Submitted:
03 December 2025
Posted:
04 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Databases and Tools
2.2. Extraction and Quantification of Flavins
3. Results and Discussion
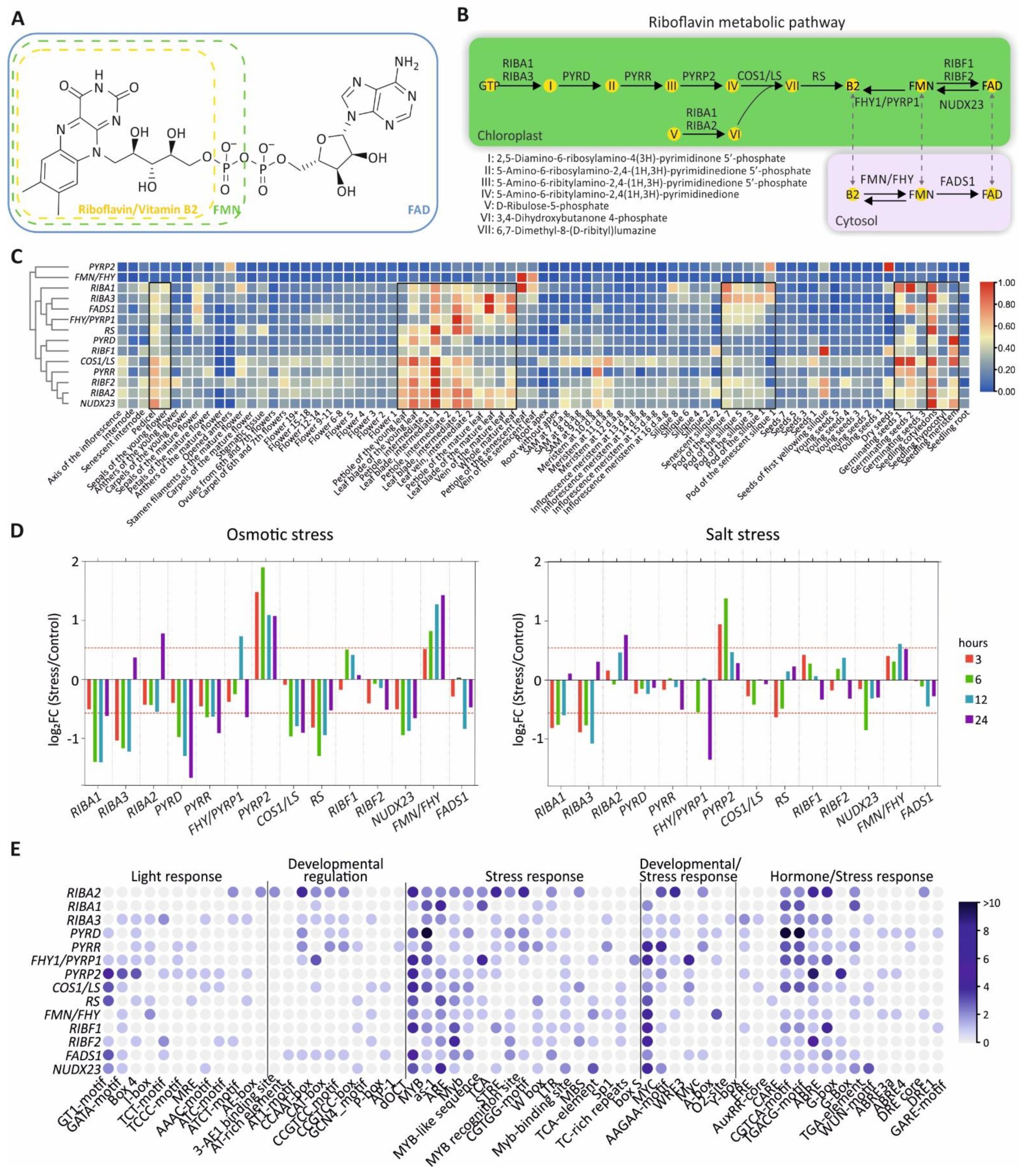
3.1. Riboflavin Metabolic Pathway
3.2. Spatiotemporal Expression of Riboflavin Metabolic Pathway Genes
3.3. Response of Riboflavin Metabolism Genes Under Under Osmotic and Salt Stress
3.4. Cis-Element Analysis of Riboflavin Pathway Gene Promoters
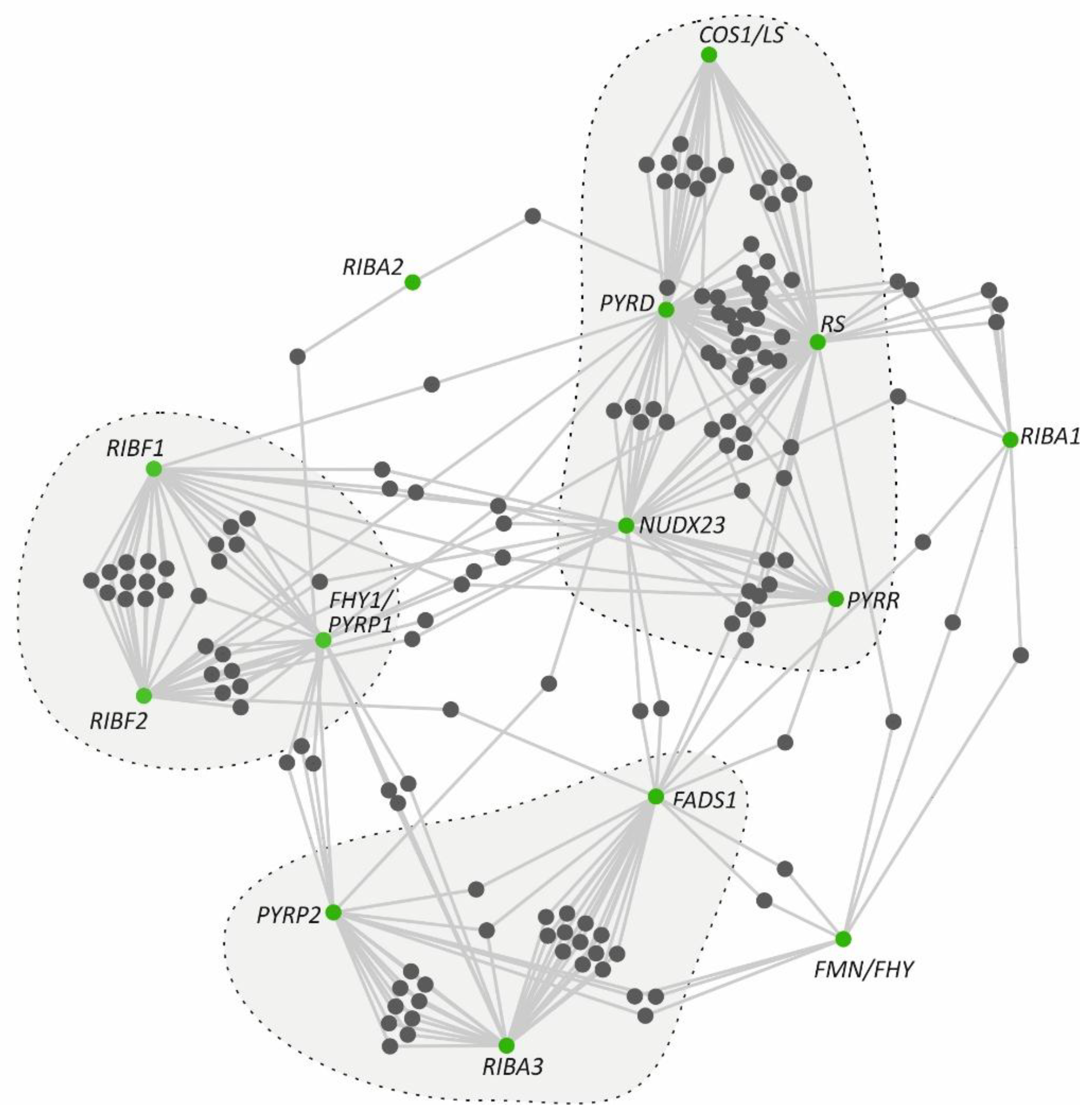
3.5. Co-Expression Hubs of Riboflavin Metabolism Pathway Genes
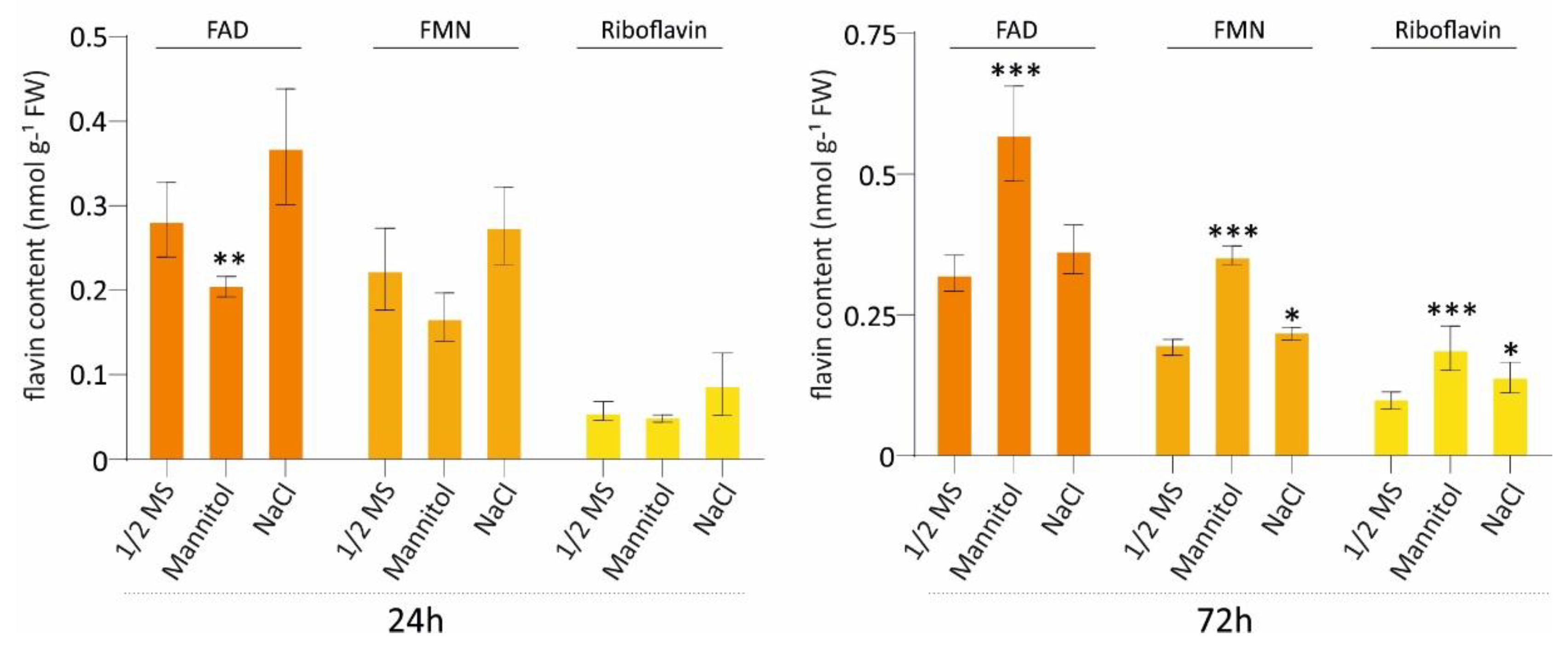
3.6. Flavin Dynamics Under Osmotic and Salt Stress Conditions
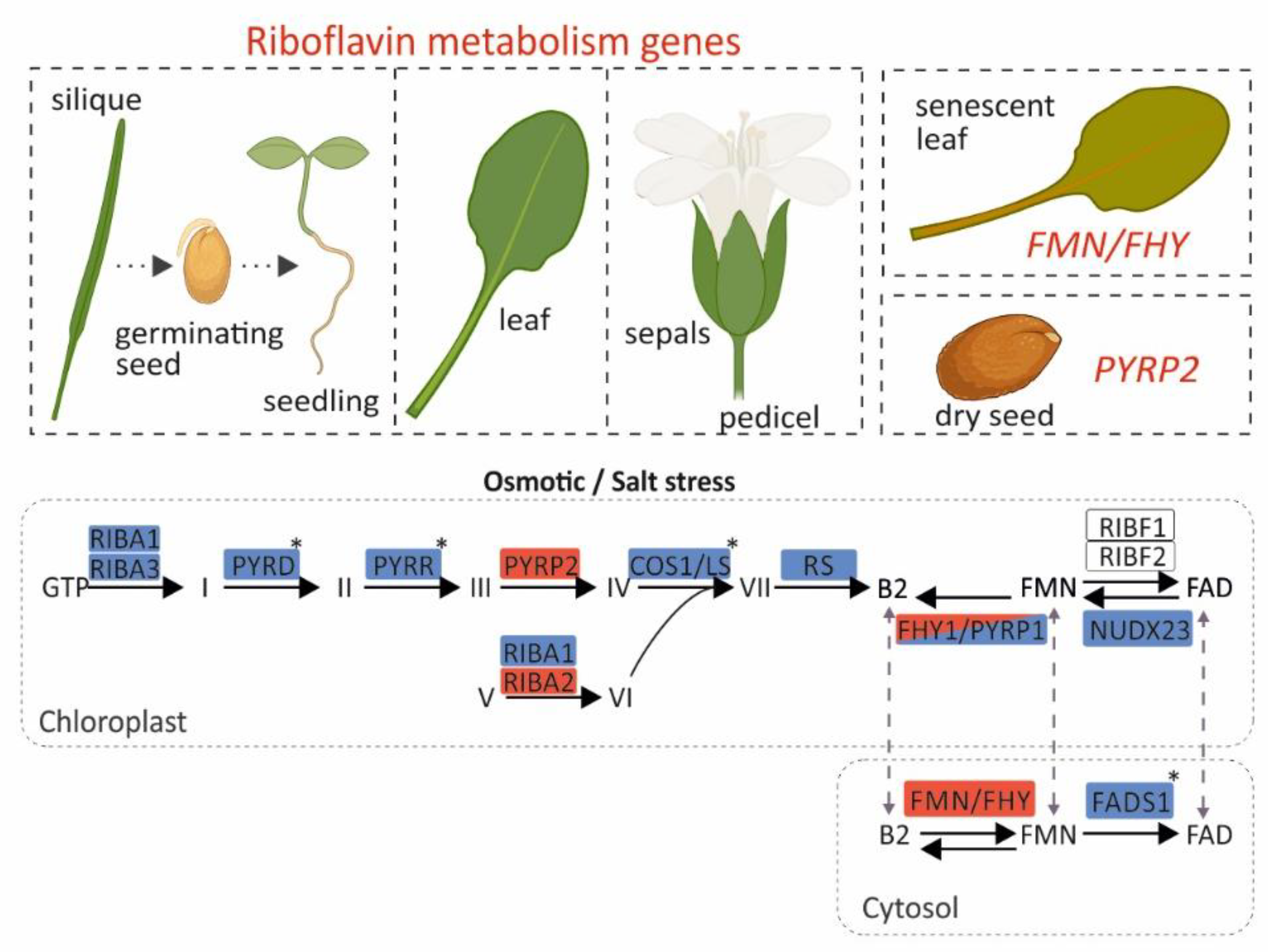
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Eggers, R.; Jammer, A.; Jha, S.; Kerschbaumer, B.; Lahham, M.; Strandback, E.; Toplak, M.; Wallner, S.; Winkler, A.; Macheroux, P. The Scope of Flavin-Dependent Reactions and Processes in Arabidopsis thaliana. Phytochemistry 2021, 189, 112822. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.G.; Croft, M.T.; Moulin, M.; Webb, M.E. Plants Need Their Vitamins Too. Curr. Opin. Plant Biol. 2007, 10, 266–275. [Google Scholar] [CrossRef]
- Yang, Y.; Saand, M.A.; Huang, L.; Abdelaal, W.B.; Zhang, J.; Wu, Y. Applications of Multi-Omics Technologies for Crop Improvement. Front. Plant Sci. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Gerdes, S.; Lerma-Ortiz, C.; Frelin, O.; Seaver, S.; Henry, C.S.; de Crécy-Lagard, V.; Hanson, A.D. Plant B Vitamin Pathways and Their Compartmentation: A Guide for the Perplexed. J. Exp. Bot. 2012, 63, 5379–5395. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Sato, Y.; Matsuura, Y.; Ishiguro-Watanabe, M. KEGG: Biological Systems Database as a Model of the Real World. Nucleic Acids Res. 2025, 53, D672–D677. [Google Scholar] [CrossRef] [PubMed]
- Klepikova, A.V.; Kasianov, A.S.; Gerasimov, E.S.; Logacheva, M.D.; Penin, A.A. A High Resolution Map of the Arabidopsis thaliana Developmental Transcriptome Based on RNA-Seq Profiling. Plant J. 2016, 88, 1058–1070. [Google Scholar] [CrossRef]
- Mergner, J.; Frejno, M.; List, M.; Papacek, M.; Chen, X.; Chaudhary, A.; Samaras, P.; Richter, S.; Shikata, H.; Messerer, M.; Lang, D. Mass-Spectrometry-Based Draft of the Arabidopsis Proteome. Nature 2020, 579, 409–414. [Google Scholar] [CrossRef]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE: A Database of Plant cis-Acting Regulatory Elements and a Portal to Tools for In Silico Analysis of Promoter Sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef]
- Waese, J.; Fan, J.; Pasha, A.; Yu, H.; Fucile, G.; Shi, R.; Cumming, M.; Kelley, L.A.; Sternberg, M.J.; Krishnakumar, V.; et al. ePlant: Visualizing and Exploring Multiple Levels of Data for Hypothesis Generation in Plant Biology. Plant Cell 2017, 29, 1806–1821. [Google Scholar] [CrossRef]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; et al. TBtools-II: A “One for All, All for One” Bioinformatics Platform for Biological Big-Data Mining. Mol. Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef]
- Hiltunen, H.M.; Illarionov, B.; Hedtke, B.; Fischer, M.; Grimm, B. Arabidopsis RIBA Proteins: Two out of Three Isoforms Have Lost Their Bifunctional Activity in Riboflavin Biosynthesis. Int. J. Mol. Sci. 2012, 13, 14086–14105. [Google Scholar] [CrossRef] [PubMed]
- Hasnain, G.; Frelin, O.; Roje, S.; Ellens, K.W.; Ali, K.; Guan, J.C.; Garrett, T.J.; de Crécy-Lagard, V.; Gregory, J.F., III; McCarty, D.R.; et al. Identification and Characterization of the Missing Pyrimidine Reductase in the Plant Riboflavin Biosynthesis Pathway. Plant Physiol. 2013, 161, 48–56. [Google Scholar] [CrossRef] [PubMed]
- Sa, N.; Rawat, R.; Thornburg, C.; Walker, K.D.; Roje, S. Identification and Characterization of the Missing Phosphatase on the Riboflavin Biosynthesis Pathway in Arabidopsis thaliana. Plant J. 2016, 88, 705–716. [Google Scholar] [CrossRef]
- Hu, H.; Ren, D.; Hu, J.; Jiang, H.; Chen, P.; Zeng, D.; Qian, Q.; Guo, L. WHITE AND LESION-MIMIC LEAF1, Encoding a Lumazine Synthase, Affects Reactive Oxygen Species Balance and Chloroplast Development in Rice. Plant J. 2021, 108, 1690–1703. [Google Scholar] [CrossRef]
- Sandoval, F.J.; Roje, S. An FMN Hydrolase Is Fused to a Riboflavin Kinase Homolog in Plants. J. Biol. Chem. 2005, 280, 38337–38345. [Google Scholar] [CrossRef]
- Rawat, R.; Sandoval, F.J.; Wei, Z.; Winkler, R.; Roje, S. An FMN Hydrolase of the Haloacid Dehalogenase Superfamily Is Active in Plant Chloroplasts. J. Biol. Chem. 2011, 286, 42091–42098. [Google Scholar] [CrossRef] [PubMed]
- Sandoval, F.J.; Zhang, Y.; Roje, S. Flavin Nucleotide Metabolism in Plants: Monofunctional Enzymes Synthesize FAD in Plastids. J. Biol. Chem. 2008, 283, 30890–30900. [Google Scholar] [CrossRef]
- Lynch, J.H.; Roje, S. A Higher Plant FAD Synthetase Is Fused to an Inactivated FAD Pyrophosphatase. J. Biol. Chem. 2022, 298, 12. [Google Scholar] [CrossRef]
- Maruta, T.; Yoshimoto, T.; Ito, D.; Ogawa, T.; Tamoi, M.; Yoshimura, K.; Shigeoka, S. An Arabidopsis FAD Pyrophosphohydrolase, AtNUDX23, Is Involved in Flavin Homeostasis. Plant Cell Physiol. 2012, 53, 1106–1116. [Google Scholar] [CrossRef]
- Tuan, P.A.; Zhao, S.; Kim, J.K.; Kim, Y.B.; Yang, J.; Li, C.H.; Kim, S.J.; Arasu, M.V.; Al-Dhabi, N.A.; Park, S.U. Riboflavin Accumulation and Molecular Characterization of cDNAs Encoding Riboflavin Biosynthesis Enzymes in Different Organs of Lycium chinense. Molecules 2014, 19, 17141–17153. [Google Scholar] [CrossRef]
- Jiadkong, K.; Fauzia, A.N.; Yamaguchi, N.; Ueda, A. Exogenous Riboflavin (Vitamin B2) Application Enhances Salinity Tolerance through the Activation of Its Biosynthesis in Rice Seedlings under Salinity Stress. Plant Sci. 2024, 339, 111929. [Google Scholar] [CrossRef]
- Martin, C.; Binda, C.; Fraaije, M.W.; Mattevi, A. The Multipurpose Family of Flavoprotein Oxidases. In The Enzymes; Academic Press: Cambridge, MA, USA, 2020; Volume 47, pp. 63–86. [Google Scholar]
- Shen, Q.; Ho, T.H. Functional Dissection of an Abscisic Acid (ABA)-Inducible Gene Reveals Two Independent ABA-Responsive Complexes Each Containing a G-Box and a Novel Cis-Acting Element. Plant Cell 1995, 7, 295–307. [Google Scholar]
- Garretón, V.; Carpinelli, J.; Jordana, X.; Holuigue, L. The as-1 Promoter Element Is an Oxidative Stress-Responsive Element and Salicylic Acid Activates It via Oxidative Species. Plant Physiol. 2002, 130, 1516–1526. [Google Scholar] [CrossRef]
- Namba, J.; Harada, M.; Shibata, R.; Toda, Y.; Maruta, T.; Ishikawa, T.; Shigeoka, S.; Yoshimura, K.; Ogawa, T. AtDREB2G Regulates Riboflavin Biosynthesis in Response to Low-Temperature Stress and Abscisic Acid Treatment in Arabidopsis thaliana. Plant Sci. 2024, 347, 112196. [Google Scholar] [CrossRef]
- Huang, D.; Wu, W.; Abrams, S.R.; Cutler, A.J. The Relationship of Drought-Related Gene Expression in Arabidopsis thaliana to Hormonal and Environmental Factors. J. Exp. Bot. 2008, 59, 2991–3007. [Google Scholar] [CrossRef]
- Zogopoulos, V.L.; Saxami, G.; Malatras, A.; Angelopoulou, A.; Jen, C.H.; Duddy, W.J.; Daras, G.; Hatzopoulos, P.; Westhead, D.R.; Michalopoulos, I. Arabidopsis Coexpression Tool: A Tool for Gene Coexpression Analysis in Arabidopsis thaliana. iScience 2021, 24, 8. [Google Scholar] [CrossRef]
- Obayashi, T.; Hibara, H.; Kagaya, Y.; Aoki, Y.; Kinoshita, K. ATTED-II v11: A Plant Gene Coexpression Database Using a Sample-Balancing Technique by Subagging of Principal Components. Plant Cell Physiol. 2022, 63, 869–881. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Lu, X.; Tao, Y.; Guo, H.; Min, W. Comparative Ionomics and Metabolic Responses and Adaptive Strategies of Cotton to Salt and Alkali Stress. Front. Plant Sci. 2022, 13, 871387. [Google Scholar] [CrossRef] [PubMed]
- Sebastián, M.; Velázquez-Campoy, A.; Medina, M. The RFK Catalytic Cycle of the Pathogen Streptococcus pneumoniae Shows Species-Specific Features in Prokaryotic FMN Synthesis. J. Enzyme Inhib. Med. Chem. 2018, 33, 842–849. [Google Scholar] [CrossRef] [PubMed]
- Sebastián, M.; Arilla-Luna, S.; Bellalou, J.; Yruela, I.; Medina, M. The Biosynthesis of Flavin Cofactors in Listeria monocytogenes. J. Mol. Biol. 2019, 431, 2762–2776. [Google Scholar] [CrossRef]
- Anoz-Carbonell, E.; Rivero, M.; Polo, V.; Velázquez-Campoy, A.; Medina, M. Human Riboflavin Kinase: Species-Specific Traits in the Biosynthesis of the FMN Cofactor. FASEB J. 2020, 34, 10871–10886. [Google Scholar] [CrossRef]
- Hanson, A.D.; Henry, C.S.; Fiehn, O.; de Crécy-Lagard, V. Metabolite damage and metabolite damage control in plants. Annu. Rev. Plant Biol. 2016, 67, 131–152. [Google Scholar] [CrossRef] [PubMed]
- Fitzpatrick, T.B. B Vitamins: An Update on Their Importance for Plant Homeostasis. Annu. Rev. Plant Biol. 2024, 75. [Google Scholar] [CrossRef] [PubMed]
- Rao, S.; Cao, H.; O’Hanna, F.J.; Zhou, X.; Lui, A.; Wrightstone, E.; Fish, T.; Yang, Y.; Thannhauser, T.; Cheng, K.; Dudavera, N.; Li, L. Nudix Hydrolase 23 Post-Translationally Regulates Carotenoid Biosynthesis in Plants. Plant Cell 2024, 36, 1868–1891. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



