Submitted:
14 January 2026
Posted:
19 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Results
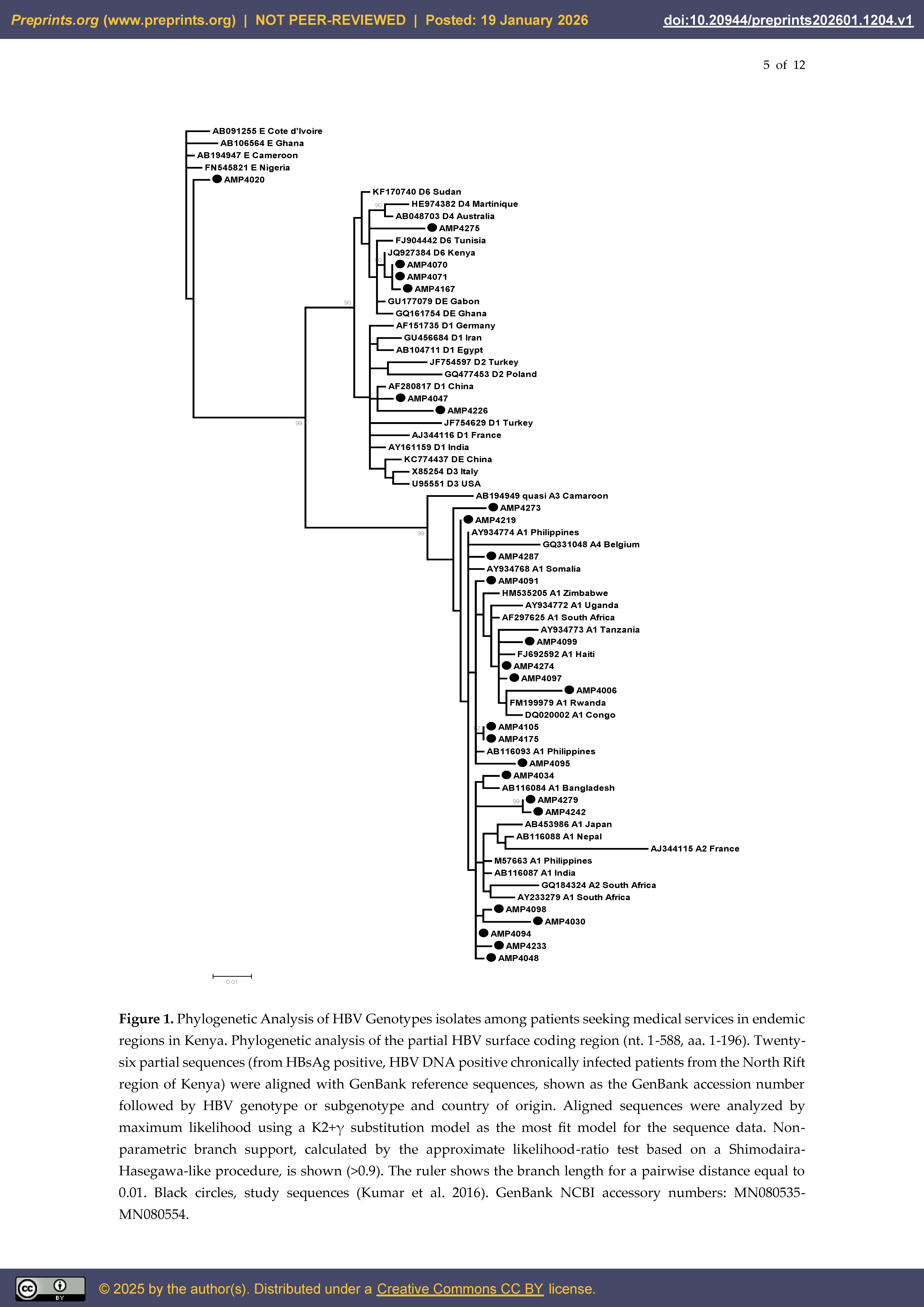
3.1. HBV Genotype A Circulating in Kenya
3.2. HBV Genotype D, E and Their Recombinants
3.3. Identification and Profile of Mutations in in Open Reading Frames of HBV Genotype D and E
3.4. Mutations Occurring in HBV Genotype A1
3.5. Mutations that Co-Existence with Each Other in Pre-Core and BCP Mutations
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BCP/PC | Basal core promoter gene/Pre-Core |
| dNTP | Deoxynucleotide Triphosphate |
| DNA | Deoxyribonucleic Acid |
| HBV | Hepatitis B virus |
| HBV/A1 | Hepatitis B virus Genotype A1 |
| HBV/D | Hepatitis B virus Genotype D |
| HBV/E | Hepatitis B virus Genotype E |
| HCC | Hepatocellular carcinoma |
| HBsAg | Hepatitis B surface antigen |
| KEMRI | Kenya Medical Research Institute’s |
| LMIC | Low- and middle-income countries |
| ORFs | Open Reading Frames |
| PCR | Polymerase Chain Reaction |
| WHO | World Health Organization |
References
- Kramvis, A; Arakawa, K; Yu, MC; Nogueira, R; Stram, DO; Kew, MC. Relationship of serological subtype, basic core promoter and precore mutations to genotypes/subgenotypes of hepatitis B virus. Journal of medical virology 2008, 80(1), 27–46. [Google Scholar] [CrossRef] [PubMed]
- World Health O. Global hepatitis report 2024: action for access in low- and middle-income countries; World Health Organization: Geneva, 2024. [Google Scholar]
- Datfar, T; Doulberis, M; Papaefthymiou, A; Hines, IN; Manzini, G. Viral Hepatitis and Hepatocellular Carcinoma: State of the Art. Pathogens 2021, 10(11), 1366. [Google Scholar] [CrossRef] [PubMed]
- Quasdorff, M; Protzer, U. Control of hepatitis B virus at the level of transcription. J Viral Hepat. 2010, 17(8), 527–36. [Google Scholar] [CrossRef]
- Giersch, K; Allweiss, L; Volz, T; Dandri, M; Lütgehetmann, M. Serum HBV pgRNA as a clinical marker for cccDNA activity. J Hepatol. 2017, 66(2), 460–2. [Google Scholar] [CrossRef]
- Prakash, K; Rydell, GE; Larsson, SB; Andersson, M; Norkrans, G; Norder, H; et al. High serum levels of pregenomic RNA reflect frequently failing reverse transcription in hepatitis B virus particles. Virol J 2018, 15(1), 86. [Google Scholar] [CrossRef]
- Zoulim, F; Chen, PJ; Dandri, M; Kennedy, PT; Seeger, C. Hepatitis B virus DNA integration: Implications for diagnostics, therapy, and outcome. J Hepatol 2024, 81(6), 1087–99. [Google Scholar] [CrossRef]
- Blum, HE; Gerok, W; Vyas, GN. The molecular biology of hepatitis B virus. Trends Genet. 1989, 5(5), 154–8. [Google Scholar] [CrossRef]
- Buckwold, VE; Xu, Z; Chen, M; Yen, TS; Ou, JH. Effects of a naturally occurring mutation in the hepatitis B virus basal core promoter on precore gene expression and viral replication. J Virol. 1996, 70(9), 5845–51. [Google Scholar] [CrossRef]
- Kumar, R. Review on hepatitis B virus precore/core promoter mutations and their correlation with genotypes and liver disease severity. World J Hepatol. 2022, 14(4), 708–18. [Google Scholar] [CrossRef] [PubMed]
- Sheldon, J; Rodès, B; Zoulim, F; Bartholomeusz, A; Soriano, V. Mutations affecting the replication capacity of the hepatitis B virus. J Viral Hepat. 2006, 13(7), 427–34. [Google Scholar] [CrossRef]
- Kramvis, A; Kew, M; François, G. Hepatitis B virus genotypes. Vaccine 2005, 23(19), 2409–23. [Google Scholar] [CrossRef]
- Yuen, MF; Lai, CL. Hepatitis B virus genotypes: natural history and implications for treatment. Expert Rev Gastroenterol Hepatol. 2007, 1(2), 321–8. [Google Scholar] [CrossRef]
- Tatematsu, K; Tanaka, Y; Kurbanov, F; Sugauchi, F; Mano, S; Maeshiro, T; et al. A genetic variant of hepatitis B virus divergent from known human and ape genotypes isolated from a Japanese patient and provisionally assigned to new genotype J. J Virol. 2009, 83(20), 10538–47. [Google Scholar] [CrossRef]
- Kramvis, A; Kew, MC. Epidemiology of hepatitis B virus in Africa, its genotypes and clinical associations of genotypes. Hepatol Res. 2007, 37(s1), S9–s19. [Google Scholar] [CrossRef] [PubMed]
- Lin, CL; Kao, JH. Hepatitis B virus genotypes and variants. Cold Spring Harb Perspect Med. 2015, 5(5), a021436. [Google Scholar] [CrossRef]
- Li, S; Wang, Z; Li, Y; Ding, G. Adaptive evolution of proteins in hepatitis B virus during divergence of genotypes. Scientific Reports 2017, 7(1), 1990. [Google Scholar] [CrossRef]
- Ochwoto, M; Chauhan, R; Gopalakrishnan, D; Chen, CY; Ng’ang’a, Z; Okoth, F; et al. Genotyping and molecular characterization of hepatitis B virus in liver disease patients in Kenya. Infect Genet Evol. 2013, 20, 103–10. [Google Scholar] [CrossRef]
- Kwange, SO; Budambula, NL; Kiptoo, MK; Okoth, F; Ochwoto, M; Oduor, M; et al. Hepatitis B virus subgenotype A1, occurrence of subgenotype D4, and S gene mutations among voluntary blood donors in Kenya. Virus Genes. 2013, 47(3), 448–55. [Google Scholar] [CrossRef]
- Langat, BK; Ochwedo, KO; Borlang, J; Osiowy, C; Mutai, A; Okoth, F; et al. Genetic diversity, haplotype analysis, and prevalence of Hepatitis B virus MHR mutations among isolates from Kenyan blood donors. PLoS One 2023, 18(11), e0291378. [Google Scholar] [CrossRef] [PubMed]
- Ochwoto, M; Kimotho, JH; Oyugi, J; Okoth, F; Kioko, H; Mining, S; et al. Hepatitis B infection is highly prevalent among patients presenting with jaundice in Kenya. BMC Infectious Diseases 2016, 16(1), 101. [Google Scholar] [CrossRef] [PubMed]
- Zirabamuzale, JT; Opio, CK; Bwanga, F; Seremba, E; Apica, BS; Colebunders, R; et al. Hepatitis B virus genotypes A and D in Uganda. J Virus Erad. 2016, 2(1), 19–21. [Google Scholar] [CrossRef]
- Forbi, JC; Dillon, M; Purdy, MA; Drammeh, BS; Tejada-Strop, A; McGovern, D; et al. Molecular epidemiology of hepatitis B virus infection in Tanzania. Journal of General Virology 2017, 98(5), 1048–57. [Google Scholar] [CrossRef] [PubMed]
- Mlewa, M; Henerico, S; Nyawale, HA; Mangowi, I; Shangali, AR; Manisha, AM; et al. The pattern change of hepatitis B virus genetic diversity in Northwestern Tanzania. Scientific Reports 2025, 15(1), 8021. [Google Scholar] [CrossRef] [PubMed]
- Mahgoub, S; Candotti, D; El Ekiaby, M; Allain, JP. Hepatitis B virus (HBV) infection and recombination between HBV genotypes D and E in asymptomatic blood donors from Khartoum, Sudan. J Clin Microbiol. 2011, 49(1), 298–306. [Google Scholar] [CrossRef]
- Yousif, M; Mudawi, H; Hussein, W; Mukhtar, M; Nemeri, O; Glebe, D; et al. Genotyping and virological characteristics of hepatitis B virus in HIV-infected individuals in Sudan. Int J Infect Dis. 2014, 29, 125–32. [Google Scholar] [CrossRef] [PubMed]
- Kurbanov, F; Tanaka, Y; Mizokami, M. Geographical and genetic diversity of the human hepatitis B virus. Hepatol Res. 2010, 40(1), 14–30. [Google Scholar] [CrossRef]
- Croagh, C; Desmond, P; Bell, S. Genotypes and viral variants in chronic hepatitis B: A review of epidemiology and clinical relevance. World Journal of Hepatology 2015, 7(3), 289–303. [Google Scholar] [CrossRef] [PubMed]
- Ito, K; Yotsuyanagi, H; Yatsuhashi, H; Karino, Y; Takikawa, Y; Saito, T; et al. Risk factors for long-term persistence of serum hepatitis B surface antigen following acute hepatitis B virus infection in Japanese adults. Hepatology 2014, 59(1), 89–97. [Google Scholar] [CrossRef]
- Organization WH. Global hepatitis report 2024: action for access in low-and middle-income countries; World Health Organization, 2024. [Google Scholar]
- Ondondo, RO; Muthusi, J; Oramisi, V; Kimani, D; Ochwoto, M; Young, P; et al. Prevalence of hepatitis B virus infection in Kenya: A study nested in the Kenya Population-based HIV Impact Assessment 2018. Plos one 2024, 19(11), e0310923. [Google Scholar] [CrossRef]
- Korir, A; Okerosi, N; Ronoh, V; Mutuma, G; Parkin, M. Incidence of cancer in N airobi, K enya (2004–2008). International journal of cancer 2015, 137(9), 2053–9. [Google Scholar] [CrossRef]
- Günther, S; Li, BC; Miska, S; Krüger, DH; Meisel, H; Will, H. A novel method for efficient amplification of whole hepatitis B virus genomes permits rapid functional analysis and reveals deletion mutants in immunosuppressed patients. J Virol. 1995, 69(9), 5437–44. [Google Scholar] [CrossRef] [PubMed]
- Osiowy, C; Giles, E; Tanaka, Y; Mizokami, M; Minuk, GY. Molecular evolution of hepatitis B virus over 25 years. J Virol. 2006, 80(21), 10307–14. [Google Scholar] [CrossRef]
- Osiowy, C; Kaita, K; Solar, K; Mendoza, K. Molecular characterization of hepatitis B virus and a 9-year clinical profile in a patient infected with genotype I. J Med Virol. 2010, 82(6), 942–8. [Google Scholar] [CrossRef]
- Kowalec, K; Minuk, GY; Børresen, ML; Koch, A; McMahon, BJ; Simons, B; et al. Genetic diversity of hepatitis B virus genotypes B6, D and F among circumpolar indigenous individuals. Journal of Viral Hepatitis 2013, 20(2), 122–30. [Google Scholar] [CrossRef] [PubMed]
- Larkin, MA; Blackshields, G; Brown, NP; Chenna, R; McGettigan, PA; McWilliam, H; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23(21), 2947–8. [Google Scholar] [CrossRef]
- Katoh, K; Misawa, K; Kuma, K; Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 2002, 30(14), 3059–66. [Google Scholar] [CrossRef]
- Kumar, S; Stecher, G; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol Biol Evol. 2016, 33(7), 1870–4. [Google Scholar] [CrossRef]
- Rozanov, M; Plikat, U; Chappey, C; Kochergin, A; Tatusova, T. A web-based genotyping resource for viral sequences. Nucleic Acids Research 2004, 32 suppl_2, W654–W9. [Google Scholar] [CrossRef]
- Ochwoto, M; Ondondo, RO; Matoke, LM; Tuitoek, G; Ogwora, EK; Omari, SW; et al. Predictors, and Trends of Hepatitis B Virus in Selected Regions of Kenya. 2025. [Google Scholar] [CrossRef]
- McNaughton, AL; Revill, PA; Littlejohn, M; Matthews, PC; Ansari, MA. Analysis of genomic-length HBV sequences to determine genotype and subgenotype reference sequences. Journal of General Virology 2020, 101(3), 271–83. [Google Scholar] [CrossRef] [PubMed]
- Toyé, RM; Loureiro, CL; Jaspe, RC; Zoulim, F; Pujol, FH; Chemin, I. The Hepatitis B Virus Genotypes E to J: The Overlooked Genotypes. Microorganisms 2023, 11(8), 1908. [Google Scholar] [CrossRef]
- Aluora, PO; Muturi, MW; Gachara, G. Seroprevalence and genotypic characterization of HBV among low risk voluntary blood donors in Nairobi, Kenya. Virol J 2020, 17(1), 176. [Google Scholar] [CrossRef]
- Kibaya, RM; Lihana, RW; Kiptoo, M; Songok, EM; Ng’ang’a, Z; Osman, S; et al. Characterization of HBV Among HBV/HIV-1 Co-Infected Injecting Drug Users from Mombasa, Kenya. Curr HIV Res. 2015, 13(4), 292–9. [Google Scholar] [CrossRef] [PubMed]
- Mabeya, SN; Ngugi, C; Lihana, RW; Khamadi, SA; Nyamache, AK. Predominance of Hepatitis B Virus Genotype A Among Treated HIV Infected Patients Experiencing High Hepatitis B Virus Drug Resistance in Nairobi, Kenya. AIDS Res Hum Retroviruses 2017, 33(9), 966–9. [Google Scholar] [CrossRef]
- Forbi, JC; Ben-Ayed, Y; Xia, GL; Vaughan, G; Drobeniuc, J; Switzer, WM; et al. Disparate distribution of hepatitis B virus genotypes in four sub-Saharan African countries. J Clin Virol. 2013, 58(1), 59–66. [Google Scholar] [CrossRef] [PubMed]
- Hundie, GB; Stalin Raj, V; Gebre Michael, D; Pas, SD; Koopmans, MP; Osterhaus, AD; et al. A novel hepatitis B virus subgenotype D10 circulating in Ethiopia. J Viral Hepat. 2017, 24(2), 163–73. [Google Scholar] [CrossRef]
- Geoghegan, JL; Duchêne, S; Holmes, EC. Comparative analysis estimates the relative frequencies of co-divergence and cross-species transmission within viral families. PLoS Pathog. 2017, 13(2), e1006215. [Google Scholar] [CrossRef] [PubMed]
- Caligiuri, P; Cerruti, R; Icardi, G; Bruzzone, B. Overview of hepatitis B virus mutations and their implications in the management of infection. World J Gastroenterol. 2016, 22(1), 145–54. [Google Scholar] [CrossRef]
- Yano, Y; Azuma, T; Hayashi, Y. Variations and mutations in the hepatitis B virus genome and their associations with clinical characteristics. World J Hepatol. 2015, 7(3), 583–92. [Google Scholar] [CrossRef]
- Baptista, M; Kramvis, A; Kew, MC. High prevalence of 1762T 1764A mutations in the basic core promoter of hepatitis B virus isolated from black Africans with hepatocellular carcinoma compared with asymptomatic carriers. Hepatology 1999, 29(3), 946–53. [Google Scholar] [CrossRef]
- Liu, CJ; Kao, JH. Core promoter mutations of hepatitis B virus and hepatocellular carcinoma: story beyond A1762T/G1764A mutations; Wiley Online Library, 2008; pp. 347–50. [Google Scholar]
- Tong, MJ; Blatt, LM; Kao, JH; Cheng, JT; Corey, WG. Basal core promoter T1762/A1764 and precore A1896 gene mutations in hepatitis B surface antigen-positive hepatocellular carcinoma: a comparison with chronic carriers. Liver International 2007, 27(10), 1356–63. [Google Scholar] [CrossRef]
- Chauhan, R; Kazim, SN; Bhattacharjee, J; Sakhuja, P; Sarin, SK. Basal core promoter, precore region mutations of HBV and their association with e antigen, genotype, and severity of liver disease in patients with chronic hepatitis B in India. Journal of medical virology 2006, 78(8), 1047–54. [Google Scholar] [CrossRef]
- Guo, X; Jin, Y; Qian, G; Tu, H. Sequential accumulation of the mutations in core promoter of hepatitis B virus is associated with the development of hepatocellular carcinoma in Qidong, China. J Hepatol. 2008, 49(5), 718–25. [Google Scholar] [CrossRef]
- Yang, Z; Zhuang, L; Lu, Y; Xu, Q; Tang, B; Chen, X. Naturally occurring basal core promoter A1762T/G1764A dual mutations increase the risk of HBV-related hepatocellular carcinoma: a meta-analysis. Oncotarget 2016, 7(11), 12525–36. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y; Chang, S; Martin, R; Flaherty, J; Mo, H; Feierbach, B. Characterization of Hepatitis B virus polymerase mutations A194T and CYEI and tenofovir disoproxil fumarate or tenofovir alafenamide resistance. J Viral Hepat. 2021, 28(1), 30–9. [Google Scholar] [CrossRef] [PubMed]
- Dahiya, M; Tai, T; Hussaini, T; Ritchie, G; Matic, N; Yoshida, EM; et al. Virological Response to Lamivudine and Tenofovir Treatment in a Mono-infected Chronic Hepatitis B Patient with Potential Tenofovir Resistance: A Case Report. J Clin Transl Hepatol. 2025, 13(1), 84–7. [Google Scholar] [CrossRef]
- Coppola, N; Onorato, L; Minichini, C; Di Caprio, G; Starace, M; Sagnelli, C; et al. Clinical significance of hepatitis B surface antigen mutants. World J Hepatol. 2015, 7(27), 2729–39. [Google Scholar] [CrossRef]
- Huang, CH; Yuan, Q; Chen, PJ; Zhang, YL; Chen, CR; Zheng, QB; et al. Influence of mutations in hepatitis B virus surface protein on viral antigenicity and phenotype in occult HBV strains from blood donors. J Hepatol. 2012, 57(4), 720–9. [Google Scholar] [CrossRef]
- Lazarevic, I; Banko, A; Miljanovic, D; Cupic, M. Immune-Escape Hepatitis B Virus Mutations Associated with Viral Reactivation upon Immunosuppression. Viruses 2019, 11(9). [Google Scholar] [CrossRef]
- Jepkemei, KB; Ochwoto, M; Swidinsky, K; Day, J; Gebrebrhan, H; McKinnon, LR; et al. Characterization of occult hepatitis B in high-risk populations in Kenya. PLoS One 2020, 15(5), e0233727. [Google Scholar] [CrossRef] [PubMed]
| PhenoP1* (1821-1843) 5’-CGGAAAGCTTATGCTCTTCTTTTTCACCTCTGCCTAATCATC -3’ PhenoP2* (1825-1804) 5’-CCGGAGAGCTCATGCTCTTCAAAAAGTTGCATGGTGCTGGTG –3’ FLG1 (F) (1826-1843) 5’- CACCTCTGCCTAATCATC –3’ FLG2 (R) (1820-1804) 5’- GTTGCATGGTGCGGTC-3’ S1f† 5’- TCCTGCTGGTGGCTCCAG-3 S1r† 5’- CGTTGACATACTTTCCAATCAA-3’ C1† (EP1) GCATGGAGACCACCGTGAAC C1† (EP2) GGAAAGAAGTCAGAAGGCAA C2† (EP3) CATAAGAGGACTCTTGGACT C2† (EP4) GGCAAAAAAGAGAGTAACTC *Günther et al., 1995(33); †Osiowy C et al., 2010 (35) |
| Code No. | Genotype (Serotype) | Pre-S2 mut (aa 5-55) | HBsAg mut (aa 1-196) | RT (pol) rt80 (incl 25 seq)—rt236/250 | BCP/PC (nt1653-1896) |
|---|---|---|---|---|---|
| AMP4020 | E(Ayw4) | - | N59S | - | G1757A |
| AMP4047 | D1(Ayw2) | - | R79H | - | G1757A, G1896A |
| AMP4070 | D6(Ayw2) | - | - | - | G1896A |
| AMP4071 | D6(Ayw2) | - | - | - | G1896A |
| AMP4167 | D6(Ayw2) | D51V | - | - | C1653T, T1753C, G1757A, A1762T, G1764A, G1896A |
| AMP4226 | D1(Ayw2) | - | I4V, T23E, L26H, S31T, D33N | - | A1752C, G1757A |
| AMP4275 | D/E(Ayw2) | - | V14A, C69stop, S104T, W182stop | V191I | 9bp ins at nt 1687, G1757A, A1762T, G1764A, G1896A |
| AMP4006 | A1(adw2) | L12Q, G102A, T104K, G130S, F134L | |||
| AMP4030 | A1(adw2)) | C76Y, G130S | |||
| AMP4034 | A1(adw2) | W36L | |||
| AMP4048 | A1(adw2) | D33N | A1762T, G1764A | ||
| AMP4095 | A1(adw2) | P46H, V177A | |||
| AMP4097 | A1(adw2) | C1766T | |||
| AMP4098 | A1(adw2) | F20S | |||
| AMP4233 | A1(adw2) | G7R, Y161F | |||
| AMP4242 | A1(adw2) | V14A, S58C, P67Q | A194T | ||
| AMP4273 | A1(adw2) | P46L, L49H | |||
| AMP4279 | A1(adw2) | V14A, S58C, P67Q | |||
| Pre-S2 mut (aa 5-55) incl 14 seq HBsAg mut (aa 1-196) incl 26 seq RT (pol) rt80 (incl 25 seq)—rt236/250 (incl 17 seq) BCP/PC (nt1653-1896) incl 12 seq | |||||
| Sample ID_Genotype | Pre-core and BCP mutations | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1753 | 1762, 1764 | 1802–1803 | 1809–1812 | 1814, 1816 | 1858 | 1862 | 1884 | 1888 | 1896 | 1899 | |
| AMP4047_D1 | T | A,G | CG | GTAC | T,G | T | G | T | G | A | G |
| AMP4070_D6 | T | A,G | CG | GCAC | A,T | T | G | T | G | A | A |
| AMP4226_D1 | T | A,G | CG | GCAC | A,G | T | G | T | G | G | G |
| AMP4071_D6 | T | A,G | CG | GCAC | A,T | T | G | T | G | A | A |
| AMP4167_D6 | C | T,A | CG | GCAC | A,A | T | G | T | G | A | A |
| AMP4275_D/E | T | T,A | CG | GCAC | A,G | T | G | T | G | A | G |
| AMP4020_E | T | A,G | CG | GCAC | A,G | C | G | Y | G | G | G |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




