Submitted:
09 January 2026
Posted:
13 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Machine Learning Approaches for Schizophrenia EEG Classification
1.2. The Generalization Problem
- Recording Equipment Heterogeneity: Different EEG systems introduce systematic differences in signal characteristics, including varying amplifier properties, electrode impedance specifications, input impedance characteristics, common-mode rejection ratios, and analog-to-digital conversion specifications. These hardware differences introduce systematic biases that classifiers may learn as spurious features.
- Protocol Variations: Differences in recording conditions (eyes-open vs. eyes-closed, task vs. rest, recording duration, environmental noise levels, time of day, subject positioning) affect signal characteristics in ways that vary systematically across sites.
- Population Differences: Variations in patient demographics, illness duration, symptom severity, medication status (type, dosage, polypharmacy), comorbidities, and other clinical factors across datasets influence EEG patterns in ways that may not reflect core disorder characteristics. If these clinical factors differ systematically between sites, classifiers may learn to discriminate based on these secondary characteristics rather than disease-relevant features.
- Diagnostic Criteria and Assessment: Differences in how schizophrenia diagnosis was established (diagnostic instruments used, clinician training, threshold for diagnosis, inclusion/exclusion criteria) may lead to heterogeneity in the patient groups labeled as “schizophrenia” across datasets.
- Preprocessing Inconsistencies: Different artifact rejection strategies, filtering approaches, and referencing schemes can alter signal properties and contribute to site-specific signatures that classifiers may exploit.
1.3. Study Objectives and Importance of Transparent Reporting
- Compare EEGNet and calibrated Random Forest performance under subject-level stratified cross-validation on the ASZED-153 training dataset.
- Evaluate generalization to an independent external dataset (Kaggle Schizophrenia EEG) acquired at a different site with different equipment and protocols.
- Quantify the domain shift between datasets using Maximum Mean Discrepancy (MMD).
- Assess whether CORAL domain adaptation can improve external dataset performance.
- Provide transparent reporting of generalization failures to inform realistic assessment of technology readiness.
2. Materials and Methods
2.1. Datasets
2.1.1. ASZED-153 Dataset (Training)
2.1.2. External Validation Dataset (Kaggle Schizophrenia EEG)
- Healthy Controls: 39 subjects without psychiatric diagnoses
- Schizophrenia Patients: 45 adolescent subjects exhibiting symptoms of schizophrenia
2.2. Preprocessing Pipeline
2.2.1. Channel Standardization
2.2.2. Signal Filtering
- Bandpass filtering: 0.5–45 Hz (4th order Butterworth filter, zero-phase implementation using filtfilt to avoid phase distortion)
- Notch filtering: 50 Hz and 60 Hz (for power line noise removal, applied based on recording location)
- Resampling: All signals resampled to 250 Hz using polyphase filtering with anti-aliasing to prevent artifacts
2.2.3. Epoch Extraction and Artifact Rejection
2.2.4. Normalization
- For EEGNet (deep learning): Raw epoch data were z-score normalized per channel to account for inter-channel amplitude differences. This normalization was computed independently for each epoch: , where and are computed from the specific epoch.
- For Random Forest (classical ML): Spectral features were standardized using training set statistics (mean and standard deviation computed from training folds), with the same transformation applied to validation and test data to prevent information leakage.
2.3. Feature Extraction for Random Forest
2.3.1. Spectral Band Power Features
- Delta (0.5–4 Hz): Associated with sleep and unconscious processes
- Theta (4–8 Hz): Linked to drowsiness, meditation, and memory processes
- Alpha (8–13 Hz): Dominant during relaxed wakefulness; often reduced in schizophrenia
- Beta (13–30 Hz): Associated with active thinking and anxiety
- Gamma (30–45 Hz): Related to cognitive processing and sensory binding
2.3.2. Coherence Features
2.3.3. Phase Lag Index (PLI)
2.3.4. Statistical Features
- Mean amplitude (normalized by standard deviation)
- Standard deviation (normalized by absolute mean)
- Skewness (asymmetry of amplitude distribution)
- Kurtosis (tailedness of amplitude distribution)
- Root-mean-square amplitude (normalized by standard deviation)
- Peak-to-peak amplitude (range, normalized by standard deviation)
2.4. Model Architectures
2.4.1. EEGNet Architecture
- 2D temporal convolution: 8 filters, kernel size (1, 64)
- Batch normalization
- Purpose: Learn temporal filters capturing frequency-specific patterns
- Depthwise 2D spatial convolution: kernel size (16, 1), depth multiplier D=2
- Batch normalization
- ELU (Exponential Linear Unit) activation
- Average pooling (1, 4) for temporal downsampling
- Dropout () for regularization
- Purpose: Learn spatial filters specific to each temporal filter
- Separable 2D convolution: 16 filters, kernel size (1, 16)
- Batch normalization
- ELU activation
- Average pooling (1, 8) for further temporal downsampling
- Dropout () for regularization
- Purpose: Learn higher-level temporal features
- Flatten layer
- Fully connected layer: Output dimension = 2 (binary classification)
- Softmax activation for probability estimates
- Optimizer: Adam (, , weight decay )
- Learning rate: 0.001 with ReduceLROnPlateau scheduler (factor=0.5, patience=5 epochs)
- Loss function: Cross-entropy with class weights inversely proportional to class frequencies
- Early stopping: Patience of 15 epochs based on validation loss
- Maximum epochs: 100
- Batch size: 32
- Gradient clipping: Max norm = 1.0 to prevent exploding gradients
2.4.2. Random Forest with Isotonic Calibration
- Number of trees: 500 (increased from default for ensemble diversity)
- Maximum depth: 15 (moderate depth to prevent overfitting)
- Minimum samples per leaf: 2 (conservative to capture subtle patterns)
- Class weights: Balanced (inversely proportional to class frequencies)
- Maximum features: (default, for decorrelation)
- Bootstrap: True (with replacement for training each tree)
- Random state: 42 (fixed for reproducibility)
- n_jobs: -1 (parallel processing using all CPU cores)
2.5. Cross-Validation Strategy
- No subject appears in both training and validation sets within a fold
- Each subject appears in exactly one validation fold
- Class balance is preserved across folds
2.6. Domain Adaptation: CORAL
- is the source feature matrix (ASZED-153 training data)
- is the target feature matrix (external validation data)
- is the source covariance matrix with regularization
- is the target covariance matrix with regularization
- is a small regularization term for numerical stability
2.7. Domain Shift Quantification: Maximum Mean Discrepancy
- Kernel: Gaussian RBF kernel
- Bandwidth: (median heuristic)
- Dimensionality reduction: PCA to 100 components before MMD computation for computational efficiency
- Sample size: Maximum 200 samples per dataset for tractable computation
2.8. Pilot Hardware Exploration: BioAmp EXG Pill
- Board dimensions: 25.4 mm × 10.0 mm
- Input channels: 1 differential channel (IN+, IN-, REF)
- Gain: Configurable (default: high gain for EEG)
- Bandwidth: Configurable bandpass filter
- Interface: Analog output compatible with standard ADCs
- Cost: Approximately $15 USD per unit
- Microcontroller: ESP32 (Espressif Systems)
- Sampling rate: 256 Hz (14-bit ADC)
- Electrode placement: Fp1 (IN+), Fp2 (IN-), ear reference (REF)
- Recording format: European Data Format (EDF)
- Software: Custom Python script for visualization and EDF export
2.9. Evaluation Metrics
- Accuracy: Proportion of correct predictions:
- AUC-ROC: Area under the Receiver Operating Characteristic curve, measuring discrimination ability across all classification thresholds
- Sensitivity (Recall): True positive rate: (proportion of schizophrenia patients correctly identified)
- Specificity: True negative rate: (proportion of healthy controls correctly identified)
- Balanced Accuracy: Average of sensitivity and specificity: (accounts for class imbalance)
- Brier Score: Mean squared error between predicted probabilities and true binary labels: , measuring calibration quality (lower is better)
- Expected Calibration Error (ECE): Average absolute difference between predicted probability and observed accuracy within 10 equally-spaced probability bins, providing another measure of calibration
2.10. Statistical Analysis
3. Results
3.1. Dataset Characteristics
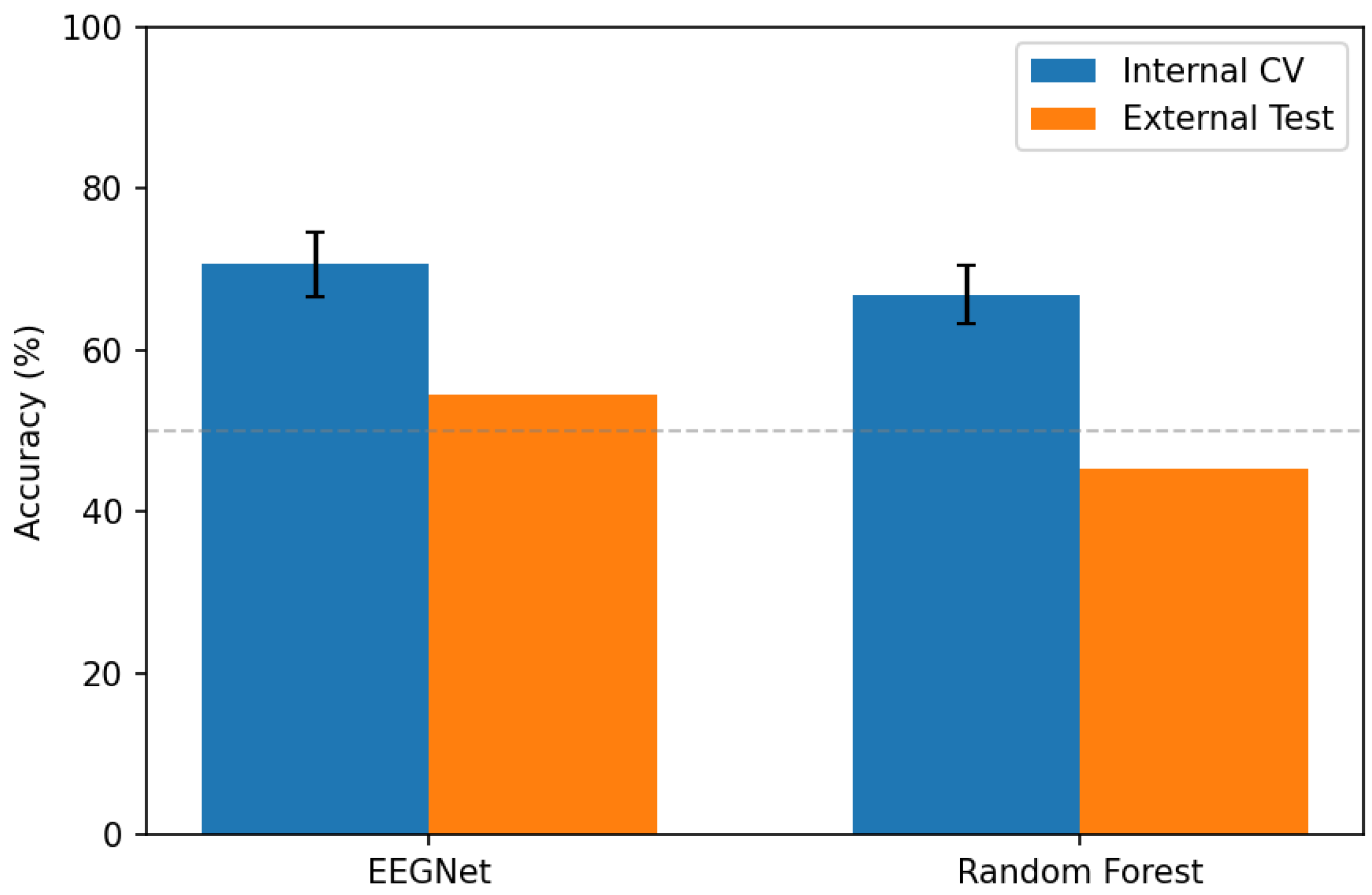
3.2. Internal Cross-Validation Performance
3.3. Per-Fold Analysis
3.4. External Dataset Generalization
- EEGNet: Accuracy dropped from 70.7% (internal CV) to 54.6% (external), representing a generalization gap of 16.1 percentage points. AUC declined from 0.814 to 0.529 (essentially chance level for binary classification). Notably, EEGNet developed extreme bias toward predicting schizophrenia (sensitivity 87.5%, specificity 16.6%), correctly identifying most patients but misclassifying most controls.
- Random Forest: Accuracy dropped from 66.8% to 45.4% (generalization gap: 21.4 percentage points). AUC of 0.424 indicates performance below chance level, suggesting systematic misclassification. Random Forest showed more balanced predictions than EEGNet but still far below acceptable clinical performance.
- RF + CORAL: Domain adaptation provided marginal improvement (1.2 percentage points accuracy increase, AUC improvement from 0.424 to 0.470). CORAL dramatically altered the classifier’s decision boundary, flipping from high sensitivity (60.2%) to high specificity (79.2%), but overall discrimination remained poor. This suggests that simple covariance alignment is insufficient to address the complex domain shift between these datasets.
3.5. Generalization Gap Analysis
- EEGNet: Gap of 16.1 percentage points vs. internal SD of 4.0 (ratio: 4.0×)
- Random Forest: Gap of 21.4 percentage points vs. internal SD of 3.6 (ratio: 5.9×)
3.6. Domain Shift Analysis
- Original MMD (ASZED-153 vs. External): 0.0914
- Post-CORAL MMD (ASZED-153 transformed vs. External): 0.0843
- MMD reduction: 7.8%
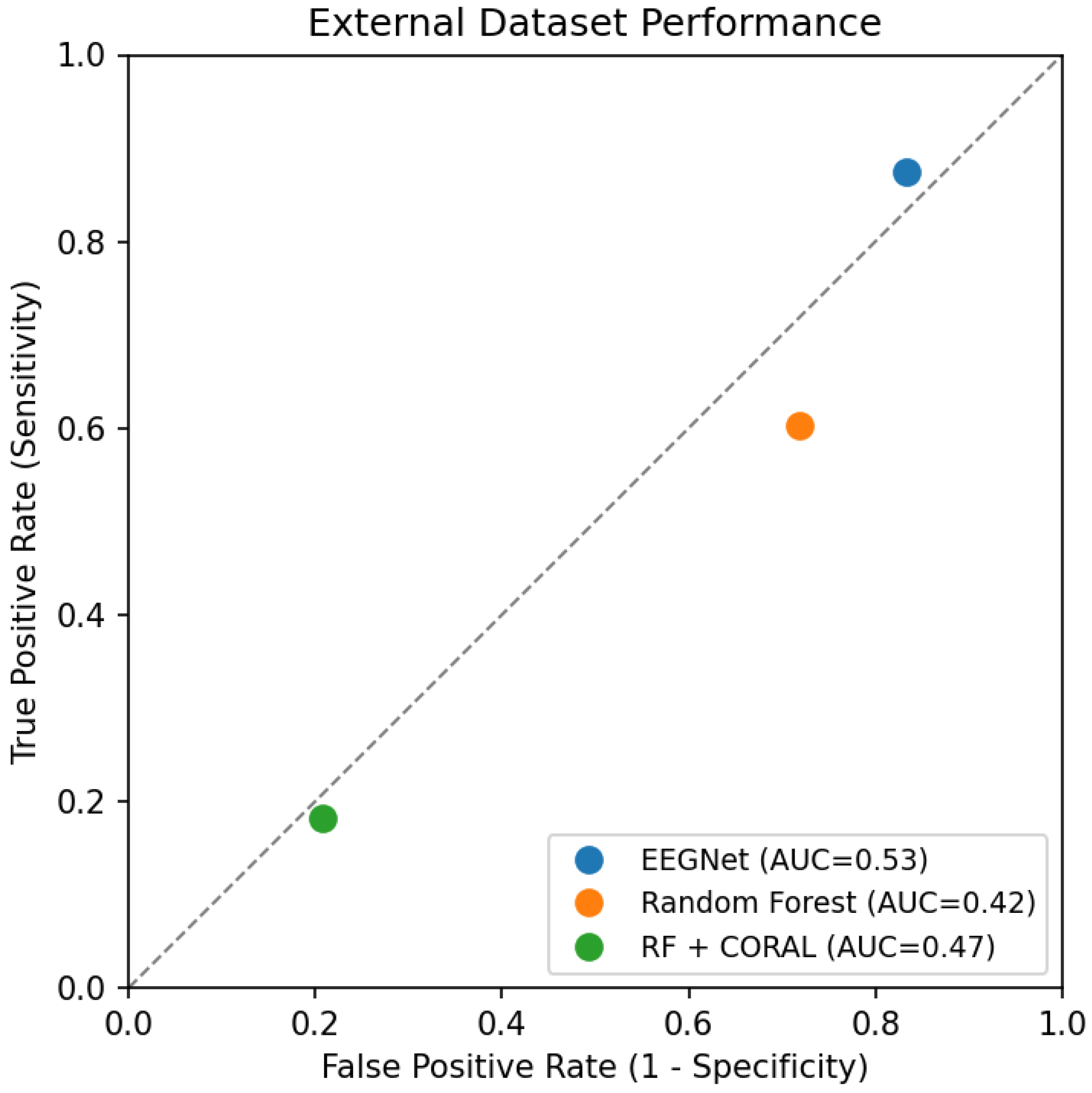
3.7. ROC Curve Analysis
- EEGNet prioritizes sensitivity (87.5%) at the cost of specificity (16.6%)
- Random Forest shows moderate sensitivity (60.2%) and poor specificity (28.2%)
- CORAL flips the trade-off, achieving reasonable specificity (79.2%) but very low sensitivity (18.2%)
3.8. Calibration Analysis
- EEGNet: Brier score increased from 0.213 (internal) to 0.350 (external), and ECE increased from approximately 0.15 (estimated from internal folds) to 0.205
- Random Forest: Brier score increased from 0.211 to 0.261, and ECE increased from approximately 0.19 to 0.266
3.9. Feature Importance Analysis
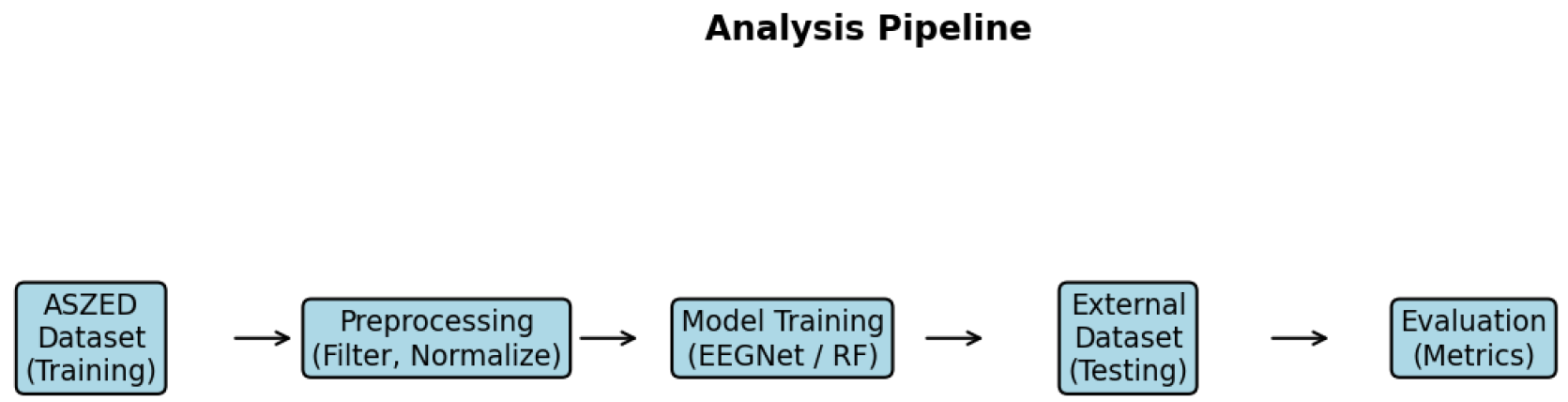
3.10. Analysis Pipeline
4. Discussion
4.1. Principal Findings
- EEGNet outperformed Random Forest during internal cross-validation on ASZED-153 (70.7% vs. 66.8% accuracy, , Cohen’s ), demonstrating that deep learning can achieve superior within-dataset performance for EEG-based psychiatric classification.
- Both approaches exhibited severe generalization difficulty on the external Kaggle dataset (EEGNet: 54.6%, Random Forest: 45.4%), with performance degrading to near or below chance level. These results represent generalization gaps of 16–21 percentage points.
- CORAL domain adaptation provided marginal improvement (1.2 percentage points for Random Forest, AUC improvement from 0.424 to 0.470) but did not approach internal validation performance levels or achieve clinically acceptable discrimination.
- MMD analysis quantitatively confirmed substantial domain shift between datasets (MMD = 0.0914, well above the 0.05 threshold for meaningful difference).
- The generalization gap far exceeded internal cross-validation variance (4–6× larger), indicating that CV-based performance estimates are overly optimistic and do not predict external performance.
- Both models showed severe miscalibration on external data, with probability estimates becoming unreliable (ECE increased from ∼0.15–0.19 to 0.21–0.27).
4.2. Interpretation of Generalization Results
4.2.1. Architecture Comparison: Deep Learning vs. Classical Machine Learning
4.2.2. Potential Sources of Domain Shift
- Recording Equipment Differences: ASZED-153 and Kaggle datasets were acquired with different EEG systems, introducing systematic hardware-related differences. Different amplifier characteristics, electrode types (passive vs. active), input impedance specifications, common-mode rejection ratios, and analog-to-digital conversion properties create systematic biases in recorded signals. Classifiers may learn these hardware signatures as “features” that correlate with class labels in the training data but do not generalize to different hardware.
- Protocol and Environmental Variations: Differences in recording protocols (exact instructions to subjects, level of supervision, duration, number of recordings per subject), environmental factors (ambient noise levels, time of day, room lighting), and subject state (alertness, anxiety level, caffeine consumption) affect EEG characteristics systematically across sites. Resting-state recordings are particularly sensitive to subject state and environmental factors.
- Population and Clinical Heterogeneity: ASZED-153 includes adult African populations, while the Kaggle dataset contains adolescent subjects of unspecified ethnicity. Age-related differences in EEG patterns (adolescents show different spectral characteristics than adults), demographic factors, cultural differences in response to experimental procedures, and differences in illness characteristics (age of onset, symptom severity, duration) introduce systematic variations. Additionally, medication status (type, dosage, polypharmacy, compliance) is unknown for both datasets but likely differs systematically.
- Diagnostic Assessment Heterogeneity: Differences in how schizophrenia diagnosis was established (specific diagnostic criteria used, diagnostic instruments, clinician training and experience, threshold for diagnosis, inclusion/exclusion criteria) may lead to heterogeneity in patient groups across datasets. The Kaggle dataset includes “adolescents exhibiting symptoms of schizophrenia,” which may include prodromal cases, first-episode psychosis, or attenuated psychosis syndrome—populations with potentially different EEG characteristics than chronic schizophrenia patients.
- Artifact and Noise Characteristics: Different recording environments and subject populations produce different types and levels of artifacts. Muscle artifacts, eye movements, and electrode artifacts may have different statistical properties across datasets, and classifiers trained with one artifact profile may fail when encountering different artifact patterns.
4.2.3. Implications for the Field
- External Validation is Essential: Internal cross-validation alone, while scientifically valid for algorithmic comparison and model selection, does not provide adequate evidence of real-world generalization or clinical readiness. Studies reporting high accuracy based solely on internal validation should be interpreted cautiously regarding clinical applicability. The field should establish standards requiring external validation on multiple independent datasets before claims of clinical utility are advanced.
- Publication of Generalization Failures: The field benefits from transparent reporting of generalization failures and negative results. Publication bias favoring positive results creates an unrealistic impression of technology readiness, delays identification of fundamental challenges, and may lead to premature clinical deployment attempts. Our findings join a growing body of work documenting cross-dataset generalization challenges in medical AI.
- Multi-Site Training Data May Be Necessary: Single-site training data may be fundamentally insufficient for developing clinically deployable classifiers. Successful translation likely requires multi-site training to learn features that generalize across acquisition conditions rather than site-specific patterns. This necessitates large-scale collaborative efforts and data-sharing initiatives, which face substantial logistical, regulatory, and financial challenges.
- Rethinking Performance Benchmarks: The community should reconsider how classifier performance is evaluated and reported. Reporting only internal CV performance without external validation provides an incomplete and potentially misleading picture. We recommend that papers report both internal CV performance (for within-dataset comparisons) and external validation performance (for generalization assessment), clearly distinguishing between these two evaluation modes.
4.3. Limitations of Domain Adaptation
- Limited Statistical Alignment: CORAL aligns only second-order statistics (covariance matrices). Higher-order distributional differences (skewness, kurtosis, multimodal structures) and complex nonlinear relationships remain unaddressed. If the domain shift involves these higher-order patterns, second-order alignment is insufficient.
- Potential Concept Shift: If the feature-label relationship differs between datasets (e.g., if alpha power has different diagnostic significance across sites due to population differences or if different EEG features are relevant in different populations), covariance alignment cannot resolve this mismatch. CORAL assumes that class-conditional distributions differ between domains but the underlying discriminative structure is preserved—an assumption that may not hold for psychiatric EEG classification across diverse populations.
- Magnitude of Shift: The large MMD value (0.0914) and dramatic performance degradation suggest a domain shift too severe for simple alignment techniques. CORAL reduced MMD by only 7.8%, indicating that most of the distributional difference remains after alignment.
- Feature Relevance: If the important features for classification differ between datasets (e.g., different frequency bands or channels carry diagnostic information), aligning feature distributions will not help if the model is attending to the wrong features for the target domain.
4.4. Comparison with Literature
4.5. Toward Improved Generalization
4.5.1. Multi-Site Training Data and Federated Learning
4.5.2. Data Harmonization and Standardization
- Standardized recording protocols (electrode montages, recording duration, instructions to subjects, environmental conditions)
- Comprehensive metadata reporting (equipment specifications, software versions, electrode types, impedance levels, medication status, illness characteristics)
- Standardized preprocessing pipelines with open-source implementations and detailed documentation
- Quality control metrics and standardized artifact rejection criteria
- Common data formats and naming conventions
4.5.3. Domain-Invariant and Meta-Learning Approaches
- Domain-adversarial neural networks [14]: Train networks to learn features that are predictive of diagnosis but uninformative about data source (site), explicitly optimizing for domain invariance
- Invariant risk minimization (IRM): Identify features with stable predictive relationships across multiple environments (datasets), focusing on causal rather than correlational features
- Meta-learning (learning to learn): Train models on multiple datasets/domains with the explicit objective of rapid adaptation to new domains with limited data
- Test-time adaptation: Adjust model parameters at test time using unlabeled target domain data to adapt to distributional shift without requiring target labels
4.5.4. Physiologically-Informed and Hybrid Approaches
- Focus on features with known physiological interpretations and documented relevance to schizophrenia pathophysiology
- Use biophysical models to generate “synthetic” training data covering diverse acquisition conditions
- Hybrid approaches combining data-driven feature learning with neuroscience-informed priors
- Incorporate known invariances (e.g., certain spectral ratios may be more robust than absolute power)
4.5.5. Low-Cost Hardware for Accessible EEG Acquisition
- Enables large-scale data collection in resource-constrained settings (low- and middle-income countries, rural areas, community mental health settings)
- Increases diversity of training data through broader geographical and demographic sampling
- Facilitates home-based or ambulatory EEG recording for longitudinal monitoring
- Reduces barriers to EEG research in underrepresented populations
- Signal quality validation: Systematic comparison with clinical-grade systems needed
- Standardization: Developing standardized acquisition protocols for low-cost hardware
- Regulatory pathway: Understanding regulatory requirements for clinical use
- Generalization across hardware types: Whether models trained on clinical-grade EEG generalize to low-cost systems (and vice versa) requires investigation
4.6. Study Limitations
- Limited Dataset Diversity: Our study utilized two datasets from specific research groups and regions. Generalization to other datasets, populations, geographic regions, and clinical settings cannot be assumed. Evaluation on additional external datasets representing diverse populations, ages, illness stages, and recording conditions would strengthen conclusions about the generality of our findings.
- Residual Preprocessing Differences: Despite our efforts to apply harmonized preprocessing, we cannot fully exclude residual preprocessing-related differences between datasets. Ideal evaluation would use data collected with identical protocols and equipment, or use raw unprocessed data with all preprocessing performed in-house.
- Limited Architectural Diversity: We compared one deep learning architecture (EEGNet) against one classical approach (Random Forest with isotonic calibration). Other architectures (recurrent neural networks, transformers, more complex CNNs, other ensemble methods) might demonstrate different generalization characteristics. However, we hypothesize similar challenges would emerge given that the fundamental issue is dataset heterogeneity rather than architecture choice.
- Single External Dataset: Generalization was evaluated on one external dataset. Evaluation on multiple independent external datasets from diverse sources would provide more robust assessment of generalization capability and help identify which factors most impact cross-site performance.
- Incomplete Clinical Metadata: Limited availability of detailed clinical metadata (medication specifics, symptom profiles, illness duration, functional status) for both datasets restricted our ability to investigate whether clinical heterogeneity contributes to generalization failure. Rich clinical phenotyping would enable more nuanced analysis of generalization across different patient subgroups.
- Resting-State Focus: We utilized only resting-state EEG data. Task-based paradigms (cognitive tasks, auditory oddball, emotional face processing) or event-related potential approaches might offer different generalization characteristics, as task-evoked responses may be more stereotyped across individuals and sites.
- Binary Classification: We focused on binary classification (schizophrenia vs. healthy controls). Real-world clinical deployment would require discrimination from other psychiatric and neurological conditions (bipolar disorder, depression, epilepsy), which presents additional challenges.
4.7. Recommendations for Future Research
- Mandatory External Validation: Journals and conferences should require external validation on at least one independent dataset for studies claiming clinical utility or readiness of EEG-based diagnostic classifiers. Reporting standards should clearly distinguish between internal CV performance (for methodological comparison) and external validation performance (for generalization assessment).
- Multi-Site Consortia: Establish multi-site research consortia with standardized protocols for EEG data collection, quality control, and phenotypic characterization in psychiatric populations. Such consortia could create benchmark datasets with held-out external test sets for fair comparison of methods.
- Public Benchmark Datasets: Develop standardized, multi-site benchmark datasets with comprehensive metadata, rigorous quality control, and held-out test sets for evaluating cross-site generalization of psychiatric EEG classifiers. These benchmarks would enable fair comparison of methods and track progress over time.
- Transparency in Negative Results: Strongly encourage publication of studies demonstrating generalization failures and negative results. These contributions provide realistic assessment of current technology readiness, identify fundamental challenges requiring solutions, and prevent wasteful duplication of failed approaches.
- Shift Focus to Robustness: Redirect research emphasis from maximizing single-dataset accuracy to developing methods that maintain reasonable performance across diverse acquisition conditions. Generalization robustness, not peak performance, should be the primary metric for clinical translation.
- Investigate Failure Modes: Systematic investigation of which factors most impact generalization (equipment type, protocol differences, population characteristics, clinical heterogeneity) through carefully controlled experiments varying one factor at a time.
- Develop Generalization Metrics: Establish standardized metrics for quantifying and reporting generalization performance, such as average performance across multiple external datasets, worst-case performance across datasets, or generalization gap (internal - external performance).
5. Conclusions
- External validation is essential and non-negotiable: Internal cross-validation, while scientifically valid for methodological comparisons, does not provide adequate evidence of clinical utility or real-world applicability. Studies relying solely on internal validation should be interpreted cautiously regarding readiness for clinical deployment.
- Architectural sophistication alone is insufficient: The similarity of generalization failure between deep learning (EEGNet) and classical machine learning (Random Forest) suggests that the challenge lies primarily in dataset heterogeneity and site-specific feature learning rather than algorithmic limitations. Solutions will require addressing data collection, harmonization, and study design.
- Current methods are not deployment-ready: Performance approaching chance level on external data definitively indicates that existing approaches require substantial development before clinical deployment is appropriate. Claims of clinical readiness should require rigorous multi-site external validation.
- Multi-site training is likely necessary: Single-site training data may be fundamentally inadequate for developing robust, generalizable classifiers suitable for clinical use across diverse healthcare settings. Large-scale collaborative efforts are needed.
- Transparent reporting of negative results is valuable: This work contributes to realistic assessment of technology readiness by transparently documenting generalization failures. Such contributions, though reporting negative findings, provide essential scientific value by identifying fundamental challenges and preventing premature clinical deployment attempts.
Institutional Review Board Statement
Data Availability Statement: ASZED-153 Dataset (Training)
Acknowledgments
References
- Tandon, R., Nasrallah, H.A., Keshavan, M.S. Schizophrenia, “just the facts” 4. Clinical features and conceptualization. Schizophrenia Research, 110(1-3):1–23, 2009.
- Marshall, M., Lewis, S., Lockwood, A., Drake, R., Jones, P., Croudace, T. Association between duration of untreated psychosis and outcome in cohorts of first-episode patients: a systematic review. Archives of General Psychiatry, 62(9):975–983, 2005.
- Boutros, N.N., Arfken, C., Galderisi, S., Warrick, J., Pratt, G., Iacono, W. The status of spectral EEG abnormality as a diagnostic test for schizophrenia. Schizophrenia Research, 99(1-3):225–237, 2008.
- Newson, J.J., Thiagarajan, T.C. EEG frequency bands in psychiatric disorders: a review of resting state studies. Frontiers in Human Neuroscience, 12:521, 2019.
- Uhlhaas, P.J., Singer, W. Abnormal neural oscillations and synchrony in schizophrenia. Nature Reviews Neuroscience, 11(2):100–113, 2010.
- Santos-Mayo, L., San-Jose-Revuelta, L.M., Arribas, J.I. A computer-aided diagnosis system with EEG based on the P3b wave during an auditory odd-ball task in schizophrenia. IEEE Transactions on Biomedical Engineering, 64(2):395–407, 2020.
- Craik, A., He, Y., Contreras-Vidal, J.L. Deep learning for electroencephalogram (EEG) classification tasks: a review. Journal of Neural Engineering, 16(3):031001, 2019.
- Lawhern, V.J., Solon, A.J., Waytowich, N.R., Gordon, S.M., Hung, C.P., Lance, B.J. EEGNet: a compact convolutional neural network for EEG-based brain–computer interfaces. Journal of Neural Engineering, 15(5):056013, 2018.
- Sun, B., Feng, J., Saenko, K. Return of frustratingly easy domain adaptation. Proceedings of the AAAI Conference on Artificial Intelligence, 30(1), 2016.
- ASZED-153: African Schizophrenia EEG Dataset. Zenodo, 2024. https://zenodo.org/records/14178398.
- Niculescu-Mizil, A., Caruana, R. Predicting good probabilities with supervised learning. Proceedings of the 22nd International Conference on Machine Learning, pages 625–632, 2005.
- Gretton, A., Borgwardt, K.M., Rasch, M.J., Schölkopf, B., Smola, A. A kernel two-sample test. Journal of Machine Learning Research, 13:723–773, 2012.
- Rieke, N., Hancox, J., Li, W., Milletari, F., Roth, H.R., Albarqouni, S., et al. The future of digital health with federated learning. NPJ Digital Medicine, 3(1):119, 2020.
- Ganin, Y., Ustinova, E., Ajakan, H., Germain, P., Larochelle, H., Laviolette, F., Marchand, M., Lempitsky, V. Domain-adversarial training of neural networks. Journal of Machine Learning Research, 17(1):2096–2030, 2016.
| Characteristic | ASZED-153 (Training) | Kaggle (External) |
|---|---|---|
| Total subjects | 153 | 84 |
| Schizophrenia patients | 76 | 45 |
| Healthy controls | 77 | 39 |
| Class balance (SZ/HC) | 0.99 | 1.15 |
| Total epochs (2-second) | 13,449 | 2,436 |
| Epochs per subject (mean) | 87.9 | 29.0 |
| Channels (standardized) | 16 | 16 |
| Sampling rate | 250 Hz | 250 Hz |
| Epoch duration | 2.0 s | 2.0 s |
| Recording condition | Resting state | Resting state |
| Model | Acc. | AUC | Sens. | Spec. | Bal. Acc. | Brier |
|---|---|---|---|---|---|---|
| EEGNet | 70.7±4.0 | 0.814±0.045 | 67.9% | 76.8% | 72.3% | 0.213 |
| Random Forest | 66.8±3.6 | 0.778±0.057 | 55.8% | 79.0% | 67.4% | 0.211 |
| Model | Fold 1 | Fold 2 | Fold 3 | Fold 4 | Fold 5 | Mean | SD |
|---|---|---|---|---|---|---|---|
| EEGNet | 72.9 | 72.6 | 67.8 | 64.5 | 75.5 | 70.7 | 4.0 |
| Random Forest | 72.0 | 63.2 | 68.0 | 62.3 | 68.7 | 66.8 | 3.6 |
| Model | Acc. | AUC | Sens. | Spec. | Brier | ECE |
|---|---|---|---|---|---|---|
| EEGNet | 54.6% | 0.529 | 87.5% | 16.6% | 0.350 | 0.205 |
| Random Forest | 45.4% | 0.424 | 60.2% | 28.2% | 0.261 | 0.266 |
| RF + CORAL | 46.6% | 0.470 | 18.2% | 79.2% | 0.262 | 0.213 |
| Rank | Feature | Importance |
|---|---|---|
| 1 | Alpha power (Pz) | 0.089 |
| 2 | Alpha power (O1) | 0.076 |
| 3 | Alpha power (O2) | 0.072 |
| 4 | Theta power (Fz) | 0.068 |
| 5 | Alpha power (P3) | 0.062 |
| 6 | Theta power (Cz) | 0.058 |
| 7 | Delta power (F3) | 0.054 |
| 8 | Alpha power (P4) | 0.051 |
| 9 | Beta power (C3) | 0.047 |
| 10 | Delta power (F4) | 0.044 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




