Submitted:
26 January 2026
Posted:
27 January 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Natural Killer Cell Biology and Development
Key Surface Markers in NK Cell Development
1.3. Natural Cytotoxicity Receptors
Metadichol: A Novel Nuclear Receptor Modulator
Materials and Methods
PBMC Isolation Protocol
RNA Isolation
cDNA Synthesis
Quantitative Real-Time PCR Analysis
Primer Sequences
Results
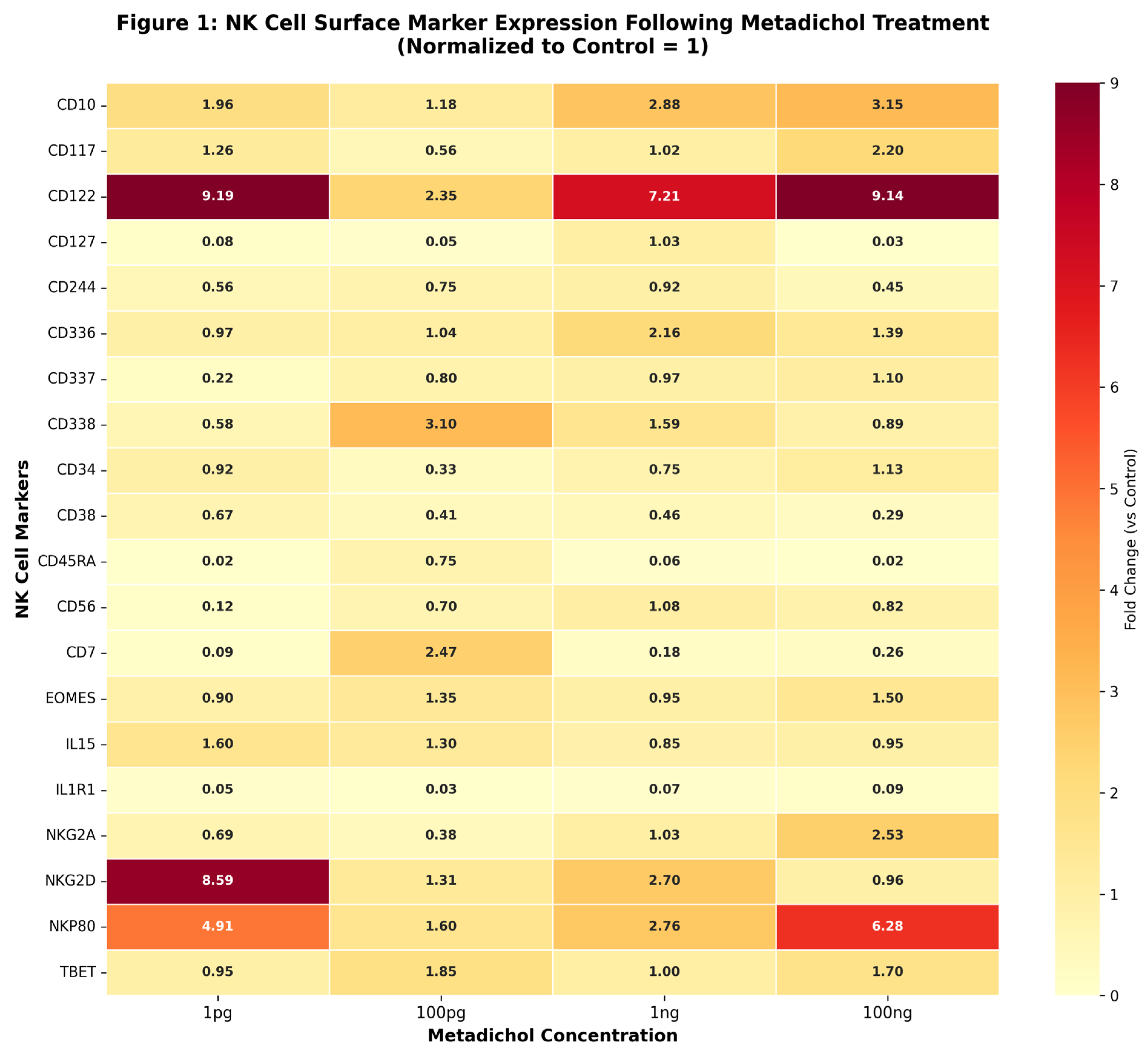
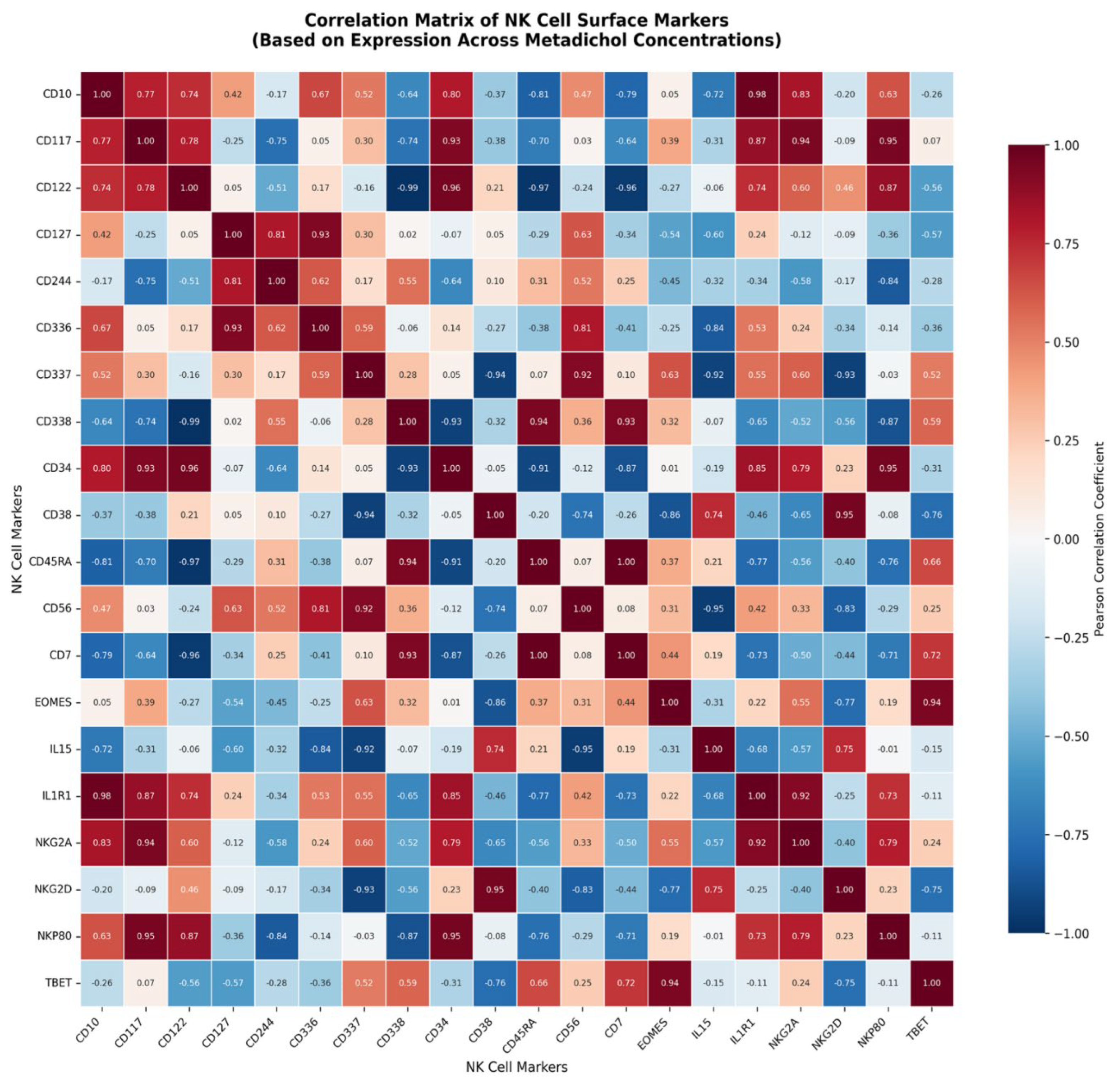
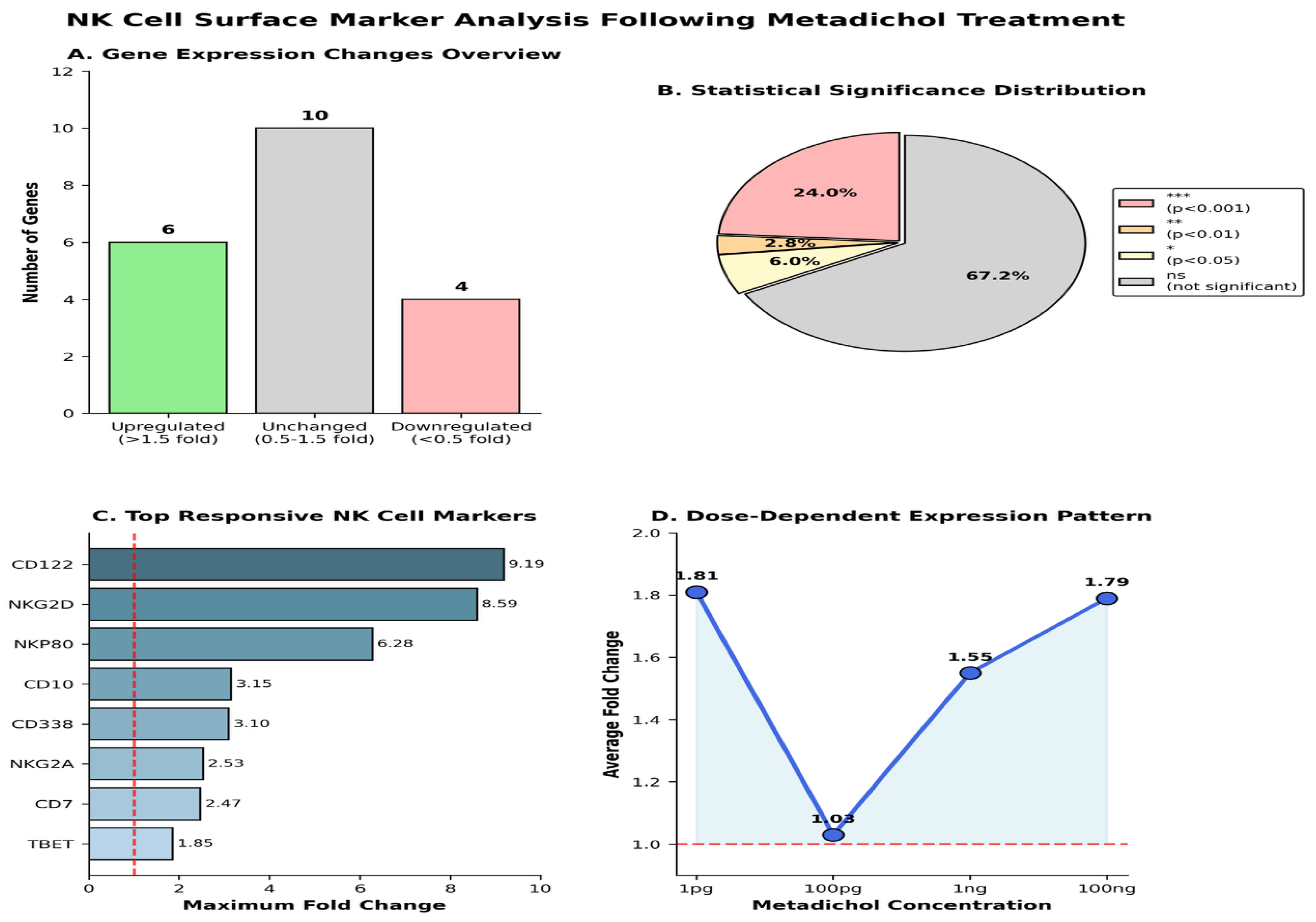
Metadichol Modulates NK Cell Surface Marker Expression
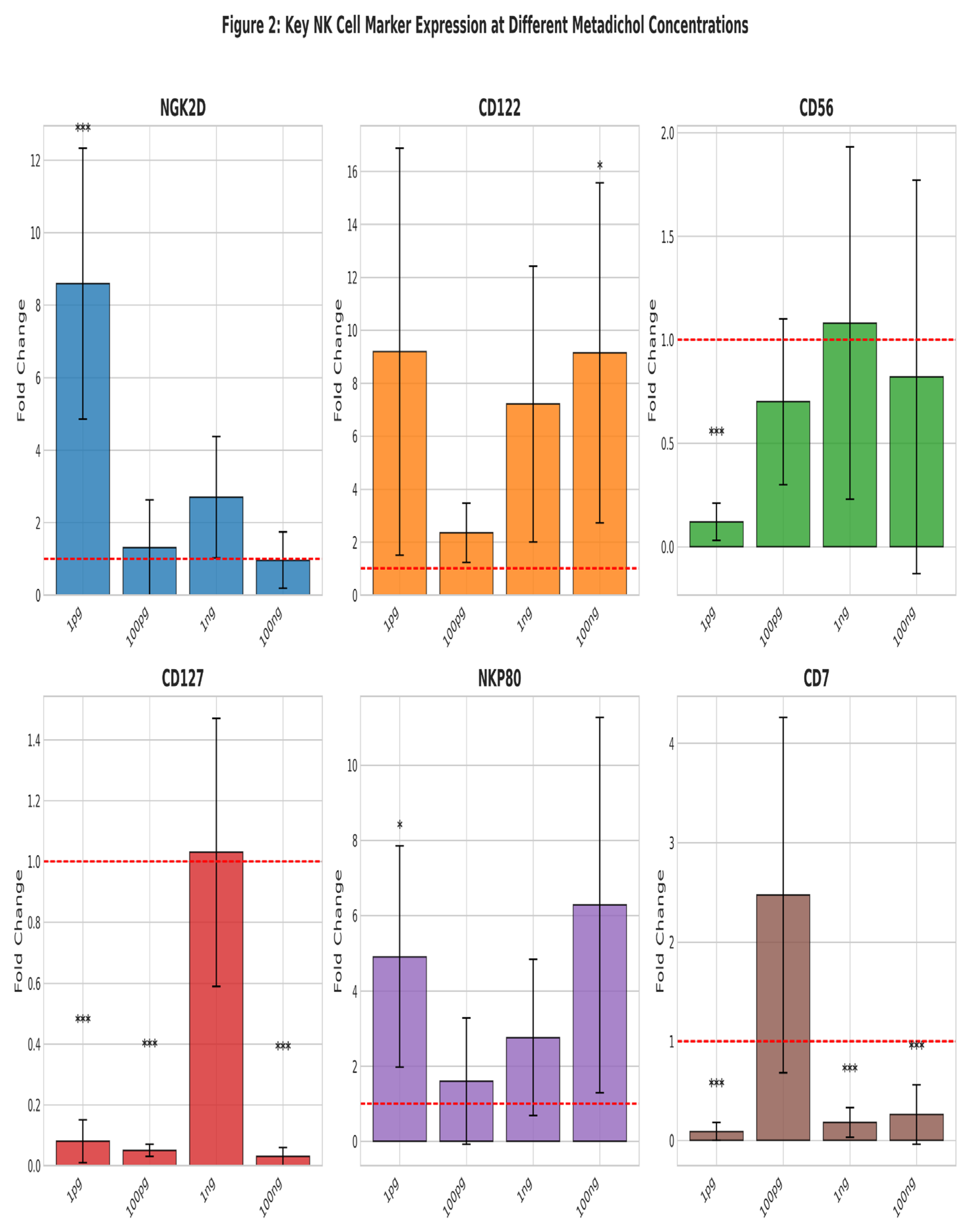
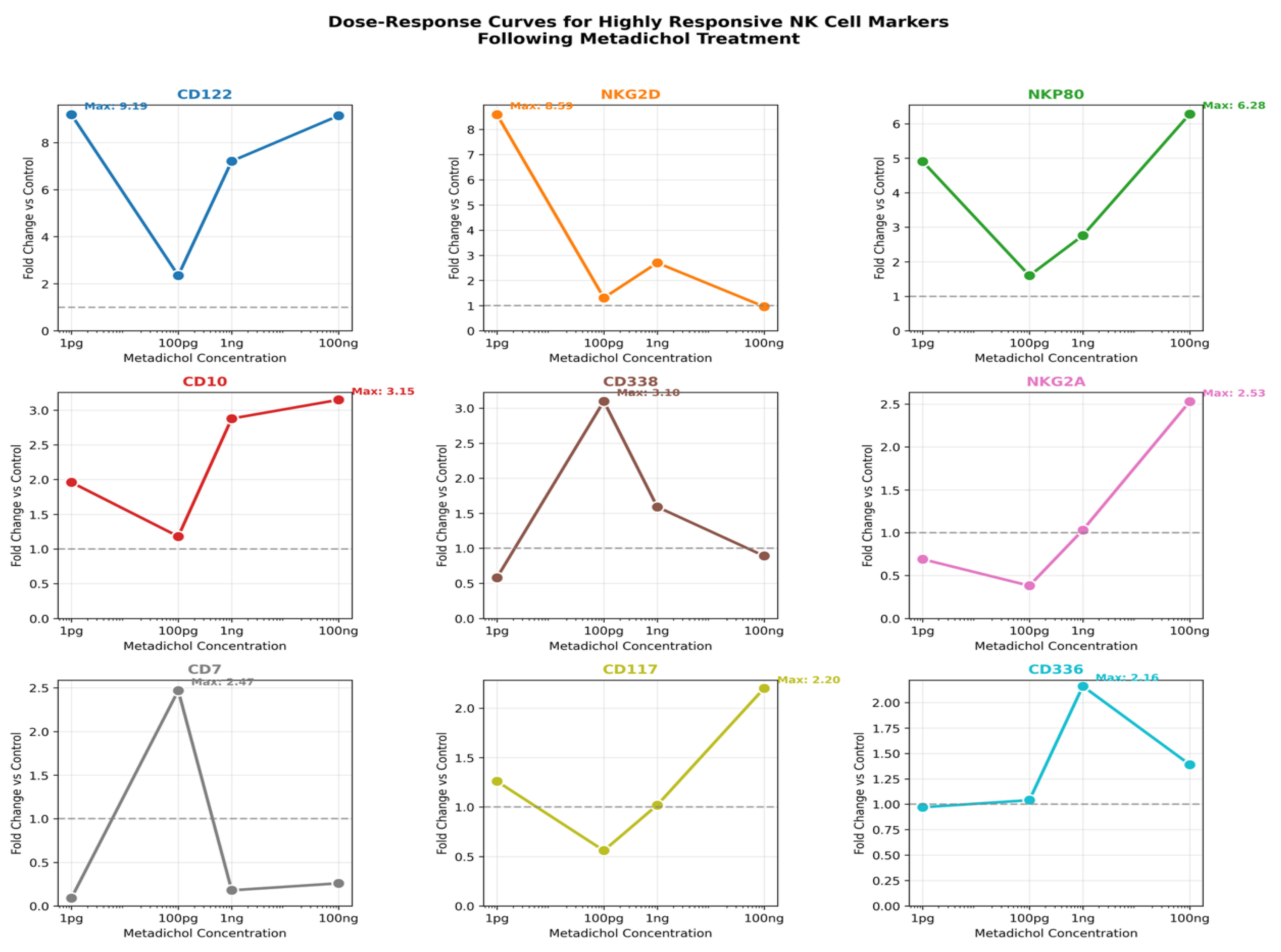
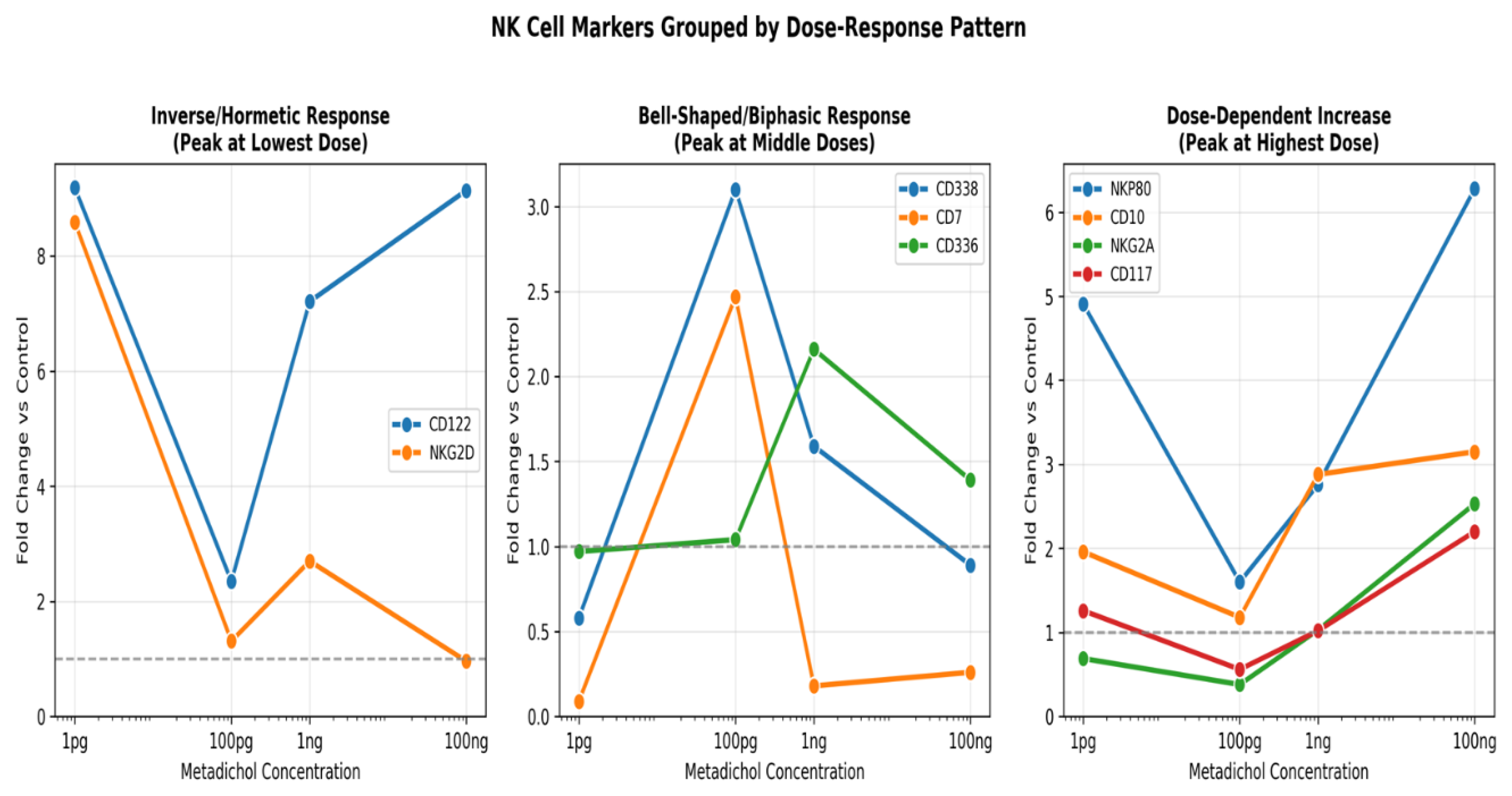
Individual Dose-Response Curves
Key Observations
- CD122 (IL-2Rβ) exhibits the highest overall response with a maximum fold change of 9.19 at 1 pg, demonstrating an inverse dose-response relationship with high expression at both 1 pg and 100 ng but reduced expression at intermediate concentrations.
- NKG2D shows a striking hormetic response pattern with peak expression (8.59-fold) at the lowest dose tested (1 pg), declining sharply at higher concentrations.
- NKP80 displays a U-shaped response curve with elevated expression at both low (1 pg: 4.91-fold) and high (100 ng: 6.28-fold) concentrations.
- CD338 uniquely peaks at 100 pg (3.10-fold), suggesting optimal activation at intermediate concentrations.
- CD10, CD117, and NKG2A demonstrate classical dose-dependent increases with maximum expression at the highest concentration (100 ng)
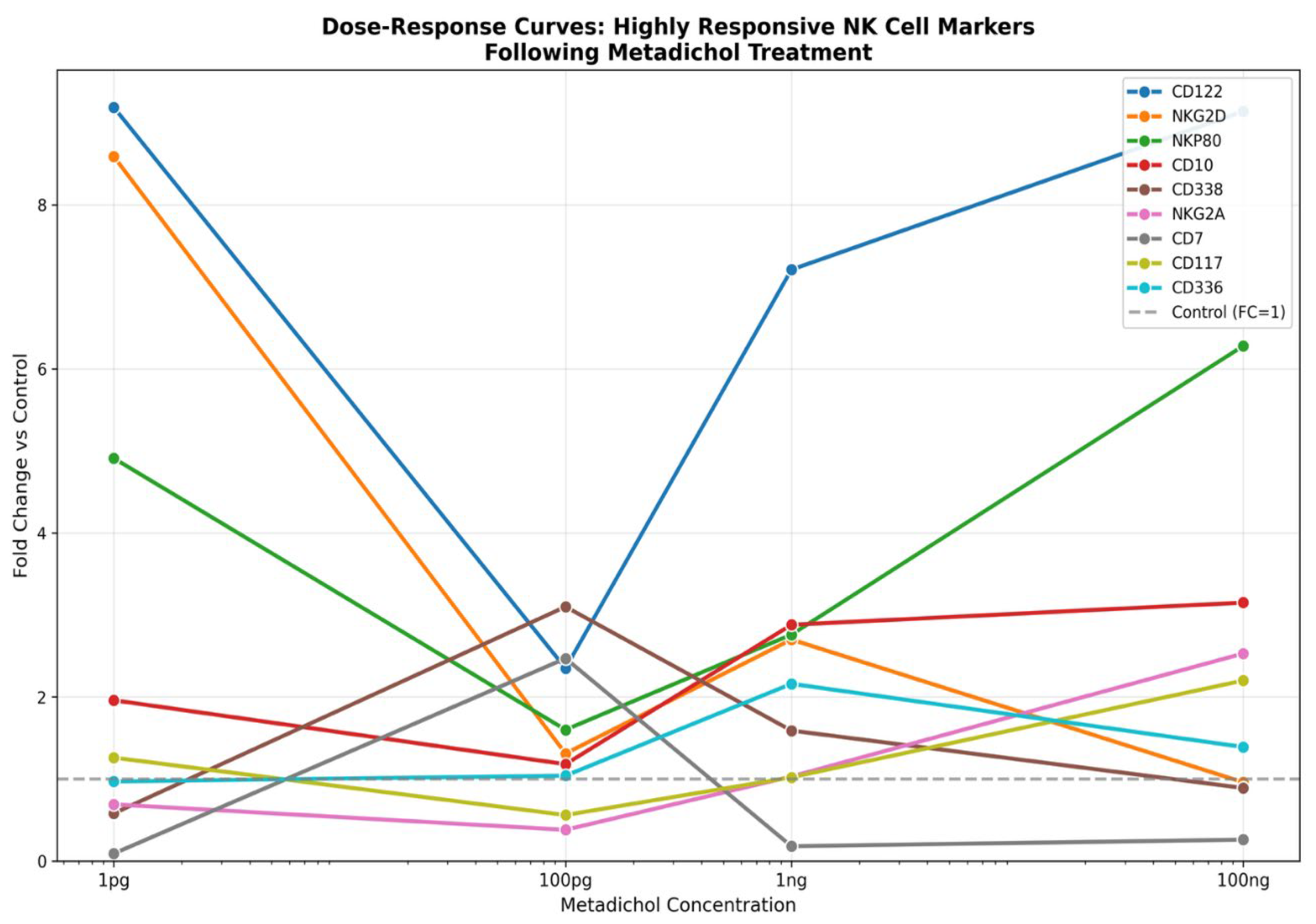
Figure 6: Dose-Response Overlay
Key Observations
- Magnitude stratification: CD122 and NKG2D clearly separate from other markers as the most responsive genes, with fold changes reaching 8–10 at their peak concentrations.
- Divergent trajectories: The overlay reveals that genes do not respond uniformly—while some markers increase monotonically with dose (CD10, CD117, NKG2A), others show inverse (NKG2D) or non-monotonic (CD122, NKP80) relationships.
- Concentration-dependent crossover: Several curves intersect at intermediate doses (100 pg–1 ng), indicating that the relative expression hierarchy among markers shifts depending on Metadichol concentration.
- Therapeutic windows: The overlay suggests that ultra-low doses (1 pg) preferentially activate cytotoxicity-associated receptors (NKG2D, CD122), while higher doses (100 ng) favor maturation markers (NKG2A, CD117).
Pattern Classification
| Pattern | Genes | Biological Interpretation |
| Inverse/Hormetic (Peak at 1 pg) | CD122, NKG2D | Ultra-sensitive signaling; possible receptor saturation or negative feedback at higher doses; suggests potent activation at picomolar concentrations |
| Bell-Shaped (Peak at 100 pg–1 ng) | CD338, CD7, CD336 | Optimal therapeutic window at intermediate concentrations; balanced receptor engagement; CD336 (NKp44) activation indicates enhanced NK cell activation state |
| Monotonic Increase (Peak at 100 ng) | NKP80, CD10, NKG2A, CD117 | Classical pharmacological response; cumulative transcriptional activation; NKG2A and CD117 upregulation suggests enhanced NK cell maturation and licensing |
Functional Implications
- Activating receptors: NKG2D (hormetic) and NKP80/CD336 (biphasic/monotonic) are activating receptors involved in tumor cell recognition. Their upregulation suggests enhanced cytotoxic potential.
- Inhibitory receptor: NKG2A (monotonic increase) is an inhibitory receptor that recognizes HLA-E. Its upregulation alongside activating receptors indicates balanced immunomodulation rather than uncontrolled activation.
- Cytokine signaling: CD122 (IL-2Rβ) is essential for IL-2 and IL-15 signaling, which drive NK cell proliferation and survival. Its hormetic response suggests maximal proliferative signaling at ultra-low Metadichol doses.
Key points
- Ultra-low dose efficacy: The hormetic responses of CD122 and NKG2D indicate that picomolar concentrations may be pharmacologically active, a finding with significant implications for therapeutic dosing.
- Pathway-specific responses: Different NK cell signaling pathways exhibit distinct concentration thresholds, suggesting engagement of multiple molecular targets.
- Balanced immunomodulation: Coordinate regulation of activating and inhibitory receptors indicates physiologically relevant immune enhancement rather than pathological hyperactivation.
- Concentration-dependent phenotypes: The overlay analysis reveals that optimal concentrations differ by target gene, necessitating careful dose selection based on desired immunological endpoints.
Discussion
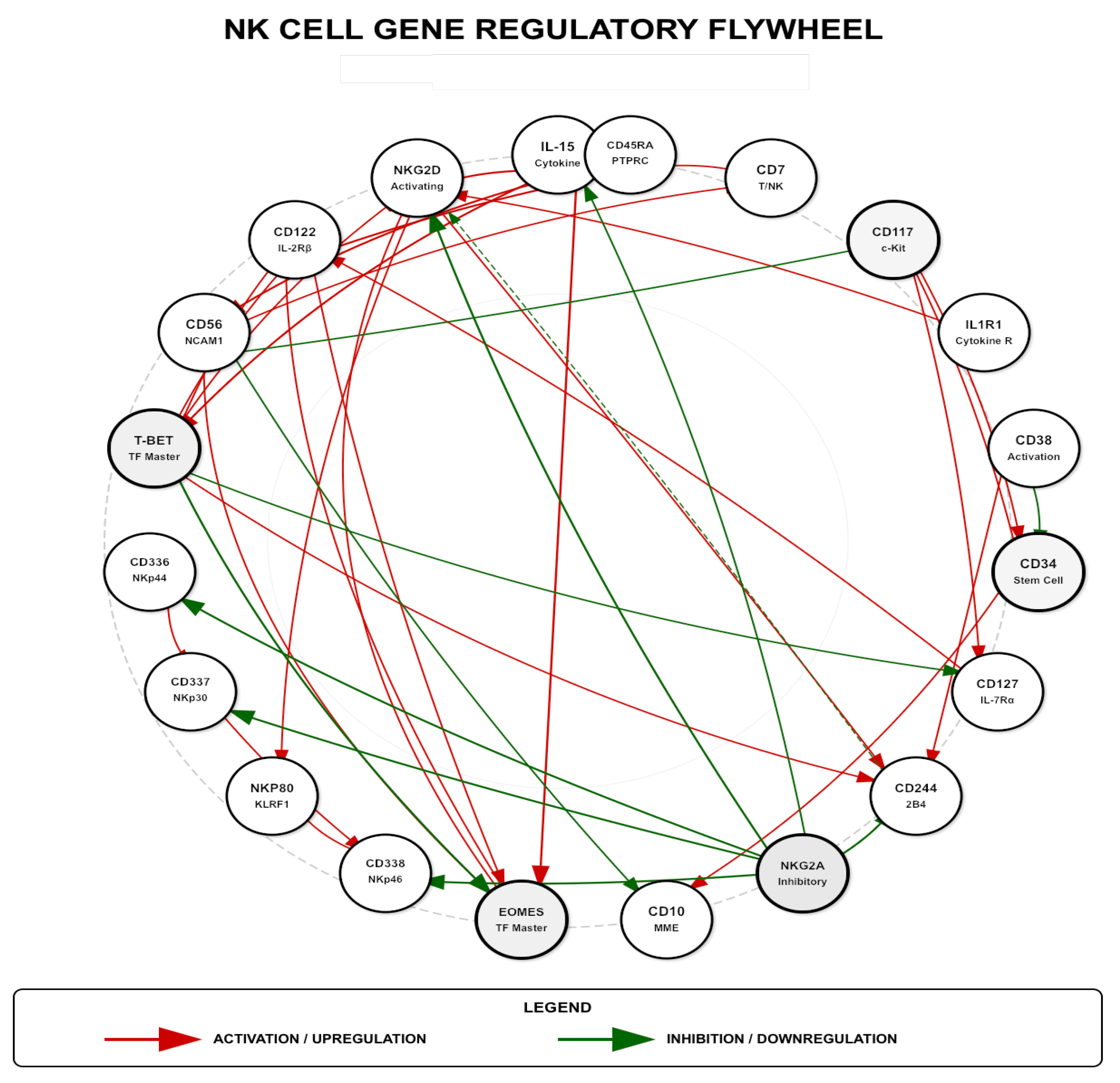
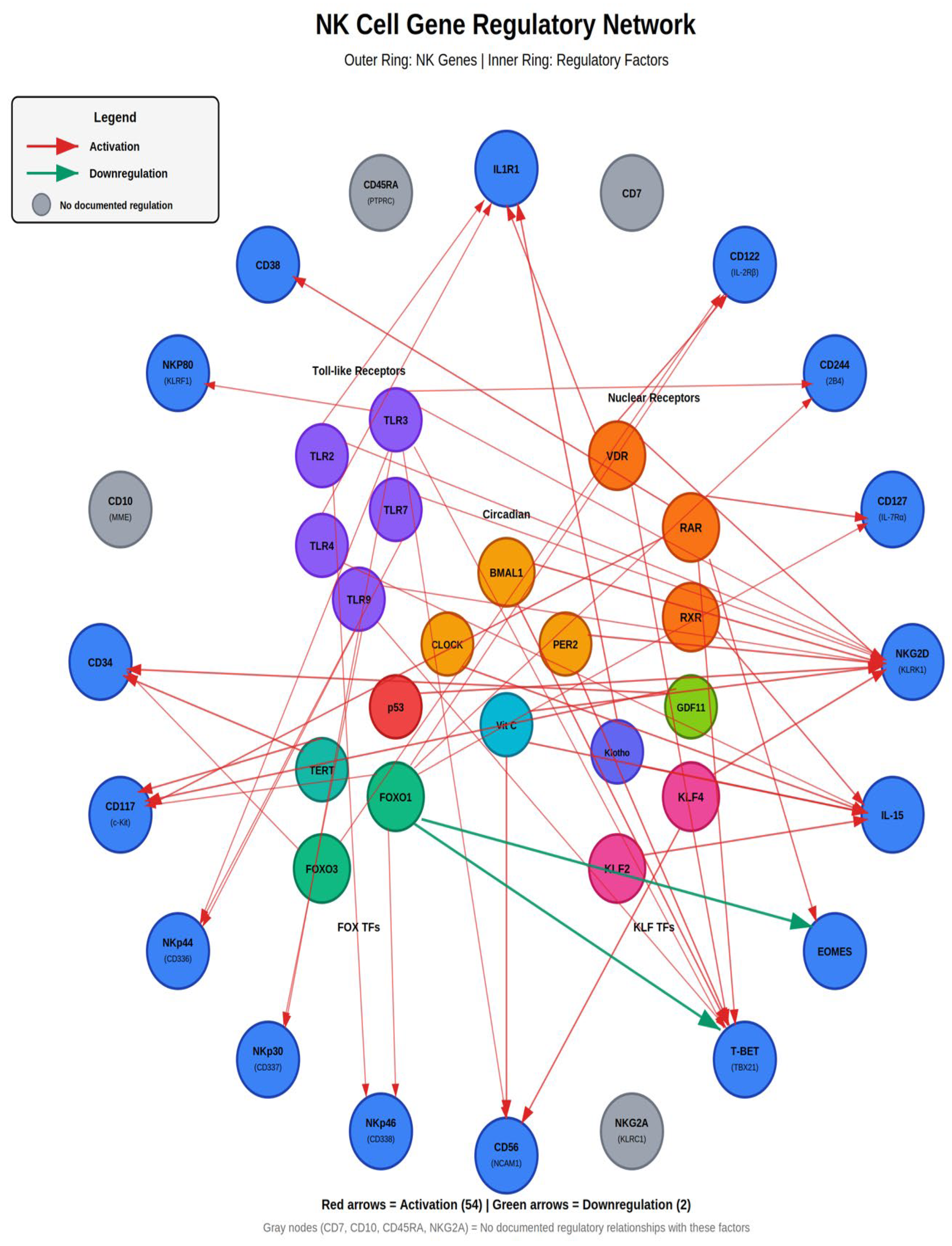
Metadichol’s Multi-Pathway Regulatory Network
NK Cell Gene Regulatory Networks
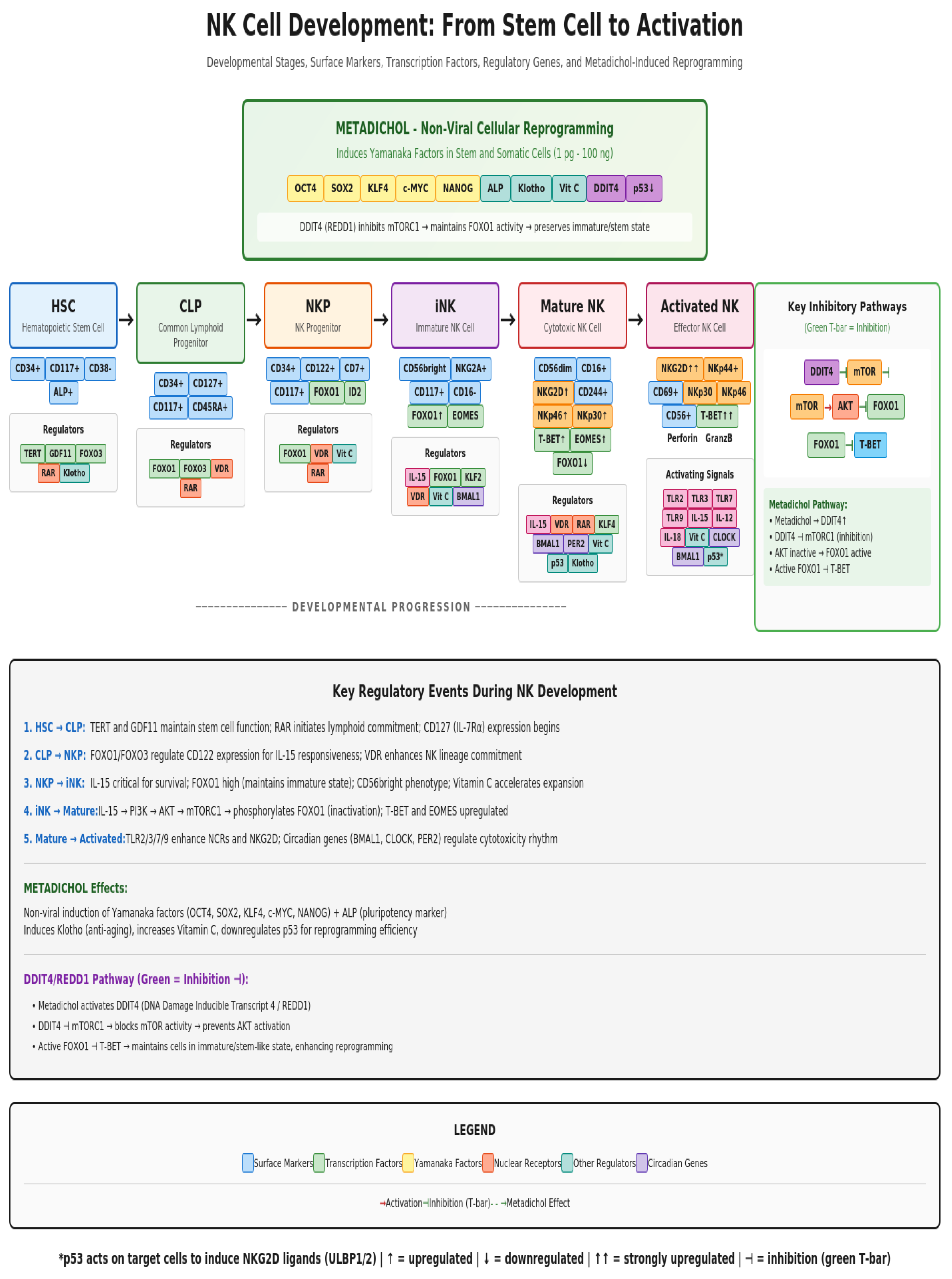
NK Cell Development Pathway
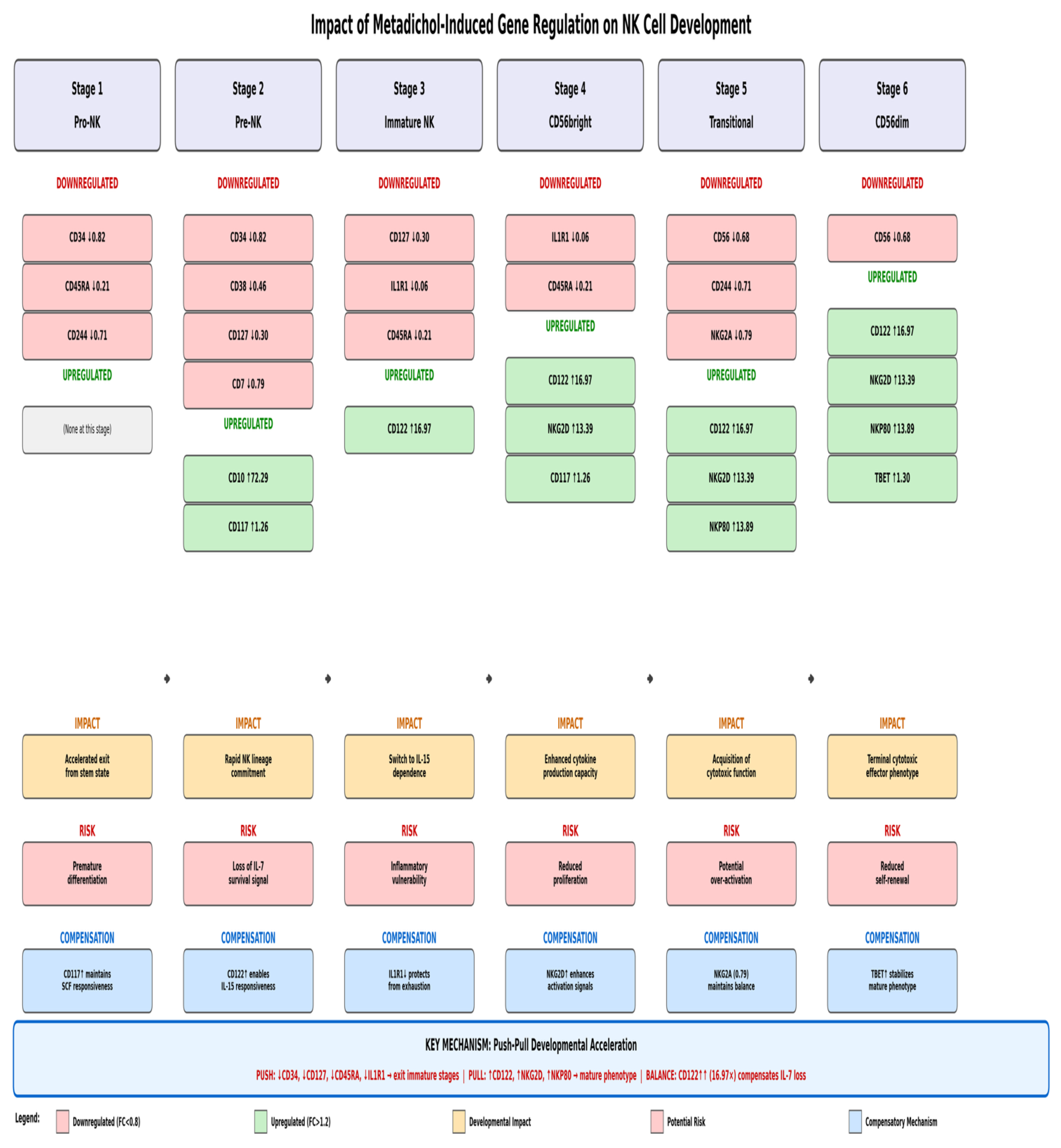
Metadichol Effects on NK Cell Developmental Stages
Push-Pull Developmental Acceleration Mechanism
Enhancement of the IL-15/CD122 Signaling Axis
Integrated Gene-Gene Interactions and Signaling Pathways:
mTOR Signaling and Metabolic Control of NK Cell Function
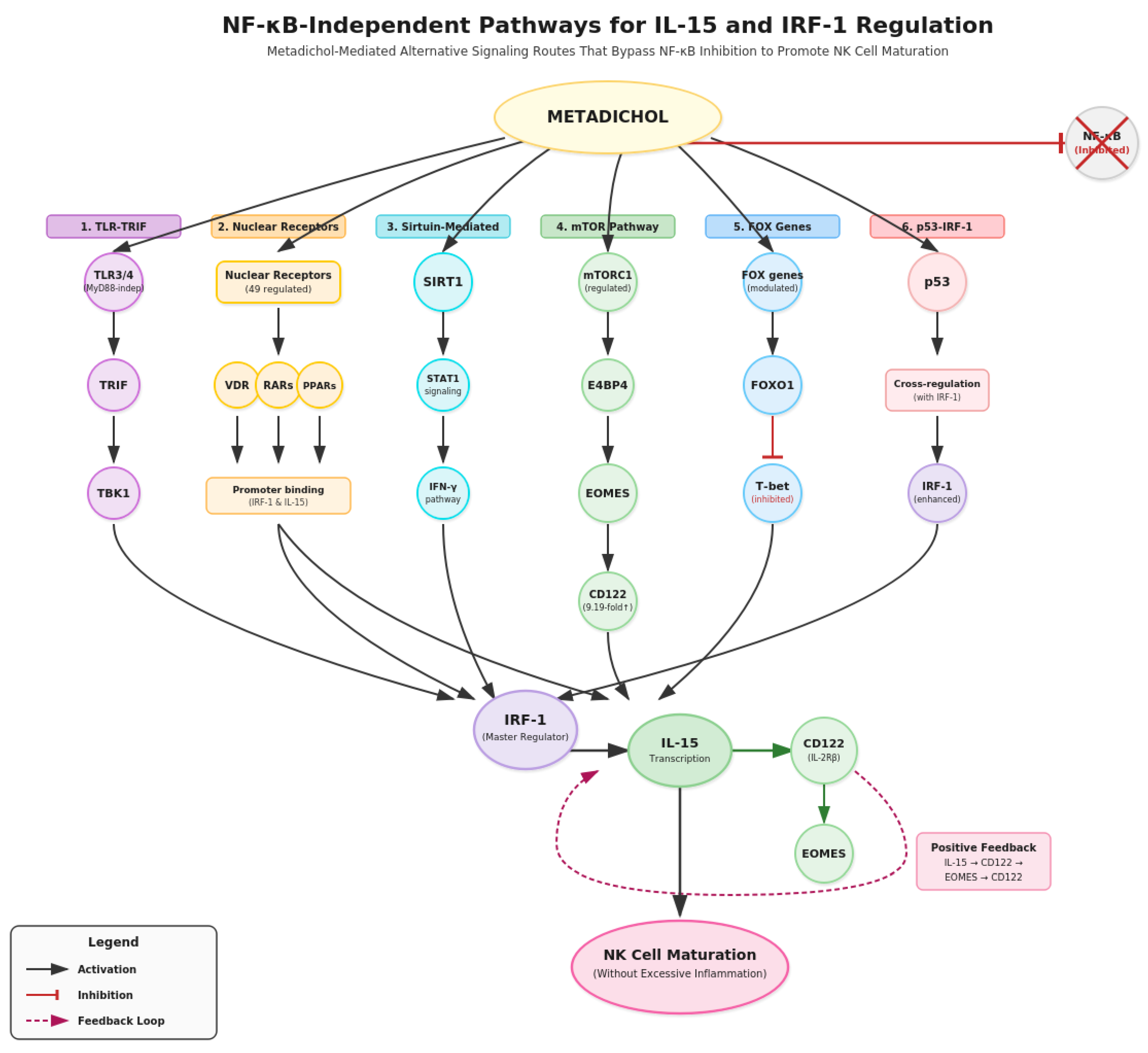
Upregulation of Activating Receptors and Cytotoxic Potential
Toll-Like Receptor Modulation and NK Cell Priming
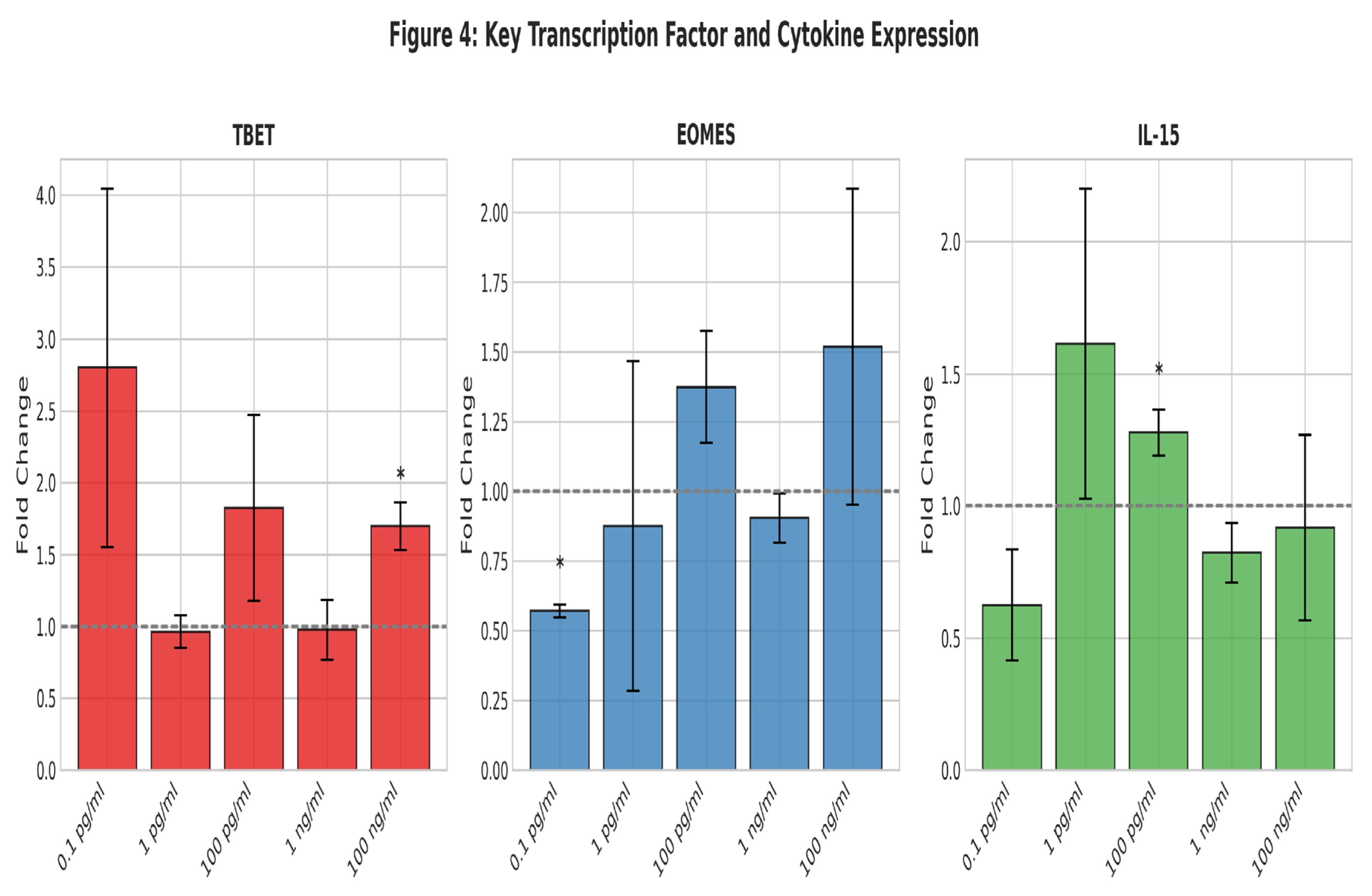
Transcription Factor Coordination: FOXO1, T-bet, and EOMES
Nuclear Receptor Signaling in NK Cell Development
Yamanaka Factors and Cellular Reprogramming Potential
Modulation of Inhibitory Receptor NKG2A
Anti-Cancer Therapeutic Implications
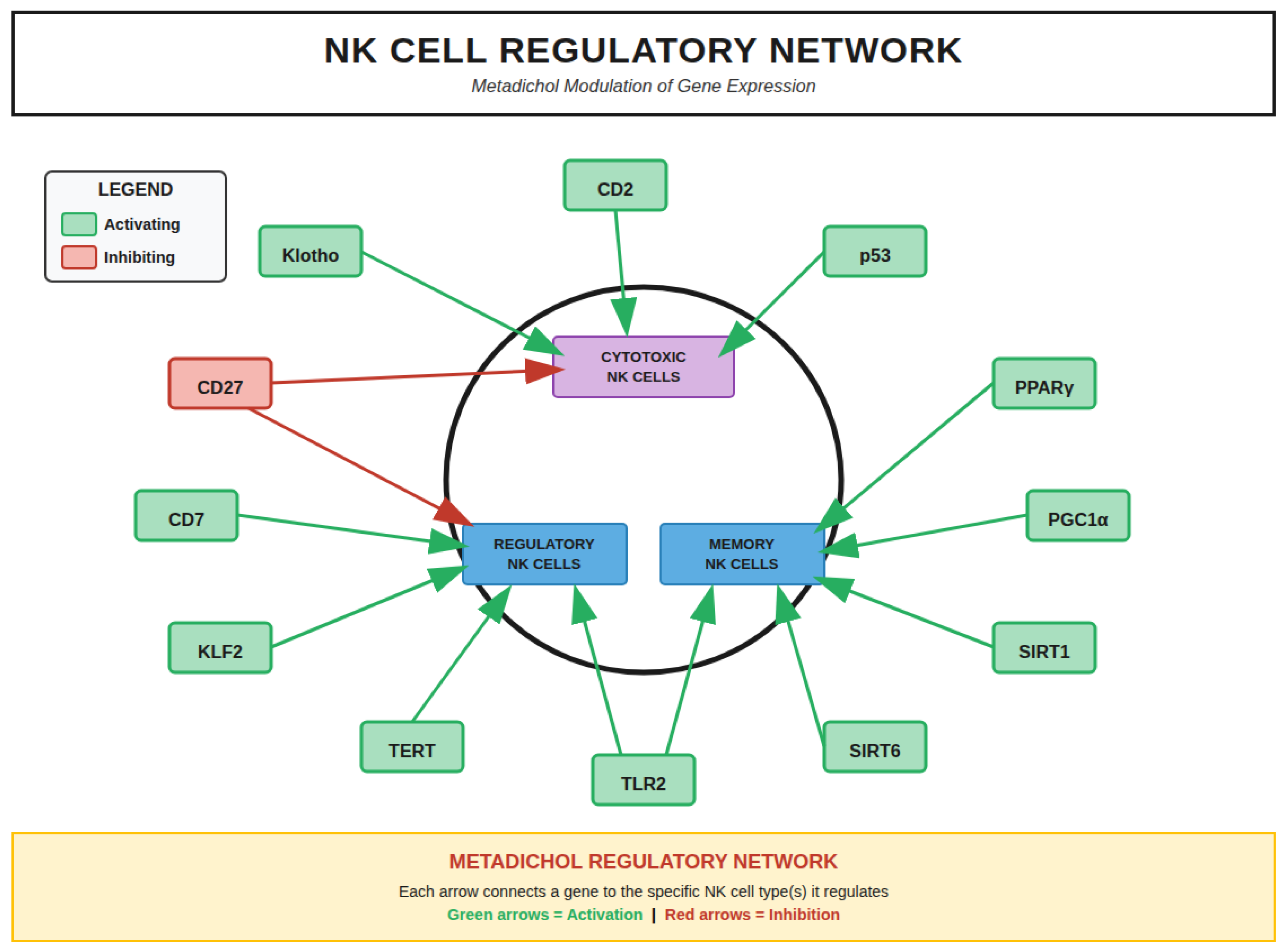
Integrated Mechanism of Metadichol Action

Conclusions
Supplementary Materials
Abbreviations
| Abbreviation | Full Name |
| CD7 | Cluster of Differentiation 7 |
| CD10 | Neprilysin/Membrane Metalloendopeptidase (MME) |
| CD34 | Hematopoietic Progenitor Cell Antigen CD34 |
| CD38 | Cyclic ADP Ribose Hydrolase |
| CD45RA | Protein Tyrosine Phosphatase Receptor Type C (naive isoform) |
| CD56 | Neural Cell Adhesion Molecule (NCAM) |
| CD117 | c-Kit; Stem Cell Factor Receptor |
| CD122 | Interleukin-2 Receptor Beta (IL-2Rβ) |
| CD127 | Interleukin-7 Receptor Alpha (IL-7Rα) |
| CD132 | Common Gamma Chain (γc) |
| CD244 | 2B4/SLAMF4 |
| CD314 | NKG2D/KLRK1 |
| CD336 | NKp44/NCR2 |
| CD337 | NKp30/NCR3 |
| IL1R1 | Interleukin-1 Receptor Type 1 |
| NCR | Natural Cytotoxicity Receptor |
| NKG2A | Natural Killer Group 2 Member A (CD159a/KLRC1) |
| NKG2D | Natural Killer Group 2 Member D (CD314/KLRK1) |
| NKp30 | Natural Killer Cell p30-Related Protein (NCR3/CD337) |
| NKp44 | Natural Killer Cell p44-Related Protein (NCR2/CD336) |
| NKp46 | Natural Killer Cell p46-Related Protein (NCR1/CD335) |
| NKp80 | Natural Killer Cell p80-Related Protein (KLRF1) |
| Abbreviation | Full Name |
| MICA | MHC Class I Chain-Related Protein A |
| MICB | MHC Class I Chain-Related Protein B |
| ULBPs | UL16-Binding Proteins (ULBP1-6) |
| Abbreviation | Full Name |
| IFN-γ | Interferon-Gamma |
| IL-2 | Interleukin-2 |
| IL-7 | Interleukin-7 |
| IL-15 | Interleukin-15 |
| SCF | Stem Cell Factor |
| TNF-α | Tumor Necrosis Factor-Alpha |
| Abbreviation | Full Name |
| ATF4 | Activating Transcription Factor 4 |
| E4BP4 | E4 Promoter-Binding Protein 4 (NFIL3) |
| EOMES | Eomesodermin |
| FOXO1 | Forkhead Box O1 |
| FOXO3 | Forkhead Box O3 |
| IRF-1 | Interferon Regulatory Factor 1 |
| KLF2 | Krüppel-Like Factor 2 |
| KLF4 | Krüppel-Like Factor 4 |
| NF-κB | Nuclear Factor Kappa-Light-Chain-Enhancer of Activated B Cells |
| OCT4 | Octamer-Binding Transcription Factor 4 |
| SOX2 | SRY-Box Transcription Factor 2 |
| STAT5 | Signal Transducer and Activator of Transcription 5 |
| T-bet | T-Box Transcription Factor 21 (TBX21) |
| TP53 | Tumor Protein P53 |
| Abbreviation | Full Name |
| JAK1 | Janus Kinase 1 |
| JAK3 | Janus Kinase 3 |
| mTOR | Mechanistic Target of Rapamycin |
| mTORC1 | Mechanistic Target of Rapamycin Complex 1 |
| mTORC2 | Mechanistic Target of Rapamycin Complex 2 |
| REDD1 | Regulated in Development and DNA Damage Responses 1 (DDIT4) |
| TBK1 | TANK-Binding Kinase 1 |
| TRIF | TIR-Domain-Containing Adapter-Inducing Interferon-β |
| Abbreviation | Full Name |
| PPAR | Peroxisome Proliferator-Activated Receptor |
| PPARγ | Peroxisome Proliferator-Activated Receptor Gamma |
| RAR | Retinoic Acid Receptor |
| VDR | Vitamin D Receptor |
| Abbreviation | Full Name |
| TLR | Toll-Like Receptor |
| TLR2 | Toll-Like Receptor 2 |
| TLR3 | Toll-Like Receptor 3 |
| TLR4 | Toll-Like Receptor 4 |
| Abbreviation | Full Name |
| ACTB | Actin Beta |
| B2M | Beta-2-Microglobulin |
| GAPDH | Glyceraldehyde-3-Phosphate Dehydrogenase |
| HPRT1 | Hypoxanthine Phosphoribosyltransferase 1 |
| Abbreviation | Full Name |
| CAR-NK | Chimeric Antigen Receptor Natural Killer Cell |
| CLP | Common Lymphoid Progenitor |
| HSC | Hematopoietic Stem Cell |
| iNK | Immature Natural Killer Cell |
| NK | Natural Killer |
| NKP | Natural Killer Progenitor |
| PBMC | Peripheral Blood Mononuclear Cell |
| Abbreviation | Full Name |
| cDNA | Complementary DNA |
| EDTA | Ethylenediaminetetraacetic Acid |
| FBS | Fetal Bovine Serum |
| PBS | Phosphate-Buffered Saline |
| qRT-PCR | Quantitative Real-Time Polymerase Chain Reaction |
| RNA | Ribonucleic Acid |
| SEM | Standard Error of the Mean |
| Abbreviation | Full Name |
| AML | Acute Myeloid Leukemia |
| DAMP | Damage-Associated Molecular Pattern |
| HLA-E | Human Leukocyte Antigen-E |
| MHC | Major Histocompatibility Complex |
References
- Arachchige, A.S.P.M. Human NK cells: From development to effector functions. J. Endotoxin Res. 2021, 27, 212–229. [Google Scholar] [CrossRef]
- Kucuksezer, U.C.; Cetin, E.A.; Esen, F.; Tahrali, I.; Akdeniz, N.; Gelmez, M.Y.; Deniz, G. The Role of Natural Killer Cells in Autoimmune Diseases. Front. Immunol. 2021, 12, 622306. [Google Scholar] [CrossRef]
- Paul, S.; Lal, G. The Molecular Mechanism of Natural Killer Cells Function and Its Importance in Cancer Immunotherapy. Front. Immunol. 2017, 8, 1124. [Google Scholar] [CrossRef] [PubMed]
- Abel, A.M.; Yang, C.; Thakar, M.S.; Malarkannan, S. Natural Killer Cells: Development, Maturation, and Clinical Utilization. Front. Immunol. 2018, 9, 1869. [Google Scholar] [CrossRef]
- Abel, A.M.; Yang, C.; Thakar, M.S.; Malarkannan, S. Natural Killer Cells: Development, Maturation, and Clinical Utilization. Front. Immunol. 2018, 9, 1869. [Google Scholar] [CrossRef]
- Cichocki, F.; Grzywacz, B.; Miller, J.S. Human NK Cell Development: One Road or Many? Front. Immunol. 2019, 10, 2078. [Google Scholar] [CrossRef] [PubMed]
- Cichocki, F.; Grzywacz, B.; Miller, J.S. Human NK Cell Development: One Road or Many? Front. Immunol. 2019, 10, 2078. [Google Scholar] [CrossRef]
- Eissens, D.N.; Spanholtz, J.; Van Der Meer, A.; Van Cranenbroek, B.; Dolstra, H.; Kwekkeboom, J.; Preijers, F.W.M.B.; Joosten, I. Defining Early Human NK Cell Developmental Stages in Primary and Secondary Lymphoid Tissues. PLOS ONE 2012, 7, e30930. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Wang, S.; Xin, J.; Wang, J.; Yao, C.; Zhang, Z. Role of NKG2D and its ligands in cancer immunotherapy. 2019, 9, 2064–2078. [Google Scholar]
- Wensveen, F.M.; Jelenčić, V.; Polić, B. NKG2D: A Master Regulator of Immune Cell Responsiveness. Front. Immunol. 2018, 9, 441. [Google Scholar] [CrossRef]
- Zhou, X.; Hao, Q.; Zhang, Q.; et al. Interleukin-2 (IL-2) interacts with IL-2 receptor beta (IL-2Rβ). Front Immunol. 2020, 11, 584983. [Google Scholar] [CrossRef]
- Liu, J.; Chen, D.; Nie, G.D.; et al. CD8+CD122+ T-cells: a newly emerging regulator with immunosuppressive function. Mediators Inflamm. 2015, 2015, 690190. [Google Scholar] [CrossRef]
- Keller, H.R.; Srivastava, S.; Hombach, A.; Spranger, S.; Cendales, L.; Bhavsar, J.D. The abundance and availability of cytokine receptor IL-2Rβ (CD122) constrain the lymphopenia-induced homeostatic proliferation. J Immunol. 2020, 204, 3227–3235. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Tian, Z.; Wei, H. Developmental and Functional Control of Natural Killer Cells by Cytokines. Front. Immunol. 2017, 8, 930. [Google Scholar] [CrossRef] [PubMed]
- Mathews, D.V.; Dong, Y.; Higginbotham, L.B.; Kim, S.C.; Breeden, C.P.; Stobert, E.A.; Jenkins, J.; Tso, J.Y.; Larsen, C.P.; Adams, A.B. CD122 signaling in CD8+ memory T cells drives costimulation-independent rejection. J. Clin. Investig. 2018, 128, 4557–4572. [Google Scholar] [CrossRef]
- Villarreal, D.O.; L’Huillier, A.; Bhardwaj, N.; et al. Targeting of CD122 enhances antitumor immunity by modulating the adaptive immune response. J Clin Invest. 2017, 127, 4307–4319. [Google Scholar] [CrossRef]
- Barros, P.O.; Berthoud, T.K.; Aloufi, N.; Angel, J.B. Soluble IL-7Rα/sCD127 in health, disease, and its potential role as a therapeutic agent. Immunotargets Ther. 2021, 10, 47–62. [Google Scholar] [CrossRef]
- Liu, W.; Putnam, A.L.; Xu-Yu, Z.; Szot, G.L.; Lee, M.R.; Zhu, S.; Gottlieb, P.A.; Kapranov, P.; Gingeras, T.R.; de St Groth, B.F.; et al. CD127 expression inversely correlates with FoxP3 and suppressive function of human CD4+ T reg cells. J. Exp. Med. 2006, 203, 1701–1711. [Google Scholar] [CrossRef] [PubMed]
- Carrette, F.; Surh, C.D. IL-7 signaling and CD127 receptor regulation in the control of T cell homeostasis. Semin. Immunol. 2012, 24, 209–217. [Google Scholar] [CrossRef]
- Galarza-Muñoz, G.; Briggs, F.B.; Evsyukova, I.; Schott-Lerner, G.; Kennedy, E.M.; Nyanhete, T.; Wang, L.; Bergamaschi, L.; Widen, S.G.; Tomaras, G.D.; et al. Human Epistatic Interaction Controls IL7R Splicing and Increases Multiple Sclerosis Risk. Cell 2017, 169, 72–84.e13. [Google Scholar] [CrossRef]
- Cottini, F.; Rodriguez, J.; Hughes, T.; Sharma, N.; Guo, L.; Lozanski, G.; Liu, B.; Cocucci, E.; Yang, Y.; Benson, D. Redefining CD56 as a Biomarker and Therapeutic Target in Multiple Myeloma. Mol. Cancer Res. 2022, 20, 1083–1095. [Google Scholar] [CrossRef]
- Van Acker, H.H.; Capsomidis, A.; Smits, E.L.; Van Tendeloo, V.F. CD56 in the Immune System: More Than a Marker for Cytotoxicity? Front. Immunol. 2017, 8, 892–892. [Google Scholar] [CrossRef]
- Poznanski, S.M.; Ashkar, A.A. Shining light on the significance of NK cell CD56 brightness. Cell. Mol. Immunol. 2018, 15, 1071–1073. [Google Scholar] [CrossRef] [PubMed]
- Gunesch, J.T.; Dixon, A.L.; Ebrahim, T.A.; Berrien-Elliott, M.M.; Tatineni, S.; Kumar, T.; Hegewisch-Solloa, E.; A Fehniger, T.; Mace, E.M. CD56 regulates human NK cell cytotoxicity through Pyk2. eLife 2020, 9, e57346. [Google Scholar] [CrossRef]
- Gunesch, J.T.; Orange, J.S. CD56/NCAM mediates cell migration of human NK cells by regulating cell polarity and actin flow. Mol Biol Cell. 2024, 35, ar45. [Google Scholar] [CrossRef]
- Ziegler, S.; Weiss, E.; Schmitt, A.-L.; Schlegel, J.; Burgert, A.; Terpitz, U.; Sauer, M.; Moretta, L.; Sivori, S.; Leonhardt, I.; et al. CD56 Is a Pathogen Recognition Receptor on Human Natural Killer Cells. Sci. Rep. 2017, 7, 5874. [Google Scholar] [CrossRef] [PubMed]
- Rabinowich, H.; Pricop, L.; Herberman, R.B.; Whiteside, T.L. Expression and function of CD7 molecule on human natural killer cells. J. Immunol. 1994, 152, 517–526. [Google Scholar] [CrossRef]
- Aandahl, E.M.; Sandberg, J.K.; Beckerman, K.P.; Taskén, K.; Moretto, W.J.; Nixon, D.F. CD7 Is a Differentiation Marker That Identifies Multiple CD8 T Cell Effector Subsets. J. Immunol. 2003, 170, 2349–2355. [Google Scholar] [CrossRef]
- Piedra-Quintero, Z.L.; Wilson, Z.; Nava, P.; Guerau-De-Arellano, M. CD38: An Immunomodulatory Molecule in Inflammation and Autoimmunity. Front. Immunol. 2020, 11, 597959. [Google Scholar] [CrossRef]
- Glaría, E.; Valledor, A.F. Roles of CD38 in the Immune Response to Infection. Cells 2020, 9, 228. [Google Scholar] [CrossRef]
- Li, W.; Li, Y.; Jin, X.; Liao, Q.; Chen, Z.; Peng, H.; Zhou, Y. CD38: A Significant Regulator of Macrophage Function. Front. Oncol. 2022, 12, 775649. [Google Scholar] [CrossRef] [PubMed]
- Polano, M.; Fabbiani, E.; Andreuzzi, E.; Di Cintio, F.; Bedon, L.; Gentilini, D.; Mongiat, M.; Ius, T.; Arcicasa, M.; Skrap, M.; et al. A New Epigenetic Model to Stratify Glioma Patients According to Their Immunosuppressive State. Cells 2021, 10, 576. [Google Scholar] [CrossRef]
- Shah, K.G.; Wu, R.; Jacob, A.; Blau, S.A.; Ji, Y.; Dong, W.; Marini, C.P.; Ravikumar, T.S.; Coppa, G.F.; Wang, P. Human Ghrelin Ameliorates Organ Injury and Improves Survival after Radiation Injury Combined with Severe Sepsis. Mol. Med. 2009, 15, 407–414. [Google Scholar] [CrossRef] [PubMed]
- Camponeschi, A.; Kläsener, K.; Sundell, T.; Lundqvist, C.; Manna, P.T.; Ayoubzadeh, N.; Sundqvist, M.; Thorarinsdottir, K.; Gatto, M.; Visentini, M.; et al. Human CD38 regulates B cell antigen receptor dynamic organization in normal and malignant B cells. J. Exp. Med. 2022, 219, e20220201. [Google Scholar] [CrossRef]
- Gurney, M.; Gornalusse, G.G.; Wang, X.; et al. CD38 contributes to human natural killer cell responses to multiple myeloma. bioRxiv. 2019. [Google Scholar] [CrossRef]
- Pulliam, T.; Jani, S.; Jing, L.; Ryu, H.; Jojic, A.; Shasha, C.; Zhang, J.; Kulikauskas, R.; Church, C.; Garnett-Benson, C.; et al. Circulating cancer-specific CD8 T cell frequency is associated with response to PD-1 blockade in Merkel cell carcinoma. Cell Rep. Med. 2024, 5, 101412. [Google Scholar] [CrossRef]
- Jiao, Y.; Yi, M.; Xu, L.; Chu, Q.; Yan, Y.; Luo, S.; Wu, K. CD38: targeted therapy in multiple myeloma and therapeutic potential for solid cancers. Expert Opin. Investig. Drugs 2020, 29, 1295–1308. [Google Scholar] [CrossRef] [PubMed]
- van de Donk, N.W.C.J.; Richardson, P.G.; Malavasi, F. CD38 antibodies in multiple myeloma: back to the future. Blood 2018, 131, 13–29. [Google Scholar] [CrossRef]
- Al Barashdi, M.A.; Ali, A.; McMullin, M.F.; Mills, K. Protein tyrosine phosphatase receptor type C (PTPRC or CD45). J. Clin. Pathol. 2021, 74, 548–552. [Google Scholar] [CrossRef]
- Krzywinska, E.; Cornillon, A.; Allende-Vega, N.; Vo, D.-N.; Rene, C.; Lu, Z.-Y.; Pasero, C.; Olive, D.; Fegueux, N.; Ceballos, P.; et al. CD45 Isoform Profile Identifies Natural Killer (NK) Subsets with Differential Activity. PLOS ONE 2016, 11, e0150434. [Google Scholar] [CrossRef]
- Henson, S.M.; E Riddell, N.; Akbar, A.N. Properties of end-stage human T cells defined by CD45RA re-expression. Curr. Opin. Immunol. 2012, 24, 476–481. [Google Scholar] [CrossRef]
- Hang, J.; Liu, Y.; Zhang, Y.; et al. The clinical implication of CD45RA+ naïve T cells in advanced pancreatic cancer. J Transl Med. 2019, 17, 97. [Google Scholar] [CrossRef]
- Tian, Y.; Babor, M.; Lane, J.; Schulten, V.; Patil, V.S.; Seumois, G.; Rosales, S.L.; Fu, Z.; Picarda, G.; Burel, J.; et al. Unique phenotypes and clonal expansions of human CD4 effector memory T cells re-expressing CD45RA. Nat. Commun. 2017, 8, 1703. [Google Scholar] [CrossRef] [PubMed]
- McGuire, D.J.; Akondy, R.S.; Yang, S.; Edupuganti, S.; Nagar, S.; Michael, G.; De Rosa, S.C.; Newell, E.W.; Farber, D.L.; Kissick, H.T.; et al. Regulation of CD45 isoforms during human effector and memory CD8 T cell differentiation: Implications for T cell nomenclature. Proc. Natl. Acad. Sci. 2025, 122, e2322982122. [Google Scholar] [CrossRef]
- Miettinen, M.; Lasota, J. KIT (CD117): A Review on Expression in Normal and Neoplastic Tissues, and Mutations and Their Clinicopathologic Correlation. Appl. Immunohistochem. Mol. Morphol. 2005, 13, 205–220. [Google Scholar] [CrossRef] [PubMed]
- Foster, B.M.; Zaidi, D.; Young, T.R.; Mobley, M.E.; Kerr, B.A. CD117/c-kit in Cancer Stem Cell-Mediated Progression and Therapeutic Resistance. Biomedicines 2018, 6, 31. [Google Scholar] [CrossRef]
- Cardoso, H.J.; Figueira, M.I.; Socorro, S. The stem cell factor (SCF)/c-KIT signalling in testis and prostate cancer. J. Cell Commun. Signal. 2017, 11, 297–307. [Google Scholar] [CrossRef]
- Rönnstrand, L. Signal transduction via the stem cell factor receptor/c-Kit. Cell. Mol. Life Sci. 2004, 61, 2535–2548. [Google Scholar] [CrossRef]
- Czechowicz, A.; Palchaudhuri, R.; Sber, M.; et al. Selective hematopoietic stem cell ablation using CD117-targeting. Nat Biotechnol. 2019, 37, 1192–1201. [Google Scholar] [CrossRef]
- Sidney, L.E.; Branch, M.J.; Dunphy, S.E.; Dua, H.S.; Hopkinson, A. Concise Review: Evidence for CD34 as a Common Marker for Diverse Progenitors. Stem Cells 2014, 32, 1380–1389. [Google Scholar] [CrossRef]
- Radu, P.; Zurzu, M.; Paic, V.; Bratucu, M.; Garofil, D.; Tigora, A.; Georgescu, V.; Prunoiu, V.; Pasnicu, C.; Popa, F.; et al. CD34—Structure, Functions and Relationship with Cancer Stem Cells. Medicina 2023, 59, 938. [Google Scholar] [CrossRef]
- AbuSamra, D.B.; Aleely, F.; Wu, Y.; et al. Not just a marker: CD34 on human hematopoietic stem/progenitor cells. Blood. 2017, 129, 1568–1578. [Google Scholar] [CrossRef]
- Hassanpour, M.; Salybekov, A.A.; Kobayashi, S.; Asahara, T. CD34 positive cells as endothelial progenitor cells in biology and medicine. Front. Cell Dev. Biol. 2023, 11, 1128134. [Google Scholar] [CrossRef]
- Merkle, F.T.; Eggan, K. Modeling Human Disease with Pluripotent Stem Cells: from Genome Association to Function. Cell Stem Cell 2013, 12, 656–668. [Google Scholar] [CrossRef] [PubMed]
- Tan, B.B.J.-W.; Chan, L.-L.; Tan, E.-K. Perceived Cognitive Changes following Chimeric Antigen Receptor T Cell Therapy in Lymphoma: Perceptual Anticipation? Biol. Blood Marrow Transplant. 2022, 29, 64. [Google Scholar] [CrossRef]
- Freud, A.G.; Keller, K.A.; Scoville, S.D.; Mundy-Bosse, B.L.; Cheng, S.; Youssef, Y.; Hughes, T.; Zhang, X.; Mo, X.; Porcu, P.; et al. NKp80 Defines a Critical Step during Human Natural Killer Cell Development. Cell Rep. 2016, 16, 379–391. [Google Scholar] [CrossRef]
- Welte, S.; Kuttruff, S.; Waldhauer, I.; Steinle, A. Mutual activation of natural killer cells and monocytes mediated by NKp80-AICL interaction. Nat. Immunol. 2006, 7, 1334–1342. [Google Scholar] [CrossRef] [PubMed]
- Morán-Plata, F.J.; Muñoz-García, N.; González-González, M.; Pozo, J.; Carretero-Domínguez, S.; Mateos, S.; Barrena, S.; Belhassen-García, M.; Lau, C.; Teixeira, M.D.A.; et al. A novel NKp80-based strategy for universal identification of normal, reactive and tumor/clonal natural killer-cells in blood. Front. Immunol. 2024, 15, 1423689. [Google Scholar] [CrossRef]
- Sun, L.; Gang, X.; Li, Z.; Zhao, X.; Zhou, T.; Zhang, S.; Wang, G. Advances in Understanding the Roles of CD244 (SLAMF4) in Immune Regulation and Associated Diseases. Front. Immunol. 2021, 12, 648182. [Google Scholar] [CrossRef]
- Agresta, L.; Hoebe, K.H.N.; Janssen, E.M. The Emerging Role of CD244 Signaling in Immune Cells of the Tumor Microenvironment. Front. Immunol. 2018, 9, 2809. [Google Scholar] [CrossRef]
- Deng, Z.; Liu, Y.; Zhou, H. Distinct roles of CD244 expression in cancer diagnosis and prognosis: A pan-cancer analysis. Heliyon 2024, 10, e28928. [Google Scholar] [CrossRef]
- Wang, X.; Xiong, H.; Ning, Z. Implications of NKG2A in immunity and immune-mediated diseases. Front. Immunol. 2022, 13, 960852. [Google Scholar] [CrossRef]
- Chen, D.G.; Xie, J.; Choi, J.; Ng, R.H.; Zhang, R.; Li, S.; Edmark, R.; Zheng, H.; Solomon, B.; Campbell, K.M.; et al. Integrative systems biology reveals NKG2A-biased immune responses correlate with protection in infectious disease, autoimmune disease, and cancer. Cell Rep. 2024, 43, 113872. [Google Scholar] [CrossRef]
- Horton, N.C.; Mathew, P.A. NKp44 and Natural Cytotoxicity Receptors as Damage-Associated Molecular Pattern Recognition Receptors. Front. Immunol. 2015, 6, 31. [Google Scholar] [CrossRef]
- Parodi, M.; Favoreel, H.; Candiano, G.; Gaggero, S.; Sivori, S.; Mingari, M.C.; Moretta, L.; Vitale, M.; Cantoni, C. NKp44-NKp44 Ligand Interactions in the Regulation of Natural Killer Cells and Other Innate Lymphoid Cells in Humans. Front. Immunol. 2019, 10, 719. [Google Scholar] [CrossRef]
- Mair, K.H.; Crossman, A.J.; Wagner, B.; Babasyan, S.; Noronha, L.; Boyd, P.; Zarlenga, D.; Stadler, M.; van Dongen, K.A.; Gerner, W.; et al. The Natural Cytotoxicity Receptor NKp44 (NCR2, CD336) Is Expressed on the Majority of Porcine NK Cells Ex Vivo Without Stimulation. Front. Immunol. 2022, 13, 767530. [Google Scholar] [CrossRef] [PubMed]
- Eisenberg, V.; Shamalov, K.; Meir, S.; Hoogi, S.; Sarkar, R.; Pinker, S.; Markel, G.; Porgador, A.; Cohen, C.J. Targeting Multiple Tumors Using T-Cells Engineered to Express a Natural Cytotoxicity Receptor 2-Based Chimeric Receptor. Front. Immunol. 2017, 8, 1212. [Google Scholar] [CrossRef]
- Murayama, Y.; Hirayama, A.V.; Houchins, J.P.; et al. NKp44-based chimeric antigen receptor effectively targets synovial sarcomas. J Immunother Cancer. 2022, 10, e004947. [Google Scholar] [CrossRef]
- Sarfraz, M.; Afzal, A.; Raza, S.M.; Bashir, S.; Madni, A.; Khan, M.W.; Ma, X.; Xiang, G. Liposomal co-delivered oleanolic acid attenuates doxorubicin-induced multi-organ toxicity in hepatocellular carcinoma. Oncotarget 2017, 8, 47136–47153. [Google Scholar] [CrossRef]
- Cui, Z.J. Deep-sea photodynamic vision at low light level — Which is more important, prosthetic retinal or apo-rhodopsin moiety? FASEB J. 2025, 39, e70470. [Google Scholar] [CrossRef] [PubMed]
- Freud, A.G.; Zhao, S.; Wei, S.; Gitana, G.M.; Molina-Kirsch, H.F.; Atwater, S.K.; Natkunam, Y. Expression of the Activating Receptor, NKp46 (CD335), in Human Natural Killer and T-Cell Neoplasia. Am. J. Clin. Pathol. 2013, 140, 853–866. [Google Scholar] [CrossRef]
- Cheminant, M.; Bruneau, J.; Malamut, G.; et al. NKp46 is a diagnostic biomarker in gastrointestinal T-cell lymphoproliferative diseases. Gut. 2019, 68, 1396–1406. [Google Scholar] [CrossRef]
- Sica, M.; Piazzolla, P.; Amparore, D.; Verri, P.; De Cillis, S.; Piramide, F.; Volpi, G.; Piana, A.; Di Dio, M.; Alba, S.; et al. 3D Model Artificial Intelligence-Guided Automatic Augmented Reality Images during Robotic Partial Nephrectomy. Diagnostics 2023, 13, 3454. [Google Scholar] [CrossRef] [PubMed]
- Gasse, P.; Mary, C.; Guenon, I.; Noulin, N.; Charron, S.; Schnyder-Candrian, S.; Schnyder, B.; Akira, S.; Quesniaux, V.F.; Lagente, V.; et al. IL-1R1/MyD88 signaling and the inflammasome are essential in pulmonary inflammation and fibrosis in mice. J. Clin. Investig. 2007, 117, 3786–3799. [Google Scholar] [CrossRef]
- Hughes, T.; Becknell, B.; Freud, A.G.; et al. IL-1β selectively expands IL-22+ immature human NK cells in secondary lymphoid tissue. Immunity. 2010, 32, 803–814. [Google Scholar] [CrossRef] [PubMed]
- Luís, J.P.; Simões, C.J.V.; Brito, R.M.M. The Therapeutic Prospects of Targeting IL-1R1 for the Modulation of Neuroinflammation in Central Nervous System Disorders. Int. J. Mol. Sci. 2022, 23, 1731. [Google Scholar] [CrossRef] [PubMed]
- Chen, I. Restoring order. Nat. Struct. Mol. Biol. 2012, 19, 267–267. [Google Scholar] [CrossRef]
- Raghavan, PR. Policosanol nanoparticles. US Patent 8,722,093, 2014. [Google Scholar]
- Raghavan, PR. Policosanol nanoparticles. US Patent 9,006,292, 2015. [Google Scholar]
- Raghavan, PR. Policosanol nanoparticles. US Patent 9,034,383, 2015. [Google Scholar]
- Raghavan, P.R. Metadichol® A Nano Lipid Emulsion that Expresses All 49 Nuclear Receptors in Stem and Somatic Cells. Arch. Clin. Biomed. Res. 2022, 7, 543–555. [Google Scholar] [CrossRef]
- Richards, J.; Danielson, A.; Stuart, R.; Richards, A.; Laurin, E. Reverse Remodeling of Methamphetamine-Associated Cardiomyopathy: An Update on Mechanisms for Recovery. Med Res. Arch. 2024, 12. [Google Scholar] [CrossRef]
- Raghavan, P.R. Beyond Rapamycin: Metadichol Represents a New Class of Multi-Target mTOR Modulators. Med Res. Arch. 2025, 13. [Google Scholar] [CrossRef]
- A Johnston, J.; Bacon, C.M.; Finbloom, D.S.; Rees, R.C.; Kaplan, D.; Shibuya, K.; Ortaldo, J.R.; Gupta, S.; Chen, Y.Q.; Giri, J.D. Tyrosine phosphorylation and activation of STAT5, STAT3, and Janus kinases by interleukins 2 and 15. Proc. Natl. Acad. Sci. 1995, 92, 8705–8709. [Google Scholar] [CrossRef] [PubMed]
- Miyazaki, T.; Kawahara, A.; Fujii, H.; Nakagawa, Y.; Minami, Y.; Liu, Z.-J.; Oishi, I.; Silvennoinen, O.; Witthuhn, B.A.; Ihle, J.N.; et al. Functional Activation of Jak1 and Jak3 by Selective Association with IL-2 Receptor Subunits. Science 1994, 266, 1045–1047. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, M.K.; Glaccum, M.; Brown, S.N.; Butz, E.A.; Viney, J.L.; Embers, M.; Matsuki, N.; Charrier, K.; Sedger, L.; Willis, C.R.; et al. Reversible Defects in Natural Killer and Memory Cd8 T Cell Lineages in Interleukin 15–Deficient Mice. J. Exp. Med. 2000, 191, 771–780. [Google Scholar] [CrossRef] [PubMed]
- Huntington, N.D.; Puthalakath, H.; Gunn, P.; Naik, E.; Michalak, E.M.; Smyth, M.J.; Tabarias, H.; A Degli-Esposti, M.; Dewson, G.; Willis, S.N.; et al. Interleukin 15–mediated survival of natural killer cells is determined by interactions among Bim, Noxa and Mcl-1. Nat. Immunol. 2007, 8, 856–863. [Google Scholar] [CrossRef]
- Freud, A.G.; Caligiuri, M.A. Human natural killer cell development. Immunol. Rev. 2006, 214, 56–72. [Google Scholar] [CrossRef]
- Freud, A.G.; Yu, J.; Caligiuri, M.A. Human natural killer cell development in secondary lymphoid tissues. Semin. Immunol. 2014, 26, 132–137. [Google Scholar] [CrossRef]
- Yu, J.; Freud, A.G.; Caligiuri, M.A. Location and cellular stages of natural killer cell development. Trends Immunol. 2013, 34, 573–582. [Google Scholar] [CrossRef]
- Marçais, A.; Cherfils-Vicini, J.; Viant, C.; Degouve, S.; Viel, S.; Fenis, A.; Rabilloud, J.; Mayol, K.; Tavares, A.; Bienvenu, J.; et al. The metabolic checkpoint kinase mTOR is essential for IL-15 signaling during the development and activation of NK cells. Nat. Immunol. 2014, 15, 749–757. [Google Scholar] [CrossRef]
- Yang, M.; Li, D.; Chang, Z.; Yang, Z.; Tian, Z.; Dong, Z. PDK1 orchestrates early NK cell development through induction of E4BP4 expression and maintenance of IL-15 responsiveness. J. Exp. Med. 2015, 212, 253–265. [Google Scholar] [CrossRef]
- Wang, F.; Meng, M.; Mo, B.; Yang, Y.; Ji, Y.; Huang, P.; Lai, W.; Pan, X.; You, T.; Luo, H.; et al. Crosstalks between mTORC1 and mTORC2 variagate cytokine signaling to control NK maturation and effector function. Nat. Commun. 2018, 9, 1–17. [Google Scholar] [CrossRef]
- Brugarolas, J.; Lei, K.; Hurley, R.L.; Manning, B.D.; Reiling, J.H.; Hafen, E.; Witters, L.A.; Ellisen, L.W.; Kaelin, W.G., Jr. Regulation of mTOR function in response to hypoxia by REDD1 and the TSC1/TSC2 tumor suppressor complex. Genes Dev. 2004, 18, 2893–2904. [Google Scholar] [CrossRef]
- Bauer, S.; Groh, V.; Wu, J.; Steinle, A.; Phillips, J.H.; Lanier, L.L.; Spies, T. Activation of NK Cells and T Cells by NKG2D, a Receptor for Stress-Inducible MICA. Science 1999, 285, 727–729. [Google Scholar] [CrossRef]
- Textor, S.; Fiegler, N.; Arnold, A.; Porgador, A.; Hofmann, T.G.; Cerwenka, A. Human NK Cells Are Alerted to Induction of p53 in Cancer Cells by Upregulation of the NKG2D Ligands ULBP1 and ULBP2. Cancer Res. 2011, 71, 5998–6009. [Google Scholar] [CrossRef] [PubMed]
- Welte, S.; Kuttruff, S.; Waldhauer, I.; Steinle, A. Mutual activation of natural killer cells and monocytes mediated by NKp80-AICL interaction. Nat. Immunol. 2006, 7, 1334–1342. [Google Scholar] [CrossRef]
- Freud, A.G.; Keller, K.A.; Scoville, S.D.; Mundy-Bosse, B.L.; Cheng, S.; Youssef, Y.; Hughes, T.; Zhang, X.; Mo, X.; Porcu, P.; et al. NKp80 Defines a Critical Step during Human Natural Killer Cell Development. Cell Rep. 2016, 16, 379–391. [Google Scholar] [CrossRef]
- Sivori, S.; Falco, M.; Della Chiesa, M.; Carlomagno, S.; Vitale, M.; Moretta, L.; Moretta, A. CpG and double-stranded RNA trigger human NK cells by Toll-like receptors: Induction of cytokine release and cytotoxicity against tumors and dendritic cells. Proc. Natl. Acad. Sci. USA 2004, 101, 10116–10121. [Google Scholar] [CrossRef]
- Girart, M.V.; Fuertes, M.B.; Domaica, C.I.; Rossi, L.E.; Zwirner, N.W. Engagement of TLR3, TLR7, and NKG2D Regulate IFN-γ Secretion but Not NKG2D-Mediated Cytotoxicity by Human NK Cells Stimulated with Suboptimal Doses of IL-12. J. Immunol. 2007, 179, 3472–3479. [Google Scholar] [CrossRef]
- Hart, O.M.; Athie-Morales, V.; O’Connor, G.M.; Gardiner, C.M. TLR7/8-Mediated Activation of Human NK Cells Results in Accessory Cell-Dependent IFN-γ Production. J. Immunol. 2005, 175, 1636–1642. [Google Scholar] [CrossRef] [PubMed]
- Gordon, S.M.; Chaix, J.; Rupp, L.J.; Wu, J.; Madera, S.; Sun, J.C.; Lindsten, T.; Reiner, S.L. The Transcription Factors T-bet and Eomes Control Key Checkpoints of Natural Killer Cell Maturation. Immunity 2012, 36, 55–67. [Google Scholar] [CrossRef] [PubMed]
- Simonetta, F.; Pradier, A.; Roosnek, E. T-bet and Eomesodermin in NK Cell Development, Maturation, and Function. Front. Immunol. 2016, 7, 241. [Google Scholar] [CrossRef]
- Deng, Y.; Kerdiles, Y.; Chu, J.; Yuan, S.; Wang, Y.; Chen, X.; Mao, H.; Zhang, L.; Zhang, J.; Hughes, T.; et al. Transcription Factor Foxo1 Is a Negative Regulator of Natural Killer Cell Maturation and Function. Immunity 2015, 42, 457–470. [Google Scholar] [CrossRef]
- Du, X.; De Almeida, P.; Manieri, N.; Nagata, D.D.A.; Wu, T.D.; Bowles, K.H.; Arumugam, V.; Schartner, J.; Cubas, R.; Mittman, S.; et al. CD226 regulates natural killer cell antitumor responses via phosphorylation-mediated inactivation of transcription factor FOXO1. Proc. Natl. Acad. Sci. USA 2018, 115, E11731–E11740. [Google Scholar] [CrossRef]
- Rao, R.R.; Li, Q.; Bupp, M.R.G.; Shrikant, P.A. Transcription Factor Foxo1 Represses T-bet-Mediated Effector Functions and Promotes Memory CD8+ T Cell Differentiation. Immunity 2012, 36, 374–387. [Google Scholar] [CrossRef]
- Rhee, E.J.; Kwon, H.; Lee, S. Regulation of natural killer cell development and function by nuclear receptor signaling. Arch Pharm Res. 2018, 41, 527–538. [Google Scholar] [CrossRef]
- Charoenngam, N.; Holick, M.F. Immunologic Effects of Vitamin D on Human Health and Disease. Nutrients 2020, 12, 2097. [Google Scholar] [CrossRef] [PubMed]
- Konjević, G.M.; Vuletić, A.M.; Mirjačić Martinović, K.M.; Larsen, A.K.; Jurišić, V.B. The role of cytokines in the regulation of NK cells in the tumor environment. Cytokine 2019, 117, 30–40. [Google Scholar] [CrossRef]
- Takahashi, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Mouse Embryonic and Adult Fibroblast Cultures by Defined Factors. Cell 2006, 126, 663–676. [Google Scholar] [CrossRef] [PubMed]
- André, P.; Denis, C.; Soulas, C.; Bourbon-Caillet, C.; Lopez, J.; Arnoux, T.; Bléry, M.; Bonnafous, C.; Gauthier, L.; Morel, A.; et al. Anti-NKG2A mAb Is a Checkpoint Inhibitor that Promotes Anti-tumor Immunity by Unleashing Both T and NK Cells. Cell 2018, 175, 1731–1743.e13. [Google Scholar] [CrossRef]
- Long, E.O.; Kim, H.S.; Liu, D.; Peterson, M.E.; Rajagopalan, S. Controlling Natural Killer Cell Responses: Integration of Signals for Activation and Inhibition. Annu. Rev. Immunol. 2013, 31, 227–258. [Google Scholar] [CrossRef]
- Braud, V.M.; Allan, D.S.J.; O'Callaghan, C.A.; Söderström, K.; D'Andrea, A.; Ogg, G.S.; Lazetic, S.; Young, N.T.; Bell, J.I.; Phillips, J.H.; et al. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 1998, 391, 795–799. [Google Scholar] [CrossRef]
- van Hall, T.; André, P.; Horowitz, A.; Ruan, D.F.; Borst, L.; Zerbib, R.; Narni-Mancinelli, E.; van der Burg, S.H.; Vivier, E. Monalizumab: inhibiting the novel immune checkpoint NKG2A. J. Immunother. Cancer 2019, 7, 263. [Google Scholar] [CrossRef] [PubMed]
- Aleman, C.; Mas, R.; Hernandez, C.; Rodeiro, I.; Cerejido, E.; Noa, M.; Capote, A.; Menendez, R.; Amor, A.; Fraga, V. A 12-month study of policosanol oral toxicity in Sprague Dawley rats. Toxicol. Lett. 1994, 70, 77–87. [Google Scholar] [CrossRef] [PubMed]
- Alemán, C.L.; Ferreiro, R.M.; Puig, M.N.; Guerra, I.R.; Ortega, C.H.; Capote, A. Carcinogenicity of policosanol in sprague dawley rats: A 24 month study. Teratog. Carcinog. Mutagen. 1994, 14, 239–249. [Google Scholar] [CrossRef] [PubMed]
| Group | Cell Type | Treatment | Concentration |
|---|---|---|---|
| 1 |
Human PBMC |
Vehicle Control | — |
| 2 |
Metadichol |
0.1pg/mL | |
| 3 | 1 pg/mL | ||
| 4 | 100 pg/mL | ||
| 5 | 1 ng/mL | ||
| 6 | 100 ng/mL |
| Test concentrations | ||||||
| RNA yield (ng/µl) | 0 | 0.1 pg/ ml | 1 pg/ ml | 100 pg/ ml | 1 ng/ ml | 100 ng/ ml |
| Human PBMC’s | 425.2 | 410.4 | 380.9 | 412.8 | 438.6 | 446.2 |
| Gene | Primers | Amplicon size | Annealing temperature | ||
| NGK2A | F | GCCTCTGTGGTAACGATAGTTGT | 118 | 65°C | |
| R | ATCCACTCCTCAGGACAATGGC | ||||
| CD38 | F | TCTTGCCCAGACTGGAGAAAGG | 100 | 65°C | |
| R | TGGACCACATCACAGGCAGCTT | ||||
| CD117 | F | CACCGAAGGAGGCACTTACACA | 121 | 50°C | |
| R | TGCCATTCACGAGCCTGTCGTA | ||||
| CD122 | F | CTGGAGAGATGGCCACGGT | 182 | 53°C | |
| R | GATGCCCAAGAGGTAGCCAG | ||||
| CD336 | F | CTGAGTCTCCATCTACCATCCC | 116 | 50°C | |
| R | TCTTGGCTACGAGGAGTCCACA | ||||
| CD337 | F | CCAGCATCTACGTGTGCAGAGT | 135 | 65°C | |
| R | GCATAGAATCCAGCCCGAAGGA | ||||
| CD338 | F | GTTCTCAGCAGCTCTTCGGCTT | 144 | 65°C | |
| R | TCCTCCAGACACACCACGGATA | ||||
| NKP80 | F | CGAGATCTGCAGACCAGACA | 274 | 49°C | |
| R | CGAGATCTGCAGACCAGACA | ||||
| CD34 | F | CCACAGGAGAAAGGCTGGAG | 184 | 62°C | |
| R | ATCTGGTAAGCAGGGCTGTG | ||||
| CD10 | F | CTTTAGTGCCCAGCAGTCCAAC | 128 | 56°C | |
| R | CACCAGTCAACGAGGTCTCCAT | ||||
| GAPDH | F | GTCTCCTCTGACTTCAACAGCG | 186 | 65°C | |
| R | ACCACCCTGTTGCTGTAGCCAA | ||||
| ILIR1 | F | GTGCTTTGGTACAGGGATTCCTG | 120 | 65°C | |
| R | CACAGTCAGAGGTAGACCCTTC | ||||
| CD127 | F | ATCGCAGCACTCACTGACCTGT | 100 | 50°C | |
| R | TCAGGCACTTTACCTCCACGAG | ||||
| CD244 | F | TCTACTGCCTGGAGGTCACCAG | 151 | 53°C | |
| R | GACCAAGCAAGACAGAGCCACT | ||||
| CD45RA | F | CATGCAGCTAGCAAGTGGTT | 264 | 50°C | |
| R | GAAGGGCTCAGAGTGGTTGT | ||||
| CD56 | F | CAGTCCATAGCCCTCCTCCA | 483 | 50°C | |
| R | CGGCTTTTCCACACAGGTTG | ||||
| NKG2D | F | AGATCTTCCCTCTCTGAGCA | 114 | 67°C | |
| R | GAGACCTCCGACCACGAATC | ||||
| CD7 | F | TGTCGGACACTGGCACCTACAC | 114 | 57°C | |
| R | TCCGAGCATCTGTGCCATCCTT | ||||
| EOMES | F | AAATGGGTGACCTGTGGCAAAGC | 101 | 67 | |
| R | CTCCTGTCTCATCCAGTGGGAA | ||||
| IL15 | F | AACAGAAGCCAACTGGGTGAATG | 157 | 67 | |
| R | AACAGAAGCCAACTGGGTGAATG | ||||
| TBET | F | ATTGCCGTGACTGCCTACCAGA | 150 | 67 | |
| R | ATTGCCGTGACTGCCTACCAGA | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




