Submitted:
25 January 2026
Posted:
27 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Protein Production
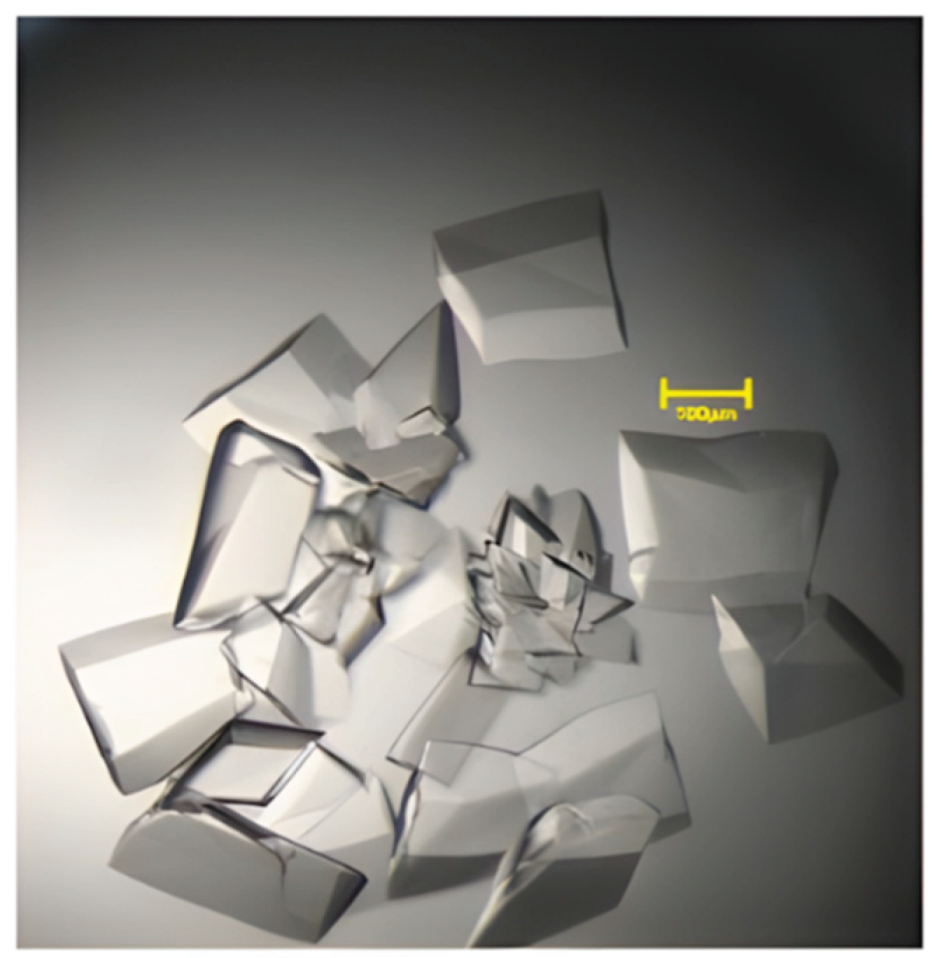
2.2. Crystallization
2.3. X-Ray Diffraction
2.4. Structure Determination
3. Results
FSH/anti-FSHα Fab/anti-kappa VHH
4. Discussion
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| cAMP CG ESI-MS |
Cyclic adenosine-monophosphate Choriogonadotropin Electrospray ionization mass spectrometry |
| FSHR GC GnRH HF LH |
FSH receptor Granulosa cells Gonadotropin-releasing hormone Hydrogen fluoride Luteinizing hormone |
| rFSH SEC |
Recombinant follicle stimulating hormone Size exclusion column |
| ST3 | α2,3-sialyltransferase |
| VHH | Anti-kappa nanobody |
References
- Ulloa-Aguirre A. Structure-function relationship of follicle-stimulating hormone and its receptor. Hum Reprod Update. 1998;4:260–83. [CrossRef]
- Leão R de BF, Esteves SC. Gonadotropin therapy in assisted reproduction: an evolutionary perspective from biologics to biotech. Clinics. 2014;69:279–93. [CrossRef]
- Bosnjak I, Bojovic V, Segvic-Bubic T, Bielen A. Occurrence of protein disulfide bonds in different domains of life: a comparison of proteins from the Protein Data Bank. Protein Engineering Design and Selection. 2014;27:65–72. [CrossRef]
- Feige MJ, Braakman I, Hendershot LM. CHAPTER 1.1. Disulfide Bonds in Protein Folding and Stability. 2018. p. 1–33. [CrossRef]
- Ulloa-Aguirre A, Timossi C, Damián-Matsumura P, Dias JA. Role of Glycosylation in Function of Follicle-Stimulating Hormone. Endocrine. 1999;11:205–16. [CrossRef]
- Zhang L, Xu H, Chen C-L, Green-Church KB, Freitas MA, Chen Y-R. Mass spectrometry profiles superoxide-induced intramolecular disulfide in the FMN-binding subunit of mitochondrial complex I. J Am Soc Mass Spectrom. 2008;19:1875–86. [CrossRef]
- Gorman JJ, Wallis TP, Pitt JJ. Protein disulfide bond determination by mass spectrometry. Mass Spectrom Rev. 2002;21:183–216. [CrossRef]
- CHMP. ANNEX I SUMMARY OF PRODUCT CHARACTERISTICS.
- Bosch E, Havelock J, Martin FS, Rasmussen BB, Klein BM, Mannaerts B, et al. Follitropin delta in repeated ovarian stimulation for IVF: a controlled, assessor-blind Phase 3 safety trial. Reprod Biomed Online. 2019;38:195–205. [CrossRef]
- Nyboe Andersen A, Nelson SM, Fauser BCJM, García-Velasco JA, Klein BM, Arce J-C, et al. Individualized versus conventional ovarian stimulation for in vitro fertilization: a multicenter, randomized, controlled, assessor-blinded, phase 3 noninferiority trial. Fertil Steril. 2017;107:387-396.e4. [CrossRef]
- US9771407.
- Tegoni M, Spinelli S, Verhoeyen M, Davis P, Cambillau C. Crystal structure of a ternary complex between human chorionic gonadotropin (hCG) and two Fv fragments specific for the α and β-subunits 1 1Edited by I. A. Wilson. J Mol Biol. 1999;289:1375–85. [CrossRef]
- Zhang Z, Lin X, Wei L, Wu Y, Xu L, Wu L, et al. A framework for Frizzled-G protein coupling and implications to the PCP signaling pathways. Cell Discov. 2024;10:3. [CrossRef]
- Bloch JS, Mukherjee S, Kowal J, Filippova E V., Niederer M, Pardon E, et al. Development of a universal nanobody-binding Fab module for fiducial-assisted cryo-EM studies of membrane proteins. Proceedings of the National Academy of Sciences. 2021;118. [CrossRef]
- Fan QR, Hendrickson WA. Structure of human follicle-stimulating hormone in complex with its receptor. Nature. 2005;433:269–77. [CrossRef]
- Lakbub JC, Shipman JT, Desaire H. Recent mass spectrometry-based techniques and considerations for disulfide bond characterization in proteins. Anal Bioanal Chem. 2018;410:2467–84. [CrossRef]
- Fox KM, Dias JA, Van Roey P. Three-Dimensional Structure of Human Follicle-Stimulating Hormone. Molecular Endocrinology. 2001;15:378–89. [CrossRef]
- Wu H, Lustbader JW, Liu Y, Canfield RE, Hendrickson WA. Structure of human chorionic gonadotropin at 2.6 å resolution from MAD analysis of the selenomethionyl protein. Structure. 1994;2:545–58. [CrossRef]
- Lapthorn AJ, Harris DC, Littlejohn A, Lustbader JW, Canfield RE, Machin KJ, et al. Crystal structure of human chorionic gonadotropin. Nature. 1994;369:455–61. [CrossRef]
- Lustbader JW, Birken S, Pileggi NF, Kolks MAG, Pollak S, Cuff ME, et al. Crystallization and characterization of human chorionic gonadotropin in chemically deglycosylated and enzymically desialylated states. Biochemistry. 1989;28:9239–43. [CrossRef]
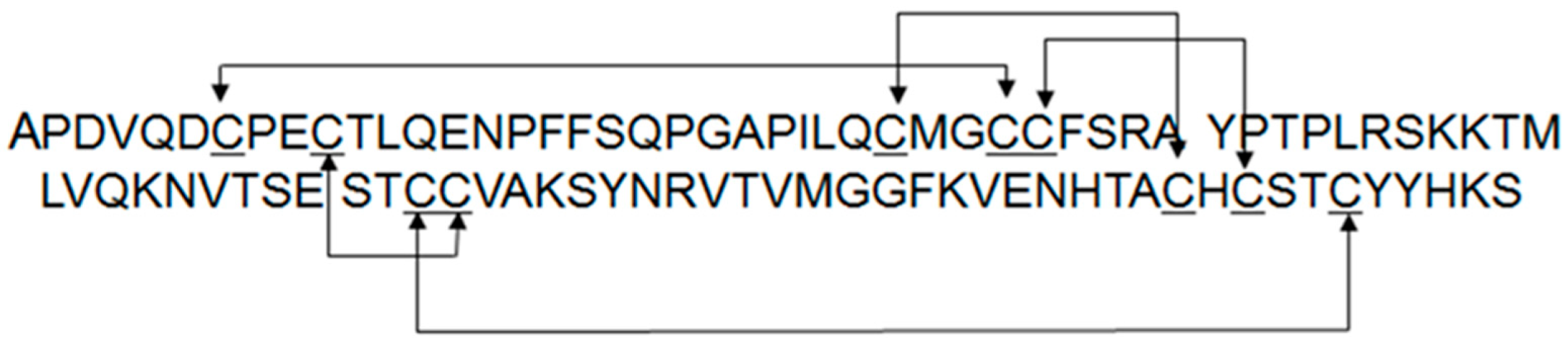
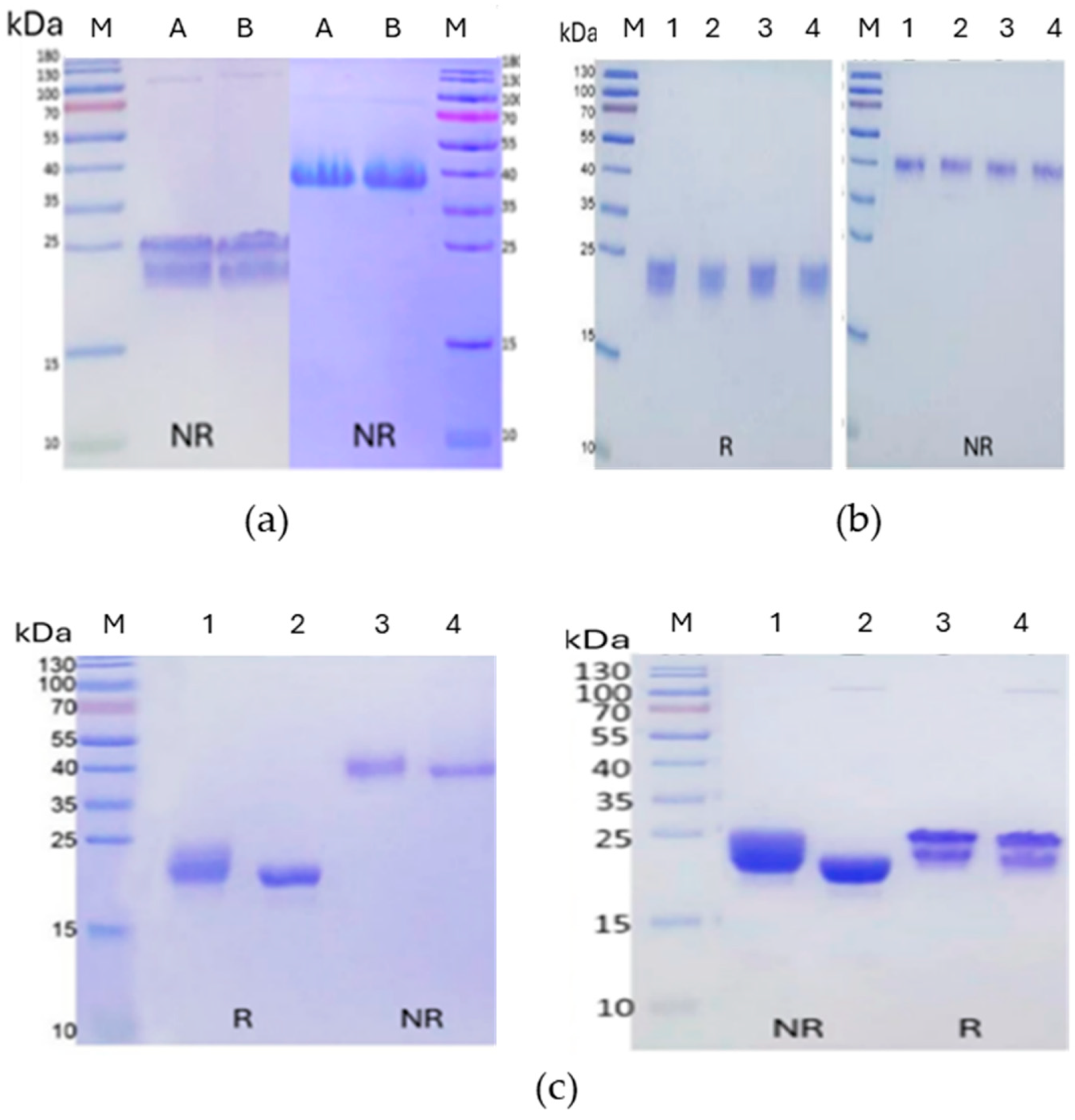
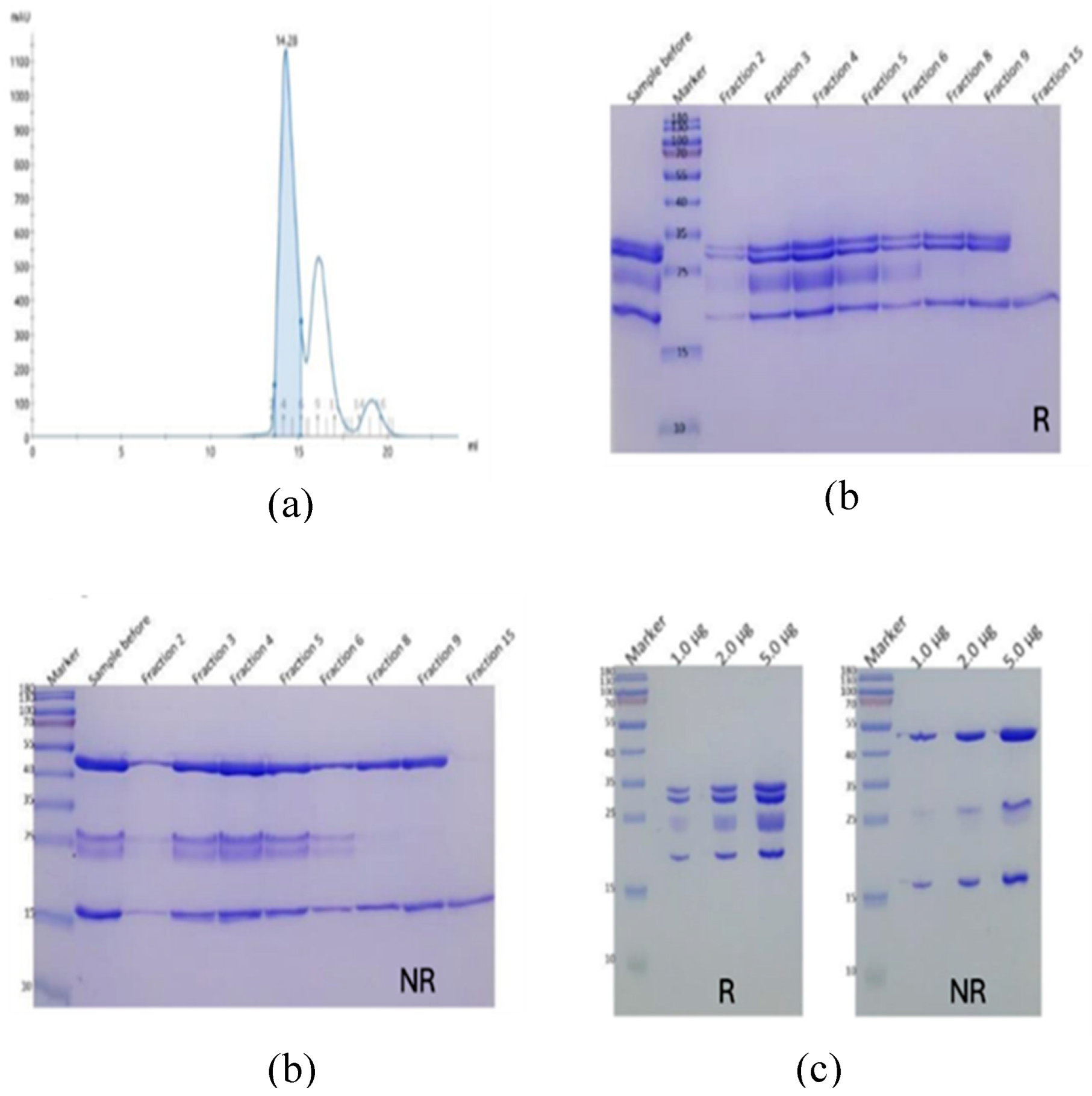

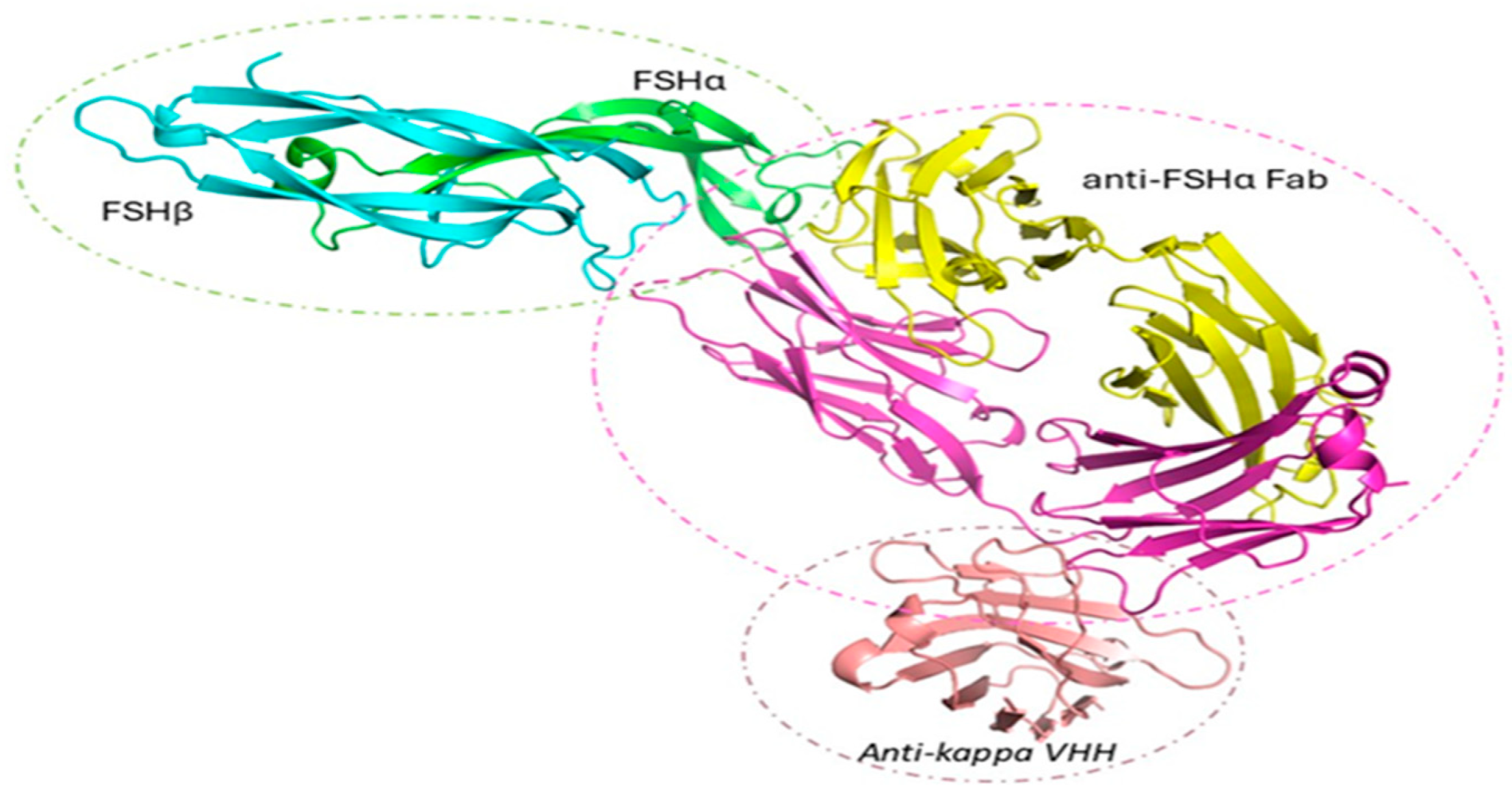

| Construct | Sequence |
|---|---|
| Anti-FSHα Fab LC |
MKHLWFFLLLVAAPRWVLSDIELTQSPDSLSVSLGQRATISCRASESVDSYGNSFMQWY QQKPGQPKLLIYRASNLEGSIPARFSGTGSRTDFTLTINPVEADDVATYYCQQSDEYPYM YTFGGGTKLEIKRTVAAPSVFIFPPSDEQLKSGTASVVCLLNNFYPREAKVQWKVDNALQSGN SQESVTEQDSKDSTYSLSSTLTLSKADYEKHKVYACEVTHQGLSSPVTKSFNRGEC |
| Anti-FSHα Fab HC |
MVLQTQVFISLLWISGAYGQVQLQQSGAELVKPGASVKLSCKASDYTFTSYWMHWVKQRP GQGLEWIGEINPTNGRTYYNEKFKSKATLTDVKSSSTAYMQLSSLTSEDSAVTTCATTYGNSFDY WGQGTTVTVSSASTKGPSVFPLAPSSKSTSGGTAALGCLVKDYFPEPVTVSWNSGALTSVHTFPA VLQSSGLYSLSSVVTVPSSSLGTQTYICNVNHKPSTNTKVDKKVEPKSCDKTHT |
| Anti-kappa VHH |
MKYLLPTAAAGLLLLAAQPAMAHHHHHHQVQLQESGGGLVQPGGSLRLSCAASGRTISRYA MSWFRQAPGKEREFVAVARRSGDGAFYADSVQGRFTVSRDDAKNTVYLQMNSLKPEDTAYYCA IDSDTFYSGSYDYGWGQGTQVTVSS |
| Data Collection & Refinement Statistics | Batch A | Batch B |
|---|---|---|
| Beamline | SSRF BL10U2 | DLS I03 |
| Wavelength (Å) | 0.9792 | 0.97625 |
| Resolution range (Å) | 158.88–2.53 (2.63–2.53) | 29.85-2.29 (2.35-2.29) |
| Space group | P2₁2₁2₁ | P2₁2₁2₁ |
| Unit cell parameters (Å; °) | a = 75.71, b = 102.47, c = 158.88; α = β = γ = 90 | a = 76.01, b = 101.10, c = 161.24; α = β = γ = 90 |
| Matthew coefficient (Å3/Da) | 3.63 | 3.65 |
| Molecules per asymmetric unit | 1 | 1 |
| Total reflections | 548805 (58681) | 739731 (40781) |
| Unique reflections | 42007 (4344) | 56196 (3969) |
| Multiplicity | 13.1 (13.5) | 13.2 (10.3) |
| Completeness (%) | 100 (100) | 98.6 (86.8) |
| Mean I/sigma(I) | 9.7 (2.3) | 33.8 (5.5) |
| Wilson B-factor | 55.49 | 50.15 |
| R-merge/R-meas/Rpim | 0.131 (1.066)/0.136 (1.109)/0.038 (0.301) | 0.044 (0.29)/0.045 (0.305)/0.012 (0.093) |
| CC1/2 | 0.996 (0.866) | 1 (0.966) |
| Reflections used in refinement | 41669 (2740) | 55871 (2341) |
| Reflections used for R-free | 2031 (128) | 2814 (131) |
| Rwork/Rfree | 0.2207/0.2705 | 0.1947 (0.2284) |
| Total no. of non-hydrogen atoms (protein) | 5978 | 6254 |
| No. of protein/solvent residues | 735/263 | 743/454 |
| RMSD bond lengths, bond angles (Å; °) | 0.009/1.15 | 0.007/0.95 |
| Ramachandran favored/ allowed/ outliers/ rotamer outliers (%) | 94.31/4.17/1.53/5.03 | 97.13/2.74/0.14/2.33 |
| Clashscore | 10.27 | 8.58 |
| Average B-factor/protein/ligands/solvent | 56.04/55.83/80.17/55.29 | 52.10/51.49/72.97/55.59 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




