Submitted:
22 January 2026
Posted:
23 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
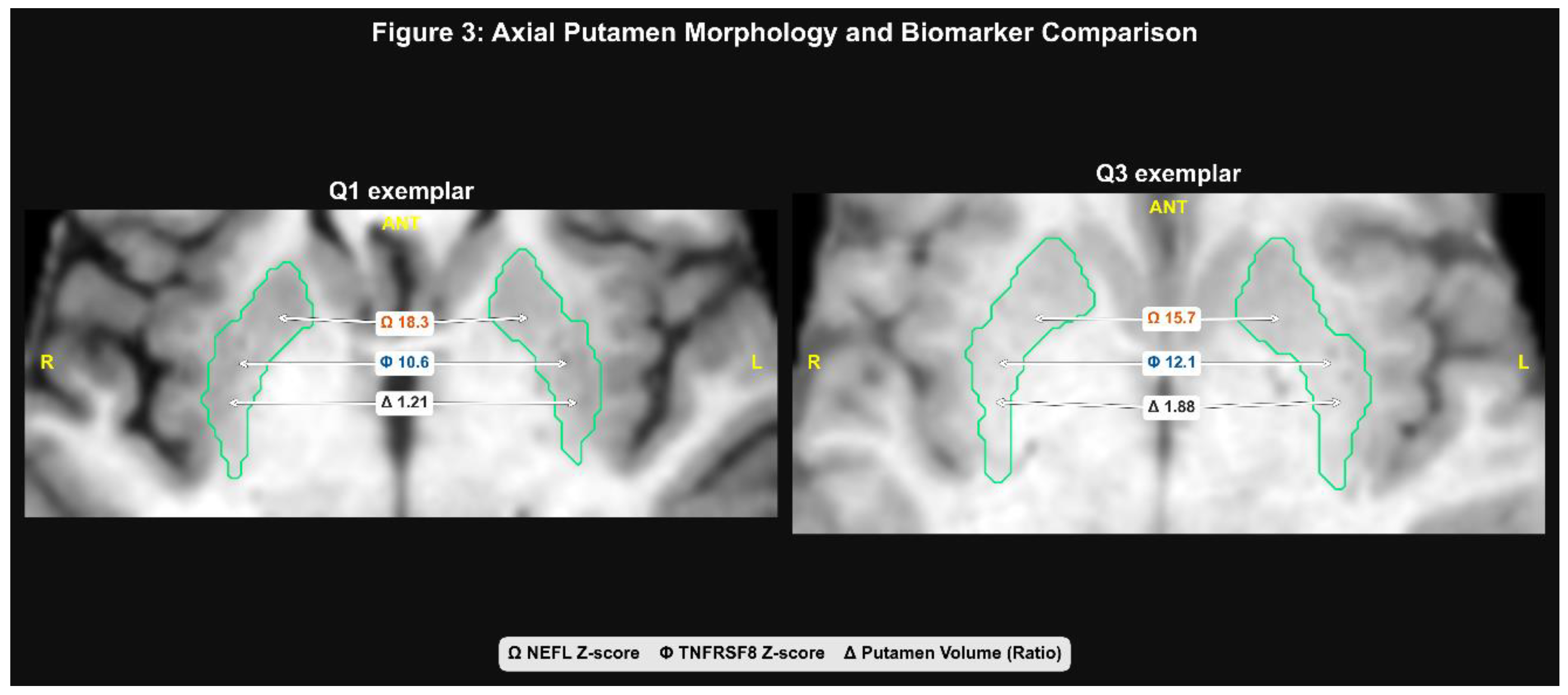
2.1. Validation of Striatal Atrophy
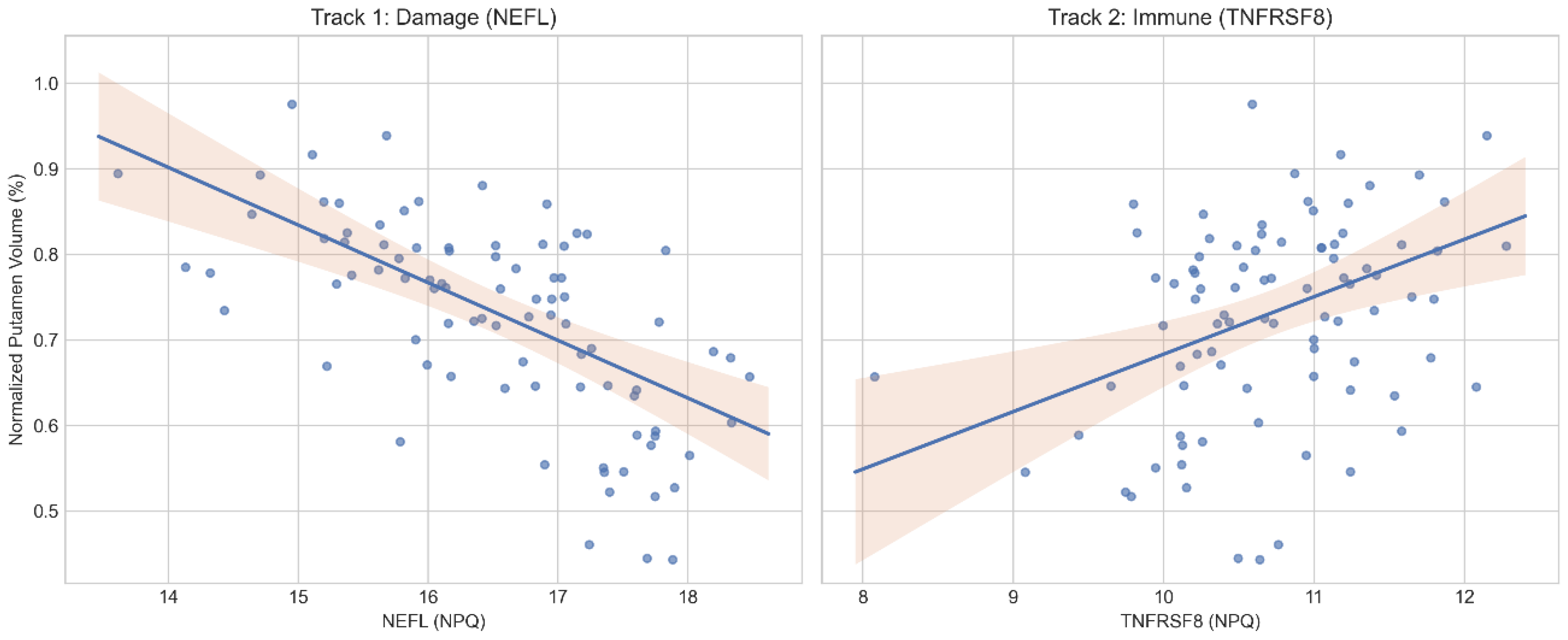
2.2. Divergent Proteomic Signatures
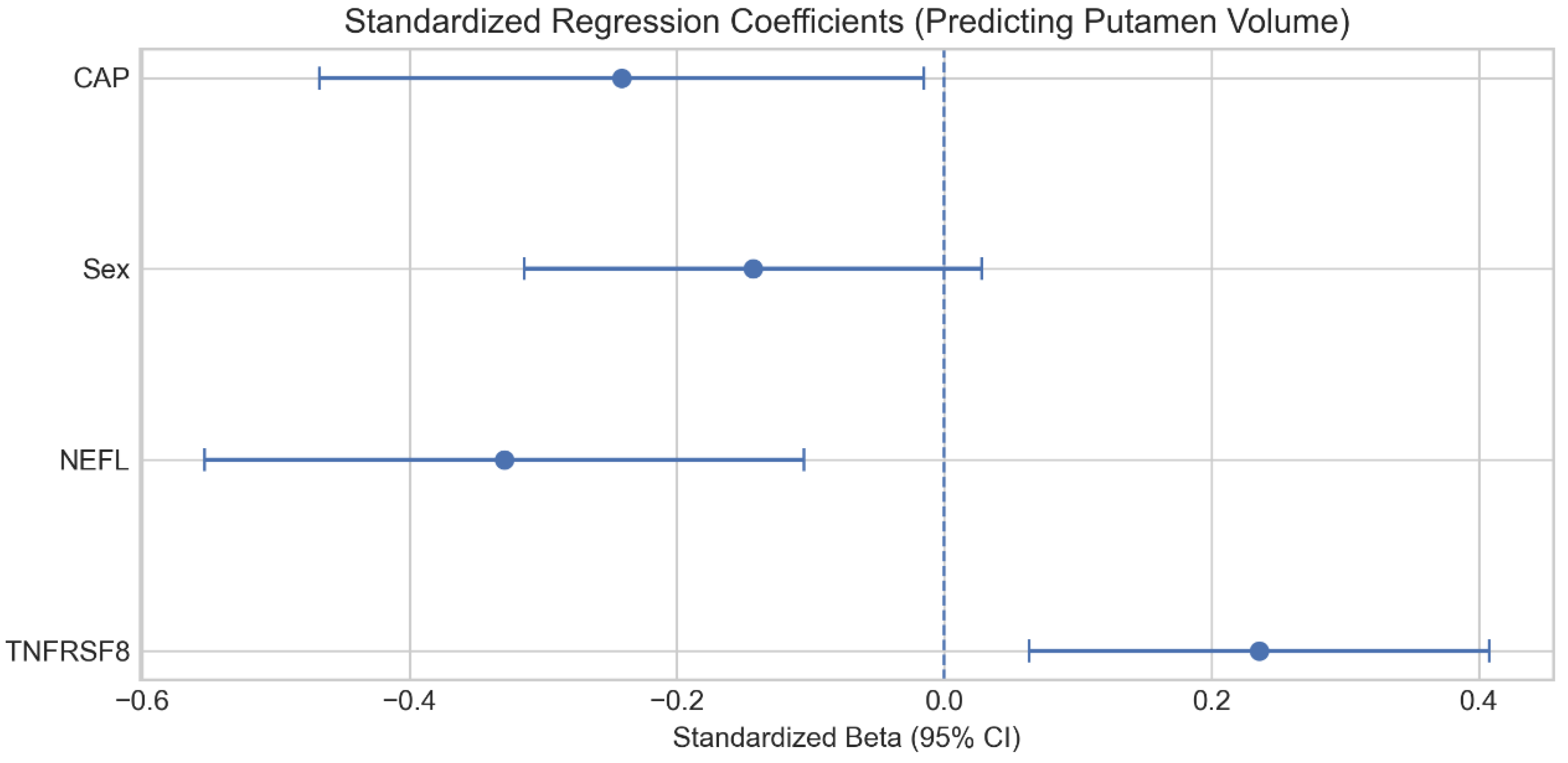
2.3. Independent Contributions to Atrophy
2.4. Mediation Analysis
3. Discussion
4. Materials and Methods
4.1. Participants
| Characteristic | Cohort (N=88) |
|---|---|
| Age, years (SD) | 39.01 (11.83) |
| Education, years (SD) | 15.29 (2.16) |
| CAP1 score (SD) | 336.48 (92.25) |
| CAG2 Repeats (SD) | 42.74 (2.93) |
| Sex, n (%) | |
| Female | 58 (65.9%) |
| Male | 30 (34.1%) |
4.2. Neuroimaging
4.3. CSF Collection and Handling
4.4. Proteomics
4.5. Proteomics
4.6. Generative Artificial Intelligence
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ADRC | Alzheimer’s Disease Research Center |
| BioSEND | BioSpecimen Exchange for Neurological Disorders |
| CAG | Cytosine-Adenine-Guanine |
| CAP | CAG-Age-Product |
| CNS | Central nervous system |
| CSF | Cerebrospinal fluid |
| FDR | False discovery rate |
| HD | Huntington’s Disease |
| HTT | Huntingtin gene |
| ICV | Intracranial volume |
| LOD | Limit of detection |
| MRI | Magnetic resonance imaging |
| NEFL | Neurofilament light |
| NPQ | NULISA Protein Quantification |
| NULISA | Next-Gen Ultra-Sensitive Immunoassay |
| OLS | Ordinary least squares |
| TNF | Tumor necrosis factor |
| VIF | Variance inflation factor |
References
- Tong, H.; Yang, T.; Xu, S.; Li, X.; Liu, L.; Zhou, G.; Yang, S.; Yin, S.; Li, X.J.; Li, S. Huntington's Disease: Complex Pathogenesis and Therapeutic Strategies. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef]
- Ehrlich, M.E. Huntington's disease and the striatal medium spiny neuron: cell-autonomous and non-cell-autonomous mechanisms of disease. Neurotherapeutics 2012, 9, 270–284. [Google Scholar] [CrossRef] [PubMed]
- Biglan, K.M.; Ross, C.A.; Langbehn, D.R.; Aylward, E.H.; Stout, J.C.; Queller, S.; Carlozzi, N.E.; Duff, K.; Beglinger, L.J.; Paulsen, J.S.; et al. Motor abnormalities in premanifest persons with Huntington's disease: the PREDICT-HD study. Mov Disord 2009, 24, 1763–1772. [Google Scholar] [CrossRef]
- Li, K.; Furr-Stimming, E.; Paulsen, J.S.; Luo, S.; Group, P.-H.I.o.t.H.S. Dynamic Prediction of Motor Diagnosis in Huntington's Disease Using a Joint Modeling Approach. J Huntingtons Dis 2017, 6, 127–137. [Google Scholar] [CrossRef] [PubMed]
- Paulsen, J.S.; Langbehn, D.R.; Stout, J.C.; Aylward, E.; Ross, C.A.; Nance, M.; Guttman, M.; Johnson, S.; MacDonald, M.; Beglinger, L.J.; et al. Detection of Huntington's disease decades before diagnosis: the Predict-HD study. J Neurol Neurosurg Psychiatry 2008, 79, 874–880. [Google Scholar] [CrossRef]
- Roze, E.; Cahill, E.; Martin, E.; Bonnet, C.; Vanhoutte, P.; Betuing, S.; Caboche, J. Huntington's Disease and Striatal Signaling. Front Neuroanat 2011, 5, 55. [Google Scholar] [CrossRef]
- Fang, Q.; Strand, A.; Law, W.; Faca, V.M.; Fitzgibbon, M.P.; Hamel, N.; Houle, B.; Liu, X.; May, D.H.; Poschmann, G.; et al. Brain-specific proteins decline in the cerebrospinal fluid of humans with Huntington disease. Mol Cell Proteomics 2009, 8, 451–466. [Google Scholar] [CrossRef]
- Byrne, L.M.; Rodrigues, F.B.; Blennow, K.; Durr, A.; Leavitt, B.R.; Roos, R.A.C.; Scahill, R.I.; Tabrizi, S.J.; Zetterberg, H.; Langbehn, D.; et al. Neurofilament light protein in blood as a potential biomarker of neurodegeneration in Huntington's disease: a retrospective cohort analysis. The Lancet Neurology 2017, 16, 601–609. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, F.B.; Byrne, L.M.; McColgan, P.; Robertson, N.; Tabrizi, S.J.; Zetterberg, H.; Wild, E.J. Cerebrospinal Fluid Inflammatory Biomarkers Reflect Clinical Severity in Huntington's Disease. PLoS One 2016, 11, e0163479. [Google Scholar] [CrossRef]
- Crotti, A.; Glass, C.K. The choreography of neuroinflammation in Huntington's disease. Trends Immunol 2015, 36, 364–373. [Google Scholar] [CrossRef]
- Feng, W.; Beer, J.C.; Hao, Q.; Ariyapala, I.S.; Sahajan, A.; Komarov, A.; Cha, K.; Moua, M.; Qiu, X.; Xu, X.; et al. NULISA: a proteomic liquid biopsy platform with attomolar sensitivity and high multiplexing. Nat Commun 2023, 14, 7238. [Google Scholar] [CrossRef] [PubMed]
- Aylward, E.H. Magnetic resonance imaging striatal volumes: a biomarker for clinical trials in Huntington's disease. Mov Disord 2014, 29, 1429–1433. [Google Scholar] [CrossRef] [PubMed]
- Ghazaleh, N.; Houghton, R.; Palermo, G.; Schobel, S.A.; Wijeratne, P.A.; Long, J.D. Ranking the Predictive Power of Clinical and Biological Features Associated With Disease Progression in Huntington's Disease. Front Neurol 2021, 12, 678484. [Google Scholar] [CrossRef] [PubMed]
- Paulsen, J.S.; Long, J.D.; Johnson, H.J.; Aylward, E.H.; Ross, C.A.; Williams, J.K.; Nance, M.A.; Erwin, C.J.; Westervelt, H.J.; Harrington, D.L.; et al. Clinical and Biomarker Changes in Premanifest Huntington Disease Show Trial Feasibility: A Decade of the PREDICT-HD Study. Front Aging Neurosci 2014, 6, 78. [Google Scholar] [CrossRef]
- Rodrigues, F.B.; Byrne, L.M.; Tortelli, R.; Johnson, E.B.; Wijeratne, P.A.; Arridge, M.; De Vita, E.; Ghazaleh, N.; Houghton, R.; Furby, H.; et al. Mutant huntingtin and neurofilament light have distinct longitudinal dynamics in Huntington's disease. Sci Transl Med 2020, 12. [Google Scholar] [CrossRef]
- Caron, N.S.; Haqqani, A.S.; Sandhu, A.; Aly, A.E.; Findlay Black, H.; Bone, J.N.; McBride, J.L.; Abulrob, A.; Stanimirovic, D.; Leavitt, B.R.; et al. Cerebrospinal fluid biomarkers for assessing Huntington disease onset and severity. Brain Commun 2022, 4, fcac309. [Google Scholar] [CrossRef]
- Muta, H.; Podack, E.R. CD30: from basic research to cancer therapy. Immunol Res 2013, 57, 151–158. [Google Scholar] [CrossRef]
- van der Weyden, C.A.; Pileri, S.A.; Feldman, A.L.; Whisstock, J.; Prince, H.M. Understanding CD30 biology and therapeutic targeting: a historical perspective providing insight into future directions. Blood Cancer J 2017, 7, e603. [Google Scholar] [CrossRef]
- Steiger, J.H. Testing Pattern Hypotheses On Correlation Matrices: Alternative Statistics And Some Empirical Results. Multivariate Behav Res 1980, 15, 335–352. [Google Scholar] [CrossRef]
- Efron, B. Estimation and Accuracy after Model Selection. J Am Stat Assoc 2014, 109, 991–1007. [Google Scholar] [CrossRef]
- Bjorkqvist, M.; Wild, E.J.; Thiele, J.; Silvestroni, A.; Andre, R.; Lahiri, N.; Raibon, E.; Lee, R.V.; Benn, C.L.; Soulet, D.; et al. A novel pathogenic pathway of immune activation detectable before clinical onset in Huntington's disease. J Exp Med 2008, 205, 1869–1877. [Google Scholar] [CrossRef]
- Ellrichmann, G.; Reick, C.; Saft, C.; Linker, R.A. The role of the immune system in Huntington's disease. Clin Dev Immunol 2013, 2013, 541259. [Google Scholar] [CrossRef]
- Li, X.; Tong, H.; Xu, S.; Zhou, G.; Yang, T.; Yin, S.; Yang, S.; Li, X.; Li, S. Neuroinflammatory Proteins in Huntington's Disease: Insights into Mechanisms, Diagnosis, and Therapeutic Implications. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef] [PubMed]
- Tai, Y.F.; Pavese, N.; Gerhard, A.; Tabrizi, S.J.; Barker, R.A.; Brooks, D.J.; Piccini, P. Microglial activation in presymptomatic Huntington's disease gene carriers. Brain 2007, 130, 1759–1766. [Google Scholar] [CrossRef] [PubMed]
- Yang, H.M.; Yang, S.; Huang, S.S.; Tang, B.S.; Guo, J.F. Microglial Activation in the Pathogenesis of Huntington's Disease. Front Aging Neurosci 2017, 9, 193. [Google Scholar] [CrossRef] [PubMed]
- Croft, M.; Benedict, C.A.; Ware, C.F. Clinical targeting of the TNF and TNFR superfamilies. Nat Rev Drug Discov 2013, 12, 147–168. [Google Scholar] [CrossRef]
- Warner, J.H.; Long, J.D.; Mills, J.A.; Langbehn, D.R.; Ware, J.; Mohan, A.; Sampaio, C. Standardizing the CAP Score in Huntington's Disease by Predicting Age-at-Onset. J Huntingtons Dis 2022, 11, 153–171. [Google Scholar] [CrossRef]
- Cain, M.K.; Zhang, Z.; Bergeman, C.S. Time and Other Considerations in Mediation Design. Educ Psychol Meas 2018, 78, 952–972. [Google Scholar] [CrossRef]
- Mackinnon, D.P.; Lockwood, C.M.; Williams, J. Confidence Limits for the Indirect Effect: Distribution of the Product and Resampling Methods. Multivariate Behav Res 2004, 39, 99. [Google Scholar] [CrossRef]
- Maxwell, S.E.; Cole, D.A. Bias in cross-sectional analyses of longitudinal mediation. Psychol Methods 2007, 12, 23–44. [Google Scholar] [CrossRef]
- Kinnunen, K.M.; Mullin, A.P.; Pustina, D.; Turner, E.C.; Burton, J.; Gordon, M.F.; Scahill, R.I.; Gantman, E.C.; Noble, S.; Romero, K.; et al. Recommendations to Optimize the Use of Volumetric MRI in Huntington's Disease Clinical Trials. Front Neurol 2021, 12, 712565. [Google Scholar] [CrossRef] [PubMed]
- DiFiglia, M.; Leavitt, B.R.; Macdonald, D.; Thompson, L.M.; Huntington's Disease Nomenclature Working, G. Towards Standardizing Nomenclature in Huntington's Disease Research. J Huntingtons Dis 2024, 13, 119–131. [Google Scholar] [CrossRef] [PubMed]
- Warner, J.H.; Sampaio, C. Modeling Variability in the Progression of Huntington's Disease A Novel Modeling Approach Applied to Structural Imaging Markers from TRACK-HD. CPT Pharmacometrics Syst Pharmacol 2016, 5, 437–445. [Google Scholar] [CrossRef]
- Henschel, L.; Conjeti, S.; Estrada, S.; Diers, K.; Fischl, B.; Reuter, M. FastSurfer - A fast and accurate deep learning based neuroimaging pipeline. Neuroimage 2020, 219, 117012. [Google Scholar] [CrossRef]
- Price, R. NINDS-supported biospecimen repositories: BioSEND and NHCDR. Alzheimers Dement 2024, 20. [Google Scholar] [CrossRef]
- Alamar Biosciences. NULISAseq CNS Disease Panel 120 (product page). 2023. [Google Scholar]
- Alamar Biosciences. Technical note: NPQ reporting and normalization. 2023. [Google Scholar]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. Journal of the Royal Statistical Society. Series B (Methodological) 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Kim, J.H. Multicollinearity and misleading statistical results. Korean J Anesthesiol 2019, 72, 558–569. [Google Scholar] [CrossRef]
- Barker, L.E.; Shaw, K.M. Best (but oft-forgotten) practices: checking assumptions concerning regression residuals. Am J Clin Nutr 2015, 102, 533–539. [Google Scholar] [CrossRef]
- Ghasemi, A.; Zahediasl, S. Normality tests for statistical analysis: a guide for non-statisticians. Int J Endocrinol Metab 2012, 10, 486–489. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Ibrahim, J.G.; Cho, H. Perturbation and Scaled Cook's Distance. Ann Stat 2012, 40, 785–811. [Google Scholar] [CrossRef]
- Imai, K.; Keele, L.; Tingley, D. A general approach to causal mediation analysis. Psychol Methods 2010, 15, 309–334. [Google Scholar] [CrossRef] [PubMed]
- MacKinnon, D.P.; Pirlott, A.G. Statistical approaches for enhancing causal interpretation of the M to Y relation in mediation analysis. Pers Soc Psychol Rev 2015, 19, 30–43. [Google Scholar] [CrossRef] [PubMed]
| Region | R_CAP1 | P_CAP | Norm_Stats | -log10(p) |
| Putamen_volmm3 | -0.64 | 1.98 × 10−11 | 0.73 (0.13) | 10.70 |
| Brainsegvol | -0.54 | 5.64 × 10−8 | 197.94 (0.91) | 7.25 |
| Accumbens-area_volmm3 | -0.48 | 1.70 × 10−6 | 0.10 (0.02) | 5.77 |
| Supratentorialvol | -0.46 | 7.18 × 10−6 | 173.48 (2.59) | 5.14 |
| Caudate_volmm3 | -0.44 | 1.41 × 10−5 | 0.53 (0.11) | 4.85 |
| Pallidum_volmm3 | -0.44 | 1.88 × 10−5 | 0.29 (0.05) | 4.73 |
| SubCortGrayVol | -0.42 | 3.83 × 10−5 | 4.74 (0.47) | 4.42 |
| ctx-rh-caudalmiddlefrontal_volmm3 | -0.38 | 2.76 × 10−4 | 0.49 (0.09) | 3.56 |
| ctx-lh-postcentral_volmm3 | -0.34 | 0.001 | 0.89 (0.13) | 2.92 |
| ctx-rh-precentral_volmm3 | -0.33 | 0.002 | 1.01 (0.14) | 2.75 |
| ctx-lh-fusiform_volmm3 | -0.32 | 0.002 | 0.66 (0.09) | 2.68 |
| ctx-lh-superiorfrontal_volmm3 | -0.31 | 0.003 | 2.07 (0.27) | 2.48 |
| ctx-lh-rostralmiddlefrontal_volmm3 | -0.29 | 0.006 | 0.97 (0.14) | 2.21 |
| ctx-rh-superiorfrontal_volmm3 | -0.28 | 0.008 | 2.27 (0.26) | 2.12 |
| ctx-rh-lateraloccipital_volmm3 | -0.27 | 0.011 | 1.07 (0.13) | 1.94 |
| Model | N_complete_case | ADJ_R2 | AIC | BIC | CV_folds | CV_RMSE | CV_MAE |
| No burden (no Age/CAP1) | 88 | 0.355 | -137.9 | -128.0 | 10 | 0.110 | 0.076 |
| Age-adjusted | 88 | 0.350 | -136.2 | -123.8 | 10 | 0.111 | 0.078 |
| CAP-adjusted | 88 | 0.380 | -140.5 | -128.1 | 10 | 0.108 | 0.075 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




