Submitted:
06 January 2026
Posted:
07 January 2026
You are already at the latest version
Abstract
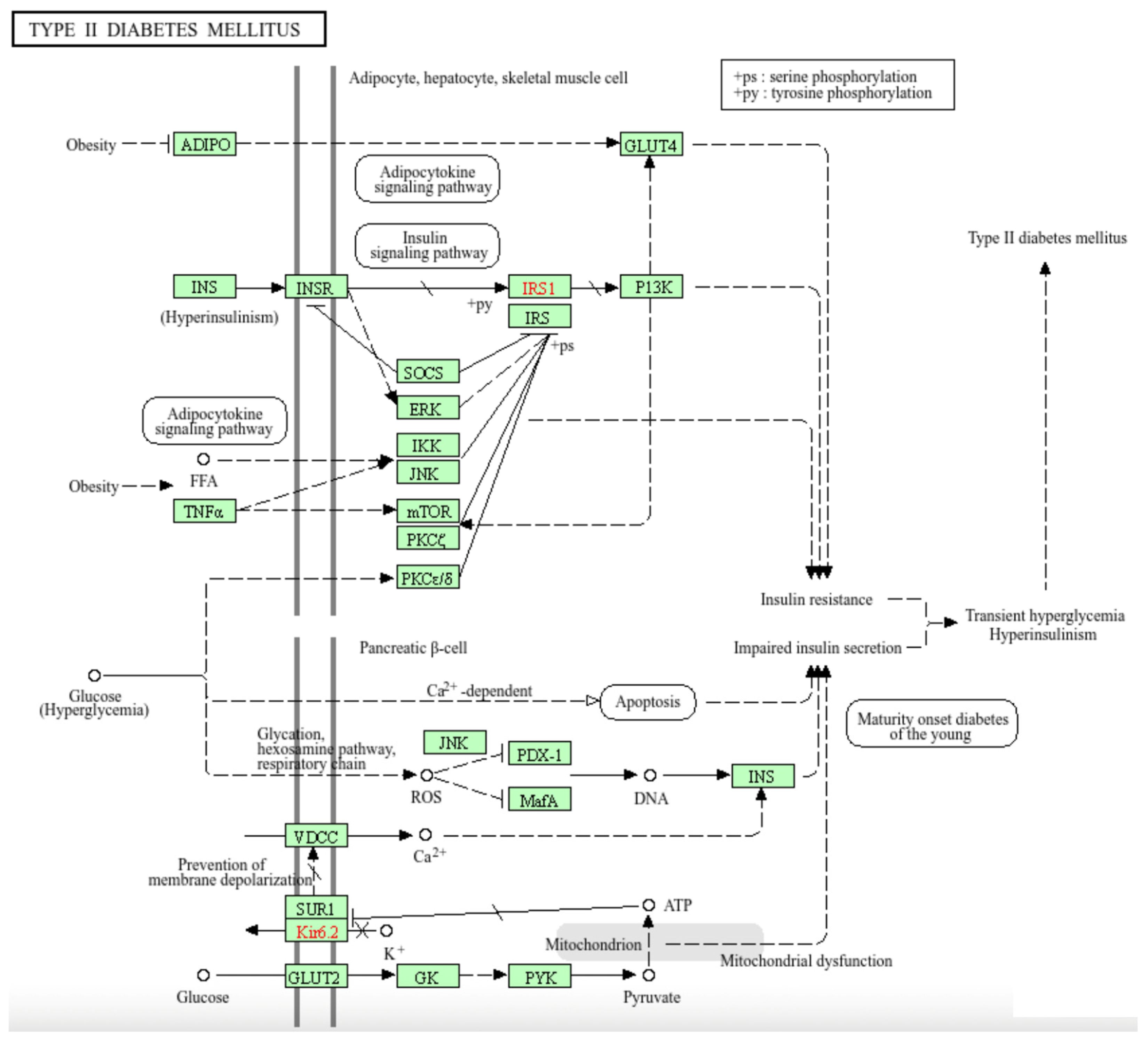
We devised a quantitative scoring function to assess the cumulative effects of nonsynonymous single nucleotide variants (SNVs) on protein-coding genes in patients with ovarian cancer (OvCa) and thyroid cancer (ThCa). The goal is to find novel candidate cancer-related genes for downstream bioinformatics analyses and wet-lab studies. With Genomic Data Commons as primary data resource, SNV information was extracted from whole-exome sequencing data from patients with these cancers. A cumulative variant scoring function, Q(G) was developed to sum up the deleterious effects of the individual SNVs on the gene G. While Q(G) can be computed using any popular functional effect analyzers such as FATHMM-XF, SIFT, PolyPhen, and CADD, we have also established an integrative scoring function iQ(G) that combines the deleterious assessments from different analyzers and demonstrated that iQ(G) is a more effective method for identifying likely cancer-related genes. Based on the iQ(G) rankings, the top three novel genes for OvCa are AHNAK2, UNC13A, and PCDHB4; and those for ThCA are PLEC, HECTD4, and CES1. Furthermore, the top 1% genes with highest iQ(G) scores for each cancer were submitted for KEGG pathway analysis. The results revealed that several genes of the CACNA1 family within the type II diabetes mellitus pathway are likely related to both OvCa and ThCa and suggested other molecular interactions that should be further studied in connection with OvCa prognosis and ThCa treatment.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Collecting, Organizing, and Extracting Information from Data Files
2.2. Functional Effect Analyzers
2.3. The Integrative Scoring Function
2.4. Bioinformatics Analyses
3. Results and Discussion
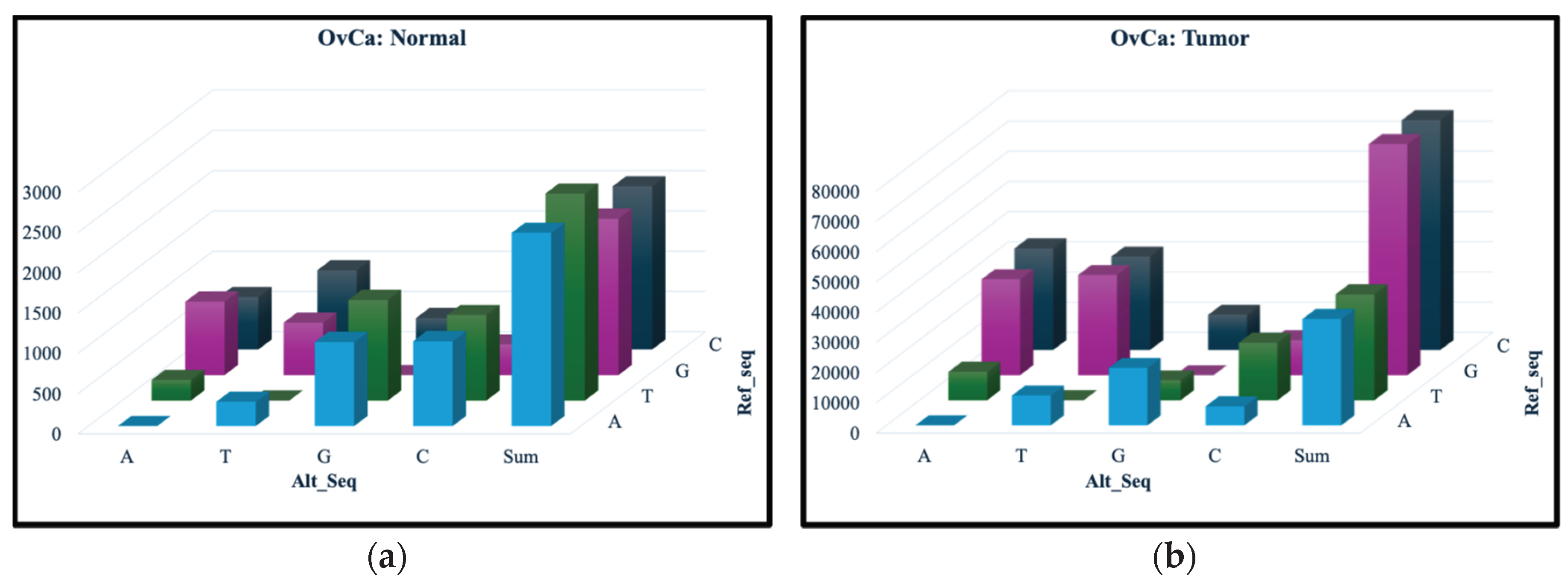
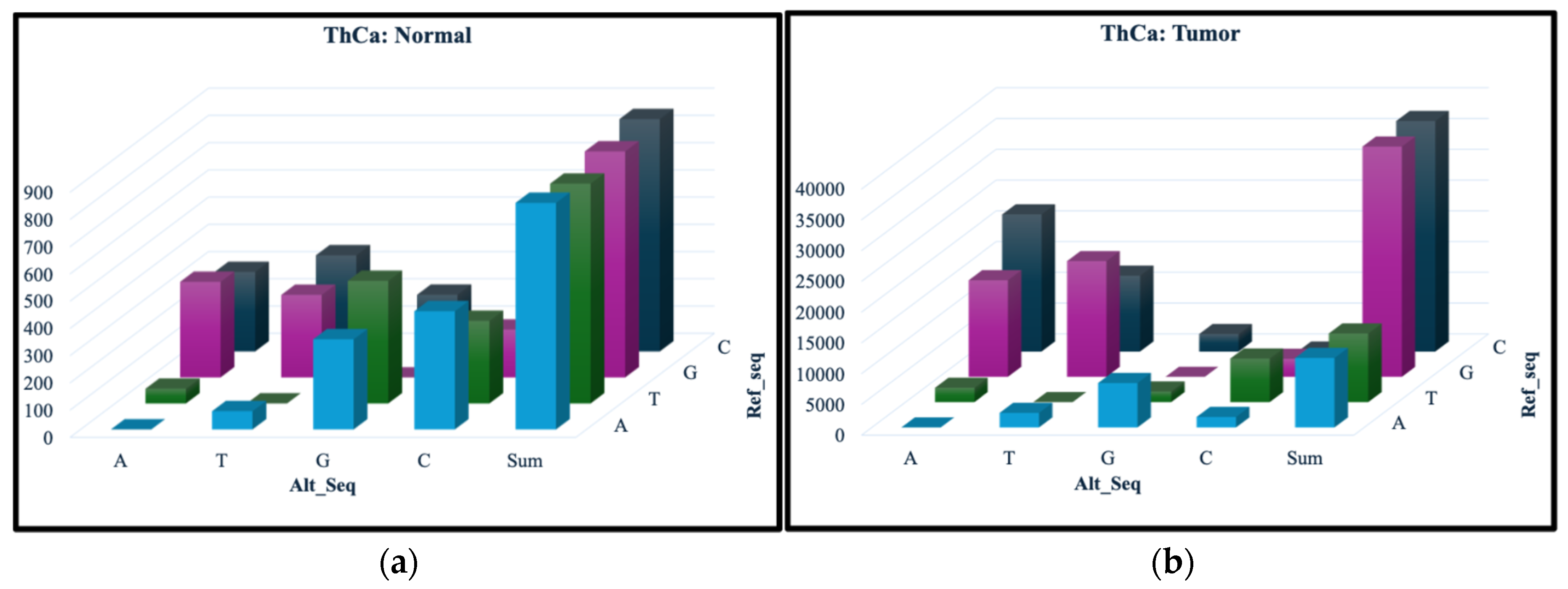
3.1. Variant Summary Statistics
3.2. Assessing the Performance of iQ(G)
3.3. Genes with Top Top iQ(G) Scores in OvCa and ThCa
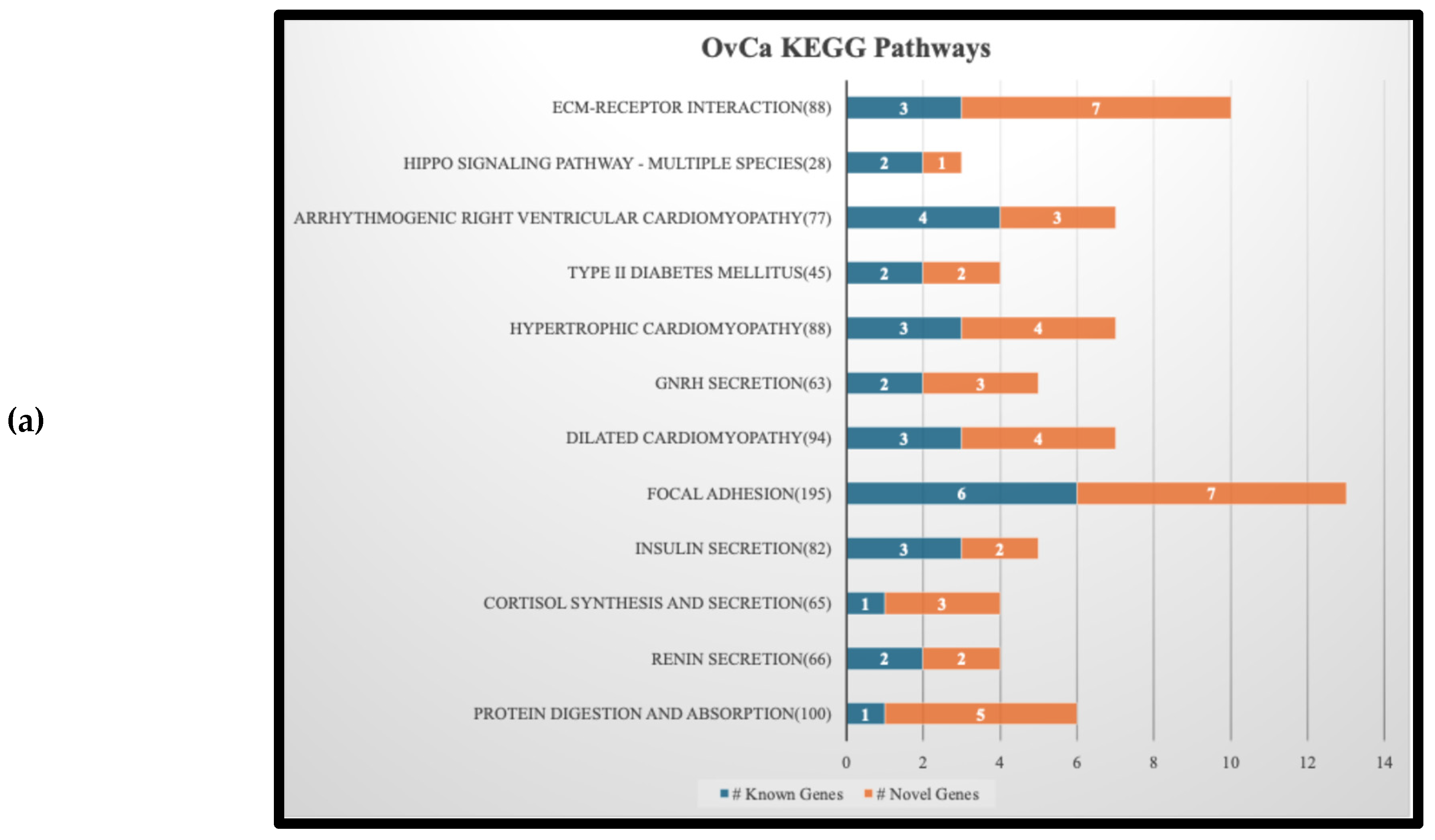
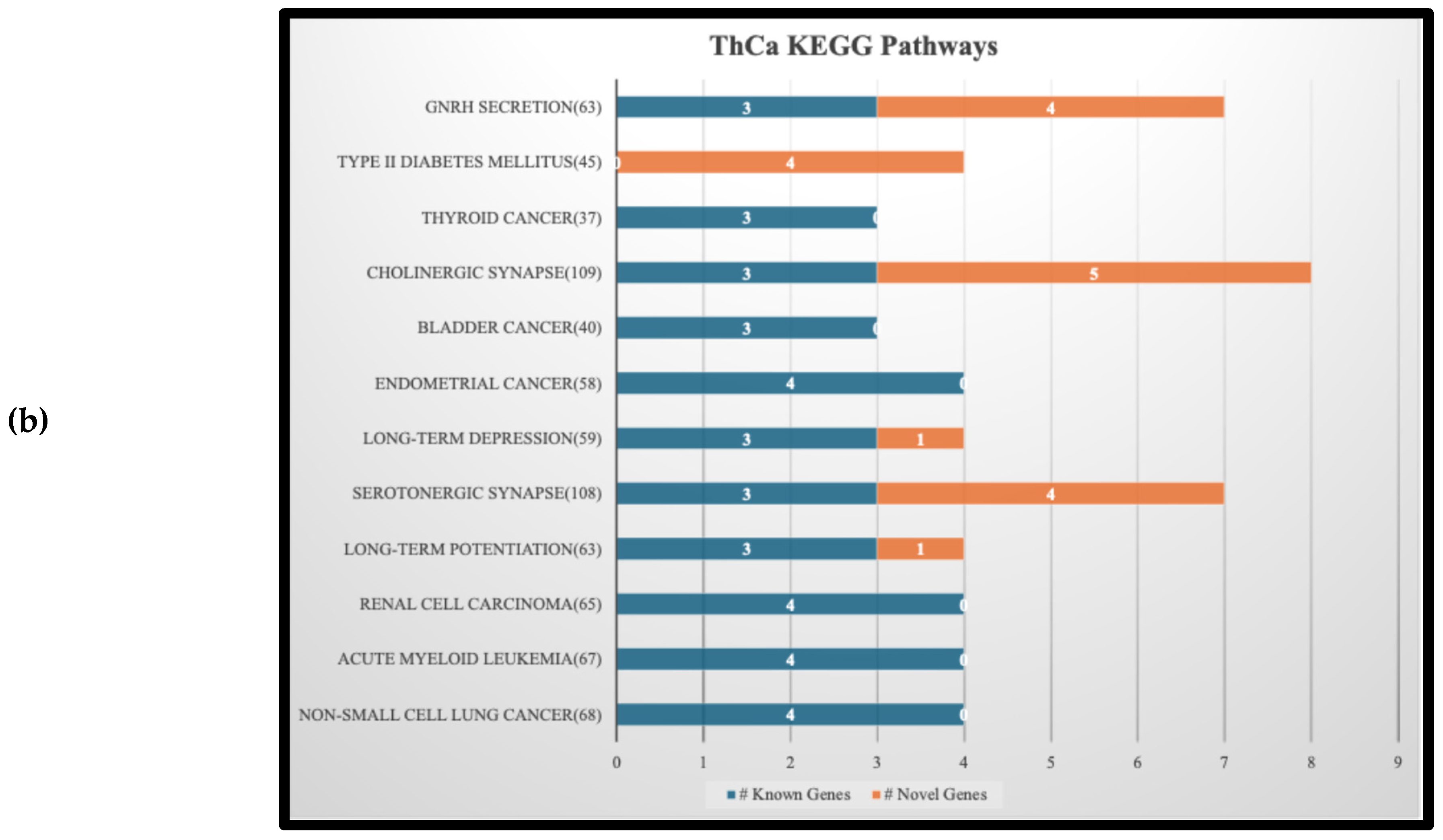
3.4. KEGG Pathway Analysis Results and Implications
4. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| OvCa | Ovarian cancer |
| ThCa | Thyroid cancer |
| SNV | Single nucleotide variant |
| VCF | Variant call format |
| FATHMM-XF | Functional analysis through hidden markov models – extended features |
| CADD | Combined annotation dependent depletion |
| SIFT | Sort intolerant from tolerant |
| PolyPhen | Polymorphism phenotyping |
| VEP | Variant effect predictor |
| KEGG | Kyoto encyclopedia of genes and genomes |
| STRING | Search tool for the retrieval of interacting genes/proteins |
| NGS | Next generation sequencing |
| GDC | Genomics data commons |
| CSQ | Consequence (data entry within VCF) |
| ROS | Reactive oxygen species |
| RAI | Radioactive Iodine |
| AML | Acute myeloid leukemia |
| DM2 | Type II diabetes mellitus |
| VDCC | Voltage-dependent calcium channel |
References
- What is ovarian cancer: Ovarian tumors and cysts. American Cancer Society. Available online: www.cancer.org/cancer/types/ovarian-cancer/about/what-is-ovarian-cancer.html (accessed on 6 March 2025).
- Ovarian cancer statistics: How common is ovarian cancer. American Cancer Society. Available online: https://www.cancer.org/cancer/types/ovarian-cancer/about/key-statistics.html (accessed on 9 May 2025).
- Modugno, F. Ovarian Cancer and High-Risk Women Symposium Presenters. Ovarian cancer and high-risk women—Implications for prevention, screening, and early detection. Gynecol. Oncol. 2003, 91, 15–31. [Google Scholar] [CrossRef] [PubMed]
- American Cancer Society What Is Thyroid Cancer? Available online: https://www.cancer.org/cancer/types/thyroid-cancer/about/what-is-thyroid-cancer.html (accessed on 29 January 2025).
- Key Statistics for Thyroid Cancer. Available online: https://www.cancer.org/cancer/types/thyroid-cancer/about/key-statistics.html (accessed on 29 January 2025).
- The causes of mutations - understanding evolution. Understanding Evolution. Available online: https://evolution.berkeley.edu/evolution-101/mechanisms-the-processes-of-evolution/the-causes-of-mutations/ (accessed on 9 September 2022).
- Ray, P.D.; Huang, B.-W.; Tsuji, Y. Reactive oxygen species (ROS) homeostasis and redox regulation in cellular signaling. Cell. Signal. 2012. Available online: https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3454471/#R1 (accessed on 9 September 2022).
- Ryu, J.Y.; Kim, H.; Lee, J.; et al. Human genes with a greater number of transcript variants tend to show biological features of housekeeping and essential genes. Mol. BioSyst. 2015, 11, 2798–2807. [Google Scholar] [CrossRef]
- PolyPhen-2 Score. Available online: https://ionreporter.thermofisher.com/ionreporter/help/GUID-57A60D00-0654-4F80-A8F9-F6B6A48D0278.html (accessed on 7 March 2024).
- Niroula, A.; Vihinen, M. How good are pathogenicity predictors in detecting benign variants? PLoS Comput. Biol. 2019, 15, e1006481. [Google Scholar] [CrossRef]
- Chen, J.; et al. Hunting for beneficial mutations: Conditioning on SIFT scores when estimating the distribution of fitness effect of new mutations. Genome Biol. Evol. 2022, 14. [Google Scholar] [CrossRef]
- Combined Annotation Dependent Depletion. CADD. Available online: https://cadd.gs.washington.edu/ (accessed on 7 March 2024).
- Rogers, M.F.; Shihab, H.A.; Mort, M.; Cooper, D.N.; Gaunt, T.R.; Campbell, C. FATHMM-XF: Enhanced accuracy in the prediction of pathogenic sequence variants via an extended feature set. Bioinformatics, 2024; submitted. [Google Scholar]
- Shihab, H.A.; Rogers, M.F.; Gough, J.; Mort, M.; Cooper, D.N.; Day, I.N.M.; Gaunt, T.R.; Campbell, C. An integrative approach to predicting the functional consequences of non-coding and coding sequence variation. Bioinformatics 2015, 31, 1536–1543. [Google Scholar] [CrossRef]
- Shihab, H.A.; Gough, J.; Cooper, D.N.; Stenson, P.D.; Barker, G.L.A.; Edwards, K.J.; Day, I.N.M.; Gaunt, T.R. Predicting the functional, molecular and phenotypic consequences of amino acid substitutions using hidden Markov models. Hum. Mutat. 2013, 34, 57–65. [Google Scholar] [CrossRef]
- Yoon, B.-J. Hidden Markov models and their applications in biological sequence analysis. Curr. Genomics 2009, 10, 402–415. Available online: www.ncbi.nlm.nih.gov/pmc/articles/PMC2766791/ (accessed on 6 March 2024). [CrossRef] [PubMed]
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Nastou, K.; Mehryary, F.; Hachilif, R.; Annika, G.L.; Fang, T.; Doncheva, N.T.; Pyysalo, S.; et al. The STRING database in 2023: Protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023, 51, D638–D646. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021, 49, D605–D612. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The STRING database in 2017: Quality-controlled protein–protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Franceschini, A.; Wyder, S.; Forslund, K.; Heller, D.; Huerta-Cepas, J.; Simonovic, M.; Roth, A.; Santos, A.; Tsafou, K.P.; et al. STRING v10: Protein–protein interaction networks, integrated over the tree of life. Nucleic Acids Res. 2015, 43, D447–D452. [Google Scholar] [CrossRef]
- Franceschini, A.; Lin, J.; von Mering, C.; Jensen, L.J. SVD-phy: Improved prediction of protein functional associations through singular value decomposition of phylogenetic profiles. Bioinformatics 2015, 31, btv696. [Google Scholar] [CrossRef] [PubMed]
- Franceschini, A.; Szklarczyk, D.; Frankild, S.; Kuhn, M.; Simonovic, M.; Roth, A.; Lin, J.; Minguez, P.; Bork, P.; von Mering, C.; et al. STRING v9.1: Protein–protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013, 41, D808–D815. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Franceschini, A.; Kuhn, M.; Simonovic, M.; Roth, A.; Minguez, P.; Doerks, T.; Stark, M.; Muller, J.; Bork, P.; et al. The STRING database in 2011: Functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res. 2011, 39, D561–D568. [Google Scholar] [CrossRef]
- Jensen, L.J.; Kuhn, M.; Stark, M.; Chaffron, S.; Creevey, C.; Muller, J.; Doerks, T.; Julien, P.; Roth, A.; Simonovic, M.; et al. STRING 8—A global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 2009, 37, D412–D416. [Google Scholar] [CrossRef] [PubMed]
- von Mering, C.; Jensen, L.J.; Kuhn, M.; Chaffron, S.; Doerks, T.; Krueger, B.; Snel, B.; Bork, P. STRING 7—Recent developments in the integration and prediction of protein interactions. Nucleic Acids Res. 2007, 35, D358–D362. [Google Scholar] [CrossRef]
- von Mering, C.; Jensen, L.J.; Snel, B.; Hooper, S.D.; Krupp, M.; Foglierini, M.; Jouffre, N.; Huynen, M.A.; Bork, P. STRING: Known and predicted protein–protein associations, integrated and transferred across organisms. Nucleic Acids Res. 2005, 33, D433–D437. [Google Scholar] [CrossRef]
- von Mering, C.; Huynen, M.; Jaeggi, D.; Schmidt, S.; Bork, P.; Snel, B. STRING: A database of predicted functional associations between proteins. Nucleic Acids Res. 2003, 31, 258–261. [Google Scholar] [CrossRef]
- Snel, B.; Lehmann, G.; Bork, P.; Huynen, M.A. STRING: A web-server to retrieve and display the repeatedly occurring neighbourhood of a gene. Nucleic Acids Res. 2000, 28, 3442–3444. [Google Scholar] [CrossRef]
- GDC. Available online: https://portal.gdc.cancer.gov/ (accessed on 30 October 2022).
- The Cancer Genome Atlas Program (TCGA). Available online: https://www.cancer.gov/ccg/research/genome-sequencing/tcga (accessed on 30 October 2022).
- McLaren, W.; Gil, L.; Hunt, S.E.; Riat, H.S.; Ritchie, G.R.; Thormann, A.; Flicek, P.; Cunningham, F. The Ensembl Variant Effect Predictor. Genome Biol. 2016, 17, 122. [Google Scholar] [CrossRef]
- Variation, Ensembl. Pathogenicity Predictions. Available online: https://useast.ensembl.org/info/genome/variation/prediction/protein_function.html (accessed on 6 March 2024).
- Kinsella, R.J.; Kähäri, A.; Haider, S.; Zamora, J.; Proctor, G.; Spudich, G.; Almeida-King, J.; Staines, D.; Derwent, P.; Kerhornou, A.; Kersey, P.; Flicek, P. Ensembl BioMarts: A hub for data retrieval across taxonomic space. Database (Oxford) 2011, bar030. [Google Scholar] [CrossRef]
- Oscanoa, J.; Sivapalan, L.; Gadaleta, E.; Dayem Ullah, A.Z.; Lemoine, N.R.; Chelala, C. SNPnexus: A web server for functional annotation of human genome sequence variation (2020 update). Nucleic Acids Res. 2020, 48, W185–W192. [Google Scholar] [CrossRef]
- Dayem Ullah, A.Z.; Oscanoa, J.; Wang, J.; Nagano, A.; Lemoine, N.; Chelala, C. SNPnexus: Assessing the functional relevance of genetic variation to facilitate the promise of precision medicine. Nucleic Acids Res. 2018, 46, W109–W113. [Google Scholar] [CrossRef] [PubMed]
- Dayem Ullah, A.Z.; Lemoine, N.R.; Chelala, C. A practical guide for the functional annotation of genetic variations using SNPnexus. Brief. Bioinform. 2013, 14, 437–447. [Google Scholar] [CrossRef]
- Dayem Ullah, A.Z.; Lemoine, N.R.; Chelala, C. SNPnexus: A web server for functional annotation of novel and publicly known genetic variants (2012 update). Nucleic Acids Res. 2012, 40, W65–W70. [Google Scholar] [CrossRef] [PubMed]
- Chelala, C.; Khan, A.; Lemoine, N.R. SNPnexus: A web database for functional annotation of newly discovered and public domain Single Nucleotide Polymorphisms. Bioinformatics 2009, 25, 655–661. [Google Scholar] [CrossRef]
- Seal, R.L.; Braschi, B.; Gray, K.; Jones, T.E.M.; Tweedie, S.; Haim-Vilmovsky, L.; Bruford, E.A. Genenames.org: The HGNC resources in 2023. Nucleic Acids Res. 2023. [Google Scholar] [CrossRef]
- HGNC Database. Available online: https://www.genenames.org (accessed on 7 March 2024).
- Stelzer, G.; Rosen, R.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; Iny Stein, T.; Nudel, R.; Lieder, I.; Mazor, Y.; et al. The GeneCards Suite: From gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinform. 2016, 54, 1.30.1–1.30.33. [Google Scholar] [CrossRef] [PubMed]
- Safran, M.; Rosen, N.; Twik, M.; BarShir, R.; Iny Stein, T.; Dahary, D.; Fishilevich, S.; Lancet, D. The GeneCards Suite. In Practical Guide to Life Science Databases, 1st ed.; Springer: Cham, Switzerland, 2022; pp. 27–56. [Google Scholar]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Matsuura, Y.; Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 2025, 53, D672–D677. [Google Scholar] [CrossRef]
- Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 2019, 28, 1947–1951. [Google Scholar] [CrossRef]
- Kanehisa, M.; Goto, S. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res. 2000, 28, 27–30. [Google Scholar] [CrossRef]
- Rappaport, N.; et al. Rational confederation of genes and diseases: NGS interpretation via GeneCards, MalaCards and VarElect. Biomed. Eng. Online 2017, 16 Suppl. 1, 72. [Google Scholar] [CrossRef]
- Rappaport, N.; et al. MalaCards: An integrated compendium for diseases and their annotation. Database (Oxford) 2013, bat018. [Google Scholar] [CrossRef] [PubMed]
- Rappaport, N.; et al. MalaCards: A comprehensive automatically-mined database of human diseases. Curr. Protoc. Bioinformatics 2014, 47, 1.24.1–1.24.19. [Google Scholar] [CrossRef]
- Rappaport, N.; et al. MalaCards: An amalgamated human disease compendium with diverse clinical and genetic annotation and structured search. Nucleic Acids Res. 2017, 45, D877–D887. [Google Scholar] [CrossRef]
- Safran, M.; et al. MalaCards - the integrated human malady compendium. In Proceedings of the ISMB 2012, Long Beach, CA, USA, 15–17 July 2012. [Google Scholar]
- Phung, H.T.; Nguyen, A.Q.; Van Nguyen, T.; Van Nguyen, T.; Nguyen, L.T.; Nguyen, K.T.; Thi Pham, H.D. Ovary metastasis from lung cancer mimicking primary ovarian cancer: A rare case report. Ann. Med. Surg. 2022, 80, 104207. [Google Scholar] [CrossRef]
- Madu, N.M.; Skinner, C.; Oyibo, S.O. Cure rates after a single dose of radioactive iodine to treat hyperthyroidism: The fixed-dose regimen. Cureus 2022, 14, e28316. [Google Scholar] [CrossRef] [PubMed]
- Chang, X.; Dong, Y. CACNA1C Is a Prognostic Predictor for Patients with Ovarian Cancer. Journal of Ovarian Research 2021, 14. [Google Scholar] [CrossRef] [PubMed]
- Jiang, A.; Jiang, Y.; Meng, Y.; Ma, M.; Qin, Z.; Chen, Y.; Fan, Y.; Li, P. M6A Modification Mediates CACNA1A Stability to Drive the Progression of Ovarian Cancer by Inhibiting Ferroptosis. Journal of Ovarian Research 2025. [Google Scholar] [CrossRef]
- Roh, E.; Noh, E.; Hwang, S.Y.; Kim, J.A.; Song, E.; Park, M.; Choi, K.M.; Baik, S.H.; Cho, G.J.; Yoo, H.J. Increased Risk of Type 2 Diabetes in Patients with Thyroid Cancer after Thyroidectomy: A Nationwide Cohort Study. J. Clin. Endocrinol. Metab. Available online. 2021, 107, e1047–e1056. [Google Scholar] [CrossRef] [PubMed]
- Oberman, B.; Khaku, A.; Camacho, F.; Goldenberg, D. Relationship between Obesity, Diabetes and the Risk of Thyroid Cancer. Am. J. Otolaryngol. Available online. 2015, 36, 535–541. [Google Scholar] [CrossRef] [PubMed]
- Xiang, J.; Su, R.; Wu, S.; Zhou, L. Construction of a Prognostic Signature for Serous Ovarian Cancer Based on Lactate Metabolism-Related Genes. Frontiers in Oncology 2022, 12. [Google Scholar] [CrossRef]
- Molenaar, R.J.; Sidana, Surbhi; Radivoyevitch, T.; Advani, A.S.; Gerds, A.T.; Carraway, H.E.; Angelini, D.E.; Kalaycio; Nazha, M.; Aziz; Adelstein, D.J.; et al. Risk of Hematologic Malignancies after Radioiodine Treatment of Well-Differentiated Thyroid Cancer. Journal of Clinical Oncology 2018, 36, 1831–1839. [Google Scholar] [CrossRef] [PubMed]
| FATHMM-XF | CADD | SIFT | PolyPhen | |
|---|---|---|---|---|
| Score Range | 0-1 | 0-99 | 0-1 | 0-1 |
| Deleterious Cutoff |
| OvCa | ThCa | |
|---|---|---|
| VCF Files | 486 | 504 |
| Unique Patients | 462 | 496 |
| Known Cancer-Related Genes | 928 | 493 |
| Unique SNVs | 222,830 | 97,373 |
| Normal only | 78 | 7 |
| Tumor only | 213,894 | 94,051 |
| Common | 8,858 | 3,315 |
| Unique Genes | 13,229 | 7,507 |
| (a) | ||||
| FATHMM | CADD | SIFT | PolyPhen | |
| OvCa | 0.6901 | 0.6234 | 0.655 | 0.6584 |
| ThCa | 0.8516 | 0.7559 | 0.8242 | 0.8335 |
| (b) | ||||
| iQ(G) | Mean (±SE) for randomly sampled human gene sets | |||
| OvCa | 0.6018 | 0.6038 (± 0.0003) | ||
| ThCa | 0.7527 | 0.8311 (± 0.0005) | ||
| (a) OvCa | (b) ThCa | |||||||
|---|---|---|---|---|---|---|---|---|
| Gene (G) | iQ(G) | #Tr | #Var | Gene (G) | iQ(G) | #Tr | #Var | |
| TP53 | 24.16 | 23 | 135 | BRAF | 30.36 | 4 | 3 | |
| TTN | 3.16 | 11 | 138 | NRAS | 4.09 | 1 | 2 | |
| CSMD3 | 2.77 | 4 | 36 | HRAS | 1.62 | 5 | 3 | |
| HMCN1 | 1.82 | 1 | 28 | TTN | 1.36 | 11 | 37 | |
| HERC2 | 1.74 | 1 | 24 | PLEC | 1.29 | 11 | 25 | |
| AHNAK2 | 1.67 | 1 | 34 | CLIP2 | 1.22 | 2 | 7 | |
| USH2A | 1.52 | 3 | 31 | CCAR1 | 1.00 | 7 | 1 | |
| UNC13A | 1.42 | 4 | 17 | HECTD4 | 0.85 | 2 | 13 | |
| CACNA1C | 1.37 | 23 | 19 | CES1 | 0.79 | 3 | 2 | |
| CSMD1 | 1.36 | 7 | 20 | MUC4 | 0.78 | 4 | 22 | |
| RYR2 | 1.35 | 5 | 34 | EPPK1 | 0.76 | 2 | 14 | |
| PCDHB4 | 1.35 | 1 | 15 | EVPL | 0.75 | 2 | 8 | |
| DNAH3 | 1.32 | 1 | 21 | RPS18 | 0.72 | 2 | 1 | |
| MYH4 | 1.29 | 1 | 17 | CACNA1C | 0.70 | 22 | 9 | |
| DNAH10 | 1.26 | 5 | 26 | TENM2 | 0.68 | 3 | 9 | |
| Gene Name | Rank | Annotations | |
|---|---|---|---|
| OvCa | AHNAK2 | 6 | Located on chromosome 14, this gene plays a role in calcium signaling; associated with non-small cell lung cancer. |
| UNC13A | 8 | Located on chromosome 19, this gene is a part of the gene family that plays a role in neurotransmitter release at synapses; identified with amyotrophic lateral sclerosis. | |
| PCDHB4 | 12 | Member of the protocadherin beta gene cluster on chromosome 5; highly suspected function includes specific cell-cell neural connections; linked to mutations on this gene are Seckel Syndrome and Autism. | |
| ThCa | PLEC | 5 | Located on chromosome 8, interlinks different elements on the cytoskeleton; mutation related diseases include muscular dystrophy and epidermolysis bullosa. |
| HECTD4 | 8 | Located on chromosome 12, this gene is involved in glucose metabolic process and homeostasis, associated to neurodevelopment disorders and seizures. | |
| CES1 | 9 | Responsible for hydrolysis or transesterification of xenobiotics (foreign synthetic chemicals); located on chromosome 16, alterations on this gene may affect drug metabolism. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




