Submitted:
10 December 2025
Posted:
11 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.1. QC, Read Alignment, and Hybrid Assembly
2.2. Single Nucleotide Polymorphisms
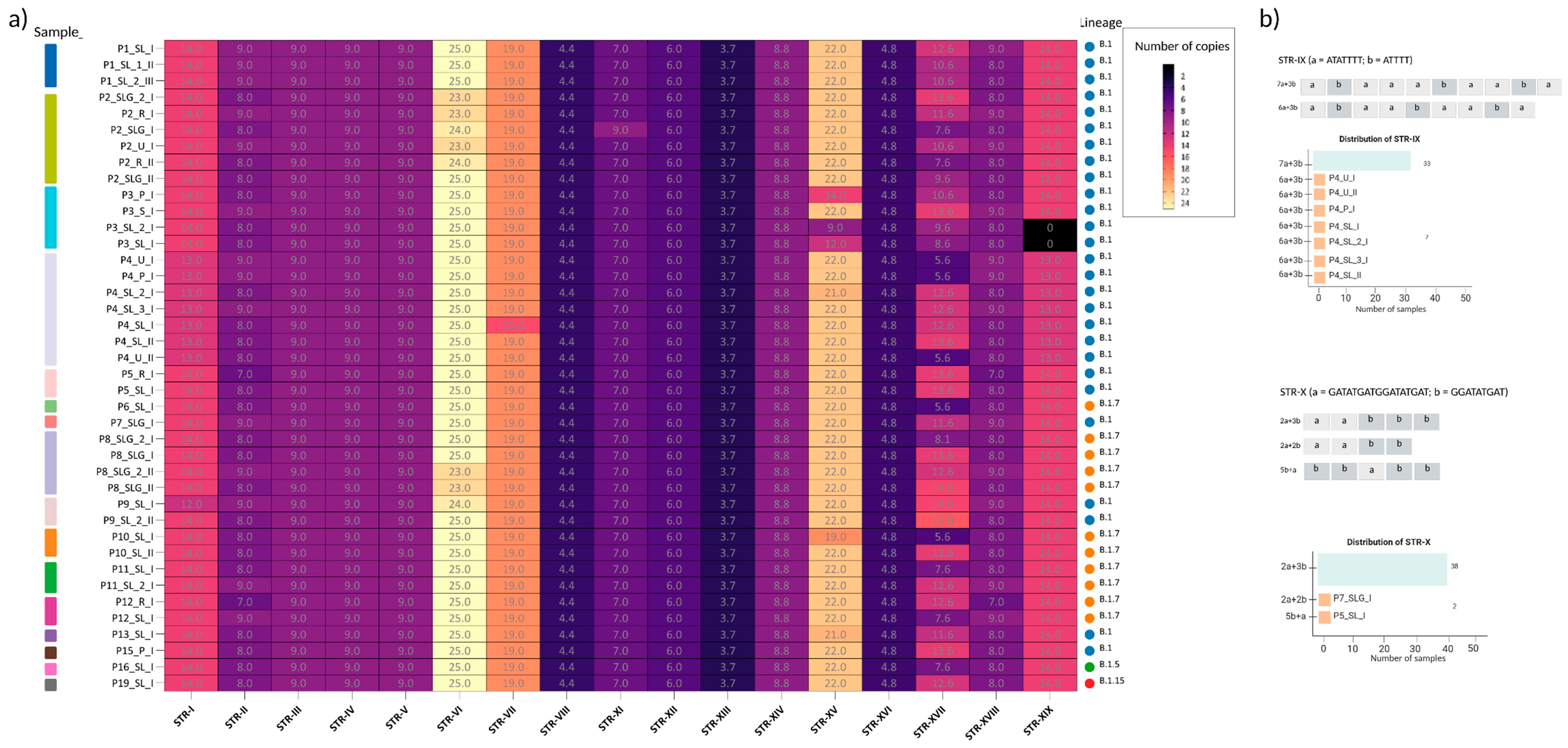
2.3. Intra-Host STRs Variation Across Matrices and Timepoints
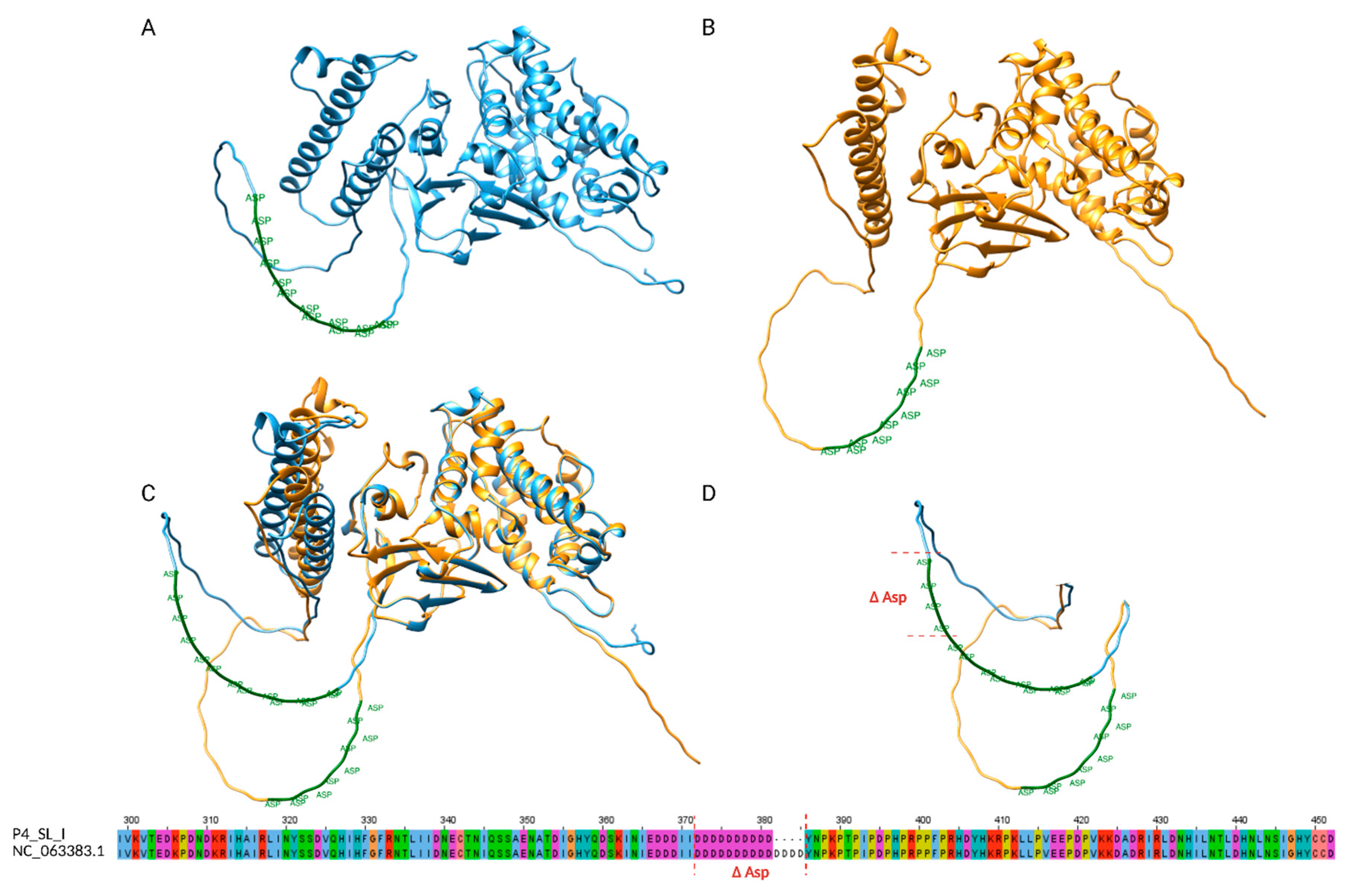
2.4. STRs and in Silico Protein Implications
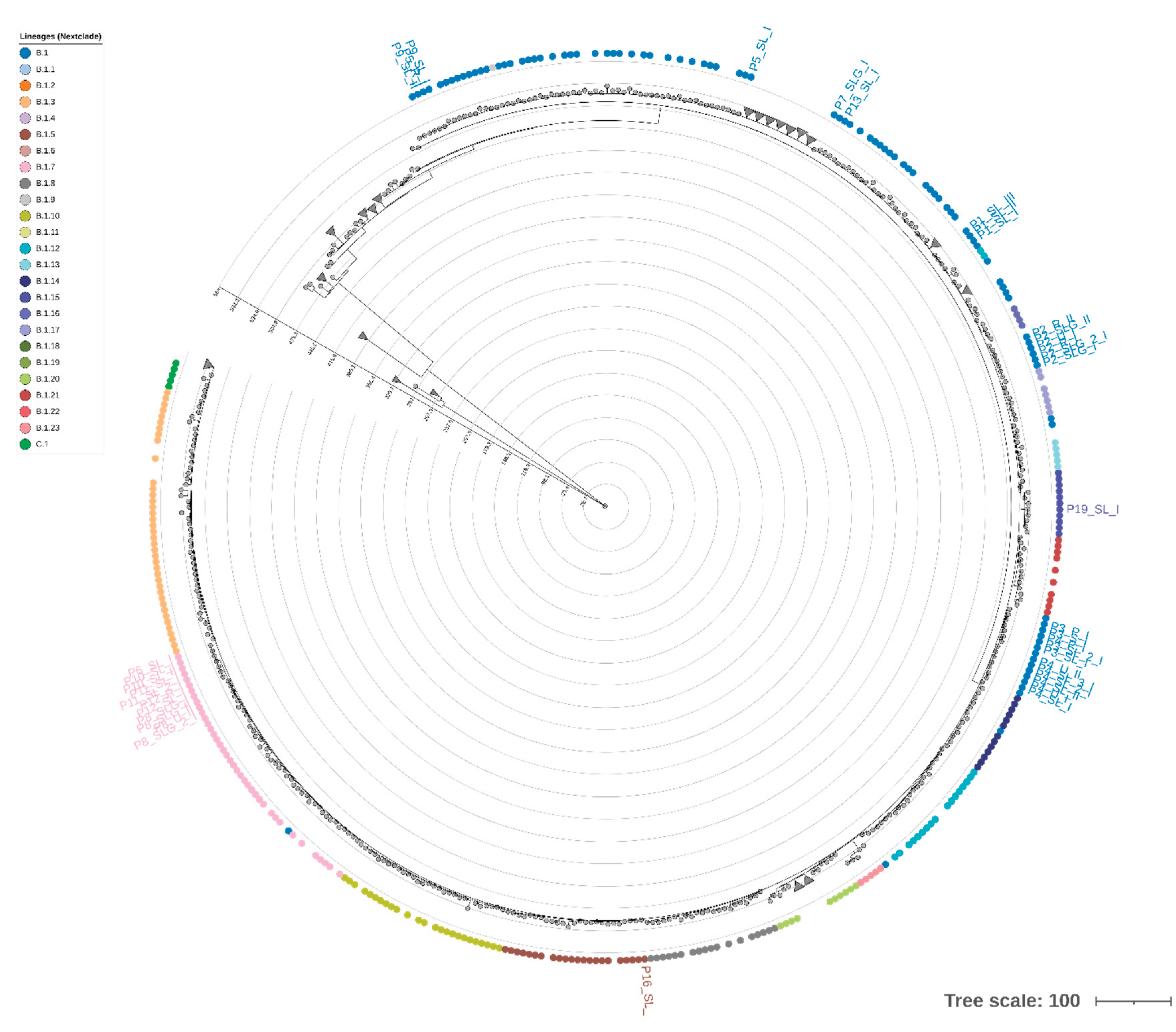
2.5. Phylogenetic Analysis and STR-Based Clustering
3. Discussion
4. Materials and Methods
4.1. DNA Extraction, Library Preparation, and Sequencing
4.2. Bioinformatic Analysis
4.2.1. Hybrid Assembly
4.2.2. STR Analysis
4.2.3. Global Dataset, Phylogeny and STR-Based Clustering Analysis
4.2.4. Protein Translation and in Silico Structural Prediction
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Reynolds, M.G.; Carroll, D.S.; Karem, K.L. Factors Affecting the Likelihood of Monkeypox’s Emergence and Spread in the Post-Smallpox Era. Curr Opin Virol. 2012, 2, 335–43. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Lloyd-Smith, J.O. Vacated Niches, Competitive Release and the Community Ecology of Pathogen Eradication. Philos Trans R Soc Lond B Biol Sci. 2013, 368, 20120150. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- World Health Organization Global Commission for the Certification of Smallpox Eradication. The Achievement of Global Eradication of Smallpox: Final Report of the Global Commission for the Certification of Smallpox Eradication; WHO: Geneva; Geneva, 1979. [Google Scholar]
- Naga, N.G.; Nawar, E.A.; Mobarak, A.A.; Faramawy, A.G.; Al-Kordy, H.M.H. Monkeypox: A Re-Emergent Virus with Global Health Implications - a Comprehensive Review. Trop Dis Travel Med Vaccines 2025, 11, 2. [Google Scholar] [CrossRef]
- 2022-25 Mpox Outbreak: Global Trends. World Health Organization: Geneva, 2025. Available online: Https://Worldhealthorg.Shinyapps.Io/Mpx_global/.
- Isidro, J.; Borges, V.; Pinto, M.; Sobral, D.; Santos, J.D.; Nunes, A.; Mixão, V.; Ferreira, R.; Santos, D.; Duarte, S.; et al. Phylogenomic Characterization and Signs of Microevolution in the 2022 Multi-Country Outbreak of Monkeypox Virus. Nature Medicine 2022, 28, 1569–1572. [Google Scholar] [CrossRef]
- Shchelkunov, SN; Totmenin, AV; Safronov, PF; Mikheev, MV; Gutorov, VV; Ryazankina, OI; Petrov, NA; Babkin, IV; Uvarova, EA; Sandakhchiev, LS; Sisler, JR; Esposito, JJ; Damon, IK; Jahrling, PB. Moss B Analysis of the Monkeypox Virus Genome Analysis of the Monkeypox Virus Genome. Virology;Virology 2002, 297 297, 172–94 172–194. [Google Scholar] [PubMed] [PubMed Central]
- Alcami, A.; Koszinowski, U.H. Viral Mechanisms of Immune Evasion. Trends in Microbiology 2000, 8, 410–418. [Google Scholar] [CrossRef] [PubMed]
- Moss, B.; Shisler, J.L. Immunology 101 at Poxvirus U: Immune Evasion Genes. Seminars in Immunology 2001, 13, 59–66. [Google Scholar] [CrossRef]
- Likos, A.M.; et al. A Tale of Two Clades: Monkeypox Viruses. Journal of General Virology 2005, 86, 2661–2672. [Google Scholar] [CrossRef]
- Doty, J.B.; et al. Assessing Monkeypox Virus Prevalence in Small Mammals at the Human–Animal Interface in the Democratic Republic of the Congo. Viruses 2017, 9, 283. [Google Scholar] [CrossRef]
- Nakazawa, Y.; et al. A Phylogeographic Investigation of African Monkeypox. Viruses 2015, 7, 2168–2184. [Google Scholar] [CrossRef]
- WHO Monkeypox Fact Sheet. Available online: https://www.who.int/news-room/fact-sheets/detail/monkeypox.
- Esteban, D.J.; Hutchinson, A.P. Genes in the Terminal Regions of Orthopoxvirus Genomes Experience Adaptive Molecular Evolution. BMC Genomics 2011, 12, 261. [Google Scholar] [CrossRef]
- Hughes, A.L.; Friedman, R. Poxvirus Genome Evolution by Gene Gain and Loss. Molecular Phylogenetics and Evolution 2005, 35, 186–195. [Google Scholar] [CrossRef]
- Kugelman, J.R. Genomic Variability and Tandem Repeats in Variola Virus and Monkeypox Virus. Emerging Infectious Diseases 2014, 20, 232–239. [Google Scholar]
- Elde, N.C.; Child, S.J.; Eickbush, M.T.; Kitzman, J.O.; Rogers, K.S.; Shendure, J.; Geballe, A.P.; Malik, H.S. Poxviruses Deploy Genomic Accordions to Adapt Rapidly against Host Antiviral Defenses. Cell 2012, 150, 831–841. [Google Scholar] [CrossRef] [PubMed]
- Yeh, T.-Y.; et al. Recombination Shapes the 2022 Monkeypox (Mpox) Outbreak. Med 2022, 3, 824–826. [Google Scholar] [CrossRef]
- Kumari, N.; Yadav, V.K.; Singh, H. Identification and Analysis of Microsatellites in Coronaviridae. Virus Research 2024, 336, 199328. [Google Scholar] [CrossRef]
- Katoh, H.; Zheng, H.; Ono, Y. others Genome-Wide Identification and Characterization of Simple Sequence Repeats in Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Scientific Reports 2020, 10, 15354. [Google Scholar] [CrossRef]
- Méndez-Tenorio, F.J.; Rojas-López, A.; Arias, C.F. Microsatellite Repeats in the Genome of Caliciviruses. Journal of Virology 2002, 76, 10638–10643. [Google Scholar] [CrossRef]
- Ding, Y.; Zhang, J.; Liu, H. others Characterization of Microsatellites in Herpes Simplex Virus Genomes. Frontiers in Microbiology 2015, 6, 1462. [Google Scholar] [CrossRef]
- Cheng, S.; Zhang, Y.; Zhao, X. others Characterization of Microsatellites in the Orf Virus Genome and Their Potential Role in Viral Evolution. Infection, Genetics and Evolution 2021, 90, 104752. [Google Scholar] [CrossRef]
- Monzón, S.; Varona, S.; Negredo, A.; Vidal-Freire, S.; Patiño-Galindo, J.A.; Ferressini-Gerpe, N.; Zaballos, A.; Orviz, E.; Ayerdi, O.; Muñoz-Gómez, A.; et al. Monkeypox Virus Genomic Accordion Strategies. Nature Communications 2024, 15, 3059. [Google Scholar] [CrossRef] [PubMed]
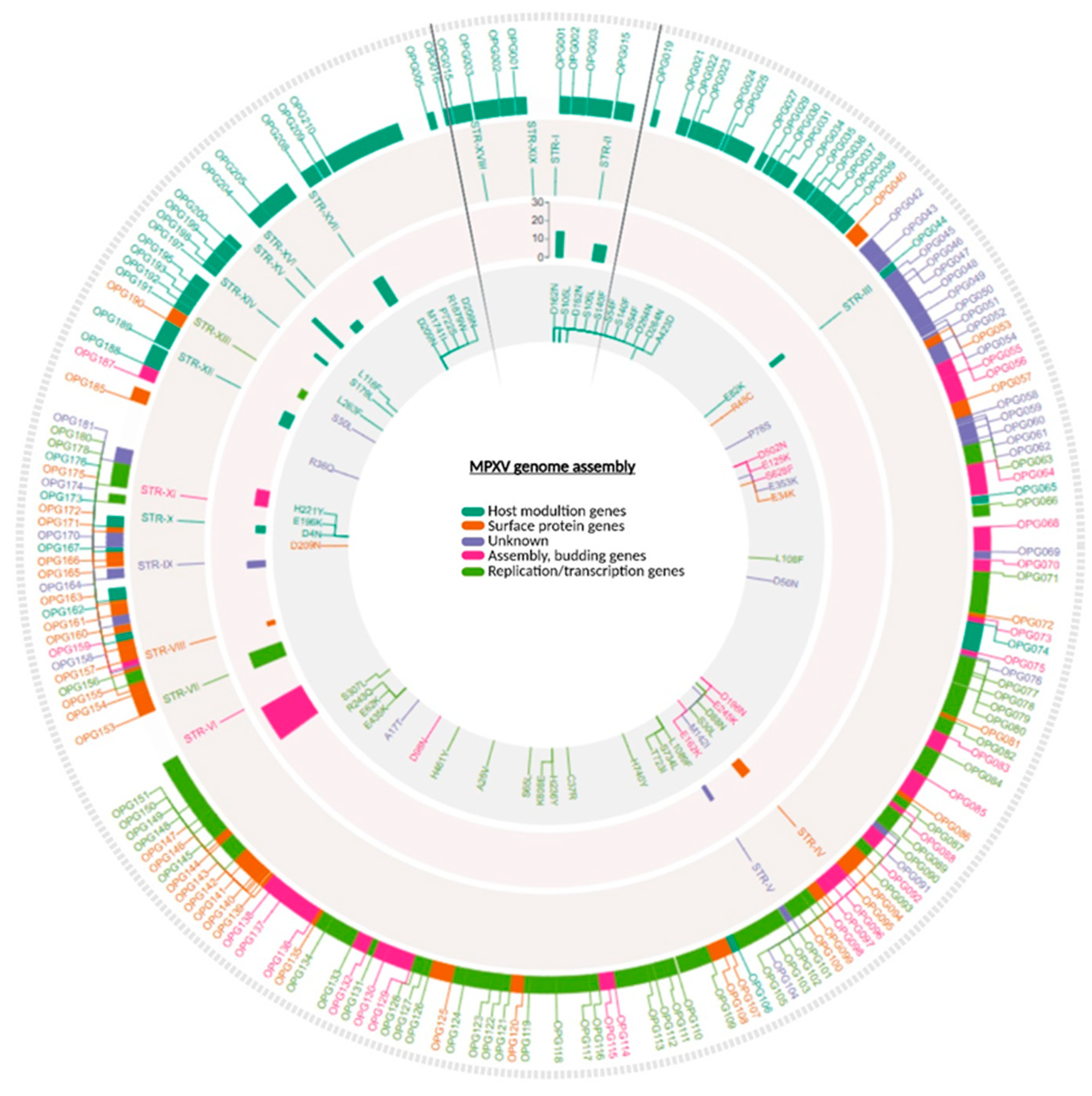
| NAME | CORRESPONDENCE WITH MONZòN ET AL. | START | END | PATTERN | # MAX REP | STR LENGTH (BP) | # SAMPLES SHARING THE STR | NEAREST GENE | RELATIVE POSITION TO THE GENE | DISTANCE IN BP | FUNCTIONAL INSIGHT |
|---|---|---|---|---|---|---|---|---|---|---|---|
| STR-I | LCR10 | 595 | 611 | [T] | 14 | 14 | 40/40 | OPG001 | Upstream | -224 | Chemokine-binding protein |
| STR-II | LCR1* | 4677 | 4801 | [TAACTAACTTATGACT] | 9 | 144 | 40/40 | OPG015 | Downstream | 37 | Ankyrin domains & immune evasion (TNF mimic) |
| STR-III | LCR12 | 28527 | 28537 | [A] | 9 | 9 | 40/40 | OPG044 | Intragene | NA | Bcl-2-like protein |
| STR-IV | LCR13 | 76082 | 76090 | [T] | 9 | 9 | 40/40 | OPG097 | Intragene | NA | Unknown |
| STR-V | LCR14 | 80844 | 80825 | [T] | 9 | 9 | 40/40 | OPG104 | Intragene | NA | Unknown |
| STR-VI | LCR5 | 133074 | 133098 | [T] | 25 | 25 | 40/40 | OPG151 | Upstream | -657 | Unknown |
| STR-VII | LCR7 | 136498 | 136555 | [ATC]n + TATGAT + [ATC]n | 19 | 57 | 40/40 | OPG153 | Intragene | NA | Key attachment/egress protein |
| STR-VIII | LCR15 | 140092 | 140131 | [ATAACAATT] | 4.4 | 39 | 40/40 | OPG159 | Intragene | NA | Unknown |
| STR-IX | LCR8 | 146835 | 146920 | [ATATTTT]n + [ATTTT]n | 10 | 64 | 40/40 | OPG170 | Upstream | -57 | Unknown |
| STR-X | LCR9* | 150544 | 150621 | [GATATGATGGATATGAT]n + [GGATATGAT]n | 5 | 62 | 40/40 | OPG176 | Intragene | NA | Bcl-2-like protein |
| STR-XI | LCR16 | 152661 | 152566 | [A] | 7 | 9 | 40/40 | OPG180 | Upstream | -31 | Unknown |
| STR-XII | LCR17 | 163169 | 163207 | [TAAC] | 6 | 24 | 40/40 | OPG188 | Upstream | -63 | Unknown |
| STR-XIII | LCR18 | 166070 | 166109 | [AATAATT] | 3.7 | 39 | 40/40 | OPG190 | Intragene | NA | IFNα/β binding protein homolog |
| STR-XIV | LCR19* | 169709 | 169761 | [CAGATA] | 8.8 | 52 | 40/40 | OPG197 | Intragene | NA | Unknown |
| STR-XV | LCR2* | 173252 | 173295 | [AT] | 22 | 43 | 40/40 | OPG200 | Upstream | -654 | Unknown |
| STR-XVI | LCR21 | 174506 | 174538 | [GATGAA] | 4.8 | 32 | 40/40 | OPG204 | Intragene | NA | Interferon decoy receptor; immune modulation |
| STR-XVII | LCR3* | 179055 | 179245 | [ATATACATT] | 16 | 144 | 40/40 | OPG208 | Downstream | 34 | Serine protease inhibitor (anti-apoptotic, potential virulence) |
| STR-XVIII | LCR4 | 192382 | 192534 | [AGTCATAAGTTAGTTA] | 9 | 144 | 40/40 | OPG015 | Upstream | -16 | Ankyrin domains & immune evasion (TNF mimic) |
| STR-XIX | LCR11* | 196608 | 196624 | [A] | 14 | 14 | 38/40 | OPG001 | Downstream | 233 | Chemokine-binding protein |
| PATIENT_ID | MATRIX | TIMEPOINT | SAMPLE_ID | COLLECTION_DATE | TRAVEL_HISTORY | QPCR |
|---|---|---|---|---|---|---|
| P1 | Skin lesion swab | I | P1_SL_I | 25/05/2022 | Mallorca | 21,00 |
| P1 | Skin lesion swab | II | P1_SL_1_II | 03/06/2022 | Mallorca | 21,51 |
| P1 | Skin lesion swab | III | P1_SL_2_III | 07/06/2022 | Mallorca | 21,00 |
| P2 | Skin lesion swab (genital) | I | P2_SLG_I | 18/07/2022 | Spain | 18.96 |
| P2 | urine | I | P2_U_I | 18/07/2022 | Spain | 25,91 |
| P2 | Rectal swab | I | P2_R_I | 18/07/2022 | Spain | 18,52 |
| P2 | Skin lesion swab (genital) | I | P2_SLG_2_I | 18/07/2022 | Spain | 17,07 |
| P2 | Skin lesion swab (genital) | II | P2_SLG_II | 21/07/2022 | Spain | 20.96 |
| P2 | Rectal swab | II | P2_R_II | 21/07/2022 | Spain | 20,63 |
| P3 | Skin lesion swab | I | P3_SL_I | 18/10/2022 | None | 14,70 |
| P3 | Pharyngeal swab | I | P3_P_I | 18/10/2022 | None | 16,01 |
| P3 | Saliva | I | P3_S_I | 18/10/2022 | None | 20,06 |
| P3 | Skin lesion swab (head) | I | P3_SL_2_I | 18/10/2022 | None | 18,99 |
| P4 | Skin lesion swab (abdomen) | I | P4_SL_I | 02/11/2022 | Bosnia-Herzegovina | 16,70 |
| P4 | Skin lesion swab (hand) | I | P4_SL_2_I | 02/11/2022 | Bosnia-Herzegovina | 14,40 |
| P4 | Skin lesion swab (head) | I | P4_SL_3_I | 02/11/2022 | Bosnia-Herzegovina | 14,00 |
| P4 | Pharyngeal swab | I | P4_P_I | 02/11/2022 | Bosnia-Herzegovina | 23,10 |
| P4 | Urine | I | P4_U_I | 02/11/2022 | Bosnia-Herzegovina | 25,80 |
| P4 | Skin lesion swab (leg) | II | P4_SL_II | 02/11/2022 | Bosnia-Herzegovina | 25,30 |
| P4 | Urine | II | P4_U_II | 02/11/2022 | Bosnia-Herzegovina | 25,76 |
| P5 | Skin lesion swab | I | P5_SL_I | 23/06/2022 | Spain | 22,19 |
| P5 | Rectal swab | I | P5_R_I | 23/06/2022 | Spain | 18,78 |
| P6 | Skin lesion swab | I | P6_SL_I | 21/07/2022 | None | 26.65 |
| P7 | Skin lesion swab (genital) | I | P7_SLG_I | 01/07/2022 | None | 21,02 |
| P8 | Skin lesion swab (genital) | I | P8_SLG_I | 15/07/2022 | None | 19,83 |
| P8 | Skin lesion swab (genital) | I | P8_SLG_2_I | 15/07/2022 | None | 15,05 |
| P8 | Skin lesion swab (genital) | II | P8_SLG_II | 21/07/2022 | None | 19,07 |
| P8 | Urine | II | P8_SLG_2_II | 21/07/2022 | None | 25,10 |
| P9 | Skin lesion swab | I | P9_SL_I | 01/07/2022 | USA | 26,67 |
| P9 | Skin lesion swab | II | P9_SL_2_II | 01/07/2022 | USA | 22,74 |
| P10 | Skin lesion swab | I | P10_SL_I | 26/07/2022 | None | 24,05 |
| P10 | Skin lesion swab | II | P10_SL_II | 29/07/2022 | None | 19,05 |
| P11 | Skin lesion swab (abdomen) | I | P11_SL_I | 05/08/2022 | None | 20.17 |
| P11 | Skin lesion swab (arm) | I | P11_SL_2_I | 05/08/2022 | None | 19,32 |
| P12 | Skin lesion swab (head) | I | P12_SL_I | 18/08/2022 | None | 18.06 |
| P12 | Rectal swab | I | P12_R_I | 18/08/2022 | None | 25,25 |
| P13 | Skin lesion swab | I | P13_SL_I | 19/09/2022 | None | 21,31 |
| P15 | Pharyngeal swab | I | P15_P_I | 08/07/2022 | None | 27,03 |
| P16 | Skin lesion swab | I | P16_SL_I | NA | NA | NA |
| P19 | Skin lesion swab | I | P19_SL_I | NA | NA | NA |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




