Submitted:
13 December 2025
Posted:
18 December 2025
You are already at the latest version
Abstract
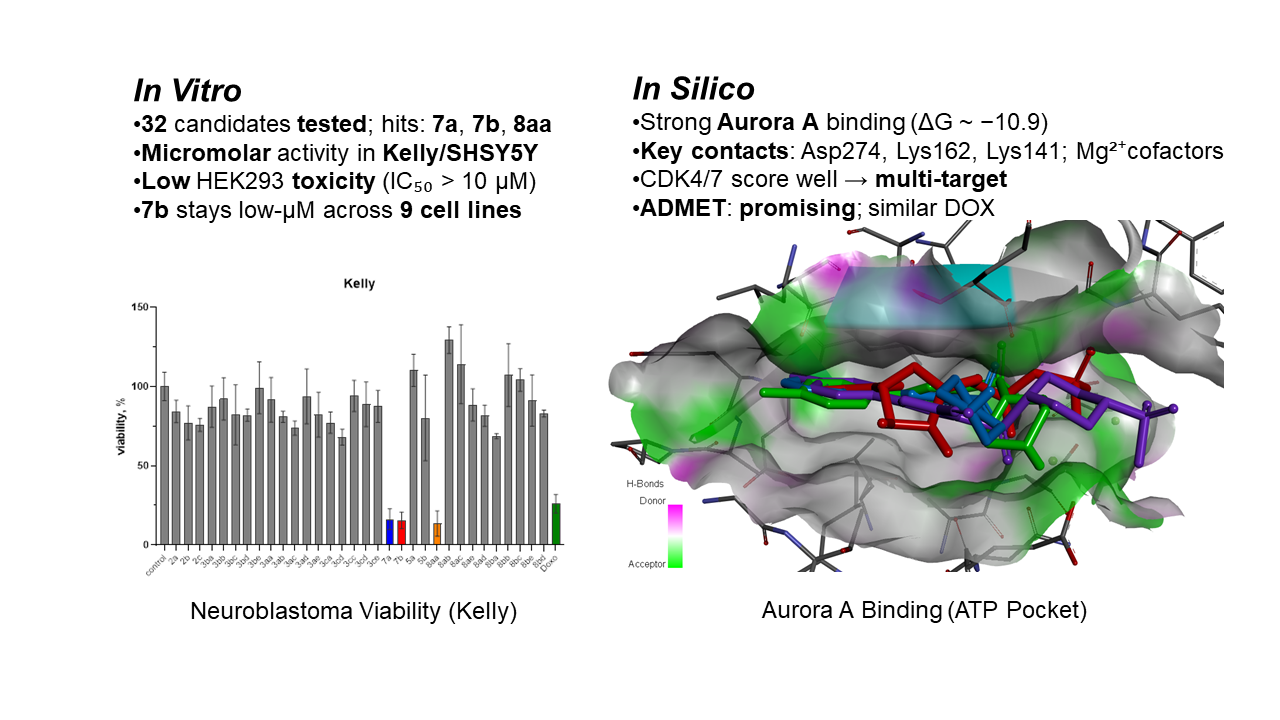
Keywords:
1. Introduction
2. Experimental Section
2.1. Chemicals Materials and Methods
2.2. Biology. In vitro Anticancer Studies.
2.3. Molecular Modeling
2.4. ADMET Evaluation
2.5. Biodegradability Study.
3. Result and Discussion
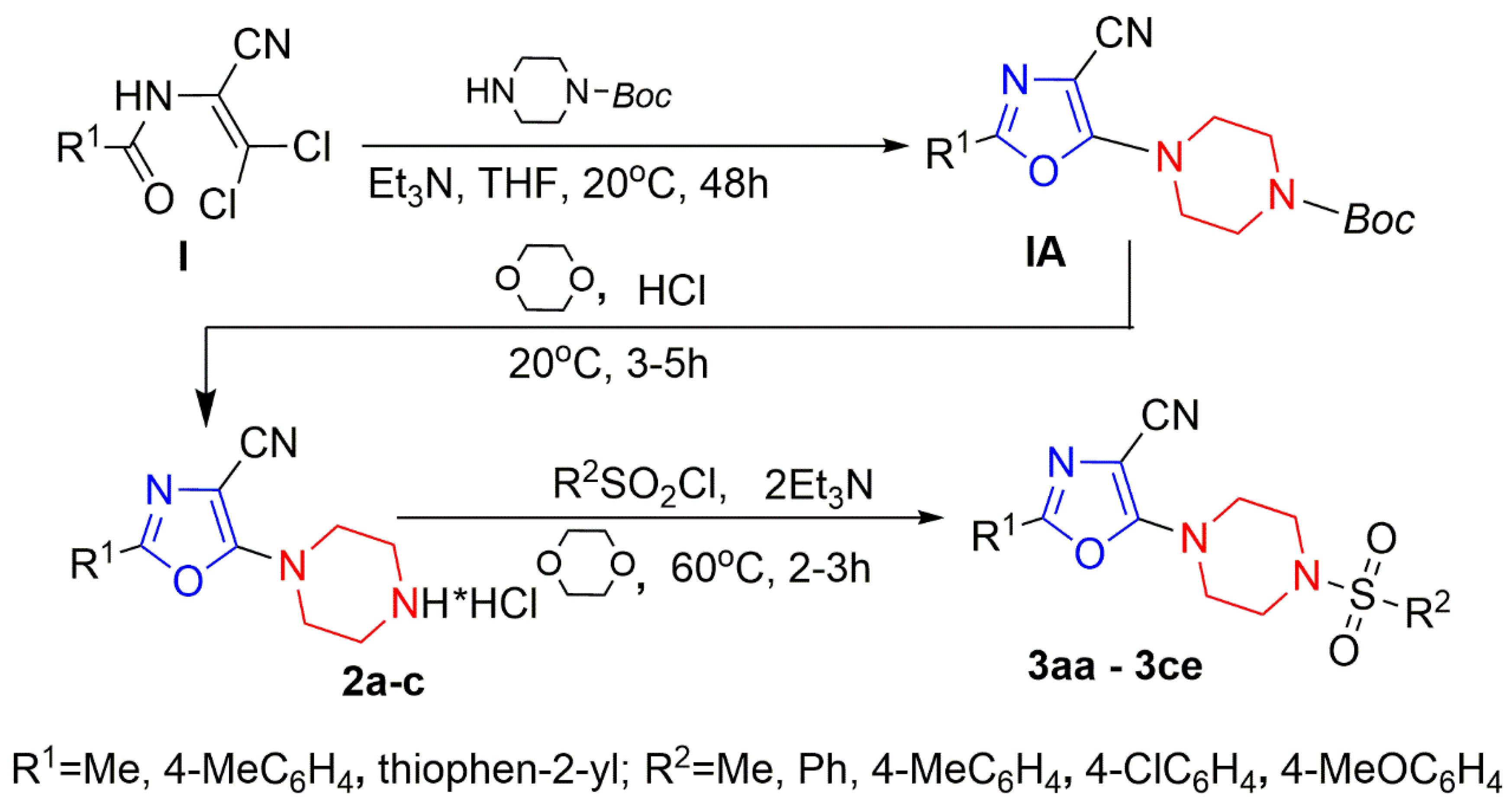
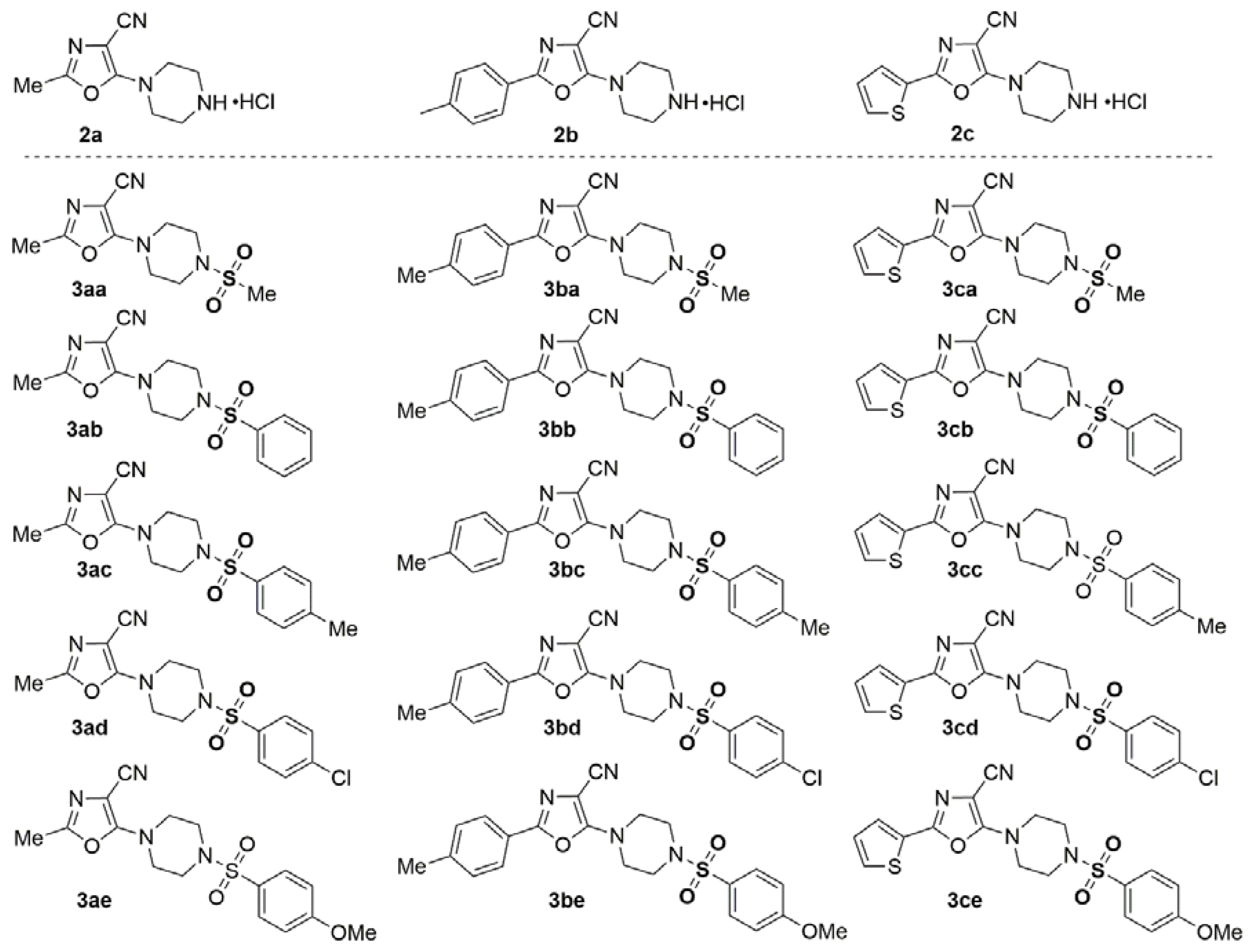
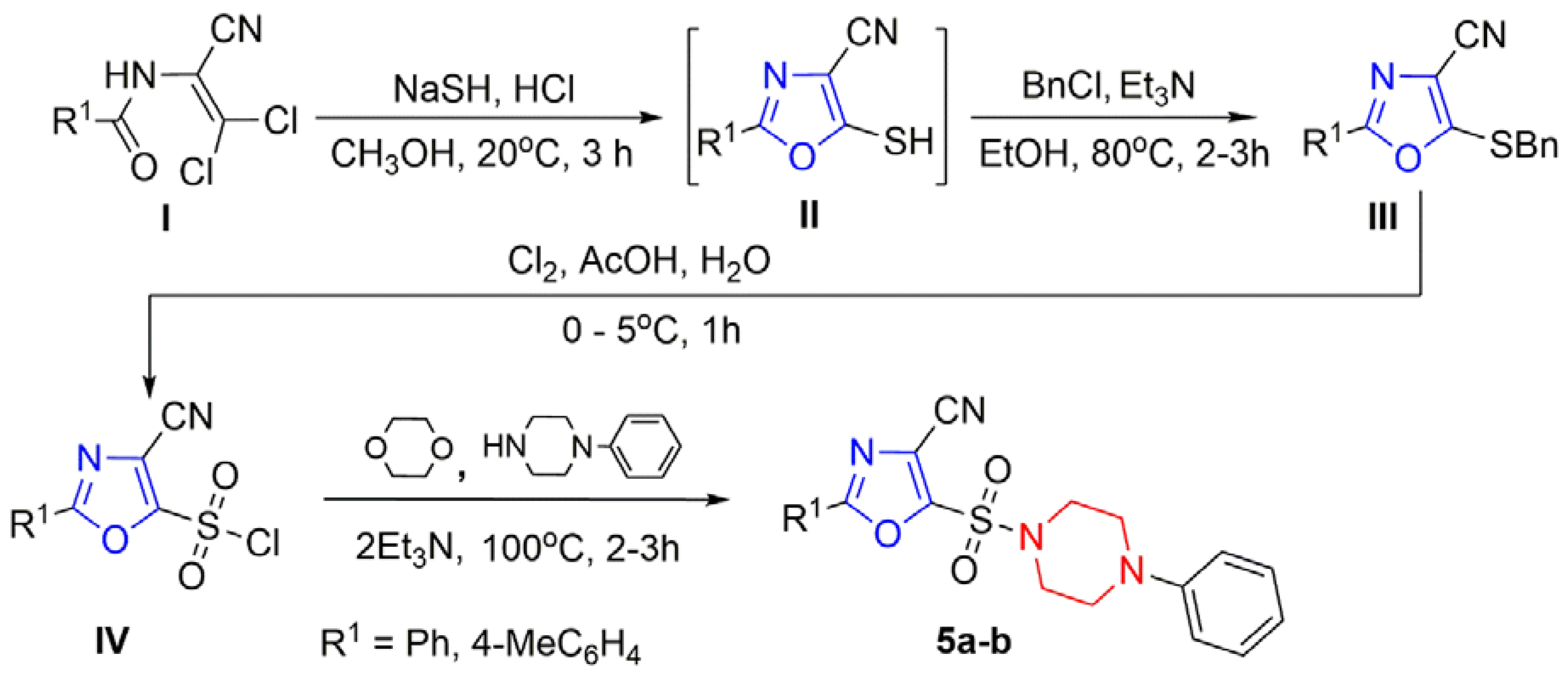
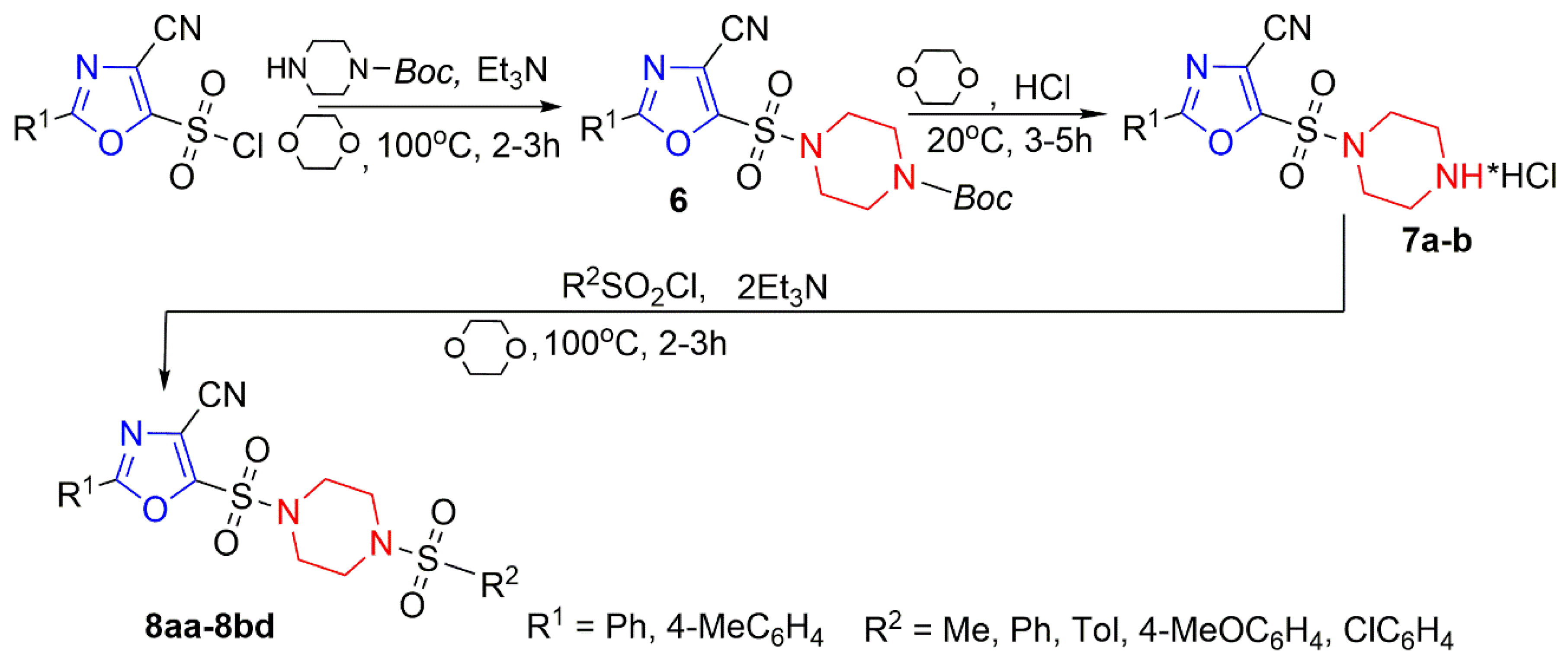
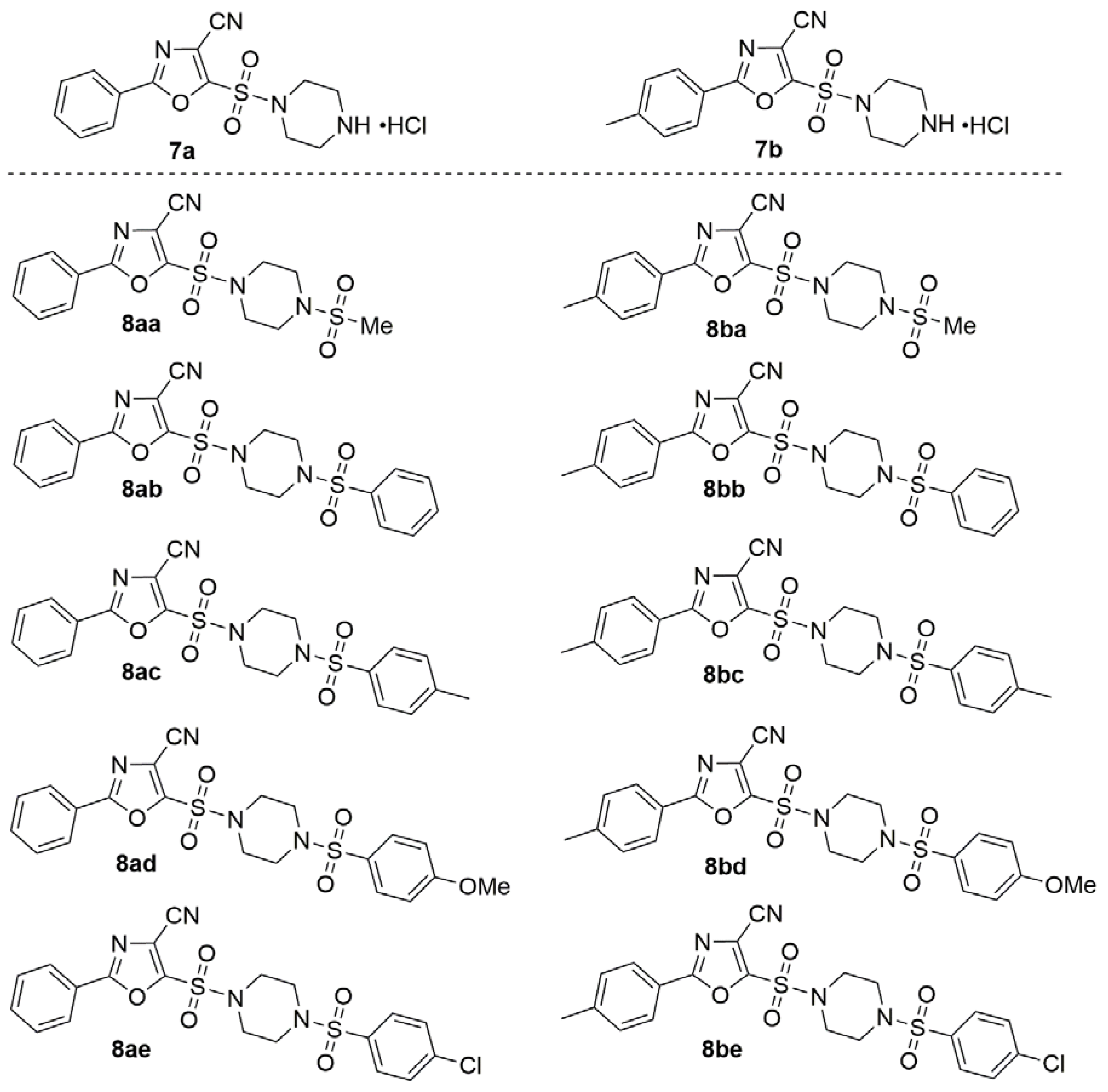
3.1. Chemistry
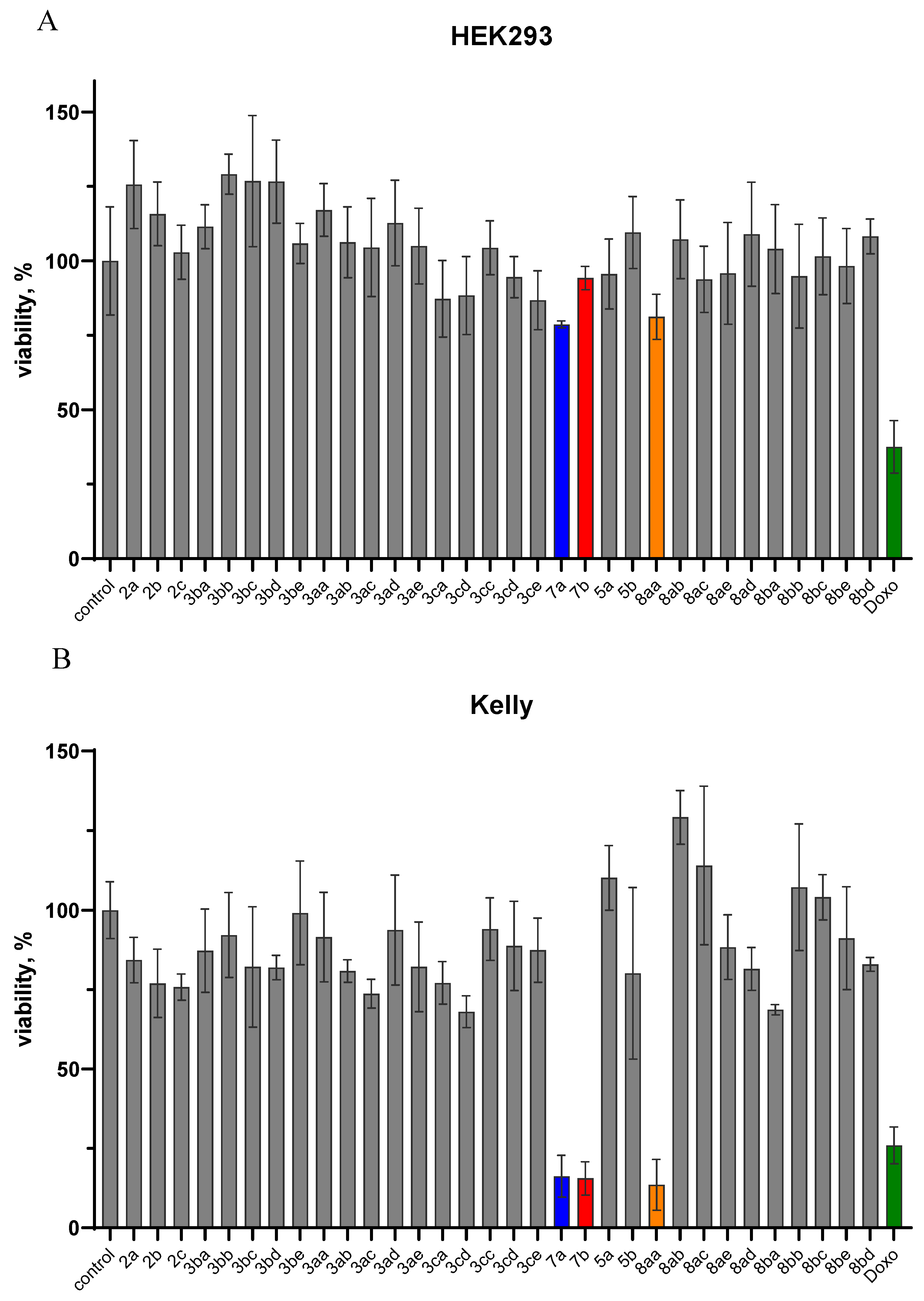
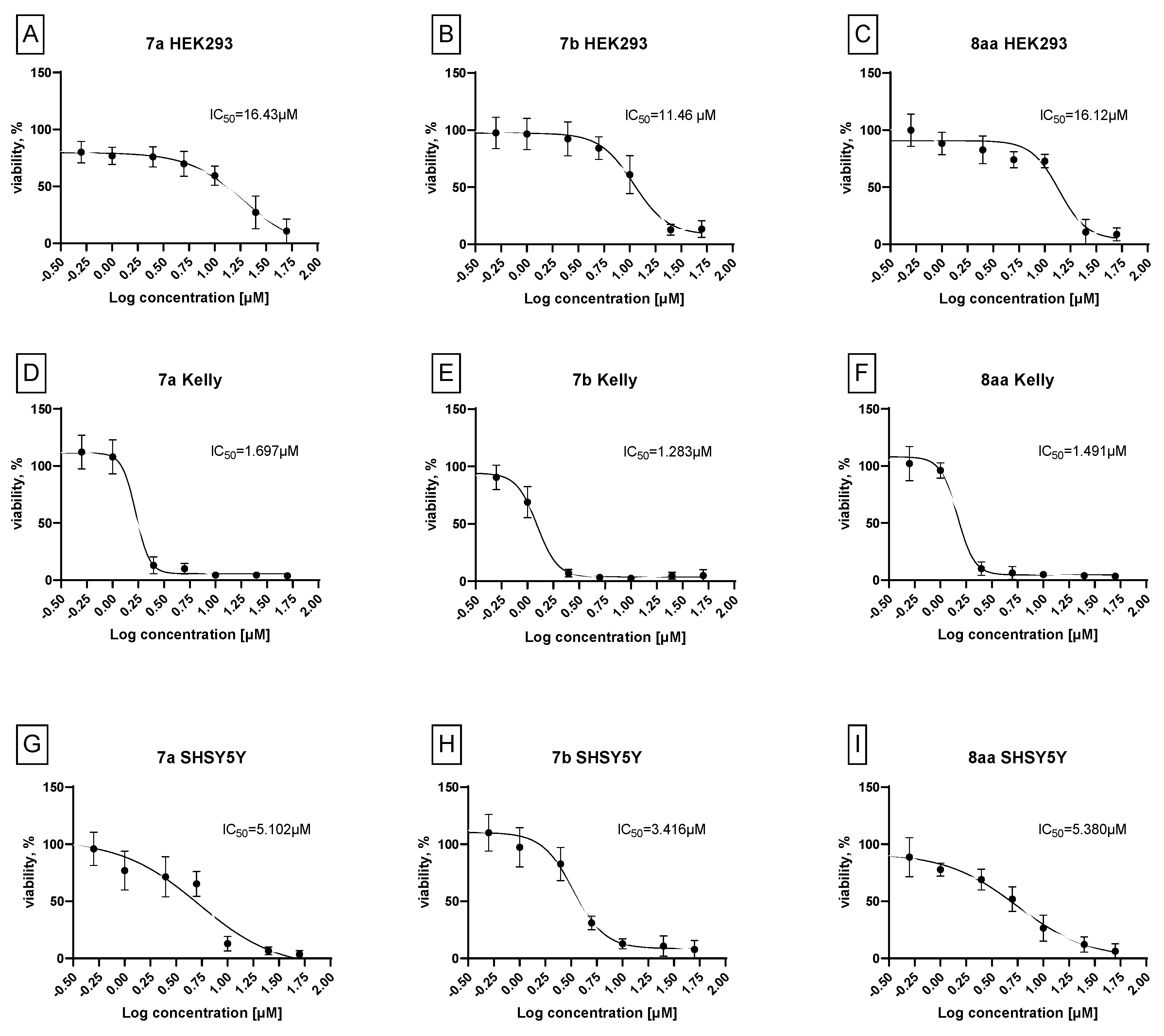
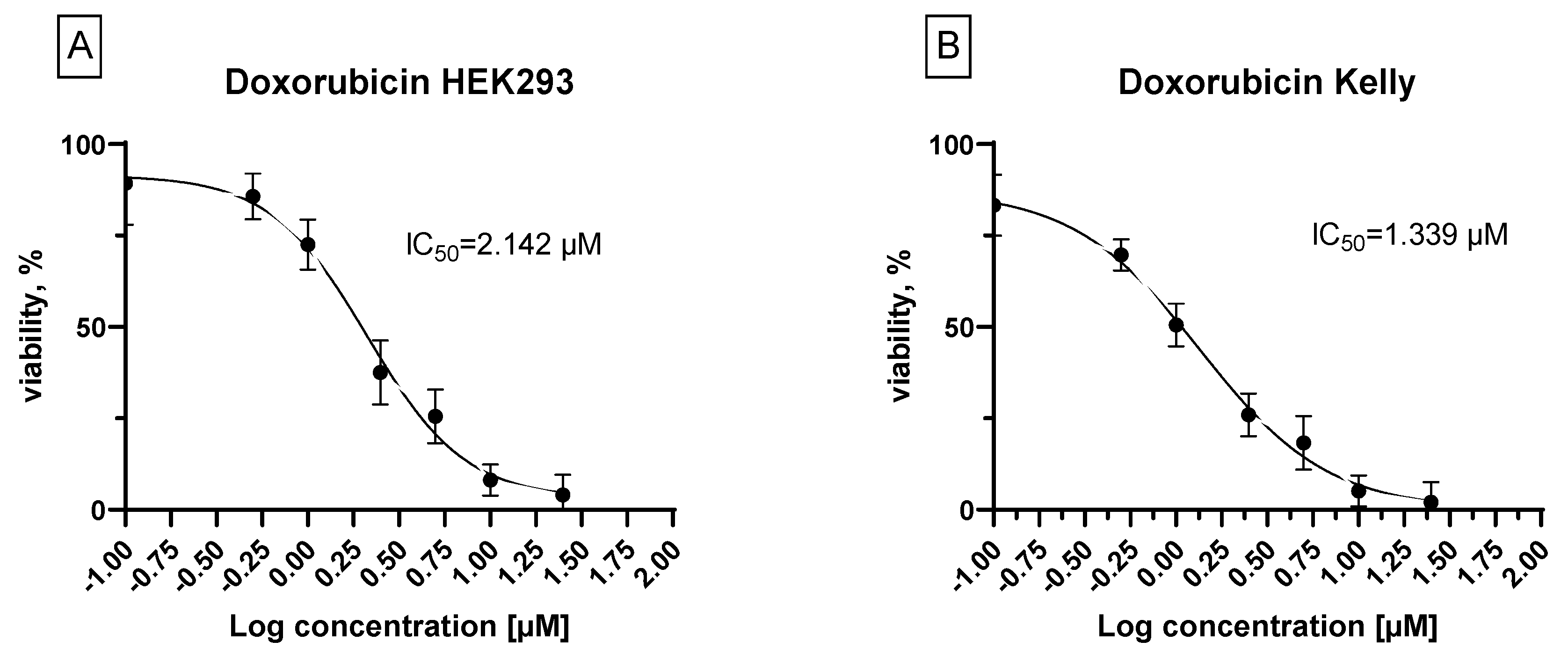
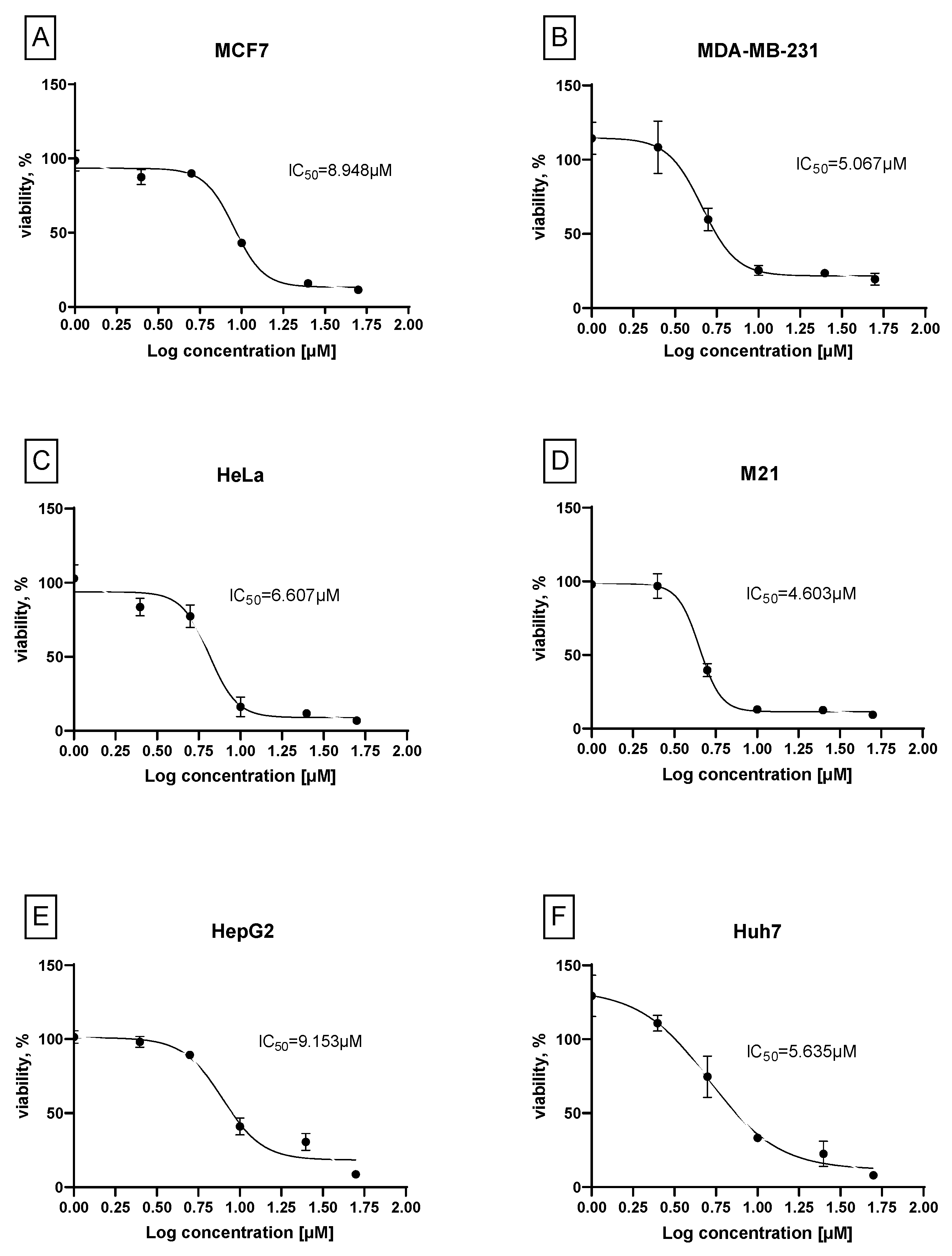
3.2. Biology. In Vitro Cytotoxicity
3.2.1. Selectivity Index
3.3. Biodegradability Study.
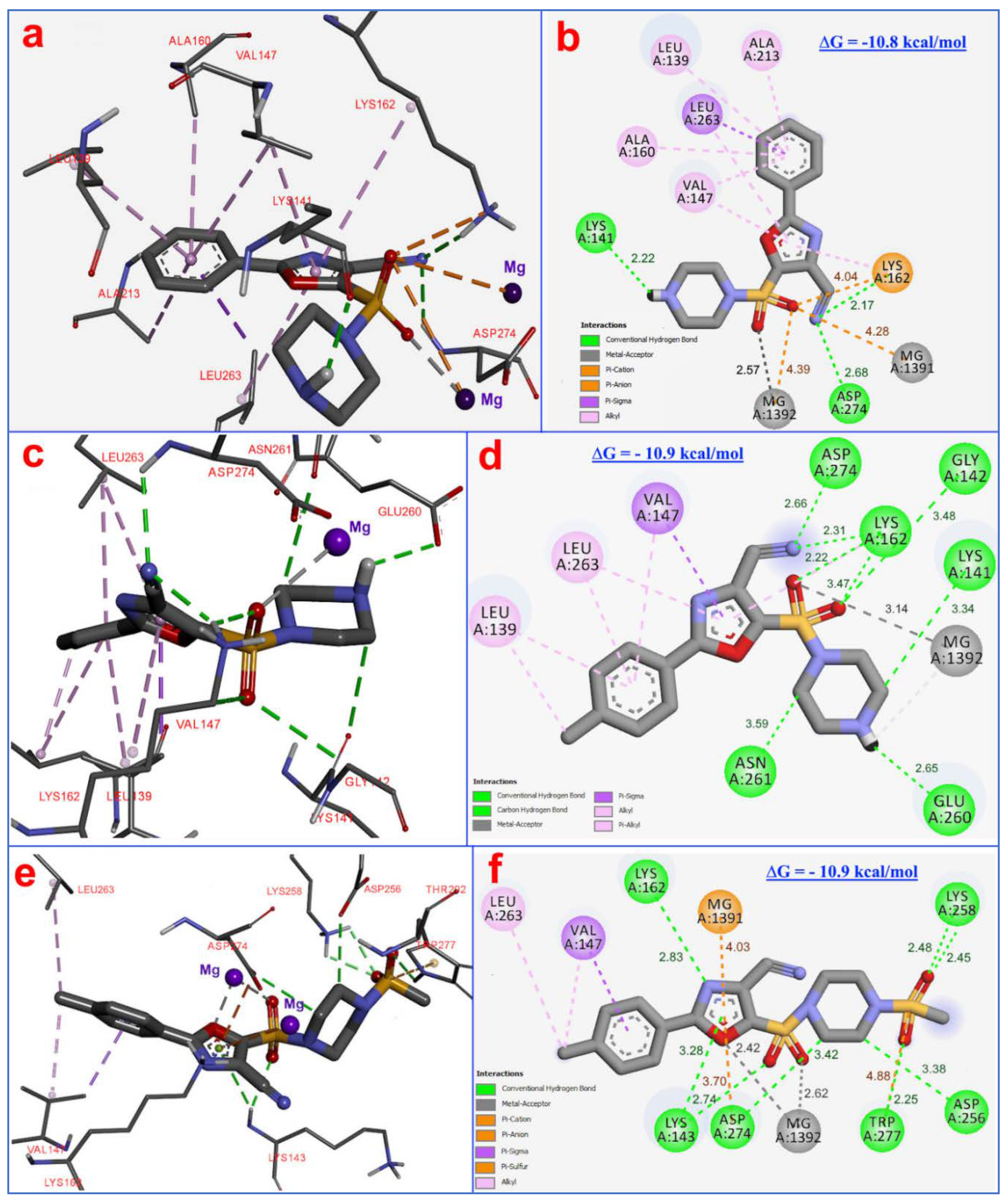
3.4. Molecular Modelling
3.5. Biology and Molecular Docking Results Discussion
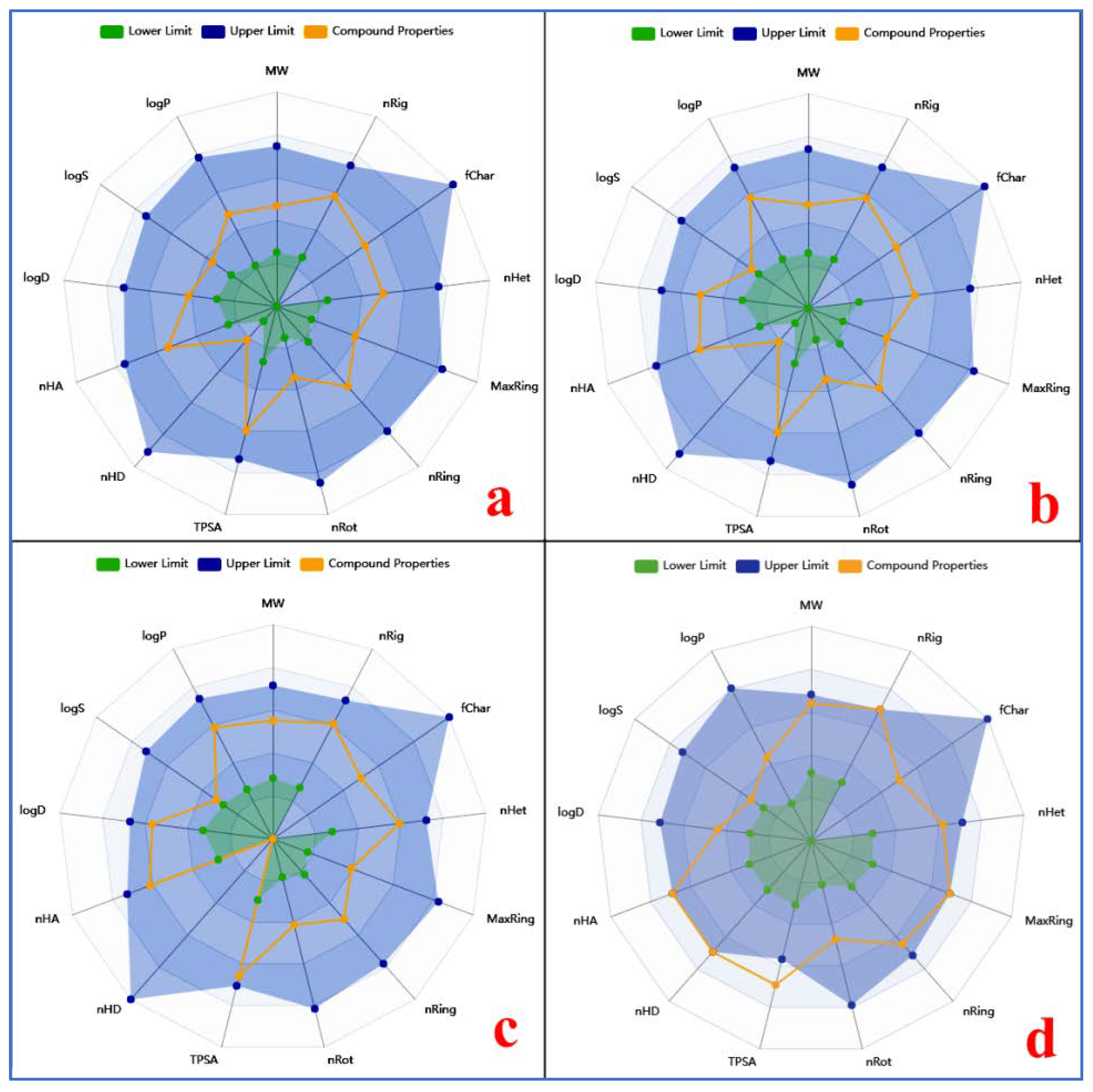
3.6. ADMET Evaluation
4. Conclusions
Supplementary Materials
Author Contributions
Acknowledgments
Conflicts of interest
Abbreviations
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J Clin 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Siegel, R.L.; Kratzer, T.B.; Giaquinto, A.N.; Sung, H.; Jemal, A. Cancer Statistics, 2025. CA Cancer J Clin 2025, 75, 10–45. [Google Scholar] [CrossRef]
- Zheng, M.; Kumar, A.; Sharma, V.; Behl, T.; Sehgal, A.; Wal, P.; Shinde, N.V.; Kawaduji, B.S.; Kapoor, A.; Anwer, Md.K.; et al. Revolutionizing Pediatric Neuroblastoma Treatment: Unraveling New Molecular Targets for Precision Interventions. Front Cell Dev Biol 2024, 12. [Google Scholar] [CrossRef]
- Maris, J.M.; Matthay, K.K. Molecular Biology of Neuroblastoma. Journal of Clinical Oncology 1999, 17, 2264–2264. [Google Scholar] [CrossRef]
- Bedoya-Reina, O.C.; Li, W.; Arceo, M.; Plescher, M.; Bullova, P.; Pui, H.; Kaucka, M.; Kharchenko, P.; Martinsson, T.; Holmberg, J.; et al. Single-Nuclei Transcriptomes from Human Adrenal Gland Reveal Distinct Cellular Identities of Low and High-Risk Neuroblastoma Tumors. Nat Commun 2021, 12, 5309. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Pacheco, M.L.; Hernández-Lemus, E.; Mejía, C. Analysis of High-Risk Neuroblastoma Transcriptome Reveals Gene Co-Expression Signatures and Functional Features. Biology (Basel) 2023, 12, 1230. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.X.; Zhao, H.; Kung, B.; Kim, D.Y.; Hicks, S.L.; Cohn, S.L.; Cheung, N.-K.; Seeger, R.C.; Evans, A.E.; Ikegaki, N. The MYCN Enigma: Significance of MYCN Expression in Neuroblastoma. Cancer Res 2006, 66, 2826–2833. [Google Scholar] [CrossRef]
- Duffy, M.J.; O’Grady, S.; Tang, M.; Crown, J. MYC as a Target for Cancer Treatment. Cancer Treat Rev 2021, 94, 102154. [Google Scholar] [CrossRef] [PubMed]
- Papadimitropoulou, A.; Makri, M.; Zoidis, G. MYC the Oncogene from Hell: Novel Opportunities for Cancer Therapy. Eur J Med Chem 2024, 267, 116194. [Google Scholar] [CrossRef]
- Hiyama, E.; Hiyama, K.; Yokoyama, T.; Ishii, T. Immunohistochemical Analysis of N-Myc Protein Expression in Neuroblastoma: Correlation with Prognosis of Patients. J Pediatr Surg 1991, 26, 838–843. [Google Scholar] [CrossRef]
- Richards, M.W.; Burgess, S.G.; Poon, E.; Carstensen, A.; Eilers, M.; Chesler, L.; Bayliss, R. Structural Basis of N-Myc Binding by Aurora-A and Its Destabilization by Kinase Inhibitors. Proceedings of the National Academy of Sciences 2016, 113, 13726–13731. [Google Scholar] [CrossRef]
- Sjostrom, S.K.; Finn, G.; Hahn, W.C.; Rowitch, D.H.; Kenney, A.M. The Cdk1 Complex Plays a Prime Role in Regulating N-Myc Phosphorylation and Turnover in Neural Precursors. Dev Cell 2005, 9, 327–338. [Google Scholar] [CrossRef] [PubMed]
- Gustafson, W.C.; Meyerowitz, J.G.; Nekritz, E.A.; Chen, J.; Benes, C.; Charron, E.; Simonds, E.F.; Seeger, R.; Matthay, K.K.; Hertz, N.T.; et al. Drugging MYCN through an Allosteric Transition in Aurora Kinase A. Cancer Cell 2014, 26, 414–427. [Google Scholar] [CrossRef]
- Sells, T.B.; Chau, R.; Ecsedy, J.A.; Gershman, R.E.; Hoar, K.; Huck, J.; Janowick, D.A.; Kadambi, V.J.; LeRoy, P.J.; Stirling, M.; et al. MLN8054 and Alisertib (MLN8237): Discovery of Selective Oral Aurora A Inhibitors. ACS Med Chem Lett 2015, 6, 630–634. [Google Scholar] [CrossRef]
- Joshi, S.; Mehra, M.; Singh, R.; Kakar, S. Review on Chemistry of Oxazole Derivatives: Current to Future Therapeutic Prospective. Egypt J Basic Appl. Sci 2023, 10, 218–239. [Google Scholar] [CrossRef]
- Chiacchio, M.A.; Lanza, G.; Chiacchio, U.; Giofrè, S. V.; Romeo, R.; Iannazzo, D.; Legnani, L. Oxazole-Based Compounds As Anticancer Agents. Curr Med Chem 2020, 26, 7337–7371. [Google Scholar] [CrossRef] [PubMed]
- Kakkar, S.; Narasimhan, B. A Comprehensive Review on Biological Activities of Oxazole Derivatives. BMC Chem 2019, 13, 16. [Google Scholar] [CrossRef]
- Nakamura, K.; Taguchi, E.; Miura, T.; Yamamoto, A.; Takahashi, K.; Bichat, F.; Guilbaud, N.; Hasegawa, K.; Kubo, K.; Fujiwara, Y.; et al. KRN951, a Highly Potent Inhibitor of Vascular Endothelial Growth Factor Receptor Tyrosine Kinases, Has Antitumor Activities and Affects Functional Vascular Properties. Cancer Res 2006, 66, 9134–9142. [Google Scholar] [CrossRef]
- Goulooze, S.C.; Cohen, A.F.; Rissmann, R. Olaparib. Br J Clin Pharmacol 2016, 81, 171–173. [Google Scholar] [CrossRef]
- Deininger, M.; Buchdunger, E.; Druker, B.J. The Development of Imatinib as a Therapeutic Agent for Chronic Myeloid Leukemia. Blood 2005, 105, 2640–2653. [Google Scholar] [CrossRef] [PubMed]
- Klempner, S.; Tran, P. Profile of Rociletinib and Its Potential in the Treatment of Non-Small-Cell Lung Cancer. Lung Cancer: Targets and Therapy 2016, 7, 91–97. [Google Scholar] [CrossRef]
- Semenyuta, I.; Kovalishyn, V.; Tanchuk, V.; Pilyo, S.; Zyabrev, V.; Blagodatnyy, V.; Trokhimenko, O.; Brovarets, V.; Metelytsia, L. 1,3-Oxazole Derivatives as Potential Anticancer Agents: Computer Modeling and Experimental Study. Comput Biol Chem 2016, 65, 8–15. [Google Scholar] [CrossRef]
- ADMETlab 3.0: Online ADMET Prediction Tool 2025.
- Fu, L.; Shi, S.; Yi, J.; Wang, N.; He, Y.; Wu, Z.; Peng, J.; Deng, Y.; Wang, W.; Wu, C.; et al. ADMETlab 3.0: An Updated Comprehensive Online ADMET Prediction Platform Enhanced with Broader Coverage, Improved Performance, API Functionality and Decision Support. Nucleic Acids Res 2024, 52, W422–W431. [Google Scholar] [CrossRef]
- Pires, D.E. V.; Blundell, T.L.; Ascher, D.B. PkCSM: Predicting Small-Molecule Pharmacokinetic and Toxicity Properties Using Graph-Based Signatures. J Med Chem 2015, 58, 4066–4072. [Google Scholar] [CrossRef]
- Assessment of Chemicals | OECD. Available online: https://www.oecd.org/en/topics/assessment-of-chemicals.html (accessed on 5 July 2024).
- Friedrich, J.; Längin, A.; Kü Mmerer, K. Comparison of an Electrochemical and Luminescence-Based Oxygen Measuring System for Use in the Biodegradability Testing According to Closed Bottle Test (OECD 301D). [CrossRef]
- Nyholm, N. The European System of Standardized Legal Tests for Assessing the Biodegradability of Chemicals. Environ Toxicol Chem 1991, 10, 1237–1246. [Google Scholar] [CrossRef]
- Gupta, A.K.; Alam, Md.M.; Alam, S.; Ansari, A.; Mishra, S.; Husain, S.Q. Production of Ethanol from Jaggery. Int J Res Appl Sci Eng Technol 2022, 10, 5042–5045. [Google Scholar] [CrossRef]
- Kachaeva, M. V.; Pilyo, S.G.; Zhirnov, V. V.; Brovarets, V.S. Synthesis, Characterization, and in Vitro Anticancer Evaluation of 2-Substituted 5-Arylsulfonyl-1,3-Oxazole-4-Carbonitriles. Med. Chem. Res. 2019, 28, 71–80. [Google Scholar] [CrossRef]
- Puhlmann, N.; Vidaurre, R.; Kümmerer, K. Designing Greener Active Pharmaceutical Ingredients: Insights from Pharmaceutical Industry into Drug Discovery and Development. Eur. J. Pharm. Sci 2024, 192, 106614. [Google Scholar] [CrossRef]
- Kapitanov, I. V.; Špulák, M.; Pour, M.; Soukup, O.; Marek, J.; Jun, D.; Novak, M.; Diz de Almeida, J.S.F.; França, T.C.C.; Gathergood, N.; et al. Sustainable Ionic Liquids-Based Molecular Platforms for Designing Acetylcholinesterase Reactivators. Chem Biol Interact 2023, 385, 110735. [Google Scholar] [CrossRef]
- Berman, H.M. The Protein Data Bank. Nucleic Acids Res 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Kim, S.; Chen, J.; Cheng, T.; Gindulyte, A.; He, J.; He, S.; Li, Q.; Shoemaker, B.A.; Thiessen, P.A.; Yu, B.; et al. PubChem 2025 Update. Nucleic Acids Res 2025, 53, D1516–D1525. [Google Scholar] [CrossRef] [PubMed]
- Diebold, M.; Schönemann, L.; Eilers, M.; Sotriffer, C.; Schindelin, H. Crystal Structure of a Covalently Linked Aurora-A–MYCN Complex. Acta Crystallogr D Struct Biol 2023, 79, 1–9. [Google Scholar] [CrossRef]
- Mou, P.K.; Yang, E.J.; Shi, C.; Ren, G.; Tao, S.; Shim, J.S. Aurora Kinase A, a Synthetic Lethal Target for Precision Cancer Medicine. Exp Mol Med 2021, 53, 835–847. [Google Scholar] [CrossRef]
- Yee, K.W.L.; Chen, H.-W.T.; Hedley, D.W.; Chow, S.; Brandwein, J.; Schuh, A.C.; Schimmer, A.D.; Gupta, V.; Sanfelice, D.; Johnson, T.; et al. A Phase I Trial of the Aurora Kinase Inhibitor, ENMD-2076, in Patients with Relapsed or Refractory Acute Myeloid Leukemia or Chronic Myelomonocytic Leukemia. Invest New Drugs 2016, 34, 614–624. [Google Scholar] [CrossRef]
- Li, Y.; Zhang, Z.-F.; Chen, J.; Huang, D.; Ding, Y.; Tan, M.-H.; Qian, C.-N.; Resau, J.H.; Kim, H.; Teh, B.T. VX680/MK-0457, a Potent and Selective Aurora Kinase Inhibitor, Targets Both Tumor and Endothelial Cells in Clear Cell Renal Cell Carcinoma. Am J Transl Res 2010, 2, 296–308. [Google Scholar]
- Vats, P.; Saini, C.; Baweja, B.; Srivastava, S.K.; Kumar, A.; Kushwah, A.S.; Nema, R. Aurora Kinases Signaling in Cancer: From Molecular Perception to Targeted Therapies. Mol Cancer 2025, 24, 180. [Google Scholar] [CrossRef]
- Rihani, A.; Vandesompele, J.; Speleman, F.; Van Maerken, T. Inhibition of CDK4/6 as a Novel Therapeutic Option for Neuroblastoma. Cancer Cell Int 2015, 15, 76. [Google Scholar] [CrossRef] [PubMed]
- Chipumuro, E.; Marco, E.; Christensen, C.L.; Kwiatkowski, N.; Zhang, T.; Hatheway, C.M.; Abraham, B.J.; Sharma, B.; Yeung, C.; Altabef, A.; et al. CDK7 Inhibition Suppresses Super-Enhancer-Linked Oncogenic Transcription in MYCN-Driven Cancer. Cell 2014, 159, 1126–1139. [Google Scholar] [CrossRef]
- Raghavendra, N.M.; Pingili, D.; Kadasi, S.; Mettu, A.; Prasad, S.V.U.M. Dual or Multi-Targeting Inhibitors: The next Generation Anticancer Agents. Eur J Med Chem 2018, 143, 1277–1300. [Google Scholar] [CrossRef]
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674. [Google Scholar] [CrossRef] [PubMed]
- Doostmohammadi, A.; Jooya, H.; Ghorbanian, K.; Gohari, S.; Dadashpour, M. Potentials and Future Perspectives of Multi-Target Drugs in Cancer Treatment: The next Generation Anti-Cancer Agents. Cell Commun. Signal. 2024, 22, 228. [Google Scholar] [CrossRef] [PubMed]
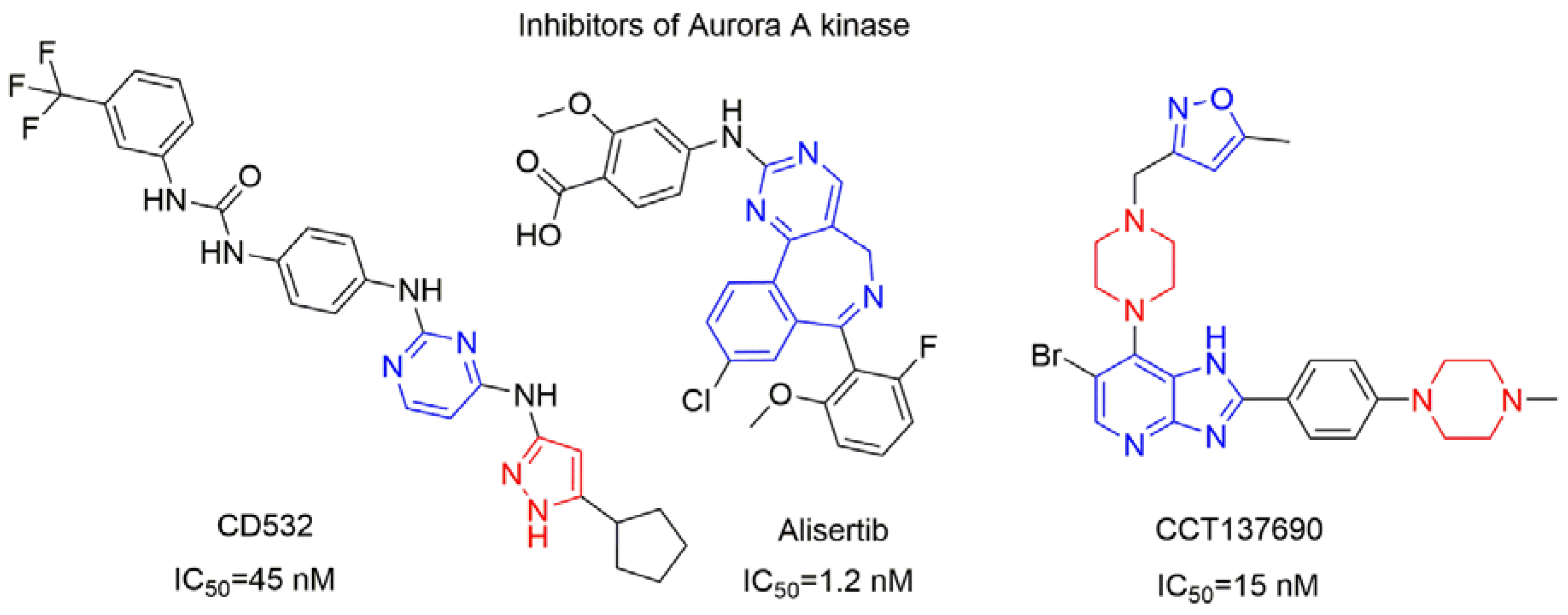
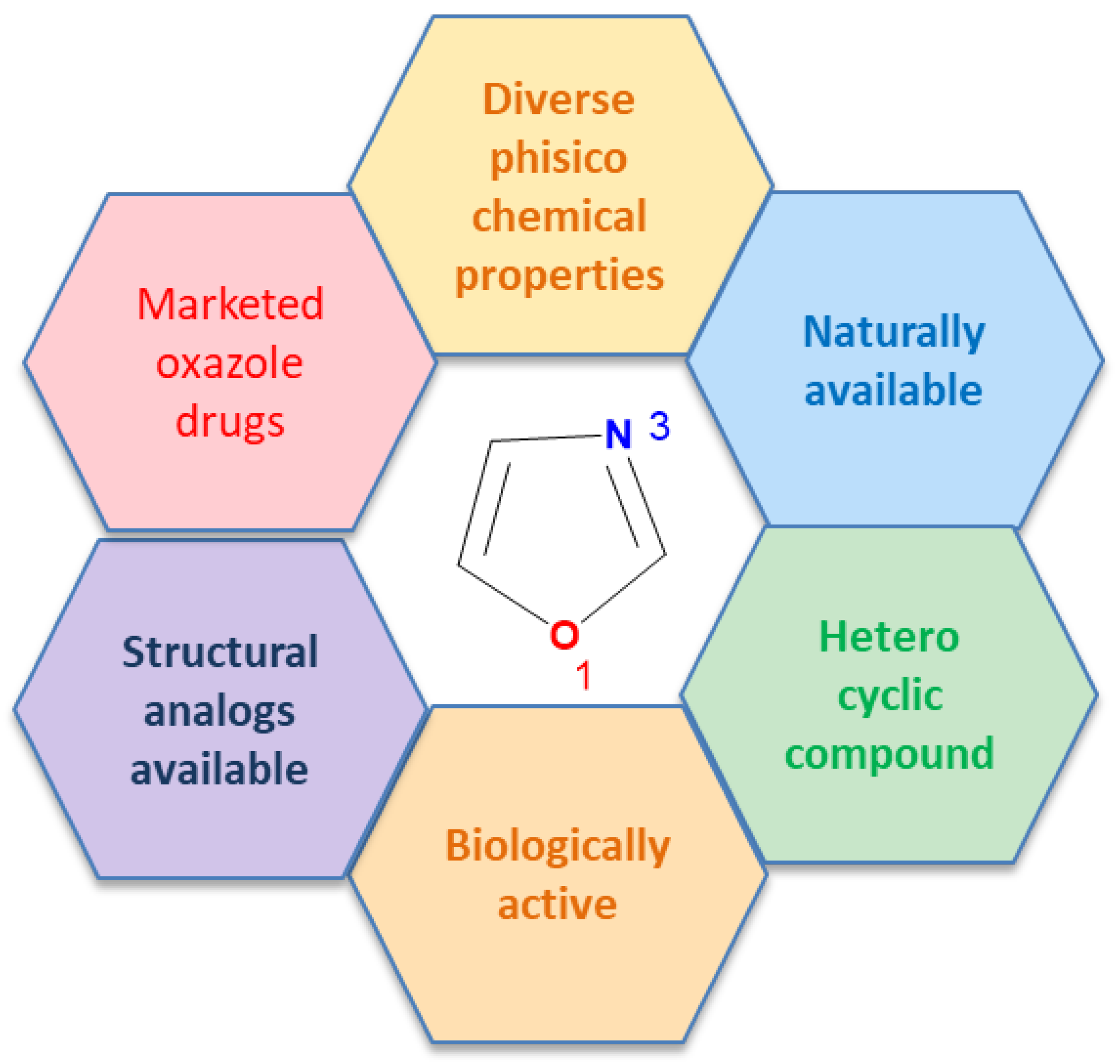
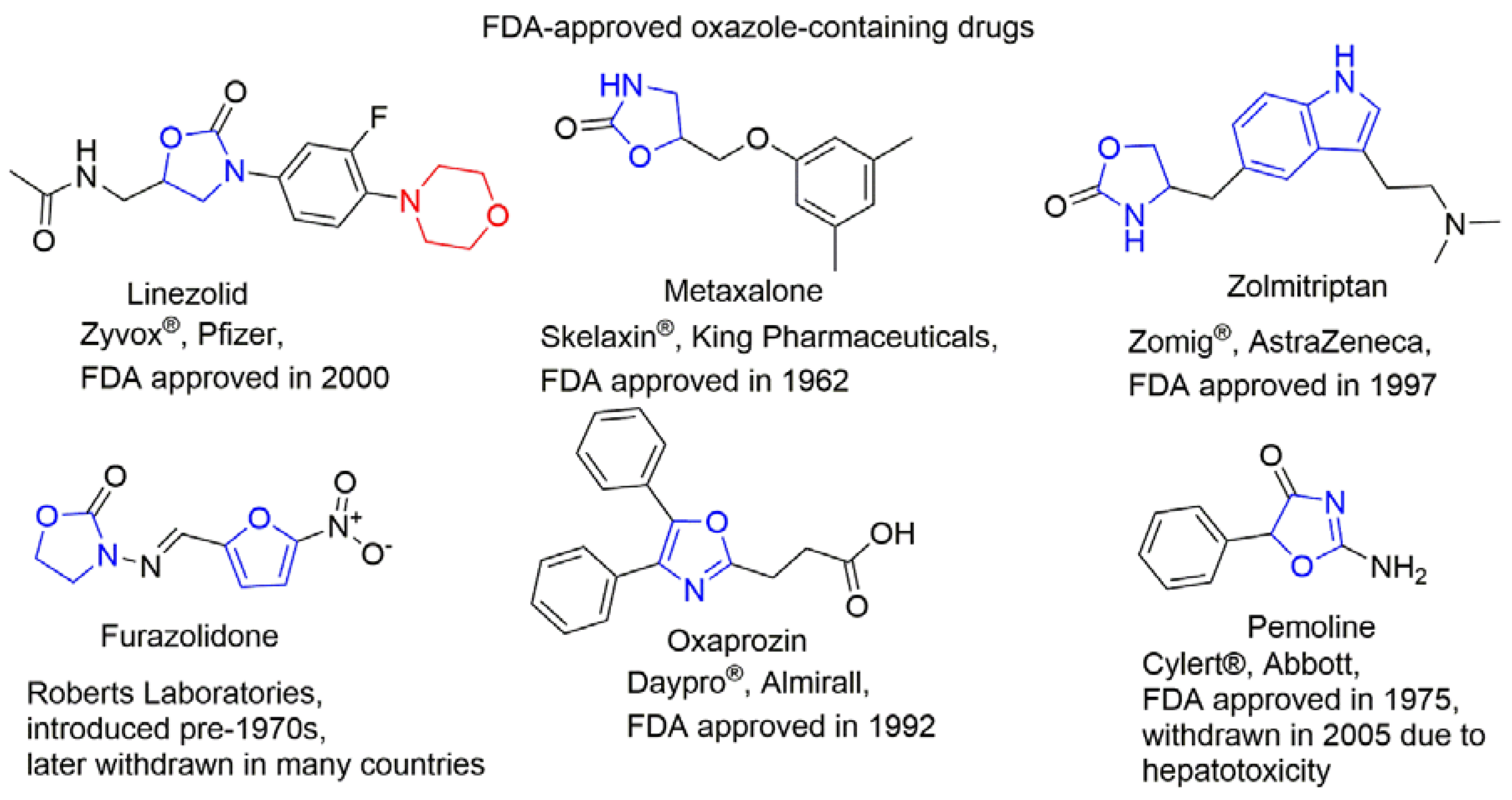

| Compounds and ligands |
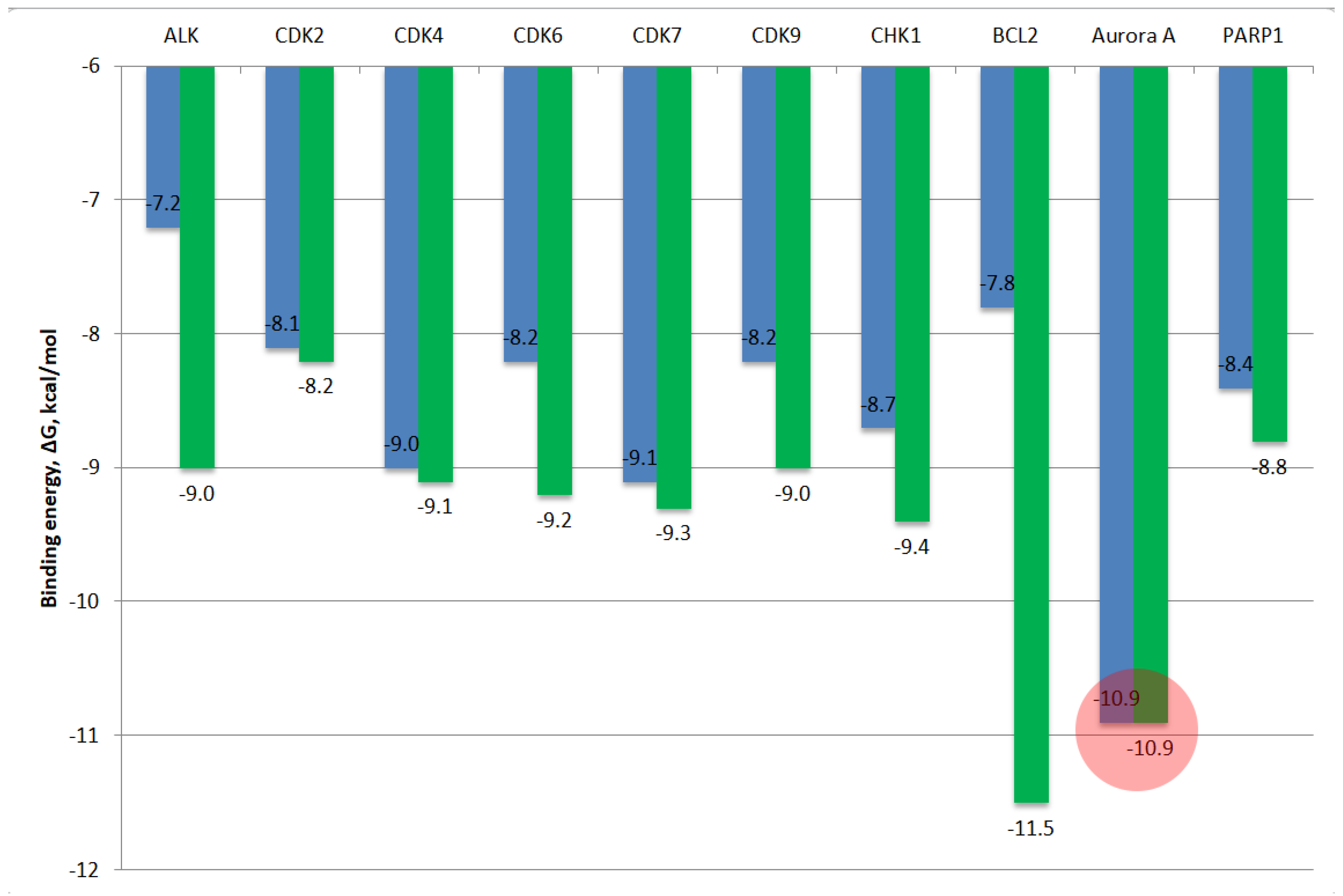
Binding energy, ∆G, kcal/mol | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| ALK | CDK2 | CDK4 | CDK6 | CDK7 | CDK9 | CHK1 | BCL2 | Aurora A/ N-MYC |
PARP1 | |
| 7a | – 7.1 | – 7.9 | – 9.0 | – 8.0 | – 8.5 | – 8.0 | – 8.9 | – 7.4 | – 10.8 | – 8.1 |
| 7b | – 7.2 | – 8.1 | – 9.0 | – 8.2 | – 9.1 | – 8.2 | – 8.7 | – 7.8 | – 10.9 | – 8.4 |
| 8aa | – 7.5 | – 8.0 | – 9.1 | – 8.1 | – 9.0 | – 8.7 | – 9.1 | – 8.0 | – 10.8 | – 8.3 |
| Crizotinib | – 9.0 | – | – | – | – | – | – | – | – | – |
| CID 57519664a | – | – 8.2 | – | – | – | – | – | – | – | – |
| Abemaciclib |
– | – | – 9.1 | – | – | – | – | – | – | – |
| CID 169552807b | – | – | – | – 9.2 | – | – | – | – | – | – |
| ATPc | – | – | – | – | – 9.3 | – | – | – | – | – |
| CID 124155204d | – | – | – | – | – | – 9.0 | – | – | – | – |
| CID 6914568e | – | – | – | – | – | – | – 9.4 | – | – | – |
| Navitoclax | – | – | – | – | – | – | – | – 11.5 | – | – |
| ADPf | – | – | – | – | – | – | – | – | – 10.9 | – |
| CID 49873226g | – | – | – | – | – | – | – | – | – | – 8.8 |
| Parameter | Compounds | |||
|---|---|---|---|---|
| 7a | 7b | 8aa | Doxorubicin | |
| Physicochemical properties | ||||
| Molecular weight, g/mol | 318.35 | 332.40 | 410.48 | 543.525 |
| Rotatable bond count | 3 | 3 | 4 | 5 |
| Hydrogen bond acceptor count | 7 | 7 | 9 | 12 |
| Hydrogen bond donor count | 1 | 1 | 0 | 7 |
| Surface area, A2 a | 127.784 | 134.149 | 157.485 | 222.081 |
| logP | 1.424 | 2.013 | 2.048 | 1.208 |
| Water solubility, log mol/L | -3.015 | -2.879 | -4.146 | -2.915 |
| Absorption | ||||
| Caco-2 permeability, log cm/s |
-5.855 | -5.47 | -4.851 | -6.77 |
| Inhibitor of P-glycoprotein | No | No | Yes | No |
| Substrate of P-glycoprotein | No | No | No | Yes |
| Distribution | ||||
| BBB permeability a | -0.822 | -0.87 | -1.474 | -1.379 |
| CNS permeability a | -3.056 | -2.621 | -2.908 | -2.846 |
| Metabolism | ||||
| CYP2D6 substrate | No | No | No | No |
| CYP3A4 substrate | Yes | Yes | Yes | No |
| CYP1A2 inhibitor | No | No | No | No |
| CYP2C19 substrate | Yes | Yes | No | No |
| CYP2C19 inhibitor | Yes | No | Yes | No |
| CYP2C9 inhibitor | No | No | Yes | No |
| CYP2D6 inhibitor | No | No | No | No |
| CYP3A4 inhibitor | Yes | Yes | Yes | No |
| Excretion | ||||
| Plasma clearance, ml/min/kg | 6.379 | 6.382 | 5.762 | 14.244 |
| Half-life of the drug, hour | 0.821 | 0.686 | 0.793 | 3.774 |
| Toxicity | ||||
| Rat Oral Acute Toxicity (LD50), mol/kg a | 2.456 | 2.471 | 2.491 | 2.408 |
| Human Hepatotoxicity a | Yes | Yes | Yes | Yes |
| Max. tolerated dose (human), log mg/kg/day a | -0.158 | -0.554 | -0.429 | 0.081 |
| a predicted using pkCSM web server. | ||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




