Submitted:
23 January 2026
Posted:
26 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Chemicals and Reagents
2.2. Cells and Culture Media
2.3. hWJ-MSCs Characterization
2.4. Conditioned Media Preparation
2.5. Conversion of rhESCs to the Naïve State
2.6. Immunofluorescence and Imaging
2.7. Real-Time Quantitative PCR (qPCR)
2.8. Statistical Analysis
3. Results
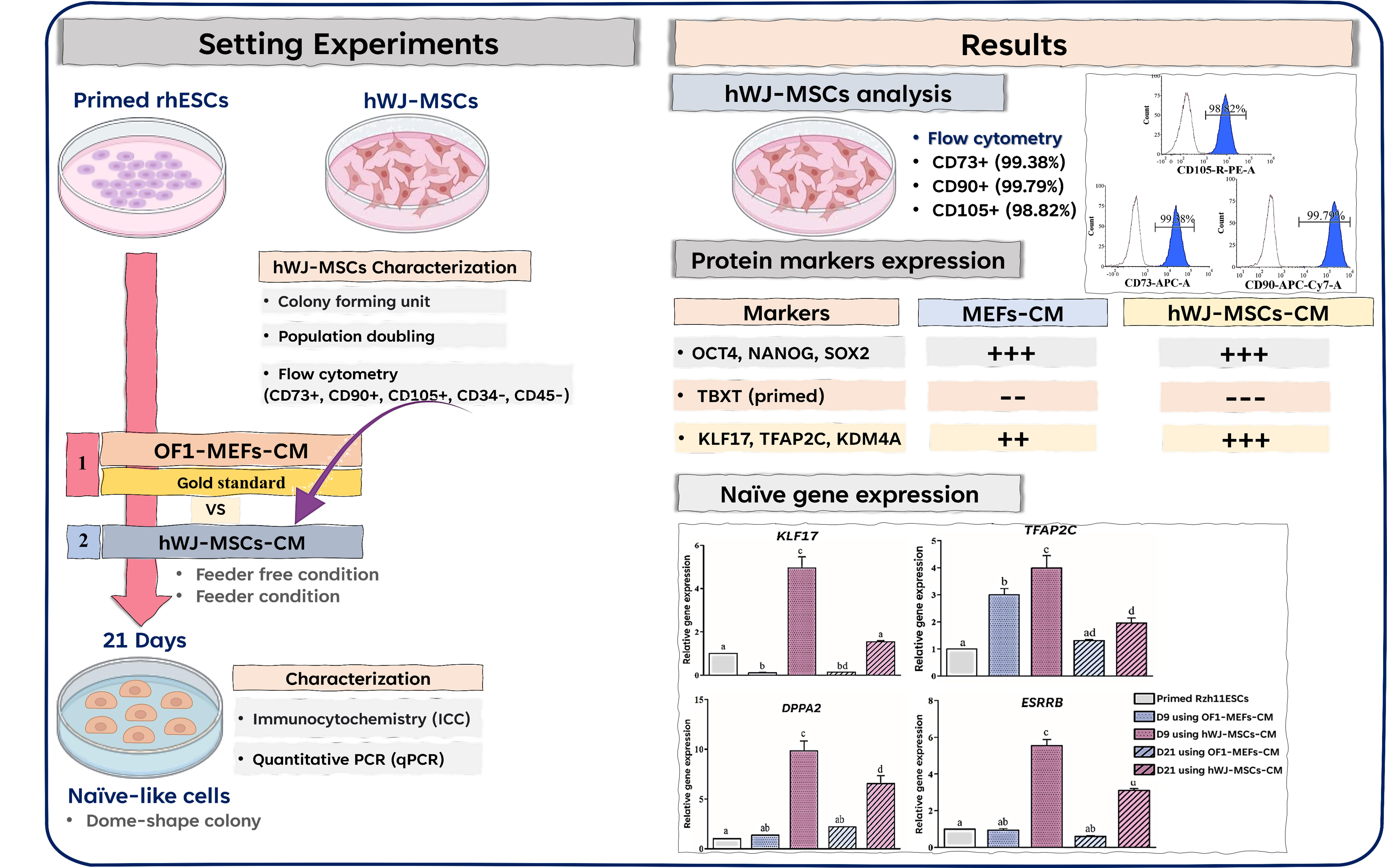
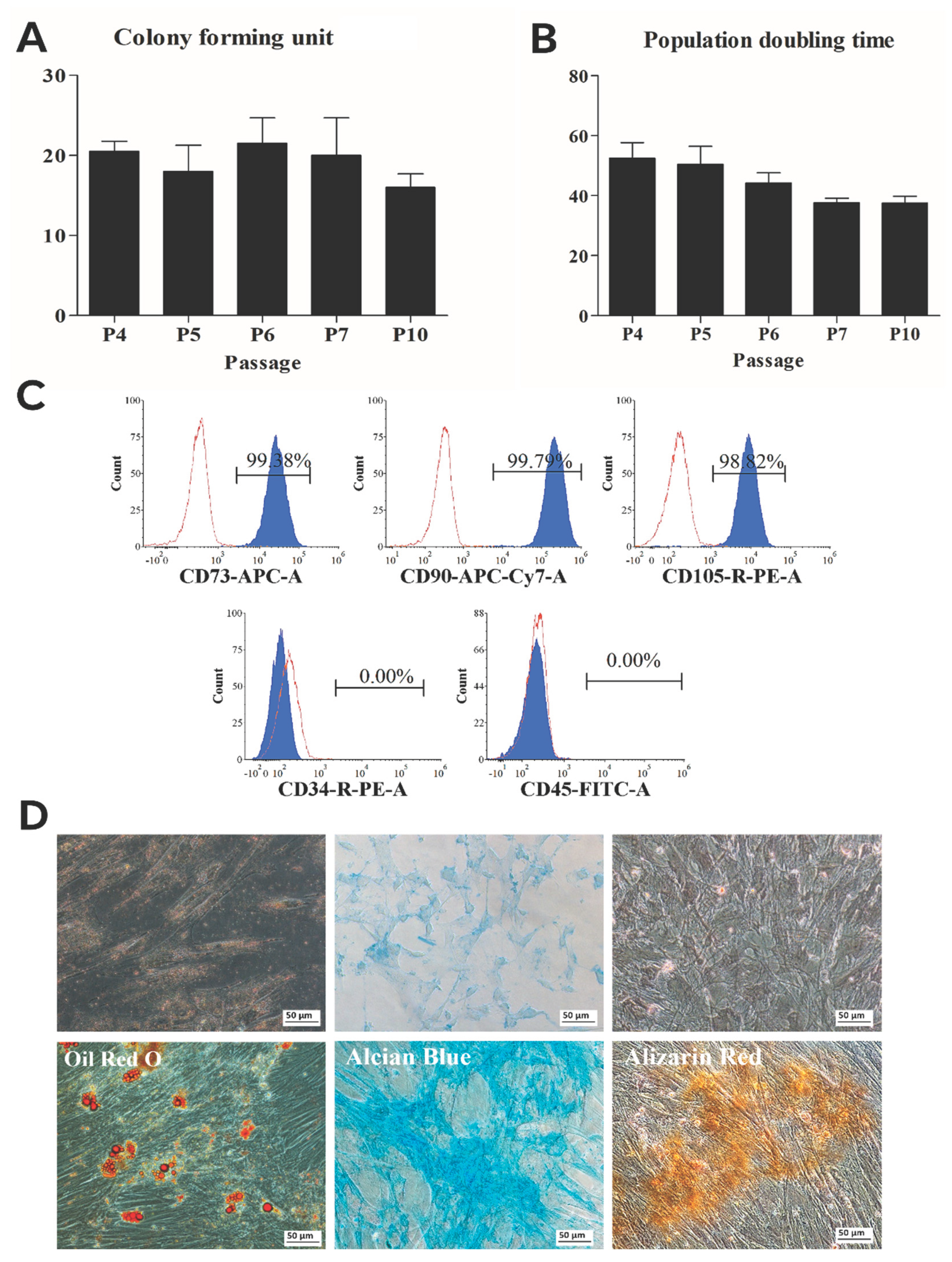
3.1. hWJ-MSC Characterization
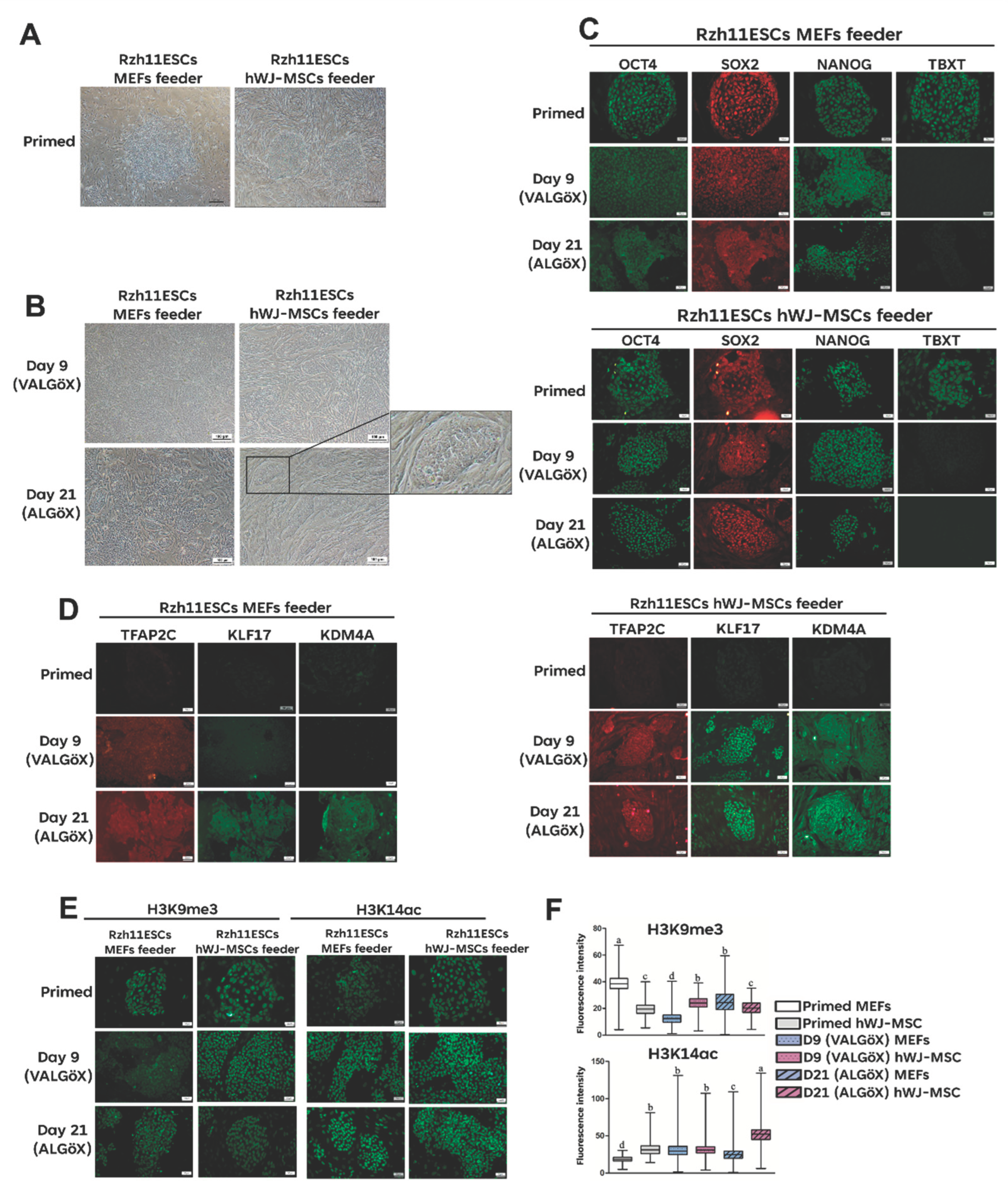
3.2. Propagation of Rzh11ESCs Using MEFs and hWJ-MSC-Derived Feeder Cells
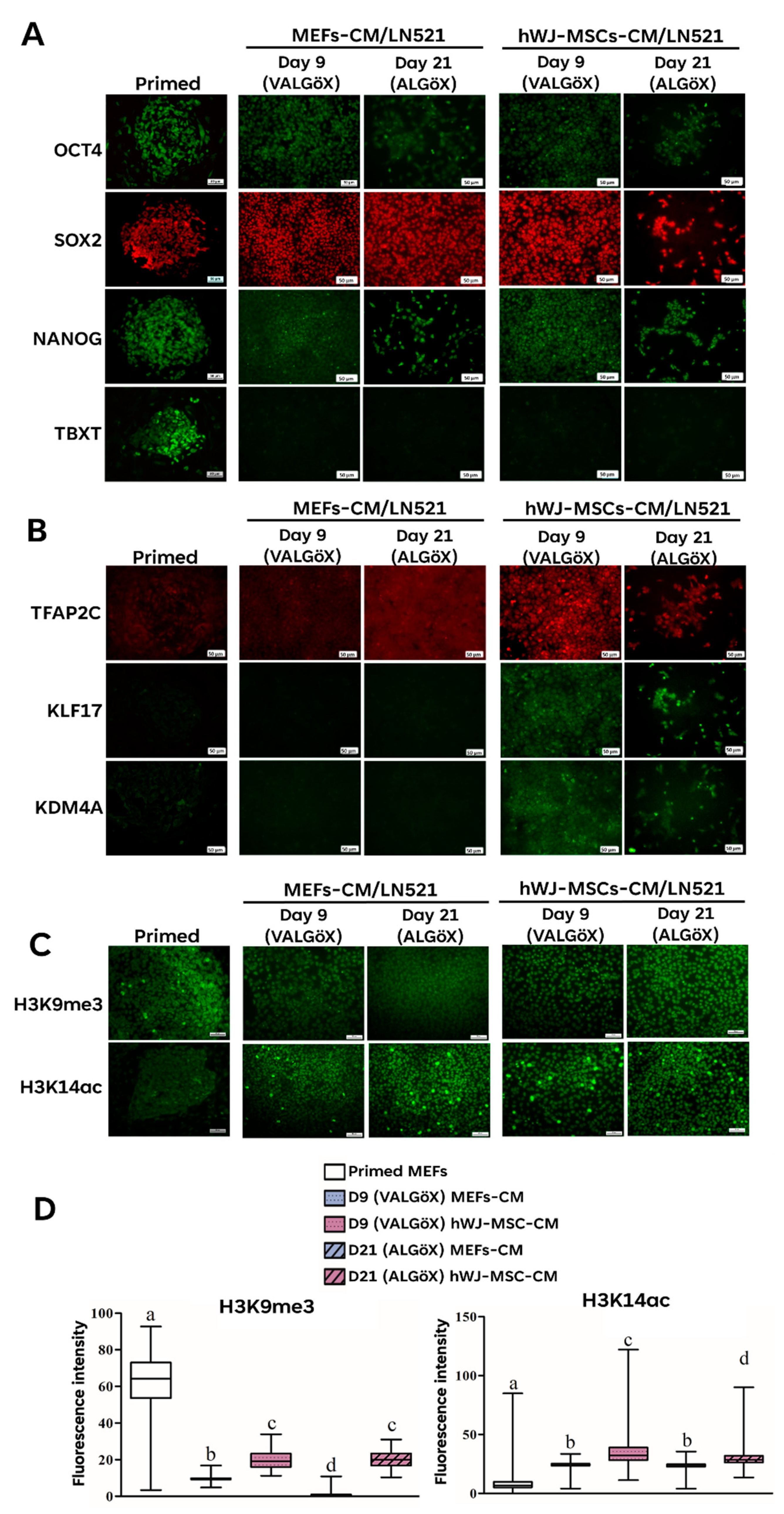
3.3. Propagation of Rzh11ESCs Using hWJ-MSC-Derived Conditioned Medium
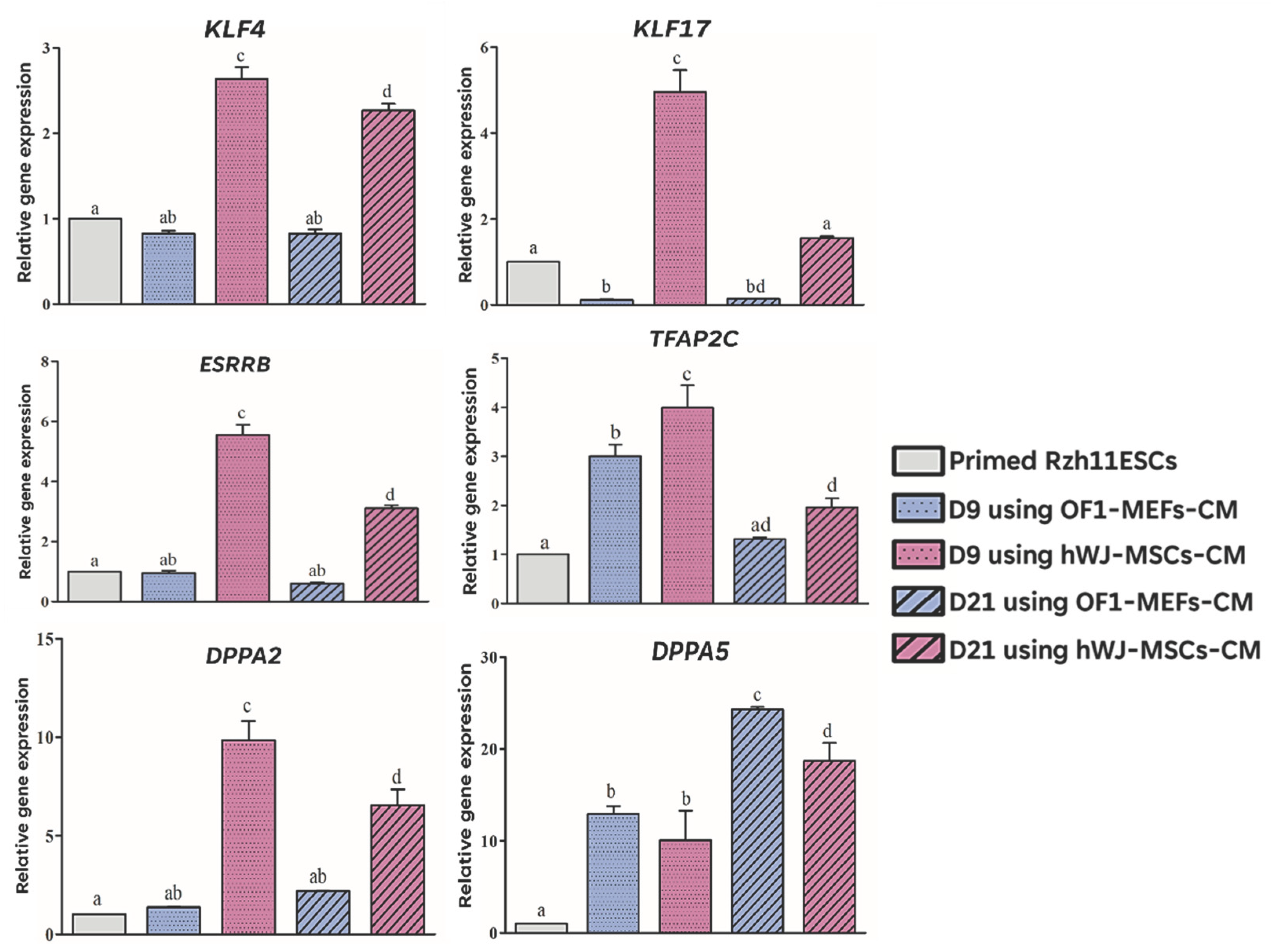
3.4. hWJ-MSCs-Derived Conditioned Medium Enhances Primed-to-Naïve State Conversion
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| bFGF | Basic fibroblast growth factor |
| CjESCs | Marmoset ESCs |
| CyESCs | Cynomolgus monkey |
| CFU | Colony-Forming Unit |
| ESCs | Embryonic stem cells |
| EpiSCs | Epiblast stem cells |
| FBS | Fetal bovine serum |
| NHPs | Non-human primates |
| HDACs | Histone deacetylases |
| hWJ-MSCs | Human Wharton’s Jelly-derived mesenchymal stem cells |
| hWJ-MSCs-CM | hWJ-MSC-conditioned media |
| KO-DMEM | Knockout Dulbecco’s modified Eagle’s medium |
| KOSR | Knockout serum replacement |
| MEFs | Mouse embryonic fibroblasts |
| MEFs-CM | MEFs-conditioned media |
| PSCs | Pluripotent stem cells |
| rhESCs | Rhesus monkey embryonic stem cells |
| RT | Room temperature |
| PFA | Paraformaldehyde |
| SAHH | S-adenosyl homocysteine hydrolase |
References
- Nichols, J.; Smith, A. Naïve and primed pluripotent states. Cell Stem Cell. 2009, 4, 487–492. [Google Scholar] [CrossRef] [PubMed]
- Weinberger, L.; Ayyash, M.; Novershtern, N.; Hanna, J.H. Dynamic stem cell states: Naïve to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell Biol. 2016, 17, 155–169. [Google Scholar] [CrossRef]
- Guo, G.; von Meyenn, F.; Santos, F.; Chen, Y.; Reik, W.; Bertone, P.; Smith, A.; Nichols, J. Naive pluripotent stem cells derived directly from isolated cells of the human inner cell mass. Stem Cell Reports 2016, 6, 437–446. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhao, C.; Hou, Z.; Yang, Y.; Bim, Y.; Wang, H.; Zhang, Y.; Gao, S. Unique molecular events during reprogramming of human somatic cells to induced pluripotent stem cells (iPSCs) at naïve state. Elife 2018, 30, 7:e29518. [Google Scholar] [CrossRef]
- Giulitti, S.; Pellegrini, M.; Zorzan, I.; Martini, P.; Gagliano, O.; Mutarelli, M.; Ziller, M.J.; Cacchiarelli, D.; Romualdi, C.; Elvassore, N.; et al. Direct generation of human naive induced pluripotent stem cells from somatic cells in microfluidics. Nat. Cell Biol. 2019, 21, 275–286. [Google Scholar] [CrossRef]
- Buckberry, S.; Liu, X.; Poppe, D.; Tan, P.J.; Sun, G.; Chen, J.; Nguyen, T.V.; de Mendoza, A.; Pflueger, J.; Frazer, T.; et al. Transient naive reprogramming corrects hiPS cells functionally and epigenetically. Nature 2023, 620, 863–872. [Google Scholar] [CrossRef] [PubMed]
- Fang, R.; Liu, K.; Zhao, Y.; Li, H.; Zhu, D.; Du, Y.; Xiang, C.; Li, X.; Liu, H.; Miao, Z.; et al. Generation of naive induced pluripotent stem cells from rhesus monkey fibroblasts. Cell Stem Cell. 2014, 15, 488–497. [Google Scholar] [CrossRef]
- Chen, Y.; Niu, Y.; Li, Y.; Ai, Z.; Kang, Y.; Shi, H.; Xiang, Z.; Yang, Z.; Tan, T.; Si, W.; et al. Generation of Cynomolgus Monkey Chimeric Fetuses using Embryonic Stem Cells. Cell Stem Cell. 2015, 17, 116–124. [Google Scholar] [CrossRef]
- Honda, H.; Kawano, Y.; Izu, H.; Choijookhuu, N.; Honsho, K.; Nakamura, T.; Yabuta, Y.; Yamamoto, T.; Takashima, Y.; Hirose, M.; et al. Discrimination of Stem Cell Status after Subjecting Cynomolgus Monkey Pluripotent Stem Cells to Naïve Conversion. Sci Rep. 2017, 7, 45285. [Google Scholar] [CrossRef]
- Bergmann, S.; Penfold, C.A.; Slatery, E.; Siriwardena, D.; Drummer, C.; Clark, S.; Strawbridge, S.E.; Kishimoto, K.; Vickers, A.; Tewary, M.; et al. Spatial profiling of early primate gastrulation in utero. Nature 2022, 609, 136–143. [Google Scholar] [CrossRef]
- Li, J.; Zhu, Q.; Cao, J.; Liu, Y.; Lu, Y.; Sun, Y.; Li, Q.; Huang, Y.; Shang, S.; Bian, X.; et al. Cynomolgus monkey embryo model captures gastrulation and early pregnancy. Cell Stem Cell 2023, 30, 362–377. [Google Scholar] [CrossRef]
- Pham, H.T.; Perold, F.; Pijoff, Y.; Doerflinger, N.; Rival-Gervier, S.; Givelet, M.; Moulin, A.; Ressaire, M.; Da Silva Fernandes, E.; Bidault, V.; et al. Efficient generation of germline chimeras in a non-rodent species using rabbit induced pluripotent stem cells. Nat. Commun. 2025, 16, 5165. [Google Scholar] [CrossRef]
- Kuo, S.C.; Chio, C.C.; Yeh, C.H.; Ma, J.T.; Liu, W.P.; Lin, M.T.; Lin, K.C.; Chang, C.P. Mesenchymal stem cell-conditioned medium attenuates the retinal pathology in amyloid-β-induced rat model of Alzheimer's disease: Underlying mechanisms. Aging Cell. 2021, 20, e13340. [Google Scholar] [CrossRef]
- da Silva Meirelles, L.; Chagastelles, P.C.; Nardi, N.B. Mesenchymal stem cells reside in virtually all post-natal organs and tissues. J. Cell Sci. 2006, 119, 2204–2213. [Google Scholar] [CrossRef]
- Chamberlain, G.; Fox, J.; Ashton, B.; Middleton, J. Concise review: mesenchymal stem cells: their phenotype, differentiation capacity, immunological features, and potential for homing. Stem Cells 2007, 25, 2739–2749. [Google Scholar] [CrossRef]
- Abbasi-Malati, Z.; Roushandeh, A.M.; Kuwahara, Y.; Roudkenar, M.H. Mesenchymal Stem Cells on Horizon: A New Arsenal of Therapeutic Agents. Stem Cell Rev Rep. 2018, 14, 484–499. [Google Scholar] [CrossRef] [PubMed]
- Acuto, S.; Lo Iacono, M.; Baiamonte, E.; Lo Re, R.; Maggio, A.; Cavalieri, V. An optimized procedure for preparation of conditioned medium from Wharton's jelly mesenchymal stromal cells isolated from umbilical cord. Front Mol Biosci. 2023, 2, 10:1273814. [Google Scholar] [CrossRef] [PubMed]
- Putkhao, K.; Kocerha, J.; Cho, I.K.; Yang, J.; Parnpai, R.; Chan, A.W. Pathogenic cellular phenotypes are germline transmissible in a transgenic primate model of Huntington's disease. Stem Cells Dev. 2013, 22, 1198–1205. [Google Scholar] [CrossRef]
- Afanassieff, M.; Tapponnier, Y.; Savatier, P. Generation of Induced Pluripotent Stem Cells in Rabbits. In duced Pluripotent Stem (iPS) Cells. Methods and Protocols, 2nd ed.; Turksen, K., Nagy, A., Eds.; Humana Press: New York, NY, USA, 2014; Volume 1357, pp. 149–172. [Google Scholar] [CrossRef]
- Somredngan, S.; Theerakittayakorn, K.; Nguyen, H.T.; Ngernsoungnern, A.; Ngernsoungnern, P.; Sritangos, P.; Ketudat-Cairns, M.; Imsoonthornruksa, S.; Keeratibharat, N.; Wongsan, R.; Rungsiwiwut, R.; Parnpai, R. The Efficiency of Neurospheres Derived from Human Wharton's Jelly Mesenchymal Stem Cells for Spinal Cord Injury Regeneration in Rats. Int J Mol Sci. 2023, 24, 3846. [Google Scholar] [CrossRef]
- Bharti, D.; Shivakumar, S.B.; Park, J.K.; Ullah, I.; Subbarao, R.B.; Park, J.S.; Lee, S.L.; Park, B.W.; Rho, G.J. Comparative analysis of human Wharton's jelly mesenchymal stem cells derived from different parts of the same umbilical cord. Cell Tissue Res. 2018, 372, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.S.; Hung, S.C.; Peng, S.T.; Huang, C.C.; Wei, H.M.; Guo, Y.J.; Fu, Y.S.; Lai, M.C.; Chen, C.C. Mesenchymal stem cells in the Wharton's jelly of the human umbilical cord. Stem Cells 2004, 22, 1330–1337. [Google Scholar] [CrossRef] [PubMed]
- Dominici, M.; Le Blanc, K.; Mueller, I.; Slaper-Cortenbach, I.; Marini, F.; Krause, D.; Deans, R.; Keating, A.; Prockop, Dj.; Horwitz, E. Minimal criteria for defining multipotent mesenchymal stromal cells. The International Society for Cellular Therapy position statement. Cytotherapy 2006, 8, 315–7. [Google Scholar] [CrossRef]
- Fong, C.Y.; Subramanian, A.; Gauthaman, K.; Venugopal, J.; Biswas, A.; Ramakrishna, S.; Bongso, A. Human umbilical cord Wharton's jelly stem cells undergo enhanced chondrogenic differentiation when grown on nanofibrous scaffolds and in a sequential two-stage culture medium environment. Stem Cell Rev Rep. 2012, 8, 195–209. [Google Scholar] [CrossRef] [PubMed]
- Bates, L.E.; Silva, J.C. Reprogramming human cells to naïve pluripotency: how close are we? Curr Opin Genet Dev. 2017, 46, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Hu, J.; Wang, Y.; Gao, S. Induction and application of human naive pluripotency. Cell Rep. 2023, 42, 112379. [Google Scholar] [CrossRef]
- Hanna, J.; Cheng, A.W.; Saha, K.; Kim, J.; Lengner, C.J.; Soldner, F.; Cassady, J.P.; Muffat, J.; Carey, B.W.; Jaenisch, R. Human embryonic stem cells with biological and epigenetic characteristics similar to those of mouse ESCs. Proc Natl Acad Sci U S A 2010, 107, 9222–9227. [Google Scholar] [CrossRef]
- Batsivari, A.; Haltalli, M.L.R.; Passaro, D.; Pospori, C.; Celso, C.L.; Bonnet, D. Dynamic responses of the haematopoietic stem cell niche to diverse stresses. Nat Cell Biol. 2020, 22, 7–17. [Google Scholar] [CrossRef]
- Theunissen, T.W.; Powell, B.E.; Wang, H.; Mitalipova, M.; Faddah, D.A.; Reddy, J.; Fan, Z.P.; Maetzel, D.; Ganz, K.; Shi, L.; et al. Systematic identification of culture conditions for induction and maintenance of naive human pluripotency. Cell Stem Cell. 2014, 15, 471–487. [Google Scholar] [CrossRef]
- Buecker, C.; Srinivasan, R.; Wu, Z.; Calo, E.; Acampora, D.; Faial, T.; Simeone, A.; Tan, M.; Swigut, T.; Wysocka, J. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell. 2014, 14, 838–853. [Google Scholar] [CrossRef]
| Gene type | Gene | Primer Sequence (5´-3´) | Annealing temperature (°C) |
|---|---|---|---|
| Housekeeping gene | GADPH | F: GGAGCGAGATCCCTCCAAAAT R: GGCTGTTGTCATACTTCTCATGG |
54 |
| Pluripotency genes | OCT4 | F: AGTGTGGTTCTGTAACCGGC R: GACCGAGGAGTACAGTGCAG |
58 |
| NANOG | F: AGTCCTGCTTGCAGTTCCAG R: TCAGGTTGCATGTTCGTGGA |
55 | |
| SOX2 | F: AACCAGCGCATGGACAGTTA R: CGAGCTGGTCATGGAGTTGT |
56 | |
| Primed specific genes | TBXT | F: CTTCAGCAAAGTCAAGCTCACC R: TGAACTGGGTCTCAGGGAAGCA |
56 |
| OTX2 | F: AAAGTGAGACCTGCCAAAAAGA R: TGGACAAGGGATCTGACAGTG |
56 | |
| Naïve specific genes | KLF4 | F: CCCTACCTCGGAGAGAGACC R: GGATGGGTCAGCGAATTGGA |
58 |
| KLF17 | F: CCTTACCGCTGCAACTACGA R: ATAGGGCCTCTCACCTGTGT |
57 | |
| TFAP2C | F: GTTCTCAGAAGAGCCAAGTCG R: TCGGCTTCACAGACATAGGC |
56 | |
| ESRRB | F: TGCCCTATGACGACAAGCTG R: TGAGCGTCACAAACTCCTCC |
58 | |
| DPPA2 | F: GTACGCCTGCAGTTTCATGC R: TCTATGCCTGGGGATGGGAA |
55 | |
| DPPA5 | F: GGTCGTGGTTTACGGTTCCT R: AGTTTGAGCATCCCTCGCTC |
56 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




