Submitted:
27 November 2025
Posted:
28 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Data Representation and Notation
2.2. Base Space Construction and Homology Computation
2.3. Fiber Construction and Fiber Homology Aggregation
2.4. Total Space Embedding and Persistent Homology
2.5. Discrete Connection and Transition Map Estimation
2.6. Holonomy, Curvature, Characteristic Class Proxies and Signature Assembly
2.7. Learning Pipeline and Software Tools
3. Theoretical Guarantees
3.1. Definition of Distances
3.2. Stability of the Discrete Connection
3.3. Stability of Holonomy
3.4. Stability of Characteristic-Class Proxies
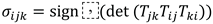
3.5. Invariance Properties
- global geometric structure,
- local feature distributions,
- transition-map organization,
- holonomy or curvature,
- cocycle parities.
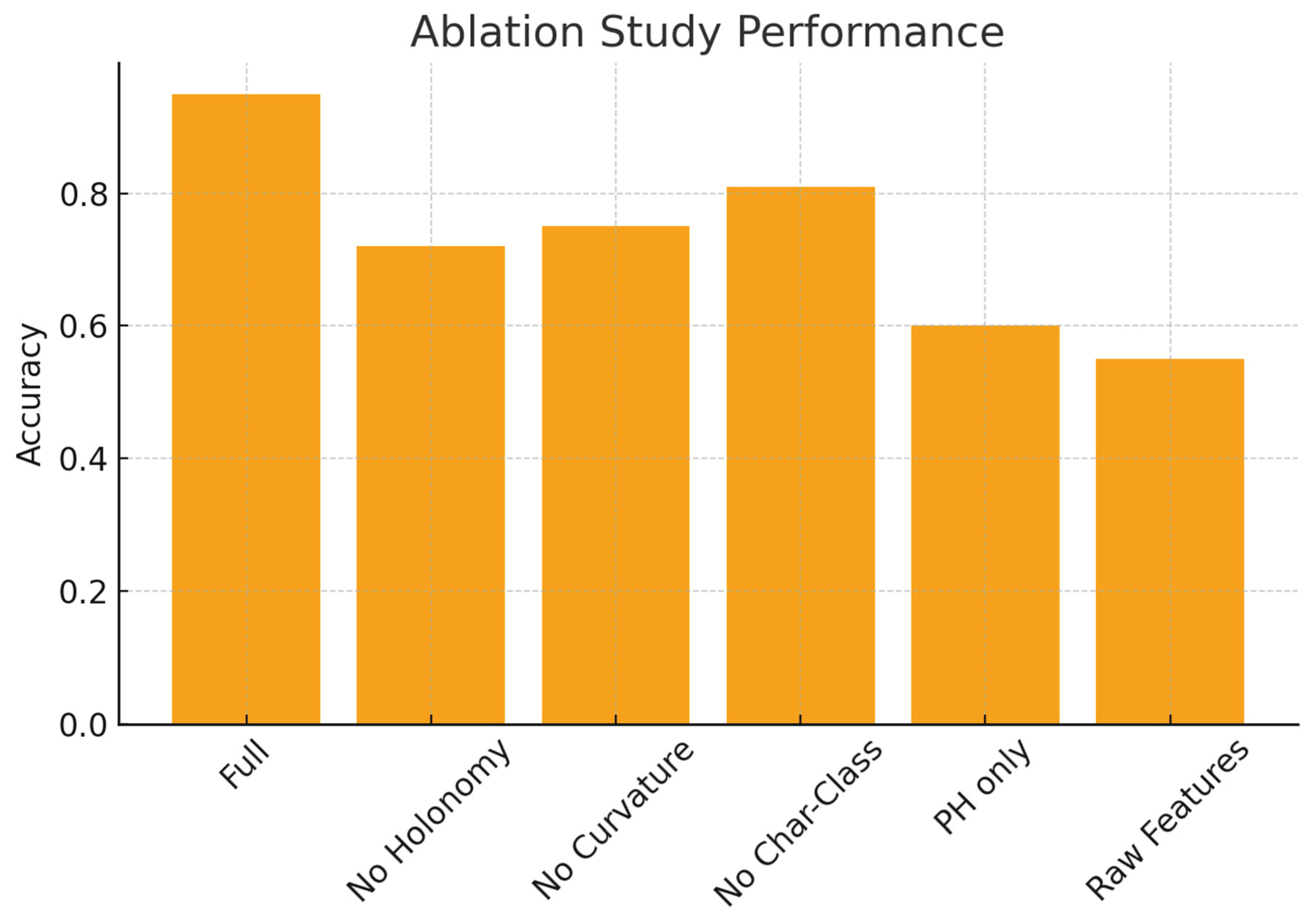
4. Experiments
5. Conclusions
Supplementary Materials
Funding
Ethics approval and consent to participate
Consent for publication
Availability of data and materials
Competing interests
Acknowledgments
Authors' contributions
Declaration of generative AI and AI-assisted technologies in the writing process
References
- Aggarwal, Manas, and Vipul Periwal. “Dory: Computation of Persistence Diagrams up to Dimension Two for Vietoris–Rips Filtrations of Large Data Sets.” Journal of Computational Science 79 (2024): 102290. [CrossRef]
- Benjamin, K., L. Mukta, G. Moryoussef, C. Uren, H. A. Harrington, U. Tillmann, and A. Barbensi. “Homology of Homologous Knotted Proteins.” Journal of the Royal Society Interface 20, no. 201 (2023): 20220727. [CrossRef]
- Bukkuri, Abhijith, Noemi Andor, and Ian K. Darcy. “Applications of Topological Data Analysis in Oncology.” Frontiers in Artificial Intelligence 4 (2021): 659037. [CrossRef]
- Chazal, Frédéric, and Bertrand Michel. “An Introduction to Topological Data Analysis: Fundamental and Practical Aspects for Data Scientists.” Frontiers in Artificial Intelligence 4 (2021): 667963. [CrossRef]
- Diksha, and S. Biswas. “Prediction of Imminent Failure Using Supervised Learning in a Fiber Bundle Model.” Physical Review E 106, no. 2–2 (2022): 025003. [CrossRef]
- Eadie, Matthew, Jianan Liao, Wafa Ageeli, Ghulam Nabi, and Nikola Krstajić. “Fiber Bundle Image Reconstruction Using Convolutional Neural Networks and Bundle Rotation in Endomicroscopy.” Sensors 23, no. 5 (2023): 2469. [CrossRef]
- El-Yaagoubi, Abdel B., Moo K. Chung, and Hernando Ombao. “Topological Data Analysis for Multivariate Time Series Data.” Entropy 25, no. 11 (2023): 1509. [CrossRef]
- Gao, Q., F. Li, Q. Wang, X. Gao, and D. Tao. “Manifold Based Multi-View K-Means.” IEEE Transactions on Pattern Analysis and Machine Intelligence 47, no. 4 (2025): 3175–82. [CrossRef]
- Hu, X., M. Zhu, Z. Feng, and L. Stanković. “Manifold-Based Shapley Explanations for High Dimensional Correlated Features.” Neural Networks 180 (2024): 106634. [CrossRef]
- Jiang, Qiangrong, and Qiang Guang. “Graph Kernels Combined with the Neural Network on Protein Classification.” Journal of Bioinformatics and Computational Biology 17, no. 5 (2019): 1950030. [CrossRef]
- Kumar, S., A. R. Shovon, and G. Deshpande. “The Robustness of Persistent Homology of Brain Networks to Data Acquisition-Related Non-Neural Variability in Resting State fMRI.” Human Brain Mapping 44, no. 13 (2023): 4637–51. [CrossRef]
- Laubenbacher, Reinhard, and Alan Hastings. “Topological Data Analysis.” Bulletin of Mathematical Biology 81, no. 7 (2019): 2051. [CrossRef]
- Maîtrejean, Driss. “Parallel Transport, Connections, and Covariant Derivatives.” In Riemannian Geometry and Geometric Analysis. Universitext. Berlin: Springer, 2008. [CrossRef]
- Maîtrejean, Driss. Equivariant Connections and Their Applications to Yang–Mills Equations. arXiv:2406.04171 [math-ph], 2024. https://arxiv.org/abs/2406.04171.
- Martino, Andrea, and Alessandro Rizzi. “(Hyper)Graph Kernels over Simplicial Complexes.” Entropy 22, no. 10 (2020): 1155. [CrossRef]
- Xu, Xinyue, Nicolas Drougard, and Raphaël N. Roy. “Topological Data Analysis as a New Tool for EEG Processing.” Frontiers in Neuroscience 15 (2021): 761703. [CrossRef]
- Nagai, Yuki, Hiroshi Ohno, and Akio Tomiya. CASK: A Gauge Covariant Transformer for Lattice Gauge Theory. arXiv:2501.16955 [hep-lat], 2025. https://arxiv.org/abs/2501.16955.
- Paulsen, Jonas, Ola Gramstad, and Philippe Collas. “Manifold Based Optimization for Single-Cell 3D Genome Reconstruction.” PLoS Computational Biology 11, no. 8 (2015): e1004396. [CrossRef]
- Qin, Zhen, Xinyu Lu, Dong Liu, Xin Nie, Yichen Yin, Jianbing Shen, and Andrew C. Loui. “Reformulating Graph Kernels for Self-Supervised Space-Time Correspondence Learning.” IEEE Transactions on Image Processing 32 (2023): 6543–57. [CrossRef]
- Sauer, Tilman. “Modeling Parallel Transport.” In Model and Mathematics: From the 19th to the 21st Century, edited by Michael Friedman and Karin Krauthausen. Trends in the History of Science. Cham: Birkhäuser, 2022. [CrossRef]
- Sekmen, A., and B. Bilgin. “Manifold-Based Approach for Neural Network Robustness Analysis.” Communications Engineering 3, no. 1 (2024): 118. [CrossRef]
- Shao, Jie, Jing Zhang, Xiaohui Huang, Rongguang Liang, and Kevin Barnard. “Fiber Bundle Image Restoration Using Deep Learning.” Optics Letters 44, no. 5 (2019): 1080–83. [CrossRef]
- Shams, Behnam, Katharina Reisch, Peter Vajkoczy, Christoph Lippert, Thomas Picht, and L. S. Fekonja. “Improved Prediction of Glioma-Related Aphasia by Diffusion MRI Metrics, Machine Learning, and Automated Fiber Bundle Segmentation.” Human Brain Mapping 44, no. 12 (2023): 4480–97. [CrossRef]
- Skaf, Yara, and Reinhard Laubenbacher. “Topological Data Analysis in Biomedicine: A Review.” Journal of Biomedical Informatics 130 (2022): 104082. [CrossRef]
- Smith, Andrew D., Gregory J. Donley, Emanuela Del Gado, and Victor M. Zavala. “Topological Data Analysis for Particulate Gels.” ACS Nano 18, no. 42 (2024): 28622–35. [CrossRef]
- Song, J., J. McNeany, Y. Wang, T. Daley, A. Stecenko, and R. Kamaleswaran. “Riemannian Manifold-Based Geometric Clustering of Continuous Glucose Monitoring to Improve Personalized Diabetes Management.” Computers in Biology and Medicine 183 (2024): 109255. [CrossRef]
- Tsamir-Rimon, M., and E. Borenstein. “A Manifold-Based Framework for Studying the Dynamics of the Vaginal Microbiome.” NPJ Biofilms and Microbiomes 9, no. 1 (2023): 102. [CrossRef]
- Wang, Bin, and Guo-Wei Wei. “Object-Oriented Persistent Homology.” Journal of Computational Physics 305 (2016): 276–99. [CrossRef]
- Vijayan, Pranav, Yash Chandak, Maheswaran M. Khapra, Srinivasan Parthasarathy, and Balaraman Ravindran. “Scaling Graph Propagation Kernels for Predictive Learning.” Frontiers in Big Data 5 (2022): 616617. [CrossRef]
- Wadhwa, Rohan R., Daniel F. K. Williamson, Aditya Dhawan, and J. G. Scott. “TDAstats: R Pipeline for Computing Persistent Homology in Topological Data Analysis.” Journal of Open Source Software 3, no. 28 (2018): 860. [CrossRef]
- Wang, Zhen, Fang Liu, Shuang Shi, Shen Xia, Fei Peng, Li Wang, Sheng Ai, and Zhi Xu. “Automatic Epileptic Seizure Detection Based on Persistent Homology.” Frontiers in Physiology 14 (2023): 1227952. [CrossRef]
- Zhang, Yunpeng, Jian Peng, Xiaoli Yuan, Lei Zhang, Dongming Zhu, Ping Hong, Jiaju Wang, Qiang Liu, and Wei Liu. “MFCIS: An Automatic Leaf-Based Identification Pipeline for Plant Cultivars Using Deep Learning and Persistent Homology.” Horticulture Research 8, no. 1 (2021): 172. [CrossRef]
- Zhang, Wei, Shen Xia, Xin Tang, Xia Zhang, Di Liang, and Yue Wang. “Topological Analysis of Functional Connectivity in Parkinson’s Disease.” Frontiers in Neuroscience 17 (2023): 1236128. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



