Submitted:
02 December 2025
Posted:
03 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
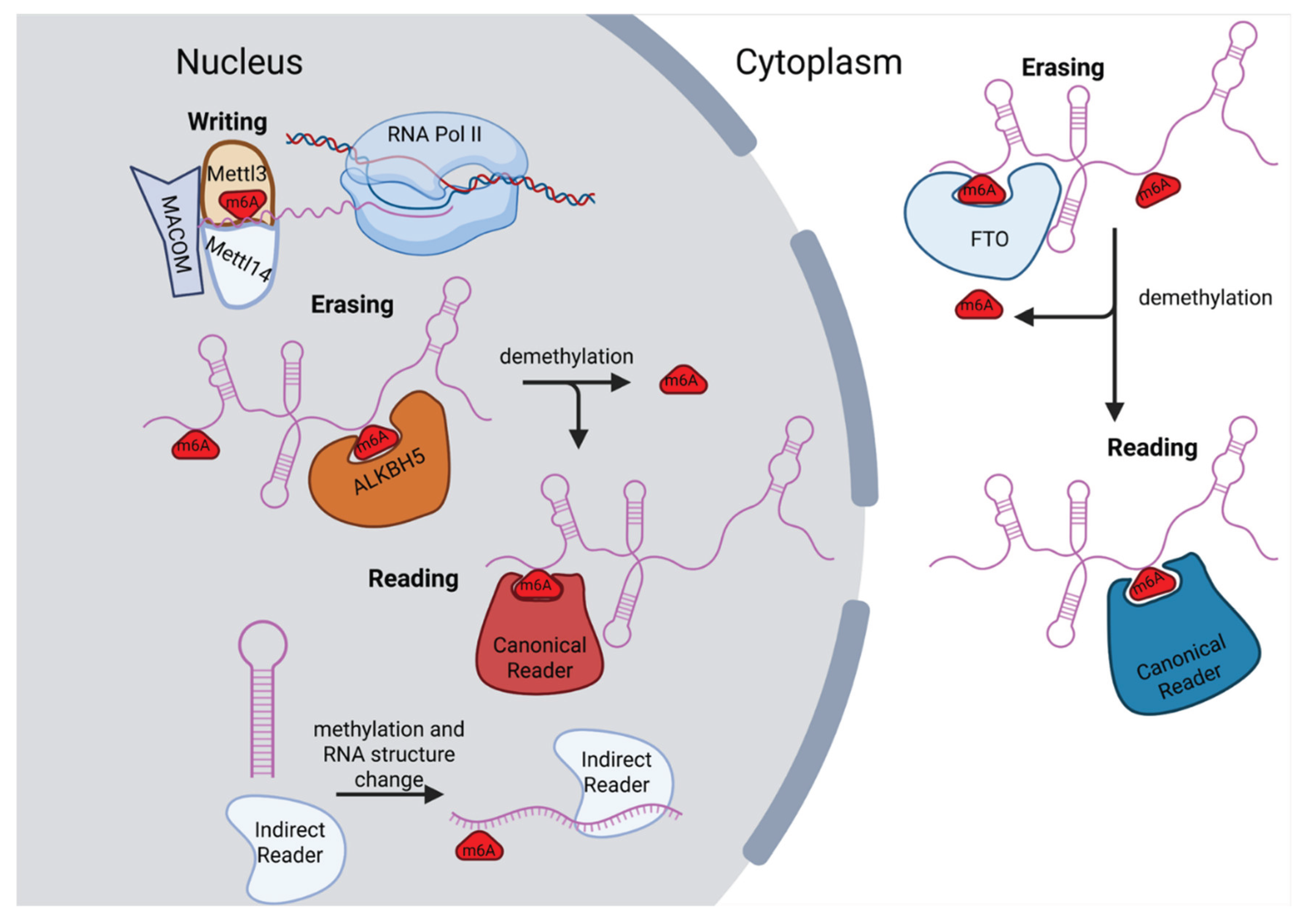
2. Writers and Erasers of m6A
3. m6A Readers and Their Interactions with Long Noncoding RNAs
4. Domain Architecture
5. Subcellular Localization
6. Consensus Sequence Binding
7. M6A Readers That Regulate lncRNAs
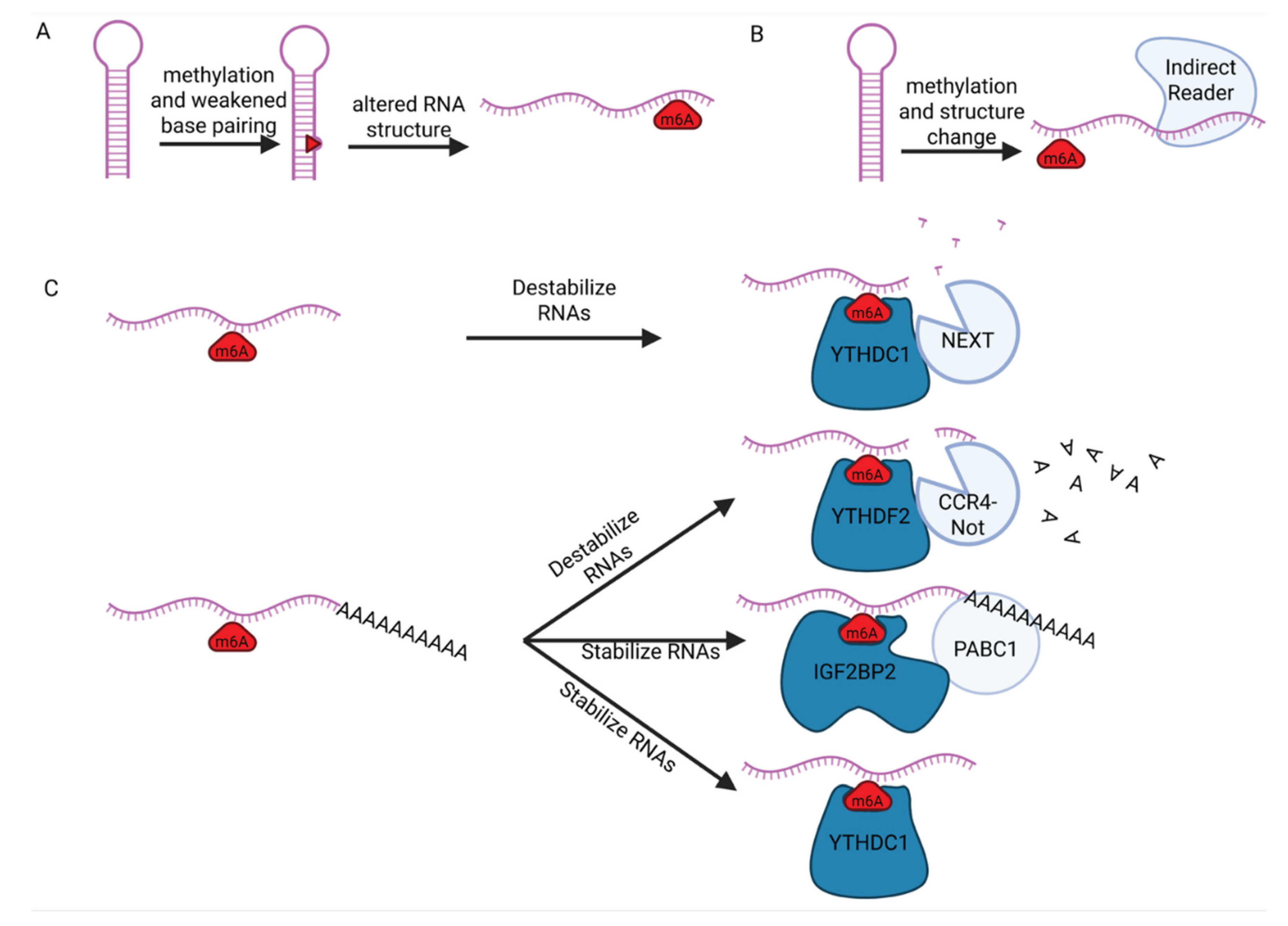
8. Role of m6A on RNA Structure and Stability
8.1. m6A Effect on RNA Structure and Accessibility to RBPs
8.2. m6A Effect on RNA Stability via RBP Recruitment
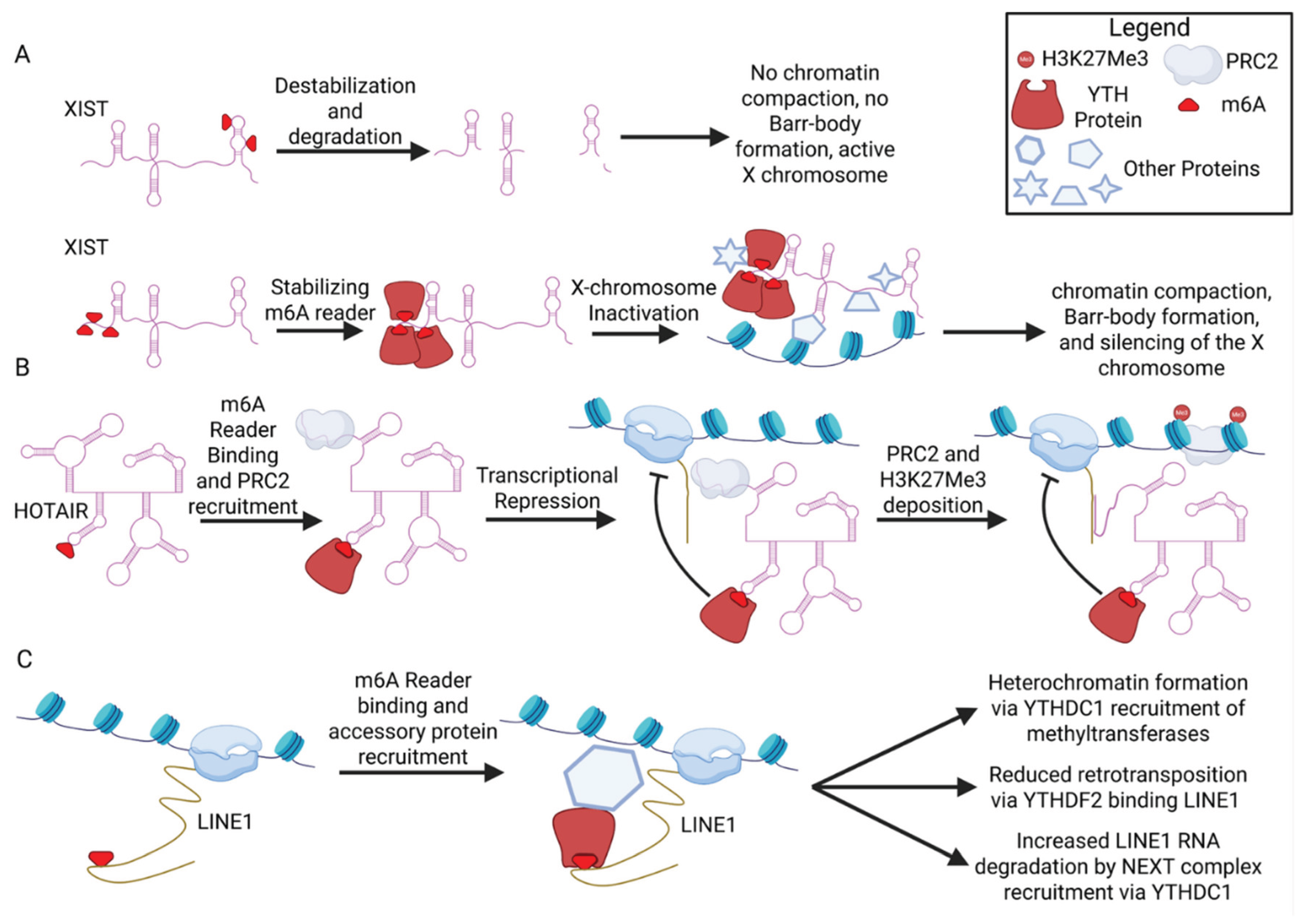
9. Involvement of m6A in lncRNA-Mediated Chromatin Repression
9.1. Xist
9.2. HOTAIR
9.3. LINE1
10. Enhancer Activation
10.1. Phase Separation
10.2. Xist Condensation of the Inactive X
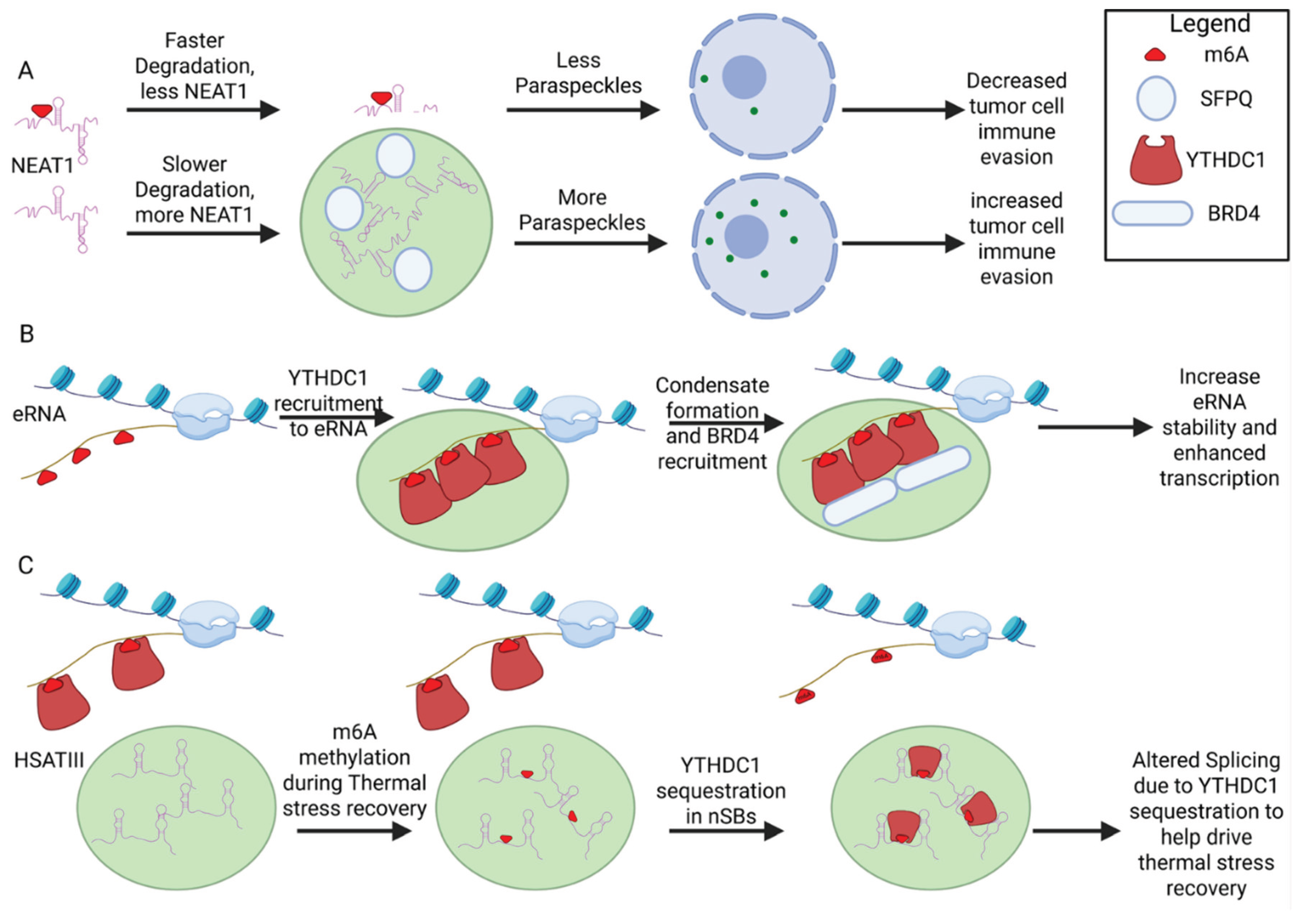
10.3. NEAT1 and Paraspeckle Formation
10.4. eRNA Mediated Transcriptional Condensates
10.5. HSATIII and Nuclear Stress Bodies
10.6. Other Example of m6A Involvement in LLPS
11. Cytoplasmic m6A-Modified lncRNAs
12. Conclusion
Funding
Authorship Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgements
Conflicts of Interests
References
- Ke, S., Pandya-Jones, A., Saito, Y., Fak, J.J., Vagbo, C.B., Geula, S., Hanna, J.H., Black, D.L., Darnell, J.E., Jr., and Darnell, R.B. (2017). m(6)A mRNA modifications are deposited in nascent pre-mRNA and are not required for splicing but do specify cytoplasmic turnover. Genes Dev 31, 990-1006. [CrossRef]
- Knuckles, P., Lence, T., Haussmann, I.U., Jacob, D., Kreim, N., Carl, S.H., Masiello, I., Hares, T., Villasenor, R., Hess, D., et al. (2018). Zc3h13/Flacc is required for adenosine methylation by bridging the mRNA-binding factor Rbm15/Spenito to the m(6)A machinery component Wtap/Fl(2)d. Genes Dev 32, 415-429. [CrossRef]
- Horiuchi, K., Kawamura, T., Iwanari, H., Ohashi, R., Naito, M., Kodama, T., and Hamakubo, T. (2013). Identification of Wilms’ tumor 1-associating protein complex and its role in alternative splicing and the cell cycle. J Biol Chem 288, 33292-33302. [CrossRef]
- He, P.C., and He, C. (2021). m(6) A RNA methylation: from mechanisms to therapeutic potential. EMBO J 40, e105977.
- Liu, J., Yue, Y., Han, D., Wang, X., Fu, Y., Zhang, L., Jia, G., Yu, M., Lu, Z., Deng, X., et al. (2013). A METTL3-METTL14 complex mediates mammalian nuclear RNA N6-adenosine methylation. Nat Chem Biol 10, 93-95. [CrossRef]
- Ruszkowska, A., Ruszkowski, M., Dauter, Z., and Brown, J.A. (2018). Structural insights into the RNA methyltransferase domain of METTL16. Sci Rep 8, 5311. [CrossRef]
- Doxtader, K.A., Wang, P., Scarborough, A.M., Seo, D., Conrad, N.K., and Nam, Y. (2018). Structural Basis for Regulation of METTL16, an S-Adenosylmethionine Homeostasis Factor. Mol Cell 71, 1001-1011 e1004. [CrossRef]
- Meiser, N., Mench, N., and Hengesbach, M. (2020). RNA secondary structure dependence in METTL3-METTL14 mRNA methylation is modulated by the N-terminal domain of METTL3. Biol Chem 402, 89-98. [CrossRef]
- Uzonyi, A., Dierks, D., Nir, R., Kwon, O.S., Toth, U., Barbosa, I., Burel, C., Brandis, A., Rossmanith, W., Le Hir, H., et al. (2023). Exclusion of m6A from splice-site proximal regions by the exon junction complex dictates m6A topologies and mRNA stability. Mol Cell 83, 237-251 e237. [CrossRef]
- Guerrero-Castillo, S., Cabrera-Orefice, A., Huynen, M.A., and Arnold, S. (2017). Identification and evolutionary analysis of tissue-specific isoforms of mitochondrial complex I subunit NDUFV3. Biochim Biophys Acta Bioenerg 1858, 208-217. [CrossRef]
- Gao, Z., Zha, X., Li, M., Xia, X., and Wang, S. (2024). Insights into the m(6)A demethylases FTO and ALKBH5: structural, biological function, and inhibitor development. Cell Biosci 14, 108. [CrossRef]
- Toh, J.D.W., Crossley, S.W.M., Bruemmer, K.J., Ge, E.J., He, D., Iovan, D.A., and Chang, C.J. (2020). Distinct RNA N-demethylation pathways catalyzed by nonheme iron ALKBH5 and FTO enzymes enable regulation of formaldehyde release rates. Proc Natl Acad Sci U S A 117, 25284-25292. [CrossRef]
- Wei, J., Liu, F., Lu, Z., Fei, Q., Ai, Y., He, P.C., Shi, H., Cui, X., Su, R., Klungland, A., et al. (2018). Differential m(6)A, m(6)A(m), and m(1)A Demethylation Mediated by FTO in the Cell Nucleus and Cytoplasm. Mol Cell 71, 973-985 e975. [CrossRef]
- Yu, F., Zhu, A.C., Liu, S., Gao, B., Wang, Y., Khudaverdyan, N., Yu, C., Wu, Q., Jiang, Y., Song, J., et al. (2023). RBM33 is a unique m(6)A RNA-binding protein that regulates ALKBH5 demethylase activity and substrate selectivity. Mol Cell 83, 2003-2019 e2006. [CrossRef]
- Li, R., Zhao, H., Huang, X., Zhang, J., Bai, R., Zhuang, L., Wen, S., Wu, S., Zhou, Q., Li, M., et al. (2023). Super-enhancer RNA m(6)A promotes local chromatin accessibility and oncogene transcription in pancreatic ductal adenocarcinoma. Nat Genet 55, 2224-2234. [CrossRef]
- Liao, S., Sun, H., and Xu, C. (2018). YTH Domain: A Family of N(6)-methyladenosine (m(6)A) Readers. Genomics Proteomics Bioinformatics 16, 99-107. [CrossRef]
- Li, F., Zhao, D., Wu, J., and Shi, Y. (2014). Structure of the YTH domain of human YTHDF2 in complex with an m(6)A mononucleotide reveals an aromatic cage for m(6)A recognition. Cell Res 24, 1490-1492. [CrossRef]
- Sikorski, V., Selberg, S., Lalowski, M., Karelson, M., and Kankuri, E. (2023). The structure and function of YTHDF epitranscriptomic m(6)A readers. Trends Pharmacol Sci 44, 335-353. [CrossRef]
- Dai, X., Wang, T., Gonzalez, G., and Wang, Y. (2018). Identification of YTH Domain-Containing Proteins as the Readers for N1-Methyladenosine in RNA. Anal Chem 90, 6380-6384. [CrossRef]
- Petri, B.J., and Klinge, C.M. (2023). m6A readers, writers, erasers, and the m6A epitranscriptome in breast cancer. J Mol Endocrinol 70. [CrossRef]
- Nicastro, G., Abis, G., Klein, P., Esteban-Serna, S., Gallagher, C., Chaves-Arquero, B., Cai, Y., Figueiredo, A.M., Martin, S.R., Patani, R., et al. (2023). Direct m6A recognition by IMP1 underlays an alternative model of target selection for non-canonical methyl-readers. Nucleic Acids Res 51, 8774-8786. [CrossRef]
- Fakhar, M., Gul, M., and Li, W. (2024). Interactive Structural Analysis of KH3-4 Didomains of IGF2BPs with Preferred RNA Motif Having m(6)A Through Dynamics Simulation Studies. Int J Mol Sci 25.
- Flamand, M.N., Tegowski, M., and Meyer, K.D. (2023). The Proteins of mRNA Modification: Writers, Readers, and Erasers. Annu Rev Biochem 92, 145-173. [CrossRef]
- Krecic, A.M., and Swanson, M.S. (1999). hnRNP complexes: composition, structure, and function. Curr Opin Cell Biol 11, 363-371.
- Wu, B., Su, S., Patil, D.P., Liu, H., Gan, J., Jaffrey, S.R., and Ma, J. (2018). Molecular basis for the specific and multivariant recognitions of RNA substrates by human hnRNP A2/B1. Nat Commun 9, 420. [CrossRef]
- Geuens, T., Bouhy, D., and Timmerman, V. (2016). The hnRNP family: insights into their role in health and disease. Hum Genet 135, 851-867.
- Li, F., and Li, W. (2024). Readers of RNA Modification in Cancer and Their Anticancer Inhibitors. Biomolecules 14.
- Hou, J., Gao, Y., Han, B., Yan, S., Wei, S., and Gao, X. (2025). Nuclear accumulation of YTHDF1 regulates mRNA splicing in the DNA damage response. Sci Adv 11, eado7660. [CrossRef]
- Zhou, J., Wan, J., Gao, X., Zhang, X., Jaffrey, S.R., and Qian, S.B. (2015). Dynamic m(6)A mRNA methylation directs translational control of heat shock response. Nature 526, 591-594.
- Kirli, K., Karaca, S., Dehne, H.J., Samwer, M., Pan, K.T., Lenz, C., Urlaub, H., and Gorlich, D. (2015). A deep proteomics perspective on CRM1-mediated nuclear export and nucleocytoplasmic partitioning. Elife 4. [CrossRef]
- Li, J., Ahmad, M., Sang, L., Zhan, Y., Wang, Y., Yan, Y., Liu, Y., Mi, W., Lu, M., Dai, Y., et al. (2023). O-GlcNAcylation promotes the cytosolic localization of the m(6)A reader YTHDF1 and colorectal cancer tumorigenesis. J Biol Chem 299, 104738.
- Wachter, K., Kohn, M., Stohr, N., and Huttelmaier, S. (2013). Subcellular localization and RNP formation of IGF2BPs (IGF2 mRNA-binding proteins) is modulated by distinct RNA-binding domains. Biol Chem 394, 1077-1090.
- Rivera Vargas, T., Boudoukha, S., Simon, A., Souidi, M., Cuvellier, S., Pinna, G., and Polesskaya, A. (2014). Post-transcriptional regulation of cyclins D1, D3 and G1 and proliferation of human cancer cells depend on IMP-3 nuclear localization. Oncogene 33, 2866-2875.
- Sterky, F.H., Ruzzenente, B., Gustafsson, C.M., Samuelsson, T., and Larsson, N.G. (2010). LRPPRC is a mitochondrial matrix protein that is conserved in metazoans. Biochem Biophys Res Commun 398, 759-764. [CrossRef]
- Nichols, R.C., Wang, X.W., Tang, J., Hamilton, B.J., High, F.A., Herschman, H.R., and Rigby, W.F. (2000). The RGG domain in hnRNP A2 affects subcellular localization. Exp Cell Res 256, 522-532.
- Lo, J., Vaeth, K.F., Bhardwaj, G., Mukherjee, N., Russ, H.A., Moore, J.K., and Taliaferro, J.M. (2024). The RNA binding protein HNRNPA2B1 regulates RNA abundance and motor protein activity in neurites. bioRxiv.
- Martino, F., Varadarajan, N.M., Perestrelo, A.R., Hejret, V., Durikova, H., Vukic, D., Horvath, V., Cavalieri, F., Caruso, F., Albihlal, W.S., et al. (2022). The mechanical regulation of RNA binding protein hnRNPC in the failing heart. Sci Transl Med 14, eabo5715.
- Xu, C., Wang, X., Liu, K., Roundtree, I.A., Tempel, W., Li, Y., Lu, Z., He, C., and Min, J. (2014). Structural basis for selective binding of m6A RNA by the YTHDC1 YTH domain. Nat Chem Biol 10, 927-929.
- Arguello, A.E., Leach, R.W., and Kleiner, R.E. (2019). In Vitro Selection with a Site-Specifically Modified RNA Library Reveals the Binding Preferences of N(6)-Methyladenosine Reader Proteins. Biochemistry 58, 3386-3395.
- Liu, B., Merriman, D.K., Choi, S.H., Schumacher, M.A., Plangger, R., Kreutz, C., Horner, S.M., Meyer, K.D., and Al-Hashimi, H.M. (2018). A potentially abundant junctional RNA motif stabilized by m(6)A and Mg(2). Nat Commun 9, 2761.
- Lu, S., Han, L., Hu, X., Sun, T., Xu, D., Li, Y., Chen, Q., Yao, W., He, M., Wang, Z., et al. (2021). N6-methyladenosine reader IMP2 stabilizes the ZFAS1/OLA1 axis and activates the Warburg effect: implication in colorectal cancer. J Hematol Oncol 14, 188. [CrossRef]
- Liu, N., Dai, Q., Zheng, G., He, C., Parisien, M., and Pan, T. (2015). N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560-564.
- Xiong, X., Feng, S., Ma, X., Liu, K., Gui, Y., Chen, B., Fan, X., Wang, F., Wang, X., and Yuan, S. (2025). hnRNPC Functions with HuR to Regulate Alternative Splicing in an m6A-Dependent Manner and is Essential for Meiosis. Adv Sci (Weinh) 12, e2412196.
- Kumar, A., Daripa, P., Penumutchu, S., Maiti, S., and Jain, N. (2025). Thermodynamic insights into N6-methyladenosine-modified ribonucleic acids and their interactions with the RNA recognition motif of heterogeneous nuclear ribonucleoprotein C. Int J Biol Macromol 312, 144210.
- Liu, N., Zhou, K.I., Parisien, M., Dai, Q., Diatchenko, L., and Pan, T. (2017). N6-methyladenosine alters RNA structure to regulate binding of a low-complexity protein. Nucleic Acids Res 45, 6051-6063. [CrossRef]
- Wang, Y., Li, Y., Toth, J.I., Petroski, M.D., Zhang, Z., and Zhao, J.C. (2014). N6-methyladenosine modification destabilizes developmental regulators in embryonic stem cells. Nat Cell Biol 16, 191-198.
- Spahr, H., Rozanska, A., Li, X., Atanassov, I., Lightowlers, R.N., Chrzanowska-Lightowlers, Z.M., Rackham, O., and Larsson, N.G. (2016). SLIRP stabilizes LRPPRC via an RRM-PPR protein interface. Nucleic Acids Res 44, 6868-6882. [CrossRef]
- Roost, C., Lynch, S.R., Batista, P.J., Qu, K., Chang, H.Y., and Kool, E.T. (2015). Structure and thermodynamics of N6-methyladenosine in RNA: a spring-loaded base modification. J Am Chem Soc 137, 2107-2115.
- Zhou, K.I., Parisien, M., Dai, Q., Liu, N., Diatchenko, L., Sachleben, J.R., and Pan, T. (2016). N(6)-Methyladenosine Modification in a Long Noncoding RNA Hairpin Predisposes Its Conformation to Protein Binding. J Mol Biol 428, 822-833.
- Wei, G. (2024). RNA m6A modification, signals for degradation or stabilisation? Biochem Soc Trans 52, 707-717. [CrossRef]
- Huang, H., Weng, H., Sun, W., Qin, X., Shi, H., Wu, H., Zhao, B.S., Mesquita, A., Liu, C., Yuan, C.L., et al. (2018). Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat Cell Biol 20, 285-295.
- Boo, S.H., and Kim, Y.K. (2020). The emerging role of RNA modifications in the regulation of mRNA stability. Exp Mol Med 52, 400-408.
- Wang, X., Lu, Z., Gomez, A., Hon, G.C., Yue, Y., Han, D., Fu, Y., Parisien, M., Dai, Q., Jia, G., et al. (2014). N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117-120.
- Du, H., Zhao, Y., He, J., Zhang, Y., Xi, H., Liu, M., Ma, J., and Wu, L. (2016). YTHDF2 destabilizes m(6)A-containing RNA through direct recruitment of the CCR4-NOT deadenylase complex. Nat Commun 7, 12626. [CrossRef]
- Park, O.H., Ha, H., Lee, Y., Boo, S.H., Kwon, D.H., Song, H.K., and Kim, Y.K. (2019). Endoribonucleolytic Cleavage of m(6)A-Containing RNAs by RNase P/MRP Complex. Mol Cell 74, 494-507 e498.
- Wu, S., Chi, C., Weng, S., Zhou, W., and Liu, Z. (2023). IGF2BP2 promotes lncRNA DANCR stability mediated glycolysis and affects the progression of FLT3-ITD + acute myeloid leukemia. Apoptosis 28, 1035-1047. [CrossRef]
- Porman, A.M., Roberts, J.T., Duncan, E.D., Chrupcala, M.L., Levine, A.A., Kennedy, M.A., Williams, M.M., Richer, J.K., and Johnson, A.M. (2022). A single N6-methyladenosine site regulates lncRNA HOTAIR function in breast cancer cells. PLoS Biol 20, e3001885.
- Lee, J.H., Wang, R., Xiong, F., Krakowiak, J., Liao, Z., Nguyen, P.T., Moroz-Omori, E.V., Shao, J., Zhu, X., Bolt, M.J., et al. (2021). Enhancer RNA m6A methylation facilitates transcriptional condensate formation and gene activation. Mol Cell 81, 3368-3385 e3369.
- Liang, D., Lin, W.J., Ren, M., Qiu, J., Yang, C., Wang, X., Li, N., Zeng, T., Sun, K., You, L., et al. (2022). m(6)A reader YTHDC1 modulates autophagy by targeting SQSTM1 in diabetic skin. Autophagy 18, 1318-1337.
- Zhang, Z., Wang, Q., Zhao, X., Shao, L., Liu, G., Zheng, X., Xie, L., Zhang, Y., Sun, C., and Xu, R. (2020). YTHDC1 mitigates ischemic stroke by promoting Akt phosphorylation through destabilizing PTEN mRNA. Cell Death Dis 11, 977.
- Liu, J., Dou, X., Chen, C., Chen, C., Liu, C., Xu, M.M., Zhao, S., Shen, B., Gao, Y., Han, D., et al. (2020). N (6)-methyladenosine of chromosome-associated regulatory RNA regulates chromatin state and transcription. Science 367, 580-586. [CrossRef]
- Wei, G., Coker, H., Rodermund, L., Almeida, M., Roach, H.L., Nesterova, T.B., and Brockdorff, N. (2025). m(6)A and the NEXT complex direct Xist RNA turnover and X-inactivation dynamics. Nat Struct Mol Biol.
- Saglam, B., and Akgul, B. (2024). An Overview of Current Detection Methods for RNA Methylation. Int J Mol Sci 25.
- Xiao, Y.L., Liu, S., Ge, R., Wu, Y., He, C., Chen, M., and Tang, W. (2023). Transcriptome-wide profiling and quantification of N(6)-methyladenosine by enzyme-assisted adenosine deamination. Nat Biotechnol 41, 993-1003.
- Zhong, Z.D., Xie, Y.Y., Chen, H.X., Lan, Y.L., Liu, X.H., Ji, J.Y., Wu, F., Jin, L., Chen, J., Mak, D.W., et al. (2023). Systematic comparison of tools used for m(6)A mapping from nanopore direct RNA sequencing. Nat Commun 14, 1906. [CrossRef]
- Sun, H., Lu, B., Zhang, Z., Xiao, Y., Zhou, Z., Xi, L., Li, Z., Jiang, Z., Zhang, J., Wang, M., et al. (2025). Mild and ultrafast GLORI enables absolute quantification of m(6)A methylome from low-input samples. Nat Methods 22, 1226-1236.
- Linder, B., Grozhik, A.V., Olarerin-George, A.O., Meydan, C., Mason, C.E., and Jaffrey, S.R. (2015). Single-nucleotide-resolution mapping of m6A and m6Am throughout the transcriptome. Nat Methods 12, 767-772.
- Chen, K., Lu, Z., Wang, X., Fu, Y., Luo, G.Z., Liu, N., Han, D., Dominissini, D., Dai, Q., Pan, T., et al. (2015). High-resolution N(6) -methyladenosine (m(6) A) map using photo-crosslinking-assisted m(6) A sequencing. Angew Chem Int Ed Engl 54, 1587-1590.
- Shen, W., Sun, H., Liu, C., Yi, Y., Hou, Y., Xiao, Y., Hu, Y., Lu, B., Peng, J., Wang, J., et al. (2024). GLORI for absolute quantification of transcriptome-wide m(6)A at single-base resolution. Nat Protoc 19, 1252-1287. [CrossRef]
- Liu, C., Sun, H., Yi, Y., Shen, W., Li, K., Xiao, Y., Li, F., Li, Y., Hou, Y., Lu, B., et al. (2023). Absolute quantification of single-base m(6)A methylation in the mammalian transcriptome using GLORI. Nat Biotechnol 41, 355-366.
- Brown, C.J., Hendrich, B.D., Rupert, J.L., Lafreniere, R.G., Xing, Y., Lawrence, J., and Willard, H.F. (1992). The human XIST gene: analysis of a 17 kb inactive X-specific RNA that contains conserved repeats and is highly localized within the nucleus. Cell 71, 527-542.
- Penny, G.D., Kay, G.F., Sheardown, S.A., Rastan, S., and Brockdorff, N. (1996). Requirement for Xist in X chromosome inactivation. Nature 379, 131-137.
- Brockdorff, N., Bowness, J.S., and Wei, G. (2020). Progress toward understanding chromosome silencing by Xist RNA. Genes Dev 34, 733-744.
- Chu, C., Zhang, Q.C., da Rocha, S.T., Flynn, R.A., Bharadwaj, M., Calabrese, J.M., Magnuson, T., Heard, E., and Chang, H.Y. (2015). Systematic discovery of Xist RNA binding proteins. Cell 161, 404-416. [CrossRef]
- Minajigi, A., Froberg, J.E., Wei, C., Sunwoo, H., Kesner, B., Colognori, D., Lessing, D., Payer, B., Boukhali, M., Haas, W., et al. (2015). Chromosomes. A comprehensive Xist interactome reveals cohesin repulsion and an RNA-directed chromosome conformation. Science 349.
- Moindrot, B., Cerase, A., Coker, H., Masui, O., Grijzenhout, A., Pintacuda, G., Schermelleh, L., Nesterova, T.B., and Brockdorff, N. (2015). A Pooled shRNA Screen Identifies Rbm15, Spen, and Wtap as Factors Required for Xist RNA-Mediated Silencing. Cell Rep 12, 562-572.
- Patil, D.P., Chen, C.K., Pickering, B.F., Chow, A., Jackson, C., Guttman, M., and Jaffrey, S.R. (2016). m(6)A RNA methylation promotes XIST-mediated transcriptional repression. Nature 537, 369-373. [CrossRef]
- Nesterova, T.B., Wei, G., Coker, H., Pintacuda, G., Bowness, J.S., Zhang, T., Almeida, M., Bloechl, B., Moindrot, B., Carter, E.J., et al. (2019). Systematic allelic analysis defines the interplay of key pathways in X chromosome inactivation. Nat Commun 10, 3129.
- Dossin, F., Pinheiro, I., Zylicz, J.J., Roensch, J., Collombet, S., Le Saux, A., Chelmicki, T., Attia, M., Kapoor, V., Zhan, Y., et al. (2020). SPEN integrates transcriptional and epigenetic control of X-inactivation. Nature 578, 455-460.
- Coker, H., Wei, G., Moindrot, B., Mohammed, S., Nesterova, T., and Brockdorff, N. (2020). The role of the Xist 5′ m6A region and RBM15 in X chromosome inactivation. Wellcome Open Res 5, 31.
- Lu, Z., Guo, J.K., Wei, Y., Dou, D.R., Zarnegar, B., Ma, Q., Li, R., Zhao, Y., Liu, F., Choudhry, H., et al. (2020). Structural modularity of the XIST ribonucleoprotein complex. Nat Commun 11, 6163.
- Wei, G., Almeida, M., Pintacuda, G., Coker, H., Bowness, J.S., Ule, J., and Brockdorff, N. (2021). Acute depletion of METTL3 implicates N (6)-methyladenosine in alternative intron/exon inclusion in the nascent transcriptome. Genome Res 31, 1395-1408. [CrossRef]
- Chang, C., Ma, G., Cheung, E., and Hutchins, A.P. (2022). A programmable system to methylate and demethylate N(6)-methyladenosine (m(6)A) on specific RNA transcripts in mammalian cells. J Biol Chem 298, 102525.
- Rinn, J.L., Kertesz, M., Wang, J.K., Squazzo, S.L., Xu, X., Brugmann, S.A., Goodnough, L.H., Helms, J.A., Farnham, P.J., Segal, E., et al. (2007). Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 129, 1311-1323.
- Balas, M.M.a.J., A. M. (2018). Exploring the mechanisms behind long noncoding RNAs and cancer. Non-coding RNA Research.
- Gupta, R.A., Shah, N., Wang, K.C., Kim, J., Horlings, H.M., Wong, D.J., Tsai, M.C., Hung, T., Argani, P., Rinn, J.L., et al. (2010). Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 464, 1071-1076. [CrossRef]
- Tsai, M.C., Manor, O., Wan, Y., Mosammaparast, N., Wang, J.K., Lan, F., Shi, Y., Segal, E., and Chang, H.Y. (2010). Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689-693.
- Somarowthu, S., Legiewicz, M., Chillon, I., Marcia, M., Liu, F., and Pyle, A.M. (2015). HOTAIR forms an intricate and modular secondary structure. Mol Cell 58, 353-361.
- Meredith, E.K., Balas, M.M., Sindy, K., Haislop, K., and Johnson, A.M. (2016). An RNA matchmaker protein regulates the activity of the long noncoding RNA HOTAIR. RNA 22, 995-1010.
- Nguyen, E.D., Balas, M.M., Griffin, A.M., Roberts, J.T., and Johnson, A.M. (2018). Global profiling of hnRNP A2/B1-RNA binding on chromatin highlights LncRNA interactions. RNA Biol, 1-13.
- Balas, M.M., Hartwick, E.W., Barrington, C., Roberts, J.T., Wu, S.K., Bettcher, R., Griffin, A.M., Kieft, J.S., and Johnson, A.M. (2021). Establishing RNA-RNA interactions remodels lncRNA structure and promotes PRC2 activity. Sci Adv 7.
- Portoso, M., Ragazzini, R., Brencic, Z., Moiani, A., Michaud, A., Vassilev, I., Wassef, M., Servant, N., Sargueil, B., and Margueron, R. (2017). PRC2 is dispensable for HOTAIR-mediated transcriptional repression. EMBO J.
- Delhaye, L., De Bruycker, E., Volders, P.J., Fijalkowska, D., De Sutter, D., Degroeve, S., Martens, L., Mestdagh, P., and Eyckerman, S. (2022). Orthogonal proteomics methods to unravel the HOTAIR interactome. Sci Rep 12, 1513.
- Wu, X.M., Mai, Y.X., Wen, Y.F., Li, Z.P., Sun, Y.X., Chen, J.J., Meng, F., Pang, F.X., Li, H.M., Pan, Y., et al. (2025). Silence of HOTAIR promotes osteogenic differentiation and accelerates distraction osteogenesis by mediating FTO ubiquitination. J Orthop Translat 50, 248-256.
- Liu, T.C., Li, H.X., Wan, Y.X., Shi, G., Zhao, Y.P., Liu, Y.F., and Fan, X.Y. (2024). METTL14-mediated upregulation of lncRNA HOTAIR represses PP1alpha expression by promoting H3K4me1 demethylation in oxycodone-treated mice. CNS Neurosci Ther 30, e14830. [CrossRef]
- Liu, J., Gao, M., He, J., Wu, K., Lin, S., Jin, L., Chen, Y., Liu, H., Shi, J., Wang, X., et al. (2021). The RNA m(6)A reader YTHDC1 silences retrotransposons and guards ES cell identity. Nature 591, 322-326.
- Chen, C., Liu, W., Guo, J., Liu, Y., Liu, X., Liu, J., Dou, X., Le, R., Huang, Y., Li, C., et al. (2021). Nuclear m(6)A reader YTHDC1 regulates the scaffold function of LINE1 RNA in mouse ESCs and early embryos. Protein Cell 12, 455-474.
- Wei, J., Yu, X., Yang, L., Liu, X., Gao, B., Huang, B., Dou, X., Liu, J., Zou, Z., Cui, X.L., et al. (2022). FTO mediates LINE1 m(6)A demethylation and chromatin regulation in mESCs and mouse development. Science 376, 968-973.
- Xiong, F., Wang, R., Lee, J.H., Li, S., Chen, S.F., Liao, Z., Hasani, L.A., Nguyen, P.T., Zhu, X., Krakowiak, J., et al. (2021). RNA m(6)A modification orchestrates a LINE-1-host interaction that facilitates retrotransposition and contributes to long gene vulnerability. Cell Res 31, 861-885.
- Li, Z., Fang, F., Zafar, M.I., Wu, X., Liu, X., Tan, X., Luo, J., Ye, Z., Xiong, C., and Li, H. (2024). RNA m(6)A modification regulates L1 retrotransposons in human spermatogonial stem cell differentiation in vitro and in vivo. Cell Mol Life Sci 81, 92.
- Chelmicki, T., Roger, E., Teissandier, A., Dura, M., Bonneville, L., Rucli, S., Dossin, F., Fouassier, C., Lameiras, S., and Bourc’his, D. (2021). m(6)A RNA methylation regulates the fate of endogenous retroviruses. Nature 591, 312-316.
- Altendorfer, E., Mundlos, S., and Mayer, A. (2025). A transcription coupling model for how enhancers communicate with their target genes. Nat Struct Mol Biol 32, 598-606. [CrossRef]
- Xiao, S., Cao, S., Huang, Q., Xia, L., Deng, M., Yang, M., Jia, G., Liu, X., Shi, J., Wang, W., et al. (2019). The RNA N(6)-methyladenosine modification landscape of human fetal tissues. Nat Cell Biol 21, 651-661.
- Xu, W., He, C., Kaye, E.G., Li, J., Mu, M., Nelson, G.M., Dong, L., Wang, J., Wu, F., Shi, Y.G., et al. (2022). Dynamic control of chromatin-associated m(6)A methylation regulates nascent RNA synthesis. Mol Cell 82, 1156-1168 e1157.
- Banani, S.F., Lee, H.O., Hyman, A.A., and Rosen, M.K. (2017). Biomolecular condensates: organizers of cellular biochemistry. Nat Rev Mol Cell Biol 18, 285-298.
- Zumbro, E., and Alexander-Katz, A. (2021). Multivalent polymers can control phase boundary, dynamics, and organization of liquid-liquid phase separation. PLoS One 16, e0245405.
- Grese, Z.R., Bastos, A.C., Mamede, L.D., French, R.L., Miller, T.M., and Ayala, Y.M. (2021). Specific RNA interactions promote TDP-43 multivalent phase separation and maintain liquid properties. EMBO Rep 22, e53632.
- Chen, C., Jia, H., Nakamura, Y., Kanekura, K., and Hayamizu, Y. (2022). Effect of Multivalency on Phase-Separated Droplets Consisting of Poly(PR) Dipeptide Repeats and RNA at the Solid/Liquid Interface. ACS Omega 7, 19280-19287. [CrossRef]
- Parker, D.M., Tauber, D., and Parker, R. (2025). G3BP1 promotes intermolecular RNA-RNA interactions during RNA condensation. Mol Cell 85, 571-584 e577.
- Ries, R.J., Zaccara, S., Klein, P., Olarerin-George, A., Namkoong, S., Pickering, B.F., Patil, D.P., Kwak, H., Lee, J.H., and Jaffrey, S.R. (2019). m(6)A enhances the phase separation potential of mRNA. Nature 571, 424-428. [CrossRef]
- Gao, Y., Pei, G., Li, D., Li, R., Shao, Y., Zhang, Q.C., and Li, P. (2019). Multivalent m(6)A motifs promote phase separation of YTHDF proteins. Cell Res 29, 767-769.
- Wang, J., Wang, L., Diao, J., Shi, Y.G., Shi, Y., Ma, H., and Shen, H. (2020). Binding to m(6)A RNA promotes YTHDF2-mediated phase separation. Protein Cell 11, 304-307.
- Liu, S.Y., Feng, Y., Wu, J.J., Zou, M.L., Sun, Z.L., Li, X., and Yuan, F.L. (2020). m(6) A facilitates YTHDF-independent phase separation. J Cell Mol Med 24, 2070-2072.
- Park, J., Wu, Y., Shao, W., Gendron, T.F., van der Spek, S.J.F., Sultanakhmetov, G., Basu, A., Castellanos Otero, P., Jones, C.J., Jansen-West, K., et al. (2023). Poly(GR) interacts with key stress granule factors promoting its assembly into cytoplasmic inclusions. Cell Rep 42, 112822.
- Chen, Y., Wan, R., Zou, Z., Lao, L., Shao, G., Zheng, Y., Tang, L., Yuan, Y., Ge, Y., He, C., et al. (2023). O-GlcNAcylation determines the translational regulation and phase separation of YTHDF proteins. Nat Cell Biol 25, 1676-1690. [CrossRef]
- Shan, T., Liu, F., Wen, M., Chen, Z., Li, S., Wang, Y., Cheng, H., and Zhou, Y. (2023). m(6)A modification negatively regulates translation by switching mRNA from polysome to P-body via IGF2BP3. Mol Cell 83, 4494-4508 e4496.
- Cheng, Y., Xie, W., Pickering, B.F., Chu, K.L., Savino, A.M., Yang, X., Luo, H., Nguyen, D.T., Mo, S., Barin, E., et al. (2021). N(6)-Methyladenosine on mRNA facilitates a phase-separated nuclear body that suppresses myeloid leukemic differentiation. Cancer Cell 39, 958-972 e958.
- Han, D., Longhini, A.P., Zhang, X., Hoang, V., Wilson, M.Z., and Kosik, K.S. (2022). Dynamic assembly of the mRNA m6A methyltransferase complex is regulated by METTL3 phase separation. PLoS Biol 20, e3001535.
- Jiang, A., Zhang, S., Wang, X., and Li, D. (2022). RBM15 condensates modulate m(6)A modification of STYK1 to promote tumorigenesis. Comput Struct Biotechnol J 20, 4825-4836. [CrossRef]
- Ge, Y., Chen, R., Ling, T., Liu, B., Huang, J., Cheng, Y., Lin, Y., Chen, H., Xie, X., Xia, G., et al. (2024). Elevated WTAP promotes hyperinflammation by increasing m6A modification in inflammatory disease models. J Clin Invest 134.
- Cai, S., Zhou, J., Luo, X., Zhang, C., Jin, S., Ren, J., and Cui, J. (2025). Phase transition of WTAP regulates m(6)A modification of interferon-stimulated genes. Elife 13.
- Cerase, A., Armaos, A., Neumayer, C., Avner, P., Guttman, M., and Tartaglia, G.G. (2019). Phase separation drives X-chromosome inactivation: a hypothesis. Nat Struct Mol Biol 26, 331-334.
- Qin, X., Long, Y., Bai, X., Cao, L., Yan, H., Zhang, K., Wang, B., and Wu, X. (2023). The disordered C terminus of ALKBH5 promotes phase separation and paraspeckles assembly. J Biol Chem 299, 105071.
- Demmerle, J., Hao, S., and Cai, D. (2023). Transcriptional condensates and phase separation: condensing information across scales and mechanisms. Nucleus 14, 2213551. [CrossRef]
- Biamonti, G., and Vourc’h, C. (2010). Nuclear stress bodies. Cold Spring Harb Perspect Biol 2, a000695.
- Pandya-Jones, A., Markaki, Y., Serizay, J., Chitiashvili, T., Mancia Leon, W.R., Damianov, A., Chronis, C., Papp, B., Chen, C.K., McKee, R., et al. (2020). A protein assembly mediates Xist localization and gene silencing. Nature 587, 145-151.
- Yamazaki, T., Yamamoto, T., and Hirose, T. (2022). Micellization: A new principle in the formation of biomolecular condensates. Front Mol Biosci 9, 974772.
- Yamazaki, T., and Hirose, T. (2015). The building process of the functional paraspeckle with long non-coding RNAs. Front Biosci (Elite Ed) 7, 1-41.
- Pisani, G., and Baron, B. (2020). NEAT1 and Paraspeckles in Cancer Development and Chemoresistance. Noncoding RNA 6.
- Mamontova, V., Trifault, B., Gribling-Burrer, A.S., Bohn, P., Boten, L., Preckwinkel, P., Gallant, P., Solvie, D., Ade, C.P., Papadopoulos, D., et al. (2024). NEAT1 promotes genome stability via m(6)A methylation-dependent regulation of CHD4. Genes Dev 38, 915-930.
- Zhang, J., Guo, S., Piao, H.Y., Wang, Y., Wu, Y., Meng, X.Y., Yang, D., Zheng, Z.C., and Zhao, Y. (2019). ALKBH5 promotes invasion and metastasis of gastric cancer by decreasing methylation of the lncRNA NEAT1. J Physiol Biochem 75, 379-389. [CrossRef]
- Guo, T., Liu, D.F., Peng, S.H., and Xu, A.M. (2020). ALKBH5 promotes colon cancer progression by decreasing methylation of the lncRNA NEAT1. Am J Transl Res 12, 4542-4549.
- Dong, F., Qin, X., Wang, B., Li, Q., Hu, J., Cheng, X., Guo, D., Cheng, F., Fang, C., Tan, Y., et al. (2021). ALKBH5 Facilitates Hypoxia-Induced Paraspeckle Assembly and IL8 Secretion to Generate an Immunosuppressive Tumor Microenvironment. Cancer Res 81, 5876-5888.
- Ninomiya, K., Adachi, S., Natsume, T., Iwakiri, J., Terai, G., Asai, K., and Hirose, T. (2020). LncRNA-dependent nuclear stress bodies promote intron retention through SR protein phosphorylation. EMBO J 39, e102729.
- Ninomiya, K., Iwakiri, J., Aly, M.K., Sakaguchi, Y., Adachi, S., Natsume, T., Terai, G., Asai, K., Suzuki, T., and Hirose, T. (2021). m(6) A modification of HSATIII lncRNAs regulates temperature-dependent splicing. EMBO J 40, e107976.
- Wang, S., Wang, Y., Li, Q., Zeng, K., Li, X., and Feng, X. (2023). RUNX1-IT1 favors breast cancer carcinogenesis through regulation of IGF2BP1/GPX4 axis. Discov Oncol 14, 42.
- Chen, L., Zhang, C., Ma, W., Huang, J., Zhao, Y., and Liu, H. (2022). METTL3-mediated m6A modification stabilizes TERRA and maintains telomere stability. Nucleic Acids Res 50, 11619-11634. [CrossRef]
- Vaid, R., Thombare, K., Mendez, A., Burgos-Panadero, R., Djos, A., Jachimowicz, D., Lundberg, K.I., Bartenhagen, C., Kumar, N., Tummler, C., et al. (2024). METTL3 drives telomere targeting of TERRA lncRNA through m6A-dependent R-loop formation: a therapeutic target for ALT-positive neuroblastoma. Nucleic Acids Res 52, 2648-2671.
- Kang, Z., Li, R., Liu, C., Dong, X., Hu, Y., Xu, L., Liu, X., Xiang, Y., Gao, L., Si, W., et al. (2024). m(6)A-modified cenRNA stabilizes CENPA to ensure centromere integrity in cancer cells. Cell 187, 6035-6054 e6027.
- Blower, M.D. (2016). Centromeric Transcription Regulates Aurora-B Localization and Activation. Cell Rep 15, 1624-1633.
- Trivedi, P., Palomba, F., Niedzialkowska, E., Digman, M.A., Gratton, E., and Stukenberg, P.T. (2019). The inner centromere is a biomolecular condensate scaffolded by the chromosomal passenger complex. Nat Cell Biol 21, 1127-1137. [CrossRef]
- Chen, S., Wang, Y., Zhang, J., Liu, B., Liu, W., Cao, G., Li, R., Li, H., Zhai, N., Song, X., et al. (2025). YTHDC1 phase separation drives the nuclear export of m(6)A-modified lncNONMMUT062668.2 through the transport complex SRSF3-ALYREF-XPO5 to aggravate pulmonary fibrosis. Cell Death Dis 16, 279.
- Liu, H., Xu, Y., Yao, B., Sui, T., Lai, L., and Li, Z. (2020). A novel N6-methyladenosine (m6A)-dependent fate decision for the lncRNA THOR. Cell Death Dis 11, 613.
- Hosono, Y., Niknafs, Y.S., Prensner, J.R., Iyer, M.K., Dhanasekaran, S.M., Mehra, R., Pitchiaya, S., Tien, J., Escara-Wilke, J., Poliakov, A., et al. (2023). Oncogenic Role of THOR, a Conserved Cancer/Testis Long Non-coding RNA. Cell 186, 4254-4255.
- Thin, K.Z., Liu, X., Feng, X., Raveendran, S., and Tu, J.C. (2018). LncRNA-DANCR: A valuable cancer related long non-coding RNA for human cancers. Pathol Res Pract 214, 801-805.
- Hu, X., Peng, W.X., Zhou, H., Jiang, J., Zhou, X., Huang, D., Mo, Y.Y., and Yang, L. (2020). IGF2BP2 regulates DANCR by serving as an N6-methyladenosine reader. Cell Death Differ 27, 1782-1794. [CrossRef]
- Wang, M., Gu, J., Zhang, X., Yang, J., Zhang, X., and Fang, X. (2021). Long Non-coding RNA DANCR in Cancer: Roles, Mechanisms, and Implications. Front Cell Dev Biol 9, 753706.
- Li, G., Ma, L., He, S., Luo, R., Wang, B., Zhang, W., Song, Y., Liao, Z., Ke, W., Xiang, Q., et al. (2022). WTAP-mediated m(6)A modification of lncRNA NORAD promotes intervertebral disc degeneration. Nat Commun 13, 1469.
- Liu, W.J., Wang, J.X., Li, Q.F., Zhang, Y.H., Ji, P.F., Jin, J.H., Zhang, Y.B., Yuan, Z.H., Feng, P., Wu, Y.F., et al. (2025). Fat mass and obesity-associated protein in mesenchymal stem cells inhibits osteoclastogenesis via lnc NORAD/miR-4284 axis in ankylosing spondylitis. World J Stem Cells 17, 98911. [CrossRef]
- Soghli, N., Yousefi, T., Abolghasemi, M., and Qujeq, D. (2021). NORAD, a critical long non-coding RNA in human cancers. Life Sci 264, 118665.
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



