Submitted:
12 January 2026
Posted:
13 January 2026
You are already at the latest version
Abstract
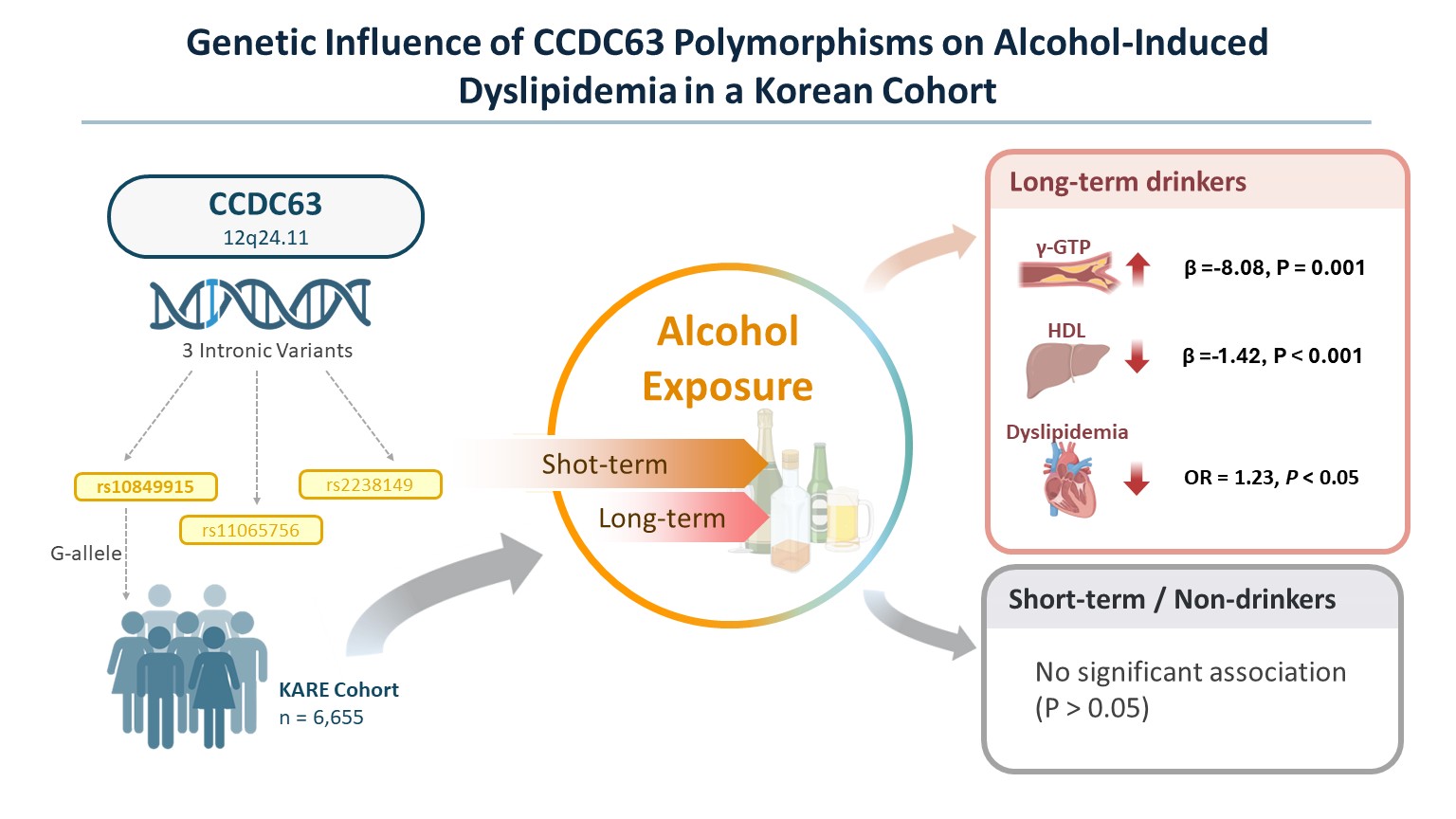
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Genetic Profiling
2.3. Data Analysis Framework
3. Results
3.1. Participant Characteristics and Lipid Profiles
3.2. Genomic Landscape of CCDC63 and Dyslipidemia Risk
3.3. Replication of rs10849915 Associations
3.4. Associations with Liver Enzymes and Lipid Markers
3.5. Distinct Genetic Effects by Alcohol Consumption
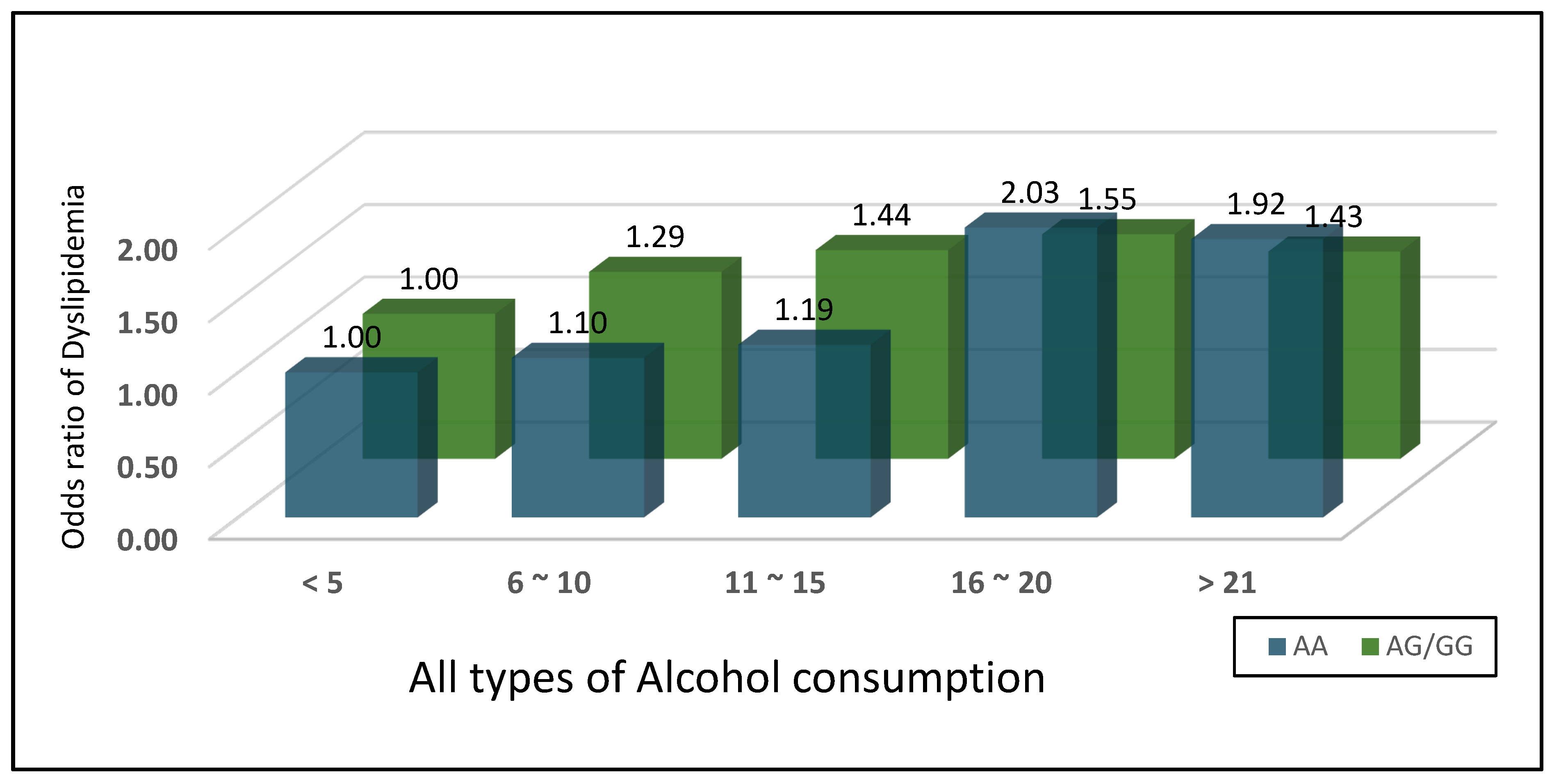
3.6. Alcohol Duration Effects Vary by Genotype
3.7. Sex-Specific Genetic Effects
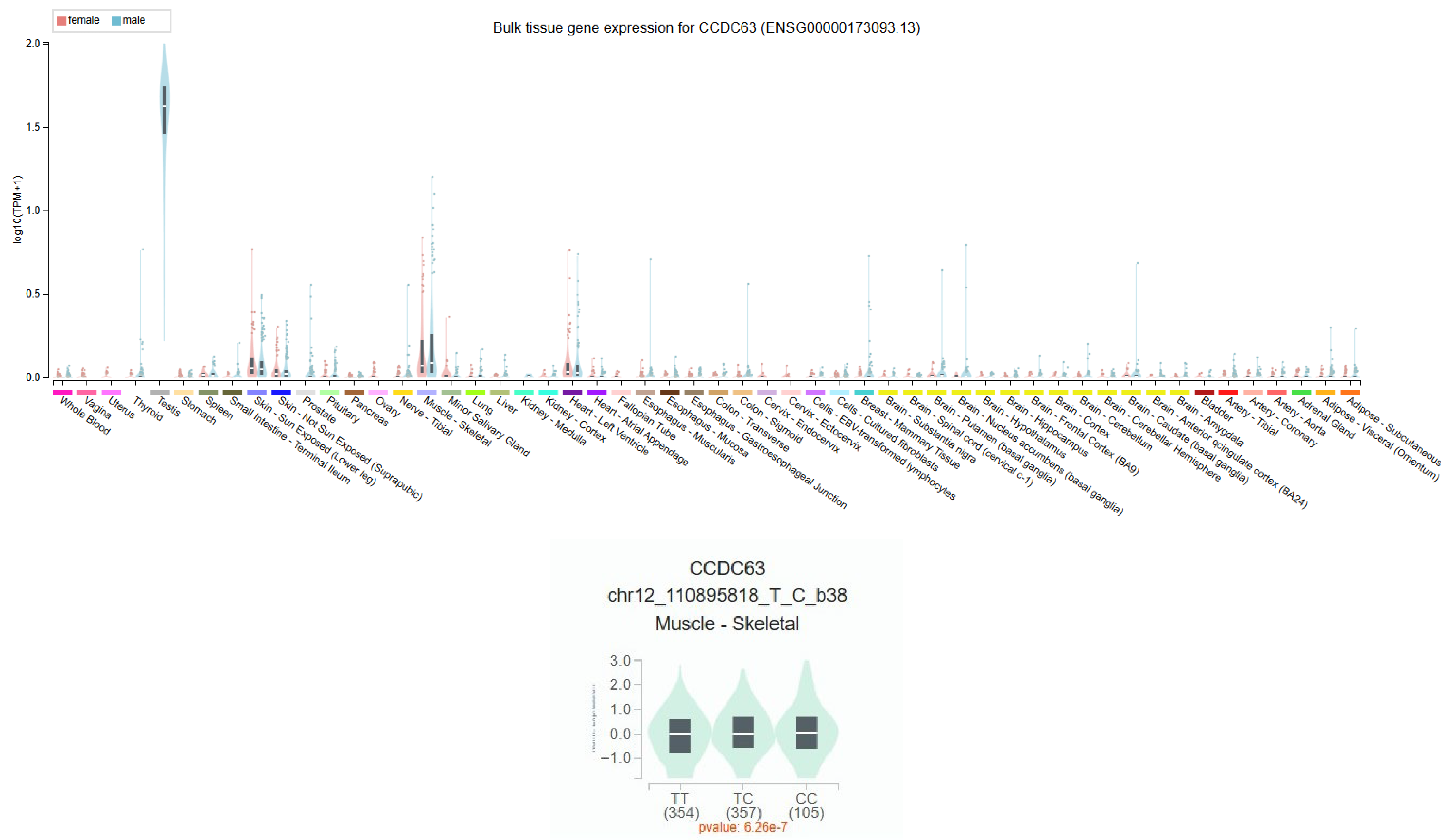
3.8. CCDC63 Tissue Expression Profile
4. Discussion
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lee, H.; Kim, S.; Son, Y.; Kim, S.; Kim, H.J.; Jo, H.; Park, J.; Lee, K.; Lee, H.; Kang, J.; et al. National trends in dyslipidemia prevalence, awareness, treatment, and control in South Korea from 2005 to 2022. Sci Rep 2025, 15, 16148. [Google Scholar] [CrossRef]
- Atherosclerosis, K. S. o. L. a. Dyslipidemia Fact Sheet in South Korea 2023. 2022. [Google Scholar]
- Klop, B.; Elte, J.W.; Cabezas, M.C. Dyslipidemia in obesity: mechanisms and potential targets. Nutrients 2013, 5, 1218–1240. [Google Scholar] [CrossRef]
- Baraona, E.; Lieber, C.S. Effects of ethanol on lipid metabolism. J Lipid Res 1979, 20, 289–315. [Google Scholar] [CrossRef]
- Brien, S.E.; Ronksley, P.E.; Turner, B.J.; Mukamal, K.J.; Ghali, W.A. Effect of alcohol consumption on biological markers associated with risk of coronary heart disease: systematic review and meta-analysis of interventional studies. BMJ 2011, 342, d636. [Google Scholar] [CrossRef] [PubMed]
- Steiner, J.L.; Lang, C.H. Alcohol, Adipose Tissue and Lipid Dysregulation. Biomolecules 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, I. Association between alcohol drinking and metabolic syndrome in Japanese male workers with diabetes mellitus. J Atheroscler Thromb 2011, 18, 684–692. [Google Scholar] [CrossRef]
- Park, H.S.; Oh, S.W.; Cho, S.I.; Choi, W.H.; Kim, Y.S. The metabolic syndrome and associated lifestyle factors among South Korean adults. Int J Epidemiol 2004, 33, 328–336. [Google Scholar] [CrossRef]
- Willer, C.J.; Schmidt, E.M.; Sengupta, S.; Peloso, G.M.; Gustafsson, S.; Kanoni, S.; Ganna, A.; Chen, J.; Buchkovich, M.L.; Mora, S.; et al. Discovery and refinement of loci associated with lipid levels. Nat Genet 2013, 45, 1274–1283. [Google Scholar] [PubMed]
- Liu, M.; Jiang, Y.; Wedow, R.; Li, Y.; Brazel, D.M.; Chen, F.; Datta, G.; Davila-Velderrain, J.; McGuire, D.; Tian, C.; et al. Association studies of up to 1.2 million individuals yield new insights into the genetic etiology of tobacco and alcohol use. Nat Genet 2019, 51, 237–244. [Google Scholar] [CrossRef]
- Clarke, T.K.; Adams, M.J.; Davies, G.; Howard, D.M.; Hall, L.S.; Padmanabhan, S.; Murray, A.D.; Smith, B.H.; Campbell, A.; Hayward, C.; et al. Genome-wide association study of alcohol consumption and genetic overlap with other health-related traits in UK Biobank (N=112 117). Mol Psychiatry 2017, 22, 1376–1384. [Google Scholar] [CrossRef] [PubMed]
- Jorgenson, E.; Thai, K.K.; Hoffmann, T.J.; Sakoda, L.C.; Kvale, M.N.; Banda, Y.; Schaefer, C.; Risch, N.; Mertens, J.; Weisner, C.; et al. Genetic contributors to variation in alcohol consumption vary by race/ethnicity in a large multi-ethnic genome-wide association study. Mol Psychiatry 2017, 22, 1359–1367. [Google Scholar] [CrossRef] [PubMed]
- Baik, I.; Cho, N.H.; Kim, S.H.; Han, B.G.; Shin, C. Genome-wide association studies identify genetic loci related to alcohol consumption in Korean men. Am J Clin Nutr 2011, 93, 809–816. [Google Scholar] [CrossRef]
- Park, B.L.; Kim, J.W.; Cheong, H.S.; Kim, L.H.; Lee, B.C.; Seo, C.H.; Kang, T.C.; Nam, Y.W.; Kim, G.B.; Shin, H.D.; et al. Extended genetic effects of ADH cluster genes on the risk of alcohol dependence: from GWAS to replication. Hum Genet 2013, 132, 657–668. [Google Scholar] [CrossRef]
- Schumann, G.; Coin, L.J.; Lourdusamy, A.; Charoen, P.; Berger, K.H.; Stacey, D.; Desrivieres, S.; Aliev, F.A.; Khan, A.A.; Amin, N.; et al. Genome-wide association and genetic functional studies identify autism susceptibility candidate 2 gene (AUTS2) in the regulation of alcohol consumption. Proc Natl Acad Sci USA 2011, 108, 7119–7124. [Google Scholar] [CrossRef]
- Kapoor, M.; Wang, J.C.; Wetherill, L.; Le, N.; Bertelsen, S.; Hinrichs, A.L.; Budde, J.; Agrawal, A.; Bucholz, K.; Dick, D.; et al. A meta-analysis of two genome-wide association studies to identify novel loci for maximum number of alcoholic drinks. Hum Genet 2013, 132, 1141–1151. [Google Scholar] [CrossRef]
- Consortium, G.T. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 2020, 369, 1318–1330. [Google Scholar] [CrossRef]
- Nica, A.C.; Montgomery, S.B.; Dimas, A.S.; Stranger, B.E.; Beazley, C.; Barroso, I.; Dermitzakis, E.T. Candidate causal regulatory effects by integration of expression QTLs with complex trait genetic associations. PLoS Genet 2010, 6, e1000895. [Google Scholar] [CrossRef]
- Zhou, H.; Kalayasiri, R.; Sun, Y.; Nunez, Y.Z.; Deng, H.W.; Chen, X.D.; Justice, A.C.; Kranzler, H.R.; Chang, S.; Lu, L.; et al. Genome-wide meta-analysis of alcohol use disorder in East Asians. Neuropsychopharmacology 2022, 47, 1791–1797. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Huang, J.; Mo, Z.; He, J.; Wang, L.; Yang, X.; Tan, A.; Chen, S.; Chen, J.; Gu, C.C.; et al. Genetic Susceptibility to Lipid Levels and Lipid Change Over Time and Risk of Incident Hyperlipidemia in Chinese Populations. Circ Cardiovasc Genet 2016, 9, 37–44. [Google Scholar] [CrossRef]
- Elisaus, P.; Williams, G.; Bourke, M.; Clough, G.; Harrison, A.; Verma, A. Factors associated with the prevalence of adolescent binge drinking in the urban areas of Greater Manchester. Eur J Public Health 2018, 28, 49–54. [Google Scholar] [CrossRef]
- Trostler, M.; Li, Y.; Plankey, M.W. Prevalence of binge drinking and associated co-factors among medical students in a U.S. Jesuit University. Am J Drug Alcohol Abuse 2014, 40, 336–341. [Google Scholar] [CrossRef] [PubMed]
- Relling, M.V.; Evans, W.E. Pharmacogenomics in the clinic. Nature 2015, 526, 343–350. [Google Scholar] [CrossRef] [PubMed]
- Cho, Y.S.; Go, M.J.; Kim, Y.J.; Heo, J.Y.; Oh, J.H.; Ban, H.J.; Yoon, D.; Lee, M.H.; Kim, D.J.; Park, M.; et al. A large-scale genome-wide association study of Asian populations uncovers genetic factors influencing eight quantitative traits. Nat Genet 2009, 41, 527–534. [Google Scholar] [CrossRef]
- Kranzler, H.R.; Zhou, H.; Kember, R.L.; Vickers Smith, R.; Justice, A.C.; Damrauer, S.; Tsao, P.S.; Klarin, D.; Baras, A.; Reid, J.; et al. Genome-wide association study of alcohol consumption and use disorder in 274,424 individuals from multiple populations. Nat Commun 2019, 10, 1499. [Google Scholar] [CrossRef]
- Gordon, D.J.; Probstfield, J.L.; Garrison, R.J.; Neaton, J.D.; Castelli, W.P.; Knoke, J.D.; Jacobs, D.R., Jr.; Bangdiwala, S.; Tyroler, H.A. High-density lipoprotein cholesterol and cardiovascular disease. Four prospective American studies. Circulation 1989, 79, 8–15. [Google Scholar] [CrossRef]
- Collins, F.S.; Varmus, H. A new initiative on precision medicine. N Engl J Med 2015, 372, 793–795. [Google Scholar] [CrossRef]
- Lieber, C.S. Metabolism of alcohol. Clin Liver Dis 2005, 9, 1–35. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Amamoto, K.; Tamaki, S.; Okamura, T.; Tsujita, Y.; Ueno, Y.; Kita, Y.; Kinoshita, M.; Ueshima, H. Genetic variation in aldehyde dehydrogenase 2 and the effect of alcohol consumption on cholesterol levels. Atherosclerosis 2002, 164, 171–177. [Google Scholar] [CrossRef]
- Wada, M.; Daimon, M.; Emi, M.; Iijima, H.; Sato, H.; Koyano, S.; Tajima, K.; Kawanami, T.; Kurita, K.; Hunt, S.C.; et al. Genetic association between aldehyde dehydrogenase 2 (ALDH2) variation and high-density lipoprotein cholesterol (HDL-C) among non-drinkers in two large population samples in Japan. J Atheroscler Thromb 2008, 15, 179–184. [Google Scholar] [CrossRef]
- Janssen, I.; Heymsfield, S.B.; Wang, Z.M.; Ross, R. Skeletal muscle mass and distribution in 468 men and women aged 18-88 yr. J Appl Physiol (1985) 2000, 89, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Baraona, E.; Abittan, C.S.; Dohmen, K.; Moretti, M.; Pozzato, G.; Chayes, Z.W.; Schaefer, C.; Lieber, C.S. Gender differences in pharmacokinetics of alcohol. Alcohol Clin Exp Res 2001, 25, 502–507. [Google Scholar] [CrossRef]
- Del Boca, F.K.; Darkes, J. The validity of self-reports of alcohol consumption: state of the science and challenges for research. Addiction 2003, 98, 1–12. [Google Scholar] [CrossRef]
- Ginsburg, G.S.; Phillips, K.A. Precision Medicine: From Science To Value. Health Aff (Millwood) 2018, 37, 694–701. [Google Scholar] [CrossRef]
- Lee, Y.J.; Lee, H.; Jang, H.B.; Yoo, M.G.; Im, S.; Koo, S.K.; Lee, H.J. The potential effects of HECTD4 variants on fasting glucose and triglyceride levels in relation to prevalence of type 2 diabetes based on alcohol intake. Arch Toxicol 2022, 96, 2487–2499. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Gable, A.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING database in 2023: protein-protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res 2023, 51, D638–D646. [Google Scholar] [CrossRef] [PubMed]
| Characteristics* | Quantitative trait analysis | Case-control analysis** | ||
|---|---|---|---|---|
| Normal | Dyslipidemia | P value*** | ||
| number of subjects | 6,655 | 2,328 | 4,327 | |
| Sex [male(%)/female(%)] | 3,177(47.7)/3,478(52.3) | 908(39.0)/1,420(61.0) | 2,269(52.4)/2,058(47.6) | <0.001 |
| Age (M years ± SD) | 52.10 ± 8.92 | 51.12 ± 9.08 | 52.68 ± 8.79 | 0.013 |
| Body mass index (BMI) (M kg/m2 ± SD) | 24.56 ± 3.14 | 23.31 ± 3.12 | 25.22 ± 2.94 | 0.003 |
| Total cholesterol (Tchl) (M ㎎/㎗ ± SD) | 186.43 ± 36.96 | 168.71 ± 19.12 | 195.97 ± 40.56 | <0.001 |
| Triglyceride (TG) (M ㎎/㎗ ± SD) | 154.99 ± 74.94 | 99.41 ± 24.44 | 184.90 ± 75.90 | <0.001 |
| High density lipoprotein cholesterol (HDL) (M ㎎/㎗ ± SD) | 43.32 ± 9.78 | 50.71 ± 7.94 | 39.35 ± 8.24 | 0.001 |
| Low density lipoprotein cholesterol (LDL) (M ㎎/㎗ ± SD) | 112.11 ± 33.36 | 98.12 ± 18.92 | 119.64 ± 36.84 | <0.001 |
| No. | SNP | A1 | A2 | Function | MAF | Dyslipidemia | ||
|---|---|---|---|---|---|---|---|---|
| (controls 2,328: cases 4,327) | ||||||||
| controls | cases | OR (95% CI) | Add P | |||||
| 1 | rs2283348 | T | C | intron | 0.3877 | 0.3768 | 0.95 (0.89~1.03) | 0.2162 |
| 2 | rs10219717 | T | C | intron | 0.3899 | 0.3798 | 0.96 (0.89~1.03) | 0.2520 |
| 3 | rs756838 | T | A | intron | 0.3901 | 0.3799 | 0.96 (0.89~1.03) | 0.2457 |
| 4 | rs2238149 | G | A | intron | 0.1554 | 0.1749 | 1.15 (1.05~1.27) | 0.0043 |
| 5 | rs10849915 | G | A | intron | 0.1645 | 0.1852 | 1.15 (1.05~1.27) | 0.0032 |
| 6 | rs11065756 | A | G | intron | 0.1634 | 0.1844 | 1.16 (1.05~1.27) | 0.0026 |
| Phenotype | Previous studies | Replication in Koreans | |||||
|---|---|---|---|---|---|---|---|
| EA | β ± SE, OR(95% CI) | P value*** | Research study | EA | β ± SE, OR(95% CI) | P value*** | |
| Alcohol consumption | G | -0.55 ± 0.05 | 1.0 X 10-23 | Inkyung Baik et al. | G | -0.31 ± 0.02 | 1.31X 10-60 |
| C | 0.04 | 4.0 X 10-28 | Eric Jorgenson et al. | NA | |||
| Traits | rs10849915 | rs11065756 | rs2238149 | |||
|---|---|---|---|---|---|---|
| Effect size | P -value | Effect size | P -value | Effect size | P -value | |
| (β±s.e) | (β±s.e) | (β±s.e) | ||||
| Alcohol-related liver enzyme levels | ||||||
| ALT (U/L) | -1.18 ± 0.51 | 0.02214 | -1.13 ± 0.52 | 0.02966 | -0.95 ± 0.53 | 0.07174 |
| AST (U/L) | -1.20 ± 0.37 | 0.00125 | -1.19 ± 0.37 | 0.00134 | -1.16 ± 0.38 | 0.00220 |
| γ-GTP (U/L) | -6.99 ± 1.29 | 5.68X10-8 | -7.24 ± 1.29 | 2.07X10-8 | -7.55 ± 1.31 | 8.67X10-8 |
| Serum lipid levels | ||||||
| HDL (㎎/㎗) | -0.87 ± 0.29 | 0.00244 | -0.91 ± 0.29 | 0.00148 | -0.79 ± 0.29 | 0.00704 |
| TG (㎎/㎗) | -3.49 ± 2.19 | 0.11080 | -3.68 ± 2.20 | 0.09409 | -3.93 ± 2.24 | 0.07923 |
| AA genotype (n=2,554) | AG/GG genotype (n=821) | |||
|---|---|---|---|---|
| OR (95% CI) | P value*** | OR (95% CI) | P value*** | |
| Sex | 0.41 (0.34~0.50) | <.001 | 1.17 (0.59~2.33) | 0.646 |
| Age | 1.01 (1.00~1.02) | 0.264 | 1.13 (1.08~1.18) | <.001 |
| BMI | 1.33 (1.29~1.37) | <.001 | 1.29 (1.15~1.44) | <.001 |
| γ-GTP | 1.00 (1.00~1.01) | <.001 | 1.00 (1.00~1.00) | <.001 |
| Alcohol consumption | ||||
| < 5 | 1(ref) | <.001 | 1(ref) | 0.05 |
| 6 ~ 10 | 1.10 (0.76~1.58) | 0.062 | 1.29 (0.70~2.40) | 0.04 |
| 11 ~ 15 | 1.19 (0.81~1.77) | 0.038 | 1.44 (0.75~2.79) | 0.03 |
| 16 ~ 20 | 2.03 (1.44~2.85) | <.001 | 1.55 (0.84~2.84) | 0.02 |
| > 21 | 1.92 (1.47~2.49) | <.001 | 1.43 (0.93~2.20) | 0.01 |
| Traits | Male | Female | ||
|---|---|---|---|---|
| Effect size | P -value | Effect size | P -value | |
| β ± SE, OR(95% CI) | β ± SE, OR(95% CI) | |||
| Dyslipidemia | 1.24 (1.07~1.43) | 0.00340 | 1.07 (0.95~1.22) | 0.27370 |
| Alcohol consumption | -0.40 ± 0.02 | 2.71 X10-74 | -0.23 ± 0.02 | 1.66 X10-22 |
| Clinical indicators of alcohol consumption | ||||
| ALT (U/L) | -1.45 ± 0.94 | 0.01223 | -0.97 ± 0.45 | 0.03138 |
| AST (U/L) | -1.98 ± 0.64 | 0.00215 | -0.51 ± 0.38 | 0.18140 |
| γ-GTP (U/L) | -13.02 ± 2.51 | 2.16 X10-7 | -1.60 ± 0.65 | 0.01461 |
| Dyslipidemia Clinical Indicators | ||||
| HDL (㎎/㎗) | -1.86 ± 0.33 | 0.00000 | 0.06 ± 0.45 | 0.90240 |
| TG (㎎/㎗) | -9.83 ± 3.57 | 0.00591 | 2.19 ± 2.55 | 0.39110 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




