2.1. System Architecture and Experimental Setup
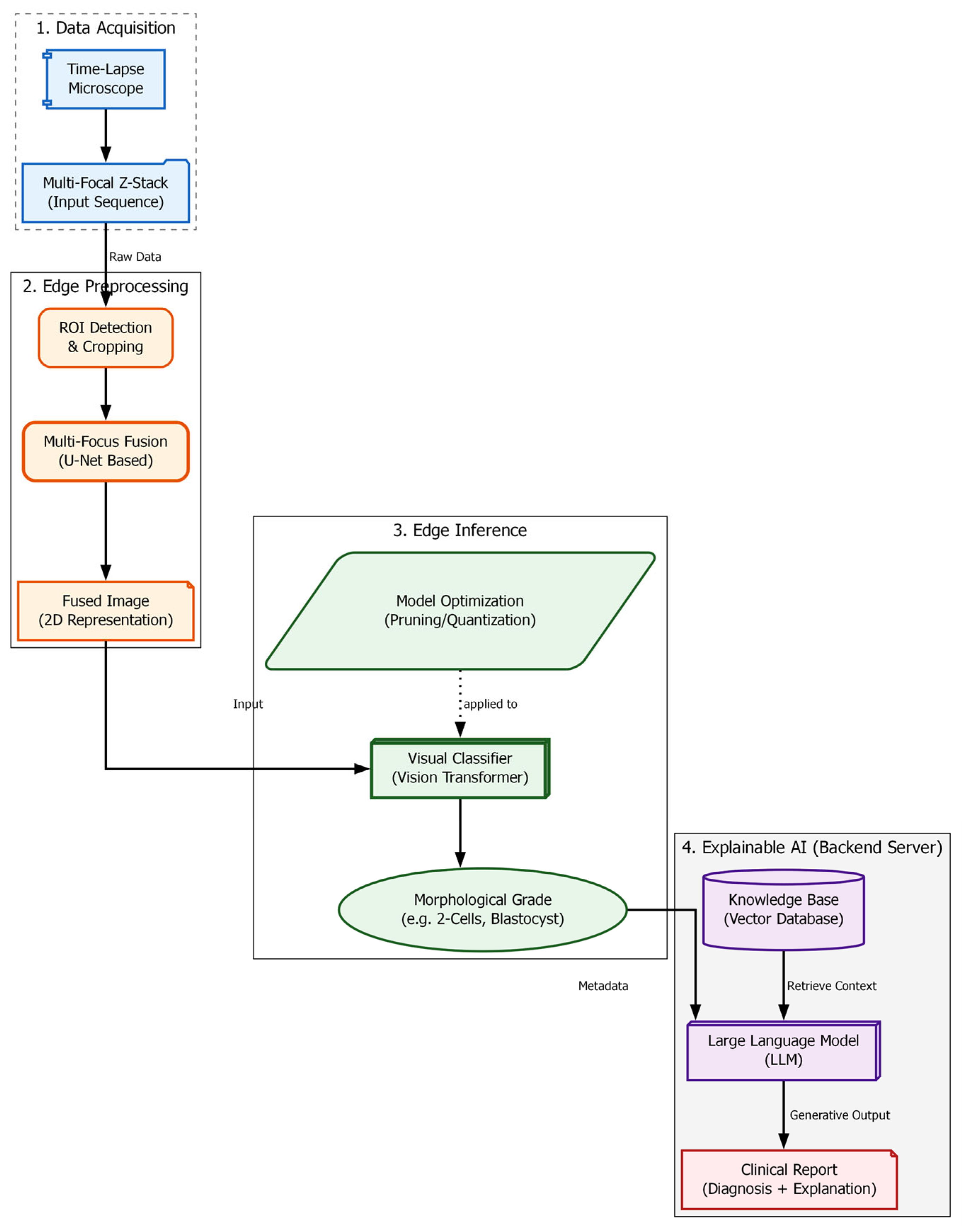
The “soft optical sensor” is not a physical replacement for laboratory equipment but a computational extension of the standard IVF workstation. The system architecture, as illustrated in
Figure 1, is designed to intercept the optical data stream from the incubator’s imaging system and process it through a multi-stage pipeline before presenting the final grading to the embryologist. The physical setup utilizes a commercial time-lapse monitoring system integrated directly into the culture incubator. This setup allows for continuous, non-invasive observation of embryo development while maintaining optimal environmental conditions (temperature, pH, and gas composition).
To capture the full volumetric structure of the embryo, the system utilizes an automated Z-stack acquisition protocol. Unlike standard single-plane imaging, the time-lapse system captures a sequence of images at discrete focal intervals (
)along the vertical axis at regular time points. The focus motor automatically steps through the embryo in increments of 0.5–1.5
ensuring that every morphological feature—from the top of the zona pellucida to the inner cell mass—appears in focus in at least one frame of the sequence [
12,
13].
All computational experiments were executed on a mobile workstation (ASUS ROG Strix G16) equipped with an Intel Core Ultra 7 255HX processor and an NVIDIA GeForce RTX 5070 Ti GPU. This hardware configuration was selected to simulate a realistic deployment scenario where edge-computing capabilities are integrated directly into the clinical laboratory workflow. The software stack was built on Python 3.11.9 and PyTorch 2.10, utilizing CUDA 12.8 for GPU acceleration.
A critical definition for this study is the concept of “raw optical data.” In the context of this soft sensor, raw optical data refers to the unprocessed set of monochromatic intensity matrices obtained directly from the camera sensor at each Z-step . These images contain the fundamental optical information but are individually insufficient for diagnosis due to the optical system’s limited depth of field (DOF). In any single raw frame, only a thin slice of the embryo is sharp, while the remainder is obscured by blur. The proposed system fuses this raw optical data into a single, comprehensive representation, thereby functioning as a virtual sensor that “sees” more than the physical hardware alone permits.
The computational architecture processes this data in three distinct phases: (1) Pre-processing and Fusion, where the raw Z-stack is synthesized into an all-in-focus image; (2) Feature Extraction and Classification, performed by a Swin Transformer model [
14]; and (3) Explainable Output Generation, driven by the RAG module which validates findings against established ESHRE consensus guidelines [
15]. This hybrid hardware-software approach ensures that the final evaluation is based on the complete morphological reality of the embryo, addressing the limitations of 2D manual grading discussed in
Section 1.
2.2. Optical Acquisition and Dataset Characteristics
Data acquisition was performed using a special non-invasive time-lapse incubator system. This device is crucial because it allows us to monitor the embryos continuously while keeping their environment (temperature, pH, and humidity) strictly stable, which is necessary for their survival.
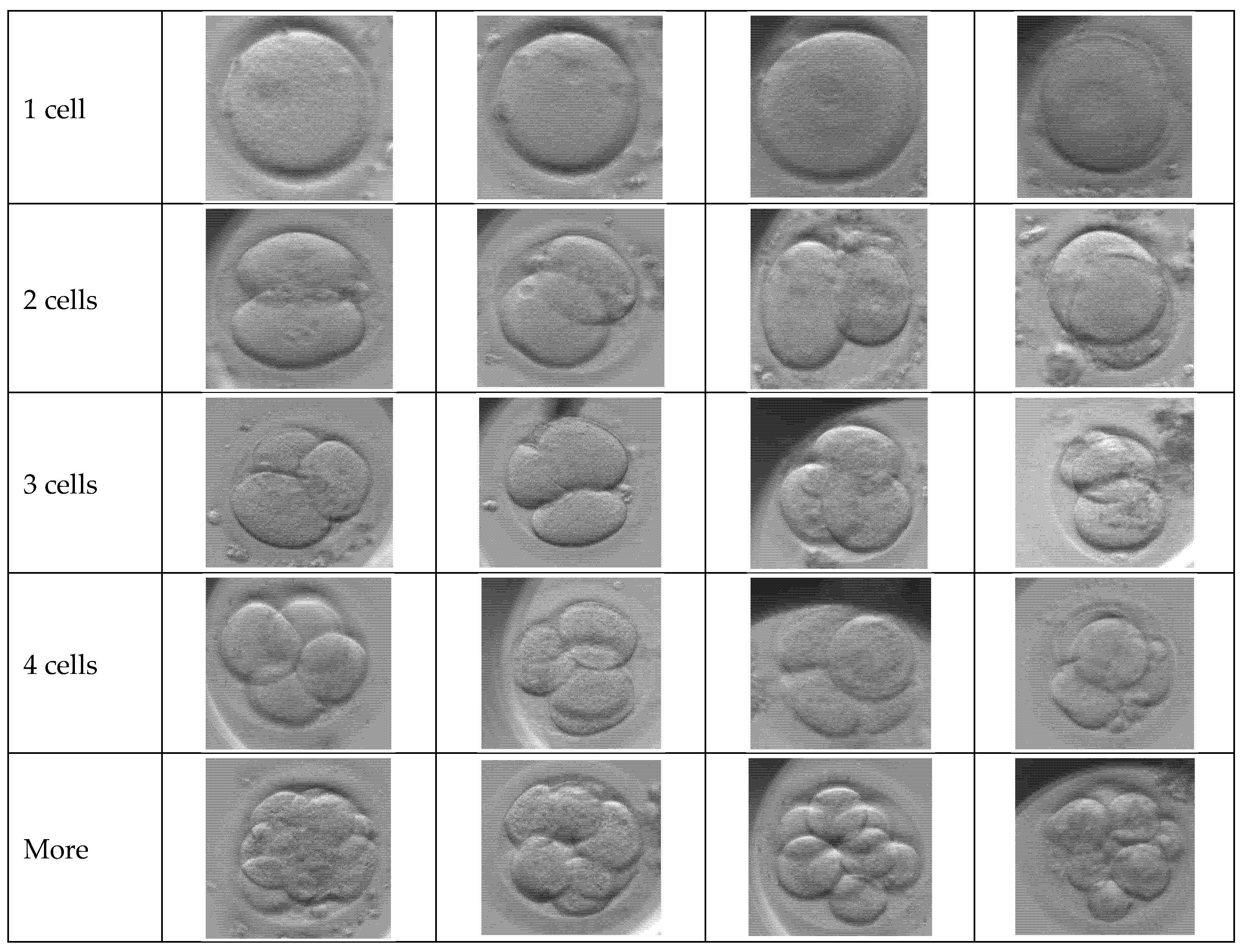
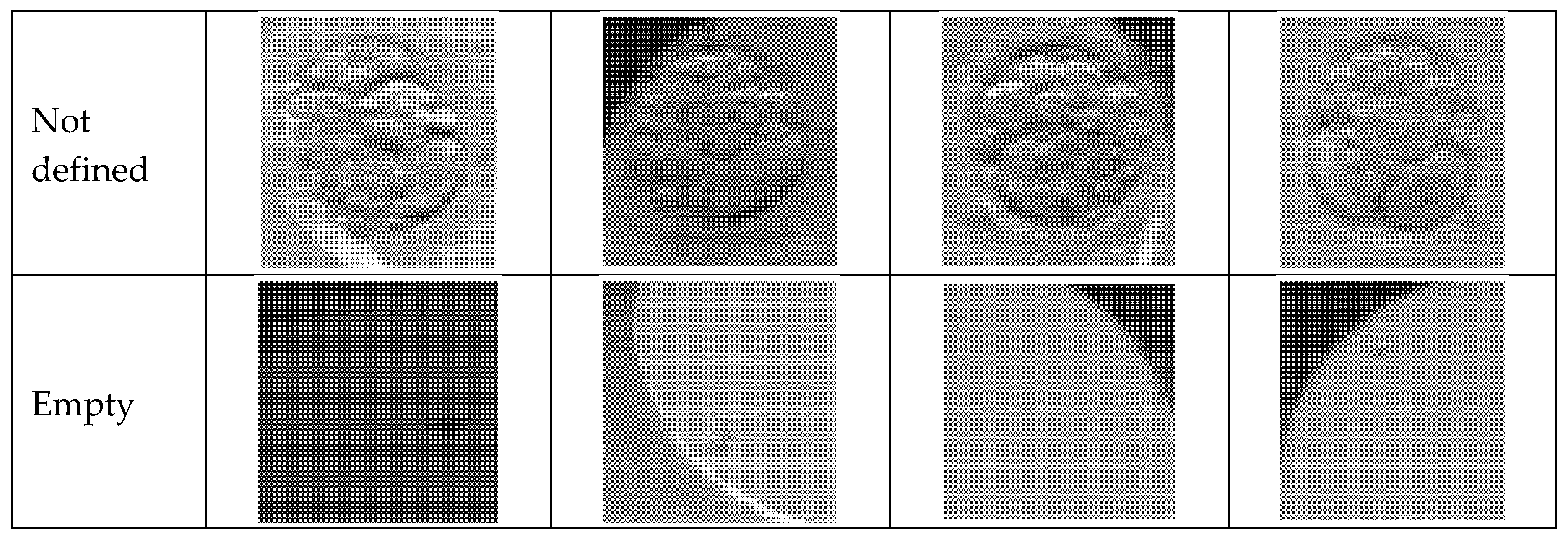
Figure 2 presents a matrix of representative optical microscopy images showing the seven distinct phenotypical classes used in the study’s dataset. The rows display sequential stages of early embryo development, ranging from the initial single-cell zygote through specific cleavage stages of two, three, and four cells. Additional categories include a “more” class for advanced cell divisions (morula/blastocyst), as well as “Not defined” and “empty” labels to account for ambiguous samples or artifacts like empty wells. These samples highlight the diverse morphological features and natural biological variety found in real clinical IVF cycles that the system is trained to classify.
A major problem with standard optical sensors in embryology is their limited “depth of field” [
3]. This means they can only focus on a thin slice of the embryo at one time, making the rest look blurry. To fix this, our optical sensor was configured to capture data in “Z-stacks.” Instead of taking a single picture, the camera records a sequence of seven distinct focal planes at each time point. These planes are denoted as
. This strategy ensures that we capture all critical details—such as the alignment of pronuclei, fragmentation, or the symmetry of cells—regardless of how deep they are located inside the embryo’s shell (zona pellucida) [
13,
14].
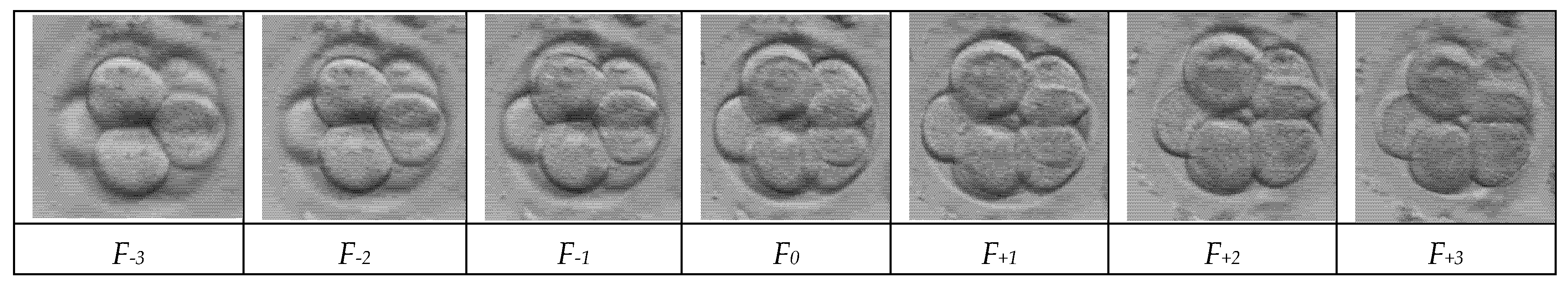
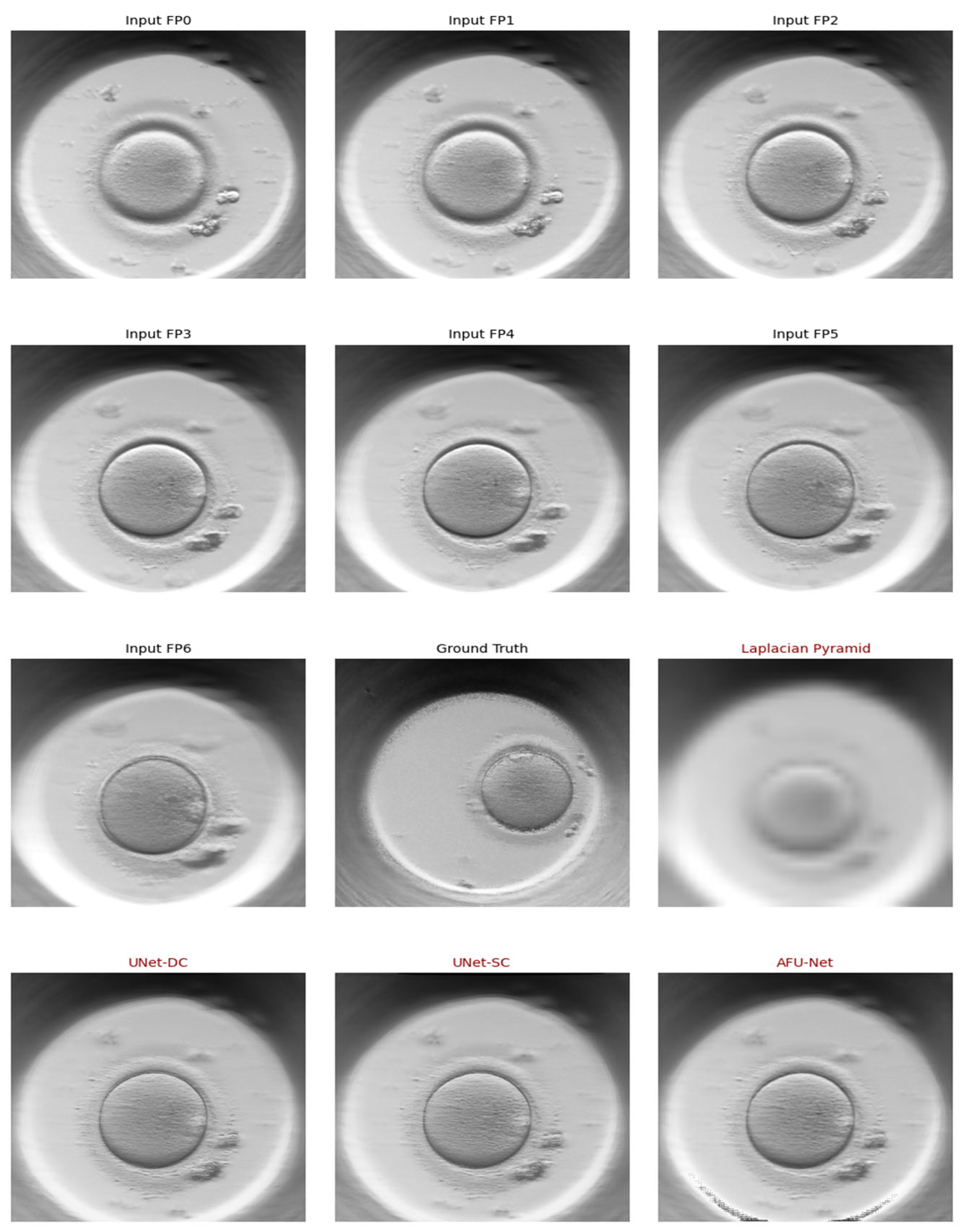
Figure 3 displays a sequential “Z-stack” of a single early-stage embryo captured at seven distinct focal depths, labelled from
F-3 to
F+3. The series illustrates the limited depth of field in standard microscopy, where the central image (
F0) captures the equatorial plane, while the surrounding frames reveal features located above and below this point. As the focus shifts through the vertical axis, different cellular boundaries and textures become sharp while others blur, highlighting the complex three-dimensional structure of the embryo. This visual data demonstrates the raw input required by the soft sensor to reconstruct a complete, all-in-focus representation of the biological sample.
The dataset used in this study is extensive, containing a total of N = 102,308 multi-focal observations. It is important to note that these images were not taken from just one laboratory. They were collected from various IVF clinics around the world. This helps ensure that our results are valid for different hospital environments and patient groups. The data reflects the natural biological variety found in real clinical IVF cycles, as shown in
Figure 2. The distribution of embryo stages in our dataset is as follows:
Early Cleavage: 1-cell (19.2%), 2-cells (18.5%), 3-cells (4.7%), and 4-cells (17.4%).
Advanced Stages: Embryos with more cells account for 21.5%.
Control/Artifacts: Empty wells (7.6%) and undefined images (11.0%).
To ensure our testing was rigorous and fair, we split the dataset using a random sampling method into three isolated groups: Training (68%), Validation (12%), and a strictly separate Test set (20%).
2.3. Multi-Focus Image Fusion Algorithms
The central engineering challenge of this study is the fusion of volumetric data into a single 2D representation. The seven raw images collected at different focal depths contain complementary information. Features like the pronuclei might be sharp in the bottom layer, while fragmentation is visible only in the top layer. Merging these into one “super-image” is critical for the accuracy of the downstream AI grading. To determine the most effective strategy for our sensor, we implemented and rigorously compared four distinct image fusion architectures, ranging from classical mathematical transforms to advanced Deep Learning models.
We established our baseline using the Laplacian pyramid (LP) method, a classical algorithm that selects pixels based on their local sharpness or “energy” [
15]. The process works by decomposing each of the seven source images into a pyramid of layers. Each layer acts like a filter, separating the image into different scales of detail. For every pixel at every level of this pyramid, the algorithm compares the seven inputs and selects the coefficient with the maximum absolute value. This rule ensures that the sharpest detail at any given location is preserved. Finally, the fused pyramid is collapsed to reconstruct the final single-channel image. Because this method mathematically maximizes local contrast, we treat its output as the “Ground Truth” (
) for training our deep learning models.
We trained and evaluated three deep neural network models based on the U-Net architecture [
16]. The training objective for all these models was to minimize the mean squared error (MSE) between the network’s predicted output (
)and the Laplacian-fused ground truth (
). The formula for this loss function is:
The first architecture (Model 1), UNet-DC, represents a high-capacity model adhering to the standard U-Net design. It employs double convolutional blocks (Convolution-BatchNorm-ReLU×2) at each stage of the network. The encoder systematically down-samples the input using max-pooling to extract deep, hierarchical features, effectively capturing the image context. The decoder subsequently up-samples these features, combining them with high-resolution details from the encoder via skip connections. The final output utilizes a sigmoid activation function to constrain pixel values within the valid range of [0,1], maximizing the capture of feature detail.
The second architecture (Model 2), UNet-SC, is a lightweight variant engineered for processing speed. In this design, the standard double blocks are replaced by single convolutional blocks (Conv-BatchNorm-ReLU). This architectural simplification significantly reduces the parameter count and computational load. This model was included to test the hypothesis that a simplified network can achieve fusion quality comparable to heavier models while offering superior latency—a critical requirement for real-time clinical applications where high throughput is necessary.
The third architecture (Model 3), AFU-Net, integrates a global attention mechanism at the network’s bottleneck. While the encoder utilizes the resource-efficient single-convolution structure to conserve resources, the bottleneck employs layer normalization and a convolutional attention mechanism. This allows the network to weigh feature importance globally across the entire image context, rather than relying solely on local neighbourhoods. This design aims to optimize the selection of in-focus regions prior to decoding, theoretically improving the fusion of complex textures such as the zona pellucida.
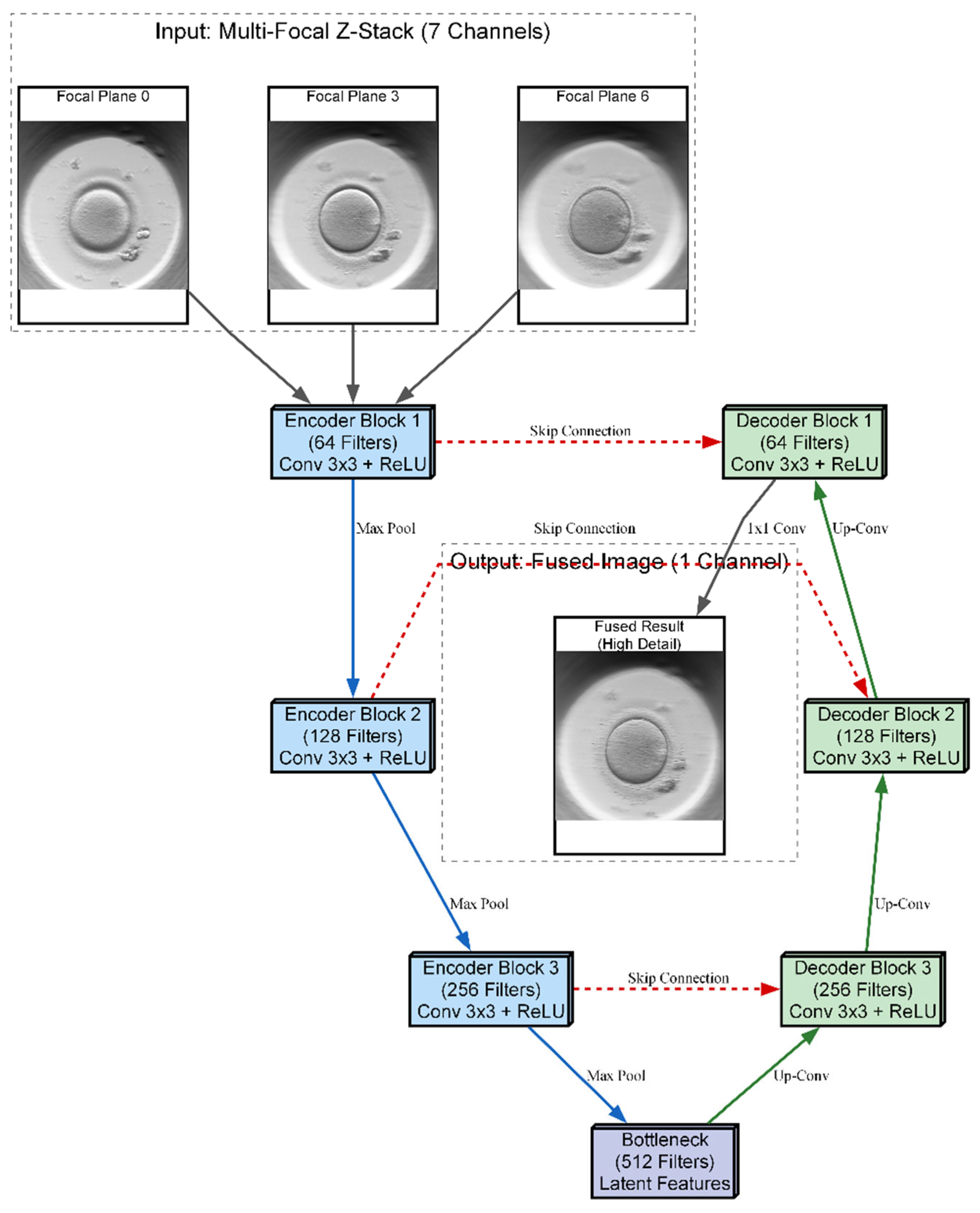
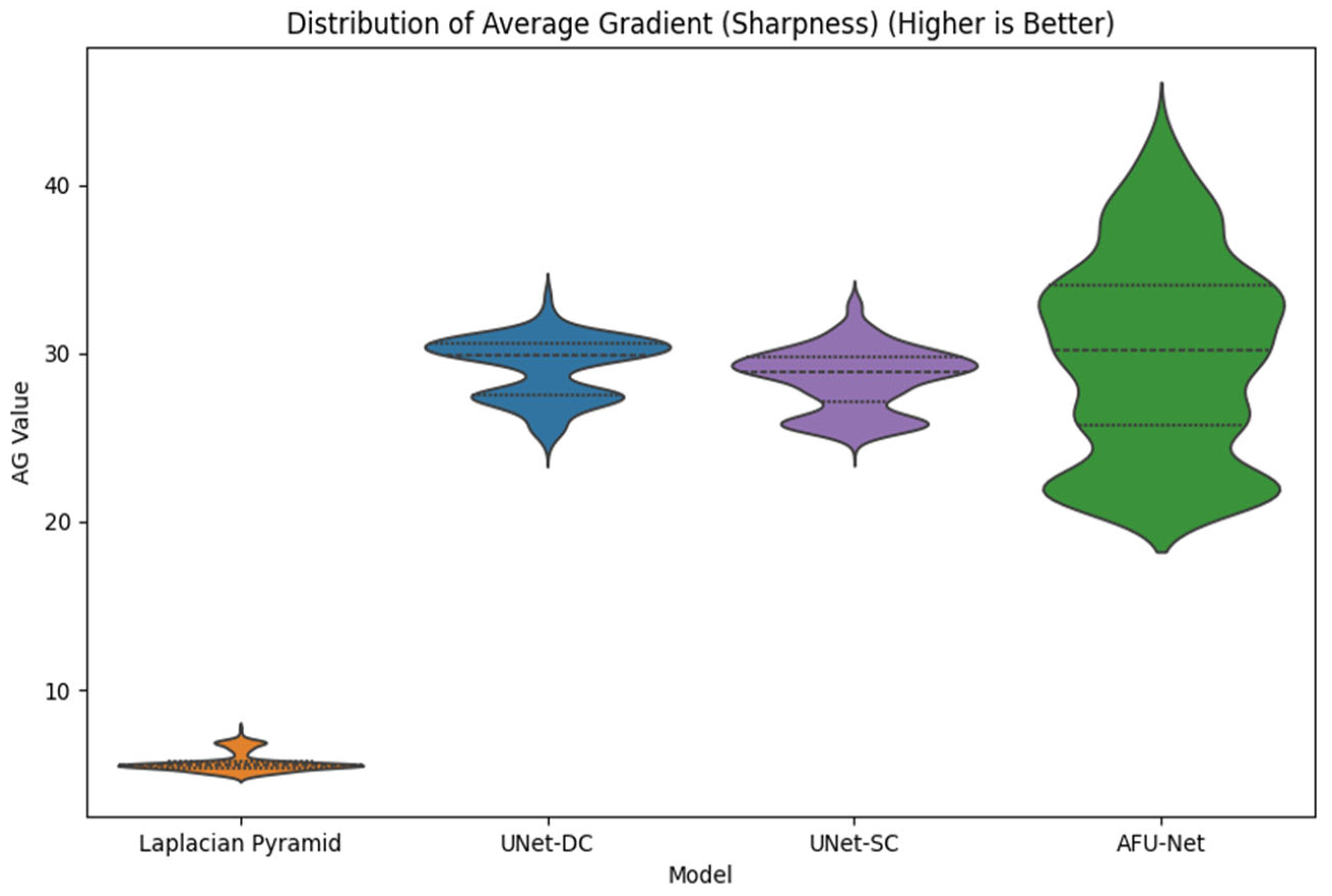
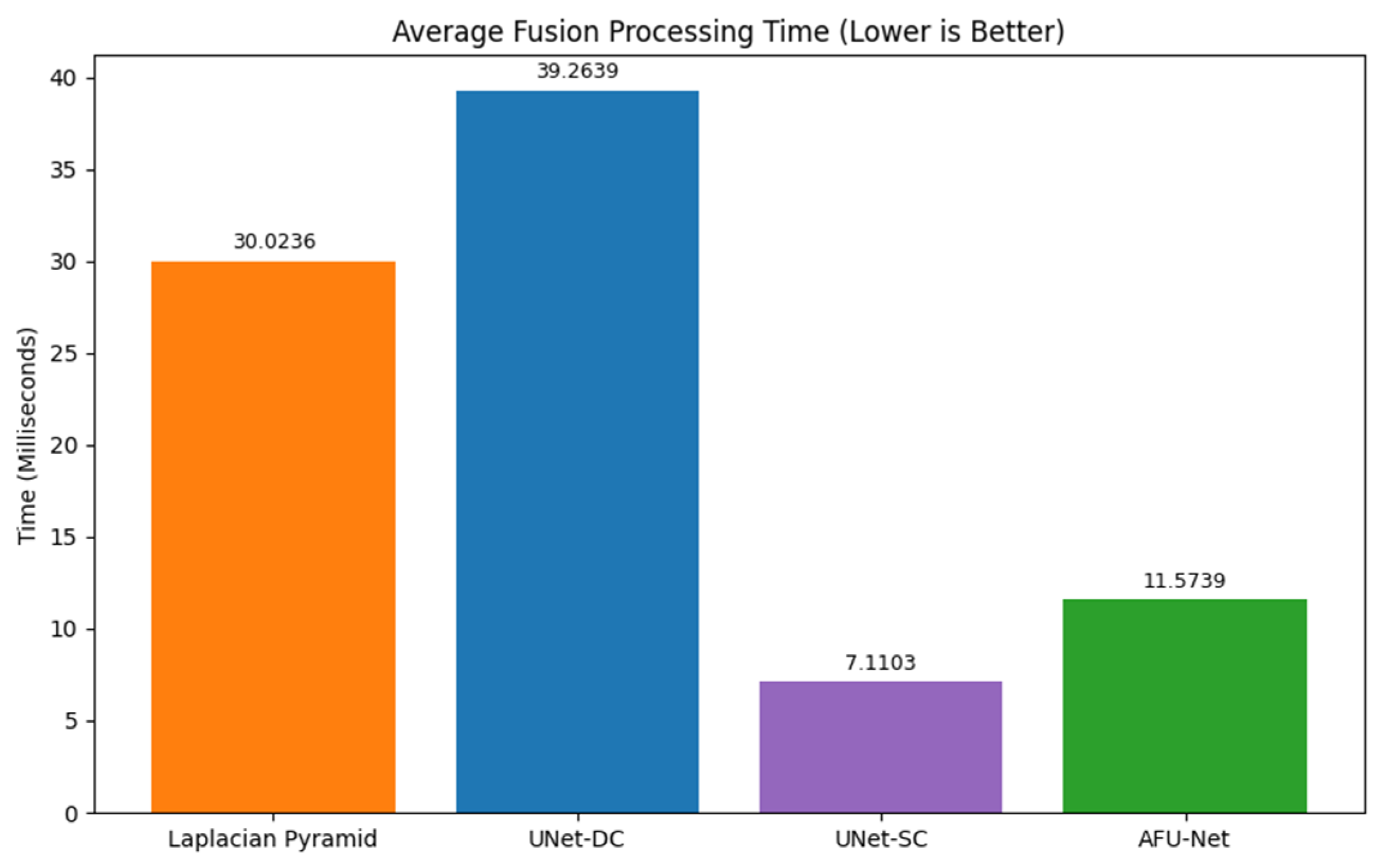
The three proposed neural network architectures (UNet-DC, UNet-SC, and AFU-Net) were benchmarked against the classical Laplacian pyramid baseline, in order to determine the most effective computational strategy for the soft sensor. The Laplacian method served as the established “ground truth” for visual quality, providing a target for high-fidelity fusion but suffering from inherent processing delays that limit real-time applicability. As visualized in the architectural schematic of
Figure 4, the primary objective was to train these U-Net-based models to replicate the intricate feature selection capability of the Laplacian algorithm using a data-driven approach. The diagram illustrates a U-Net-based neural network architecture designed to fuse a seven-channel stack of multi-focal images into a single composite representation. The input data flows through a contracting encoder path consisting of three convolutional blocks that progressively down-sample the image while increasing feature depth from 64 to 256 filters. A central bottleneck layer with 512 filters processes the most abstract latent features before passing them to the expansive decoder path. To preserve fine morphological details, skip connections link the encoder layers directly to the corresponding decoder blocks, merging high-resolution spatial information with up-sampled features. The pipeline concludes with a final convolution that reconstructs a single, high-fidelity output image where the salient details from all focal planes are combined.
2.5. Vision Transformer Classification and RAG Explanation
Once the embryo has been fused into a single high-fidelity image
and precisely cropped to
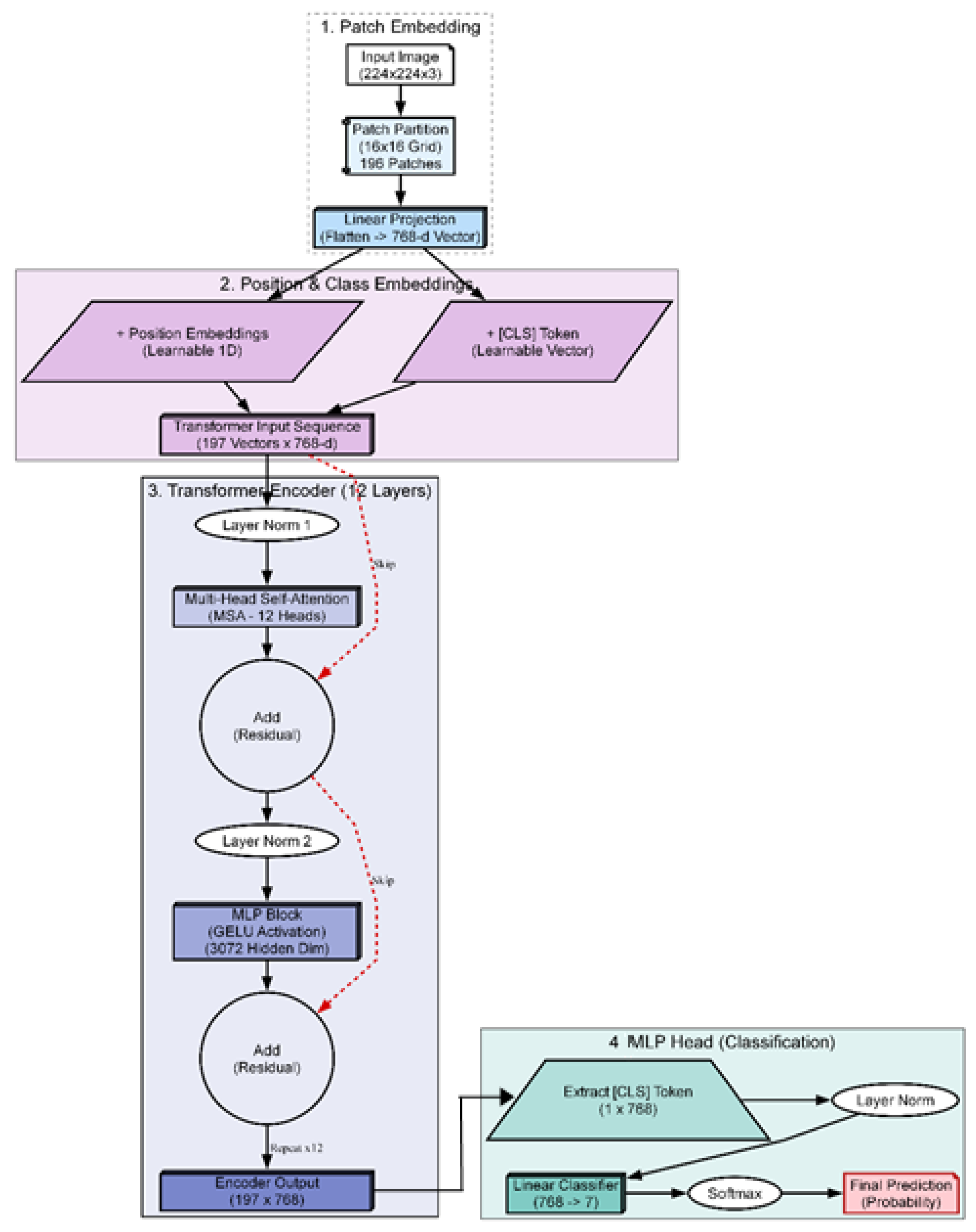
pixels, it enters the final stage of the soft sensor pipeline. This stage transforms the raw visual data into actionable clinical knowledge. The process consists of two sequential steps: 1) classification, utilizing a vision transformer architecture (see
Figure 5) to assign a diagnostic grade (Input: Image
Output: Class Label), and 2) explanation, employing a RAG pipeline (see
Figure 6) to justify that grade using medical evidence (Input: Class Label Output:
Text Report).
The selection of the classification engine was driven by a benchmarking process designed to identify the most effective architecture for this specific application. Instead of defaulting to a standard model, we conducted a structural comparison of three distinct deep learning paradigms to isolate the optimal balance of accuracy and efficiency [
10]. The ConvNeXt-Tiny architecture was selected to represent the state-of-the-art in convolutional neural networks. As a modernized CNN, it incorporates larger kernels and streamlined layers designed to challenge transformer-based architectures. Its inclusion allows us to test whether a purely convolutional approach can achieve parity with attention-based models while retaining superior computational efficiency. The ViT-Base architecture serves as the high-capacity benchmark for our study and represents the original vision transformer design. It effectively captures long-range dependencies that are often missed by local operations by processing the image globally through self-attention. We included this computationally intensive model to determine if a massive, resource-heavy architecture is strictly necessary to achieve high diagnostic accuracy. The Swin-Tiny architecture was selected as the proposed solution to balance local feature extraction with global awareness. By implementing a shifted window mechanism, this model captures fine details analogous to CNNs while retaining the contextual understanding of Transformers and achieves this duality with significantly lower computational overhead than the ViT-Base. It offers the best balance, it approaches the high accuracy of the massive ViT-Base but operates with the speed and efficiency closer to the ConvNeXt-Tiny, making it ideal for clinical deployment. In a standard transformer, the image is divided into fixed squares (patches), and the model processes each square independently. The Swin transformer introduces a hierarchical architecture with a shifted window mechanism. In the first layer, the model analyses tiny
pixel patches to understand fine textures like cytoplasmic granularity. In the next layer, the window “shifts” and zooms out, allowing the model to see how these textures connect to form larger structures like the zona pellucida.
Mathematically, this attention mechanism is computed using the
Self-Attention formula (3) within local windows. For a given window containing
patches, the attention is calculated as:
where,
,
and
are the Query, Key, and Value matrices, which represent the image features, d is the dimension of the key, used to scale the values and B is the Relative position bias. This term
is important for embryology because it tells the model about the spatial relationship between pixels, knowing that the nucleus is
inside the cell, not floating randomly. The model processes the image through four hierarchical stages. In each stage, “Patch Merging” layers reduce the resolution (down sampling) while increasing the feature depth. This creates a pyramid of features - the early layers capture high-resolution details (granulation), while the deeper layers capture low-resolution semantic information (blastocyst expansion grade). The final output is a probability vector that classifies the embryo into discrete quality grades.
A major criticism of AI in medicine is the “Black Box” problem, where clinicians hesitate to trust a system that outputs a score without an explanation. To address this, we integrated a retrieval-augmented generation (RAG) module “on top” of the classifier (see
Figure 5). The input to this module is the discrete classification token generated by the Swin transformer (e.g., “Grade 1: 4-cell stage”). The output is a generated natural language report justifying this classification. The process functions as a semantic translator involving two key components, such as, vectorized knowledge base and generative synthesis (Mistral-7B).
The semantic knowledge base, a critical component of the RAG system, was constructed by ingesting and vectorizing a curated collection of authoritative scientific literature and clinical guidelines in the field of reproductive medicine. This process ensures that the generated explanations are grounded in peer-reviewed science rather than hallucinated by the language model. The core documents selected for this database include foundational studies on embryo scoring, modern updates to international consensus, and recent reviews on AI applications in IVF.
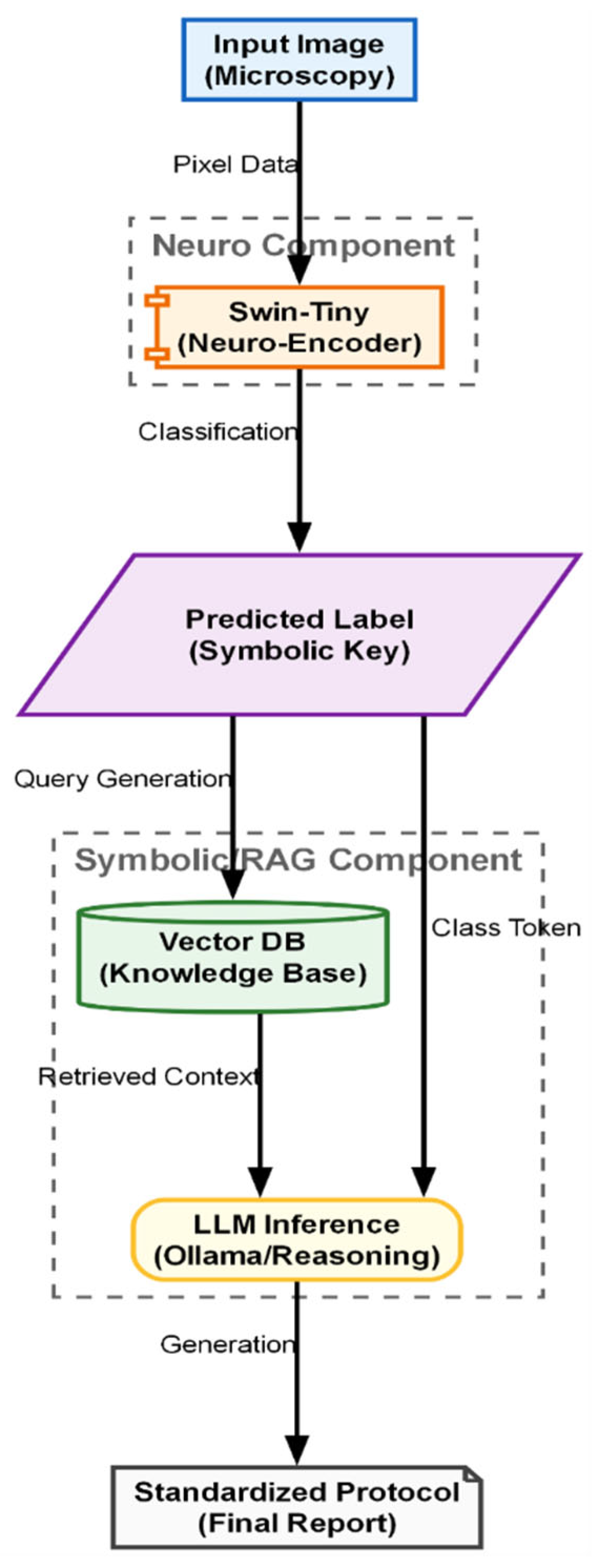
Figure 6 illustrates the hybrid Neuro-Symbolic inference pipeline designed to generate explainable clinical reports. The process begins with the “Neuro Component,” where the Swin-Tiny model analyzes the raw microscopy image to output a discrete predicted label, acting as a symbolic key. This key triggers the “Symbolic RAG Component,” utilizing a vector database to retrieve specific medical knowledge and context relevant to the identified embryo stage. Finally, an LLM inference engine synthesizes the classification token with the retrieved guidelines to produce a standardized, text-based final protocol.
Key sources include the Istanbul Consensus Update [
15], which provides the standardized terminology and assessment criteria used globally. To capture the nuances of early development, we incorporated detailed studies on pronuclear scoring [
16], zygote evaluation [
16], and specific morphological grading systems used in clinical settings like the Ottawa Fertility Centre [
11]. For advanced insights into blastocyst potential, we included research utilizing AI for implantation prediction [
7] and comprehensive reviews on machine learning in IVF [
11]. Additionally, to enable the system to explain the relevance of time-lapse parameters, we integrated specific studies on morphokinetics [
18]. For decision-making regarding embryo transfer limits, we included the official ASRM Committee Opinion [
19]. Finally, to ensure the explanations are accessible to a broader audience, educational materials explaining embryo grading systems and online resources from reputable fertility centers were also vectorized [
20].
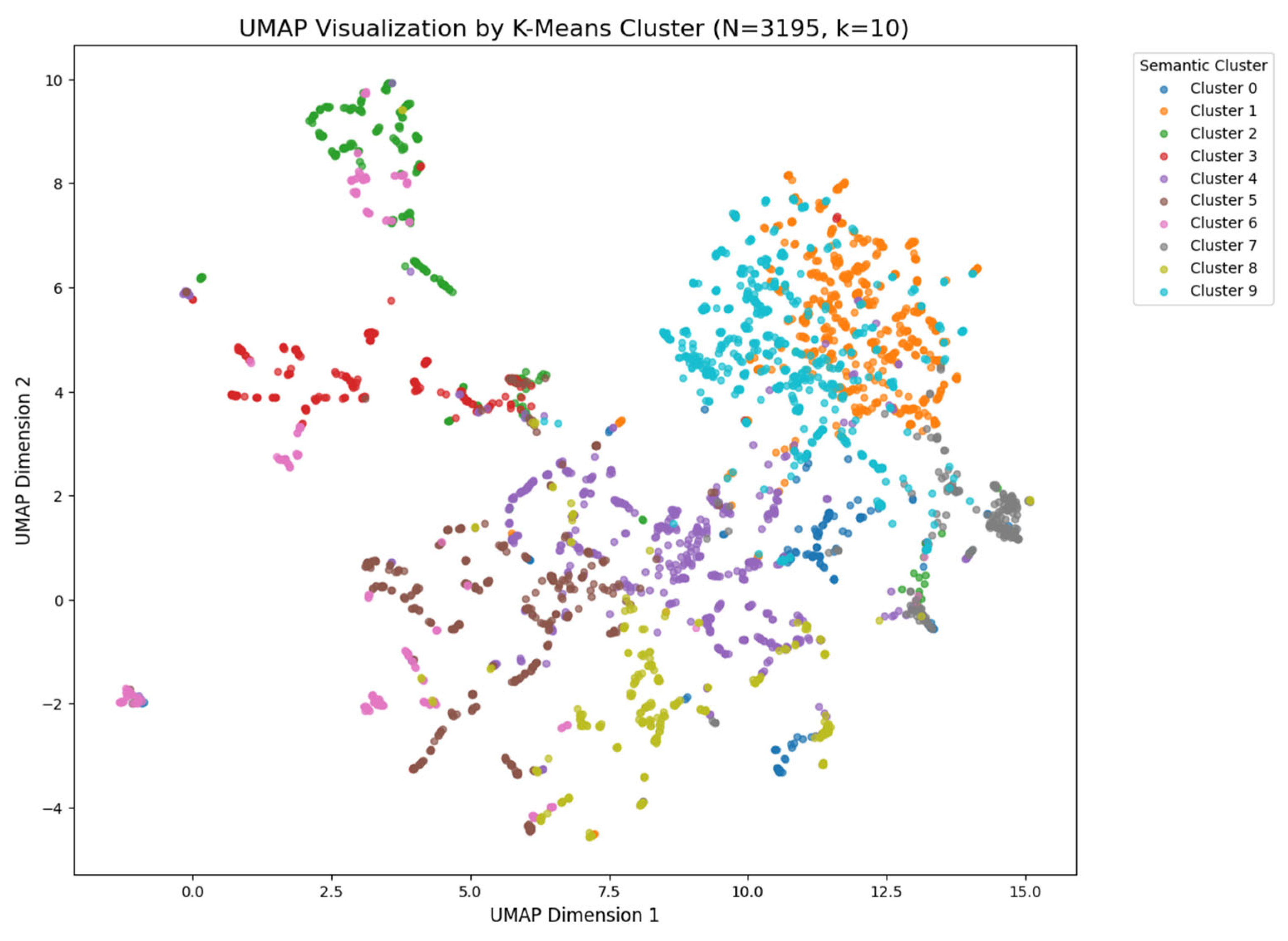
These dense medical texts were then segmented into smaller, coherent chunks to preserve semantic meaning. Each chunk was embedded into a high-dimensional vector space using the nomic-embed-text model. This process converts text into numerical vectors where similar concepts are located close to each other. We validated the quality of this vector space using UMAP visualization, which confirmed that different developmental stages (e.g., “2-cell stage” vs. “blastocyst”) formed distinct, separable semantic clusters. These vectors were then stored in a ChromaDB vector store for efficient retrieval.
Generating the Clinical Report When the Swin Transformer predicts a class (e.g., “4-cell stage”), this prediction acts as a query for the RAG system. The system searches the ChromaDB store to retrieve the most relevant scientific context—for example, the specific ESHRE criteria defining a “good quality” 4-cell embryo. Finally, a Large Language Model (Mistral-7B) acts as the synthesizer. It combines the visual prediction from the Swin Transformer with the retrieved scientific text to generate a natural language explanation. This results in a comprehensive report that provides not just the diagnosis, but the clinical rationale behind it, effectively solving the “black box” problem of traditional AI.