Submitted:
25 December 2025
Posted:
25 December 2025
You are already at the latest version
Abstract
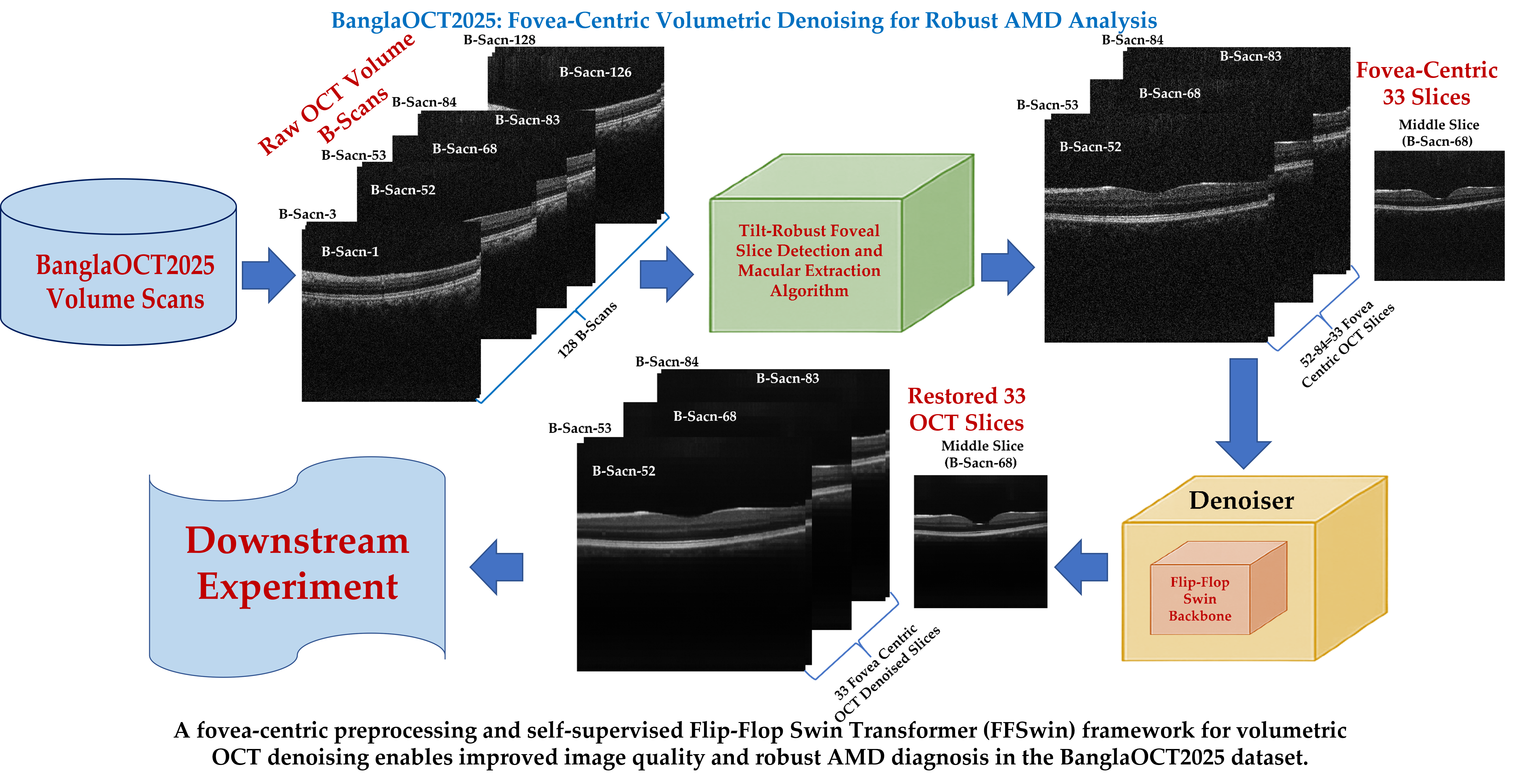
Keywords:
1. Introduction
2. Materials and Methods
2.1. Dataset: BanglaOCT2025
2.1.1. Dataset Acquisition
2.1.2. Ground Truth Labeling
- Initial Classification: Two specialists (from SBMCH and KMCH) independently labeled 573 patients (857 individual scans) into categories including Dry AMD, Wet AMD, and other retinal conditions.
- Conflict Resolution: While classifications were generally consistent, disagreements arose in specific early-stage cases. These contentious cases were reviewed by a third specialist from NIOH.
2.2. Constraint-Based Fovea-Centric Volume Extraction
- Segment retinal tissue in the central region of each slice.
- Compute a column-wise centroid (one per A-scan) to detect the shallowest point.
- Define a slice-level pit metric using the minimum centroid height.
- Apply a clinical penalty to enforce the expected anatomical slice range.
- Extract the fovea and its 16 neighboring slices on each side.
2.2.1. Tilt-Robust Foveal Slice Detection and Macular Extraction Algorithm
- Parameters:
- (Adjacent slices each side)
- (Total slices per volume)
- , (preferred foveal range)
- (Penalty for out-of-range slices)
- Procedure:
- Initialize metric dictionary
-
For each sliceto
- Load image
- Extract central region (35% width):
- Apply Gaussian blur:
- Compute Otsu threshold
- Segment tissue:
- Compute column centroids
- If or :
- Else:
- Find foveal slice:
- Calculate range: ,
- If : adjust range leftward
- If : adjust range rightward
- Copy slices through to output folder
2.2.2. Design Rationale and Robustness Analysis
2.2.3. Parameter Summary
| Parameters | Value and Rationale |
|---|---|
| ) | 128 (standard NIDEK protocol) |
| ) | 1.5mm |
| Search range | 5 slices) |
| ) | 200 (empirically validated on the BanglaOCT2025 dataset) |
| Central width | 35% of image (focus on anatomically relevant region) |
| Gaussian kernel | , balances noise reduction and edge preservation) |
| Threshold method | Otsu’s adaptive threshold (robust to brightness variation) |
2.2.4. Validation and Error Handling
- Fallback mechanism: If no valid images are found, defaults to slice 64 (midpoint of preferred range)
- Boundary checking: Ensures extracted sub-volume stays within 1–128 range
- Empty folder detection: Skips folders without valid OCT images
- Numerical stability: prevents division by zero in centroid calculations
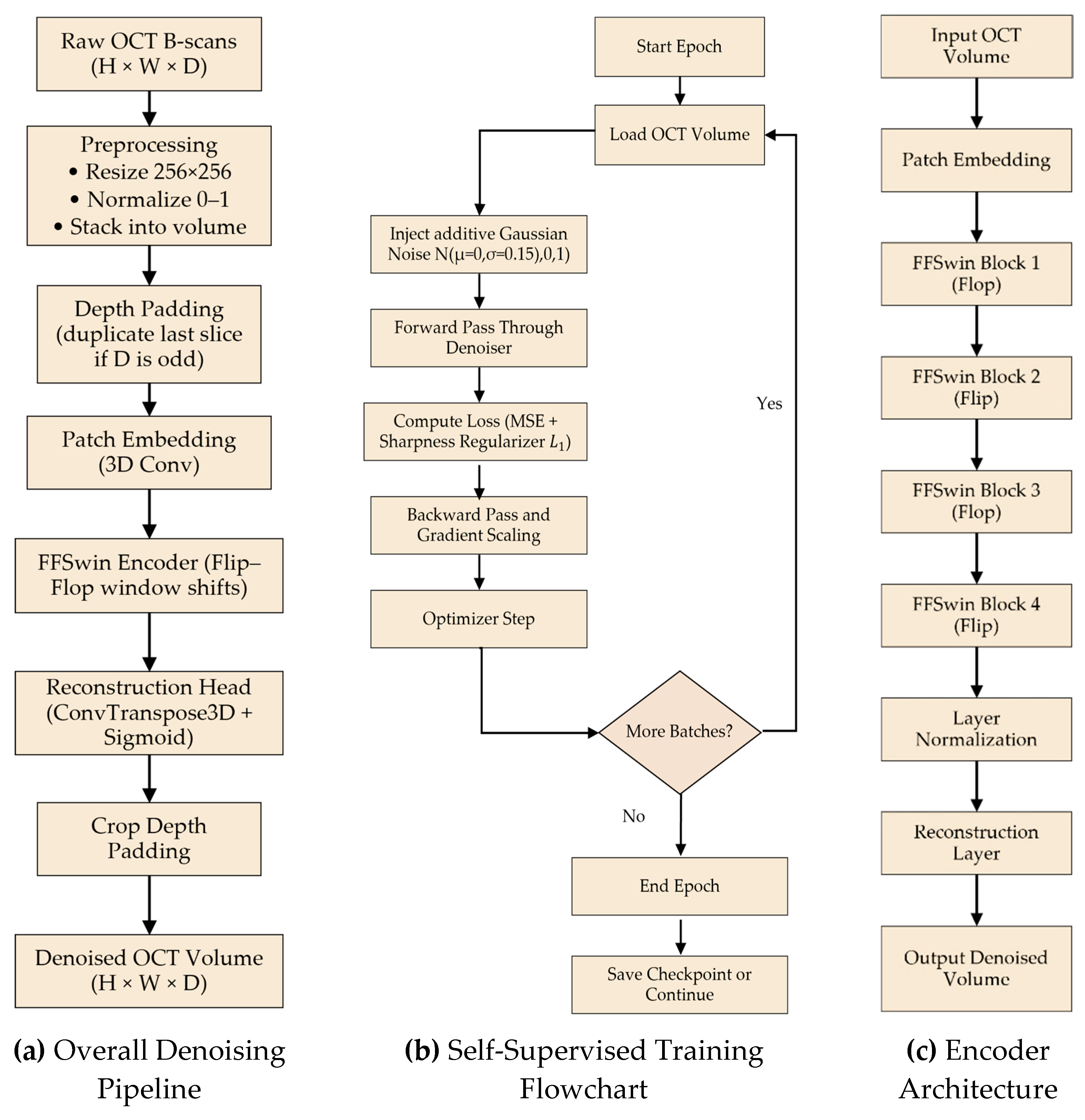
2.3. Self-Supervised Volumetric Denoising Framework Using FFSwin Backbone
2.3.1. Theoretical Premise: 3D Spatio-Temporal Consistency
- Anatomical Continuity: Retinal layers (e.g., RPE, ILM) and pathologies (e.g., Drusen) are physically continuous structures. If a feature exists at coordinates in slice , it likely exists near in slice and
- Noise Independence: Speckle noise is an interference pattern that is stochastic. A noise granule at in slice has no correlation with the pixel at in slice
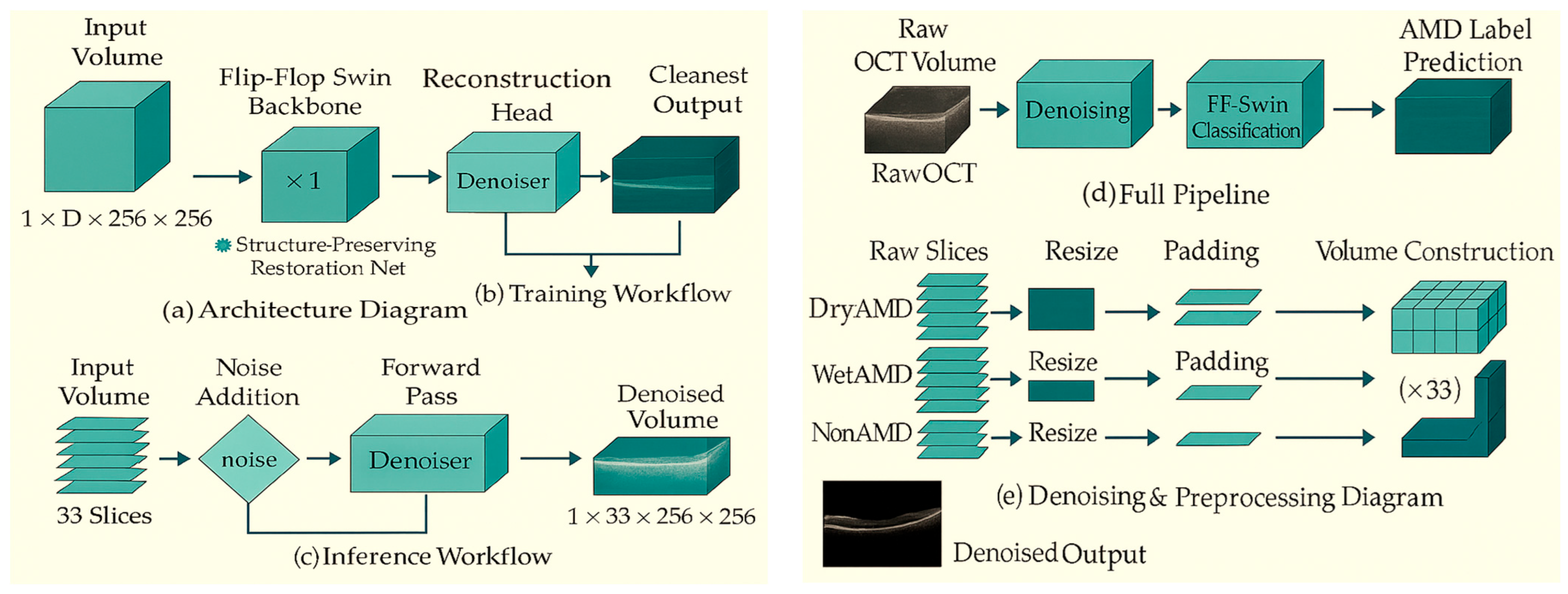
2.3.2. Network Backbone: The "Flip-Flop" Abstraction
- Intra-Slice Attention (Flop Mode): The model attends to local patches within the 2D plane (). This allows the network to learn texture and edge definitions within a single B-scan.
- Inter-Slice Attention (Flip Mode): The attention window is shifted along the -axis (Depth). This forces the model to aggregate information from co-located patches in adjacent slices ).
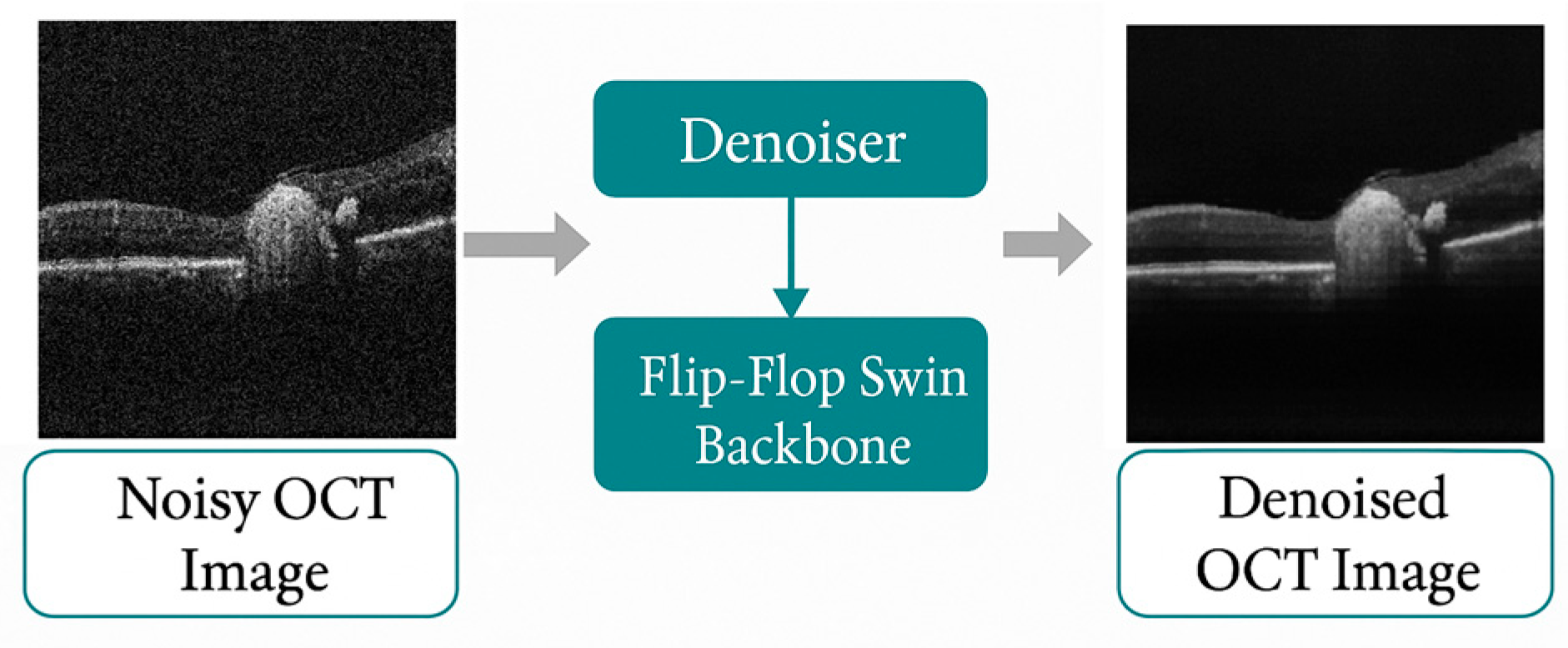
2.3.3. Self-Supervised Training Strategy
2.3.4. Volumetric Patch Embedding and Context Modeling
2.3.5. Reconstruction Loss Function
2.3.6. High-Level Architecture (Decoder-Free Design)
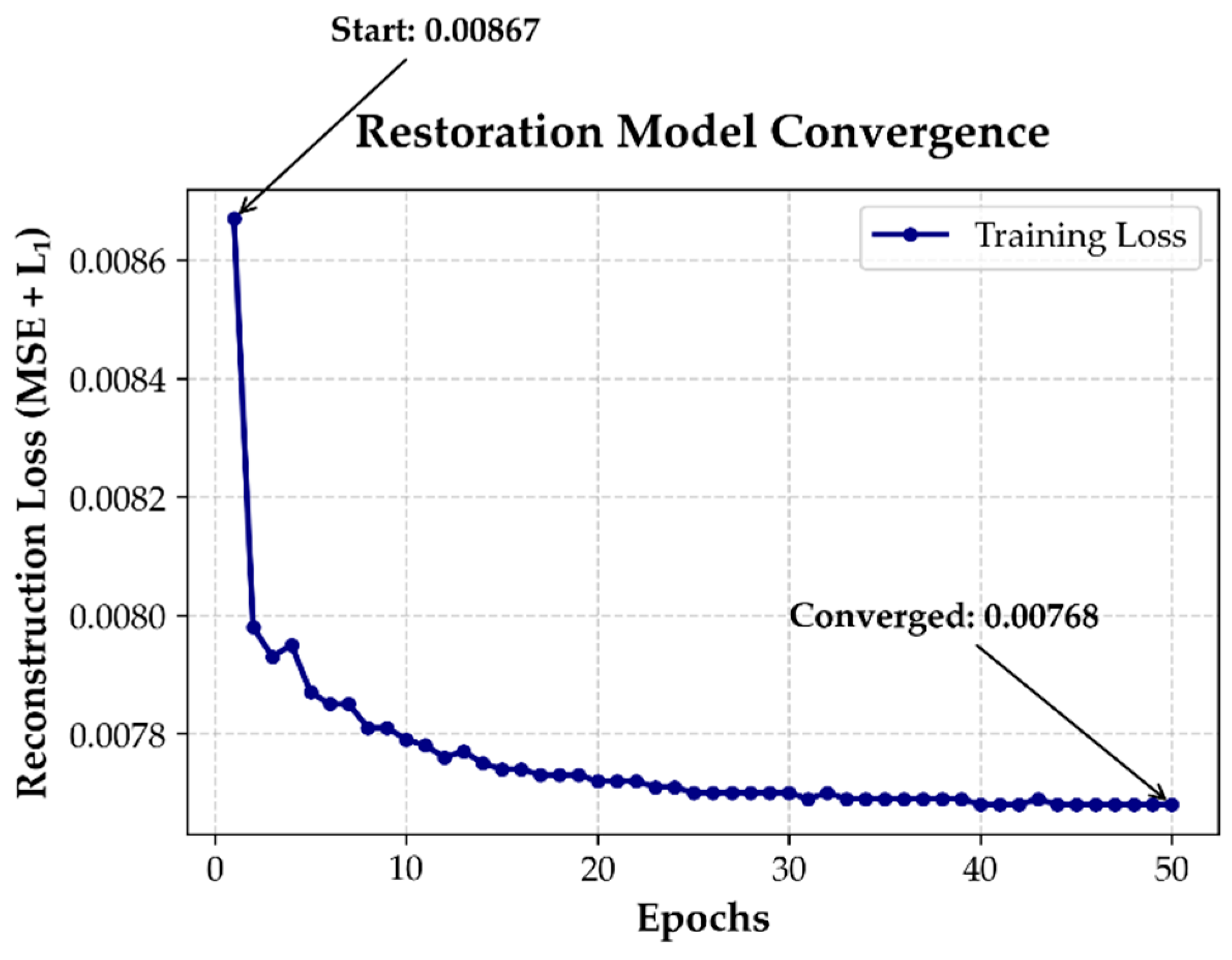
2.3.7. Training Protocol and Convergence Analysis
2.3.8. Inference Pipeline
3. Results
3.1. BanglaOCT2025 Characteristics and Clinical Composition
3.2. Evaluation of Constraint-Based Fovea-Centric Volume Extraction
3.2.1. Robustness of Automated Foveal Slice Detection
3.2.2. Standardization of Macular Sub-Volumes
3.2.3. Role of Fovea-Centric Extraction in Downstream Analysis
3.3. Self-Supervised Volumetric Restoration Framework Using FFSwin Backbone
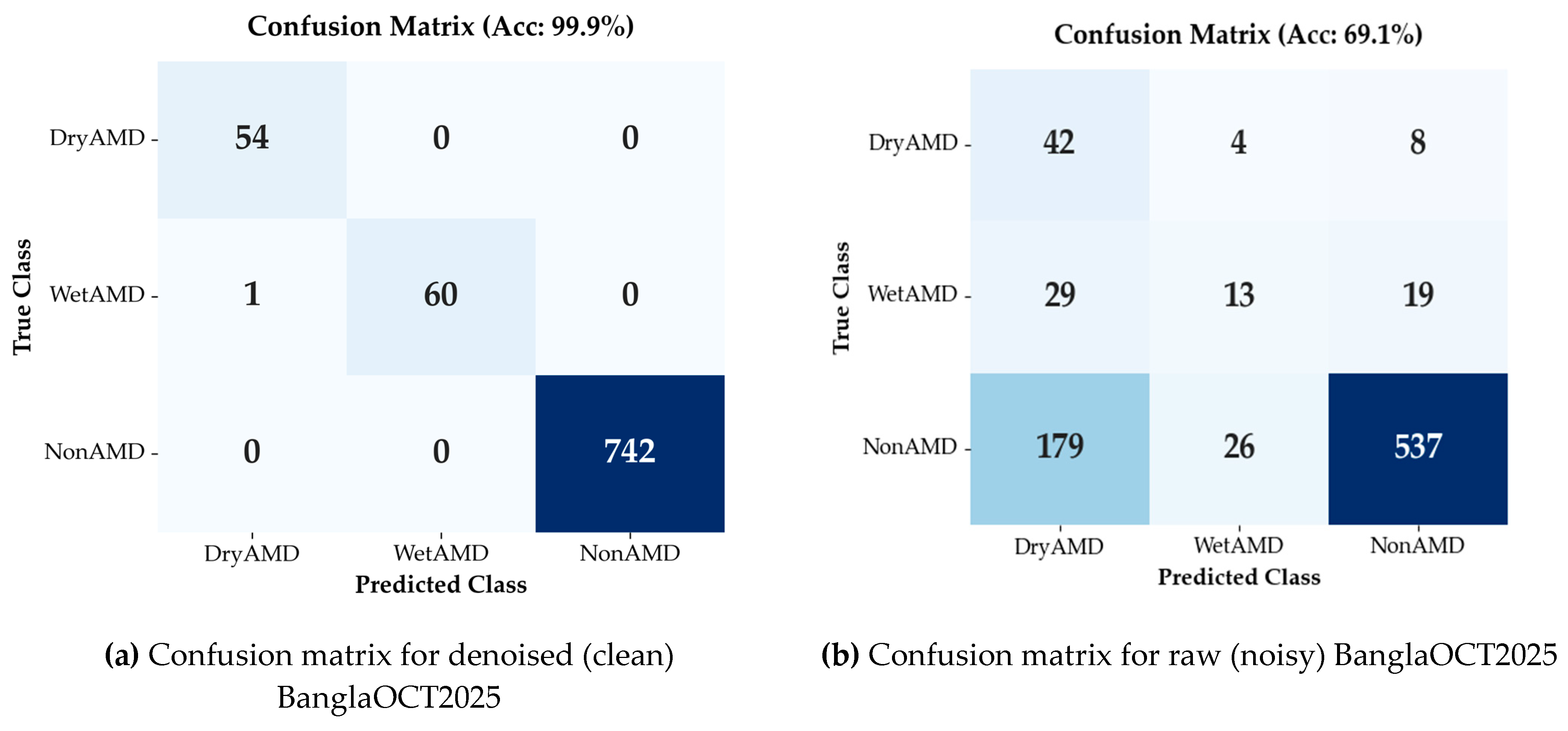
3.3.1. Classification Performance on BanglaOCT2025 Dataset
3.3.2. Denoising Effectiveness via Downstream Diagnostic Task
3.3.3. Class-Imbalance–Aware Analysis
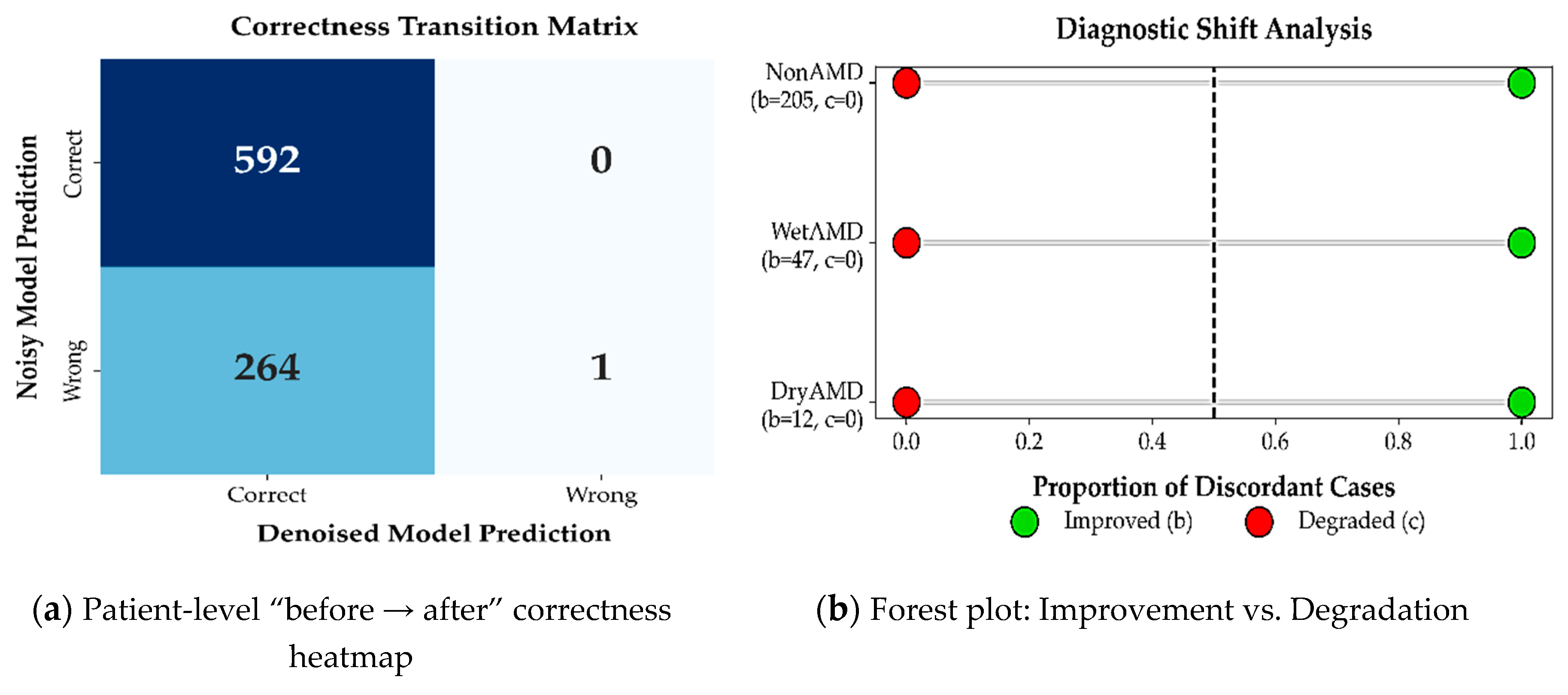
3.3.4. McNemar’s Test for Paired Diagnostic Outcomes
3.3.5. Performance Evaluation
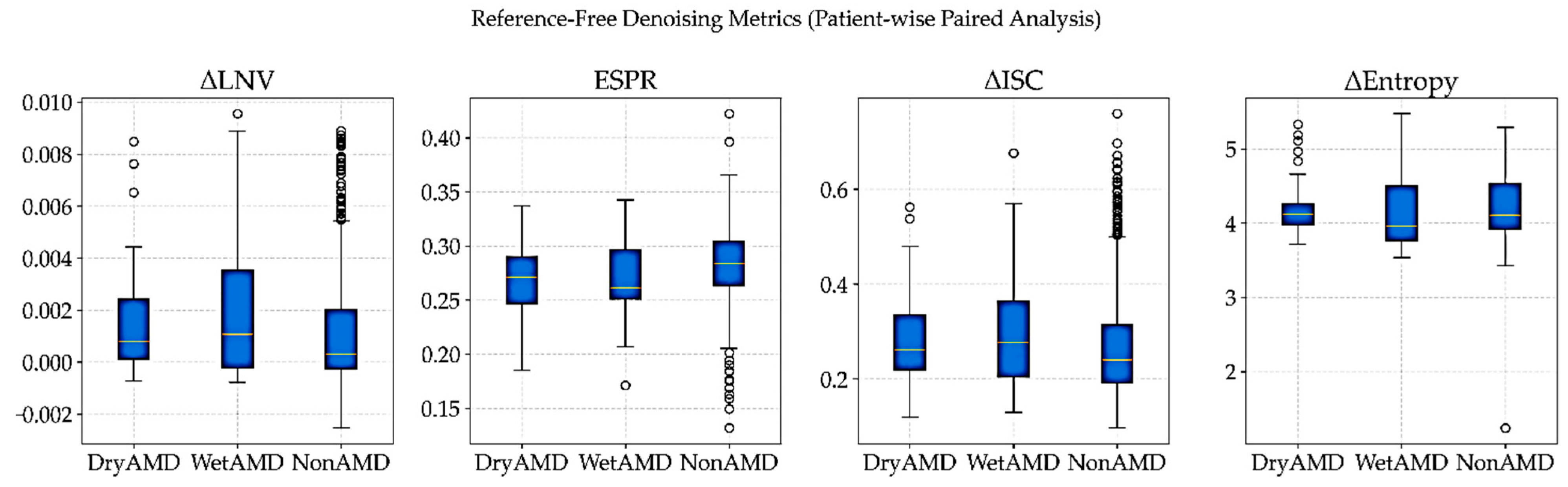
3.3.6. Reference-Free Evaluation of Denoising on Real OCT Volumes
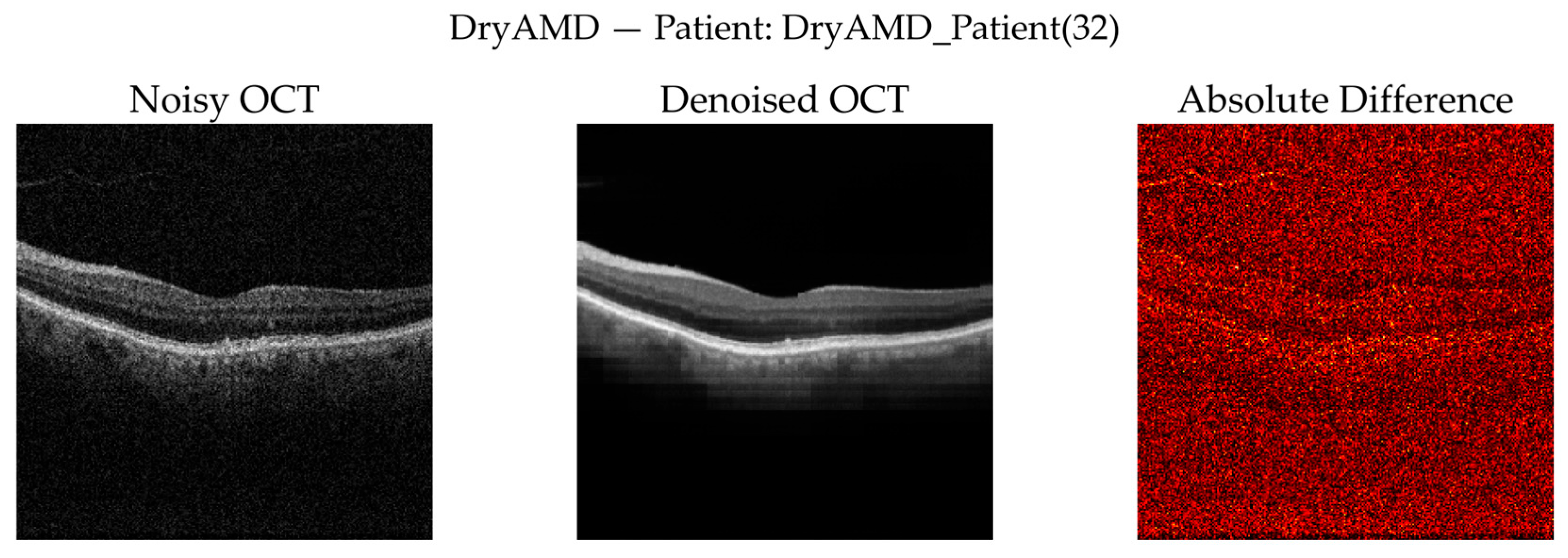
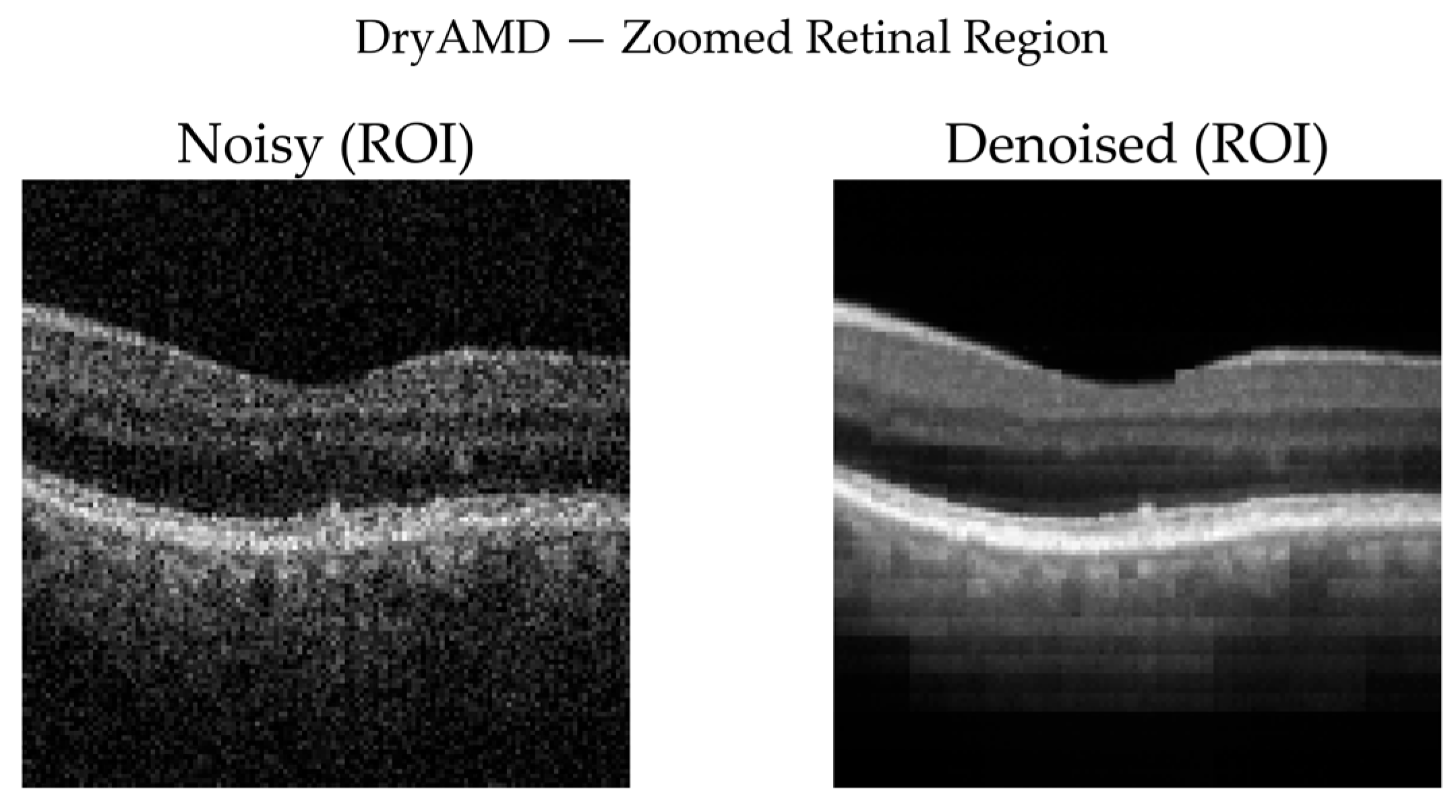
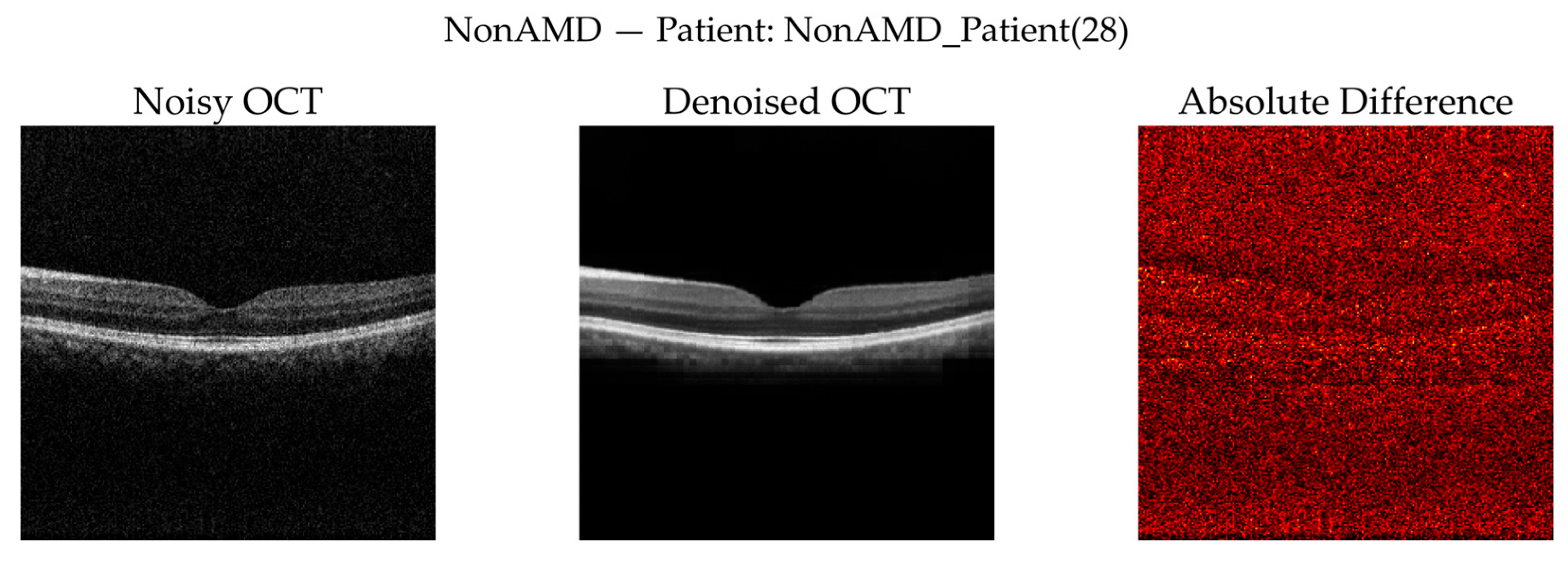
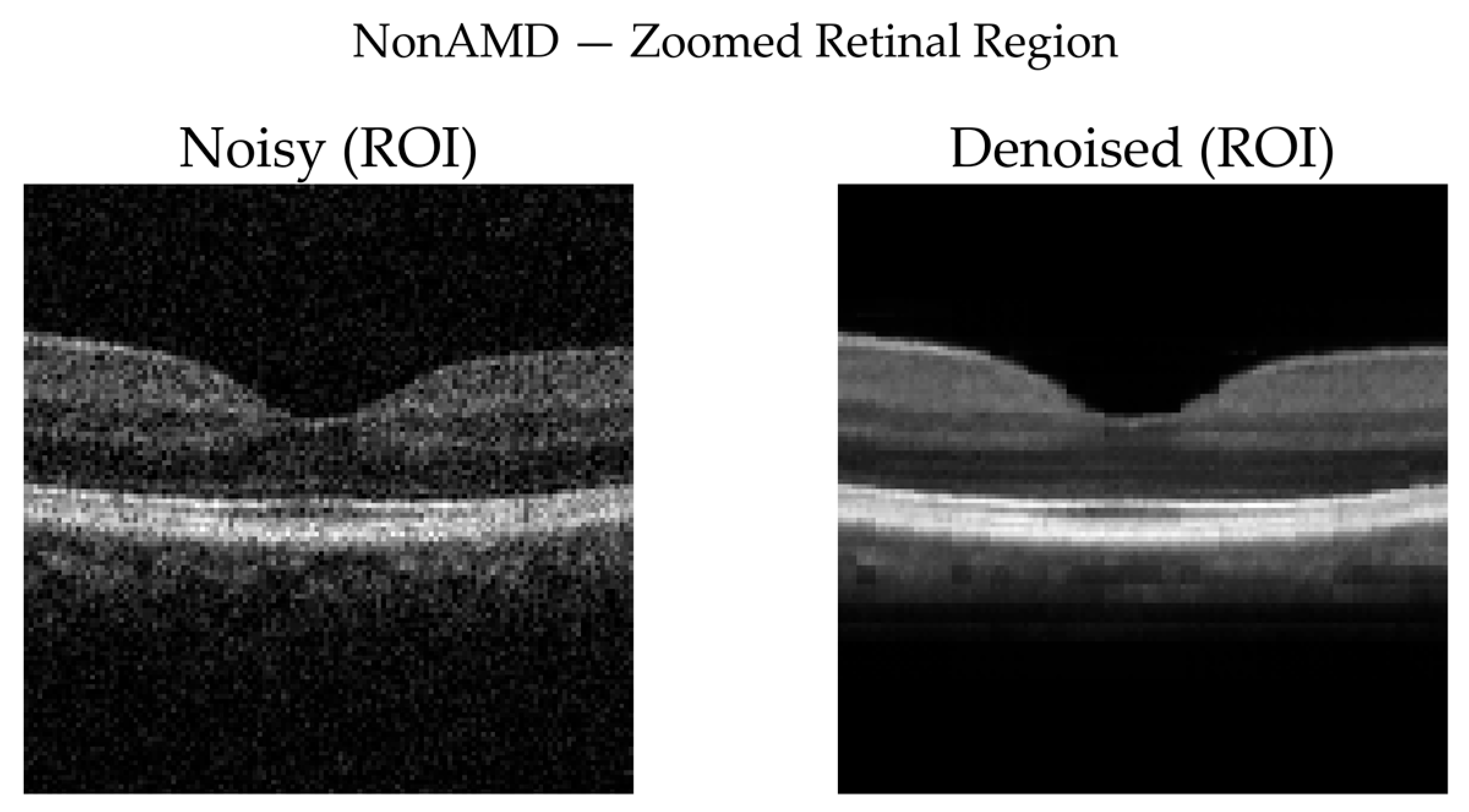
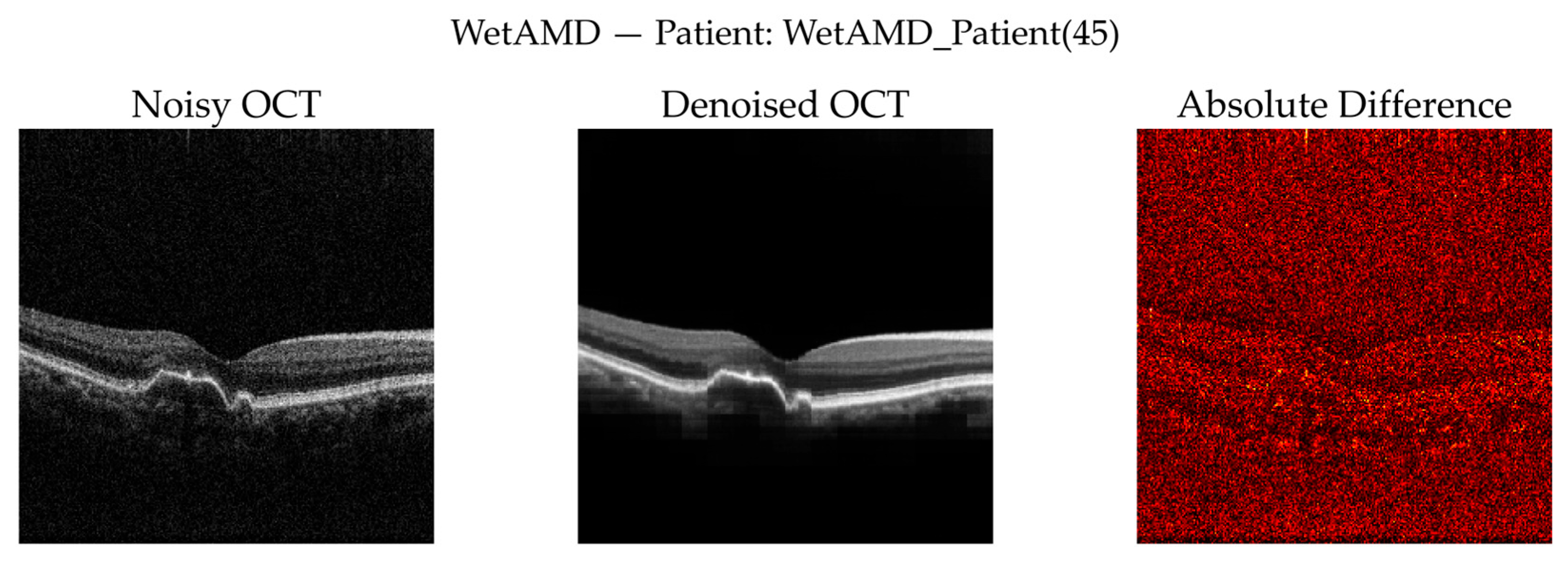
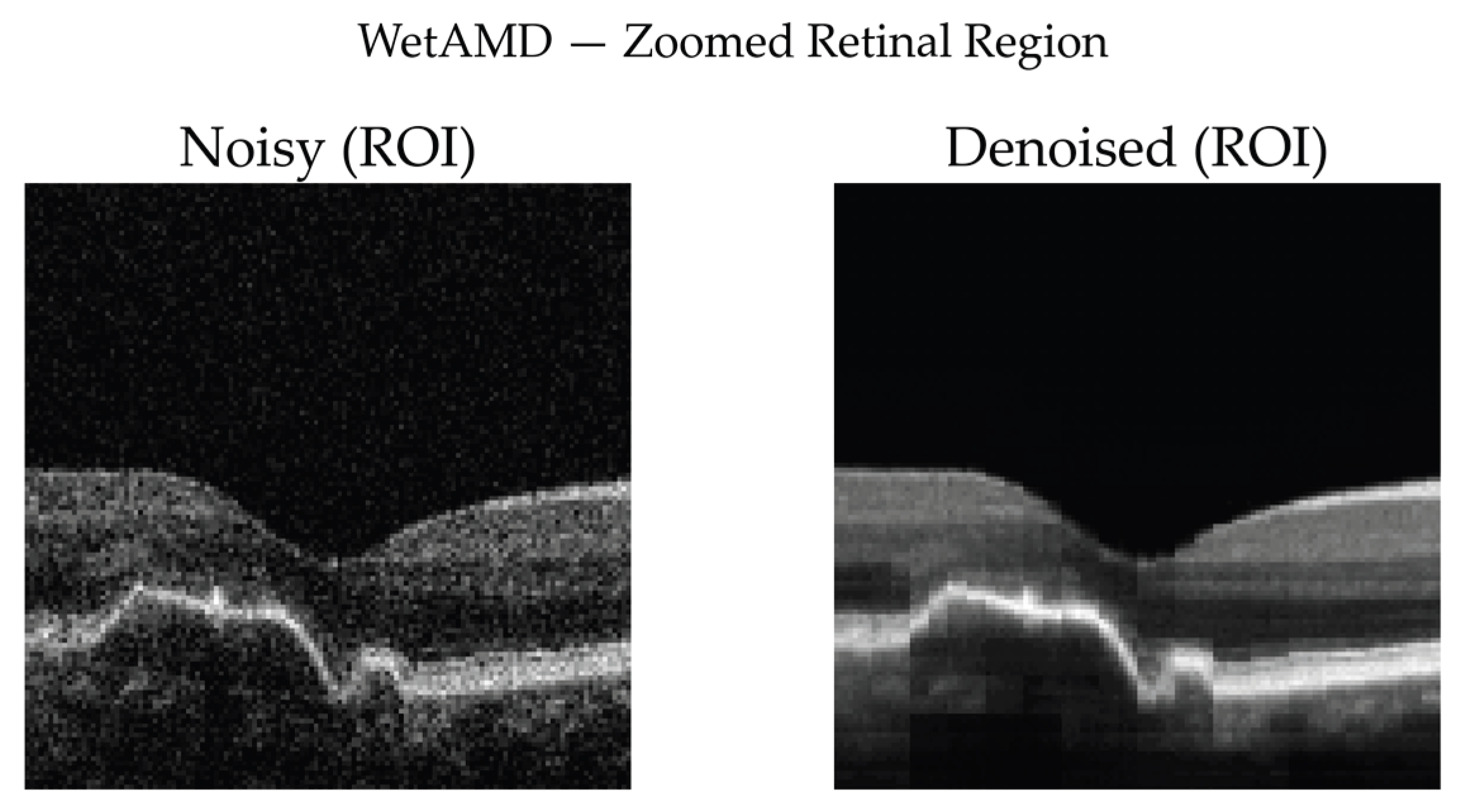
3.3.7. Qualitative Visual Assessment of Denoising Performance
3.3.8. Why Quantitative Metrics (PSNR, SSIM, MSE) Are Not Included
- No available clean ground truth for real OCT volumes: Self-supervised denoising cannot be directly benchmarked using reference-based metrics.
- The purpose of denoising is functional, not comparative: The FFSwin denoiser is used as a preprocessing backbone for AMD classification.
- Indirect validation through diagnostic accuracy: Our classifier trained on denoised volumes achieves 99.88% accuracy, which strongly indicates structural preservation and useful noise suppression.
- Novel dataset (BanglaOCT2025): No public baselines exist for fair cross-model comparison.
4. Discussion
4.1. Principal Findings
4.2. Comparison with Existing OCT Datasets and Processing Paradigms
4.3. Clinical Relevance of Self-Supervised Volumetric Denoising
4.4. Interpretation of Diagnostic Performance Gains
4.5. Class Imbalance and Robustness Considerations
4.6. Clinical and Practical Implications
4.7. Limitations
4.8. Future Directions
5. Conclusions
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
| 1 | FFSwin Architecture is used here as backbone. Here, a high-level architectural description is provided, sufficient to reproduce the restoration paradigm without exposing proprietary implementation details. |
References
- Huang, D.; Swanson, E.A.; Lin, C.P.; Schuman, J.S.; Stinson, W.G.; Chang, W.; Hee, M.R.; Flotte, T.; Gregory, K.; Puliafito, C.A.; et al. Optical Coherence Tomography. Science (1979) 1991, 254, 1178–1181. [Google Scholar] [CrossRef]
- Drexler, W.; Fujimoto, J.G. Optical Coherence Tomography: Technology and Applications; Springer Science & Business Media, 2008; ISBN 3540775501. [Google Scholar]
- Schmidt-Erfurth, U.; Waldstein, S.M. A Paradigm Shift in Imaging Biomarkers in Neovascular Age-Related Macular Degeneration. Prog Retin Eye Res 2016, 50, 1–24. [Google Scholar] [CrossRef] [PubMed]
- Farsiu, S.; Chiu, S.J.; O’Connell, R. V.; Folgar, F.A.; Yuan, E.; Izatt, J.A.; Toth, C.A. Quantitative Classification of Eyes with and without Intermediate Age-Related Macular Degeneration Using Optical Coherence Tomography. Ophthalmology 2014, 121, 162–172. [Google Scholar] [CrossRef]
- Li, M.; Huang, K.; Xu, Q.; Yang, J.; Zhang, Y.; Ji, Z.; Xie, K.; Yuan, S.; Liu, Q.; Chen, Q. OCTA-500: A Retinal Dataset for Optical Coherence Tomography Angiography Study. Med Image Anal 2024, 93, 103092. [Google Scholar] [CrossRef]
- Melinščak, M.; Radmilović, M.; Vatavuk, Z.; Lončarić, S. Annotated Retinal Optical Coherence Tomography Images (AROI) Database for Joint Retinal Layer and Fluid Segmentation. Automatika : časopis za automatiku, mjerenje, elektroniku, računarstvo i komunikacije 2021, 62, 375–385. [Google Scholar] [CrossRef]
- Wagner-Schuman, M.; Dubis, A.M.; Nordgren, R.N.; Lei, Y.; Odell, D.; Chiao, H.; Weh, E.; Fischer, W.; Sulai, Y.; Dubra, A.; et al. Race- and Sex-Related Differences in Retinal Thickness and Foveal Pit Morphology. Invest Ophthalmol Vis Sci 2011, 52, 625–634. [Google Scholar] [CrossRef]
- Esteva, A.; Robicquet, A.; Ramsundar, B.; Kuleshov, V.; DePristo, M.; Chou, K.; Cui, C.; Corrado, G.; Thrun, S.; Dean, J. A Guide to Deep Learning in Healthcare. Nature Medicine 2019 25:1 2019, 25, 24–29. [Google Scholar] [CrossRef] [PubMed]
- NIDEK CO.,LTD. Available online: https://www.nidek-intl.com/ (accessed on 7 November 2025).
- Logiciel NAVIS-EX - NIDEK France. Available online: https://www.nidek.fr/en/logiciel-navis-ex/logiciel-navis-ex-2/ (accessed on 16 December 2025).
- Ishikawa, H.; Stein, D.M.; Wollstein, G.; Beaton, S.; Fujimoto, J.G.; Schuman, J.S. Macular Segmentation with Optical Coherence Tomography. Invest Ophthalmol Vis Sci 2005, 46, 2012–2017. [Google Scholar] [CrossRef]
- Ahlers, C.; Golbaz, I.; Stock, G.; Fous, A.; Kolar, S.; Pruente, C.; Schmidt-Erfurth, U. Time Course of Morphologic Effects on Different Retinal Compartments after Ranibizumab Therapy in Age-Related Macular Degeneration. Ophthalmology 2008, 115, e39–e46. [Google Scholar] [CrossRef]
- Mylonas, G.; Ahlers, C.; Malamos, P.; Golbaz, I.; Deak, G.; Schütze, C.; Sacu, S.; Schmidt-Erfurth, U. Comparison of Retinal Thickness Measurements and Segmentation Performance of Four Different Spectral and Time Domain OCT Devices in Neovascular Age-Related Macular Degeneration. British Journal of Ophthalmology 2009, 93, 1453–1460. [Google Scholar] [CrossRef]
- Liu, Y.Y.; Chen, M.; Ishikawa, H.; Wollstein, G.; Schuman, J.S.; Rehg, J.M. Automated Macular Pathology Diagnosis in Retinal OCT Images Using Multi-Scale Spatial Pyramid and Local Binary Patterns in Texture and Shape Encoding. Med Image Anal 2011, 15, 748–759. [Google Scholar] [CrossRef] [PubMed]
- Goodman, J.W. Some Fundamental Properties of Speckle*. JOSA Vol. 66(Issue 11 66), pp. 1145-1150 1976 1145–1150. [CrossRef]
- Mehdizadeh, M.; MacNish, C.; Xiao, D.; Alonso-Caneiro, D.; Kugelman, J.; Bennamoun, M. Deep Feature Loss to Denoise OCT Images Using Deep Neural Networks. J Biomed Opt 2021, 26, 046003. [Google Scholar] [CrossRef]
- Li, F.; Wu, Q.; Jia, B.; Yang, Z. Speckle Noise Removal in OCT Images via Wavelet Transform and DnCNN. Applied Sciences 2025, Vol. 15 15, 6557 2025 6557. [Google Scholar] [CrossRef]
- Bepery, C.; Rahaman, G.M.A.; Debnath, R.; Saha, S. Forward Autoencoder Approach for Denoising Retinal OCT Images. 2025 International Conference on Electrical, Computer and Communication Engineering, ECCE, 2025 2025. [Google Scholar] [CrossRef]
- Ahmed, H.; Zhang, Q.; Donnan, R.; Alomainy, A. Transformer Enhanced Autoencoder Rendering Cleaning of Noisy Optical Coherence Tomography Images. Journal of Medical Imaging 2024, 11, 34008. [Google Scholar] [CrossRef]
- Özkan, A.; Stoykova, E.; Sikora, T.; Madjarova, V. Denoising OCT Images Using Steered Mixture of Experts with Multi-Model Inference. arXiv 2024, arXiv:2402.12735. [Google Scholar] [CrossRef]
- Weigert, M.; Schmidt, U.; Boothe, T.; Müller, A.; Dibrov, A.; Jain, A.; Wilhelm, B.; Schmidt, D.; Broaddus, C.; Culley, S.; et al. Content-Aware Image Restoration: Pushing the Limits of Fluorescence Microscopy. Nature Methods 2018, 15:12 15, 1090–1097. [Google Scholar] [CrossRef] [PubMed]
- Download NAVIS-EX by NIDEK CO., LTD. Available online: https://navis-ex.software.informer.com/download/ (accessed on 7 November 2025).
- RS-330 - NIDEK France. Available online: https://www.nidek.fr/en/oct-angiographie/rs-330/ (accessed on 16 December 2025).
- RS-3000 Advance 2 - NIDEK France. Available online: https://www.nidek.fr/en/oct-angiographie/rs-3000-advance-2/ (accessed on 16 December 2025).
- RS-330 Manual| ManualsLib. Available online: https://www.manualslib.com/manual/3952363/Nidek-Medical-Rs-330.html#manual (accessed on 16 December 2025).
- Hee, M.R.; Izatt, J.A.; Swanson, E.A.; Huang, D.; Schuman, J.S.; Lin, C.P.; Puliafito, C.A.; Fujimoto, J.G. Optical Coherence Tomography of the Human Retina. Archives of Ophthalmology 1995, 113, 325–332. [Google Scholar] [CrossRef]
- Ting, D.S.W.; Cheung, C.Y.L.; Lim, G.; Tan, G.S.W.; Quang, N.D.; Gan, A.; Hamzah, H.; Garcia-Franco, R.; Yeo, I.Y.S.; Lee, S.Y.; et al. Development and Validation of a Deep Learning System for Diabetic Retinopathy and Related Eye Diseases Using Retinal Images From Multiethnic Populations With Diabetes. JAMA 2017, 318, 2211–2223. [Google Scholar] [CrossRef]
- Landis, J.R.; Koch, G.G. The Measurement of Observer Agreement for Categorical Data. Biometrics 1977, 33, 159. [Google Scholar] [CrossRef] [PubMed]
- Network, D.R.C.R. Relationship between Optical Coherence Tomography–Measured Central Retinal Thickness and Visual Acuity in Diabetic Macular Edema. Ophthalmology 2007, 114, 525–536. [Google Scholar] [CrossRef]
- Chiu, S.J.; Li, X.T.; Nicholas, P.; Izatt, J.A.; Toth, C.A.; Chiu, S.J.; Farsiu, S. Automatic Segmentation of Seven Retinal Layers in SDOCT Images Congruent with Expert Manual Segmentation. Optics Express Vol. 18(Issue 18 18), pp. 19413-19428 2010 19413–19428. [CrossRef]
- Garvin, M.K.; Abràmoff, M.D.; Wu, X.; Russell, S.R.; Burns, T.L.; Sonka, M. Automated 3-D Intraretinal Layer Segmentation of Macular Spectral-Domain Optical Coherence Tomography Images. IEEE Trans Med Imaging 2009, 28, 1436–1447. [Google Scholar] [CrossRef] [PubMed]
- Goodman, J.W. Statistical Properties of Laser Speckle Patterns; 1975; pp. 9–75. [Google Scholar] [CrossRef]
- Schmitt, J.M.; Xiang, S.H.; Yung, K.M. Speckle in Optical Coherence Tomography. J Biomed Opt 1999, 4, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Fang, L.; Li, S.; McNabb, R.P.; Nie, Q.; Kuo, A.N.; Toth, C.A.; Izatt, J.A.; Farsiu, S. Fast Acquisition and Reconstruction of Optical Coherence Tomography Images via Sparse Representation. IEEE Trans Med Imaging 2013, 32, 2034–2049. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Lin, Y.; Cao, Y.; Hu, H.; Wei, Y.; Zhang, Z.; Lin, S.; Guo, B. Swin Transformer: Hierarchical Vision Transformer Using Shifted Windows. In Proceedings of the Proceedings of the IEEE/CVF international conference on computer vision, 2021; pp. 10012–10022. [Google Scholar]
- Hatamizadeh, A.; Tang, Y.; Nath, V.; Yang, D.; Myronenko, A.; Landman, B.; Roth, H.R.; Xu, D. Unetr: Transformers for 3d Medical Image Segmentation. In Proceedings of the Proceedings of the IEEE/CVF winter conference on applications of computer vision, 2022; pp. 574–584. [Google Scholar]
- Lehtinen, J.; Munkberg, J.; Hasselgren, J.; Laine, S.; Karras, T.; Aittala, M.; Aila, T. Noise2Noise: Learning Image Restoration without Clean Data. 35th International Conference on Machine Learning, ICML 2018 2018, 7, 4620–4631. [Google Scholar]
- Krull, A.; Buchholz, T.-O.; Jug, F. Noise2Void - Learning Denoising From Single Noisy Images. 2019, 2129–2137. [Google Scholar]
- Batson, J.; Royer, L. Noise2Self: Blind Denoising by Self-Supervision 2019, 524–533.
- Zhao, H.; Gallo, O.; Frosio, I.; Kautz, J. Loss Functions for Image Restoration With Neural Networks. IEEE Trans Comput Imaging 2017, 3, 47–57. [Google Scholar] [CrossRef]
- Goodfellow, I.; Bengio, Y.; Courville, A.; Bengio, Y. Deep Learning; MIT press: Cambridge, 2016; Vol. 1. [Google Scholar]
- Perona, P.; Malik, J. Scale-Space and Edge Detection Using Anisotropic Diffusion. IEEE Trans Pattern Anal Mach Intell 1990, 12, 629–639. [Google Scholar] [CrossRef]
- Sobel, I.; Feldman, G. A 3×3 Isotropic Gradient Operator for Image Processing; Stanford Artificial Intelligence Project (SAIL): Stanford, CA, USA, 1968. [Google Scholar]
- Shannon, C.E. A Mathematical Theory of Communication. The Bell system technical journal 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Wang, Z.; Bovik, A.C. Mean Squared Error: Love It or Leave It? A New Look at Signal Fidelity Measures. IEEE Signal Process Mag 2009, 26, 98–117. [Google Scholar] [CrossRef]
| OCT Machine Model | Patients in NAVIS-EX* | Valid Patients | Valid Scans** | Scans for BanglaOCT2025 | Slices in BanglaOCT2025 |
|---|---|---|---|---|---|
| Nidek RS-330 Duo 2 | 1071 | 738 | 1128 | 1147 | 146816 |
| Nidek RS-3000 Advance | 348 | 333 | 530 | 438 | 56064 |
| Total | 1419 | 1071 | 1658 | 1585 | 202880 |
| Particulars | Quantity |
|---|---|
| Total patients | 1419 |
| Valid patients | 1071 |
| Scans from both eyes or multi scans from single eye | 1658 |
| Discard scans due to image acquisition issues | 73 |
| Considered scans for BanglaOCT2025 | 1585 |
| Considered 2D OCT slices for BanglaOCT2025 | 202880 |
| Patients for ground truth labelling in BanglaOCT2025 | 573 |
| Scans in BanglaOCT2025 without ground truth labelling | 728 |
| Scans for doctor labelling in BanglaOCT2025 | 857 |
| Dry AMD | 54 |
| Wet AMD | 61 |
| Non-AMD | 742 |
| Age Range | No. of Patients | Ground Truth Labelled 573 Patients | ||
|---|---|---|---|---|
| No. of Patients | Dry AMD | Wet AMD | ||
| 5-10.5 | 6 | 0 | 0 | 0 |
| 11-20.5 | 45 | 0 | 0 | 0 |
| 21-30.5 | 96 | 0 | 0 | 0 |
| 31-40.5 | 186 | 0 | 0 | 0 |
| 41-45.5 | 105 | 0 | 0 | 0 |
| 46-50.5 | 141 | 81 | 4 | 4 |
| 51-55.5 | 160 | 160 | 11 | 11 |
| 56-60.5 | 125 | 125 | 9 | 9 |
| 61-65.5 | 98 | 98 | 6 | 10 |
| 66-70.5 | 70 | 70 | 17 | 15 |
| 71-75.5 | 26 | 26 | 2 | 9 |
| 76-80.5 | 9 | 9 | 5 | 2 |
| 81-85.5 | 4 | 4 | 0 | 1 |
| Total | 1071 | 573 | 54 | 61 |
| Gender | Total Patients | Ground Truth Labelling |
Dry AMD | Wet AMD | Total AMD |
|---|---|---|---|---|---|
| Male | 658 | 349 | 31 | 36 | 67 |
| Female | 413 | 224 | 23 | 25 | 48 |
| Total | 1071 | 573 | 54 | 61 | 115 |
| Precision | Recall | F1-Score | Support | |
|---|---|---|---|---|
| DryAMD | 0.17 | 0.78 | 0.28 | 54 |
| WetAMD | 0.30 | 0.21 | 0.25 | 61 |
| NonAMD | 0.95 | 0.72 | 0.82 | 742 |
| Precision | Recall | F1-Score | Support | |
|---|---|---|---|---|
| DryAMD | 0.98 | 1.00 | 0.99 | 54 |
| WetAMD | 1.00 | 0.98 | 0.99 | 61 |
| NonAMD | 1.00 | 1.00 | 1.00 | 742 |
| Clean Correct: Yes | Clean Correct: No | |
|---|---|---|
| Noisy Correct: YES | 592 → a | 0 → c |
| Noisy Correct: No | 264 → b | 1→ d |
| Measurement Parameters | Value |
|---|---|
| b (improved) | 264 |
| c (degraded) | 0 |
| 262.0038 | |
| 0.000000 | |
| 264 | |
| 0.000000 | |
| Exact binomial p-value | (Essentially 0) |
| Class | Condition | TP | FN | FP | TN | Sensitivity | Specificity |
|---|---|---|---|---|---|---|---|
| DryAMD | Noisy | 42 | 12 | 208 | 595 | 0.7778 | 0.7407 |
| DryAMD | Denoised | 54 | 0 | 1 | 802 | 1 | 0.9988 |
| WetAMD | Noisy | 13 | 48 | 30 | 766 | 0.2131 | 0.9623 |
| WetAMD | Denoised | 60 | 1 | 0 | 796 | 0.9836 | 1 |
| NonAMD | Noisy | 537 | 205 | 27 | 88 | 0.7237 | 0.7652 |
| NonAMD | Denoised | 742 | 0 | 0 | 115 | 1 | 1 |
| Metric | Noisy Data | Denoised Data | Δ Improvement |
|---|---|---|---|
| Overall Accuracy | 0.6908 | 0.9988 | 0.308 |
| Balanced Accuracy | 0.5715 | 0.9945 | 0.423 |
| Macro Precision | 0.4742 | 0.9939 | 0.5197 |
| Macro Recall | 0.5715 | 0.9945 | 0.423 |
| Macro F1-score | 0.4496 | 0.9942 | 0.5446 |
| Weighted F1-score | 0.7472 | 0.9988 | 0.2516 |
| MCC | 0.2912 | 0.9952 | 0.704 |
| Cohen’s Kappa | 0.2426 | 0.9952 | 0.7526 |
| Class | Noisy Recall | Denoised Recall |
|---|---|---|
| DryAMD | 0.7778 | 1 |
| WetAMD | 0.2131 | 0.9836 |
| NonAMD | 0.7237 | 1 |
| Class | Δ LNV | ESPR | Δ ISC | Δ Entropy |
|---|---|---|---|---|
| DryAMD | 0.0015 | 0.269 | 0.2859 | 4.183 |
| WetAMD | 0.0018 | 0.2684 | 0.2963 | 4.1497 |
| NonAMD | 0.0012 | 0.2829 | 0.2695 | 4.2225 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




