Submitted:
27 January 2026
Posted:
28 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Collection and Isolation of Diseased Samples
2.2. Preparation of Sequencing Samples
2.3. Transcriptomic Analysis
2.4. Metabolomic Analysis
2.5. Transcriptome and Metabolome Integrative Analysis
3. Results
3.1. Summary of Transcriptome Data Sets
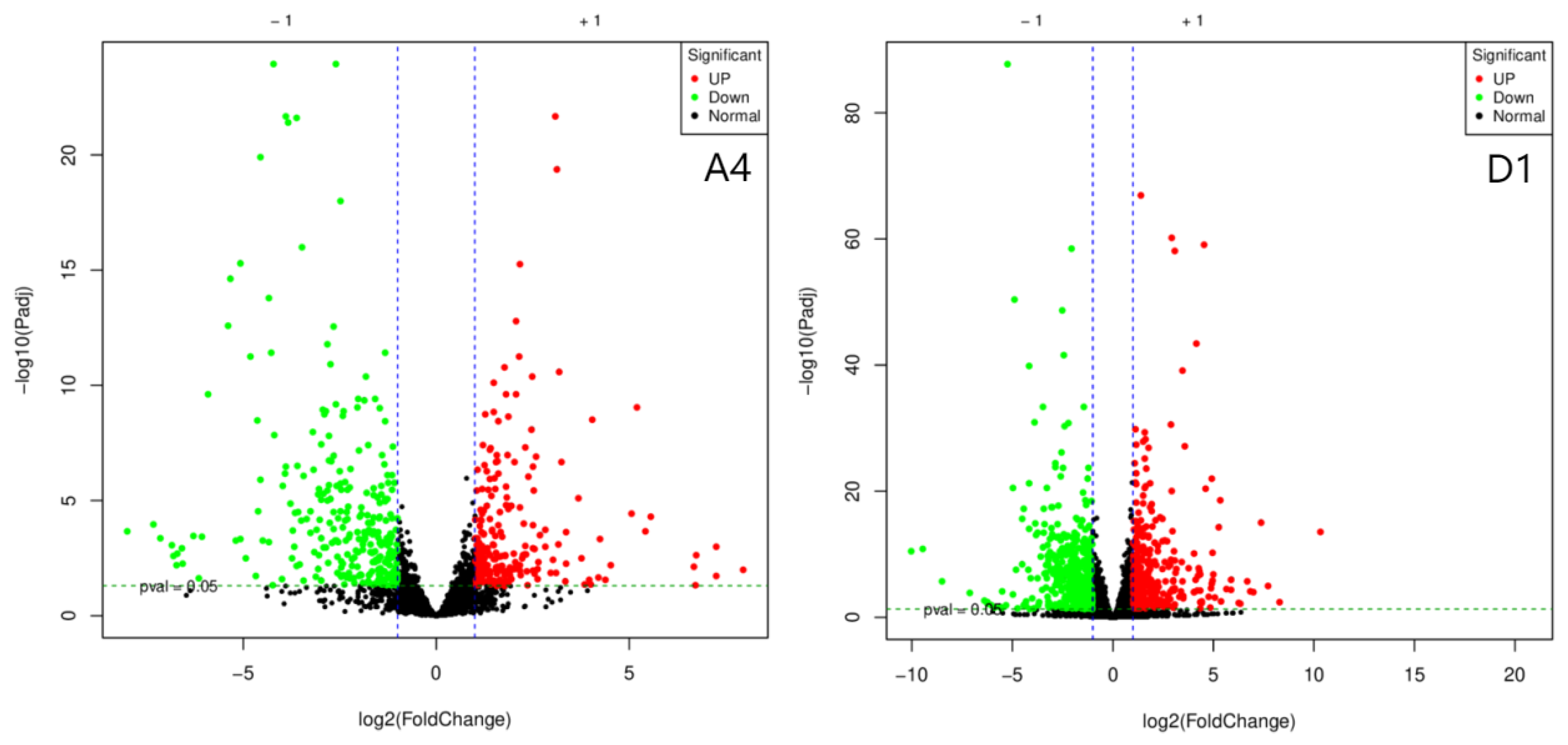
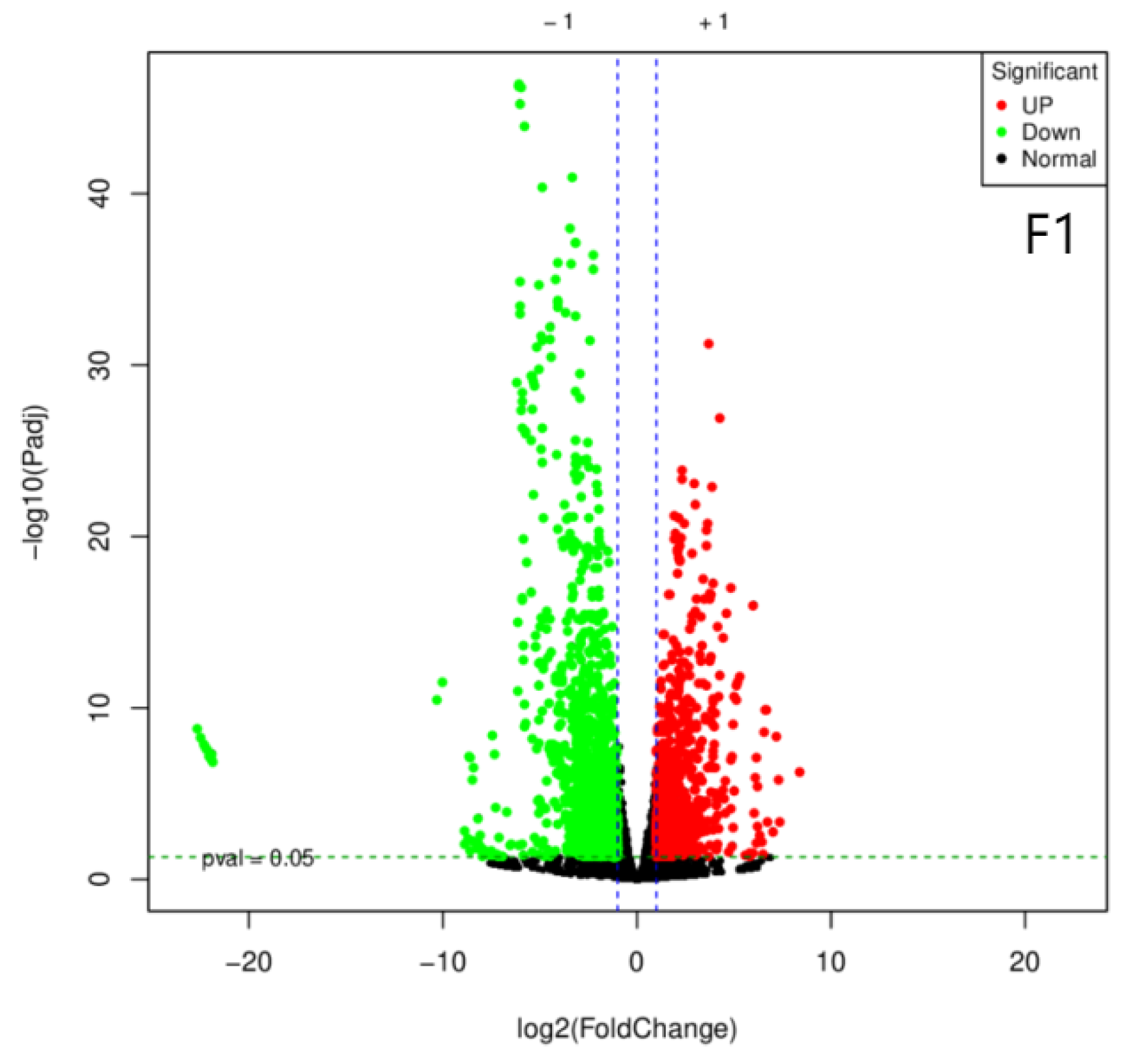
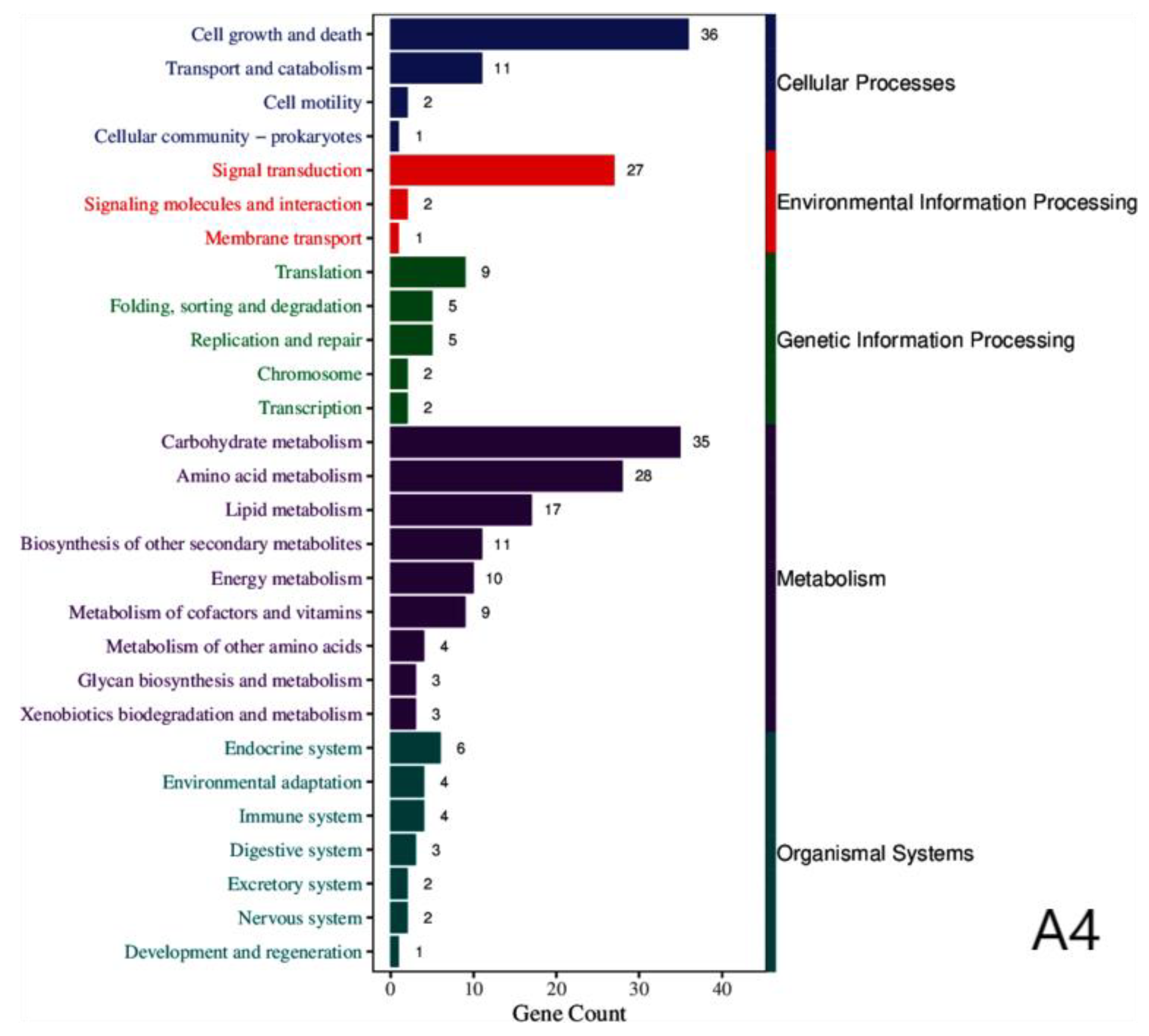
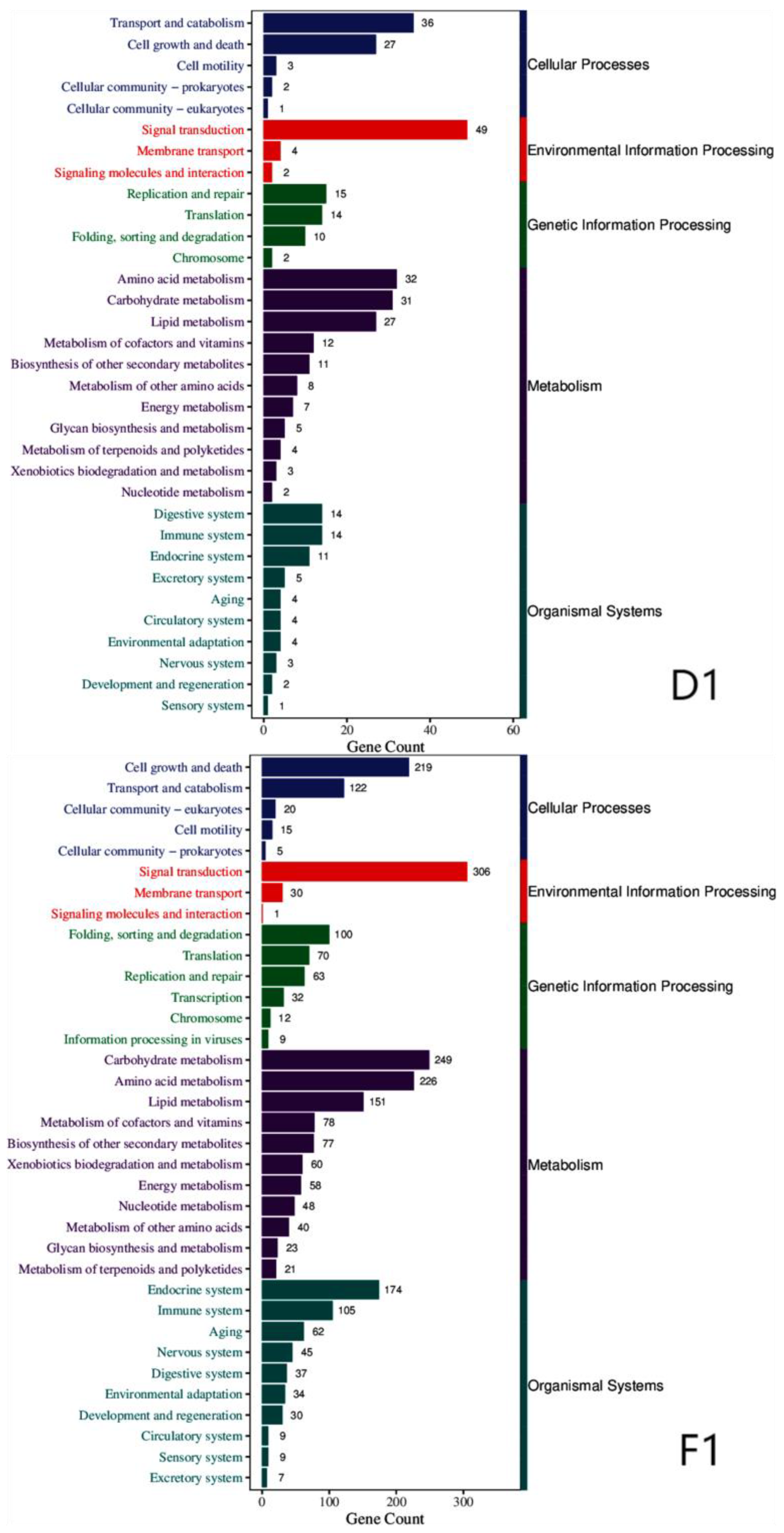
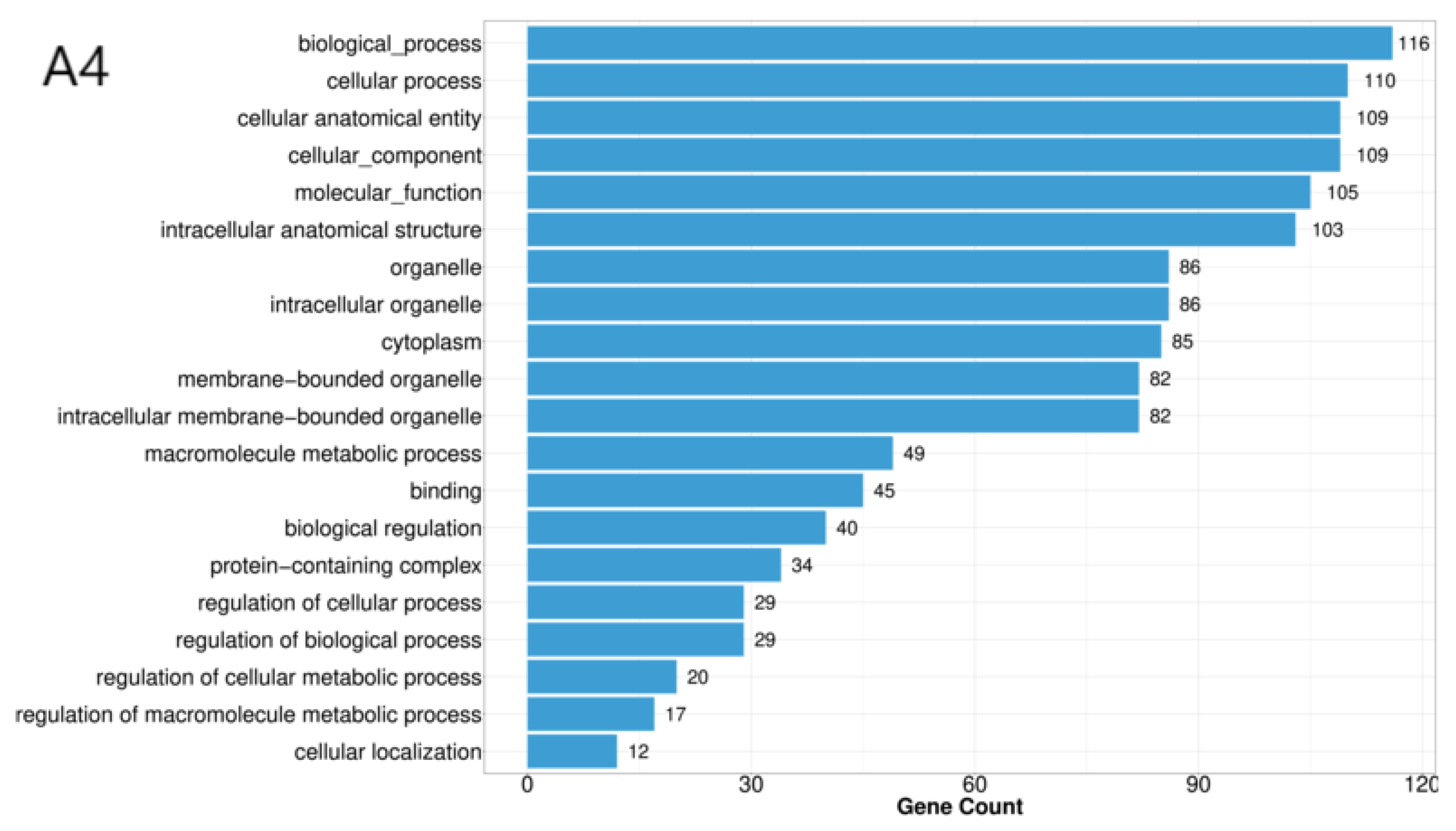
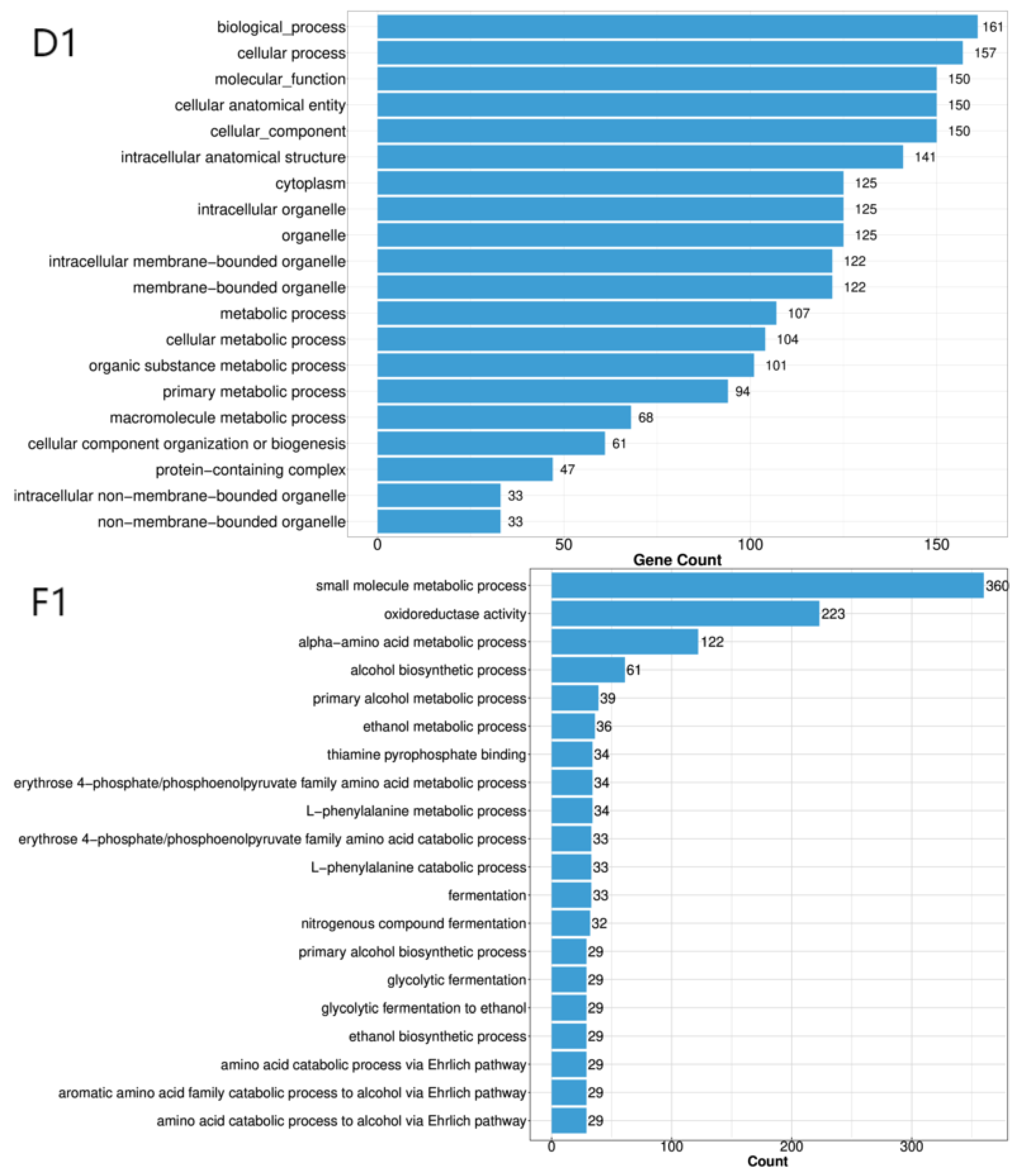
3.2. Annotation of Differentially Expressed Genes
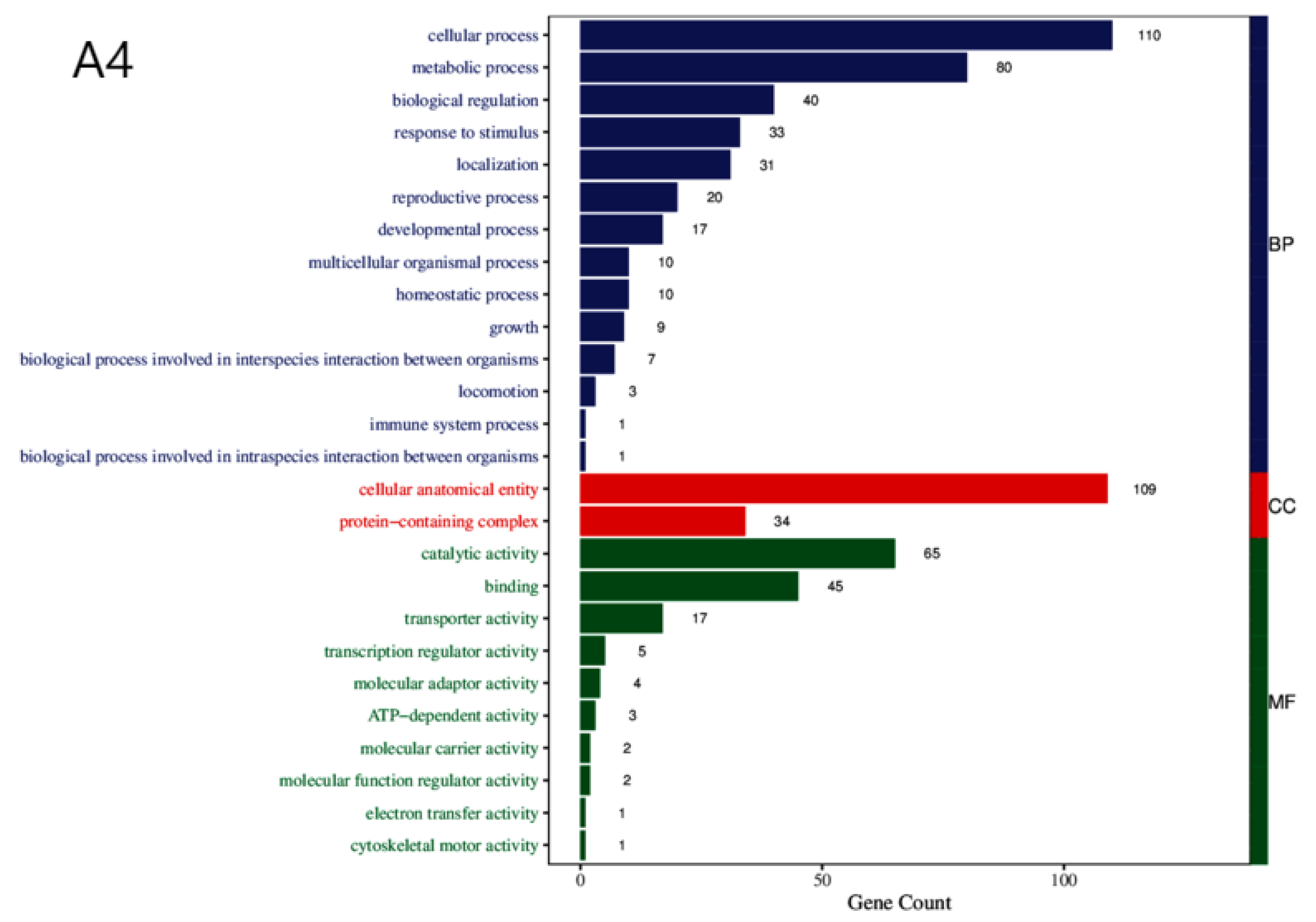
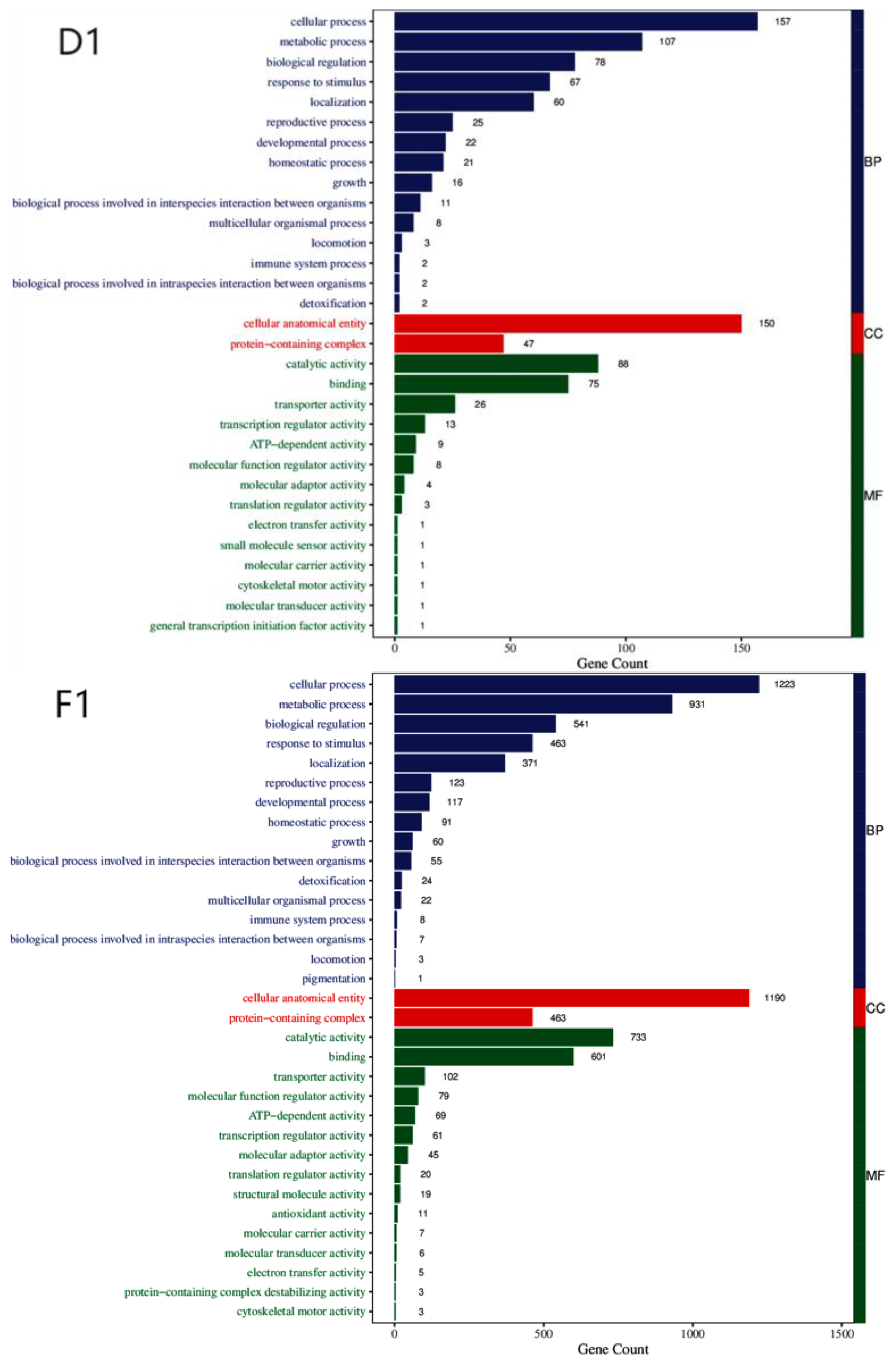
3.3. Enrichment Analysis of DEGs
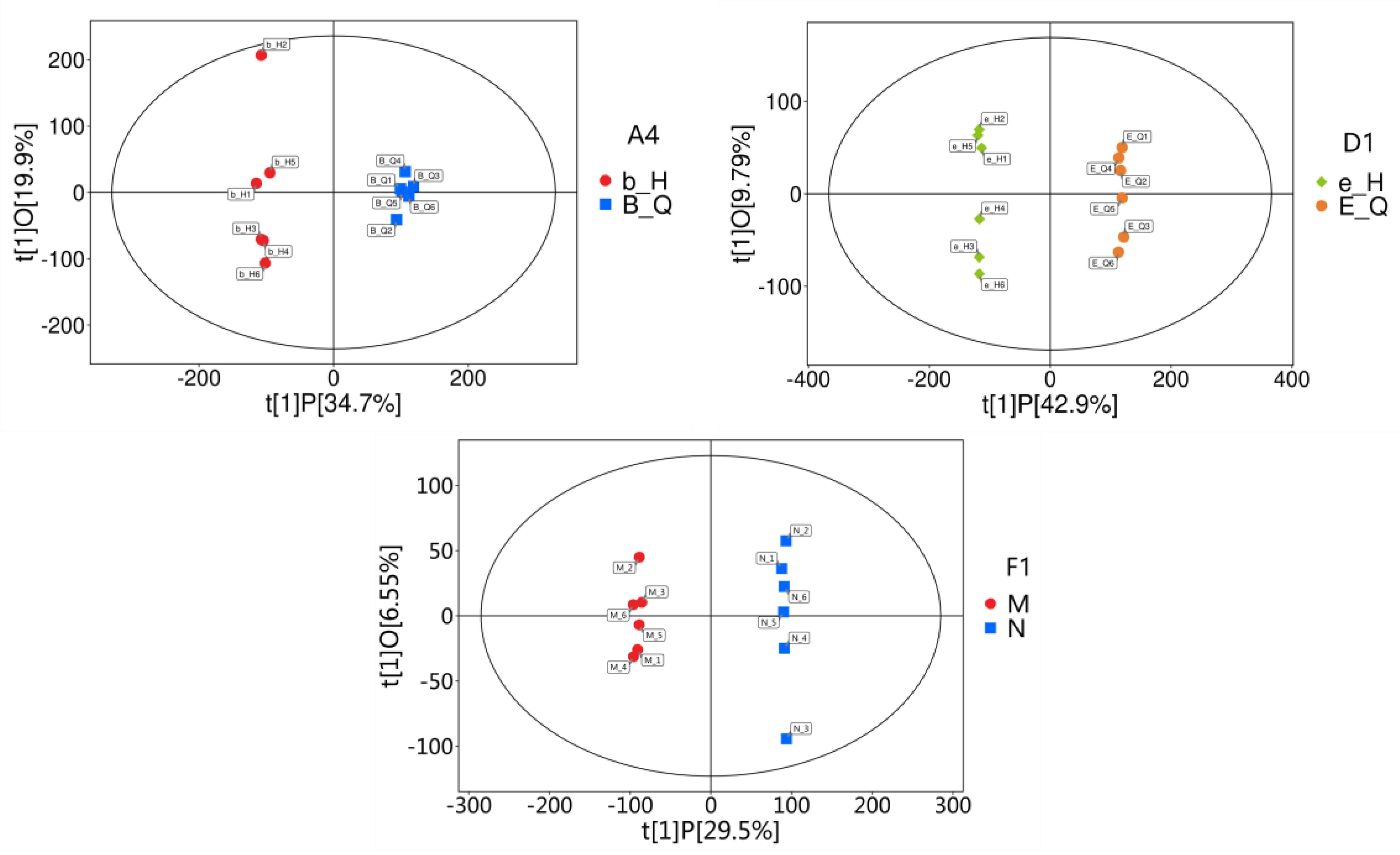
3.4. Orthogonal Partial Least Squares-Discriminant Analysis (OPLS-DA) of Metabolomic Data
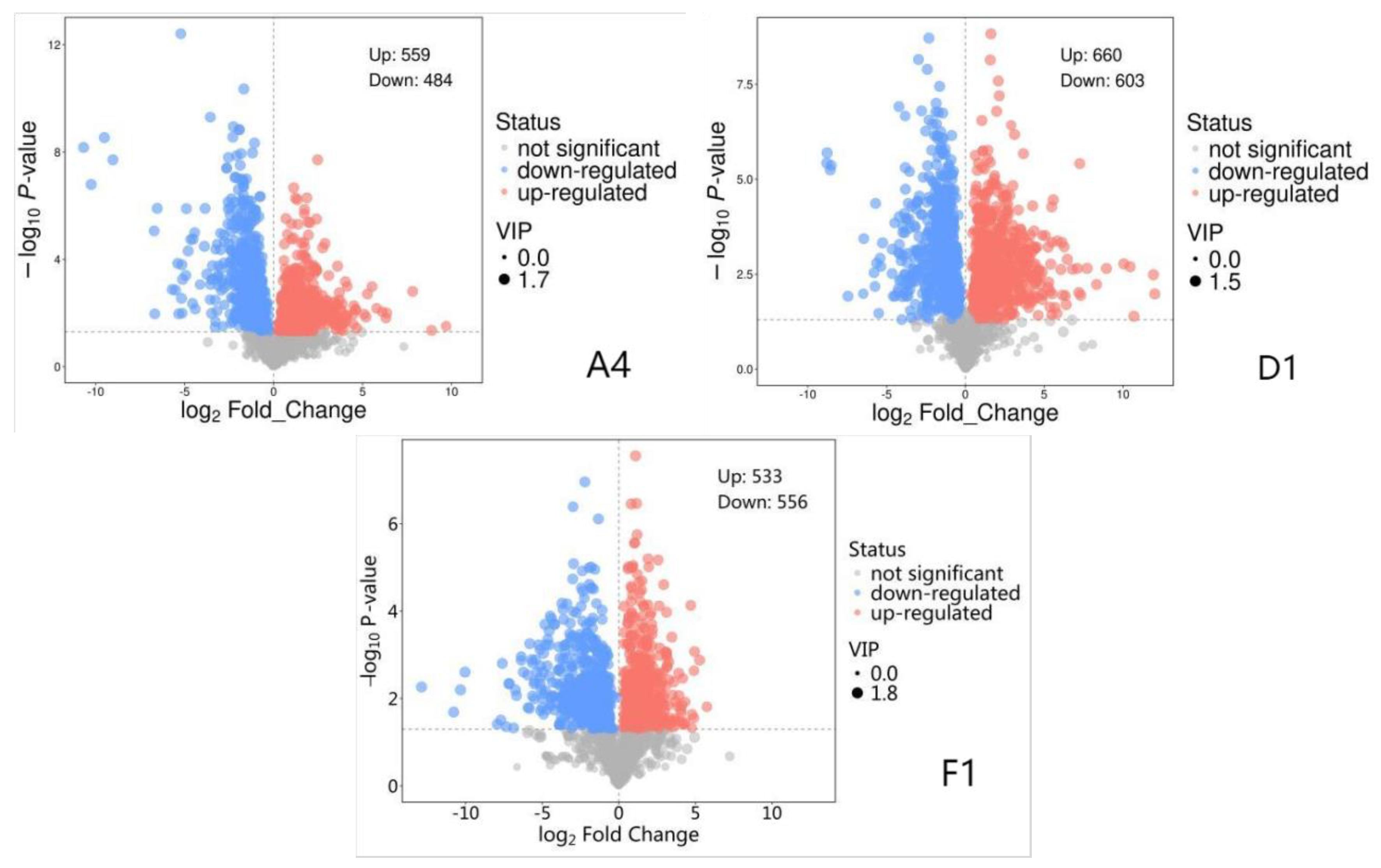
3.5. Identification of Differentially Abundant Metabolites (DAMs)
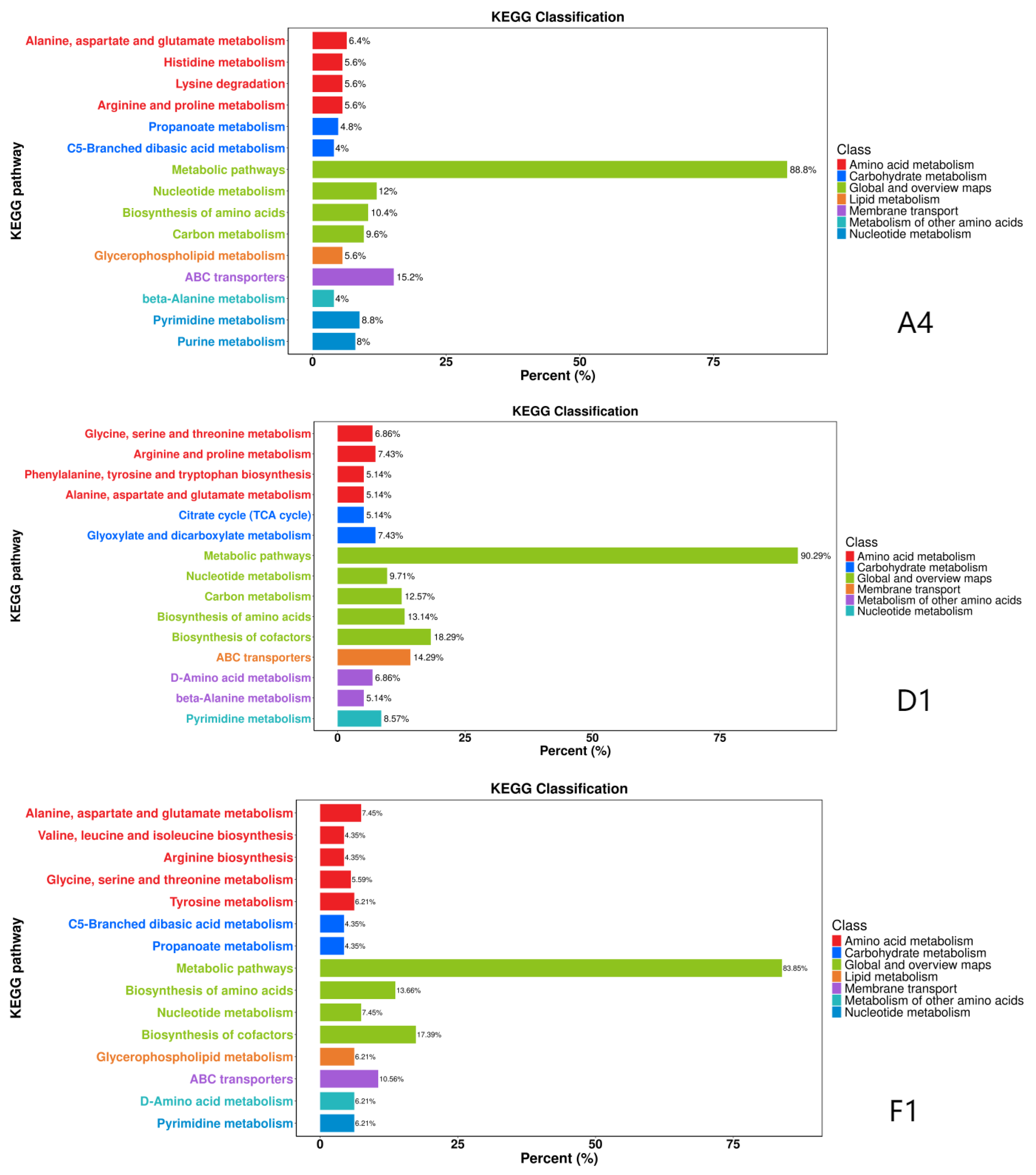
3.6. KEGG Pathway Annotation and Enrichment Analysis of DAMs
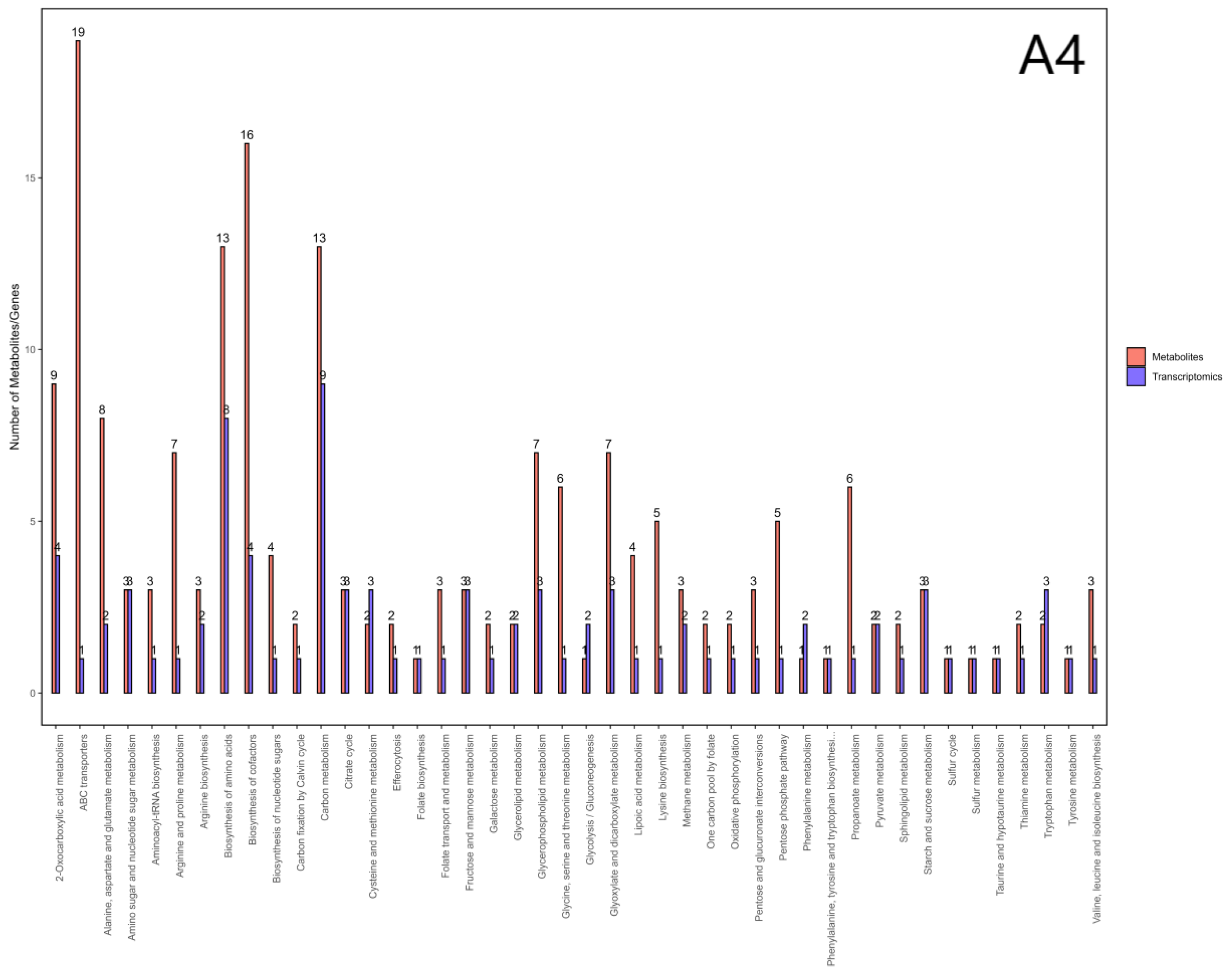
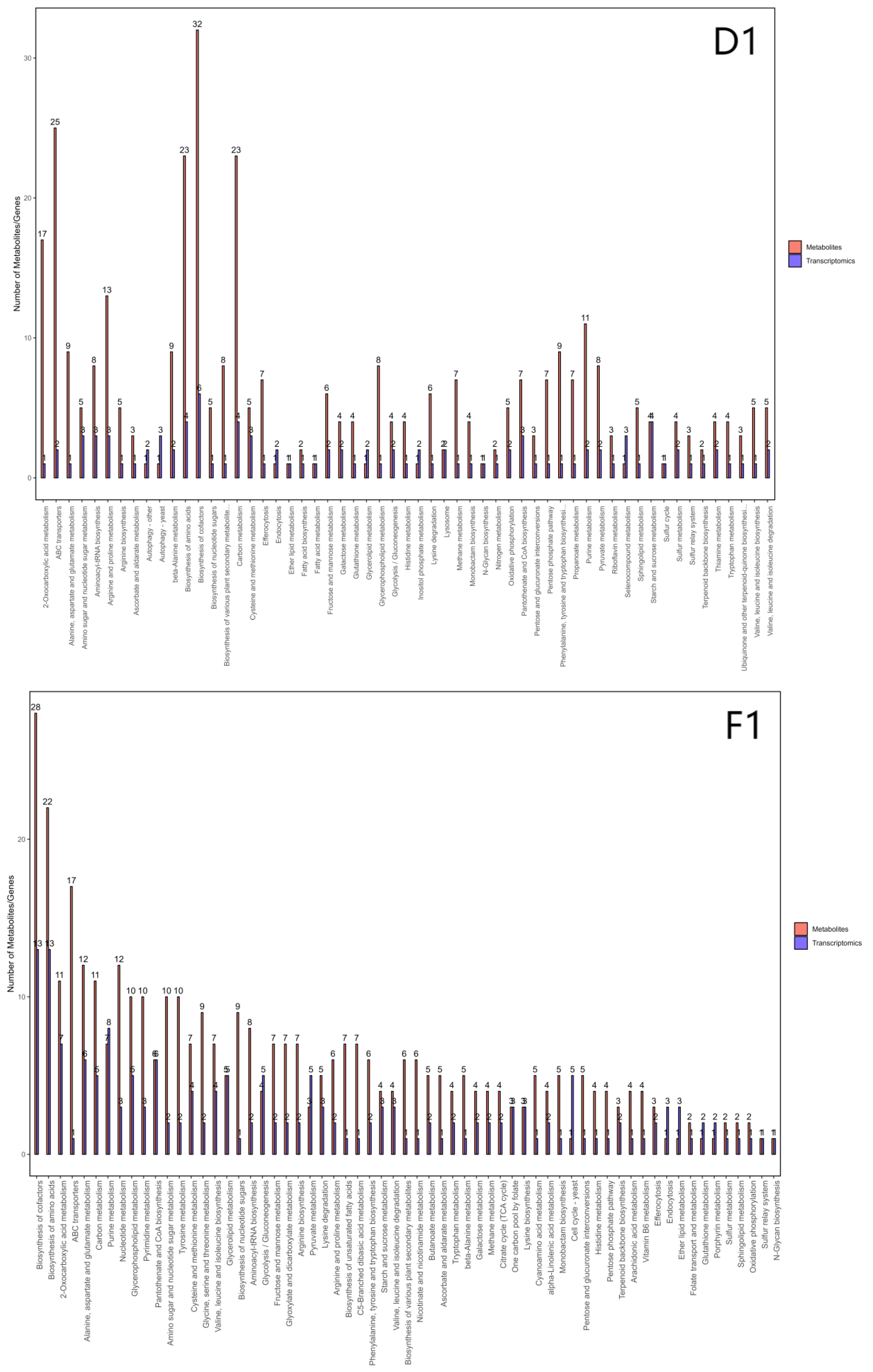
3.7. Integrated Analysis of Transcriptomics and Metabolomics
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
AI Statement
References
- Liu, Y. Study on the Antibacterial Effect of Volatile Organic Compounds from Ceratocystis fimbriata on Two Leaf Spot Diseases. Ph.D. Thesis, Yunnan Agricultural University, Kunming, China, 2024. [Google Scholar]
- Liu, Y.; He, Y.; Ruan, X. Ceratocystis fimbriata——A Pathogen Damaging Multiple Crops with Wide Distribution. J. Yunnan Agric. Univ. 2003, 18, 408–412. [Google Scholar]
- Zou, L.; Jiang, Y.; Yu, L.; et al. Morphological Studies on Ceratocystis fimbriata Isolated from Pomegranate, Taro and Sweet Potato. J. Yunnan Univ. (Nat. Sci. Ed.) 2008, 30 (Suppl. 1), 81–85. [Google Scholar]
- Yu, L.; Gao, L.; Guo, J.; et al. Effects of Ceratocystis fimbriata Infection on Antioxidant Enzyme Activities in Sweet Potato Storage Roots. Chin. J. Eco-Agric. 2011, 19, 141–145. [Google Scholar] [CrossRef]
- Sailapura, E. N.; Kulkarni, S.; Shantaraju, S.; et al. Molecular Characterization and Genetic Variation in Ceratocystis fimbriata Ell. & Halst. on Pomegranate. J. Fungi 2022, 8, 1276. [Google Scholar]
- Alizadeh, M.; Azaryan, A.; Ghaleh, M. R. K.; et al. Host and Geographical Distribution of the Plant-Pathogenic Fungus Ceratocystis fimbriata (Ascomycota) Worldwide. Nova Hedwigia 2024, 118, 1–24. [Google Scholar] [CrossRef]
- Ploetz, R. C.; Hulcr, J.; Wingfield, M. J.; et al. Destructive Tree Diseases Associated with Ambrosia and Bark Beetles: Black-Swan Events in Tree Pathology? Plant Dis. 2013, 97, 856–872. [Google Scholar] [CrossRef] [PubMed]
- Alfenas, A. C.; Ferreira, M. A.; et al. Global Spread of Ceratocystis Pathogens: Pathways and Impacts. Annu. Rev. Phytopathol. 2023, 61, 301–325. [Google Scholar]
- Zheng, X.; Xu, B.; He, P.; et al. The Pathogen of Pomegranate Wilt in Sichuan Province. Mycosystema 2012, 31, 523–530. [Google Scholar]
- Yuan, Y.; Liu, X.; Hu, X. Preliminary Study on Fungicide Screening and Synergistic Effects for Control of Pomegranate Wilt. Agrochemicals 2018, 57, 71–74. [Google Scholar]
- Huang, Q.; Lu, W.; Fan, J.; et al. Discovery of Pomegranate Wilt in Yunnan Province. Acta Phytopathol. Sin. 2004, 34, 95–96. [Google Scholar]
- 12. Harrington, T. C. Ecology and Evolution of Mycophagous Bark Beetles and Their Fungal Partners. In Bark Beetles: Biology and Ecology of Native and Invasive Species; Vega, F. E., Hofstetter, R. W., Eds.; Academic Press: Cambridge, MA, USA, 2015; pp. 257–291. ISBN 9780124171565. [Google Scholar]
- Chaudhari, V. G. Studies on Wilt-Complex Disease of Pomegranate. Ph.D. Thesis, Mahatma Phule Krishi Vidyapeeth, Rahuri, India, 2015. [Google Scholar]
- Halfeld-Vieira, B. A.; Zilli, J. E.; Nechet, K. L.; Pereira, G. M. D.; Souza, G. R. First Record of Ceratocystis fimbriata on Carapa guianensis. New Dis. Rep. 2012, 26, 13. [Google Scholar] [CrossRef]
- Kenna, G. H. J.; Mullins, E.; Doohan, F. M. The Resilience of Fusarium Pathogens: A Review of the Role of Chlamydospores and Persistent Propagules in Disease Dynamics. Mol. Plant Pathol. 2023, 24, 1023–1040. [Google Scholar]
- Ding, Z.; Peng, J.; Zhang, X. Correlation Analysis of Transcriptome and Metabolome Reveals Association between Amino-Acid Biosynthesis and Chlamydospore Formation in Fusarium oxysporum f. sp. cubense. In Proceedings of the 2024 Annual Academic Conference of the Mycological Society of China, Beijing, China, 2024. [Google Scholar]
- Zhang, H.; Wu, Q.; Cao, L.; et al. Integrated Transcriptomic and Metabolomic Analysis Reveals Key Genes Involved in Fusarium oxysporum Chlamydospore Formation. Front. Microbiol. 2022, 13, 845731. [Google Scholar]
- Son, H.; Kim, M. G.; Min, K.; et al. A Metabolomic Approach to Understand Fusarium graminearum Development. Nat. Commun. 2021, 12, 3816. [Google Scholar]
- Rangel, L. I.; Spanner, R. E.; Bolton, M. D.; et al. Targeting Fungal Dormancy: A New Strategy for Disease Control. Phytopathology 2022, 112, 1029–1038. [Google Scholar]
- Pereira, C.; Fernandes, Â.; Barros, L.; et al. Natural Compounds as Inhibitors of Fungal Chlamydospore Formation. J. Fungi 2023, 9, 588. [Google Scholar]
- Hu, Z.; Chang, X.; Dai, T.; et al. GC-MS-Based Metabolomic Analysis of Botrytis cinerea. J. Instrum. Anal. 2017, 36, 633–639. [Google Scholar]
- Kim, K. T.; Lee, Y. H. RNA-Seq-Based Transcriptome Analysis of Fungal Development. Methods Mol. Biol. 2022, 2480, 173–192. [Google Scholar]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A Revolutionary Tool for Transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A. M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Grabherr, M. G.; Haas, B. J.; Yassour, M.; et al. Trinity: Reconstructing a Full-Length Transcriptome without a Genome from RNA-Seq Data. Nat. Biotechnol. 2011, 29, 644–652. [Google Scholar] [CrossRef]
- Anders, S.; Huber, W. Differential Expression Analysis for Sequence Count Data. Genome Biol. 2010, 11, R106. [Google Scholar] [CrossRef] [PubMed]
- Pang, Z.; Chong, J.; Zhou, G.; et al. MetaboAnalystR 3.0: Toward an Optimized Workflow for Global Metabolomics. Nat. Protoc. 2021, 16, 4615–4633. [Google Scholar] [CrossRef]
- Smedsgaard, J.; Nielsen, J. Metabolite Profiling of Fungi and Yeast: From Phenotype to Metabolome by MS and Informatics. Fungal Genet. Biol. 2005, 42, 981–992. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Zhang, H.; Wang, L. Optimization of Metabolite Extraction and LC-MS/MS Detection Methods for Fungi. Acta Microbiol. Sin. 2020, 60, 1650–1658. [Google Scholar]
- Want, E. J.; Wilson, I. D.; Gika, H.; et al. Global Metabolic Profiling Procedures for Urine Using UPLC-MS. Nat. Protoc. 2010, 5, 1005–1018. [Google Scholar] [CrossRef]
| GO categories | Functional annotation |
| BP | Cellular process, metabolic process, biological regulation, response to stimulus, localization, reproductive process, developmental process, multicellular organismal process, homeostasis process, growth, biological process involved in interspecies interaction between organisms, locomotion, immune system process, biological process involved in intraspecies interaction between organisms. |
| CC | Cellular anatomical entity, protein-containing complex. |
| MF | Catalytic activity, binding, transporter activity, transcription regulator activity, molecular adaptor activity, ATP-dependent activity, molecular carrier activity, molecular function regulator activity, electron transfer activity, cytoskeletal motor activity. |
| KEGG categories | KEGG pathway annotation |
| Cellular processes | Cell growth and death, transport and catabolism, cell motility, cellular community-prokaryotes |
| Environmental information processing | Signal transduction, signaling molecules and interactions, membrane transport |
| Genetic information processing | Replication and repair, translation, folding, sorting and degradation, chromosome |
| Metabolism | Carbohydrate metabolism, amino acid metabolism, lipid metabolism, biosynthesis of other secondary metabolites, energy metabolism, metabolism of cofactors and vitamins, metabolism of other amino acids, glycan biosynthesis and metabolism, xenobiotics biodegradation and metabolism. |
| Organismal systems | Endocrine system, environmental adaptation, immune system, digestive system, excretory system, nervous system, development and regeneration. |
| KEGG pathway | Strain A4 | Strain D1 | Strain F1 |
|---|---|---|---|
| Starch and sucrose metabolism | √ | √ | √ |
| Glycerophospholipid metabolism | √ | ||
| Pentose and glucuronate interconversions | √ | √ | |
| Fructose and mannose metabolism | √ | ||
| Lysine degradation | √ | √ | |
| Glycine, serine and threonine metabolism | √ | ||
| Arginine biosynthesis | √ | ||
| Ubiquinone and other terpenoid-quinone biosynthesis | √ | ||
| Carbohydrate digestion and absorption | √ | √ | |
| Plant-pathogen interaction | √ | ||
| Tropane, piperidine and pyridine alkaloid biosynthesis | √ | ||
| Isoquinoline alkaloid biosynthesis | √ | √ | √ |
| Nitrogen metabolism | √ | ||
| Sphingolipid metabolism | √ | ||
| Neomycin, kanamycin and gentamicin biosynthesis | √ | ||
| Streptomycin biosynthesis | √ | ||
| Retinol metabolism | √ | ||
| Novobiocin biosynthesis | √ | ||
| Neuroactive ligand-receptor interaction | √ | ||
| Biosynthesis of unsaturated fatty acids | √ | ||
| Lysosome | √ | ||
| PI3K-Akt signaling pathway | √ | ||
| FoxO signaling pathway | √ | ||
| Steroid biosynthesis | √ | ||
| Fanconi anemia pathway | √ | ||
| Homologous recombination | √ | ||
| beta-Alanine metabolism | √ | ||
| Longevity regulating pathway-worm | √ | ||
| Pantothenate and CoA biosynthesis | √ | √ | |
| Tyrosine metabolism | √ | √ | |
| Ferroptosis | √ | ||
| Selenocompound metabolism | √ | ||
| Mineral absorption | √ | ||
| RIG-I-like receptor signaling pathway | √ | ||
| Cell adhesion molecules | √ | ||
| Glycolysis/ Gluconeogesis | √ | ||
| Biosynthesis of amino acids | √ | ||
| Purine metabolism | √ | ||
| 2-Oxocarboxylic acid metabolism | √ | ||
| Cysteine and methionine metabolism | √ | ||
| Alanine, aspartate and glutamato metabolism | √ | ||
| Pyruvate metabolism | √ | ||
| Steroid biosynthesis | √ | ||
| Valine, leucine and isoleucine biosynthesis | √ | ||
| Glutathione metabolism | √ | ||
| Glycerolipid metabolism | √ | ||
| Ether lipid metabolism | √ | ||
| Fatty acid degradation | √ | ||
| Butanoate metabolism | √ | ||
| lsoquinoine alkaloid biosynthesis | √ | ||
| Riboflavin metabolism | √ | ||
| Caffeine metabolism | √ |
| KEGG ID | Pathway Name |
|---|---|
| sce00010 | Glycolysis / Gluconeogenesis |
| sce00030 | Pentose phosphate pathway |
| sce00040 | Pentose and glucuronate interconversions |
| sce00051 | Fructose and mannose metabolism |
| sce00052 | Galactose metabolism |
| sce00190 | Oxidative phosphorylation |
| sce00220 | Arginine biosynthesis |
| sce00250 | Alanine, aspartate and glutamate metabolism |
| sce00270 | Cysteine and methionine metabolism |
| sce00290 | Valine, leucine and isoleucine biosynthesis |
| sce00330 | Arginine and proline metabolism |
| sce00380 | Tryptophan metabolism |
| sce00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
| sce00500 | Starch and sucrose metabolism |
| sce00520 | Amino sugar and nucleotide sugar metabolism |
| sce00561 | Glycerolipid metabolism |
| sce00564 | Glycerophospholipid metabolism |
| sce00600 | Sphingolipid metabolism |
| sce00620 | Pyruvate metabolism |
| sce00680 | Methane metabolism |
| sce00920 | Sulfur metabolism |
| sce00970 | Aminoacyl-tRNA biosynthesis |
| sce01200 | Carbon metabolism |
| sce01210 | 2-Oxocarboxylic acid metabolism |
| sce01230 | Biosynthesis of amino acids |
| sce01240 | Biosynthesis of cofactors |
| sce01250 | Biosynthesis of nucleotide sugars |
| sce02010 | ABC transporters |
| sce04148 | Efferocytosis |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




