Results
Competitive nucleosome reconstitution assay. The relative affinities of the α- and β-globin gene DNA for histones was determined by comparing their ability to reconstitute nucleosomes in vitro, using the 5S rRNA gene from
Xenopus borealis as a reference. Sequences from this gene have been shown to reconstitute nucleosomes with a well-characterized affinity [
23,
24]. We adopted a nucleosome reconstitution procedure utilizing progressive dilution from high salt [
21,
22,
25]. To confirm that this protocol produced DNA-histone complexes with a critical property expected for nucleosomes, we tested the products for protection of an approximately 140 bp fragment from digestion by micrococcal nuclease. Two DNA fragments from the
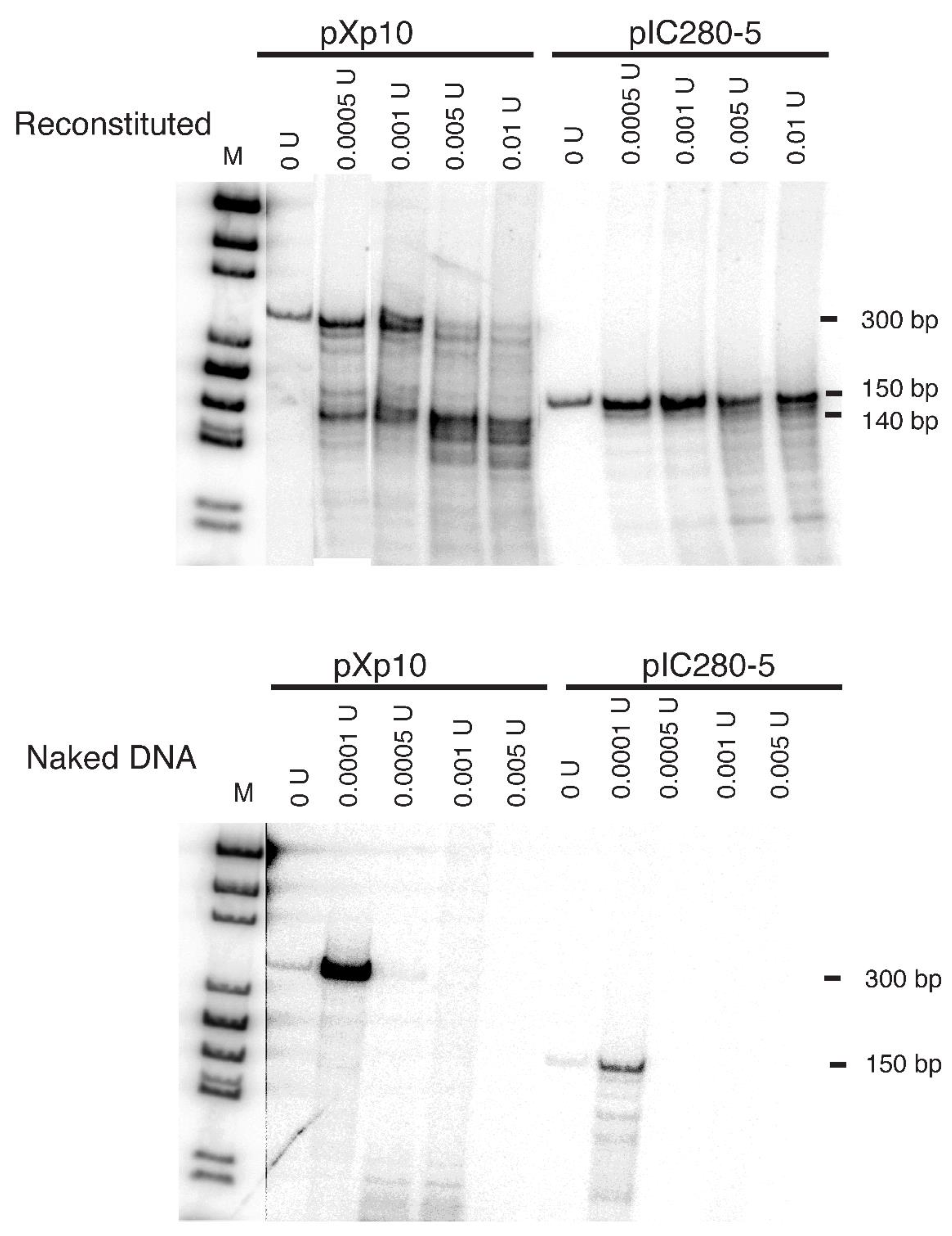
Xenopus borealis 5S rRNA gene, 300bp from plasmid pXp10 and 150bp from plasmid pIC280-5, were used as the templates for nucleosome reconstitution. Both fragments contain a nucleosome-positioning sequence. Digestion of the reconstitution products with increasing amounts of micrococcal nuclease showed protection of the expected size fragments in the range from 140 to 150 bp for both of the 5S rRNA gene templates (
Figure 1, top panel). The protection is observed over a 20-fold range of enzyme activity, and it is dependent on the presence of histones in the reconstitution mix, since no protection is seen in their absence (
Figure 1, bottom panel). Random binding between histones and DNA should have produced a heterogeneous mixture of products with no preferred sizes. The observed preferred size around 140 bp argues that the histone-DNA complexes are in nucleosomes.
In order to compare affinities of different DNAs for histones in nucleosome formation, we employed a technique derived from the competitive nucleosome reconstitution method of Shrader and Crothers [
25]. In this procedure, a small quantity of a labeled test fragment is assayed for its ability to form nucleosomes with purified core histones in the presence of a large amount of competitor DNA. We used a 300 bp fragment encompassing the
Xenopus borealis 5S rRNA gene as a positive control for the ability of this method to yield nucleosomes. As shown in
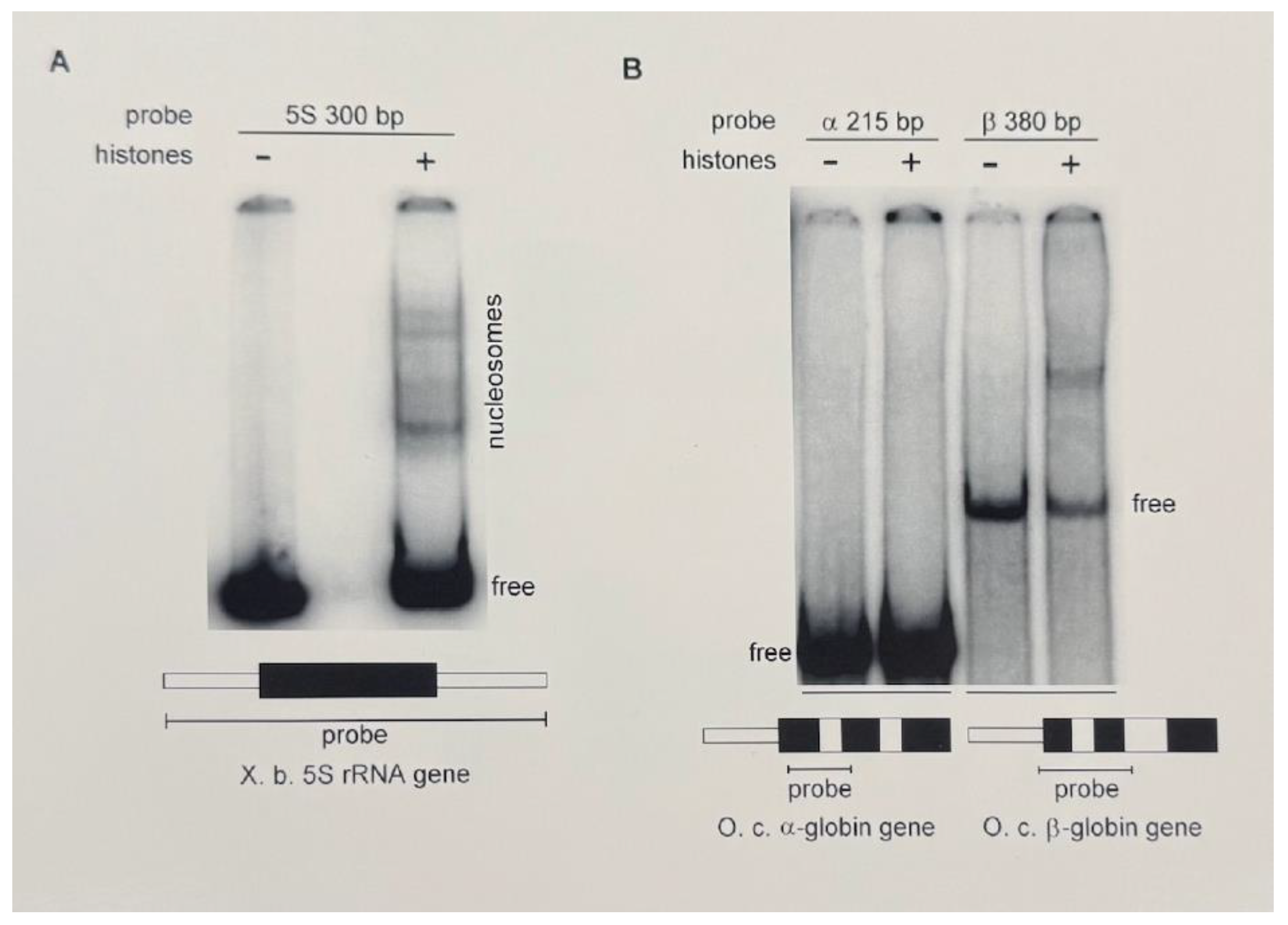
Figure 2A, a small fraction of the total labeled probe was reconstituted with histones to yield six discrete complexes. These complexes may represent both mono- and di-nucleosomes in several preferred phases along the template, each with a unique mobility. The formation of these complexes requires at least a 1:1 mass ratio of competitor calf thymus DNA to histones; at lower amounts of competitor DNA we obtain a non-specific aggregate containing all of the labeled probe (data not shown). The level of reconstitution on this template was used to normalize the relative affinities of other DNA sequences for core histones in subsequent experiments.
Sequences around the CpG island of the rabbit α-globin gene have a lower affinity for core histones than do sequences from the β-globin gene. DNA restriction fragments from the 5’ ends of the rabbit (Oryctolagus cuniculus) α- and β-globin genes were used in this competitive nucleosome reconstitution in order to compare their abilities to form specific complexes with core histones. Using a 2:1 mass ratio of calf thymus DNA to histones, a single discrete complex was formed with the β-globin DNA probe, whereas no complex was evident with the α-globin DNA probe (
Figure 2B). This comparison suggests that the CpG island DNA from the α-globin gene has an inherently lower affinity for core histones in this in vitro competitive reconstitution method.
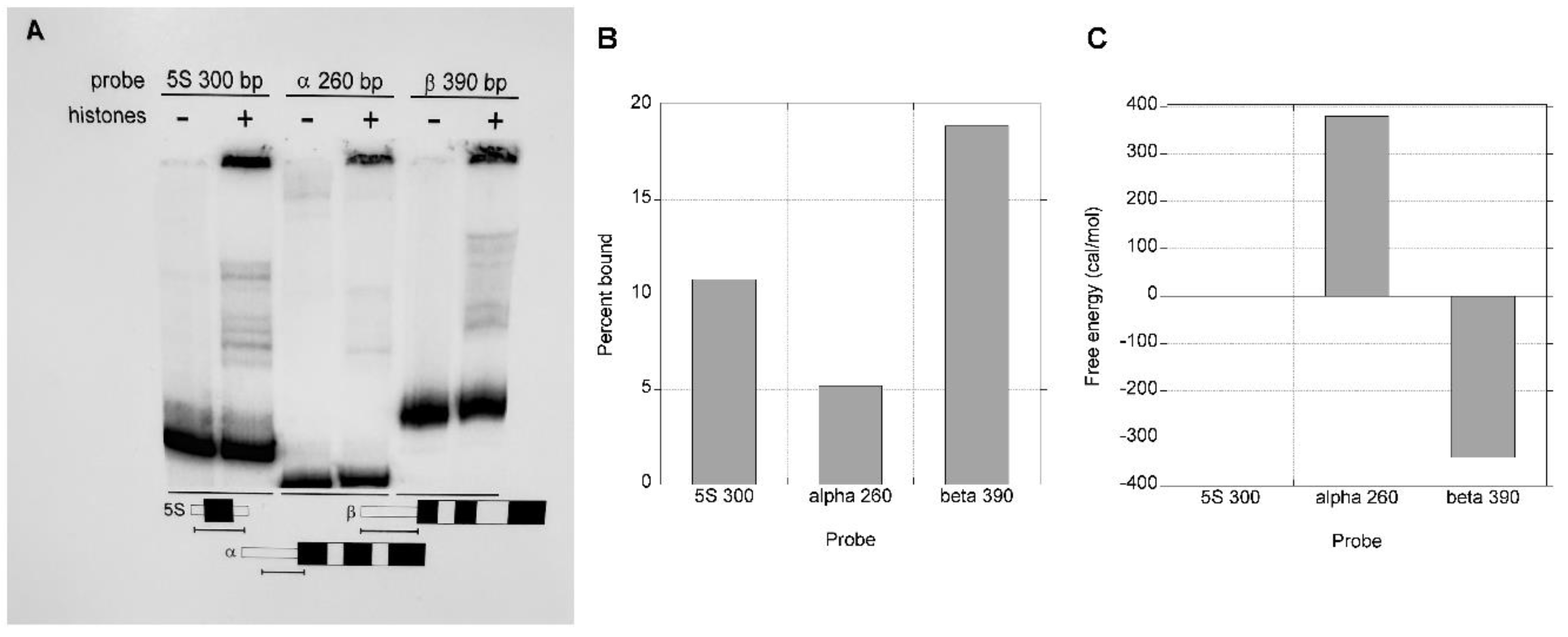
The experiment was repeated with less competitor DNA to allow some nucleosome formation on the α-globin gene DNA template and thereby facilitate a quantitative comparison (
Figure 3A). At a 1:1 mass ratio of calf thymus DNA to histones, the affinity for the β-globin gene fragment was 3-4 times greater than that for the α-globin gene fragment (
Figure 3A and 3B), consistent with the conclusion that the rabbit α-globin gene CpG island has a reduced ability to reconstitute into nucleosomes relative to β-globin gene non-CpG island DNA. The probes in this experiment are from different regions of the genes than those used in
Figure 2B, indicating that these differential affinities for nucleosome reconstitution may reflect general properties of the DNA rather than a particular sequence
per se.
We applied the free energy expression of Shrader and Crothers [
25] to our results in order to ground them with what is known regarding the inherent abilities of various artificial and naturally occurring sequences to reconstitute into nucleosomes. This expression translates the relative proportions of two DNA fragments that form a complex with histones to a difference in the free energy of reconstitution. The levels of reconstitution shown in
Figure 3B translate to a free energy of reconstitution on the α-globin gene template of about 750 cal mol-1 relative to reconstitution on the β-globin gene template (
Figure 3C). Interestingly, the 390 bp β-globin fragment appears to have a higher affinity for core histones than the
X. borealis 5S rRNA gene, with a free energy of about 350 cal mol-1 lower than that of the 5S rRNA gene.
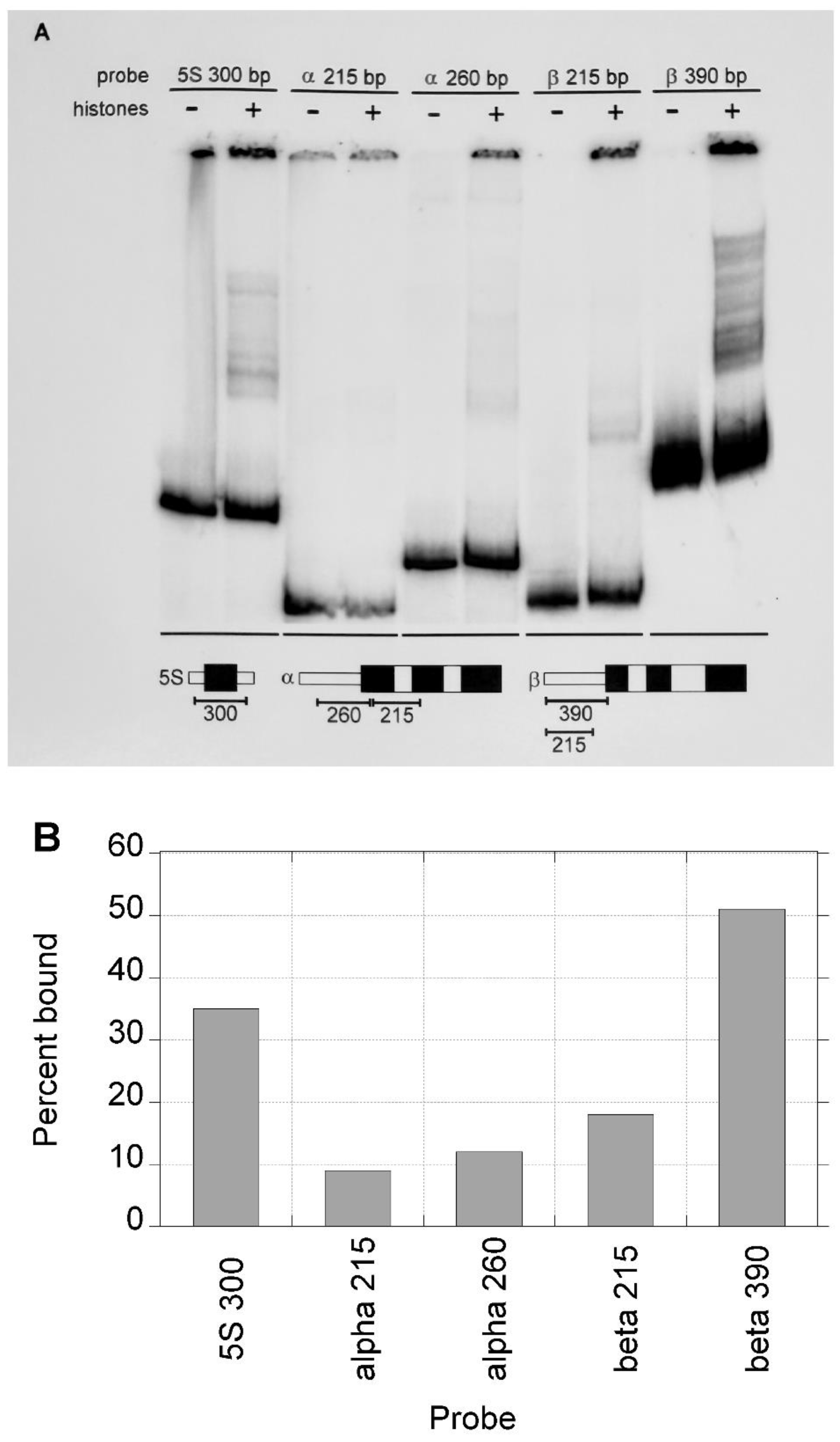
DNA fragments of the same size show a lower affinity for nucleosome reconstitution on CpG island versus non-CpG island DNA. One concern about these initial results is that the higher affinity of the β-globin gene sequence for histones, compared to that of the α-globin gene sequence, could be due to the greater length of the β-globin gene fragment assayed rather than inherent properties of the DNA sequence. To test this possibility, DNA fragments of identical size (215 bp) from the α-globin and β-globin genes were assayed along with the fragments employed in
Figure 3A. Each of the fragments yielded complexes with core histones to varying degrees (
Figure 4A). The 5S 300 bp, α-globin 260 bp and β-globin 390 bp fragments had the same relative levels of reconstitution as seen previously. The affinity for core histones of the α-globin gene 215 bp fragment was 2-fold lower than that of the same sized fragment from the β-globin gene. This is consistent with a lower affinity of CpG island DNA for histones, but the difference is less than that seen previously (
Figure 4A and 4B).
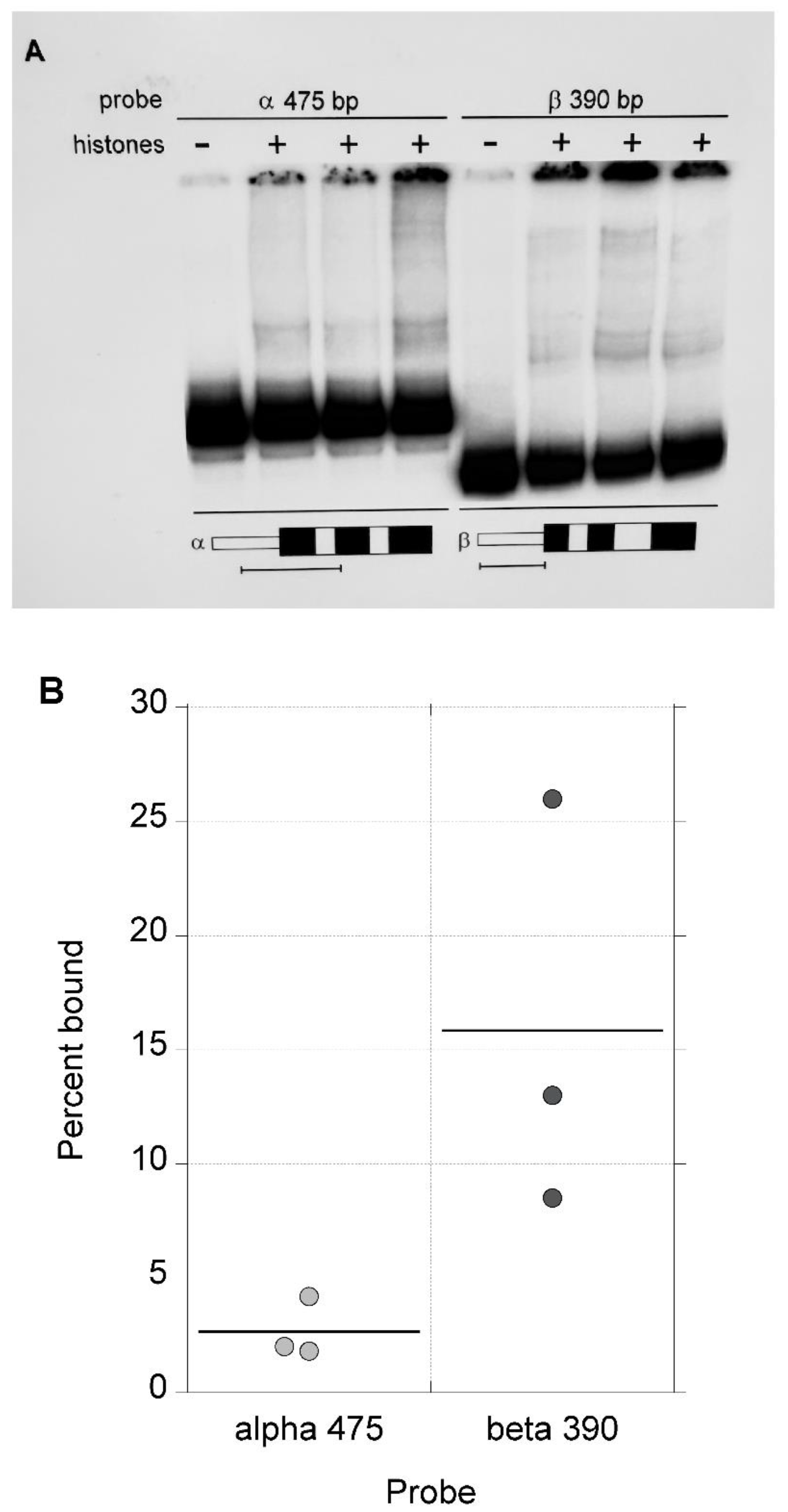
Next, two relatively long DNA fragments were compared in the competitive nucleosome reconstitution assay, a large 475 bp fragment from the α-globin gene containing the sequences from both the 215 bp and 260 bp fragments and the β-globin gene 390 bp fragment used previously (
Figure 5A). The average level of reconstitution with the 475 bp α-globin gene fragment was considerably less than that obtained with the β-globin gene 390 bp sequence (
Figure 5B); this difference is significant by the Kruskal-Wallis Rank Sum Test (p<0.05). The differences in fraction bound translates to a free energy of reconstitution of about 1000 cal mol-1 for the 475 bp α-globin gene fragment relative to the 390 bp β-globin gene fragment. Thus, a relatively large DNA fragment from the α-globin gene CpG island (475 bp) showed a lower level of nucleosome reconstitution than the 390 bp non-CpG island fragment from the β-globin gene.
Longer DNA fragments reconstitute nucleosomes more avidly, but the affinity of CpG island DNA is less than that of non-CpG island DNA. The results presented so far show that both short and long DNA fragments (sufficient to form 1 or 2 nucleosomes, respectively) from the α-globin CpG island form nucleosomes much less readily than do comparably sized fragments from the β-globin gene, which lacks a CpG island. Since the difference in affinities for histones is greater for longer DNA fragments, it appears that the β-globin gene sequence increases its avidity for histones as fragment length increases more so than the α-globin DNA does. There are two possible explanations for this observation. There may be a discrete high-affinity site for nucleosome formation within the 3’ end of the large 390 bp β-globin fragment that is not included in the short 215 bp fragment, whereas there is no such sequence in the analogous region of the α-globin gene. Alternatively, the nature of the β-globin non-CpG island DNA sequence may be such that it contains periodic sequence motifs of favorable topology that increase the ability of DNA to reconstitute into nucleosomes as more sequence is included. In contrast, the α-globin CpG island DNA sequence may not contain such motifs, making it a relatively poor overall template for nucleosome reconstitution that does not improve as more sequence is included.
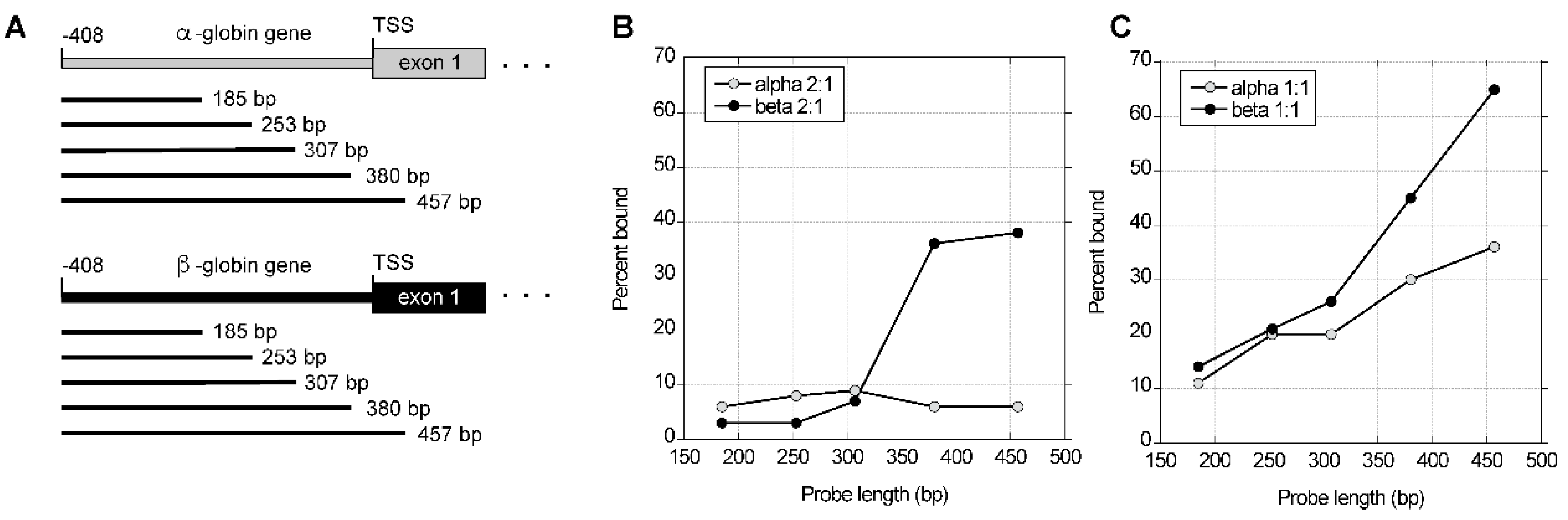
These possibilities were tested by measuring the affinities for core histones of an ordered series of fragments from the α-globin gene CpG island and β-globin gene non-CpG island DNA. Competitive nucleosome reconstitution reactions were performed on a series of PCR fragments ranging in length from 185 bp to 457 bp from the rabbit α- and β-globin genes (
Figure 6A). The endpoints of the PCR fragments were the same relative to the cap site of the respective gene. Using a 2:1 mass ratio of calf thymus DNA to histones, the affinity of β-globin gene non-CpG island sequences for histones increased with longer DNA fragments (
Figure 6B). In contrast, very little of the α-globin gene CpG-island DNA reconstituted into nucleosomes, and that amount did not change as the length of the DNA fragment increased (
Figure 6B). When the amount of competitor DNA was reduced to give a 1:1 mass ratio of calf thymus DNA to histones, the amount of CpG island probe bound to histones did increase with longer DNA fragments, but an even greater increase in the affinities of the non-CpG island probes was observed for longer DNA fragments (
Figure 6C).
These observations are congruent with the results obtained from restriction fragment probes, and they are consistent with the conclusion that the rabbit α-globin gene CpG island sequence is a relatively poor substrate for nucleosome reconstitution. Also, the gradual increase in nucleosome formation as the length of the β-globin gene probe was increased suggests that it is not driven by a discrete high-affinity site, but rather by more dispersed signals.