Submitted:
29 November 2025
Posted:
05 December 2025
You are already at the latest version
Abstract
Background: Nasal Staphylococcus aureus is a major source of community and hospital associated staphylococcal infections, including Methicillin-resistant S. aureus (MRSA). Screening of MRSA nasal colonization is important in the prevention and control of infection and may provide useful information to guide antimicrobial therapy.Objective: The study aims to determine the prevalence of nasal colonization by S. aureus and investigated its antimicrobial resistance profile to methicillin and glycopeptides in hospital and non-hospital settings in Maputo, Mozambique. Methods: A cross-sectional study was conducted by collecting nasal specimens of symptomatic outpatients and asymptomatic students at Eduardo Mondlane University in Maputo. Using culture techniques on Mannitol salt and HiCrome™ Rapid MRSA agar, S. aureus was isolated based on the colonial characteristics and confirmed by Gram staining, catalase, and Microgen®Staph tests. The antibiotic susceptibility test was done using Kirby-Buer disk diffusion method on Salt Muellar Hinton agar for methicillin (cefoxitin), vancomycin and teicoplanin antibiotics. Results: A total of 50 (39.1%) S. aureus isolates were obtained from 128 collected nasal samples, in which 104 (31.3%) were from university students and 24 (7.8%) from outpatient’s form university clinic. The isolates showed a low overall 7.8% resistance to cefoxitin and, 14.1% and 11.7% presumptive resistance to teicoplanin and vancomycin, respectively. Conclusion: Methicillin and glycopeptide resistant S. aureus were highly prevalent in asymptomatic students. These observations call for strategies to prevent S. aureus spread to more vulnerable populations where the consequences can be severe.
Keywords:
Introduction
Materials and Methods
Study Type and Population
Isolation and Characterization of S. aureus
Antimicrobial Susceptibility Test
Statistical Analysis
Results
| Category | Sample Number (N) |
S. aureus | CoN Staphylococcus | P value* | ||
|---|---|---|---|---|---|---|
| n | (%) | n | (%) | |||
| Overall | 128 | 50 | 39.1 | 54 | 42.2 | - |
| Type of participants | ||||||
| UEM clinic Outpatients | 24 | 10 | 7.8 | 11 | 8.6 | 0.684 |
| DBC-UEM Students | 104 | 40 | 31.3 | 43 | 33.6 | |
| Sex | ||||||
| Female | 81 | 23 | 18.0 | 41 | 32.0 | 0.005 |
| Male | 47 | 27 | 21.1 | 13 | 10.2 | |
| Age group (years) | ||||||
| ≤17 | 2 | 2 | 1.6 | 0 | 0.0 | 0.525 |
| 18-30 | 116 | 42 | 32.8 | 51 | 39.8 | |
| 31-45 | 4 | 3 | 2.3 | 1 | 0.8 | |
| 46-60 | 3 | 2 | 1.6 | 0 | 0.0 | |
| ≥61 | 3 | 1 | 0.8 | 2 | 1.6 | |
| CoN – Coagulase-Negative Staphylococcus; *Chi-square test | ||||||
| Antibiotics | Overall Resistance (n=128) | Outpatientes (n=24) | Students (n=124) | |||
| n | (%) | n | (%) | n | (%) | |
| Cefoxitin (30µg) | 10 | 7.8 | 1 | 4.2 | 9 | 7.3 |
| Vancomycin (30µg) | 15 | 11.7 | 4 | 16.7 | 11 | 8.9 |
| Teicoplanin (30µg) | 18 | 14.1 | 6 | 25.0 | 12 | 9.7 |
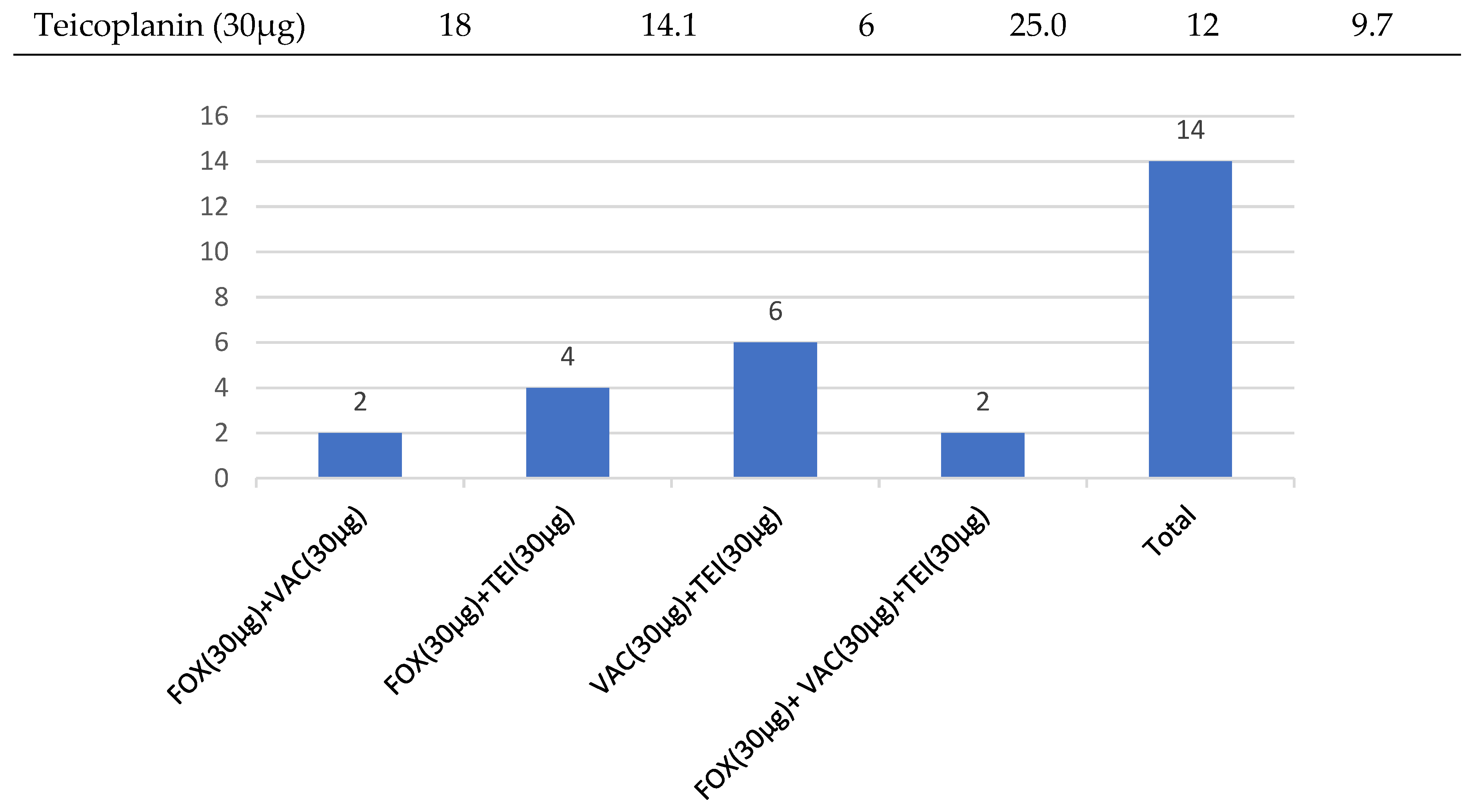
Discussion
Acknowledgements
Author Contributions
Disclosure
References
- Thampi N, Showler A, Burry L, Bai AD, Steinberg M, Ricciuto DR, et al. Multicenter study of health care cost of patients admitted to hospital with Staphylococcus aureus bacteremia: Impact of length of stay and intensity of care. Am J Infect Control. 2015 Jul;43(7):739–44. [CrossRef]
- Sakr A, Brégeon F, Mège JL, Rolain JM, Blin O. Staphylococcus aureus Nasal Colonization: An Update on Mechanisms, Epidemiology, Risk Factors, and Subsequent Infections. Front Microbiol. 2018 Oct 8;9. [CrossRef]
- Bartlett AH, Hulten KG. Staphylococcus aureus Pathogenesis. Pediatric Infectious Disease Journal. 2010 Sep;29(9):860–1.
- Tong SYC, Davis JS, Eichenberger E, Holland TL, Fowler VG. Staphylococcus aureus Infections: Epidemiology, Pathophysiology, Clinical Manifestations, and Management. Clin Microbiol Rev. 2015 Jul;28(3):603–61. [CrossRef]
- Kluytmans J, van Belkum A, Verbrugh H. Nasal carriage of Staphylococcus aureus: epidemiology, underlying mechanisms, and associated risks. Clin Microbiol Rev. 1997 Jul;10(3):505–20. [CrossRef]
- Wertheim HF, Melles DC, Vos MC, van Leeuwen W, van Belkum A, Verbrugh HA, et al. The role of nasal carriage in Staphylococcus aureus infections. Lancet Infect Dis. 2005 Dec;5(12):751–62. [CrossRef]
- Wertheim HF, Vos MC, Ott A, van Belkum A, Voss A, Kluytmans JA, et al. Risk and outcome of nosocomial Staphylococcus aureus bacteraemia in nasal carriers versus non-carriers. The Lancet. 2004 Aug;364(9435):703–5. [CrossRef]
- Nouwen J, Schouten J, Schneebergen P, Snijders S, Maaskant J, Koolen M, et al. Staphylococcus aureus Carriage Patterns and the Risk of Infections Associated with Continuous Peritoneal Dialysis. J Clin Microbiol. 2006 Jun;44(6):2233–6. [CrossRef]
- Krismer B, Peschel A. Does Staphylococcus Aureus Nasal Colonization Involve Biofilm Formation? Future Microbiol. 2011 May 17;6(5):489–93. [CrossRef]
- Zacher AT, Mirza K, Thieme L, Nietzsche S, Senft C, Schwarz F. Biofilm formation of Staphylococcus aureus on various implants used for surgical treatment of destructive spondylodiscitis. Sci Rep. 2024 Aug 21;14(1):19364. [CrossRef]
- Murray CJL, Ikuta KS, Sharara F, Swetschinski L, Robles Aguilar G, Gray A, et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. The Lancet. 2022 Feb;399(10325):629–55. [CrossRef]
- Foster TJ. Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. Vol. 41, FEMS Microbiology Reviews. Oxford University Press; 2017. p. 430–49. [CrossRef]
- Gnanamani A, Hariharan P, Paul-Satyaseela M. Staphylococcus aureus: Overview of Bacteriology, Clinical Diseases, Epidemiology, Antibiotic Resistance and Therapeutic Approach. In: Frontiers in Staphylococcus aureus. InTech; 2017. [CrossRef]
- Adeoye-Isijola M, Olajuyigbe O, Adebola K, Coopoosamy R, Afolayan A. Vancomycin intermediate resistant Staphylococcus aureus in the nasal cavity of asymptomatic individuals: A potential public health challenge. Afr Health Sci. 2020;20(3):1109–17. [CrossRef]
- Conceição T, de Lencastre H, Aires-de-Sousa M. Carriage of Staphylococcus aureus among Portuguese nursing students: A longitudinal cohort study over four years of education. PLoS One. 2017 Nov 30;12(11):e0188855. [CrossRef]
- Schaumburg F, Alabi AS, Peters G, Becker K. New epidemiology of Staphylococcus aureus infection in Africa. Vol. 20, Clinical Microbiology and Infection. Blackwell Publishing Ltd; 2014. p. 589–96. [CrossRef]
- Lee AS, Huttner BD, Catho G, Harbarth S. Methicillin-Resistant Staphylococcus aureus: An Update on Prevention and Control in Acute Care Settings. Vol. 35, Infectious Disease Clinics of North America. W.B. Saunders; 2021. p. 931–52. [CrossRef]
- Lawal OU, Ayobami O, Abouelfetouh A, Mourabit N, Kaba M, Egyir B, et al. A 6-Year Update on the Diversity of Methicillin-Resistant Staphylococcus aureus Clones in Africa: A Systematic Review. Vol. 13, Frontiers in Microbiology. Frontiers Media S.A.; 2022. [CrossRef]
- Ezeh CK, Eze CN, Dibua MEU, Emencheta SC. A meta-analysis on the prevalence of resistance of Staphylococcus aureus to different antibiotics in Nigeria. Antimicrob Resist Infect Control. 2023 Apr 25;12(1):40. [CrossRef]
- Diop M, Bassoum O, Ndong A, Wone F, Ghogomu Tamouh A, Ndoye M, et al. Prevalence of multidrug-resistant bacteria in healthcare and community settings in West Africa: systematic review and meta-analysis. BMC Infect Dis. 2025 Feb 28;25(1):292. [CrossRef]
- Hasanpour AH, Sepidarkish M, Mollalo A, Ardekani A, Almukhtar M, Mechaal A, et al. The global prevalence of methicillin-resistant Staphylococcus aureus colonization in residents of elderly care centers: a systematic review and meta-analysis. Antimicrob Resist Infect Control. 2023 Jan 29;12(1):4. [CrossRef]
- Abdulgader SM, Shittu AO, Nicol MP, Kaba M. Molecular epidemiology of Methicillin-resistant Staphylococcus aureus in Africa: a systematic review. Front Microbiol. 2015 Apr 30;6.
- van der Meeren BT, Millard PS, Scacchetti M, Hermans MH, Hilbink M, Concelho TB, et al. Emergence of methicillin resistance and P anton- V alentine leukocidin positivity in hospital- and community-acquired S taphylococcus aureus infections in B eira, M ozambique. Tropical Medicine & International Health. 2014 Feb 11;19(2):169–76.
- Vubil D, Garrine M, Ruffing U, Acácio S, Sigaúque B, Alonso PL, et al. Molecular Characterization of Community Acquired Staphylococcus aureus Bacteremia in Young Children in Southern Mozambique, 2001–2009. Front Microbiol. 2017 May 4;8. [CrossRef]
- Garrine M, Quintó L, Costa SS, Messa A, Massinga AJ, Vubil D, et al. Epidemiology and clinical presentation of community-acquired Staphylococcus aureus bacteraemia in children under 5 years of age admitted to the Manhiça District Hospital, Mozambique, 2001–2019. European Journal of Clinical Microbiology & Infectious Diseases. 2023 May 18;42(5):653–9. [CrossRef]
- Onanuga A, Tc T. Nasal carriage of multi-drug resistant Staphylococcus aureus in healthy inhabitants of Amassoma in Niger delta region of Nigeria. Vol. 11, African Health Sciences. 2011.
- von Eiff C, Becker K, Machka K, Stammer H, Peters G. Nasal Carriage as a Source of Staphylococcus aureus Bacteremia. New England Journal of Medicine. 2001 Jan 4;344(1):11–6. [CrossRef]
- Robotham J V., Graves N, Cookson BD, Barnett AG, Wilson JA, Edgeworth JD, et al. Screening, isolation, and decolonisation strategies in the control of meticillin resistant Staphylococcus aureus in intensive care units: Cost effectiveness evaluation. BMJ (Online). 2011 Oct 15;343(7827). [CrossRef]
- Lee AS, de Lencastre H, Garau J, Kluytmans J, Malhotra-Kumar S, Peschel A, et al. Methicillin-resistant Staphylococcus aureus. Nat Rev Dis Primers. 2018 May 31;4(1):18033.
- Abebe A, Birhanu A. Methicillin Resistant Staphylococcus aureus: Molecular Mechanisms Underlying Drug Resistance Development and Novel Strategies to Combat. Infect Drug Resist. 2023 Dec;Volume 16:7641–62. [CrossRef]
- Tarai B, Das P, Kumar D. Recurrent Challenges for Clinicians: Emergence of Methicillin-resistant Staphylococcus aureus, Vancomycin Resistance, and Current Treatment Options. J Lab Physicians. 2013 Jul 7;5(02):071–8. [CrossRef]
- Roy PC, Shaheduzzaman Md, Sultana N, Jahid IK. Comparative Antibiotic Sensitivity Pattern of Hospital and Community Acquired <i>Staphylococcus aureus</i> Isolates of Jessore, Bangladesh. J Biosci Med (Irvine). 2015;03(10):17–23.
- K D. In-Vitro Activity of Teicoplanin against Clinical Methicillin Resistant Staphylococcus aureus Isolates. Open Access Journal of Microbiology & Biotechnology. 2017 Mar;2(1).
- Ennab R, Al-Momani W, Al-Titi R, Elayan A. Antibiotic Profile of Pathogenic Bacteria Isolated from Postsurgical Site Infections in Public Hospitals in Northern Jordan. Infect Drug Resist. 2022;15:359–66. [CrossRef]
- Brown DFJ, Wootton M, Howe RA. Antimicrobial susceptibility testing breakpoints and methods from BSAC to EUCAST. Journal of Antimicrobial Chemotherapy. 2016 Jan;71(1):3–5. [CrossRef]
- Techasupaboon T, Vasikasin V, Varothai N, Raknaisil N, Nasomsong W. Staphylococcus aureus nasal carriage and bloodstream infection among conventional hemodialysis patients in Thailand: a prospective multicenter cohort study. BMC Res Notes. 2022 Dec 1;15(1). [CrossRef]
- Kuehnert MJ, Kruszon-Moran D, Hill HA, McQuillan G, McAllister SK, Fosheim G, et al. Prevalence of Staphylococcus aureus Nasal Colonization in the United States, 2001–2002. J Infect Dis. 2006 Jan 15;193(2):172–9.
- Chigbu C. O, Ezeronye OU. Antibiotic resistant Staphylococcus aureus in Abia State of Nigeria. Afr J Biotechnol. 2003 Oct 31;2(10):374–8. [CrossRef]
- Rongpharpi SR, Hazarika NK, Kalita H. The prevalence of nasal carriage of Staphylococcus aureus among healthcare workers at a tertiary care hospital in Assam with special reference to MRSA. Journal of Clinical and Diagnostic Research. 2013 Feb 1;7(2):257–60.
- Mukhiya RK, Shrestha A, Rai SK, Panta K, Singh R, Rai G, et al. Prevalence of Methicillin-Resistant Staphylococcus aureus in Hospitals of Kathmandu Valley. Nepal J Sci Technol. 2013 Mar 10;13(2):185–90. [CrossRef]
- Adesida SA, Abioye OA, Bamiro BS, Brai BIC, Smith SI, Amisu KO, et al. Associated risk factors and pulsed field gel electrophoresis of nasal isolates of Staphylococcus aureus from medical students in a tertiary hospital in Lagos, Nigeria. Brazilian Journal of Infectious Diseases. 2007 Feb;11(1). [CrossRef]
- Abie S, Tiruneh M, Abebe W. Methicillin-resistant Staphylococcus aureus nasal carriage among janitors working in hospital and non-hospital areas: a comparative cross-sectional study. Ann Clin Microbiol Antimicrob. 2020 Dec 1;19(1). [CrossRef]
- Taqveem A, Rasool MH, Aslam B, Mujahid F, Ibrar A, Ejaz H, et al. Methicillin-Resistant Staphylococcus aureus among Students: Nasal Carriage Rate, Contributing Factors, and Antimicrobial Susceptibility. Medicina (B Aires). 2024 Sep 27;60(10):1590. [CrossRef]
- Schaumburg F, Nurjadi D, Dike AE, Ojurongbe O, Kolawole DO, Kun JF, et al. Clonal expansion accounts for an excess of antimicrobial resistance in Staphylococcus Adesola O.
- Kinabo GD, van der Ven A, Msuya LJ, Shayo AM, Schimana W, Ndaro A, et al. Dynamics of nasopharyngeal bacterial colonisation in HIV -exposed young infants in T anzania. Tropical Medicine & International Health. 2013 Mar 16;18(3):286–95.
- Ceccarelli D, Mondlane J, Sale M, Salvia AM, Folgosa E, Cappuccinelli P, et al. Sporadic methicillin resistance in community acquired Staphylococcus aureus in Mozambique. Vol. 28, THE NEW MICROBIOLOGICA. 2005.
- Chen CJ, Huang YC. New epidemiology of Staphylococcus aureus infection in Asia. Clinical Microbiology and Infection. 2014 Jul;20(7):605–23. [CrossRef]
- Olonitola OS, Olayinka BO, Sani FD. Antibiotic susceptibility of <i>Staphylococcus aureus</i> isolates from a Nigerian Federal Medical Center. Cameroon Journal of Experimental Biology. 2008 Feb 27;3(2). [CrossRef]
- Rijal KR, Pahari N, Shrestha BK, Nepal AK, Paudel B, Mahato P, et al. Prevalence of methicillin resistant Staphylococcus aureus in school children of Pokhara. Nepal Med Coll J. 2008 Sep;10(3):192–5.
- Olayinka BO, Olayinka AT, Obajuluwa AF, Onaolapo JA, Olurinola PF. Absence of meca gene in methicillin-resistant staphylococcus aureus isolates. Afr J Infect Dis. 2010 Jun 2;3(2). [CrossRef]
- Fey PD, Saïd-Salim B, Rupp ME, Hinrichs SH, Boxrud DJ, Davis CC, et al. Comparative Molecular Analysis of Community- or Hospital-Acquired Methicillin-Resistant Staphylococcus aureus. Antimicrob Agents Chemother. 2003 Jan;47(1):196–203. [CrossRef]
- Shariati A, Dadashi M, Moghadam MT, van Belkum A, Yaslianifard S, Darban-Sarokhalil D. Global prevalence and distribution of vancomycin resistant, vancomycin intermediate and heterogeneously vancomycin intermediate Staphylococcus aureus clinical isolates: a systematic review and meta-analysis. Sci Rep. 2020 Jul 29;10(1):12689. [CrossRef]
- Belete MA, Gedefie A, Alemayehu E, Debash H, Mohammed O, Gebretsadik D, et al. The prevalence of vancomycin-resistant Staphylococcus aureus in Ethiopia: a systematic review and meta-analysis. Antimicrob Resist Infect Control. 2023 Aug 30;12(1):86. [CrossRef]
- Humphries RM, Abbott AN, Hindler JA. Understanding and Addressing CLSI Breakpoint Revisions: a Primer for Clinical Laboratories. J Clin Microbiol. 2019 Jun;57(6). [CrossRef]
| Antimicrobial agente | Inhibition/Diameter zone (mm) | ||
| R (resistant) | I (intermediate) | S (sensitive) | |
| Cefoxitin (30µg) | ≤ 21 | - | ≥ 22 |
| Vancomycin (30µg) | - | - | ≥15 |
| Teicoplanin (30µg) | - | - | ≥14 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





