Submitted:
23 November 2025
Posted:
26 November 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Implementation of EfficientNetB4 for multiclass skin disease classification
- Comprehensive performance evaluation across six distinct skin conditions
- Analysis of class-wise performance and identification of challenging cases
- Discussion of practical implications for clinical deployment
2. Related Work
3. Methodology
3.1. Data Preprocessing
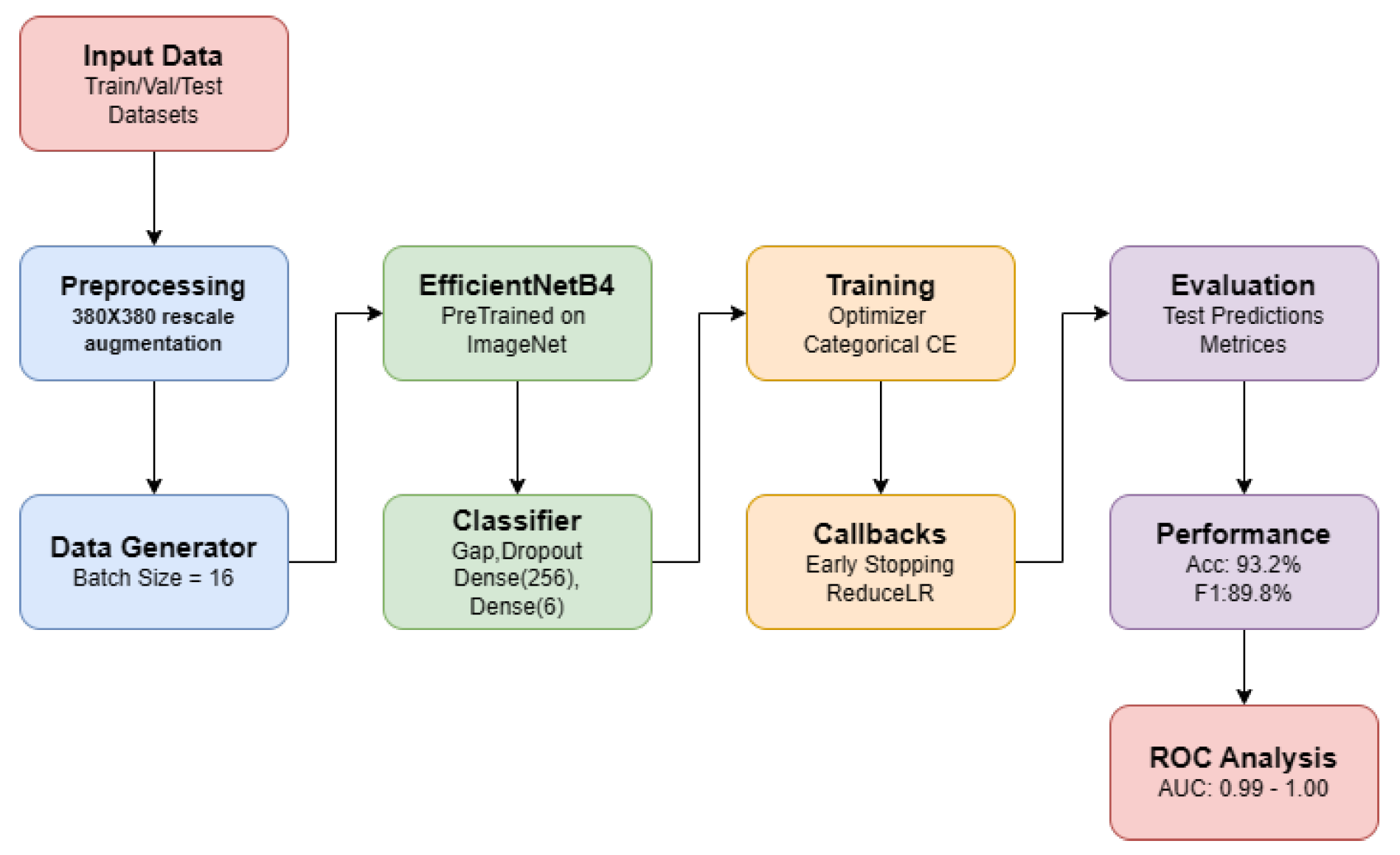
3.2. Model Architecture
- EfficientNetB4 base (pre-trained on ImageNet, without top layer)
- Global Average Pooling layer
- Dropout layer (rate = 0.5) for regularization
- Dense layer with 256 units and ReLU activation
- Output Dense layer with 6 units and softmax activation
3.3. Training Configuration
- Optimizer: Adam with default learning rate (0.001)
- Loss function: Categorical cross-entropy
- Batch size: 16
- Maximum epochs: 30
- Early stopping with a patience of 5 epochs, monitoring validation loss
- Learning rate reduction with a patience of 2 epochs and a factor of 0.3
- Model checkpoint to save the best-performing model based on validation accuracy
4. Experimental Results
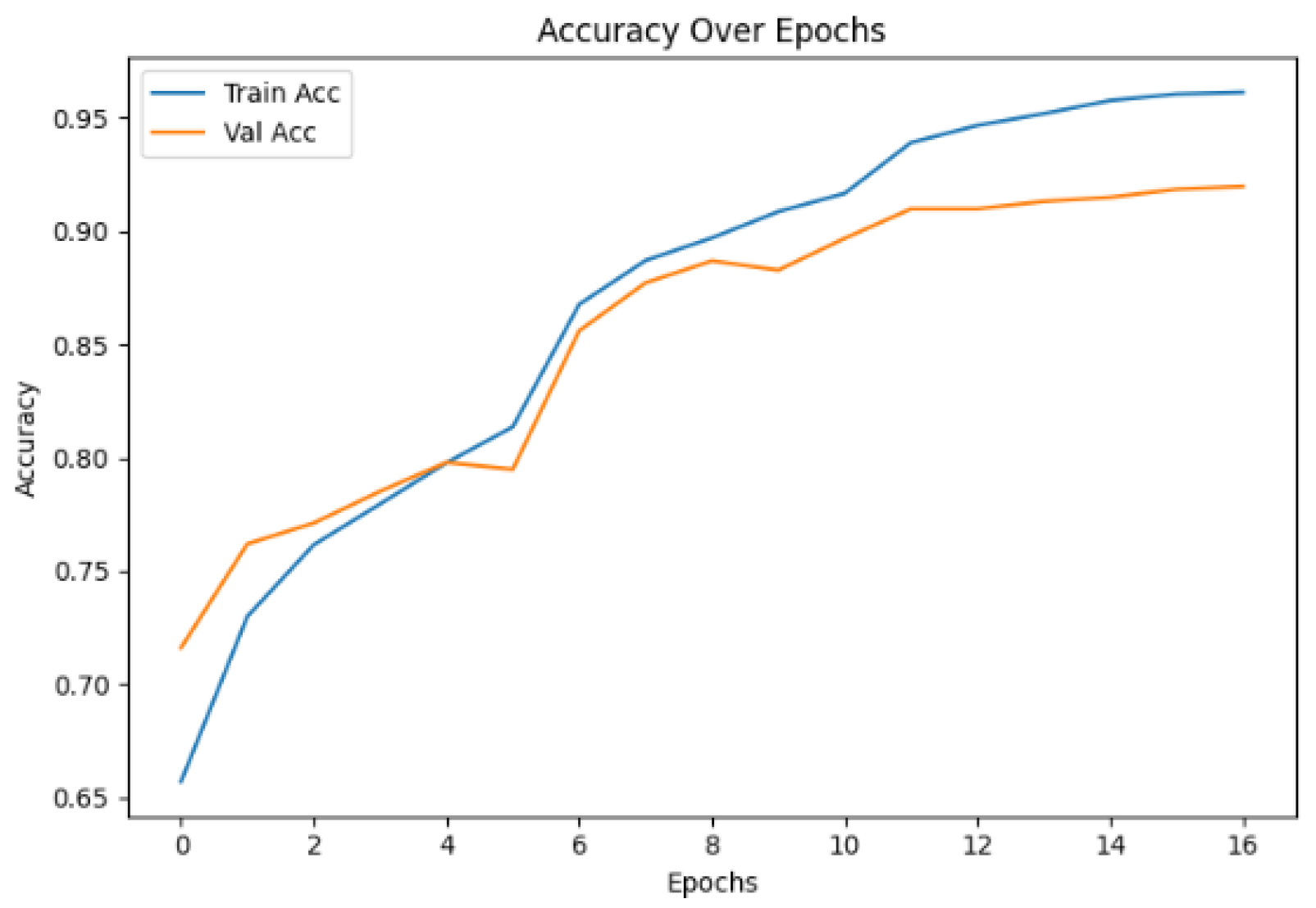
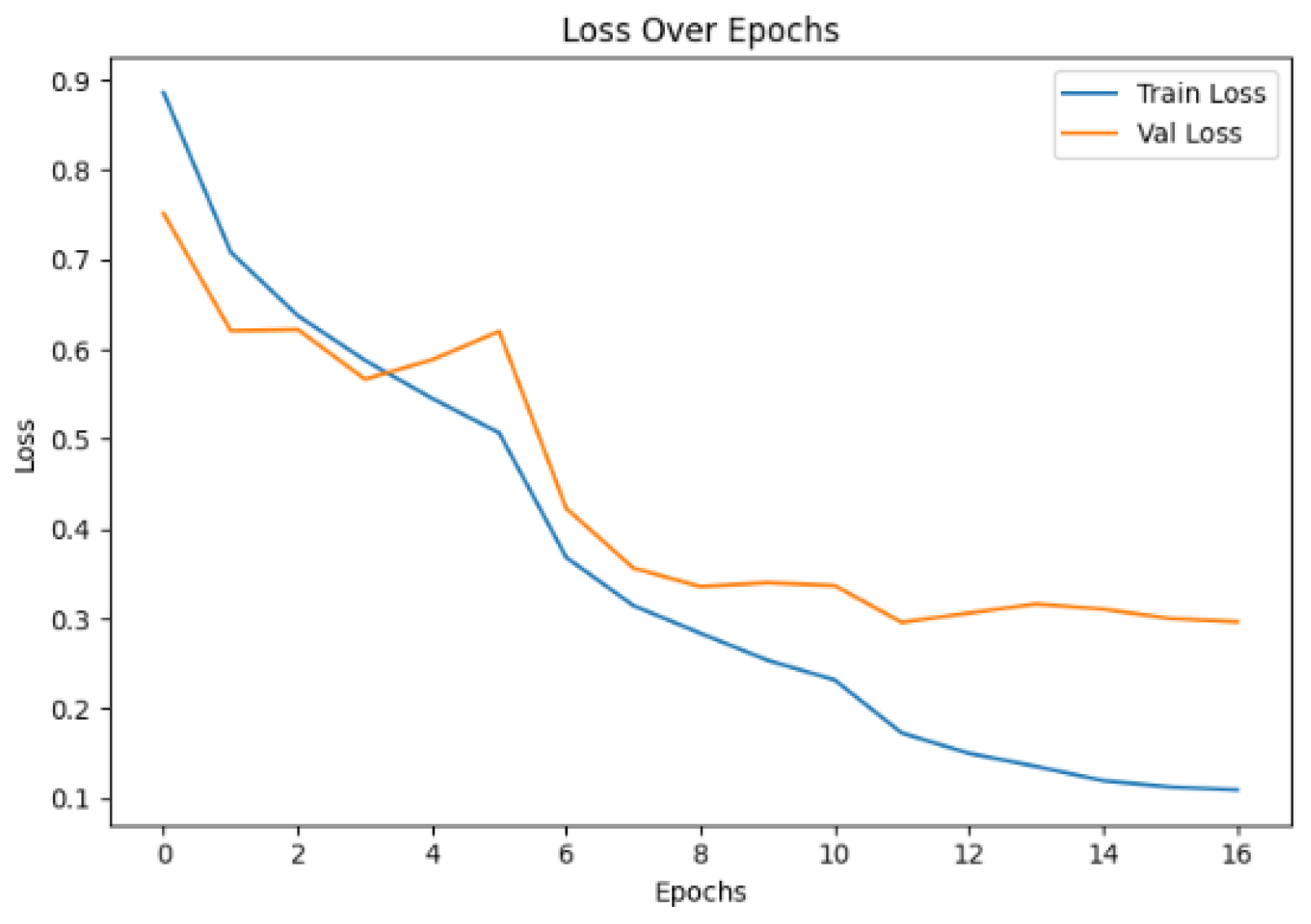
4.1. Training Performance
4.2. Test Set Performance
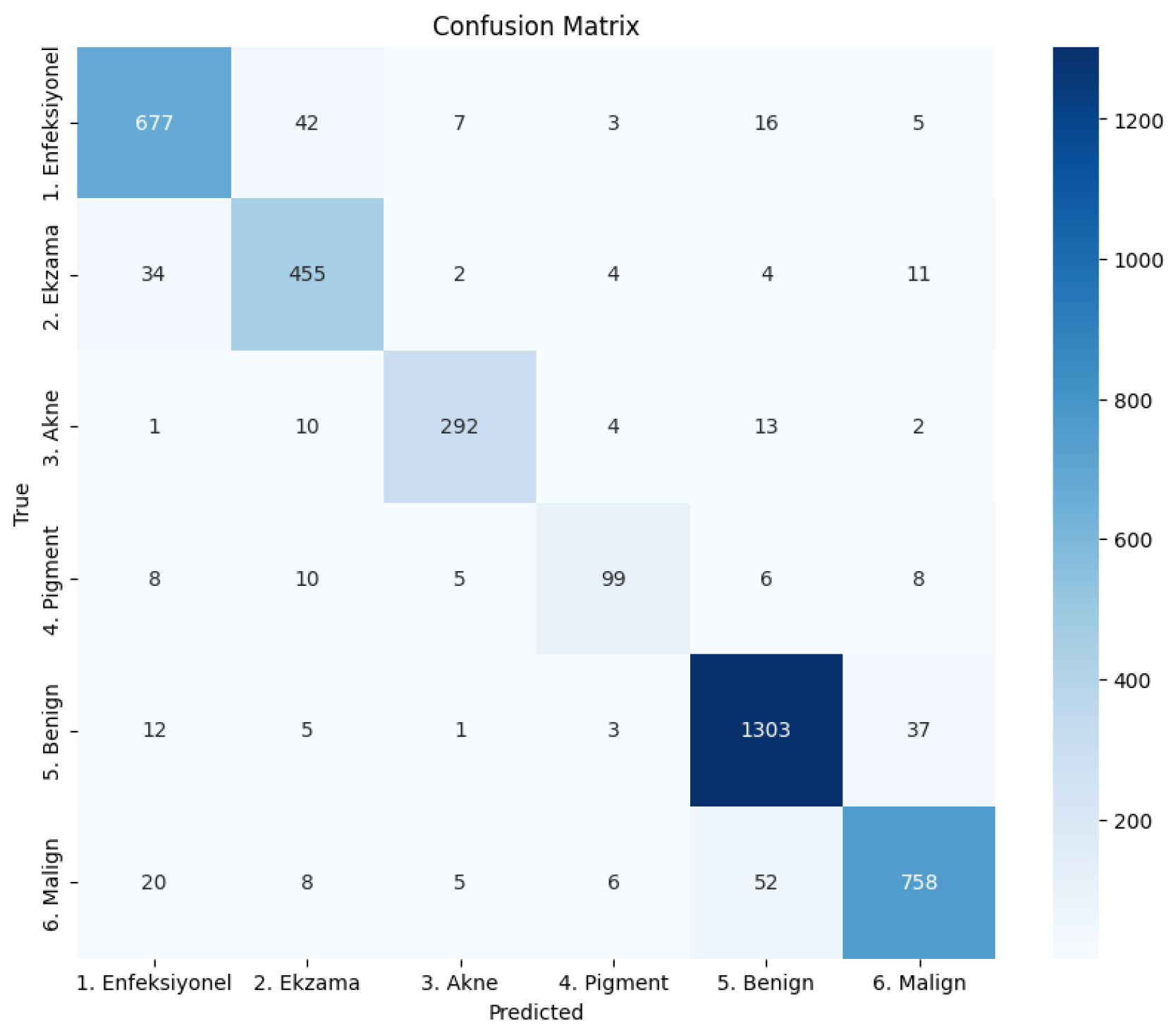
4.3. Confusion Matrix
- The Benign class has the highest number of correctly classified instances (1303) with minimal misclassifications.
- There is some confusion between Ekzama and Enfeksiyonel categories, with 34 instances of Ekzama misclassified as Enfeksiyonel and 42 instances of Enfeksiyonel misclassified as Ekzama.
- The Pigment class shows significant misclassifications, particularly with Malign (8 instances) and Enfeksiyonel (8 instances), consistent with its lower recall value.
- Misclassifications between Benign and Malign are relatively low (52 Malign samples misclassified as Benign, and 37 Benign samples misclassified as Malign), which is encouraging given the clinical importance of distinguishing between these two categories.
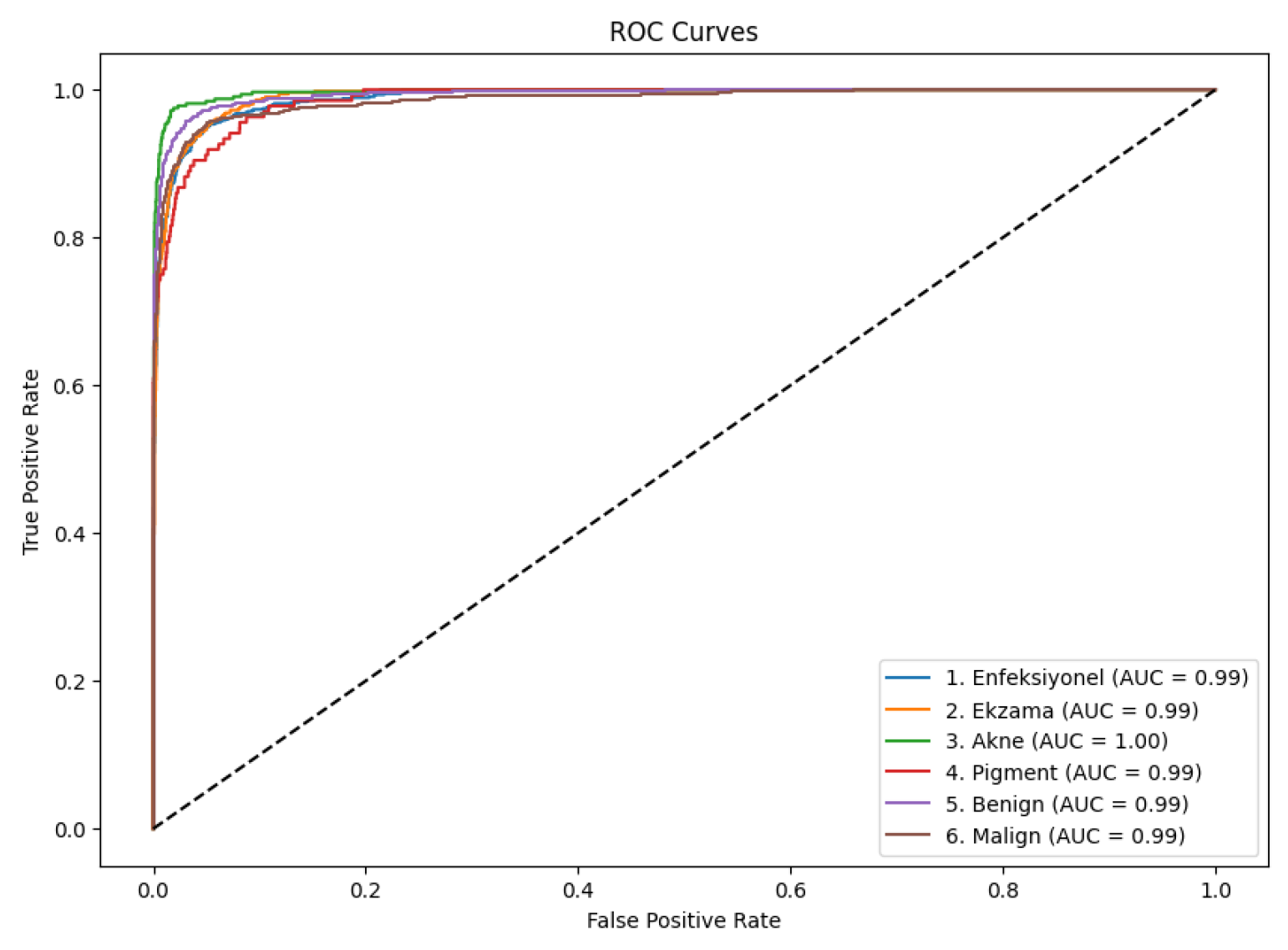
4.4. ROC Curve
5. Results
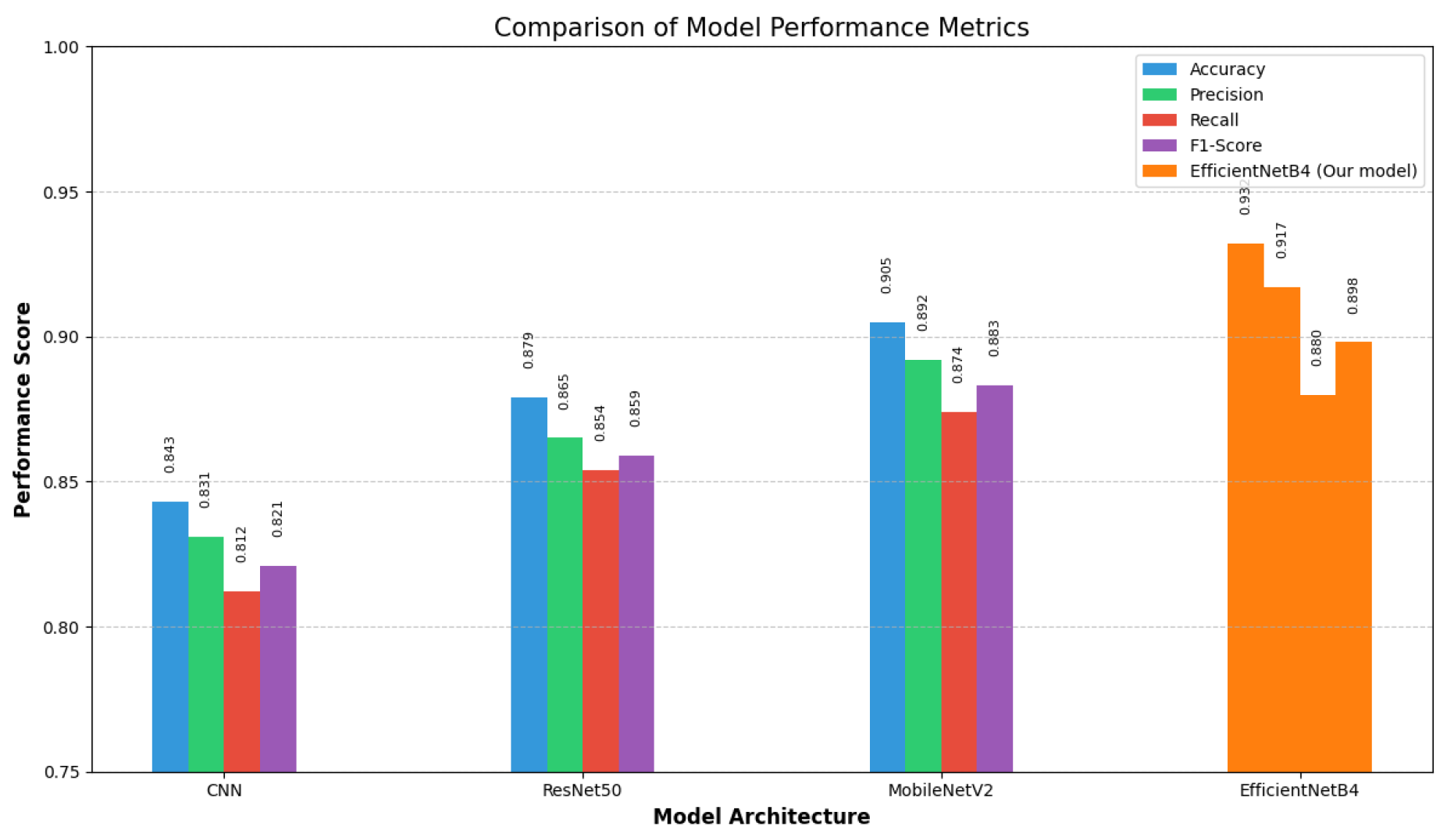
6. Comparative Analysis
7. Conclusions
References
- Hay, R. J.; et al. The global burden of skin disease in 2010: an analysis of the prevalence and impact of skin conditions. Journal of Investigative Dermatology 2014, 134, 1527–1534. [Google Scholar] [CrossRef]
- Argenziano, G.; et al. Dermoscopy of pigmented skin lesions: Results of a consensus meeting via the Internet. Journal of the American Academy of Dermatology 2003, 48, 679–693. [Google Scholar] [CrossRef]
- N. C. F. Codella et al., “Skin lesion analysis toward melanoma detection: A challenge at the 2017 International Symposium on Biomedical Imaging (ISBI), hosted by the International Skin Imaging Collaboration (ISIC),” in IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), 2018, pp. 168-172.
- G. Litjens et al., “A survey on deep learning in medical image analysis,” Medical Image Analysis, vol. 42, pp. 60-88, 2017.
- M. Tan and Q. V. Le, “EfficientNet: Rethinking model scaling for convolutional neural networks,” in Proceedings of the 36th International Conference on Machine Learning (ICML), 2019, pp. 6105-6114.
- L. Yu et al., “Automated melanoma recognition in dermoscopy images via very deep residual networks,” IEEE Transactions on Medical Imaging, vol. 36, no. 4, pp. 994-1004, 2017.
- A. Esteva et al., “Dermatologist-level classification of skin cancer with deep neural networks,” Nature, vol. 542, no. 7639, pp. 115-118, 2017.
- K. He, X. K. He, X. Zhang, S. Ren, and J. Sun, “Deep residual learning for image recognition,” in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2016, pp. 770-778.
- K. Simonyan and A. Zisserman, “Very deep convolutional networks for large-scale image recognition,” in International Conference on Learning Representations (ICLR), 2015.
- A. G. Howard et al. MobileNets: Efficient convolutional neural networks for mobile vision applications. arXiv, 2017. arXiv:1704.04861.
- H. C. Shin et al., “Deep convolutional neural networks for computer-aided detection: CNN architectures, dataset characteristics and transfer learning,” IEEE Transactions on Medical Imaging, vol. 35, no. 5, pp. 1285-1298, 2016.
- J. Hu, L. J. Hu, L. Shen, and G. Sun, “Squeeze-and-excitation networks,” in Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), 2018, pp. 7132-7141.
- P. Tschandl, C. P. Tschandl, C. Rosendahl, and H. Kittler, “The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions,” Scientific Data, vol. 5, no. 1, pp. 1-9, 2018.
- N. Gessert, T. Sentker, F. Madesta, R. Schmitz, H. Kniep, I. Baltruschat, R.Werner, and A. Schlaefer. Skin lesion diagnosis using ensembles, unscaled multi-crop evaluation and loss weighting. arXiv, 2018. arXiv:1808.01694.
- X. He, K. X. He, K. Zhao, and X. Chu, “AutoML: A survey of the state-of-the-art,” Knowledge-Based Systems, vol. 212, p. 106622, 2021.
| Category | Train | Val | Test | Total |
|---|---|---|---|---|
| Enfeksiyonel | 592 | 155 | 750 | 1497 |
| Ekzama | 420 | 98 | 510 | 1028 |
| Akne | 256 | 65 | 322 | 643 |
| Pigment | 110 | 26 | 136 | 272 |
| Benign | 1150 | 280 | 1361 | 2791 |
| Malign | 700 | 159 | 849 | 1708 |
| Class | Precision | Recall | F1-score | Support |
|---|---|---|---|---|
| Enfeksiyonel | 0.93 | 0.90 | 0.92 | 750 |
| Ekzama | 0.89 | 0.89 | 0.89 | 510 |
| Akne | 0.94 | 0.91 | 0.92 | 322 |
| Pigment | 0.85 | 0.73 | 0.79 | 136 |
| Benign | 0.96 | 0.96 | 0.96 | 1361 |
| Malign | 0.93 | 0.89 | 0.91 | 849 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




