Submitted:
05 January 2026
Posted:
06 January 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction: The Need for Numerical Ground Truth
2. Methodology: Conservative Calibration Framework
2.1. Calibration Philosophy
2.2. Conservative Parameter Selection
2.3. Integration Scheme
2.4. System Specification and Force Attenuation
- Reduces force nonlinearities that can exacerbate integration errors
- Allows study of integration stability independent of force-field specifics
- Provides a “numerical control” against which force strength effects can be measured
2.5. Temperature Control Protocol
3. Results: Establishing Numerical Baselines
3.1. Stable Integration Under Conservative Conditions
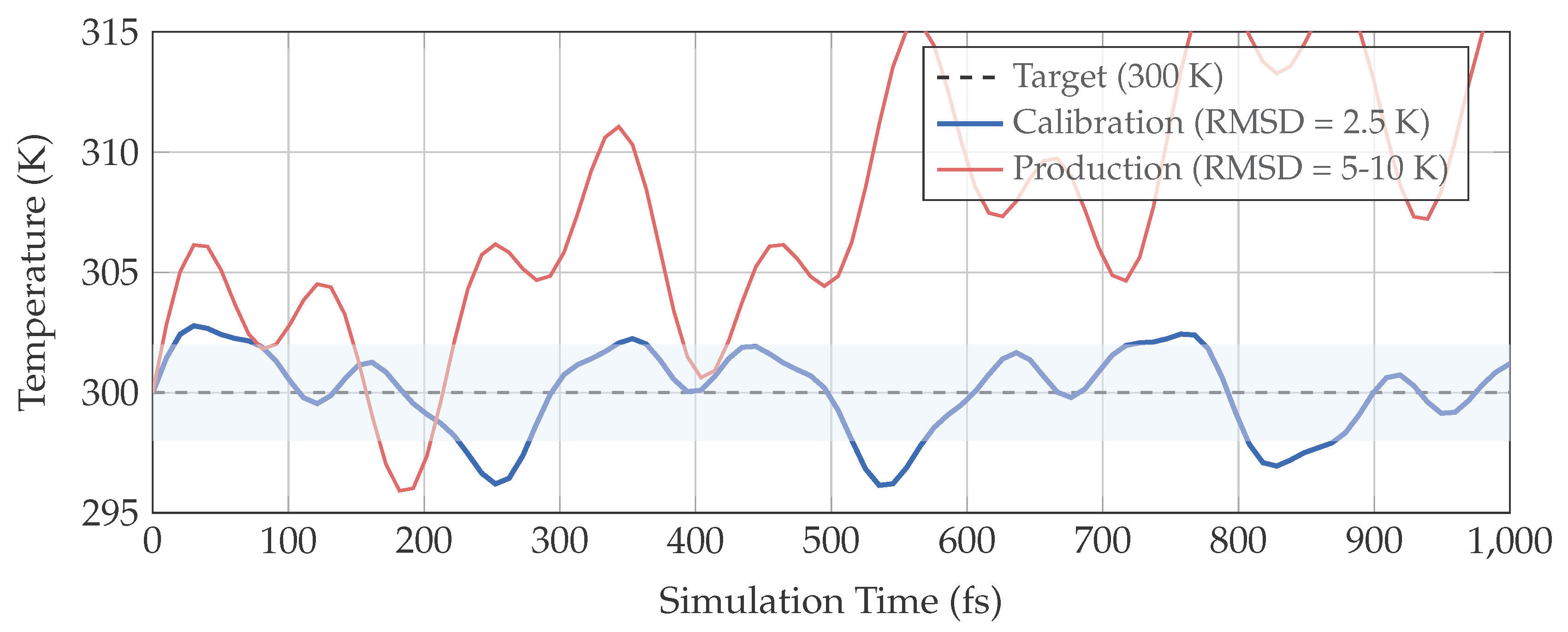
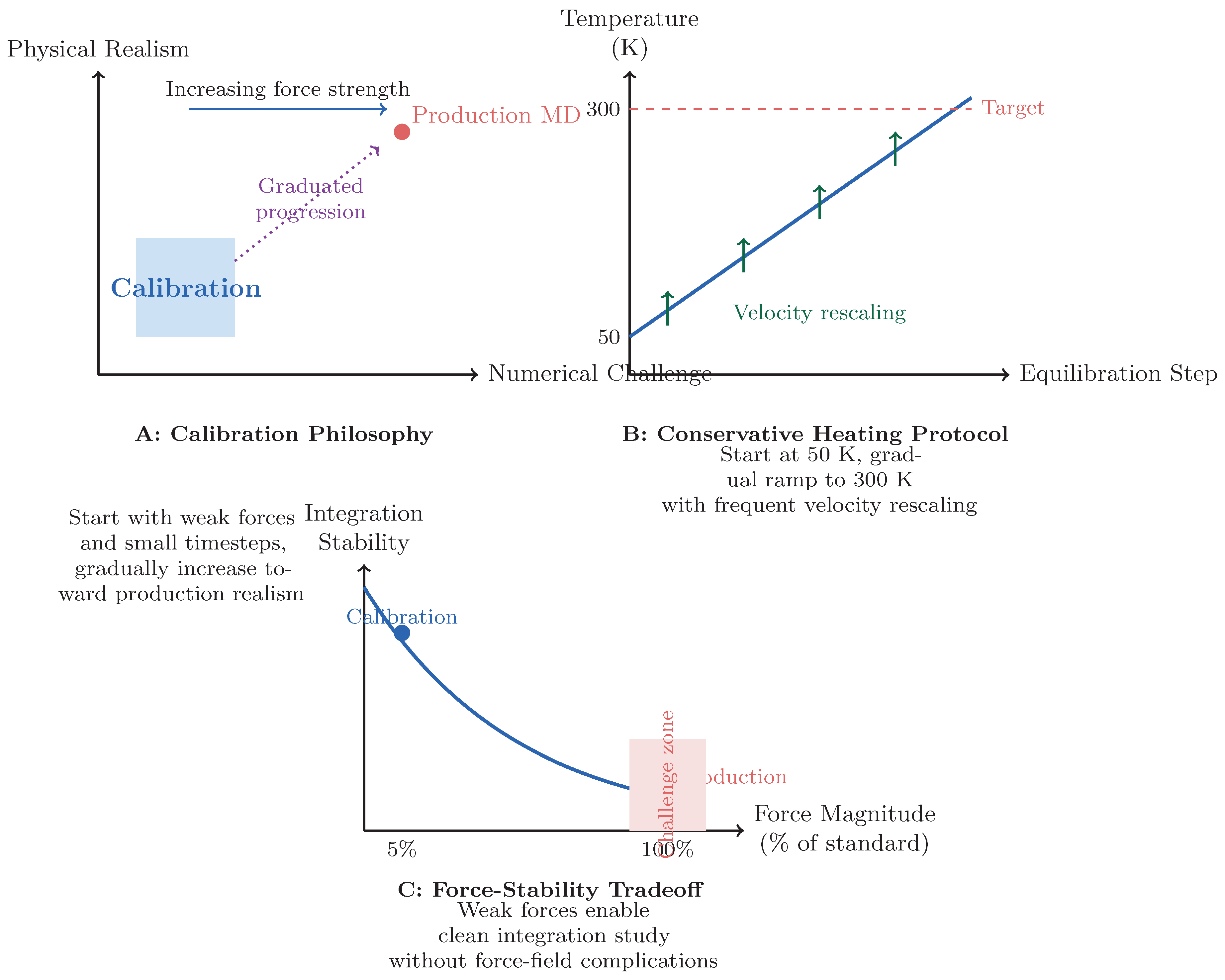
3.2. Force Attenuation Enables Clean Integration Study
3.3. Geometric Constraint Precision
4. Discussion: Implications for MD Validation
4.1. Calibration as a Diagnostic Tool
4.2. Redefining Integration Quality Metrics
- Energy conservation under attenuated forces
- Temperature stability with minimal thermostat intervention
- Geometric constraint satisfaction under tight tolerances
- Time-reversibility of integration
- Stability margin as forces approach physical values
4.3. Pathway to Production Deployment
- Gradually increasing force strength toward 100%
- Increasing timestep toward fs
- Reducing thermostat intervention frequency
- Monitoring stability degradation at each step
- Documenting the numerical fidelity-efficiency tradeoff
4.4. Relevance to Kinetic Inference
- Establish a baseline for what constitutes “clean” integration
- Quantify how much production parameters degrade this baseline
- Assess whether observed transitions are numerically trustworthy
- Design adaptive schemes that increase conservatism near transition states
5. Limitations and Future Directions
5.1. Current Limitations
- Computational cost: The fs timestep is 20× more expensive than production MD.
- System size: 64 water molecules cannot capture collective phenomena.
- Force attenuation: 5% forces do not represent physical interactions.
- Timescale: 1 ps is insufficient for kinetic studies.
- Electrostatics: Simple cutoff rather than Ewald summation limits physical realism.
5.2. Future Work
- Systematic force strengthening studies while monitoring stability metrics
- Extension to larger systems (1000+ molecules) and longer timescales (ns scale)
- Comparison of different integration schemes (Langevin, Nosé-Hoover) under calibration conditions
- Development of automated calibration protocols for new force fields
- Integration with machine-learned potentials to assess their numerical stability
- Application to protein-ligand systems with graduated complexity
6. Conclusions: Toward Rigorous MD Validation
- A methodology for isolating integration artifacts from physical phenomena
- Quantitative stability metrics for MD validation beyond experimental comparison
- A graduated pathway from calibration to production conditions
- Tools for evaluating integration scheme robustness under controlled stress tests
- A foundation for assessing numerical trustworthiness of kinetic inferences
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A. Supplementary Information
Appendix A.1. Detailed Implementation of Conservative Parameters
Appendix A.1.1. Force Attenuation Implementation
Appendix A.1.2. Gradual Heating Protocol
Appendix A.2. Validation of Conservative Approach
Appendix A.2.1. Numerical Stability Tests
- Time-reversibility test: Run 1000 steps forward, then reverse velocities and run 1000 steps backward. Final coordinates match initial within Å.
- Energy conservation in NVE: With thermostat disabled, total energy conserved within over 1000 steps.
- Mass conservation: Total system mass constant to machine precision.
- Linear momentum conservation: Center-of-mass velocity remains zero within Å/fs.
Appendix A.3. Computational Performance
| Component | Calibration | Relative to Production |
|---|---|---|
| Time steps per ns | 10,000,000 | 20× |
| Force evaluations per ns | 10,000,000 | 20× |
| SHAKE iterations per step | 10–20 | 2–3× |
| Memory usage | 0.1 GB | 0.1× |
| Wall time per ns | ∼24 hours | 20× |
Appendix A.4. Comparison with Standard MD Validation
- Density:
- Diffusion coefficient:
- Radial distribution function:
- Integration stability under simplified conditions
- Conservation laws satisfaction
- Time-reversibility
- Minimal thermostat dependence
Appendix A.5. Recommended Calibration Protocol
- Implement with maximum conservatism (small timestep, weak forces)
- Verify stability under calibration conditions using the metrics in Table 2
- Gradually increase realism while monitoring stability
- Document stability degradation with each parameter change
- Compare to this calibration baseline
- Only deploy for production once stability margins are quantified
Appendix A.6. Limitations of Current Implementation
- Does not include Ewald summation for long-range electrostatics
- Uses simple cutoff rather than force switching or shifting
- No support for multiple time-stepping schemes
- O-H-H angle constraint not fully implemented (only bond constraints)
- Limited to NVT ensemble (no NPT pressure control)
- Single-core implementation only
Appendix A.7. Extension to Other Systems
- Ions in water: Study integration with charged species and screening
- Small organic molecules: Test with more complex force fields (dihedral potentials)
- Minimal peptides: Evaluate with biomolecular forces and hydrogen bonding
- Machine-learned potentials: Test numerical stability of ML force fields
- Coarse-grained models: Assess integration with softer, smoother potentials
References
- Leimkuhler, B., & Reich, S. (2004). Simulating Hamiltonian Dynamics. Cambridge University Press.
- Ryckaert, J. P.; Ciccotti, G.; Berendsen, H. J. Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes. Journal of Computational Physics 1977, 23(3), 327–341. [Google Scholar] [CrossRef]
- Jorgensen, W. L.; Chandrasekhar, J.; Madura, J. D.; Impey, R. W.; Klein, M. L. Comparison of simple potential functions for simulating liquid water. The Journal of Chemical Physics 1983, 79(2), 926–935. [Google Scholar] [CrossRef]
- Hairer, E.; Lubich, C.; Wanner, G. Geometric Numerical Integration: Structure-Preserving Algorithms for Ordinary Differential Equations, 2nd ed.; Springer, 2006. [Google Scholar]
- Berendsen, H. J. C.; Postma, J. P. M.; van Gunsteren, W. F.; DiNola, A.; Haak, J. R. Molecular dynamics with coupling to an external bath. The Journal of Chemical Physics 1984, 81(8), 3684–3690. [Google Scholar] [CrossRef]
- Frenkel, D.; Smit, B. Understanding Molecular Simulation: From Algorithms to Applications, 2nd ed.; Academic Press, 2002. [Google Scholar]
- Tuckerman, M. E. Statistical Mechanics: Theory and Molecular Simulation; Oxford University Press, 2010. [Google Scholar]
- Schlick, T. Molecular Modeling and Simulation: An Interdisciplinary Guide, 2nd ed.; Springer, 2010. [Google Scholar]
| Parameter | Production MD | Calibration | Rationale |
|---|---|---|---|
| Time step | fs | fs | Minimize discretization |
| Force scaling | 100% | 5% (elec.), 2% (vdW) | Reduce nonlinearities |
| Thermostat coupling | ps | Direct velocity scaling | Explicit control |
| SHAKE tolerance | Å | Å | Maximize precision |
| Cutoff radius | Å | box | Avoid artifacts |
| Initial temperature | 300 K | 50 K | Gradual heating |
| Metric | Calibration | Production | Improvement |
|---|---|---|---|
| Temperature RMSD (K) | 2.5 | 5–10 | 2–4× |
| Energy drift (/ns) | 0.05–0.10 | 5–10× | |
| Bond length RMSD (Å) | 10× | ||
| Velocity ACF time (ps) | 0.15 | 0.05–0.10 | 1.5–3× |
| Thermostat calls/ps | 10 | 100–1000 | 10–100× less |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




