Submitted:
27 November 2025
Posted:
28 November 2025
You are already at the latest version
Abstract
This paper proposes a novel scheme for the origin of RNA replicase based on the replication-first stable complex evolution (SCE) model, also known as the stable complex encoding (SCE) model, and attempts to derive this scheme from the metabolism-first graded autocatalysis replication domain (GARD) model, thereby theoretically integrating the two hypotheses of the origin of life: replication-first and metabolism-first. Currently, although the replication-first model has made some progress in the artificial selection of RNA replicase, it has yet to achieve a true breakthrough. Meanwhile, metabolism-first models such as the CAS (Collectively Autocatalytic Set) and its graph version RAF (Reflexively Autocatalytic and Food-generated) models, have conducted in-depth research into the origin of metabolic networks but have failed to address the critical transformation issue from metabolism to RNA replication. This paper argues that these two hypotheses should mutually support each other: By introducing oligonucleotide assemblies and expanding the concept of composomes in the GARD model, this paper attempts to understand the general evolutionary mechanism of enzymes, thereby addressing the long-standing neglect of enzymatic catalysis in metabolism-first theories. This integrated scheme not only provides new theoretical support for the evolution of RNA replicase but also offers important insights into solving the key transition problem from chemical evolution to biological evolution.
Keywords:
1. Introduction
2. From Mutual Catalytic Networks to Replicases
3. Polymer and Metabolic Expansion of GARD
4. Repeated Assembly of Replicases
5. From Compositional Information to Sequence Information
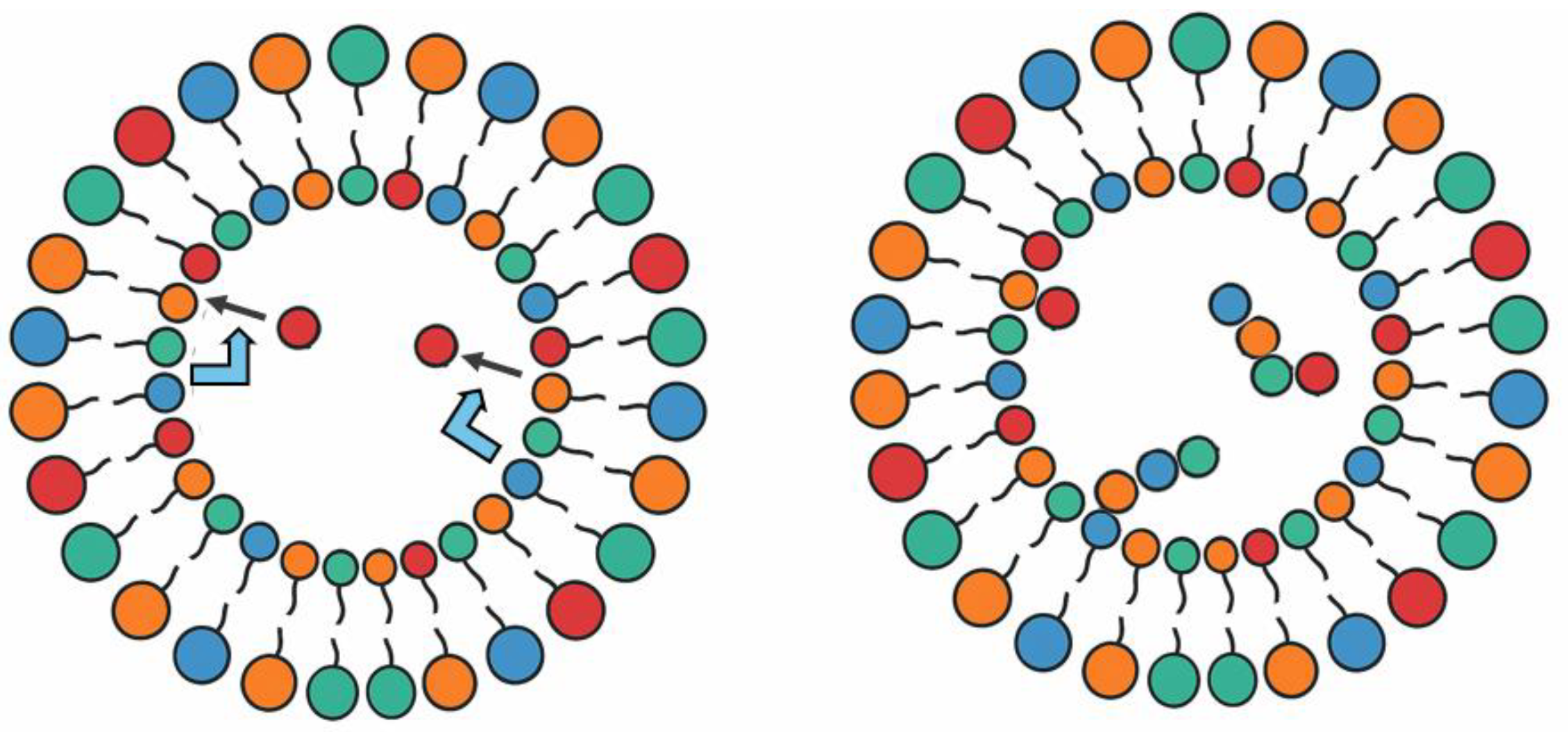
6. Attractor Properties and Selectivity of Replication Systems
7. Conclusions
Abbreviations
| GARD: graded autocatalysis replication domain |
| M-GARD: metabolic GARD |
| P-GARD: polymer GARD |
| CAS: Collectively Autocatalytic Set |
| RAF: reflexively autocatalytic and food-generated |
| SCE: stable complex evolution (encoding) |
References
- Falk, R.; Lazcano, A. The forgotten dispute: A.I. Oparin and H.J. Muller on the origin of life. History and philosophy of the life sciences 2012, 34, 373–390. [Google Scholar]
- Woese, C.R. The Genetic Code: the Molecular basis for Genetic Expression. 1967: Harper & Row.
- Orgel, L.E. Evolution of the genetic apparatus. Journal of molecular biology 1968, 38, 381–393. [Google Scholar] [CrossRef]
- Crick, F.H. The origin of the genetic code. Journal of molecular biology 1968, 38, 367–379. [Google Scholar] [CrossRef]
- Orgel, L.E. Prebiotic chemistry and the origin of the RNA world. Critical reviews in biochemistry and molecular biology 2004, 39, 99–123. [Google Scholar] [PubMed]
- Robertson, M.P.; Joyce, G.F. The Origins of the RNA World. Cold Spring Harb. Perspect. Biol. 2012, 4, a003608. [Google Scholar] [CrossRef] [PubMed]
- Bernhardt, H.S. The RNA world hypothesis: the worst theory of the early evolution of life (except for all the others)a. Biol. Direct 2012, 7, 23. [Google Scholar] [CrossRef] [PubMed]
- Higgs, P.G.; Lehman, N. The RNA World: molecular cooperation at the origins of life. Nat. Rev. Genet. 2015, 16, 7–17. [Google Scholar] [CrossRef]
- Lancet, D.; Zidovetzki, R.; Markovitch, O. Systems protobiology: origin of life in lipid catalytic networks. J. R. Soc. Interface 2018, 15, 20180159. [Google Scholar] [CrossRef]
- Orgel, L.E. Molecular replication. Nature 1992, 358, 203–209. [Google Scholar] [CrossRef]
- Segré, D.; Lancet, D. Composing life. Embo Rep. 2000, 1, 217–222. [Google Scholar] [CrossRef]
- Segré, D.; Ben-Eli, D.; Lancet, D. Compositional genomes: Prebiotic information transfer in mutually catalytic noncovalent assemblies. Proc. Natl. Acad. Sci. USA 2000, 97, 4112–4117. [Google Scholar] [CrossRef]
- Markovitch, O.; Lancet, D. Excess Mutual Catalysis Is Required for Effective Evolvability. Artif. Life 2012, 18, 243–266. [Google Scholar] [CrossRef]
- Kahana, A.; Segev, L.; Lancet, D. Attractor dynamics drives self-reproduction in protobiological catalytic networks. Cell Rep. Phys. Sci. 2023, 4. [Google Scholar] [CrossRef]
- Mulkidjanian, A.Y.; Galperin, M.Y. Physico-Chemical and Evolutionary Constraints for the Formation and Selection of First Biopolymers: Towards the Consensus Paradigm of the Abiogenic Origin of Life. Chem. Biodivers. 2007, 4, 2003–2015. [Google Scholar] [CrossRef] [PubMed]
- Hordijk, W. Autocatalytic sets: From the origin of life to the economy. BioScience 2013, 63, 877–881. [Google Scholar]
- Deng, S. The origin of genetic and metabolic systems: Evolutionary structuralinsights. Heliyon 2023, 9, e14466. [Google Scholar] [CrossRef]
- Lancet, D.; Segrè, D.; Kahana, A. Twenty years of “lipid world”: a fertile partnership with David Deamer. Life 2019, 9, 77. [Google Scholar] [CrossRef] [PubMed]
- Guseva, E.; Zuckermann, R.N.; Dill, K.A. Foldamer hypothesis for the growth and sequence differentiation of prebiotic polymers. Proc. Natl. Acad. Sci. USA 2017, 114, E7460–E7468. [Google Scholar] [CrossRef] [PubMed]
- Derr, J.; Manapat, M.L.; Rajamani, S.; Leu, K.; Xulvi-Brunet, R.; Joseph, I.; Nowak, M.A.; Chen, I.A. Prebiotically plausible mechanisms increase compositional diversity of nucleic acid sequences. Nucleic Acids Res. 2012, 40, 4711–4722. [Google Scholar] [CrossRef]
- Egel, R. Life’s Order, Complexity, Organization, and Its Thermodynamic–Holistic Imperatives. Life 2012, 2, 323–363. [Google Scholar] [CrossRef]
- Åqvist, J.; et al. Entropy and enzyme catalysis. Accounts of chemical research 2017, 50, 199–207. [Google Scholar] [CrossRef]
- Bowman, J.C.; Hud, N.V.; Williams, L.D. The Ribosome Challenge to the RNA World. J. Mol. Evol. 2015, 80, 143–161. [Google Scholar] [CrossRef]
- Shan, S.-O.; Kravchuk, A.V.; Piccirilli, J.A.; Herschlag, D. Defining the Catalytic Metal Ion Interactions in the Tetrahymena Ribozyme Reaction. Biochemistry 2001, 40, 5161–5171. [Google Scholar] [CrossRef]
- Klein, D.J.; Moore, P.B.; Steitz, T.A. The contribution of metal ions to the structural stability of the large ribosomal subunit. RNA 2004, 10, 1366–1379. [Google Scholar] [CrossRef]
- Steitz, T.A. A mechanism for all polymerases. Nature 1998, 391, 231–232. [Google Scholar] [CrossRef]
- Ferris, J.P.; Hill, A.R.; Liu, R.; Orgel, L.E. Synthesis of long prebiotic oligomers on mineral surfaces. Nature 1996, 381, 59–61. [Google Scholar] [CrossRef] [PubMed]
- Adamala, K.; Engelhart, A.E.; Szostak, J.W. Generation of Functional RNAs from Inactive Oligonucleotide Complexes by Non-enzymatic Primer Extension. J. Am. Chem. Soc. 2015, 137, 483–489. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Higgs, P.G. Comparison of the Roles of Nucleotide Synthesis, Polymerization, and Recombination in the Origin of Autocatalytic Sets of RNAs. Astrobiology 2011, 11, 895–906. [Google Scholar] [CrossRef] [PubMed]
- Briones, C.; Stich, M.; Manrubia, S.C. The dawn of the RNA World: Toward functional complexity through ligation of random RNA oligomers. RNA 2009, 15, 743–749. [Google Scholar] [CrossRef]
- Wu, M.; Higgs, P.G. Compositional Inheritance: Comparison of Self-assembly and Catalysis. Discov. Life 2008, 38, 399–418. [Google Scholar] [CrossRef]
- Kahana, A.; Lancet, D.; Palmai, Z. Micellar Composition Affects Lipid Accretion Kinetics in Molecular Dynamics Simulations: Support for Lipid Network Reproduction. Life 2022, 12, 955. [Google Scholar] [CrossRef] [PubMed]
- Yaniv, R.; Lancet, D. Life's Emergence by Protocellular Mutually Catalytic Networks; CRC Press, 2023; pp. 239–263. [Google Scholar]
- Sharov, A.A. Coenzyme world model of the origin of life. Biosystems 2016, 144, 8–17. [Google Scholar] [CrossRef]
- Hunding, A.; Kepes, F.; Lancet, D.; Minsky, A.; Norris, V.; Raine, D.; Sriram, K.; Root-Bernstein, R. Compositional complementarity and prebiotic ecology in the origin of life. BioEssays 2006, 28, 399–412. [Google Scholar] [CrossRef]
- Eigen, M. Selforganization of matter and the evolution of biological macromolecules. Sci. Nat. 1971, 58, 465–523. [Google Scholar] [CrossRef]
- Shapiro, R. A Simpler Origin for Life. Sci. Am. 2007, 296, 46–53. [Google Scholar] [CrossRef]
- Nowak, M.A.; Ohtsuki, H. Prevolutionary dynamics and the origin of evolution. Proc. Natl. Acad. Sci. 2008, 105, 14924–14927. [Google Scholar] [CrossRef]
- Engelhart, A.E.; Hud, N.V. Primitive Genetic Polymers. Cold Spring Harb. Perspect. Biol. 2010, 2, a002196–a002196. [Google Scholar] [CrossRef]
- Hud, N.V.; et al. The origin of RNA and “my grandfather’s axe”. Chemistry & biology 2013, 20, 466–474. [Google Scholar]
- Lai, Y.-C.; Liu, Z.; Chen, I.A. Encapsulation of ribozymes inside model protocells leads to faster evolutionary adaptation. Proc. Natl. Acad. Sci. USA 2021, 118, e2025054118. [Google Scholar] [CrossRef] [PubMed]
- Segré, D.; Ben-Eli, D.; Deamer, D.W.; Lancet, D. The Lipid World. Discov. Life 2001, 31, 119–145. [Google Scholar] [CrossRef] [PubMed]
- Erastova, V.; Degiacomi, M.T.; Fraser, D.G.; Greenwell, H.C. Mineral surface chemistry control for origin of prebiotic peptides. Nat. Commun. 2017, 8, 2033. [Google Scholar] [CrossRef]
- Vasas, V.; Szathmáry, E.; Santos, M. Lack of evolvability in self-sustaining autocatalytic networks constraints metabolism-first scenarios for the origin of life. Proc. Natl. Acad. Sci. USA 2010, 107, 1470–1475. [Google Scholar] [CrossRef]
- Shenhav, B.; Bar-Even, A.; Kafri, R.; Lancet, D. Polymer Gard: Computer Simulation of Covalent Bond Formation in Reproducing Molecular Assemblies. Discov. Life 2005, 35, 111–133. [Google Scholar] [CrossRef]
- Ohtsuki, H.; Nowak, M.A. Prelife catalysts and replicators. Proc. R. Soc. B: Biol. Sci. 2009, 276, 3783–3790. [Google Scholar] [CrossRef] [PubMed]
- Anet, F.A. The place of metabolism in the origin of life. Current opinion in chemical biology 2004, 8, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Higgs, P.G. The origin of life is a spatially localized stochastic transition. Biol. Direct 2012, 7, 42. [Google Scholar] [CrossRef] [PubMed]
- Fendler, J.H. Polymerized Surfactant Vesicles: Novel Membrane Mimetic Systems. Science 1984, 223, 888–894. [Google Scholar] [CrossRef]
- Cech, T.R. A model for the RNA-catalyzed replication of RNA. Proc. Natl. Acad. Sci. USA 1986, 83, 4360–4363. [Google Scholar] [CrossRef]
- Segré, D.; Shenhav, B.; Kafri, R.; Lancet, D. The Molecular Roots of Compositional Inheritance. J. Theor. Biol. 2001, 213, 481–491. [Google Scholar] [CrossRef]
- Vasas, V.; Fernando, C.; Santos, M.; Kauffman, S.; Szathmáry, E. Evolution before genes. Biol. Direct 2012, 7, 1. [Google Scholar] [CrossRef]
- Higgs, P.G. Chemical Evolution and the Evolutionary Definition of Life. J. Mol. Evol. 2017, 84, 225–235. [Google Scholar] [CrossRef]
- Zhuang, X.; Bartley, L.E.; Babcock, H.P.; Russell, R.; Ha, T.; Herschlag, D.; Chu, S. A Single-Molecule Study of RNA Catalysis and Folding. Science 2000, 288, 2048–2051. [Google Scholar] [CrossRef]
- Fedor, M.J. Structure and function of the hairpin ribozyme. J. Mol. Biol. 2000, 297, 269–291. [Google Scholar] [CrossRef]
- Lafontaine, D.A.; Norman, D.G.; Lilley, D.M.J. The structure and active site of the Varkud satellite ribozyme. Biochem. Soc. Trans. 2002, 30, 1170–1175. [Google Scholar] [CrossRef]
- Haslinger, C.; Stadler, P.F. RNA Structures with Pseudo-knots: Graph-theoretical, Combinatorial, and Statistical Properties. Bull. Math. Biol. 1999, 61, 437–467. [Google Scholar] [CrossRef] [PubMed]
- Schuster, P.; Fontana, W.; Stadler, P.F.; Hofacker, I.L. From sequences to shapes and back: a case study in RNA secondary structures. Proc. R. Soc. B: Biol. Sci. 1994, 255, 279–284. [Google Scholar] [CrossRef]
- Schuster, P. Genotypes with phenotypes: Adventures in an RNA toy world. Biophys. Chem. 1997, 66, 75–110. [Google Scholar] [CrossRef] [PubMed]
- Kun, Á.; Szilágyi, A.; Könnyű, B.; Boza, G.; Zachar, I.; Szathmáry, E. The dynamics of the RNA world: insights and challenges. Ann. New York Acad. Sci. 2015, 1341, 75–95. [Google Scholar] [CrossRef] [PubMed]
- Kuhn, H.; Kuhn, C. Diversified World: Drive to Life's Origin?! Angew. Chem. Int. Ed. Engl. 2003, 42, 262–266. [Google Scholar] [CrossRef]
- Deamer, D.W. Origin of life: What everyone needs to know®. 2020: Oxford University Press.
- Nakayama, T.J.; A Rodrigues, F.; Neumaier, N.; Marcolino-Gomes, J.; Molinari, H.B.C.; Santiago, T.R.; Formighieri, E.F.; Basso, M.F.; Farias, J.R.B.; Emygdio, B.M.; et al. Insights into soybean transcriptome reconfiguration under hypoxic stress: Functional, regulatory, structural, and compositional characterization. PLOS ONE 2017, 12, e0187920. [Google Scholar] [CrossRef]
- Baum, D.A.; Peng, Z.; Dolson, E.; Smith, E.; Plum, A.M.; Gagrani, P. The ecology–evolution continuum and the origin of life. J. R. Soc. Interface 2023, 20, 20230346. [Google Scholar] [CrossRef]
- Kahana, A.; Lancet, D. Self-reproducing catalytic micelles as nanoscopic protocell precursors. Nat. Rev. Chem. 2021, 5, 870–878. [Google Scholar] [CrossRef]
- Marahiel, M.A. Working outside the protein-synthesis rules: insights into non-ribosomal peptide synthesis. Journal of peptide science: an official publication of the European Peptide Society 2009, 15, 799–807. [Google Scholar] [CrossRef]
- Markovitch, O.; Lancet, D. Multispecies population dynamics of prebiotic compositional assemblies. J. Theor. Biol. 2014, 357, 26–34. [Google Scholar] [CrossRef]
- Baum, D.A. The origin and early evolution of life in chemical composition space. J. Theor. Biol. 2018, 456, 295–304. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente, I.M.; et al. Attractor metabolic networks. PLoS One 2013, 8, e58284. [Google Scholar] [CrossRef] [PubMed]
- James, A.; Swann, K.; Recce, M. Cell Behaviour as a Dynamic Attractor in the Intracellular Signalling System. J. Theor. Biol. 1999, 196, 269–288. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.; Eichler, G.; Bar-Yam, Y.; Ingber, D.E. Cell Fates as High-Dimensional Attractor States of a Complex Gene Regulatory Network. Phys. Rev. Lett. 2005, 94, 128701. [Google Scholar] [CrossRef]
- Bui, T.T.; Selvarajoo, K. Attractor Concepts to Evaluate the Transcriptome-wide Dynamics Guiding Anaerobic to Aerobic State Transition in Escherichia coli. Sci. Rep. 2020, 10, 5878. [Google Scholar] [CrossRef] [PubMed]
- Ghaffarizadeh, A.; Podgorski, G.J.; Flann, N.S. Applying attractor dynamics to infer gene regulatory interactions involved in cellular differentiation. Biosystems 2017, 155, 29–41. [Google Scholar] [CrossRef]
- Kashiwagi, A.; Urabe, I.; Kaneko, K.; Yomo, T. Adaptive Response of a Gene Network to Environmental Changes by Fitness-Induced Attractor Selection. PLOS ONE 2006, 1, e49. [Google Scholar] [CrossRef]
- Kauffman, S.A. Requirements for evolvability in complex systems: orderly components and frozen dynamics. Physica D 1990, 42, 135–152. [Google Scholar] [CrossRef]
- Eigen, M.; Schuster, P. , The hypercycle: a principle of natural self-organization part C: the realistic hypercycle. The Science of Nature 1978, 65, 341–369. [Google Scholar] [CrossRef]
- Kahana, A.; Lancet, D. Protobiotic Systems Chemistry Analyzed by Molecular Dynamics. Life 2019, 9, 38. [Google Scholar] [CrossRef] [PubMed]
- Clarke, B.L.; Jeffries, C. Chemical reaction networks with finite attractor regions. J. Chem. Phys. 1985, 82, 3107–3117. [Google Scholar] [CrossRef]
- Kauffman, S.A. The Origins of Order: Self-Organization and Selection in Evolution; Oxford University Press: Oxford, UK, 1993. [Google Scholar]
- Ruelle, D. Small random perturbations of dynamical systems and the definition of attractors. Commun. Math. Phys. 1981, 82, 137–151. [Google Scholar] [CrossRef]
- Hou, X.; et al. Attractor–a new turning point in drug discovery. Drug Design, Development and Therapy 2019, 13, 2957–2968. [Google Scholar] [CrossRef]
- Kasperski, A.; Kasperska, R. Study on attractors during organism evolution. Sci. Rep. 2021, 11, 9637. [Google Scholar] [CrossRef]
- Davila-Velderrain, J.; Martinez-Garcia, J.C.; Alvarez-Buylla, E.R. Modeling the epigenetic attractors landscape: toward a post-genomic mechanistic understanding of development. Front. Genet. 2015, 6, 160. [Google Scholar] [CrossRef]
- Lian, H.; He, S.; Chen, C.; Yan, X. Flow Cytometric Analysis of Nanoscale Biological Particles and Organelles. Annu. Rev. Anal. Chem. 2019, 12, 389–409. [Google Scholar] [CrossRef]
- Borhani, D.W.; Shaw, D.E. The future of molecular dynamics simulations in drug discovery. J. Comput. Mol. Des. 2012, 26, 15–26. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



