Submitted:
19 December 2025
Posted:
22 December 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Aim and Review Type
2.2. Conceptual Framework
2.3. Literature Identification
2.4. Eligibility Criteria
2.5. Qualitative Appraisal of Evidence
2.6. Synthesis Strategy
3. Mechanistic Synthesis
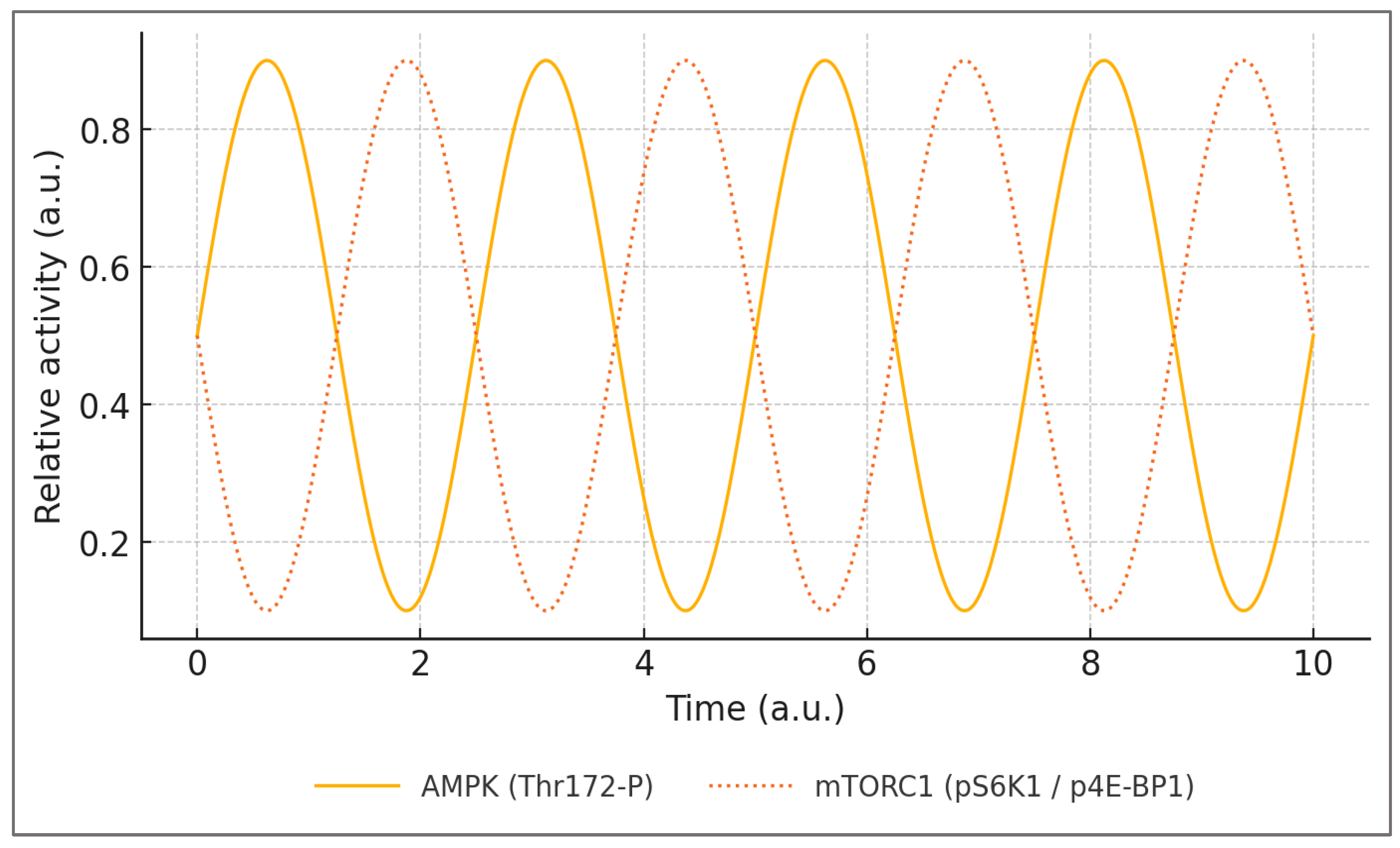
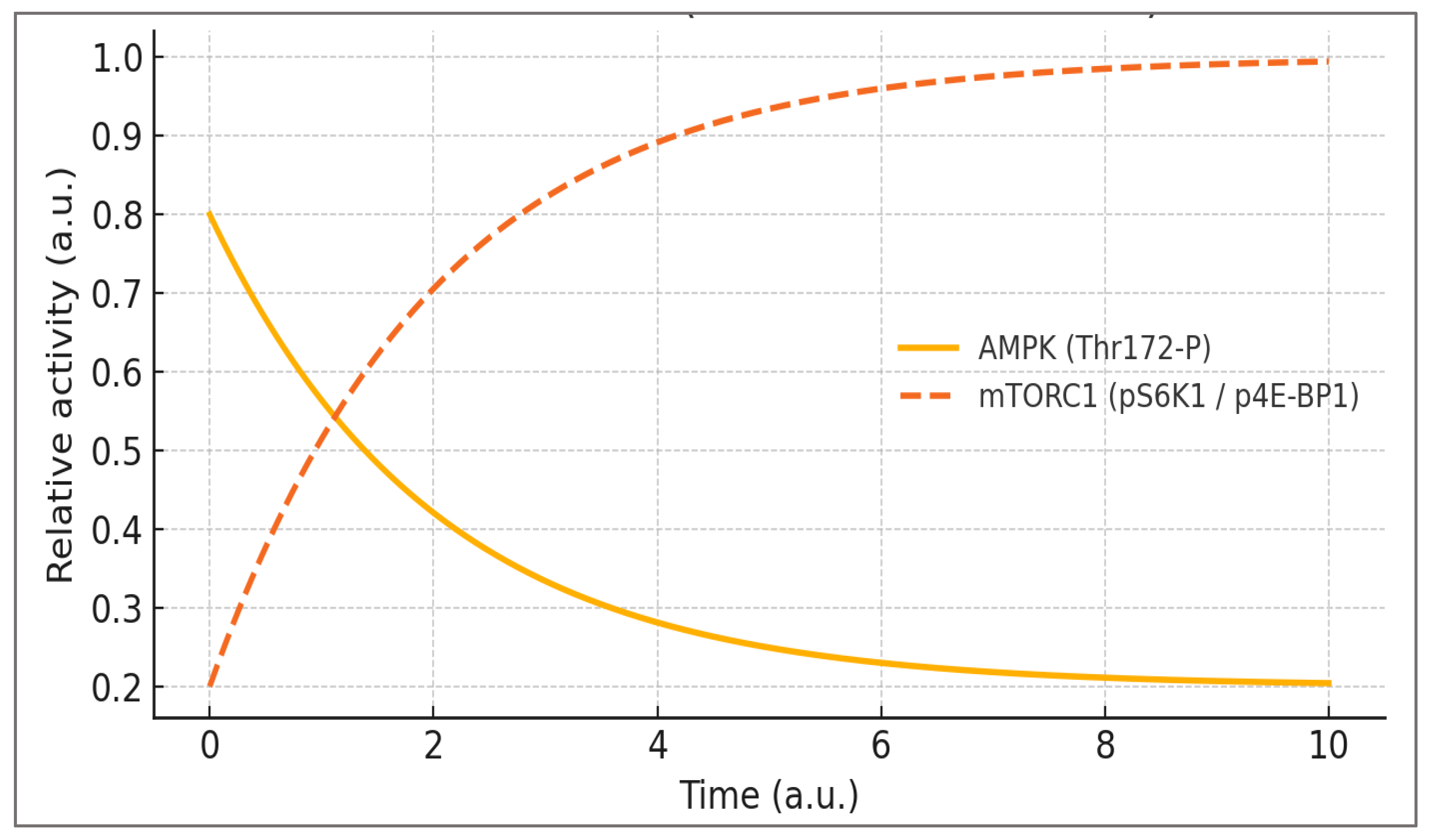
3.1. Nutritional Signaling and the AMPK–mTOR Axis

3.2. Molecular Parallels Between Performance Enhancement and Tumorigenesis
3.3. Epigenetic Modulation Under Metabolic Stress
3.4. Case Contexts: Elite Sport and Doping Paradigms
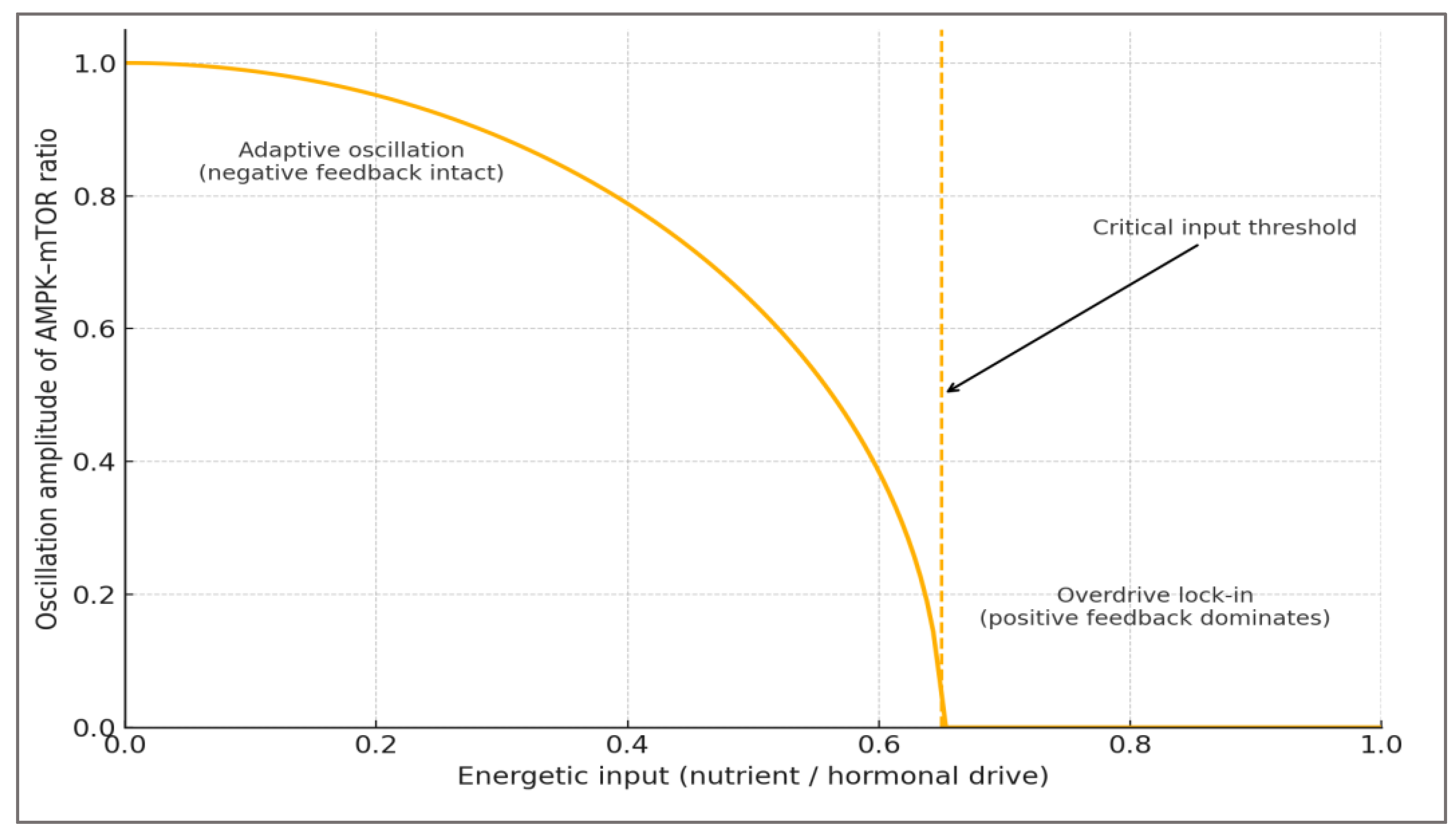
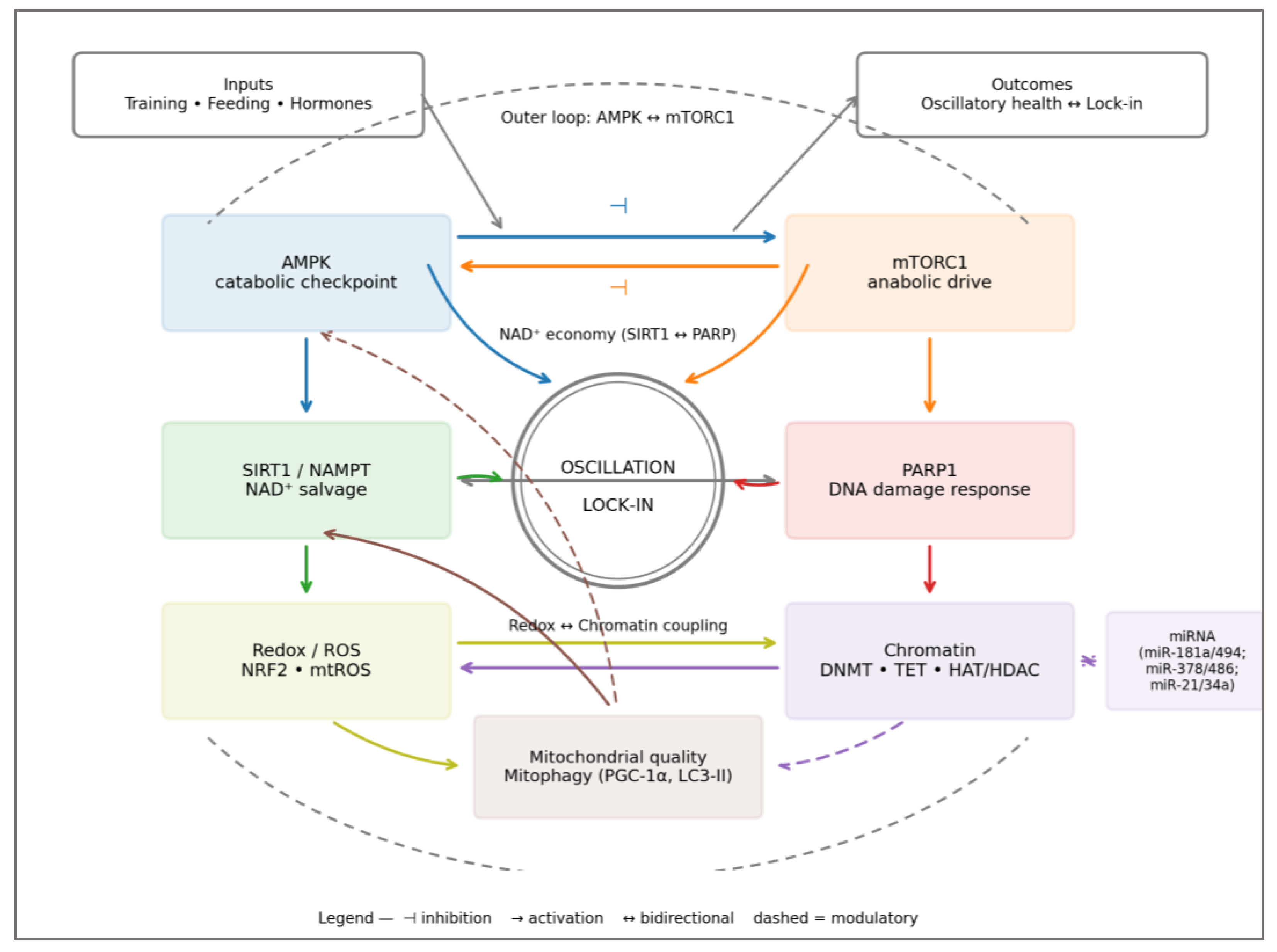
3.5. The Metabolic Overdrive Model
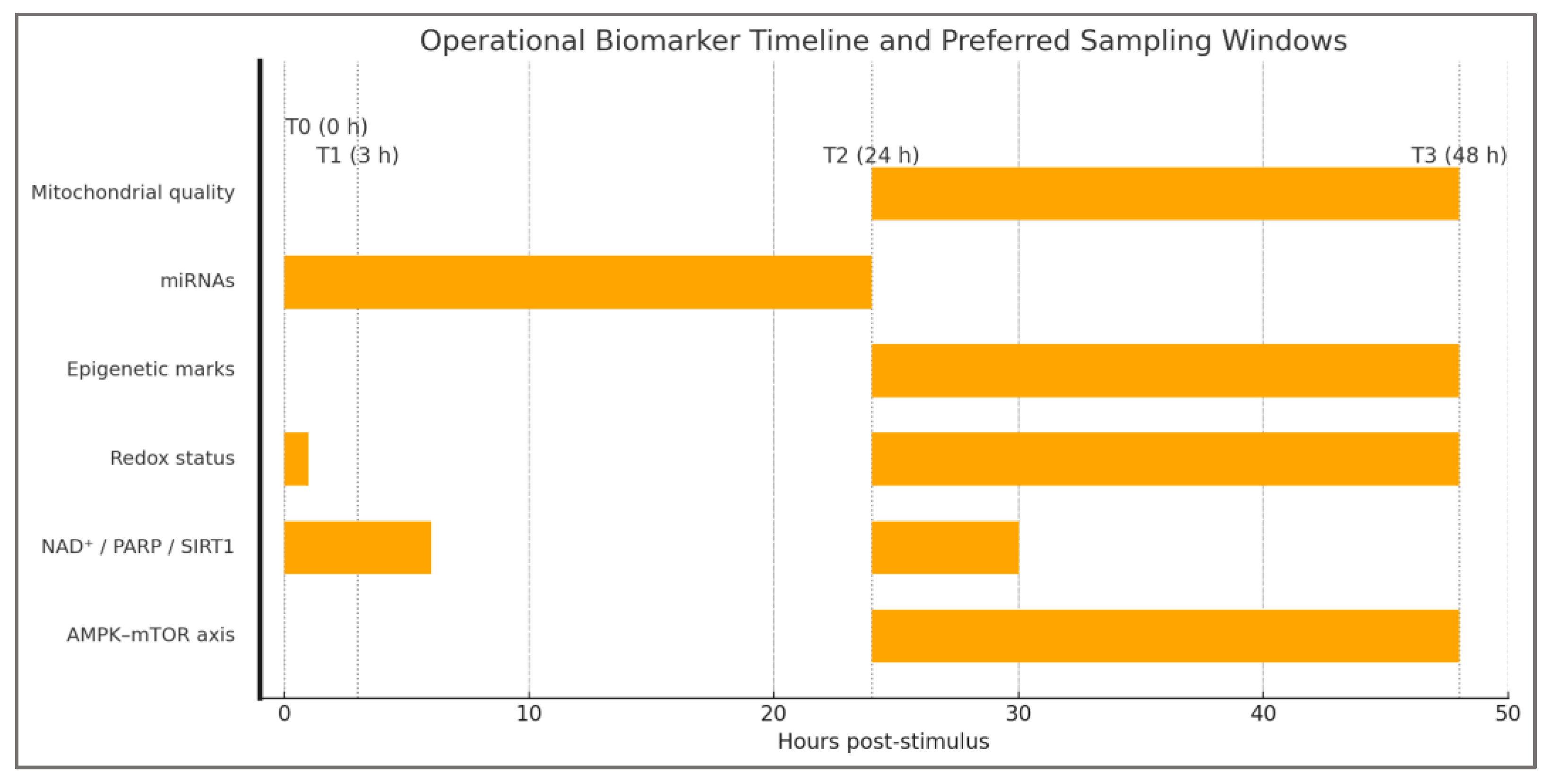
3.6. Testable Predictions and Biomarkers
| Mechanistic axis |
Primary human readouts (assay & specimen) |
Predicted direction in overdrive | Evidence base (cell/animal/human) |
Reversibility signal (periodic AMPK) |
Preferred sampling window |
| AMPK–mTOR balance | AMPK Thr172-P (WB/ELISA; muscle or PBMCs), p-S6K1 (Thr389) / p-4E-BP1 (Ser65) (WB), LC3-II/p62 (autophagic flux with lysosomal inhibition) | ↓ AMPK Thr172-P; ↑ p-S6K1/4E-BP1; ↓ autophagic flux | Strong (cells/rodents) + human acute & training blocks |
↑ AMPK Thr172-P; normalization of p-S6K1; ↑ LC3-II | 24–48 h post-stimulus, spanning feeding |
| NAD+ economy (SIRT1–PARP) | NAD+/NADH (enzymatic cycling; muscle/PBMCs), PARylation (WB), SIRT1 activity (fluor./ELISA) | ↓ NAD+; ↑ PARylation; ↓ SIRT1 activity | Moderate–strong (cells/rodents) + human acute (exercise/irradiation) |
↑ NAD+; ↓ PARylation; ↑ SIRT1 | 0–6 h and 24 h post-stimulus |
| Redox status | 8-oxo-dG (DNA damage), GSH/GSSG, TBARS/MDA, SOD2/Catalase (WB/activity) | ↑ oxidative adducts; ↑ inflammatory redox tone | Strong across models; robust human acute | ↓ adducts; normalization of GSH/GSSG | Immediately post + 24–48 h |
| Epigenetic layer | H3K9ac (ChIP-WB), 5mC/5hmC (LC-MS/MS; targeted bisulfite), TET activity, DNMT1; target loci: PGC-1α, TFAM, PDK4 | ↑ H3K9ac; ↓ TET/5hmC at oxidative loci; ↑ DNMT1; methylation drift | Moderate (cells/rodents) + human acute/longitudinal |
↓ H3K9ac; ↑ 5hmC/TET upon AMPK restoration | 24–72 h (miRNAs earlier) |
| miRNA remodeling | Plasma/serum miR-181a/494 (oxidative), miR-378/486 (hypertrophic), miR-21/34a (fibro-oncogenic) | ↑ miR-21/34a; rigidified profile | Human: acute + training periods |
Shift back toward oxidative profile (miR-181a/494) | 0–24 h |
| Mitochondrial quality | PGC-1α, MFN2, DRP1, mtDNA damage, LC3-II | ↓ mitophagy; ↑ DRP1; ↑ mtROS | Robust experimental; emerging human data | ↑ mitophagy; ↑ PGC-1α; reduced mtROS | 24–72 h |

4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of interest
References
- González, A.; Hall, M.N.; Lin, S.C.; Hardie, D.G. AMPK and TOR: The Yin and Yang of Cellular Nutrient Sensing and Growth Control. Cell Metab. 2020, 31, 472–492. [Google Scholar] [CrossRef] [PubMed]
- Perry, C.G.R.; Hawley, J.A. Molecular Basis of Exercise-Induced Skeletal Muscle Mitochondrial Biogenesis: Historical Advances, Current Knowledge, and Future Challenges. Cold Spring Harb. Perspect. Med. 2018, 8, a029686. [Google Scholar] [CrossRef] [PubMed]
- Saxton, R.A.; Sabatini, D.M. mTOR signaling in growth, metabolism, and disease. Cell 2017, 169, 361–371. [Google Scholar] [CrossRef] [PubMed]
- Mennitti, C.; Farina, G.; Imperatore, A.; De Fonzo, G.; Gentile, A.; La Civita, E.; Carbone, G.; De Simone, R.R.; Di Iorio, M.R.; Tinto, N.; et al. How Does Physical Activity Modulate Hormone Responses? Biomolecules 2024, 14, 1418. [Google Scholar] [CrossRef]
- Esteves, J.V.; Stanford, K.I. Exercise as a tool to mitigate metabolic disease. Am. J. Physiol. Cell Physiol. 2024, 327, C587–C598. [Google Scholar] [CrossRef]
- Camera, D.M.; Smiles, W.J.; Hawley, J.A. Exercise-induced skeletal muscle signaling pathways and the regulation of metabolism. Biochem. J. 2016, 473, 2319–2354. [Google Scholar] [CrossRef]
- Mentch, S.J.; Locasale, J.W. One-carbon metabolism and epigenetics: Understanding the specificity of the methyl donor pathway. Nat. Rev. Mol. Cell Biol. 2016, 17, 561–571. [Google Scholar] [CrossRef]
- Seaborne, R.A.; Strauss, J.; Cocks, M.; Shepherd, S.O.; O’Brien, T.D.; van Someren, K.A.; Bell, P.G.; Murgatroyd, C.; Morton, J.P.; Stewart, C.E. Human Skeletal Muscle Possesses an Epigenetic Memory of Hypertrophy. Sci. Rep. 2018, 8, 1898. [Google Scholar] [CrossRef]
- Barres, R.; Zierath, J.R. The role of diet and exercise in the transgenerational epigenetic landscape of T2DM. Nat. Rev. Endocrinol. 2016, 12, 441–451. [Google Scholar] [CrossRef]
- Mihaylova, M.M.; Shaw, R.J. The AMPK signaling pathway coordinates cell growth, autophagy, and metabolism. Nat. Cell Biol. 2011, 13, 1016–1023. [Google Scholar] [CrossRef]
- Sharples, A.P. Skeletal Muscle Possesses an Epigenetic Memory of Exercise: Role of Nucleus Type-Specific DNA Methylation. Function 2021, 2, zqab047. [Google Scholar] [CrossRef] [PubMed]
- Powers, S.K.; Deminice, R.; Ozdemir, M.; Yoshihara, T.; Bomkamp, M.P.; Hyatt, H. Exercise-induced oxidative stress: Friend or foe? J. Sport Health Sci. 2020, 9, 415–425. [Google Scholar] [CrossRef] [PubMed]
- Clemente-Suárez, V.J.; Martín-Rodríguez, A.; Redondo-Flórez, L.; Ruisoto, P.; Navarro-Jiménez, E.; Ramos-Campo, D.J.; Tornero-Aguilera, J.F. Metabolic Health, Mitochondrial Fitness, Physical Activity, and Cancer. Cancers 2023, 15, 814. [Google Scholar] [CrossRef] [PubMed]
- Khamoui, A.V.; Park, B.-S.; Kim, D.-H.; Yeh, M.-C.; Oh, S.-L.; Elam, M.L.; Jo, E.; Arjmandi, B.H.; Salazar, G.; Grant, S.C.; et al. Aerobic and resistance training dependent skeletal muscle plasticity in the colon-26 murine model of cancer cachexia. Metabolism 2016, 65, 685–698. [Google Scholar] [CrossRef]
- Herzig, S.; Shaw, R.J. AMPK: Guardian of Metabolism and Mitochondrial Homeostasis. Nat. Rev. Mol. Cell Biol. 2018, 19, 121–135. [Google Scholar] [CrossRef]
- Karasawa, T.; Choi, R.H.; Meza, C.A.; Maschek, J.A.; Cox, J.E.; Funai, K. Skeletal muscle PGC-1α remodels mitochondrial phospholipidome but does not alter energy efficiency for ATP synthesis. bioRxiv 2024. [Google Scholar] [CrossRef]
- Goul, C.; Peruzzo, R.; Zoncu, R. The molecular basis of nutrient sensing and signalling by mTORC1 in metabolism regulation and disease. Nat. Rev. Mol. Cell Biol. 2023, 24, 857–875. [Google Scholar] [CrossRef]
- Martinez-Lopez, N.; Mattar, P.; Toledo, M.; Aoun, M.L.; Sharma, M.; McIntire, L.B.; Gunther-Cummins, L.; Macaluso, F.P.; Aguilan, J.T.; Sidoli, S.; et al. mTORC2–NDRG1–CDC42 axis couples fasting to mitochondrial fission. Nat. Cell Biol. 2023, 25, 989–1003. [Google Scholar] [CrossRef]
- Knudsen, J.R.; Blaschke, S.; et al. Exercise increases phosphorylation of the putative mTORC2 activity marker NDRG1 Thr346 in human skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 2022, 322, E465–E475. [Google Scholar] [CrossRef]
- Li, J.; Wang, Z.; Li, C.; Song, Y.; Wang, Y.; Bo, H.; Zhang, Y. Impact of Exercise and Aging on Mitochondrial Homeostasis in Skeletal Muscle: Roles of ROS and Epigenetics. Cells 2022, 11, 2086. [Google Scholar] [CrossRef]
- Jaguri, A.; Nahar, P.; Srinivasan, S.; et al. Exercise Metabolome: Insights for Health and Performance. Metabolites 2023, 13, 694. [Google Scholar] [CrossRef]
- Bojarczuk, A.; Egorova, E.S.; Dzitkowska-Zabielska, M.; et al. Genetics of Exercise and Diet-Induced Fat Loss Efficiency: A Systematic Review. J. Sports Sci. Med. 2024, 23, 236–257. [Google Scholar] [CrossRef] [PubMed]
- Schwalm, C.; Jamart, C.; Benoit, N.; Naslain, D.; Premont, C.; Prevet, J.; Van Thienen, R.; Deldicque, L.; Francaux, M. Activation of Autophagy in Human Skeletal Muscle Is Dependent on Exercise Intensity and AMPK Activation. FASEB J. 2015, 29, 3515–3526. [Google Scholar] [CrossRef] [PubMed]
- Mouchiroud, L.; et al. The NAD+/Sirtuin pathway modulates longevity through mitochondrial UPR. Cell 2013, 154, 430–441. [Google Scholar] [CrossRef] [PubMed]
- Powers, S.K.; Morton, A.B.; Ahn, B.; Smuder, A.J. Redox control of skeletal muscle atrophy. Free Radic. Biol. Med. 2016, 98, 208–217. [Google Scholar] [CrossRef]
- Fanti, M.; Longo, V.D. Nutrition, GH/IGF-1 Signaling, and Cancer. Endocr. Relat. Cancer 2024, 31, e230048. [Google Scholar] [CrossRef]
- Jewell, J.L.; Kim, Y.C.; Russell, R.C.; Yu, F.X.; Park, H.W.; Plouffe, S.W.; Tagliabracci, V.S.; Guan, K.L. Metabolism. Differential regulation of mTORC1 by leucine and glutamine. Science 2015, 347, 194–198. [Google Scholar] [CrossRef]
- Zubrzycki, I.Z.; Ossowski, Z.; Przybylski, S.; Wiacek, M.; Clarke, A.; Trąbka, B. Supplementation with Silk Amino Acids Improves Physiological Parameters Defining Stamina in Elite Fin-Swimmers. J. Int. Soc. Sports Nutr. 2014, 11, 57. [Google Scholar] [CrossRef]
- Yang, L.; Courneya, K.S.; Friedenreich, C.M. The Physical Activity and Cancer Control (PACC) framework: Update on the evidence, guidelines, and future research priorities. Br. J. Cancer 2024, 131, 957–969. [Google Scholar] [CrossRef]
- McDermott, K.T.; Noake, C.; Wolff, R.; Bauld, L.; Espina, C.; Foucaud, J.; Steindorf, K.; Thorat, M.A.; Weijenberg, M.P.; Schüz, J.; et al. Digital interventions to moderate physical inactivity and/or nutrition in young people: A Cancer Prevention Europe overview of systematic reviews. Front. Digit. Health 2023, 5, 1185586. [Google Scholar] [CrossRef]
- Cantó, C.; Menzies, K.J.; Auwerx, J. NAD(+) Metabolism and the Control of Energy Homeostasis: A Balancing Act between Mitochondria and the Nucleus. Cell Metab. 2015, 22, 31–53. [Google Scholar] [CrossRef]
- Longo, V.D.; Panda, S. Fasting, circadian rhythms, and time-restricted feeding in healthy lifespan. Cell Metab. 2016, 23, 1048–1059. [Google Scholar] [CrossRef] [PubMed]
- Hargreaves, M.; Spriet, L.L. Skeletal Muscle Energy Metabolism during Exercise. Nat. Metab. 2020, 2, 817–828. [Google Scholar] [CrossRef] [PubMed]
- Sirago, G.; Maggi, L.; et al. Mammalian Target of Rapamycin (mTOR) Signaling at the Crossroad of Multicellular Muscle Adaptation. Int. J. Mol. Sci. 2022, 23, 13823. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Wei, Y.; Feng, Y.; Zhang, S.; Ma, N.; Wang, K.; Tan, P.; Zhao, Y.; Zhao, J.; Ma, X. Arginine Regulates Skeletal Muscle Fiber Type Formation via mTOR Signaling Pathway. Int. J. Mol. Sci. 2024, 25, 6184. [Google Scholar] [CrossRef]
- Fu, W.; Wu, G. Targeting mTOR for Anti-Aging and Anti-Cancer Therapy. Molecules 2023, 28, 3157. [Google Scholar] [CrossRef]
- Fingar, D.C.; Blenis, J. Target of rapamycin (TOR): An integrator of nutrient and growth factor signals and coordinator of cell growth and cell cycle progression. Oncogene 2004, 23, 3151–3171. [Google Scholar] [CrossRef]
- Mănescu, D.C. Computational Analysis of Neuromuscular Adaptations to Strength and Plyometric Training: An Integrated Modeling Study. Sports 2025, 13, 298. [Google Scholar] [CrossRef]
- Clemmons, D.R. Metabolic actions of insulin-like growth factor-I in normal physiology and diabetes. Endocrinol. Metab. Clin. N. Am. 2012, 41, 425–443. [Google Scholar] [CrossRef]
- Galifi, C.A.; Colalillo, S.; Bertuzzi, A.; Bianchini, M.; Silletti, S. Insulin-Like Growth Factor-1 Receptor Crosstalk with Integrins in Cancer Progression. Endocr. Relat. Cancer 2023, 30, ERC-23-0031. [Google Scholar] [CrossRef]
- Tran, D.; Bergholz, J.; Zhang, H.; He, H.; Wang, Y.; Zhang, Y.; Li, Q.; Kirkland, J.L.; Xiao, Z.; Wang, C.; et al. Insulin-like growth factor-1 regulates the SIRT1–p53 pathway in cellular senescence. Aging Cell 2014, 13, 669–678. [Google Scholar] [CrossRef] [PubMed]
- Guan, Y.; Yan, Z. Molecular Mechanisms of Exercise and Healthspan. Cells 2022, 11, 872. [Google Scholar] [CrossRef] [PubMed]
- Jacko, D.; Schaaf, K.; Masur, L.; Windoffer, H.; Aussieker, T.; Schiffer, T.; Zacher, J.; Bloch, W.; Gehlert, S. Repeated and Interrupted Resistance Exercise Induces the Desensitization and Re-Sensitization of mTOR-Related Signaling in Human Skeletal Muscle Fibers. Int. J. Mol. Sci. 2022, 23, 5431. [Google Scholar] [CrossRef] [PubMed]
- Maciejczyk, M.; Palka, T.; Wiecek, M.; Masel, S.; Szygula, Z. The Effects of Intermittent Hypoxic Training on Anaerobic Performance in Young Men. Appl. Sci. 2024, 14, 676. [Google Scholar] [CrossRef]
- Kang, J.S.; Lee, C.; Kim, S.; et al. Baf155 Regulates Skeletal Muscle Metabolism via HIF-1α Signalling. PLoS Biol. 2023, 21, e3002192. [Google Scholar] [CrossRef]
- Yin, W.; Noguchi, C.T. The Role of Erythropoietin in Metabolic Regulation. Cells 2025, 14, 280. [Google Scholar] [CrossRef]
- Man, M.C.; Ganera, C.; Bărbuleţ, G.D.; Krzysztofik, M.; Panaet, A.E.; Cucui, A.I.; Tohănean, D.I.; Alexe, D.I. The Modifications of Haemoglobin, Erythropoietin Values and Running Performance While Training at Mountain vs. Hilltop vs. Seaside. Int. J. Environ. Res. Public Health 2021, 18, 9486. [Google Scholar] [CrossRef]
- Zhou, Z.; Chen, C.; Teo, E.-C.; Zhang, Y.; Huang, J.; Xu, Y.; Gu, Y. Intracellular Oxidative Stress Induced by Physical Exercise in Adults: Systematic Review and Meta-Analysis. Antioxidants 2022, 11, 1751. [Google Scholar] [CrossRef]
- Meng, Q.; Su, C.-H. The Impact of Physical Exercise on Oxidative and Nitrosative Stress: Balancing the Benefits and Risks. Antioxidants 2024, 13, 573. [Google Scholar] [CrossRef]
- Sabharwal, S.S.; Schumacker, P.T. Mitochondrial ROS in cancer: Initiators, amplifiers or an Achilles’ heel? Nat. Rev. Cancer 2014, 14, 709–721. [Google Scholar] [CrossRef]
- Sullivan, L.B.; Chandel, N.S. Mitochondrial reactive oxygen species and cancer. Cancer Metab. 2014, 2, 17. [Google Scholar] [CrossRef] [PubMed]
- Barba, I.; Carrillo-Bosch, L.; Seoane, J. Targeting the Warburg Effect in Cancer: Where Do We Stand? Int. J. Mol. Sci. 2024, 25, 3142. [Google Scholar] [CrossRef] [PubMed]
- Maddocks, O.D.K.; Labuschagne, C.F.; Adams, P.D.; Vousden, K.H. Serine metabolism supports the methionine cycle and DNA/RNA methylation through de novo ATP synthesis in cancer cells. Mol. Cell 2016, 61, 210–221. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Chauhan, A.; Dai, Y. Connections between metabolism and epigenetics: From basic mechanisms to clinical implications. Front. Pharmacol. 2022, 13, 935536. [Google Scholar] [CrossRef]
- Pouikli, A.; Tessarz, P. Metabolism and chromatin: A dynamic duo that regulates development and ageing. BioEssays 2021, 43, e2000273. [Google Scholar] [CrossRef]
- Xie, N.; Zhang, L.; Wang, J.; Zhao, Y.; Dong, Y. NAD+ metabolism: Pathophysiologic mechanisms and therapeutic potential. Signal Transduction and Targeted Therapy 2020, 5, 227. [Google Scholar] [CrossRef]
- Mentch, S.J.; Locasale, J.W. One-carbon metabolism and epigenetics: Understanding the specificity of the methyl donor pathway. Nat. Rev. Mol. Cell Biol. 2016, 17, 561–571. [Google Scholar] [CrossRef]
- Janke, R.; Dodson, A.E.; Rine, J. Metabolism and epigenetics. Annu. Rev. Cell Dev. Biol. 2015, 31, 473–496. [Google Scholar] [CrossRef]
- Barrès, R.; Yan, J.; Egan, B.; Treebak, J.T.; Rasmussen, M.; Fritz, T.; Caidahl, K.; Krook, A.; O’Gorman, D.J.; Zierath, J.R. Acute exercise remodels promoter methylation in human skeletal muscle. Cell Metab. 2012, 15, 405–411. [Google Scholar] [CrossRef]
- Steinberg, G.R.; Kemp, B.E. AMPK in health and disease. Physiol. Rev. 2019, 89, 1025–1078. [Google Scholar] [CrossRef]
- Seaborne, R.A.; Strauss, J.; Cocks, M.; Shepherd, S.O.; O’Brien, T.D.; van Someren, K.A.; Bell, P.G.; Murgatroyd, C.; Morton, J.P.; Stewart, C.E. Human skeletal muscle possesses an epigenetic memory of hypertrophy. Sci. Rep. 2018, 8, 1898. [Google Scholar] [CrossRef] [PubMed]
- Barrès, R.; Zierath, J.R. The role of diet and exercise in the transgenerational epigenetic landscape of T2DM. Nat. Rev. Endocrinol. 2016, 12, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; Costello, J. DNA methylation: An epigenetic mark of cellular memory. Exp. Mol. Med. 2017, 49, e322. [Google Scholar] [CrossRef] [PubMed]
- Baylin, S.B.; Jones, P.A. A decade of exploring the cancer epigenome—Biological and translational implications. Nat. Rev. Cancer 2011, 11, 726–734. [Google Scholar] [CrossRef]
- Shvedunova, M.; Akhtar, A. Modulation of cellular processes by histone and non-histone lysine acetylation: Mechanisms, targets, and physiological significance. Nat. Rev. Mol. Cell Biol. 2022, 23, 304–326. [Google Scholar] [CrossRef]
- Cantó, C.; Sauve, A.A.; Bai, P. Crosstalk between poly(ADP-ribose) polymerase and sirtuin enzymes. Mol. Aspects Med. 2013, 34, 1168–1201. [Google Scholar] [CrossRef]
- Afzal, M.; Greco, F.; Quinzi, F.; Scionti, F.; Maurotti, S.; Montalcini, T.; Mancini, A.; Buono, P.; Emerenziani, G.P. The Effect of Physical Activity/Exercise on miRNA Expression and Function in Non-Communicable Diseases—A Systematic Review. Int. J. Mol. Sci. 2024, 25, 6813. [Google Scholar] [CrossRef]
- Humińska-Lisowska, K. Dopamine in Sports: A Narrative Review on the Genetic and Epigenetic Factors Shaping Personality and Athletic Performance. Int. J. Mol. Sci. 2024, 25, 11602. [Google Scholar] [CrossRef]
- Grieb, A.; Schmitt, A.; Fragasso, A.; Widmann, M.; Mattioni Maturana, F.; Burgstahler, C.; Erz, G.; Schellhorn, P.; Nieß, A.M.; Munz, B. Skeletal Muscle MicroRNA Patterns in Response to a Single Bout of Exercise in Females: Biomarkers for Subsequent Training Adaptation? Biomolecules 2023, 13, 884. [Google Scholar] [CrossRef]
- Quintanilha, B.J.; Reis, B.Z.; Duarte, G.B.S.; Cozzolino, S.M.F.; Rogero, M.M. Nutrimiromics: Role of microRNAs and nutrition in modulating inflammation and chronic diseases. Nutrients 2017, 9, 1168. [Google Scholar] [CrossRef]
- Radom-Aizik, S.; et al. Effects of exercise training on microRNA expression in young and older men. Cell Physiol. Biochem. 2022, 29, 831–842. [Google Scholar] [CrossRef]
- Fu, J.; Zhang, X.; Li, H.; Liu, Y.; Wang, X. MicroRNA-34 Family in Cancers: Role, Mechanism, and Therapeutic Potential. Cancers 2023, 15, 4723. [Google Scholar] [CrossRef] [PubMed]
- Xiao, M.; Yang, H.; Xu, W.; Ma, S.; Lin, H.; Zhu, H.; Liu, L.; Liu, Y.; Yang, C.; Xu, Y.; et al. Inhibition of α-KG–dependent histone and DNA demethylases by fumarate and succinate that are accumulated in mutations of FH and SDH tumor suppressors. Genes Dev. 2012, 26, 1326–1338. [Google Scholar] [CrossRef] [PubMed]
- Walzik, D.; Jonas, W.; Joisten, N.; Belen, S.; Wüst, R.C.I.; Guillemin, G.; Zimmer, P. Tissue-Specific Effects of Exercise as NAD+-Boosting Strategy: Current Knowledge and Future Perspectives. Acta Physiol. 2023, 237, e13921. [Google Scholar] [CrossRef]
- Horvath, S.; Raj, K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat. Rev. Genet. 2018, 19, 371–384. [Google Scholar] [CrossRef]
- Mănescu, D.C.; Mănescu, A.M. Artificial Intelligence in the Selection of Top-Performing Athletes for Team Sports: A Proof-of-Concept Predictive Modeling Study. Appl. Sci. 2025, 15, 9918. [Google Scholar] [CrossRef]
- Mănescu, D.C. Big Data Analytics Framework for Decision-Making in Sports Performance Optimization. Data 2025, 10, 116. [Google Scholar] [CrossRef]
- Humińska-Lisowska, K.; Chmielowiec, K.; Strońska-Pluta, A.; Chmielowiec, J.; Suchanecka, A.; Masiak, J.; Michałowska-Sawczyn, M.; Boroń, A.; Cięszczyk, P.; Grzywacz, A. Epigenetic Analysis of the Dopamine Transporter Gene DAT1 with a Focus on Personality Traits in Athletes. Int. J. Mol. Sci. 2023, 24, 8931. [Google Scholar] [CrossRef]
- Kostrzewa-Nowak, D.; Nowak, R. Differential Th Cell-Related Immune Responses in Young Physically Active Men after an Endurance Effort. J. Clin. Med. 2020, 9, 1795. [Google Scholar] [CrossRef]
- Fernández-Lázaro, D.; Mielgo-Ayuso, J.; Caballero-García, A.; Calleja-González, J.; Herrador-Pérez, M.; Armegod-Benítez, M.; Crespo-Otín, S.; Carratalá-Tejada, R. Adequacy of an Altitude Fitness Program (Living and Training at Moderate Altitude) Supplemented with Intermittent Hypoxic Training: Effects on Sports Performance, Blood Biomarkers and Safety Profiles in Elite Athletes. Int. J. Environ. Res. Public Health 2022, 19, 9095. [Google Scholar] [CrossRef]
- Maciejczyk, M.; Palka, T.; Wiecek, M.; Szymura, J.; Kusmierczyk, J.; Bawelski, M.; Masel, S.; Szygula, Z. Effects of Intermittent Hypoxic Training on Aerobic Capacity and Second Ventilatory Threshold in Untrained Men. Appl. Sci. 2023, 13, 9954. [Google Scholar] [CrossRef]
- Goliniewski, J.; Czuba, M.; Płoszczyca, K.; Chalimoniuk, M.; Gajda, R.; Niemaszyk, A.; Kaczmarczyk, K.; Langfort, J. The Impact of Normobaric Hypoxia and Intermittent Hypoxic Training on Cardiac Biomarkers in Endurance Athletes: A Pilot Study. Int. J. Mol. Sci. 2024, 25, 4584. [Google Scholar] [CrossRef] [PubMed]
- Andersen, A.B.; Bejder, J.; Bonne, T.C.; Nordsborg, N.B. Contemporary Blood Doping—Performance, Mechanism, and Detection. Scand. J. Med. Sci. Sports 2024, 34, e14243. [Google Scholar] [CrossRef] [PubMed]
- Gordeuk, V.R.; Key, N.S.; Prchal, J.T. Re-Evaluation of Hematocrit as a Determinant of Thrombotic Risk in Erythrocytosis. Haematologica 2019, 104, 653–658. [Google Scholar] [CrossRef]
- Chen, Z.; Han, F.; Du, Y.; Shi, H.; Zhou, W. Hypoxic Microenvironment in Cancer: Molecular Mechanisms and Therapeutic Interventions. Signal Transduct. Target. Ther. 2023, 8, 70. [Google Scholar] [CrossRef]
- Yasuoka, Y.; Izumi, Y.; Sands, J.M.; Kawahara, K.; Nonoguchi, H. Progress in the Detection of Erythropoietin in Blood, Urine, and Tissue. Molecules 2023, 28, 4446. [Google Scholar] [CrossRef]
- Nagel, M.J.; Jarrard, C.P.; Lalande, S. Effect of a Single Session of Intermittent Hypoxia on Erythropoietin and Oxygen-Carrying Capacity. Int. J. Environ. Res. Public Health 2020, 17, 7257. [Google Scholar] [CrossRef]
- Mănescu, D.C. Alimentaţia în fitness şi bodybuilding; Editura ASE: Bucharest, Romania, 2010. [Google Scholar]
- Hurst, P.; Kavussanu, M.; Davies, R.; Dallaway, N.; Ring, C. Use of Sport Supplements and Doping Substances by Athletes: Prevalence and Relationships. J. Clin. Med. 2024, 13, 7132. [Google Scholar] [CrossRef]
- Park, J. The mechanisms of anabolic steroids, selective androgen receptor modulators and resistance exercise in skeletal muscle hypertrophy. Korean J. Sports Med. 2022, 40, 67–77. [Google Scholar] [CrossRef]
- Waskiw-Ford, M.; Hodson, N.; Fung, H.J.W.; West, D.W.D.; Apong, P.; Bashir, R.; Moore, D.R. Essential Amino Acid Ingestion Facilitates Leucine Retention and Attenuates Myofibrillar Protein Breakdown following Bodyweight Resistance Exercise in Young Adults in a Home-Based Setting. Nutrients 2022, 14, 3532. [Google Scholar] [CrossRef]
- Sivalokanathan, S.; Małek, Ł.A.; Malhotra, A. The Cardiac Effects of Performance-Enhancing Medications: Caffeine vs. Anabolic Androgenic Steroids. Diagnostics 2021, 11, 324. [Google Scholar] [CrossRef]
- Nielsen, J.L.; Rasmussen, J.J.; Frandsen, M.N.; Fredberg, J.; Brandt-Jacobsen, N.H.; Aagaard, P.; Kistorp, C. Higher Myonuclei Density in Muscle Fibers Persists Among Former Users of Anabolic Androgenic Steroids. J. Clin. Endocrinol. Metab. 2024, 109. [Google Scholar] [CrossRef] [PubMed]
- Herlitz, L.C.; Markowitz, G.S.; Farris, A.B. Development of Focal Segmental Glomerulosclerosis after Anabolic Steroid Abuse. J. Am. Soc. Nephrol. 2010, 21, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Brennan, B.P.; Kanayama, G.; Hudson, J.I.; Pope, H.G., Jr. Human Growth Hormone Abuse in Male Weightlifters. Am. J. Addict. 2011, 20, 9–13. [Google Scholar] [CrossRef] [PubMed]
- Hoshino, D.; Yoshida, Y.; Holloway, G.P.; Lally, J.; Hatta, H.; Bonen, A. Clenbuterol, a β2-Adrenergic Agonist, Reciprocally Alters PGC-1α and RIP140 and Reduces Fatty Acid and Pyruvate Oxidation in Rat Skeletal Muscle. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2012, 302, R373–R384. [Google Scholar] [CrossRef]
- Wiacek, M.; Trąbka, B.; Tomasiuk, R.; Zubrzycki, I.Z. Changes in Health-Related Parameters Associated with Sports Performance Enhancement Drugs. Int. J. Sports Med. 2023, 44, 206–214. [Google Scholar] [CrossRef]
- Nowak, R.; Turkiewicz, K.; Sławiński, M.; Clark, J.S.C.; Szylińska, A.; Proia, P.; Jodko, Ł.; Wojciuk, B.; Sulżyc-Bielicka, V.; Kostrzewa-Nowak, D. The Impact of Endurance Exercise on Routine Laboratory Parameters in Young Trained Individuals. J. Clin. Med. 2025, 14, 5703. [Google Scholar] [CrossRef]
- Mănescu, A.M.; Grigoroiu, C.; Smîdu, N.; Dinciu, C.C.; Mărgărit, I.R.; Iacobini, A.; Mănescu, D.C. Biomechanical Effects of Lower Limb Asymmetry During Running: An OpenSim Computational Study. Symmetry 2025, 17, 1348. [Google Scholar] [CrossRef]
- Furrer, R.; Handschin, C. Molecular Aspects of the Exercise Response and Training Adaptation in Skeletal Muscle. Free Radic. Biol. Med. 2024, 223, 53–68. [Google Scholar] [CrossRef]
- Zubrzycki, I.Z.; Wiacek, M.; Trąbka, B.; Ossowski, Z. Anabolic Steroids in a Contest Preparation of the Top World-Class Bodybuilder. Int. J. Med. Pharm. Case Rep. 2015, 5, 1–8. [Google Scholar] [CrossRef]
- Plaza-Díaz, J.; Izquierdo, D.; Torres-Martos, Á.; Baig, A.T.; Aguilera, C.M.; Ruiz-Ojeda, F.J. Impact of Physical Activity and Exercise on the Epigenome in Skeletal Muscle and Effects on Systemic Metabolism. Biomedicines 2022, 10, 126. [Google Scholar] [CrossRef]
- Clemente-Suárez, V.J.; Bustamante-Sánchez, Á.; Mielgo-Ayuso, J.; Martínez-Guardado, I.; Martín-Rodríguez, A.; Tornero-Aguilera, J.F. Antioxidants and Sports Performance. Nutrients 2023, 15, 2371. [Google Scholar] [CrossRef]
- Lee, J.; Kim, D.; Kim, C. Resistance Training for Glycemic Control, Muscular Strength, and Lean Body Mass in Old Type 2 Diabetic Patients: A Meta-Analysis. Diabetes Ther. 2017, 8, 459–473. [Google Scholar] [CrossRef] [PubMed]
- Bıçakçı, B.; Cięszczyk, P.; Humińska-Lisowska, K. Genetic Determinants of Endurance: A Narrative Review on Elite Athlete Status and Performance. Int. J. Mol. Sci. 2024, 25, 13041. [Google Scholar] [CrossRef] [PubMed]
- Toniolo, L.; Concato, M.; Giacomello, E. Resveratrol, a Multitasking Molecule That Improves Skeletal Muscle Health. Nutrients 2023, 15, 3413. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Zhang, H.; Zhou, H.; Liu, Q.; Xu, W.; Chen, X. Impact of Exercise and Aging on Mitochondrial Function via Epigenetic Modifications. Cells 2022, 11, 2086. [Google Scholar] [CrossRef]
- Williams, A.; Wadsworth, D.D.; Geetha, T. Exercise, Epigenetics, and Body Composition: Molecular Connections. Cells 2025, 14, 1553. [Google Scholar] [CrossRef]
- Zeng, Z.; Centner, C.; Gollhofer, A.; König, D. Effects of Dietary Strategies on Exercise-Induced Oxidative Stress: A Narrative Review of Human Studies. Antioxidants 2021, 10, 542. [Google Scholar] [CrossRef]
- Soto, D.A.S.; Pérez, D.I.V.; Souza, M.R.d.; Reis, M.L.G.; Almeida, N.R.; Miarka, B.; Aedo-Muñoz, E.; Oyarzo, A.I.A.; Sillero-Quintana, M.; Queiroz, A.C.C.; et al. Thermogenic Activation of Adipose Tissue by Caffeine During Strenuous Exercising and Recovery: A Double-Blind Crossover Study. Metabolites 2025, 15, 517. [Google Scholar] [CrossRef]
- Kolodziej, F.; O’Halloran, K.D. Re-Evaluating the Oxidative Phenotype: Can Endurance Exercise Save the Western World? Antioxidants 2021, 10, 609. [Google Scholar] [CrossRef]
- Moreno-Villanueva, M.; Kramer, A.; Hammes, T.; Venegas-Carro, M.; Thumm, P.; Bürkle, A.; Gruber, M. Influence of Acute Exercise on DNA Repair and PARP Activity before and after Irradiation in Lymphocytes from Trained and Untrained Individuals. Int J Mol Sci. 2019, 20, 2999. [Google Scholar] [CrossRef]
- Yasemi, M.; Jolicoeur, M. Modelling Cell Metabolism: A Review on Constraint-Based Steady-State and Kinetic Approaches. Processes 2021, 9, 322. [Google Scholar] [CrossRef]
- Mănescu, A.M.; Hangu, S.Ș.; Mănescu, D.C. Nutritional Supplements for Muscle Hypertrophy: Mechanisms and Morphology—Focused Evidence. Nutrients 2025, 17, 3603. [Google Scholar] [CrossRef]
- Imperatore, A.; Mennitti, C.; De Fonzo, G.; Amitrano, R.; Gentile, A.; Calvanese, M.; Iafusco, F.; Coppola, S.; Digno, M.; Borrelli, P.; et al. Toward a Multi-Trait Genetic Panel Targeting Training, Rehabilitation, and Chronic Disease Prevention: A Narrative Review. Genes 2025, 16, 1309. [Google Scholar] [CrossRef]
- Volkova, S.; Matos, M.R.A.; Mattanovich, M.; Marín de Mas, I. Metabolic Modelling as a Framework for Metabolomics Data Integration and Analysis. Metabolites 2020, 10, 303. [Google Scholar] [CrossRef]
| Pathway / Component | Nutritional or Energetic Trigger | Upstream Regulator(s) | Primary Molecular Effect | Downstream Target(s) | Adaptive Role (Physiological Context) | Potential Dysregulation (Overdrive State) |
| AMPK | Energy deficit, exercise, fasting | AMP/ATP ratio, LKB1 | Activates catabolic fluxes; inhibits mTORC1 | ACC, ULK1, PGC-1α | Enhances mitochondrial biogenesis, endurance, autophagy | Chronic suppression → anabolic dominance, insulin resistance |
| mTORC1 | Amino acids (leucine), insulin, IGF-1 | PI3K–AKT, Rheb, Rag GTPases | Stimulates protein synthesis via p70S6K and 4E-BP1 | Ribosomal proteins, eIF4E | Promotes muscle growth and recovery | Persistent activation → oxidative stress, epigenetic drift |
| SIRT1 | NAD+ levels, caloric restriction | AMPK, NAMPT | Deacetylates transcriptional regulators (PGC-1α, FOXO) | PGC-1α, p53, NF-κB | Increases oxidative metabolism and stress resistance | NAD+ depletion → loss of redox control and mitochondrial decline |
| IGF-1/AKT axis | Protein intake, GH, insulin | GH/IGF-1 signaling | Activates mTORC1, inhibits AMPK | FOXO, TSC2, GSK3β | Supports hypertrophy and tissue regeneration | Chronic activation → reduced autophagy, oncogenic signaling |
| Leucine–AMPK–mTOR crosstalk | Branched-chain amino acids | Ragulator complex | Coordinates anabolism–catabolism balance | mTORC1, AMPK | Fine-tunes training–fuel coupling, metabolic flexibility | Excess BCAA intake → insulin desensitization, ROS accumulation |
| NAD+/PARP balance | Oxidative load, DNA repair demand | PARP1, SIRT1 | Competes for NAD+ substrate | SIRT1, PGC-1α | Links redox balance to mitochondrial maintenance | PARP hyperactivation → NAD+ depletion, energy collapse |
| FOXO transcription factors | Energy deficit, oxidative stress | AMPK, SIRT1, AKT | Regulate autophagy, antioxidant enzymes | Catalase, MnSOD, LC3 | Antioxidant defense and longevity | AKT-mediated inhibition → reduced oxidative protection |
| Mechanistic domain | Representative trigger or intervention |
Physiological driver (Elite sport) |
Oncogenic analogue (Cancer biology) |
Key molecular mediator(s) | Adaptive outcome | Pathological counterpart (Overdrive state) |
| Anabolic signaling (mTOR/AKT) | Resistance training, growth hormone, anabolic steroids | Transient anabolic activation for muscle hypertrophy and recovery | PI3K–AKT–mTOR hyperactivation promoting uncontrolled cell proliferation | mTORC1, AKT, S6K1, 4E-BP1, TSC2 | Enhanced protein synthesis and tissue remodeling | Persistent anabolic signaling, autophagy suppression, tumor-like growth |
| Hypoxia and angiogenesis | Altitude exposure, EPO doping, hypoxic training | HIF-1α activation and VEGF upregulation for capillary and mitochondrial biogenesis | Tumor hypoxia, neovascularization | HIF-1α, VEGF, EPO, PGC-1α | Improved oxygen transport and energy efficiency | Pathological angiogenesis, oxidative stress, DNA instability |
| Oxidative stress and redox signaling | Intense endurance training, overreaching, nutrient excess | ROS-mediated activation of antioxidant defense and mitochondrial adaptation | Chronic ROS accumulation and oxidative DNA damage | NRF2, NF-κB, SIRT1, p53 | Adaptive hormesis, redox-dependent signaling plasticity | Genomic instability, lipid peroxidation, mitochondrial dysfunction |
| Autophagy–UPR balance | Caloric restriction, intermittent fasting, recovery phases | Controlled proteostasis and organelle turnover | Autophagy dysregulation and ER stress adaptation | AMPK, LC3, ATF4, GRP78 | Protein quality control and metabolic recycling | Proteotoxic stress, ER overload, resistance to apoptosis |
| Epigenetic reprogramming | Nutrient timing, methyl donor availability, supplementation | Transient changes in DNA methylation and histone acetylation for gene regulation | Stable oncogenic methylation, miRNA dysregulation | DNMT1, HDACs, TETs, miR-21 | Flexible gene expression, adaptive remodeling | Epigenetic drift, transcriptional instability, dedifferentiation |
| Hormonal and growth factor signaling | GH/IGF-1 therapy, insulin use, anabolic stacking | Enhanced recovery, anabolic sensitivity | Endocrine-driven tumor growth, hyperinsulinemia | IGF-1R, STAT5, ERK1/2 | Accelerated regeneration, anabolic drive | Endocrine disruption, persistent proliferative signaling |
| Metabolic reprogramming (Warburg-like effect) | High-intensity training, glucose or amino acid loading | Glycolytic shift for ATP and biosynthetic precursor supply | Aerobic glycolysis in tumors (Warburg effect) | HK2, PKM2, LDHA, MYC | Efficient energy flux for adaptation | Persistent glycolysis, one-carbon imbalance, redox exhaustion |
| Epigenetic layer | Primary metabolic cofactor or pathway |
Key enzymes/ regulators |
Physiological role (adaptive) | Pathological outcome (overdrive state) | Representative molecular targets | Evidence from elite sport |
Potential interventions/ modulators |
| DNA methylation | SAM / one-carbon metabolism |
DNMT1, TET1–3, MTHFR, MTR | Hypermethylation of antioxidant genes; hypomethylation of anabolic/proliferative loci; methylation drift |
PGC-1α, PDK4, TFAM, FOXO3, MYC |
Endurance athletes show promoter hypomethylation of PGC-1α and TFAM after repeated training cycles | Folate and methionine balance, caloric restriction, AMPK activation |
|
| Histone acetylation | Acetyl-CoA, NAD+ / AMPK–SIRT1 axis | SIRT1, p300/CBP, GCN5, HDACs | Chromatin compaction under energy deficit; regulation of repair and mitochondrial genes | Hyperacetylation and persistent transcription of anabolic programs |
H3K9ac, H4K16ac, PGC-1α, p53 |
Training fasted or under caloric restriction enhances SIRT1 activity and deacetylation | Resveratrol, curcumin, exercise–fasting cycles |
| Histone and DNA demethylation | α-KG / TCA flux | TETs, Jumonji demethylases | Removal of repressive marks; mitochondrial-nuclear communication | α-KG depletion, succinate/fumarate inhibition of demethylases | H3K27me3, H3K9me3, IDH2, SDH | Overreaching phases show transient TET up-regulation linked to oxidative flux | Endurance exercise, antioxidant restoration |
| NAD+-dependent regulation | NAD+ salvage, PARP vs. sirtuins | PARP1, SIRT1, NAMPT | Balanced DNA repair and mitochondrial biogenesis | PARP overactivation → NAD+ depletion, SIRT1 silencing, redox collapse |
PGC-1α, FOXO, PARP1, SIRT1 |
Overtraining decreases muscle NAD+ and SIRT1, paralleling redox fatigue | Caloric restriction, AMPK activators, NAD+ precursors |
| MicroRNA regulation | Energy/redox-dependent transcription | miR-1, miR-21, miR-34a, miR-486, miR-494 | Fine-tuning of hypertrophy, oxidative capacity, stress defense | Aberrant miRNA expression driving fibrosis and oncogenic signaling |
miR-21, miR-34a, miR-133a, miR-206 |
Endurance ↑ miR-181a, miR-494; resistance ↑ miR-378, miR-486; overtraining ↑ miR-21 | Training modulation, antioxidant support |
| One-carbon and methyl donor flux | Methionine–folate cycle | MAT2A, BHMT, SHMT1/2 | Maintenance of methyl balance for DNA/histone regulation | SAM/SAH imbalance, global hypermethylation, redox-linked drift | DNMT3A, HMTs, MTHFD1L | High-protein diets alter plasma methionine and SAM/SAH ratios in athletes | Controlled protein intake, B-vitamin support |
| System component | Primary molecular driver | Key sensors or enzymes |
Physiological feedback (adaptive) |
Pathological feedback (overdrive state) | Representative molecular signature | Potential modulators / countermeasures |
| Mitochondrial ROS generation | Electron transport flux, NADH/NAD+ ratio | Complex I–III, NOX, SOD2 | ROS act as signaling molecules activating AMPK and antioxidant genes | Chronic ROS accumulation leads to mtDNA damage, lipid peroxidation, and nuclear stress signaling | ↑ ROS, ↑ SOD2, ↑ NRF2 | Endurance training, antioxidant periodization, redox-adaptive nutrition |
| NAD+ metabolism | NAMPT salvage pathway, PARP activity | NAMPT, PARP1, SIRT1 | Balanced NAD+ use supports DNA repair and mitochondrial biogenesis | PARP overactivation depletes NAD+, silencing SIRT1 and impairing repair | ↓ NAD+, ↓ SIRT1, ↑ PARylation | Caloric restriction, resveratrol, niacinamide, AMPK activation |
| AMPK–SIRT1–PGC-1α axis | Energy sensing, NAD+/AMP ratio | AMPK, SIRT1, PGC-1α | Enhances oxidative metabolism, mitophagy, and chromatin integrity | Collapse of AMPK–SIRT1 feedback causes autophagy failure and metabolic rigidity | ↑ PGC-1α, ↑ LC3-II, ↓ FOXO | Exercise-induced AMPK activation, fasting cycles |
| DNA and histone modifications | SAM/SAH ratio, α-KG availability | DNMTs, TETs, HDACs, HATs | Dynamic methylation/acetylation maintains gene expression flexibility | Methylation drift and histone hyperacetylation stabilize maladaptive transcription | ↑ H3K9ac, ↓ TET activity, ↑ DNMT1 | Balanced methyl donor intake, one-carbon flux restoration |
| Inflammatory redox signaling | NF-κB, NLRP3, cytokine ROS loops | NF-κB, IL-6, TNF-α | Transient activation supports repair and immune remodeling | Chronic activation sustains oxidative stress and metabolic block | ↑ NF-κB, ↑ IL-6, ↑ TNF-α | Polyphenols, omega-3s, anti-inflammatory recovery |
| Epigenetic memory and drift | ROS/NAD+-dependent enzyme regulation | SIRT1, PARP1, DNMT1, miRNAs | Transient chromatin remodeling encodes adaptive responses | Persistent oxidative stress leads to irreversible epigenetic drift | Hypomethylated oncogenes, hyperacetylated histones | Controlled recovery, antioxidant therapy, NAD+ support |
| System-level outcome | Redox–epigenetic coupling | Integrated signaling through AMPK–mTOR–SIRT1 | Self-limiting oscillation ensures resilience | Feedback saturation locks system in pathological anabolism | ↓ AMPK, ↑ mTOR, ↑ ROS | Nutritional periodization, training load modulation |
| Regulatory axis | Adaptive (oscillatory) state |
Overdrive (lock-in) state |
Representative molecular indicators |
Functional outcome (physiological/ clinical) |
Evidence or relevance in elite sport | Reversibility potential |
| Nutritional signaling (AMPK–mTOR) | Alternating activation maintains energy balance and anabolic–catabolic cycling; mTOR activity transient and self-limiting. | Chronic nutrient or hormonal stimulation suppresses AMPK; mTORC1 locked in constitutive activation. | Adaptive: ↑ AMPK (Thr172-P), oscillatory ↑ mTOR (Ser2448-P); Overdrive: ↑ S6K1, ↓ AMPK. | Efficient recovery, balanced hypertrophy, metabolic flexibility. | Observed in endurance vs. bulking athletes; chronic protein or insulin use suppresses AMPK signaling. | High — restored through fasting, caloric periodization, AMPK activators. |
| Redox homeostasis (ROS–NAD+ balance) | Controlled ROS pulses activate NRF2–SIRT1 defense; NAD+ recycling sustains redox tone. | ROS accumulation exceeds detox capacity; PARP activation depletes NAD+, silencing SIRT1. | Adaptive: ↑ NRF2, ↑ SIRT1, stable NAD+/NADH; Overdrive: ↑ PARP1, ↓ NAD+, ↑ oxidative adducts. | Enhanced mitochondrial efficiency and repair capacity. | Seen in overtraining and altitude exposure; redox imbalance linked to delayed recovery and fatigue. | Moderate — restored with NAD+ boosters, antioxidant periodization. |
| Epigenetic regulation (chromatin remodeling) | Reversible histone deacetylation and DNA demethylation maintain transcriptional flexibility; stress memory remains functional. | Hyperacetylation and aberrant methylation fix anabolic programs; transcriptional noise accumulates. | Adaptive: ↑ TET1–3, ↑ HDACs; Overdrive: ↑ DNMT1, ↑ HATs, ↑ H3K9ac. | Improved adaptation memory, genomic stability. | Exercise-induced hypomethylation of PGC-1α and TFAM vs. hypermethylation under chronic overload. | Variable — reversible early, lost under prolonged overload. |
| Mitochondrial dynamics | Balanced fusion–fission and autophagic recycling sustain bioenergetic quality. | Hyperpolarization, impaired mitophagy, ROS leakage disrupt energy homeostasis. | Adaptive: ↑ PGC-1α, ↑ MFN2, ↑ LC3-II; Overdrive: ↑ DRP1, ↑ ROS. | Optimal endurance and energy turnover. | Endurance training enhances mitophagy; chronic supplementation or doping impairs mitochondrial turnover. | Moderate — restored through recovery and redox normalization. |
| System-level behavior | Oscillatory negative feedback ensures renewal after stress; system remains dynamically stable. | Positive feedback loops reinforce anabolic and oxidative stress; system enters chaotic saturation. | Adaptive: oscillatory AMPK/mTOR ratio; Overdrive: flattened rhythm, ↑ entropy. | Resilient adaptation, sustained performance. | Elite athletes under chronic load exhibit reduced HRV and hormonal adaptability—biomarkers of systemic lock-in. | Low — requires full metabolic reset through deloading and restoration cycles. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




