Submitted:
23 January 2026
Posted:
26 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
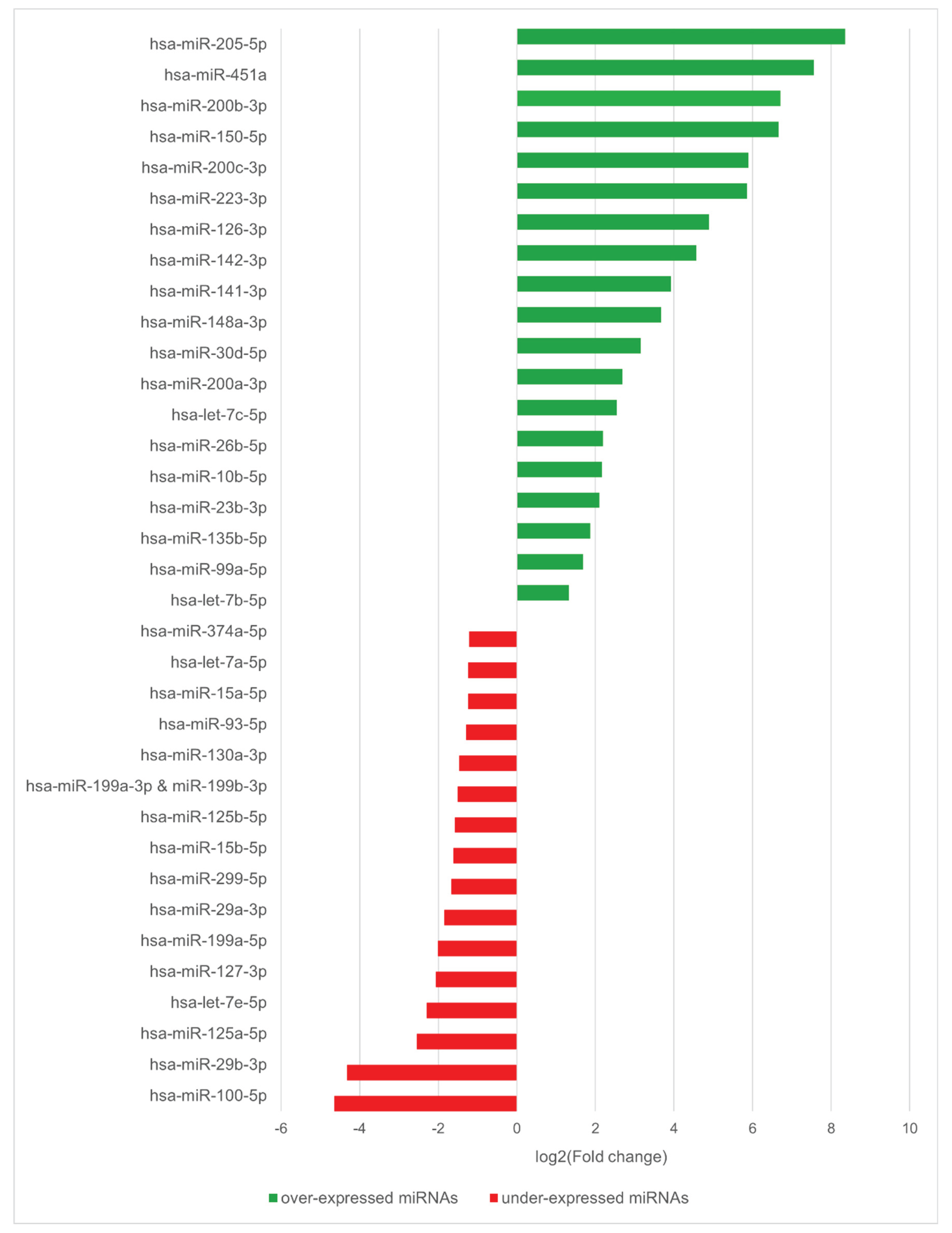
2.1. Identification of Deregulated miRNAs in CPO Patients
2.2. Validation of miRNA Dysregulation and Target Gene Expression in Primary CPO Cell Cultures
3. Discussion
4. Materials and Methods
4.1. Specimen Collection
4.2. Isolation and Primary Culture of Human Palatal Fibroblasts
4.3. RNA Isolation
4.4. miRNA Expression Profiling
4.5. Reverse Transcription
4.6. Real-Time PCR Amplifications
4.7. Transfection with miRNA Inhibitors and Mimic
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CL/P | Cleft Lip with or without cleft Palate |
| CLO | Cleft Lip Only |
| CPO | Cleft Palate Only |
| DE | Differential Expression |
| EMT | Epithelial to Mesenchymal Transition |
| FBS | Fetal Bovine Serum |
| HEPM | Human Embryonic Palate Mesenchymal |
| miRNA | microRNA |
| nsCL/P | non-syndromic Cleft Lip with or without cleft Palate |
| nsCPO | non-syndromic form of Cleft Palate Only |
| RT-qPCR | Reverse Transcription-quantitative Polymerase Chain Reaction |
| SNP | Single Nucleotide Polymorphisms |
References
- Tolarova, M.M.; Cervenka, J. Classification and birth prevalence of orofacial clefts. Am J Med Genet 1998, 75, 126–137. [Google Scholar] [CrossRef]
- Robinson, K.; Curtis, S.W.; Leslie, E.J. The heterogeneous genetic architectures of orofacial clefts. Trends Genet 2024, 40, 410–421. [Google Scholar] [CrossRef]
- Martinelli, M.; Palmieri, A.; Carinci, F.; Scapoli, L. Non-syndromic Cleft Palate: An Overview on Human Genetic and Environmental Risk Factors. Front Cell Dev Biol 2020, 8, 592271. [Google Scholar] [CrossRef] [PubMed]
- Leslie, E.J.; Liu, H.; Carlson, J.C.; Shaffer, J.R.; Feingold, E.; Wehby, G.; Laurie, C.A.; Jain, D.; Laurie, C.C.; Doheny, K.F.; et al. A Genome-wide Association Study of Nonsyndromic Cleft Palate Identifies an Etiologic Missense Variant in GRHL3. American journal of human genetics 2016, 98, 744–754. [Google Scholar] [CrossRef]
- Seelan, R.S.; Pisano, M.M.; Greene, R.M. MicroRNAs as epigenetic regulators of orofacial development. Differentiation 2022, 124, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microRNA target sites in mammalian mRNAs. Elife 2015, 4. [Google Scholar] [CrossRef]
- Shi, C.; Jiao, P.; Chen, Z.; Ma, L.; Yao, S. Exploring the roles of noncoding RNAs in craniofacial abnormalities: A systematic review. Dev Biol 2024, 505, 75–84. [Google Scholar] [CrossRef]
- Im, H.; Song, Y.; Kim, J.K.; Park, D.K.; Kim, D.S.; Kim, H.; Shin, J.O. Molecular Regulation of Palatogenesis and Clefting: An Integrative Analysis of Genetic, Epigenetic Networks, and Environmental Interactions. International journal of molecular sciences 2025, 26. [Google Scholar] [CrossRef] [PubMed]
- Burg, M.L.; Chai, Y.; Yao, C.A.; Magee, W., 3rd; Figueiredo, J.C. Epidemiology, Etiology, and Treatment of Isolated Cleft Palate. Front Physiol 2016, 7, 67. [Google Scholar] [CrossRef]
- Suzuki, A.; Li, A.; Gajera, M.; Abdallah, N.; Zhang, M.; Zhao, Z.; Iwata, J. MicroRNA-374a, -4680, and -133b suppress cell proliferation through the regulation of genes associated with human cleft palate in cultured human palate cells. BMC Med Genomics 2019, 12, 93. [Google Scholar] [CrossRef]
- Wang, S.; Sun, C.; Meng, Y.; Zhang, B.; Wang, X.; Su, Y.; Shi, L.; Zhao, E. A pilot study: Screening target miRNAs in tissue of nonsyndromic cleft lip with or without cleft palate. Exp Ther Med 2017, 13, 2570–2576. [Google Scholar] [CrossRef] [PubMed]
- Lan, Y.; Xu, J.; Jiang, R. Cellular and Molecular Mechanisms of Palatogenesis. Current topics in developmental biology 2015, 115, 59–84. [Google Scholar] [CrossRef] [PubMed]
- Jia, S.; Zhou, J.; Fanelli, C.; Wee, Y.; Bonds, J.; Schneider, P.; Mues, G.; D’Souza, R.N. Small-molecule Wnt agonists correct cleft palates in Pax9 mutant mice in utero. Development 2017, 144, 3819–3828. [Google Scholar] [CrossRef]
- Li, C.; Lan, Y.; Krumlauf, R.; Jiang, R. Modulating Wnt Signaling Rescues Palate Morphogenesis in Pax9 Mutant Mice. Journal of dental research 2017, 96, 1273–1281. [Google Scholar] [CrossRef]
- Almaidhan, A.; Cesario, J.; Landin Malt, A.; Zhao, Y.; Sharma, N.; Choi, V.; Jeong, J. Neural crest-specific deletion of Ldb1 leads to cleft secondary palate with impaired palatal shelf elevation. BMC Dev Biol 2014, 14, 3. [Google Scholar] [CrossRef]
- Won, H.J.; Won, H.S.; Shin, J.O. Increased miR-200c levels disrupt palatal fusion by affecting apoptosis, cell proliferation, and cell migration. Biochem Biophys Res Commun 2023, 664, 43–49. [Google Scholar] [CrossRef]
- Sandhu, N.; Rana, S.; Meena, K. Nuclear receptor subfamily 5 group A member 2 (NR5A2): role in health and diseases. Mol Biol Rep 2021, 48, 8155–8170. [Google Scholar] [CrossRef]
- Kimura, H.; Miyashita, H.; Suzuki, Y.; Kobayashi, M.; Watanabe, K.; Sonoda, H.; Ohta, H.; Fujiwara, T.; Shimosegawa, T.; Sato, Y. Distinctive localization and opposed roles of vasohibin-1 and vasohibin-2 in the regulation of angiogenesis. Blood 2009, 113, 4810–4818. [Google Scholar] [CrossRef]
- Beaty, T.H.; Ruczinski, I.; Murray, J.C.; Marazita, M.L.; Munger, R.G.; Hetmanski, J.B.; Murray, T.; Redett, R.J.; Fallin, M.D.; Liang, K.Y.; et al. Evidence for gene-environment interaction in a genome wide study of nonsyndromic cleft palate. Genet Epidemiol 2011, 35, 469–478. [Google Scholar] [CrossRef]
- Wang, Y.C.; Juan, H.C.; Wong, Y.H.; Kuo, W.C.; Lu, Y.L.; Lin, S.F.; Lu, C.J.; Fann, M.J. Protogenin prevents premature apoptosis of rostral cephalic neural crest cells by activating the alpha5beta1-integrin. Cell Death Dis 2013, 4, e651. [Google Scholar] [CrossRef] [PubMed]
- Leslie, E.J.; Carlson, J.C.; Shaffer, J.R.; Buxo, C.J.; Castilla, E.E.; Christensen, K.; Deleyiannis, F.W.B.; Field, L.L.; Hecht, J.T.; Moreno, L.; et al. Association studies of low-frequency coding variants in nonsyndromic cleft lip with or without cleft palate. Am J Med Genet A 2017, 173, 1531–1538. [Google Scholar] [CrossRef] [PubMed]
- Siewert, A.; Hoeland, S.; Mangold, E.; Ludwig, K.U. Combining genetic and single-cell expression data reveals cell types and novel candidate genes for orofacial clefting. Sci Rep 2024, 14, 26492. [Google Scholar] [CrossRef]
- Letra, A.; Menezes, R.; Fonseca, R.F.; Govil, M.; McHenry, T.; Murphy, M.J.; Hennebold, J.D.; Granjeiro, J.M.; Castilla, E.E.; Orioli, I.M.; et al. Novel cleft susceptibility genes in chromosome 6q. Journal of dental research 2010, 89, 927–932. [Google Scholar] [CrossRef] [PubMed]
- de Araujo, T.K.; Secolin, R.; Felix, T.M.; de Souza, L.T.; Fontes, M.I.; Monlleo, I.L.; de Souza, J.; Fett-Conte, A.C.; Ribeiro, E.M.; Xavier, A.C.; et al. A multicentric association study between 39 genes and nonsyndromic cleft lip and palate in a Brazilian population. J Craniomaxillofac Surg 2016, 44, 16–20. [Google Scholar] [CrossRef] [PubMed]
- Balcells, I.; Cirera, S.; Busk, P.K. Specific and sensitive quantitative RT-PCR of miRNAs with DNA primers. BMC Biotechnol 2011, 11, 70. [Google Scholar] [CrossRef]
- Busk, P.K. A tool for design of primers for microRNA-specific quantitative RT-qPCR. BMC Bioinformatics 2014, 15, 29. [Google Scholar] [CrossRef]
- Ye, J.; Coulouris, G.; Zaretskaya, I.; Cutcutache, I.; Rozen, S.; Madden, T.L. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinformatics 2012, 13, 134. [Google Scholar] [CrossRef]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
| miRNA | ΔΔCt | Student’s t | df | p value | Fold change | 95% CI | |
|---|---|---|---|---|---|---|---|
| mir205-5p | -5.5 | -6.04 | 9.35 | 0.0001* | 45.39 | 10.96 | 187.96 |
| mir200c-3p | -2.46 | -3.44 | 12 | 0.005* | 5.51 | 1.87 | 16.22 |
| mir125a-5p | 7.47 | 2.95 | 6.02 | 0.025* | 0.01 | 7.70E-05 | 0.41 |
| mir29b-3p | -0.76 | -1.1 | 10.93 | 0.293 | 1.69 | 0.59 | 4.84 |
| mir100-5p | -0.47 | -0.91 | 8.13 | 0.386 | 1.38 | 0.61 | 3.13 |
| Gene | Potential regulator | Score (miRDB) | ΔΔCt | Student's t | df | P value | Fold change | 95% CI | |
|---|---|---|---|---|---|---|---|---|---|
| AXIN2 | miR-205-5p | 89 | 0.96 | 1.64 | 11.8 | 0.126 | 0.51 | 0.21 | 1.24 |
| CCNJ | miR-205-5p | 97 | -0.07 | -0.11 | 8.43 | 0.911 | 1.05 | 0.41 | 2.66 |
| CDK19 | miR-205-5p | 99 | 1.19 | 2.05 | 6.11 | 0.085 | 0.44 | 0.17 | 1.17 |
| JADE1 | miR-205-5p | 89 | 0.23 | 0.49 | 8.51 | 0.633 | 0.86 | 0.42 | 1.76 |
| PAX9 | miR-205-5p | 91 | 3.65 | 3.03 | 11.65 | 0.011* | 0.08 | 0.01 | 0.5 |
| HIPK3 | miR-200c-3p | 100 | 0.36 | 2.28 | 11.53 | 0.042* | 0.78 | 0.62 | 0.99 |
| NR5A2 | miR-200c-3p | 100 | 2.58 | 3.26 | 6.35 | 0.016* | 0.17 | 0.04 | 0.63 |
| RECK | miR-200c-3p | 99 | 0.81 | 1.79 | 7.62 | 0.112 | 0.57 | 0.27 | 1.18 |
| TBK1 | miR-200c-3p | 93 | 2.68 | 4.28 | 7.72 | 0.003* | 0.16 | 0.06 | 0.43 |
| VASH2 | miR-200c-3p | 100 | 1.29 | 2.45 | 10.81 | 0.033* | 0.41 | 0.18 | 0.91 |
| ZEB1 | miR-200c-3p | 100 | 0.12 | 0.61 | 9.46 | 0.554 | 0.92 | 0.67 | 1.26 |
| DVL3 | miR-125a-5p | 64 | -2.97 | -7.82 | 9.15 | <0.001* | 7.82 | 4.32 | 14.16 |
| MAP6 | miR-125a-5p | 87 | -1.36 | -3.27 | 8.86 | 0.010* | 2.56 | 1.33 | 4.91 |
| PRSS35 | miR-125a-5p | 86 | 0.33 | 0.41 | 8.16 | 0.69 | 0.79 | 0.22 | 2.87 |
| PRTG | miR-125a-5p | 95 | -1.51 | -3.29 | 10.3 | 0.008* | 2.85 | 1.41 | 5.79 |
| TNFSF4 | miR-125a-5p | 90 | -1.11 | -1.14 | 8.27 | 0.288 | 2.16 | 0.46 | 10.19 |
| Inhibited miRNA | Gene | ΔΔCt | Student’s t | df | p value | Fold change | 95% CI | |
|---|---|---|---|---|---|---|---|---|
| miR-205-5p | AXIN2 | -0.20 | -1.11 | 1 | 0.467 | 1.15 | 0.23 | 5.88 |
| CCNJ | 0.19 | 0.36 | 1 | 0.779 | 0.87 | 0.01 | 96.44 | |
| CDK19 | -0.001 | -0.004 | 1 | 0.997 | 1.00 | 0.29 | 3.42 | |
| JADE1 | -0.42 | -0.75 | 1 | 0.592 | 1.34 | 0.01 | 182.88 | |
| PAX9 | 0.40 | 1.91 | 1 | 0.307 | 0.76 | 0.12 | 4.77 | |
| miR-200c-3p | HIPK3 | 0.44 | 1.76 | 6 | 0.129 | 0.74 | 0.48 | 1.13 |
| NR5A2 | 0.50 | 1.36 | 6 | 0.222 | 0.71 | 0.38 | 1.32 | |
| TBK1 | 0.76 | 2.84 | 6 | 0.030* | 0.59 | 0.38 | 0.93 | |
| VASH2 | -0.21 | -1.13 | 6 | 0.301 | 1.16 | 0.85 | 1.58 | |
| Gene | ΔΔCt | Student’s t | df | p value | Fold change | 95% CI | |
|---|---|---|---|---|---|---|---|
| DLV3 | 0.34 | 1.04 | 4 | 0.356 | 0.79 | 0.42 | 1.48 |
| MAP6 | -0.09 | -0.42 | 6 | 0.689 | 1.06 | 0.74 | 1.53 |
| PRSS35 | 0.56 | 2.61 | 6 | 0.040* | 0.68 | 0.47 | 0.98 |
| PRTG | 0.59 | 4.88 | 6 | 0.003* | 0.67 | 0.54 | 0.82 |
| TNFSF4 | 0.69 | 2.12 | 6 | 0.078 | 0.62 | 0.35 | 1.08 |
| Patient | Gender | Age (months) | Case classification | Nanostring® | Validation |
|---|---|---|---|---|---|
| 1 | Female | 12 | Sporadic | X | |
| 2 | Male | 12 | Sporadic | X | |
| 3 | Male | 11 | Sporadic | X | |
| 4 | Female | 11 | Sporadic | X | |
| 5 | Female | 7 | Sporadic | X | X |
| 6 | Male | 10 | Sporadic | X | X |
| 7 | Male | 5 | Family history of CPO | X | X |
| 8 | Male | 4 | Sporadic | X | X |
| 9 | Male | 6 | Sporadic | X | X |
| 10 | Male | 4 | Sporadic | X | X |
| 11 | Male | 6 | Sporadic | X | X |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




