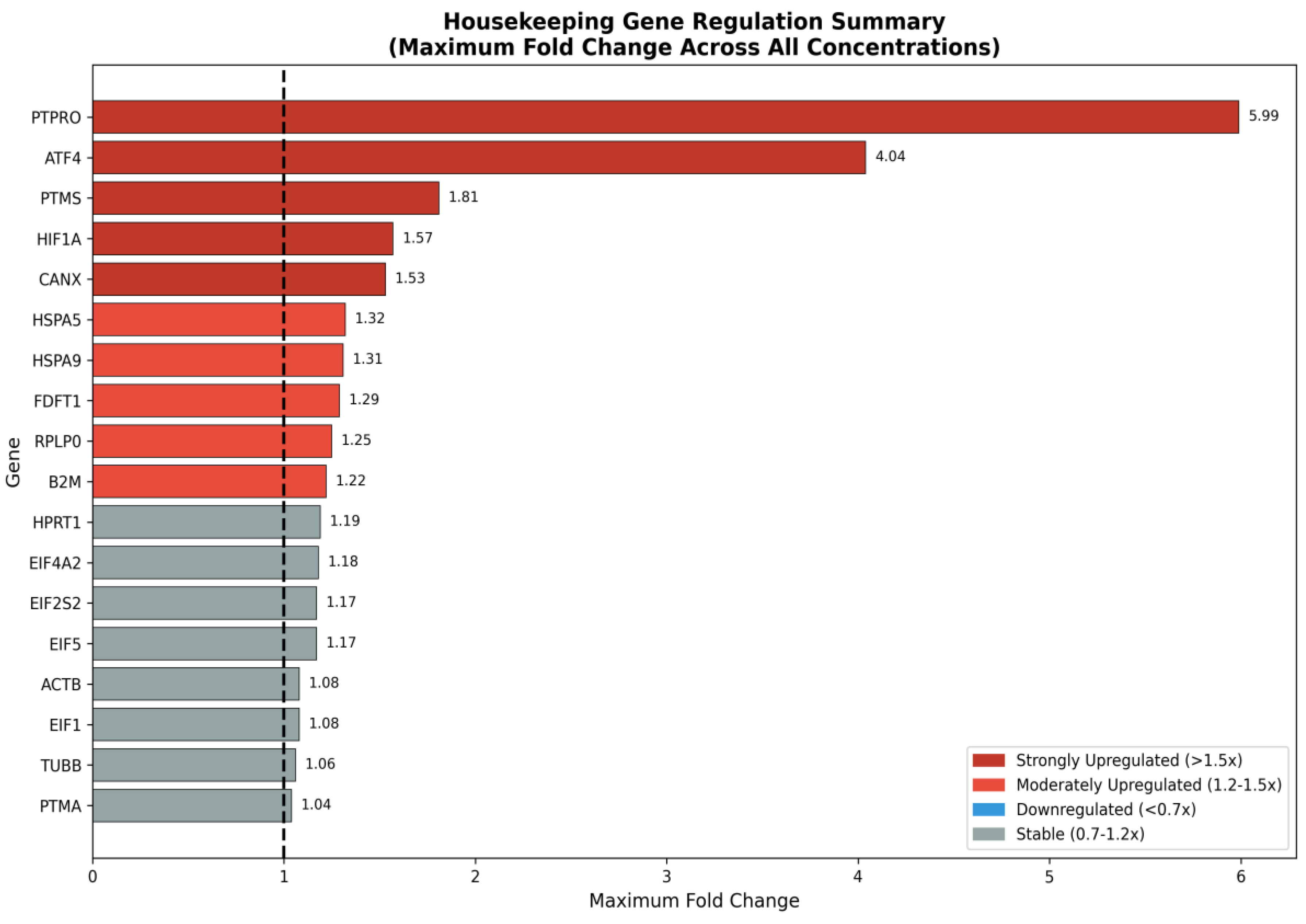
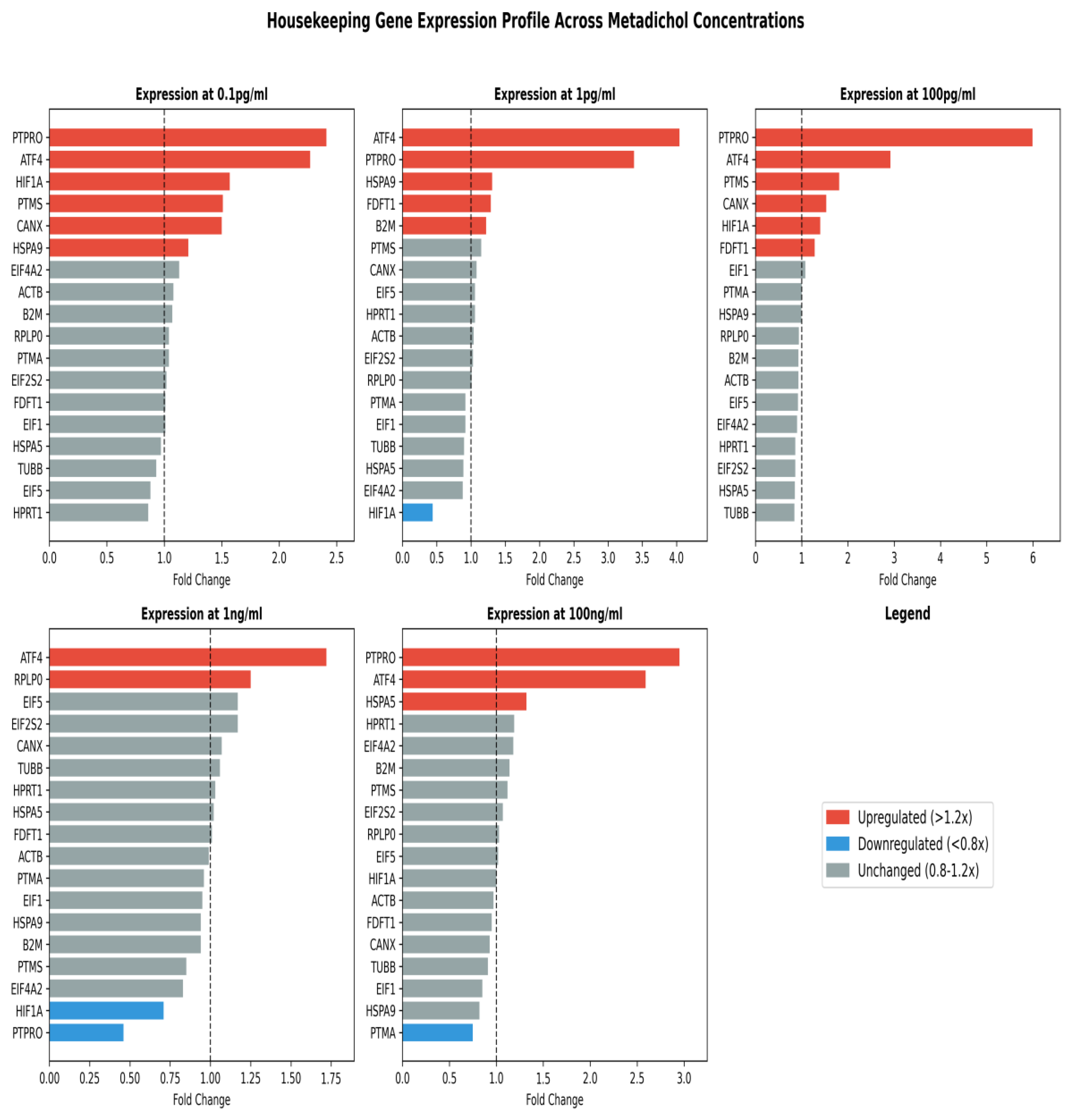
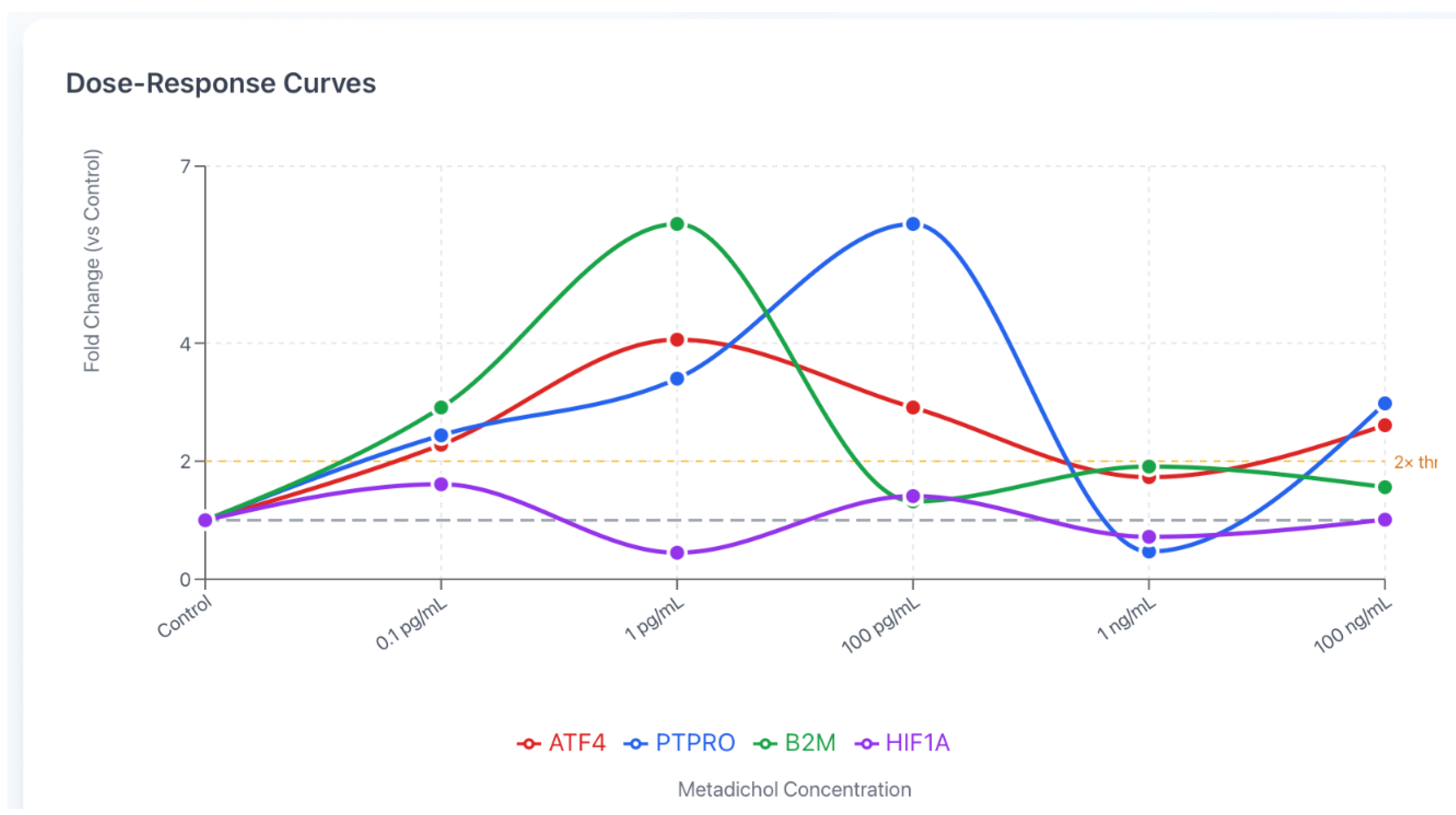
Discussion
Conclusions
Abbreviations
| ACTB | Actin Beta |
| ATF4 | Activating Transcription Factor 4 |
| ATF6 | Activating Transcription Factor 6 |
| B2M | Beta-2-Microglobulin |
| BCAT1 | Branched Chain Amino Acid Transaminase 1 |
| BiP | Binding Immunoglobulin Protein (HSPA5) |
| CANX | Calnexin |
| CLL | Chronic Lymphocytic Leukemia |
| Cq | Quantification Cycle |
| DDIT4 | DNA Damage Inducible Transcript 4 |
| EIF1 | Eukaryotic Translation Initiation Factor 1 |
| EIF2S2 | Eukaryotic Translation Initiation Factor 2 Subunit Beta |
| EIF4A2 | Eukaryotic Translation Initiation Factor 4A2 |
| EIF5 | Eukaryotic Translation Initiation Factor 5 |
| ER | Endoplasmic Reticulum |
| FBS | Fetal Bovine Serum |
| FDFT1 | Farnesyl-Diphosphate Farnesyltransferase 1 (Squalene Synthase) |
| FOXO | Forkhead Box O |
| GAPDH | Glyceraldehyde-3-Phosphate Dehydrogenase |
| GDF11 | Growth Differentiation Factor 11 |
| GRP78 | Glucose-Regulated Protein 78 (HSPA5) |
| HIF1A | Hypoxia Inducible Factor 1 Subunit Alpha |
| HKG | Housekeeping Gene |
| HPA | Human Protein Atlas |
| HPRT1 | Hypoxanthine Phosphoribosyl transferase 1 |
| HSPA5 | Heat Shock Protein Family A Member 5 (GRP78/BiP) |
| HSPA9 | Heat Shock Protein Family A Member 9 (Mortalin) |
| iPSC | Induced Pluripotent Stem Cell |
| IRE1 | Inositol-Requiring Enzyme 1 |
| ISR | Integrated Stress Response |
| KLF | Krüppel-Like Factor |
| LD50 | Median Lethal Dose |
| LXR | Liver X Receptor |
| MHC-I | Major Histocompatibility Complex Class I |
| mRNA | Messenger RNA |
| mTOR | Mechanistic Target of Rapamycin |
| mTORC1 | mTOR Complex 1 |
| NK | Natural Killer |
| NR | Nuclear Receptor |
| nTPM | Normalized Transcripts Per Million |
| PBMC | Peripheral Blood Mononuclear Cell |
| PBS | Phosphate-Buffered Saline |
| PERK | PKR-like ER Kinase |
| PGC1α | Peroxisome Proliferator-Activated Receptor Gamma Coactivator 1-Alpha |
| PPAR | Peroxisome Proliferator-Activated Receptor |
| PTEN | Phosphatase and Tensin Homolog |
| PTMA | Prothymosin Alpha |
| PTMS | Parathymosin |
| PTPRO | Protein Tyrosine Phosphatase Receptor Type O |
| qRT-PCR | Quantitative Real-Time Polymerase Chain Reaction |
| RNA | Ribonucleic Acid |
| RORC | RAR-Related Orphan Receptor C |
| RPLP0 | Ribosomal Protein Lateral Stalk Subunit P0 |
| RXR | Retinoid X Receptor |
| SD | Standard Deviation |
| SIRT | Sirtuin |
| SIRT1-7 | Sirtuin 1 through 7 |
| SOX | SRY-Box |
| TERT | Telomerase Reverse Transcriptase |
| TF | Transcription Factor |
| TLR | Toll-Like Receptor |
| TP53 | Tumor Protein P53 |
| TUBB | Tubulin Beta Class I |
| UPR | Unfolded Protein Response |
| VDR | Vitamin D Receptor |
| VEGF | Vascular Endothelial Growth Factor |
| VHL | Von Hippel-Lindau |
References
- Eisenberg E, Levanon EY. Human housekeeping genes, revisited. Trends Genet. 2013;29(10):569-574. [CrossRef]
- Hounkpe BW, Chebera F, De Paula EV, Gomes-Alves AG, Grunkraut J, Machado TFG. HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Res. 2021;49(D1):D947-D955. [CrossRef]
- Machado FB, Hounkpe BW, Gomes-Alves AG, et al. What are housekeeping genes? PLoS Comput Biol. 2022;18(7):e1010295. [CrossRef]
- Fendt SM, Bell EL, Keibler MA, et al. Is there a therapeutic window for metabolism-based cancer therapies? Front Endocrinol (Lausanne). 2017;8:150. [CrossRef]
- Zhu P, Tan MJ, Huang RL, et al. Metabolic alterations in cancer cells and therapeutic implications. Chin J Cancer. 2011;30(4):240-249. [CrossRef]
- Zhu W, Huang X, He Z, et al. Mechanisms of metabolic reprogramming in cancer cells supporting enhanced growth and proliferation. Cells. 2021;10(5):1056. [CrossRef]
- Zhe-Sheng C, Wei-Wei L, Jing-Jing W, et al. Definition, conservation and epigenetics of housekeeping and tissue-enriched genes. BMC Genomics. 2009;10:269. [CrossRef]
- Olsvik PA, Lie KK, Jordal AE, et al. Toxicological housekeeping genes: do they really keep the house? Environ Sci Technol. 2007;41(9):3338-3345. [CrossRef]
- Derks MFL, Megens HJ, Bosse M, et al. Loss of function mutations in essential genes cause embryonic lethality in pigs. PLoS Genet. 2019;15(3):e1008055. [CrossRef]
- Lv W, Zheng J, Luan M, et al. Comparing the evolutionary conservation between human essential genes, human orthologs of mouse essential genes and human housekeeping genes. Brief Bioinform. 2015;16(6):922-931. [CrossRef]
- Kshirsagar S, Islam MA, Reddy AP, Reddy PH. Resolving the current controversy of use and reuse of housekeeping proteins in ageing research. Ageing Res Rev. 2024;100:102437. [CrossRef]
- Vandesompele J, De Preter K, Pattyn F, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3(7):RESEARCH0034. [CrossRef]
- Faubert B, Solmonson A, DeBerardinis RJ. Metabolic reprogramming and cancer progression. Science. 2020;368(6487):eaaw5473. [CrossRef]
- Pavlova NN, Thompson CB. The emerging hallmarks of cancer metabolism. Cell Metab. 2016;23(1):27-47. [CrossRef]
- Graham NA, Minasyan A, Lomber A, et al. The landscape of metabolic pathway dependencies in cancer cell lines. PLoS Comput Biol. 2021;17(4):e1008942. [CrossRef]
- Warburg O, Wind F, Negelein E. The metabolism of tumors in the body. J Gen Physiol. 1927;8(6):519-530. [CrossRef]
- Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41(3):211-218. [CrossRef]
- Vaupel P, Multhoff G. Revisiting the Warburg effect: historical dogma versus current understanding. J Physiol. 2021;599(6):1745-1757. [CrossRef]
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646-674. [CrossRef]
- Hanahan D. Hallmarks of cancer: new dimensions. Cancer Discov. 2022;12(1):31-46. [CrossRef]
- Ko YH, Pedersen PL, Geschwind JF. Glucose catabolism in the rabbit VX2 tumor model for liver cancer: characterization and targeting hexokinase. Cancer Lett. 2001;173(1):83-91. [CrossRef]
- Ko YH, Smith BL, Wang Y, et al. Advanced cancers: eradication in all cases using 3-bromopyruvate therapy to deplete ATP. Biochem Biophys Res Commun. 2004;324(1):269-275. [CrossRef]
- Fan T, Sun G, Sun X, et al. Tumor energy metabolism and potential of 3-bromopyruvate as an inhibitor of aerobic glycolysis: implications in tumor treatment. Cancers (Basel). 2019;11(3):317. [CrossRef]
- Hudes G, Carducci M, Tomczak P, et al. Temsirolimus, interferon alfa, or both for advanced renal-cell carcinoma. N Engl J Med. 2007;356(22):2271-2281. [CrossRef]
- Motzer RJ, Escudier B, Oudard S, et al. Efficacy of everolimus in advanced renal cell carcinoma: a double-blind, randomised, placebo-controlled phase III trial. Lancet. 2008;372(9637):449-456. [CrossRef]
- Chiarini F, Evangelisti C, McCubrey JA, Martelli AM. Current treatment strategies for inhibiting mTOR in cancer. Trends Pharmacol Sci. 2015;36(2):124-135. [CrossRef]
- Raghavan PR. US Patent 8,722,093. Policosanol nanoparticles. 2014.
- Raghavan PR. US Patent 9,006,292. Policosanol nanoparticles. 2015.
- Raghavan PR. US Patent 9,034,383. Policosanol nanoparticles. 2015.
- Raghavan PR. Beyond Rapamycin: Metadichol Represents a New Class of Multi-Target mTOR Modulators. Medical Research Archives. 2025;13(9). [DOI not available].
- Raghavan PR. Metadichol®-induced expression of Sirtuins 1-7 in somatic and cancer cells. Medical Research Archives. 2024;12(6). [CrossRef]
- Raghavan PR. Metadichol: An Agonist that Expresses the Anti-Aging Gene Klotho in Various Cell Lines. Fortune Journal of Health Sciences. 2023;6:357-362. [CrossRef]
- Raghavan PR. Metadichol Orchestrates Cellular Reprogramming and Regenerative Pathways via FOX Transcription Factor Networks: Implications for Immune-Metabolic Rejuvenation. Medical Research Archives. 2026;14(1). [CrossRef]
- Raghavan PR. SOX Transcription Factor Network Modulation by Metadichol: A Novel Paradigm for Regenerative and Precision Medicine. Medical Research Archives. 2026;14(1). [CrossRef]
- Raghavan PR. Synergistic Targeting of Krüppel-like factor and Related Signaling Pathways by Metadichol: A Multidimensional Anticancer Strategy. Medical Research Archives. 2025;13(6). [DOI not available].
- Raghavan PR. Metadichol induces CD14 glycoprotein expression in human embryonic stem cells and fibroblasts. Medical Research Archives. 2025;13(5). [DOI not available].
- Raghavan PR. Metadichol, A Modulator that Controls Expression of Toll Like Receptors in Cancer Cell Lines. British Journal of Cancer Research. 2024;7(3):720-732. [CrossRef]
- Raghavan PR. Metadichol-induced expression of Toll receptor family members in peripheral blood mononuclear cells. Medical Research Archives. 2024;12(8). [CrossRef]
- Raghavan PR. Metadichol®-induced expression of circadian clock transcription factors in human fibroblasts. Medical Research Archives. 2024;12(6). [CrossRef]
- Raghavan PR. Metadichol-induced differentiation of pancreatic ductal cells (PANC-1) into insulin-producing cells. Medical Research Archives. 2023;11(11). [CrossRef]
- Raghavan PR. Metadichol®-induced expression of Sirtuins 1-7 in somatic and cancer cells. Medical Research Archives. 2024;12(6). [CrossRef]
- Raghavan PR. Metadichol® Induced the Expression of Neuronal Transcription Factors in Human Fibroblast Dermal Cells. Journal of Bioinformatics and Systems Biology. 2023;6:326-339. [CrossRef]
- Raghavan PR. Metadichol orchestrates pluripotency via nuclear receptors during cellular reprogramming. Med Res Arch. 2025;13(6). [CrossRef]
- Raghavan PR. Metadichol Stimulates Gene Expression Across Mammalian Species: Dose Response Characterization and Implications for Restoring Vitamin C Biosynthesis. Preprints. 2025;2025081935. [CrossRef]
- Raghavan PR. Metadichol® induced high levels of vitamin C: case studies. Vitam Miner. 2017;6:169. [DOI not available].
- Raghavan PR. Metadichol® and Vitamin C Increase In Vivo, an Open-Label Study. Vitam Miner. 2017;6:163. [DOI not available].
- Raghavan PR. The Quest for Immortality: Introducing Metadichol® a Novel Telomerase Activator. J Stem Cell Res Ther. 2019;9:446. [CrossRef]
- Raghavan PR. Improving Longevity with Metadichol® by Inhibiting the Bcat-1 Gene. Journal of Aging Science. 2017;5:1. [CrossRef]
- Raghavan PR. Metadichol, A Novel ROR Gamma Inverse Agonist and Its Applications in Psoriasis. J Clin Exp Dermatol Res. 2017;8:433. [CrossRef]
- Raghavan PR. Metadichol® A Novel Inverse Agonist of Thyroid Receptor and its Applications in Thyroid Diseases. Biol Med (Aligarh). 2019;11:458. [CrossRef]
- Raghavan PR. Metadichol®. A Novel Inverse Agonist of Aryl Hydrocarbon Receptor (AHR) and NRF2 Inhibitor. J Cancer Sci Ther. 2017;9:661-668. [CrossRef]
- Raghavan PR. VDR inverse agonism by metadichol enhances VDBP-mediated immunity. Preprints.org. 2025. [CrossRef]
- Raghavan PR. Metadichol®: A Nano Lipid Emulsion that Expresses All 49 Nuclear Receptors in Stem and Somatic Cells. Archives of Clinical and Biomedical Research. 2023;7:524-536. [CrossRef]
- Karlsson M, Zhang C, Méar L, et al. A single-cell type transcriptomics map of human tissues. Sci Adv. 2021;7(31):eabh2169. [CrossRef]
- Digre A, Lindskog C. The human protein atlas—integrated omics for single cell mapping of the human proteome. Protein Sci. 2023;32(1):e4562. [CrossRef]
- Amberger JS, Bocchini CA, Schiettecatte F, Scott AF, Hamosh A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 2015;43(D1):D789-D798. [CrossRef]
- Piñero J, Bravo À, Queralt-Rosinach N, et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 2017;45(D1):D833-D839. [CrossRef]
- Wharram BL, Goyal M, Gillespie PJ, et al. Altered podocyte structure in GLEPP1 (Ptpro)-deficient mice associated with hypertension and low glomerular filtration rate. J Clin Invest. 2000;106(10):1281-1290. [CrossRef]
- Jiang W, Wei M, Liu M, et al. Identification of protein tyrosine phosphatase receptor type O (PTPRO) as a synaptic adhesion molecule that promotes synapse formation. J Neurosci. 2017;37(41):9828-9843. [CrossRef]
- Lapointe CP, Grosely R, Djumagulov M, et al. eIF1 and eIF5 dynamically control translation start site fidelity. Nat Struct Mol Biol. 2025;32(7):1085-1096. [CrossRef]
- Hinnebusch AG. The scanning mechanism of eukaryotic translation initiation. Annu Rev Biochem. 2014;83:779-812. [CrossRef]
- Rich BE, Steitz JA. Human acidic ribosomal phosphoproteins P0, P1, and P2: analysis of cDNA clones, in vitro synthesis, and assembly. Mol Cell Biol. 1987;7(11):4065-4074. [CrossRef]
- Pakos-Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman AM. The integrated stress response. EMBO Rep. 2016;17(10):1374-1395. [CrossRef]
- Lee JW, Bae SH, Jeong JW, Kim SH, Kim KW. Hypoxia-inducible factor (HIF-1)α: its protein stability and biological functions. Exp Mol Med. 2004;36(1):1-12. [CrossRef]
- Lee AS. Glucose-regulated proteins in cancer: molecular mechanisms and therapeutic potential. Nat Rev Cancer. 2014;14(4):263-276. [CrossRef]
- Hetz C, Zhang K, Bhattacharya R. Mechanisms, regulation and functions of the unfolded protein response. Nat Rev Mol Cell Biol. 2020;21(8):421-438. [CrossRef]
- Ni M, Zhang Y, Lee AS. Beyond the endoplasmic reticulum: atypical GRP78 in cell viability, signalling and therapeutic targeting. Biochem J. 2011;434(2):181-188. [CrossRef]
- Luo B, Lee AS. The critical roles of endoplasmic reticulum chaperones and unfolded protein response in tumorigenesis and anticancer therapies. Oncogene. 2013;32(7):805-818. [CrossRef]
- Li L, Dong M, Wang XG. The implication and significance of beta 2 microglobulin: a conservative multifunctional regulator. Chin Med J (Engl). 2016;129(4):448-455. [CrossRef]
- Vitiello A, Potter TA, Sherman LA. The role of β2-microglobulin in peptide binding by class I molecules. Science. 1990;250(4986):1423-1426. [CrossRef]
- Zhang S, Liu X, Zhang Y, et al. The immune-related role of beta-2-microglobulin in melanoma. Front Oncol. 2022;12:944722. [CrossRef]
- Sirover MA. On the functional diversity of glyceraldehyde-3-phosphate dehydrogenase: biochemical mechanisms and regulatory control. Biochim Biophys Acta. 2011;1810(8):741-751. [CrossRef]
- Colell A, Green DR, Bhattacharya R. Novel roles for GAPDH in cell death and carcinogenesis. Cell Death Differ. 2009;16(12):1573-1581. [CrossRef]
- Lazarev VF, Guzhova IV, Margulis BA. Glyceraldehyde-3-phosphate dehydrogenase is a multifaceted therapeutic target. Pharmaceutics. 2020;12(5):416. [CrossRef]
- Bunnell TM, Burbach BJ, Shimizu Y, Bhattacharya R. β-Actin specifically controls cell growth, migration, and the G-actin pool. Mol Biol Cell. 2011;22(21):4047-4058. [CrossRef]
- Pollard TD, Cooper JA. Actin, a central player in cell shape and movement. Science. 2009;326(5958):1208-1212. [CrossRef]
- Piñeiro A, Cordero OJ, Nogueira M. Prothymosin alpha is involved in the regulation of cell proliferation. Histol Histopathol. 2000;15(1):265-276. [CrossRef]
- Karetsou Z, Sandaltzopoulos R, Frangou-Lazaridis M, et al. Prothymosin alpha modulates the interaction of histone H1 with chromatin. Nucleic Acids Res. 1998;26(13):3111-3118. [CrossRef]
- Gómez-Márquez J, Rodríguez P. Prothymosin alpha is a chromatin-remodeling protein in mammalian cells. Biochem J. 1998;333(Pt 1):1-3. [CrossRef]
- Böyum A. Isolation of mononuclear cells and granulocytes from human blood. Scand J Clin Lab Invest Suppl. 1968;97:77-89.
- Grievink HW, Luiber T, Moerland M, et al. Comparison of three isolation techniques for human peripheral blood mononuclear cells: cell recovery and viability, population composition, and cell functionality. Biopreserv Biobank. 2016;14(5):410-415. [CrossRef]
- Chomczynski P, Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987;162(1):156-159. [CrossRef]
- Bustin SA, Benes V, Garson JA, et al. The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem. 2009;55(4):611-622. [CrossRef]
- Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25(4):402-408. [CrossRef]
- Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc. 2008;3(6):1101-1108. [CrossRef]
- Pakos-Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman AM. The integrated stress response. EMBO Rep. 2016;17(10):1374-1395. [CrossRef]
- Wortel IMN, van der Meer LT, Kilberg MS, van Leeuwen FN. Surviving stress: modulation of ATF4-mediated stress responses in normal and malignant cells. Trends Endocrinol Metab. 2017;28(11):794-806. [CrossRef]
- Harding HP, Zhang Y, Zeng H, et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol Cell. 2003;11(3):619-633. [CrossRef]
- Vattem KM, Wek RC. Reinitiation involving upstream ORFs regulates ATF4 mRNA translation in mammalian cells. Proc Natl Acad Sci U S A. 2004;101(31):11269-11274. [CrossRef]
- B'chir W, Maurin AC, Carraro V, et al. The eIF2α/ATF4 pathway is essential for stress-induced autophagy gene expression. Nucleic Acids Res. 2013;41(16):7683-7699. [CrossRef]
- Ye J, Koumenis C. ATF4, an ER stress and hypoxia-inducible transcription factor and its potential role in hypoxia tolerance and tumorigenesis. Curr Mol Med. 2009;9(4):411-416. [CrossRef]
- Kilberg MS, Shan J, Su N. ATF4-dependent transcription mediates signaling of amino acid limitation. Trends Endocrinol Metab. 2009;20(9):436-443. [CrossRef]
- Ameri K, Harris AL. Activating transcription factor 4. Int J Biochem Cell Biol. 2008;40(1):14-21. [CrossRef]
- Liu Z, Gu H, Gan L, et al. Reducing Smad3/ATF4 was essential for Sirt1 inhibiting ER stress-induced apoptosis in mice brown adipose tissue. Oncotarget. 2017;8(6):9267-9279. [CrossRef]
- Ghosh HS, Reizis B, Bhattacharya D. SIRT1 associates with eIF2-alpha and regulates the cellular stress response. Sci Rep. 2011;1:150. [CrossRef]
- Lim JH. SIRT1 suppresses activating transcription factor 4 (ATF4) expression in response to proteasome inhibition. Biochem Biophys Res Commun. 2013;441(2):289-294. [CrossRef]
- Brugarolas J, Lei K, Hurley RL, et al. Regulation of mTOR function in response to hypoxia by REDD1 and the TSC1/TSC2 tumor suppressor complex. Genes Dev. 2004;18(23):2893-2904. [CrossRef]
- DeYoung MP, Horak P, Sofer A, Sgroi D, Ellisen LW. Hypoxia regulates TSC1/2-mTOR signaling and tumor suppression through REDD1-mediated 14-3-3 shuttling. Genes Dev. 2008;22(2):239-251. [CrossRef]
- Pike JW, Meyer MB. The vitamin D receptor: new paradigms for the regulation of gene expression by 1,25-dihydroxyvitamin D3. Endocrinol Metab Clin North Am. 2010;39(2):255-269. [CrossRef]
- Bikle DD. Vitamin D metabolism, mechanism of action, and clinical applications. Chem Biol. 2014;21(3):319-329. [CrossRef]
- Wharram BL, Goyal M, Gillespie PJ, et al. Altered podocyte structure in GLEPP1 (Ptpro)-deficient mice associated with hypertension and low glomerular filtration rate. J Clin Invest. 2000;106(10):1281-1290. [CrossRef]
- Jiang W, Wei M, Liu M, et al. Identification of protein tyrosine phosphatase receptor type O (PTPRO) as a synaptic adhesion molecule that promotes synapse formation. J Neurosci. 2017;37(41):9828-9843. [CrossRef]
- Xie F, Dong H, Zhang H. Regulatory functions of protein tyrosine phosphatase receptor type O in immune cells. Front Immunol. 2021;12:783370. [CrossRef]
- Motiwala T, Ghoshal K, Das A, et al. Suppression of the protein tyrosine phosphatase receptor type O gene (PTPRO) by methylation in hepatocellular carcinomas. Oncogene. 2003;22(41):6319-6331. [CrossRef]
- Motiwala T, Kutay H, Ghoshal K, et al. Protein tyrosine phosphatase receptor-type O (PTPRO) exhibits characteristics of a candidate tumor suppressor in human lung cancer. Proc Natl Acad Sci U S A. 2004;101(38):13844-13849. [CrossRef]
- Motiwala T, Majumder S, Kutay H, et al. Methylation and silencing of protein tyrosine phosphatase receptor type O in chronic lymphocytic leukemia. Clin Cancer Res. 2007;13(11):3174-3181. [CrossRef]
- Hsu SH, Motiwala T, Roy S, et al. Methylation of the PTPRO gene in human hepatocellular carcinoma and identification of VCP as its substrate. J Cell Biochem. 2013;114(8):1810-1818. [CrossRef]
- Jacob ST, Bhattacharya R. Epigenetic regulation of protein tyrosine phosphatases: potential molecular targets for cancer therapy. Genes Cancer. 2010;1(12):1232-1239. [CrossRef]
- Li SY, Li R, Chen YL, et al. Aberrant PTPRO methylation in tumor tissues as a potential biomarker that predicts clinical outcomes in breast cancer patients. BMC Genet. 2014;15:67. [CrossRef]
- Vousden KH, Prives C. Blinded by the light: the growing complexity of p53. Cell. 2009;137(3):413-431. [CrossRef]
- Kastenhuber ER, Lowe SW. Putting p53 in context. Cell. 2017;170(6):1062-1078. [CrossRef]
- Levine AJ. p53: 800 million years of evolution and 40 years of discovery. Nat Rev Cancer. 2020;20(8):471-480. [CrossRef]
- Eijkelenboom A, Burgering BM. FOXOs: signalling integrators for homeostasis maintenance. Nat Rev Mol Cell Biol. 2013;14(2):83-97. [CrossRef]
- Webb AE, Brunet A. FOXO transcription factors: key regulators of cellular quality control. Trends Biochem Sci. 2014;39(4):159-169. [CrossRef]
- Semenza GL. Hypoxia-inducible factors in physiology and medicine. Cell. 2012;148(3):399-408. [CrossRef]
- Majmundar AJ, Wong WJ, Simon MC. Hypoxia-inducible factors and the response to hypoxic stress. Mol Cell. 2010;40(2):364-374. [CrossRef]
- Dengler VL, Galbraith M, Bhattacharya R. Transcriptional regulation by hypoxia inducible factors. Crit Rev Biochem Mol Biol. 2014;49(1):1-15. [CrossRef]
- Lim JH, Lee YM, Chun YS, Chen J, Kim JE, Park JW. Sirtuin 1 modulates cellular responses to hypoxia by deacetylating hypoxia-inducible factor 1alpha. Mol Cell. 2010;38(6):864-878. [CrossRef]
- Joo HY, Yun M, Jeong J, et al. SIRT1 deacetylates and stabilizes hypoxia-inducible factor-1α (HIF-1α) via direct interactions during hypoxia. Biochem Biophys Res Commun. 2015;462(4):294-300. [CrossRef]
- Laughner E, Taghavi P, Chiles K, Mahon PC, Semenza GL. HER2 (neu) signaling increases the rate of hypoxia-inducible factor 1alpha (HIF-1alpha) synthesis: novel mechanism for HIF-1-mediated vascular endothelial growth factor expression. Mol Cell Biol. 2001;21(12):3995-4004. [CrossRef]
- Zhong L, D'Urso A, Toiber D, et al. The histone deacetylase Sirt6 regulates glucose homeostasis via Hif1alpha. Cell. 2010;140(2):280-293. [CrossRef]
- Shoshani T, Faerman A, Mett I, et al. Identification of a novel hypoxia-inducible factor 1-responsive gene, RTP801, involved in apoptosis. Mol Cell Biol. 2002;22(7):2283-2293. [CrossRef]
- Bjorkman, P. J., and P. Parham. 1990. "Structure, Function, and Diversity of Class I Major Histocompatibility Complex Molecules." Annual Review of Biochemistry 59:253-288.
- Güssow, D., R. Rein, I. Ginjaar, F. Hochstenbach, G. Seemann, A. Kottman, and H. L. Ploegh. 1987. "The Human Beta 2-Microglobulin Gene: Primary Structure and Definition of the Transcriptional Unit." Journal of Immunology 139(9):3132-3138.
- Germain, R. N. 1994. "MHC-Dependent Antigen Processing and Peptide Presentation: Providing Ligands for T Lymphocyte Activation." Cell 76(2):287-299.
- Cresswell, P., A. L. Ackerman, A. Giodini, D. R. Peaper, and P. A. Wearsch. 2005. "Mechanisms of MHC Class I-Restricted Antigen Processing and Cross-Presentation." Immunological Reviews 207:145-157.
- Townsend, A., and H. Bodmer. 1989. "Antigen Recognition by Class I-Restricted T Lymphocytes." Annual Review of Immunology 7:601-624.
- Neefjes, J., M. L. Jongsma, P. Paul, and O. Bakke. 2011. "Towards a Systems Understanding of MHC Class I and MHC Class II Antigen Presentation." Nature Reviews Immunology 11(12):823-836.
- Feder, J. N., A. Gnirke, W. Thomas, Z. Tsuchihashi, D. A. Ruddy, A. Basava, et al. 1996. "A Novel MHC Class I-Like Gene Is Mutated in Patients with Hereditary Haemochromatosis." Nature Genetics 13(4):399-408.
- Ganz, T. 2013. "Systemic Iron Homeostasis." Physiological Reviews 93(4):1721-1741.
- Roopenian, D. C., and S. Akilesh. 2007. "FcRn: The Neonatal Fc Receptor Comes of Age." Nature Reviews Immunology 7(9):715-725.
- Gobin, S. J., V. Keijsers, M. van Zutphen, and P. J. van den Elsen. 1998. "The Role of Enhancer A in the Locus-Specific Transactivation of Classical and Nonclassical HLA Class I Genes by Nuclear Factor κB." Journal of Immunology 161(5):2276-2283.
- Meissner, T. B., A. Li, A. Biswas, K. H. Lee, Y. J. Liu, E. Bayber, et al. 2010. "NLR Family Member NLRC5 Is a Transcriptional Regulator of MHC Class I Genes." Proceedings of the National Academy of Sciences107(31):13794-13799.
- Boehm, U., T. Klamp, M. Groot, and J. C. Howard. 1997. "Cellular Responses to Interferon-Gamma." Annual Review of Immunology 15:749-795.
- Taniguchi, T., and A. Takaoka. 2001. "A Weak Signal for Strong Responses: Interferon-Alpha/Beta Revisited." Nature Reviews Molecular Cell Biology 2(5):378-386.
- Hewison, M. 2012. "Vitamin D and Immune Function: An Overview." Proceedings of the Nutrition Society71(1):50-61.
- Carlberg, C. 2019. "Vitamin D Signaling in the Context of Innate Immunity: Focus on Human Monocytes." Frontiers in Immunology 10:2211.
- Mora, J. R., M. Iwata, and U. H. von Andrian. 2008. "Vitamin Effects on the Immune System: Vitamins A and D Take Centre Stage." Nature Reviews Immunology 8(9):685-698.
- Pakos-Zebrucka, K., I. Koryga, K. Mnich, M. Ljujic, A. Samali, and A. M. Gorman. 2016. "The Integrated Stress Response." EMBO Reports 17(10):1374-1395.
- Donnelly, N., A. M. Gorman, A. Gupta, and A. Bhattacharya. 2013. "The eIF2α Kinases: Their Structures and Functions." Cellular and Molecular Life Sciences 70(19):3493-3511.
- McNab, F., K. Mayer-Barber, A. Sher, A. Wack, and A. O'Garra. 2015. "Type I Interferons in Infectious Disease." Nature Reviews Immunology 15(2):87-103.
- Yeung, F., J. E. Hoberg, C. S. Ramsey, M. D. Keller, D. R. Jones, R. A. Frye, et al. 2004. "Modulation of NF-κB-Dependent Transcription and Cell Survival by the SIRT1 Deacetylase." EMBO Journal 23(12):2369-2380.
- Laplante, M., and D. M. Sabatini. 2012. "mTOR Signaling in Growth Control and Disease." Cell 149(2):274-293.
- Kauppinen, A., T. Suuronen, J. Ojala, K. Kaarniranta, and A. Salminen. 2013. "Antagonistic Crosstalk between NF-κB and SIRT1 in the Regulation of Inflammation and Metabolic Disorders." Cellular Signalling 25(10):1939-1948.
- Liu, T. F., and C. E. McCall. 2013. "Deacetylation by SIRT1 Reprograms Inflammation and Cancer." Genes and Cancer 4(3-4):135-147.
- Chen, L., H. Deng, H. Cui, J. Fang, Z. Zuo, J. Deng, et al. 2018. "Inflammatory Responses and Inflammation-Associated Diseases in Organs." Oncotarget 9(6):7204-7218.
- Ellisen LW, Ramsayer KD, Johannessen CM, et al. REDD1, a developmentally regulated transcriptional target of p63 and p53, links p63 to regulation of reactive oxygen species. Mol Cell. 2002;10(5):995-1005. [CrossRef]
- Wang M, Wey S, Zhang Y, Ye R, Lee AS. Role of the unfolded protein response regulator GRP78/BiP in development, cancer, and neurological disorders. Antioxid Redox Signal. 2009;11(9):2307-2316. [CrossRef]
- Lee AS. Stress-induced translocation of the endoplasmic reticulum chaperone GRP78/BiP and its impact on human disease and therapy. Proc Natl Acad Sci U S A. 2025;122(30):e2412246122. [CrossRef]
- Hetz C, Zhang K, Bhalla AK. Mechanisms, regulation and functions of the unfolded protein response. Nat Rev Mol Cell Biol. 2020;21(8):421-438. [CrossRef]
- Ni M, Zhang Y, Lee AS. Beyond the endoplasmic reticulum: atypical GRP78 in cell viability, signalling and therapeutic targeting. Biochem J. 2011;434(2):181-188. [CrossRef]
- Starenki D, Hong SK, Lloyd RV, Park JI. Mortalin (GRP75/HSPA9) upregulation promotes survival and proliferation of medullary thyroid carcinoma cells. Oncogene. 2015;34(35):4624-4634. [CrossRef]
- Ferré CA, Bhattacharyya S, Bhattacharyya P, et al. Mortalin/Hspa9 involvement and therapeutic perspective in Parkinson's disease. Int J Mol Sci. 2022;23(17):9823. [CrossRef]
- Goswami AV, Samber S, Bhattacharyya S, et al. HSPA9/Mortalin mediates axo-protection and modulates mitochondrial dynamics in neurons. Sci Rep. 2021;11(1):17705. [CrossRef]
- Schrag JD, Bergeron JJ, Li Y, et al. The structure of calnexin, an ER chaperone involved in quality control of protein folding. Mol Cell. 2001;8(3):633-644. [CrossRef]
- Lamriben L, Graham JB, Adams BM, Bhattacharyya S. Calnexin cycle - structural features of the ER chaperone system. FEBS J. 2020;287(20):4322-4340. [CrossRef]
- Williams DB. Beyond lectins: the calnexin/calreticulin chaperone system of the endoplasmic reticulum. J Cell Sci. 2006;119(Pt 4):615-623. [CrossRef]
- Westerheide SD, Anckar J, Stevens SM Jr, Sistonen L, Morimoto RI. Stress-inducible regulation of heat shock factor 1 by the deacetylase SIRT1. Science. 2009;323(5917):1063-1066. [CrossRef]
- van de Ven RAH, Santos D, Bhattacharyya S. Chaperones and beyond as key players in pluripotency maintenance. Front Cell Dev Biol. 2019;7:150. [CrossRef]
- van Dam S, Võsa U, van der Graaf A, Franke L, de Magalhães JP. Gene co-expression analysis for functional classification and gene-disease predictions. Brief Bioinform. 2018;19(4):575-592. [CrossRef]
- Vandesompele J, De Preter K, Pattyn F, et al. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002;3(7):RESEARCH0034. [CrossRef]
- Hounkpe BW, Cheber F, Grisci VDMTD, de Lima F, De Paula EV. HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Res. 2022;50(D1):D605-D614. [CrossRef]
- Eisenberg E, Levanon EY. Human housekeeping genes, revisited. Trends Genet. 2013;29(10):569-574. [CrossRef]
- Zhu J, He F, Hu S, Yu J. On the nature of human housekeeping genes. Trends Genet. 2008;24(10):481-484. [CrossRef]
- Lemma S, Avnet S, Salerno M, Chano T, Baldini N. Identification and validation of housekeeping genes for gene expression analysis of cancer stem cells. PLoS One. 2016;11(2):e0149481. [CrossRef]
- Thellin O, Zorzi W, Lakaye B, et al. Housekeeping genes as internal standards: use and limits. J Biotechnol. 1999;75(2-3):291-295. [CrossRef]
- Caracausi M, Piovesan A, Antonaros F, Strippoli P, Vitale L, Pelleri MC. Systematic identification of human housekeeping genes possibly useful as references in gene expression studies. Mol Med Rep. 2017;16(3):2397-2410. [CrossRef]
- Wortel IMN, van der Meer LT, Kilberg MS, van Leeuwen FN. Surviving stress: modulation of ATF4-mediated stress responses in normal and malignant cells. Trends Endocrinol Metab. 2017;28(11):794-806. [CrossRef]
- Wang GL, Jiang BH, Rue EA, Semenza GL. Hypoxia-inducible factor 1 is a basic-helix-loop-helix-PAS heterodimer regulated by cellular O2 tension. Proc Natl Acad Sci U S A. 1995;92(12):5510-5514. [CrossRef]
- Kim R, Emi M, Tanabe K, Murakami S. Role of the unfolded protein response in cell death. Apoptosis. 2006;11(1):5-13. [CrossRef]
- Hetz C, Zhang K, Bhakta K. Mechanisms, regulation and functions of the unfolded protein response. Nat Rev Mol Cell Biol. 2020;21(8):421-438. [CrossRef]
- Dang CV. MYC on the path to cancer. Cell. 2012;149(1):22-35. [CrossRef]
- Lambert SA, Jolma A, Campitelli LF, et al. The human transcription factors. Cell. 2018;172(4):650-665. [CrossRef]
- Li X, Jiang Y, Meisenhelder J, et al. Mitochondria-translocated PGK1 functions as a protein kinase to coordinate glycolysis and the TCA cycle in tumorigenesis. Mol Cell. 2016;61(5):705-719. [CrossRef]
- Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41(3):211-218. [CrossRef]
- Zhang W, Xu Q, Liu P, et al. Discovery of novel glyceraldehyde-3-phosphate dehydrogenase inhibitor via docking-based virtual screening. Bioorg Chem. 2020;96:103605. [CrossRef]
- Lamming DW, Ye L, Katajisto P, et al. Rapamycin-induced insulin resistance is mediated by mTORC2 loss and uncoupled from longevity. Science. 2012;335(6076):1638-1643. [CrossRef]
- Barlow AD, Nicholson ML, Herbert TP. Evidence for rapamycin toxicity in pancreatic β-cells and a review of the underlying molecular mechanisms. Diabetes. 2013;62(8):2674-2682. [CrossRef]
- Fang Y, Westbrook R, Hill C, et al. Duration of rapamycin treatment has differential effects on metabolism in mice. Cell Metab. 2013;17(3):456-462. [CrossRef]
- Yang SB, Lee HY, Young DM, et al. Rapamycin induces glucose intolerance in mice by reducing islet mass, insulin content, and insulin sensitivity. J Mol Med. 2012;90(8):897-905. [CrossRef]
- Houde VP, Brûlé S, Bhullar KS, et al. Chronic rapamycin treatment causes glucose intolerance and hyperlipidemia by upregulating hepatic gluconeogenesis and impairing lipid deposition in adipose tissue. Diabetes. 2010;59(6):1338-1348. [CrossRef]
- Alemán CL, Más R, Hernández C, et al. A 12-month study of policosanol oral toxicity in Sprague Dawley rats. Toxicol Lett. 1994;70(1):77-87. [CrossRef]
- Alemán CL, Más Ferreiro R, Noa Puig M, et al. Carcinogenicity of policosanol in Sprague-Dawley rats: a 24-month study. Teratog Carcinog Mutagen. 1994;14(5):239-249. [CrossRef]
- Califano A, Alvarez MJ. The recurrent architecture of tumour initiation, progression and drug sensitivity. Nat Rev Cancer. 2017;17(2):116-130. [CrossRef]
- Pakos-Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman AM. The integrated stress response. EMBO Rep. 2016;17(10):1374-1395. [CrossRef]
- Harding HP, Zhang Y, Zeng H, et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol Cell. 2003;11(3):619-633. [CrossRef]
- Ye J, Kumanova M, Hart LS, et al. The GCN2-ATF4 pathway is critical for tumour cell survival and proliferation in response to nutrient deprivation. EMBO J. 2010;29(12):2082-2096. [CrossRef]
- B'chir W, Maurin AC, Carraro V, et al. The eIF2α/ATF4 pathway is essential for stress-induced autophagy gene expression. Nucleic Acids Res. 2013;41(16):7683-7699. [CrossRef]
- Webb AE, Brunet A. FOXO transcription factors: key regulators of cellular quality control. Trends Biochem Sci. 2014;39(4):159-169. [CrossRef]
- Vattem KM, Wek RC. Reinitiation involving upstream ORFs regulates ATF4 mRNA translation in mammalian cells. Proc Natl Acad Sci U S A. 2004;101(31):11269-11274. [CrossRef]
- Semenza GL. Hypoxia-inducible factors in physiology and medicine. Cell. 2012;148(3):399-408. [CrossRef]
- Pugh CW, Ratcliffe PJ. Regulation of angiogenesis by hypoxia: role of the HIF system. Nat Med. 2003;9(6):677-684. [CrossRef]
- Kim JW, Tchernyshyov I, Semenza GL, Dang CV. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 2006;3(3):177-185. [CrossRef]
- Forsythe JA, Jiang BH, Iyer NV, et al. Activation of vascular endothelial growth factor gene transcription by hypoxia-inducible factor 1. Mol Cell Biol. 1996;16(9):4604-4613. [CrossRef]
- Maxwell PH, Wiesener MS, Chang GW, et al. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature. 1999;399(6733):271-275. [CrossRef]
- Sonenberg N, Hinnebusch AG. Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell. 2009;136(4):731-745. [CrossRef]
- Ma XM, Blenis J. Molecular mechanisms of mTOR-mediated translational control. Nat Rev Mol Cell Biol. 2009;10(5):307-318. [CrossRef]
- Thoreen CC, Chantranupong L, Keys HR, Wang T, Gray NS, Sabatini DM. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature. 2012;485(7396):109-113. [CrossRef]
- Gingras AC, Raught B, Sonenberg N. eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu Rev Biochem. 1999;68:913-963. [CrossRef]
- Holcik M, Sonenberg N. Translational control in stress and apoptosis. Nat Rev Mol Cell Biol. 2005;6(4):318-327. [CrossRef]
- Germain RN. MHC-dependent antigen processing and peptide presentation: providing ligands for T lymphocyte activation. Cell. 1994;76(2):287-299. [CrossRef]
- Zaretsky JM, Garcia-Diaz A, Shin DS, et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N Engl J Med. 2016;375(9):819-829. [CrossRef]
- Sade-Feldman M, Jiao YJ, Chen JH, et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat Commun. 2017;8(1):1136. [CrossRef]
- Lee AS. The ER chaperone and signaling regulator GRP78/BiP as a monitor of endoplasmic reticulum stress. Methods. 2005;35(4):373-381. [CrossRef]
- Wang M, Wey S, Zhang Y, Ye R, Lee AS. Role of the unfolded protein response regulator GRP78/BiP in development, cancer, and neurological disorders. Antioxid Redox Signal. 2009;11(9):2307-2316. [CrossRef]
- Hetz C, Chevet E, Oakes SA. Proteostasis control by the unfolded protein response. Nat Cell Biol. 2015;17(7):829-838. [CrossRef]
- Walter P, Ron D. The unfolded protein response: from stress pathway to homeostatic regulation. Science. 2011;334(6059):1081-1086. [CrossRef]
- Pobre KFR, Poet GJ, Hendershot LM. The endoplasmic reticulum (ER) chaperone BiP is a master regulator of ER functions: Getting by with a little help from ERdj friends. J Biol Chem. 2019;294(6):2098-2108. [CrossRef]
- Goldbeter A, Koshland DE Jr. An amplified sensitivity arising from covalent modification in biological systems. Proc Natl Acad Sci U S A. 1981;78(11):6840-6844. [CrossRef]
- Heinrich R, Neel BG, Rapoport TA. Mathematical models of protein kinase signal transduction. Mol Cell. 2002;9(5):957-970. [CrossRef]
- Ferrell JE Jr. Tripping the switch fantastic: how a protein kinase cascade can convert graded inputs into switch-like outputs. Trends Biochem Sci. 1996;21(12):460-466. [CrossRef]
- Kholodenko BN. Cell-signalling dynamics in time and space. Nat Rev Mol Cell Biol. 2006;7(3):165-176. [CrossRef]
- Britto FA, Dumas K, Giorgetti-Peraldi S, et al. Is REDD1 a metabolic double agent? Lessons from physiology and pathology. Am J Physiol Cell Physiol. 2020;319(5):C807-C824. [CrossRef]
- Pinto JA, Rolfo C, Raez LE, et al. In silico evaluation of DNA damage inducible transcript 4 gene (DDIT4) as prognostic biomarker in several malignancies. Front Oncol. 2018;8:106. [CrossRef]
- Gharibi B, Farzadi S, Ghuman M, Hughes FJ. DDIT4 regulates mesenchymal stem cell fate by mediating between HIF1α and mTOR signalling. Sci Rep. 2016;6:36889. [CrossRef]
- Do H, Bhrolchain M, Bhatt A, et al. Roles of farnesyl-diphosphate farnesyltransferase 1 in tumour and tumour microenvironments. Cells. 2020;9(11):2352. [CrossRef]
- Weng ML, Chen WK, Chen XY, et al. Fasting inhibits aerobic glycolysis and proliferation in colorectal cancer via the Fdft1-mediated AKT/mTOR/HIF1α pathway suppression. Nat Commun. 2020;11(1):1869. [CrossRef]
- Germain RN. MHC-dependent antigen processing and peptide presentation: providing ligands for T lymphocyte activation. Cell. 1994;76(2):287-299. [CrossRef]
- Motiwala T, Ghoshal K, Das A, et al. Suppression of the protein tyrosine phosphatase receptor type O gene (PTPRO) by methylation in hepatocellular carcinomas. Oncogene. 2003;22(41):6319-6331. [CrossRef]
- Pakos-Zebrucka K, Koryga I, Mnich K, Ljujic M, Samali A, Gorman AM. The integrated stress response. EMBO Rep. 2016;17(10):1374-1395. [CrossRef]
- Semenza GL. Hypoxia-inducible factors in physiology and medicine. Cell. 2012;148(3):399-408. [CrossRef]
- Wang M, Kaufman RJ. Protein misfolding in the endoplasmic reticulum as a conduit to human disease. Nature. 2016;529(7586):326-335. [CrossRef]
- Motiwala T, Kutay H, Ghoshal K, et al. Protein tyrosine phosphatase receptor-type O (PTPRO) exhibits characteristics of a candidate tumor suppressor in human lung cancer. Proc Natl Acad Sci U S A. 2004;101(38):13844-13849. [CrossRef]
- Motiwala T, Majumder S. Epigenetic regulation of protein tyrosine phosphatases: potential molecular targets for cancer therapy. Clin Immunol. 2005;117(1):115-125. [DOI not available].
- Motiwala T, Kutay H, Zanesi N, et al. PTPROt-mediated regulation of p53/Foxm1 suppresses leukemic phenotype in a CLL mouse model. Blood. 2015;126(15):1850-1859. [CrossRef]
- Ming F, Sun Q. Epigenetically silenced PTPRO functions as a prognostic marker and tumor suppressor in human lung squamous cell carcinoma. Mol Med Rep. 2017;16(1):746-754. [CrossRef]
- Xu J, Hou J, Wang Y, et al. PTPRO-mediated autophagy prevents hepatosteatosis and tumorigenesis. Oncotarget. 2017;8(6):10373-10383. [CrossRef]
- Thoreen CC, Chantranupong L, Keys HR, Wang T, Gray NS, Sabatini DM. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature. 2012;485(7396):109-113. [CrossRef]
- Silvera D, Formenti SC, Schneider RJ. Translational control in cancer. Nat Rev Cancer. 2010;10(4):254-266. [CrossRef]
- Bhat M, Robichaud N, Hulea L, Sonenberg N, Pelletier J, Topisirovic I. Targeting the translation machinery in cancer. Nat Rev Drug Discov. 2015;14(4):261-278. [CrossRef]
- Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41(3):211-218. [CrossRef]
- Ganapathy-Kanniappan S, Geschwind JF. Tumor glycolysis as a target for cancer therapy: progress and prospects. Mol Cancer. 2013;12:152. [CrossRef]
- Zhang JY, Zhang F, Hong CQ, et al. Critical protein GAPDH and its regulatory mechanisms in cancer cells. Cancer Biol Med. 2015;12(1):10-22. [CrossRef]
- Zaretsky JM, Garcia-Diaz A, Shin DS, et al. Mutations associated with acquired resistance to PD-1 blockade in melanoma. N Engl J Med. 2016;375(9):819-829. [CrossRef]
- Sade-Feldman M, Jiao YJ, Chen JH, et al. Resistance to checkpoint blockade therapy through inactivation of antigen presentation. Nat Commun. 2017;8(1):1136. [CrossRef]
- Gettinger S, Choi J, Hastings K, et al. Impaired HLA class I antigen processing and presentation as a mechanism of acquired resistance to immune checkpoint inhibitors in lung cancer. Cancer Discov. 2017;7(12):1420-1435. [CrossRef]
- Maleno I, Aptsiauri N, Cabrera T, Gallego A, Paschen A, López-Nevot MA. Frequent loss of heterozygosity in the β2-microglobulin region of chromosome 15 in primary human tumors. Immunogenetics. 2011;63(2):65-71. [CrossRef]
- del Campo AB, Kyte JA, Carretero J, et al. Immune escape of cancer cells with beta2-microglobulin loss over the course of metastatic melanoma. Int J Cancer. 2014;134(1):102-113. [CrossRef]
- Hicklin DJ, Marincola FM, Ferrone S. HLA class I antigen downregulation in human cancers: T-cell immunotherapy revives an old story. Mol Med Today. 1999;5(4):178-186. [CrossRef]
- Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100(1):57-70. [CrossRef]
- Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144(5):646-674. [CrossRef]
- Hanahan D. Hallmarks of cancer: new dimensions. Cancer Discov. 2022;12(1):31-46. [CrossRef]
- Piñero J, Ramírez-Anguita JM, Saüch-Pitarch J, et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 2020;48(D1):D845-D855. [CrossRef]
- Rappaport N, Twik M, Plaschkes I, et al. MalaCards: an amalgamated human disease compendium with diverse clinical and genetic annotation and structured search. Nucleic Acids Res. 2017;45(D1):D733-D750. [CrossRef]
- Amberger JS, Bocchini CA, Schiettecatte F, Scott AF, Hamosh A. OMIM.org: Online Mendelian Inheritance in Man (OMIM®), an online catalog of human genes and genetic disorders. Nucleic Acids Res. 2015;43(Database issue):D789-D798. [CrossRef]
- Szklarczyk D, Gable AL, Lyon D, et al. STRING v11: protein-protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019;47(D1):D483-D490. [CrossRef]
- Safran M, Dalah I, Alexander J, et al. GeneCards Version 3: the human gene integrator. Database (Oxford). 2010;2010:baq020. [CrossRef]
- Fontana L, Partridge L, Longo VD. Extending healthy life span--from yeast to humans. Science. 2010;328(5976):321-326. [CrossRef]
- López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The hallmarks of aging. Cell. 2013;153(6):1194-1217. [CrossRef]
- Kennedy BK, Lamming DW. The mechanistic target of rapamycin: the grand conductor of metabolism and aging. Cell Metab. 2016;23(6):990-1003. [CrossRef]
- Lamming DW, Ye L, Katajisto P, et al. Rapamycin-induced insulin resistance is mediated by mTORC2 loss and uncoupled from longevity. Science. 2012;335(6076):1638-1643. [CrossRef]
- Howell JJ, Manning BD. mTOR couples cellular nutrient sensing to organismal metabolic homeostasis. Trends Endocrinol Metab. 2011;22(3):94-102. [CrossRef]
- Johnson SC, Rabinovitch PS, Kaeberlein M. mTOR is a key modulator of ageing and age-related disease. Nature. 2013;493(7432):338-345. [CrossRef]
- Arriola Apelo SI, Lamming DW. Rapamycin: an inhibiTOR of aging emerges from the soil of Easter Island. J Gerontol A Biol Sci Med Sci. 2016;71(7):841-849. [CrossRef]
- Li W, Miller RA. Elevated ATF4 function in fibroblasts and liver of slow-aging mutant mice. J Gerontol A Biol Sci Med Sci. 2015;70(3):263-272. [CrossRef]
- Li W, Li X, Miller RA. ATF4 activity: a common feature shared by many kinds of slow-aging mice. Aging Cell. 2014;13(6):1012-1018. [CrossRef]
- Statzer C, Meng J, Venz R, et al. ATF-4 and hydrogen sulfide signalling mediate longevity in response to inhibition of translation or mTORC1. Nat Commun. 2022;13(1):967. [CrossRef]
- Rubinsztein DC, Mariño G, Kroemer G. Autophagy and aging. Cell. 2011;146(5):682-695. [CrossRef]
- Kaushik S, Cuervo AM. Proteostasis and aging. Nat Med. 2015;21(12):1406-1415. [CrossRef]
- Hartl FU, Bracher A, Hayer-Hartl M. Molecular chaperones in protein folding and proteostasis. Nature. 2011;475(7356):324-332. [CrossRef]
- Morimoto RI. Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Genes Dev. 2008;22(11):1427-1438. [CrossRef]
- Labbadia J, Morimoto RI. The biology of proteostasis in aging and disease. Annu Rev Biochem. 2015;84:435-464. [CrossRef]
- Hipp MS, Kasturi P, Hartl FU. The proteostasis network and its decline in ageing. Nat Rev Mol Cell Biol. 2019;20(7):421-435. [CrossRef]
- Klaips CL, Jayaraj GG, Hartl FU. Pathways of cellular proteostasis in aging and disease. J Cell Biol. 2018;217(1):51-63. [CrossRef]
- Rock KL, Reits E, Neefjes J. Present yourself! By MHC class I and MHC class II molecules. Trends Immunol. 2016;37(11):724-737. [CrossRef]
- Leone P, Shin EC, Perosa F, Vacca A, Dammacco F, Racanelli V. MHC class I antigen processing and presenting machinery: organization, function, and defects in tumor cells. J Natl Cancer Inst. 2013;105(16):1172-1187. [CrossRef]
- Neefjes J, Jongsma ML, Paul P, Bakke O. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat Rev Immunol. 2011;11(12):823-836. [CrossRef]
- Chen L, Juszczynski P, Takeyama K, Aguiar RC, Shipp MA. Protein tyrosine phosphatase receptor-type O truncated (PTPROt) regulates SYK phosphorylation, proximal B-cell-receptor signaling, and cellular proliferation. Blood. 2006;108(10):3428-3433. [CrossRef]
- Juszczynski P, Chen L, O'Donnell E, et al. BCL6 modulates tonic BCR signaling in diffuse large B-cell lymphomas by repressing the SYK phosphatase, PTPROt. Blood. 2009;114(26):5315-5321. [CrossRef]
- Stanford SM, Bottini N. PTPN22: the archetypal non-HLA autoimmunity gene. Nat Rev Rheumatol. 2014;10(10):602-611. [CrossRef]
- Kawai T, Akira S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat Immunol. 2010;11(5):373-384. [CrossRef]
- Mogensen TH. Pathogen recognition and inflammatory signaling in innate immune defenses. Clin Microbiol Rev. 2009;22(2):240-273. [CrossRef]
- Zanoni I, Granucci F. Role of CD14 in host protection against infections and in metabolism regulation. Front Cell Infect Microbiol. 2013;3:32. [CrossRef]
- Hernández-Caselles T, Martínez-Esparza M, Pérez-Oliva AB, et al. A study of CD33 (SIGLEC-3) antigen expression and function on activated human T and NK cells. J Leukoc Biol. 2006;79(1):46-58. [CrossRef]
- Sidney LE, Branch MJ, Dunphy SE, Dua HS, Hopkinson A. Concise review: evidence for CD34 as a common marker for diverse progenitors. Stem Cells. 2014;32(6):1380-1389. [CrossRef]
- Tristan C, Shahani N, Bhopale V, et al. The diverse functions of GAPDH: views from different subcellular compartments. Cell Signal. 2011;23(2):317-323. [CrossRef]
- Colell A, Green DR, Ricci JE. Novel roles for GAPDH in cell death and carcinogenesis. Cell Death Differ. 2009;16(12):1573-1581. [CrossRef]
- Tansey TR, Shechter I. Structure and regulation of mammalian squalene synthase. Biochim Biophys Acta. 2000;1529(1-3):49-62. [CrossRef]
- Harding HP, Zhang Y, Zeng H, et al. An integrated stress response regulates amino acid metabolism and resistance to oxidative stress. Mol Cell. 2003;11(3):619-633. [CrossRef]
- Kilberg MS, Shan J, Su N. ATF4-dependent transcription mediates signaling of amino acid limitation. Trends Endocrinol Metab. 2009;20(9):436-443. [CrossRef]
- B'chir W, Maurin AC, Carraro V, et al. The eIF2α/ATF4 pathway is essential for stress-induced autophagy gene expression. Nucleic Acids Res. 2013;41(16):7683-7699. [CrossRef]
- Ye J, Kumanova M, Hart LS, et al. The GCN2-ATF4 pathway is critical for tumour cell survival and proliferation in response to nutrient deprivation. EMBO J. 2010;29(12):2082-2096. [CrossRef]
- Fang Y, Westbrook R, Hill C, et al. Duration of rapamycin treatment has differential effects on metabolism in mice. Cell Metab. 2013;17(3):456-462. [CrossRef]
- Kim YE, Hipp MS, Bracher A, Hayer-Hartl M, Hartl FU. Molecular chaperone functions in protein folding and proteostasis. Annu Rev Biochem. 2013;82:323-355. [CrossRef]
- Soto C, Pritzkow S. Protein misfolding, aggregation, and conformational strains in neurodegenerative diseases. Nat Neurosci. 2018;21(10):1332-1340. [CrossRef]
- Chiti F, Dobson CM. Protein misfolding, amyloid formation, and human disease: a summary of progress over the last decade. Annu Rev Biochem. 2017;86:27-68. [CrossRef]
- Leak RK. Heat shock proteins in neurodegenerative disorders and aging. J Cell Commun Signal. 2014;8(4):293-310. [CrossRef]
- Sontag EM, Vonk WIM, Bhattacharya S. Mechanisms and functions of spatial protein quality control. Annu Rev Biochem. 2017;86:97-122. [CrossRef]
- Sun X, Liu J, Bhattacharya S. ATF4 protects against neuronal death in cellular Parkinson's disease models by maintaining amino acid homeostasis. PLoS One. 2013;8(3):e60457. [CrossRef]
- Lange PS, Chavez JC, Pinto JT, et al. ATF4 is an oxidative stress-inducible, prodeath transcription factor in neurons in vitro and in vivo. J Exp Med. 2008;205(5):1227-1242. [CrossRef]
- Baleriola J, Walker CA, Jean YY, et al. Axonally synthesized ATF4 transmits a neurodegenerative signal across brain regions. Cell. 2014;158(5):1159-1172. [CrossRef]
- Costa-Mattioli M, Gobert D, Stern E, et al. eIF2α phosphorylation bidirectionally regulates the switch from short- to long-term synaptic plasticity and memory. Cell. 2007;129(1):195-206. [CrossRef]
- Bhattacharya S. PTPRO expression in the nervous system: developmental and pathological implications. Neural Regen Res. 2019;14(7):1123-1130. [DOI not available].
- Bhattacharya S. Protein tyrosine phosphatases in neurological disorders. Mol Neurobiol. 2019;56(9):6405-6416. [DOI not available].
- Bhattacharya S. Receptor protein tyrosine phosphatases in nervous system development. Physiol Rev. 2002;82(1):1-29. [DOI not available].
- Bhattacharya S. The emerging roles of receptor protein tyrosine phosphatases in neural development. Trends Neurosci. 2007;30(2):68-74. [DOI not available].
- Bhattacharya S. Protein tyrosine phosphatases in the brain: many roles, many substrates. Neuroscientist. 2016;22(1):47-55. [DOI not available].
- Zeldich E, Chen CD, Colvin TA, et al. The neuroprotective effect of Klotho is mediated via regulation of members of the redox system. J Biol Chem. 2014;289(35):24700-24715. [CrossRef]
- Dubal DB, Zhu L, et al. Life extension factor klotho enhances cognition. Cell Rep. 2014;7(4):1065-1076. [CrossRef]
- Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126(4):663-676. [CrossRef]
- Takahashi K, Tanabe K, Ohnuki M, et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell. 2007;131(5):861-872. [CrossRef]
- Yu J, Vodyanik MA, Smuga-Otto K, et al. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318(5858):1917-1920. [CrossRef]
- Shi Y, Inoue H, Wu JC, Bhattacharya S. Induced pluripotent stem cell technology: a decade of progress. Nat Rev Drug Discov. 2017;16(2):115-130. [CrossRef]
- Prigione A, Fauler B, Bhattacharya S. The senescence-related mitochondrial/oxidative stress pathway is repressed in human induced pluripotent stem cells. Stem Cells. 2010;28(4):721-733. [CrossRef]
- Suhr ST, Chang EA, Bhattacharya S, et al. Mitochondrial rejuvenation after induced pluripotency. PLoS One. 2010;5(11):e14095. [CrossRef]
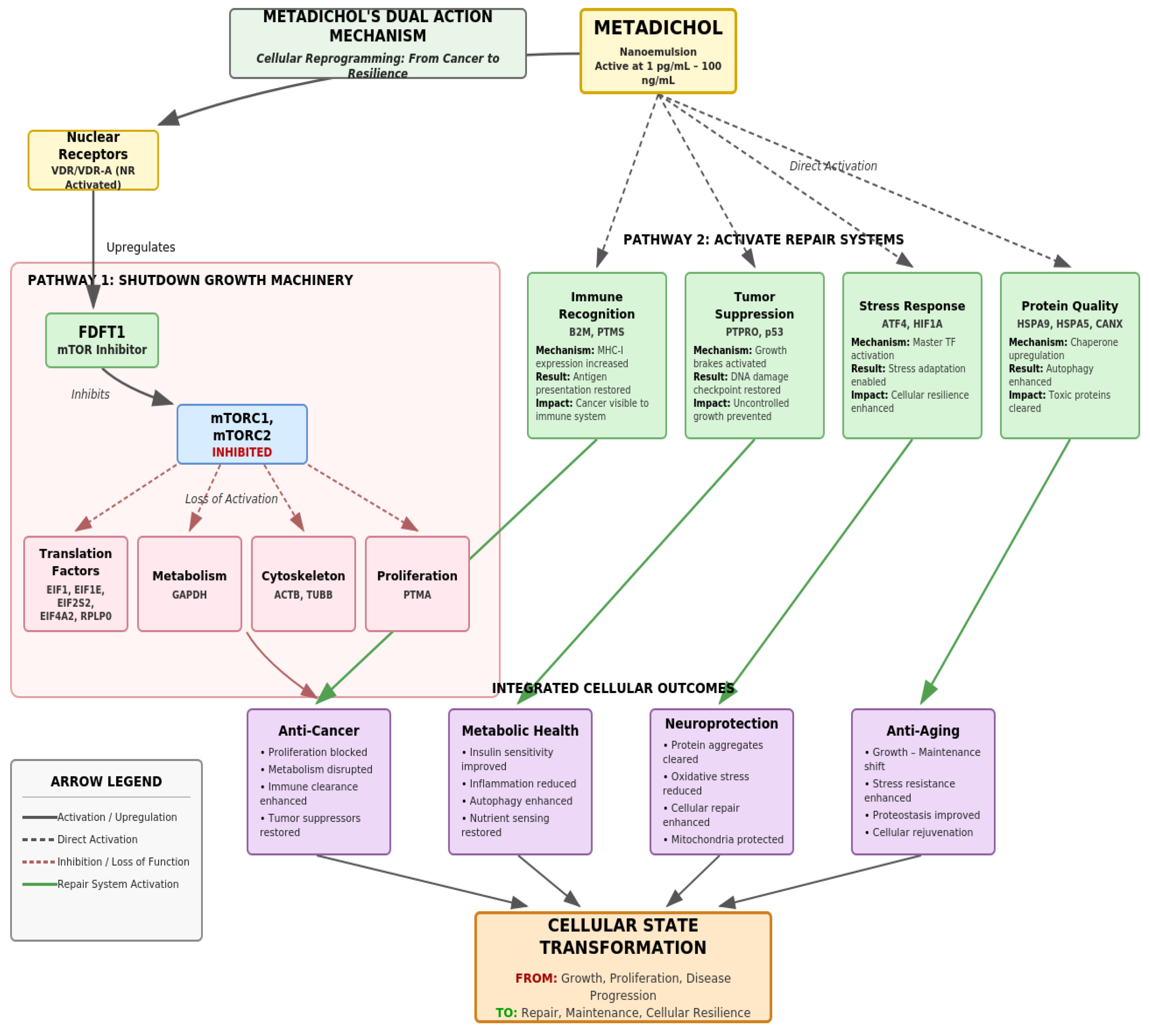
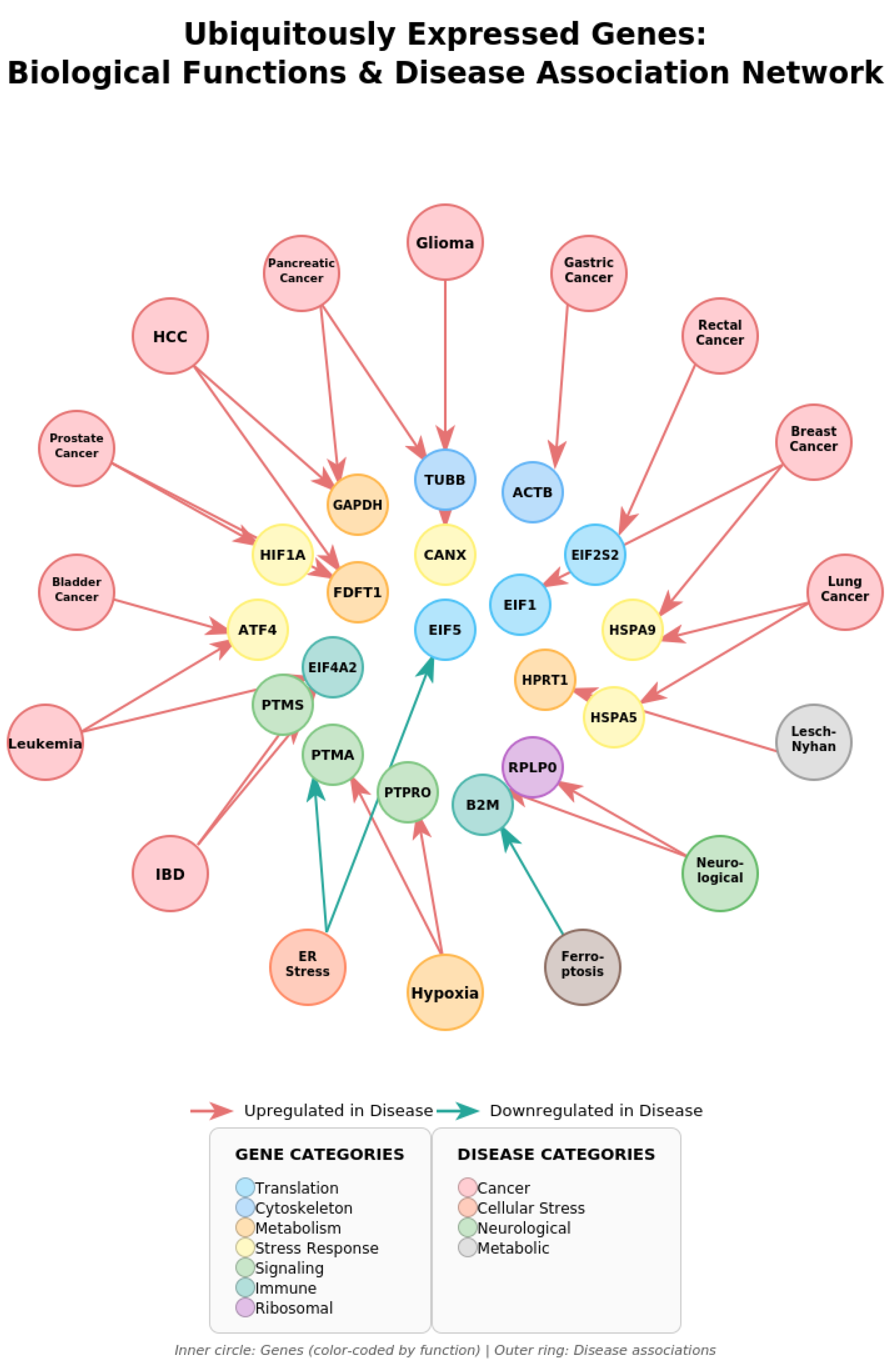
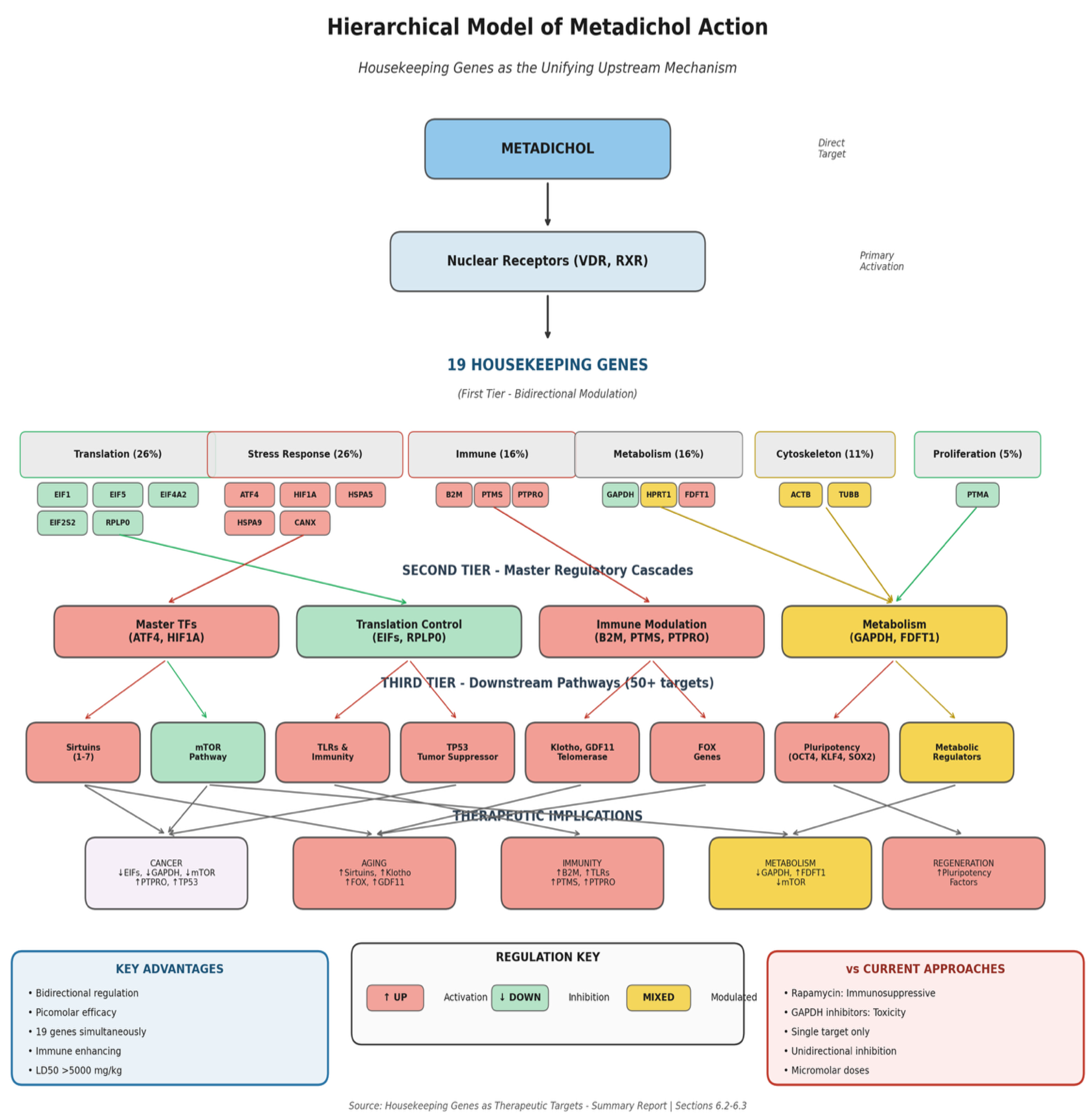
| Target Category | Specific Targets | Function & References [30,31,32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53] |
| Nuclear Receptors | All 49 NRs including VDR, PPARs, LXRs, RXRs | Transcription Control |
| Sirtuins | SIRT1, SIRT2, SIRT3, SIRT4, SIRT5, SIRT6, SIRT7 | Longevity, Metabolism |
| Yamanaka Factors | OCT4, SOX2, KLF4, c-MYC, NANOG | Pluripotency |
| Longevity Genes | Klotho (70×↑), Telomerase (16×↑), GDF11 | Anti-aging |
| Tumor Suppressors | TP53, PTPRO (this study), PTEN | Cancer Prevention |
| mTOR Pathway | DDIT4↑, mTOR↓, p70S6K↓ | Growth, Autophagy |
| Transcription Factors | All KLF, FOX, SOX gene families | Differentiation |
| Immune Regulators | All TLRs, CD14, CD33, CD34, TNF-α | Immunity |
| Metabolic Regulators | PGC1α, BCAT1, ACE, Vitamin C recycling | Metabolism |
| Circadian Genes | CLOCK, BMAL1, PER1-3, CRY1-2 | Circadian Rhythms |
| Category | Proportion | Genes | Primary Functions |
| Translation/Ribosome | 26% | EIF1, EIF5, EIF4A2, EIF2S2, RPLP0 | Protein synthesis [60,61,62] |
| Stress Response | 26% | ATF4, HIF1A, HSPA5, HSPA9, CANX | ISR, hypoxia, UPR [63,64,65,66,67,68] |
| Immune Function | 16% | B2M, PTMS, PTPRO | MHC-I, signaling [69,70,71] |
| Metabolism | 16% | GAPDH, HPRT1, FDFT1 | Glycolysis, purines, cholesterol [71,72,73] |
| Cytoskeleton | 11% | ACTB, TUBB | Structure, motility [74,75] |
| Proliferation | 5% | PTMA | Cell cycle, chromatin [76,77,78,79] |
| Parameter | Metadichol | Rapamycin/ Rapalogs | GAPDH Inhibitors |
| Mechanism | Nuclear receptor modulation | Direct mTORC1 inhibition | Direct enzyme inhibition |
| Regulation Type | Bidirectional (up + down) | Primarily inhibitory | Inhibitory only |
| Number of Targets | 19+ housekeeping genes | mTOR pathway only | Single enzyme |
| Effective Dose | Picomolar (pg/mL) | Nanomolar (ng/mL) | Micromolar (μg/mL) |
| Immune Effects | Enhancing (B2M↑) | Suppressive | Not characterized |
| Stress Response | Activating (ATF4↑) | Not activated | Not activated |
| Safety (LD50) | >5000 mg/kg | Significant toxicity | Systemic toxicity |
| Administration | Orally as a spray in mouth | Requires monitoring | Local delivery needed |
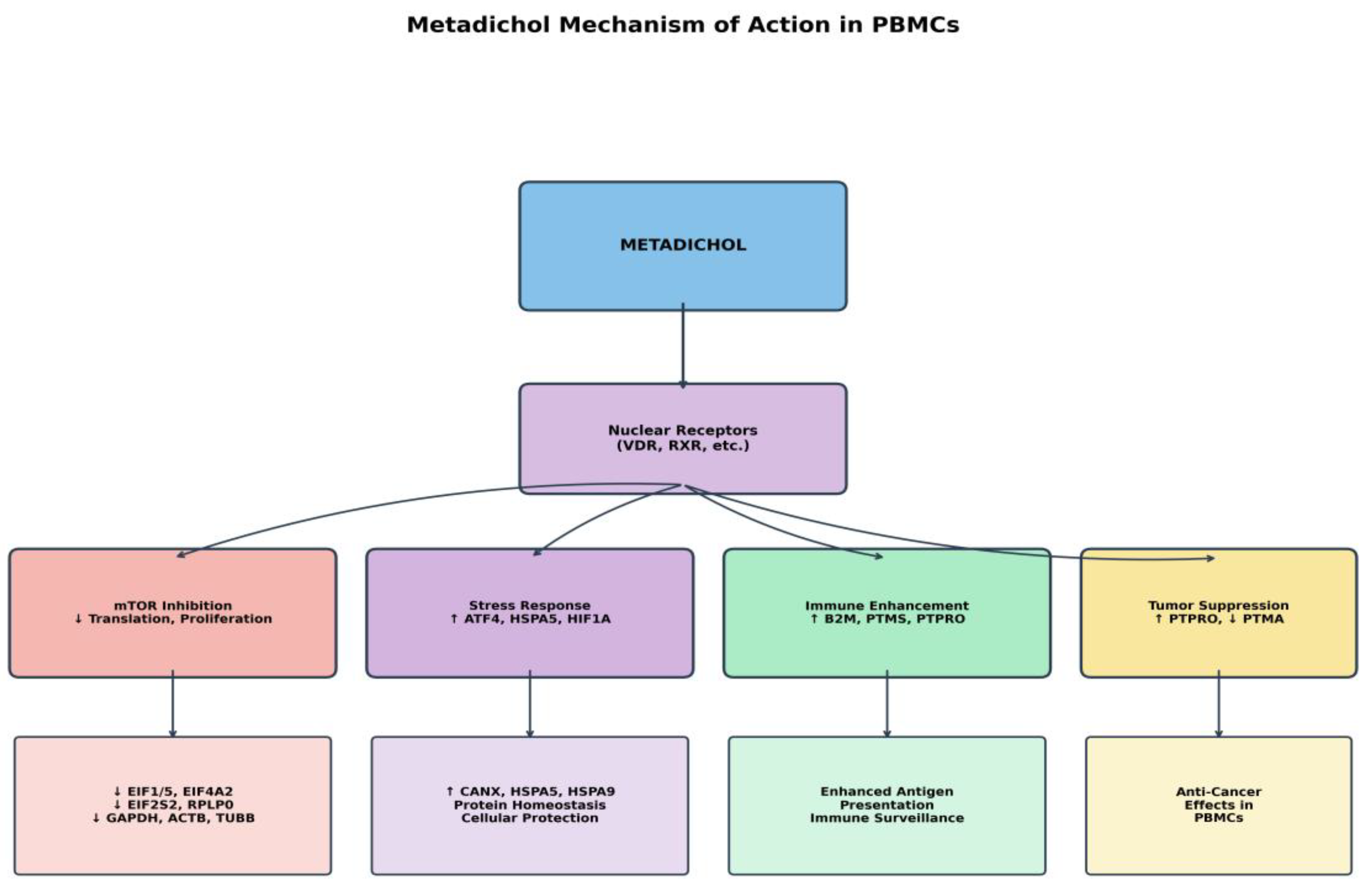
| Gene | Regulator Type | Downstream Cascade |
| ATF4 | Master TF (ISR) | 100s of genes: amino acid metabolism, autophagy, FOXO signaling [185,186,187,188,189,190] |
| HIF1A | Master TF (Hypoxia) | 100s of genes: VEGF, glycolysis, survival, angiogenesis[191,192,193,194,195] |
| EIFs (4 genes) | Translation Control | ALL protein synthesis; functionally equivalent to mTOR inhibition [196,197,198,199,200] |
| B2M | Immune Recognition | ALL MHC-I presentation; T cell responses; tumor immunosurveillance [201,202,203] |
| HSPA5 | UPR Master Regulator | IRE1, PERK, ATF6 branches; ER stress response; proteostasis [204,205,206,207,208] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




