Submitted:
06 October 2025
Posted:
07 October 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Related Works
2.1. Sparse Topological Initialization Methods
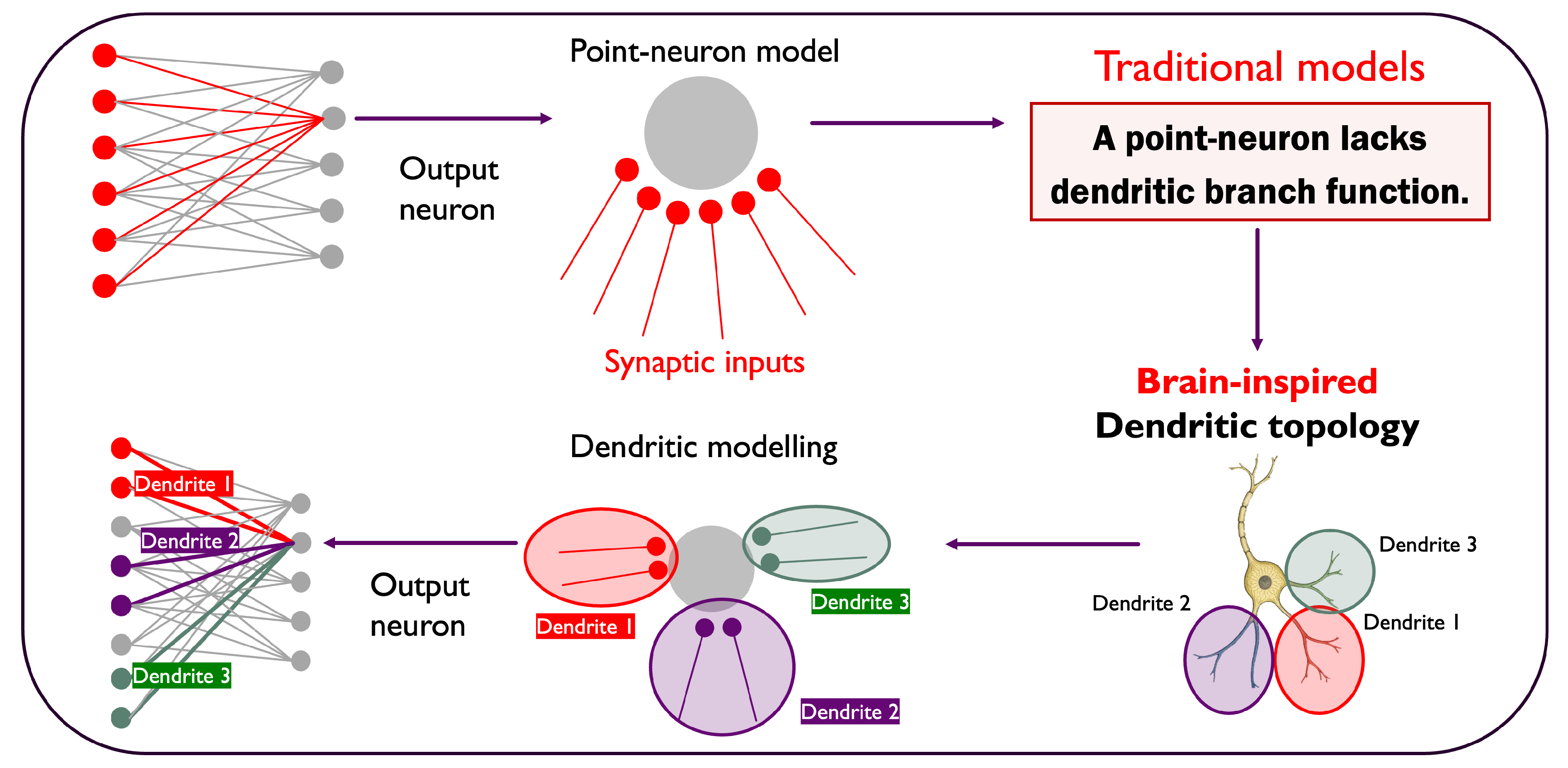
3. The Dendritic Network Model
3.1. Biological Inspiration and Principles
3.2. Parametric Specification
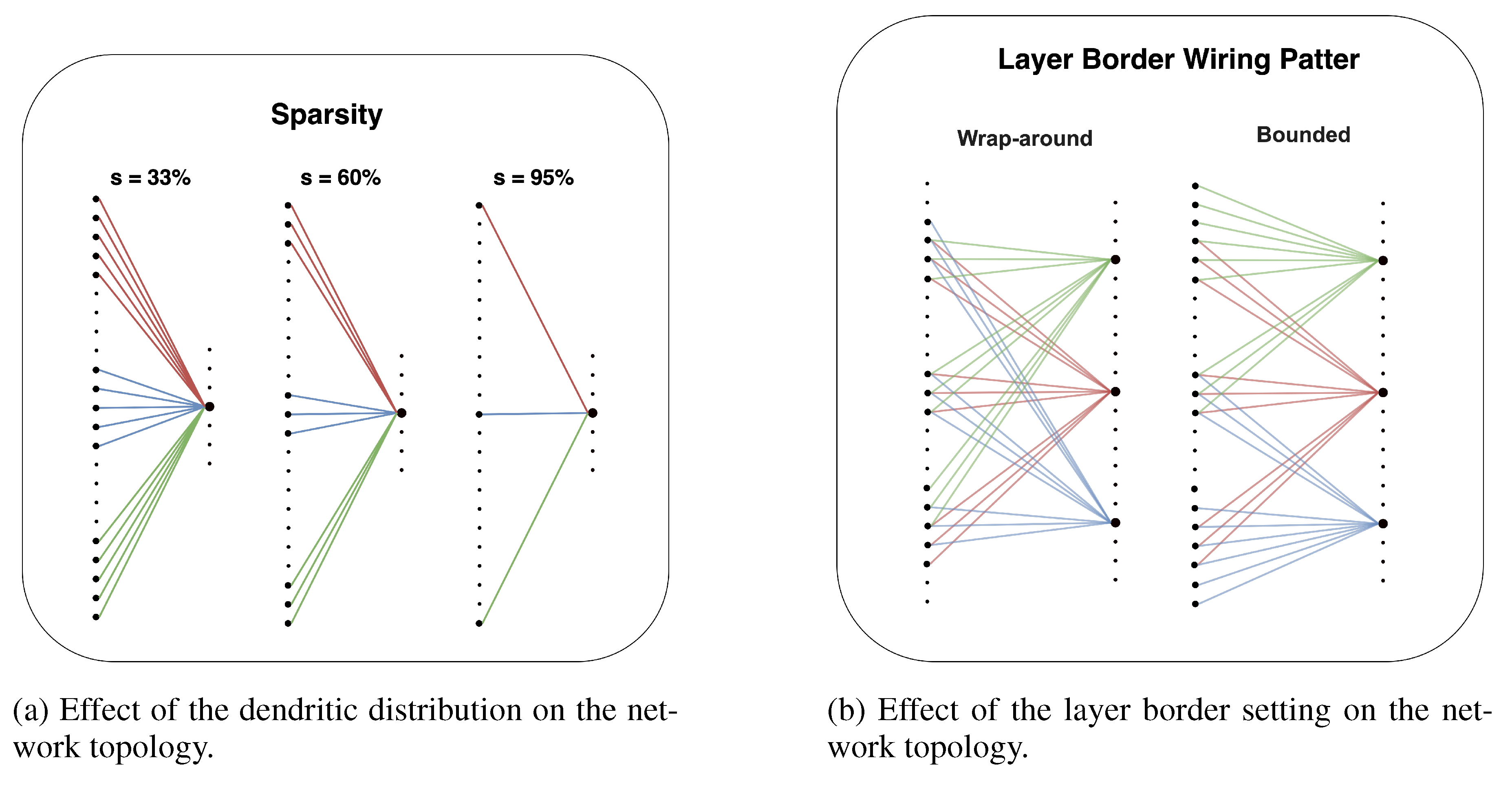
Sparsity (s)
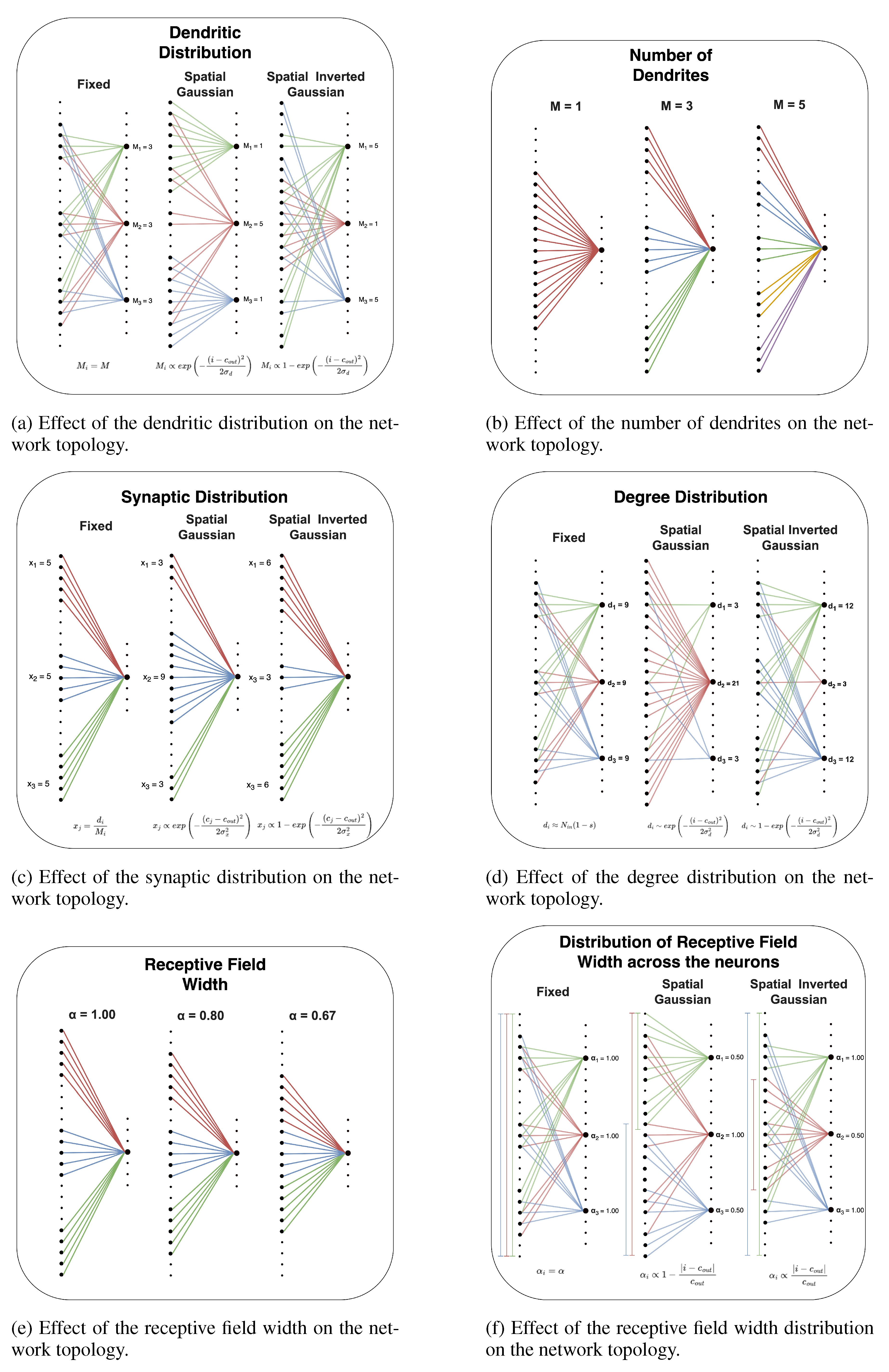
Dendritic Distribution
Receptive Field Width Distribution
Degree Distribution
Synaptic Distribution
Layer Border Wiring Pattern
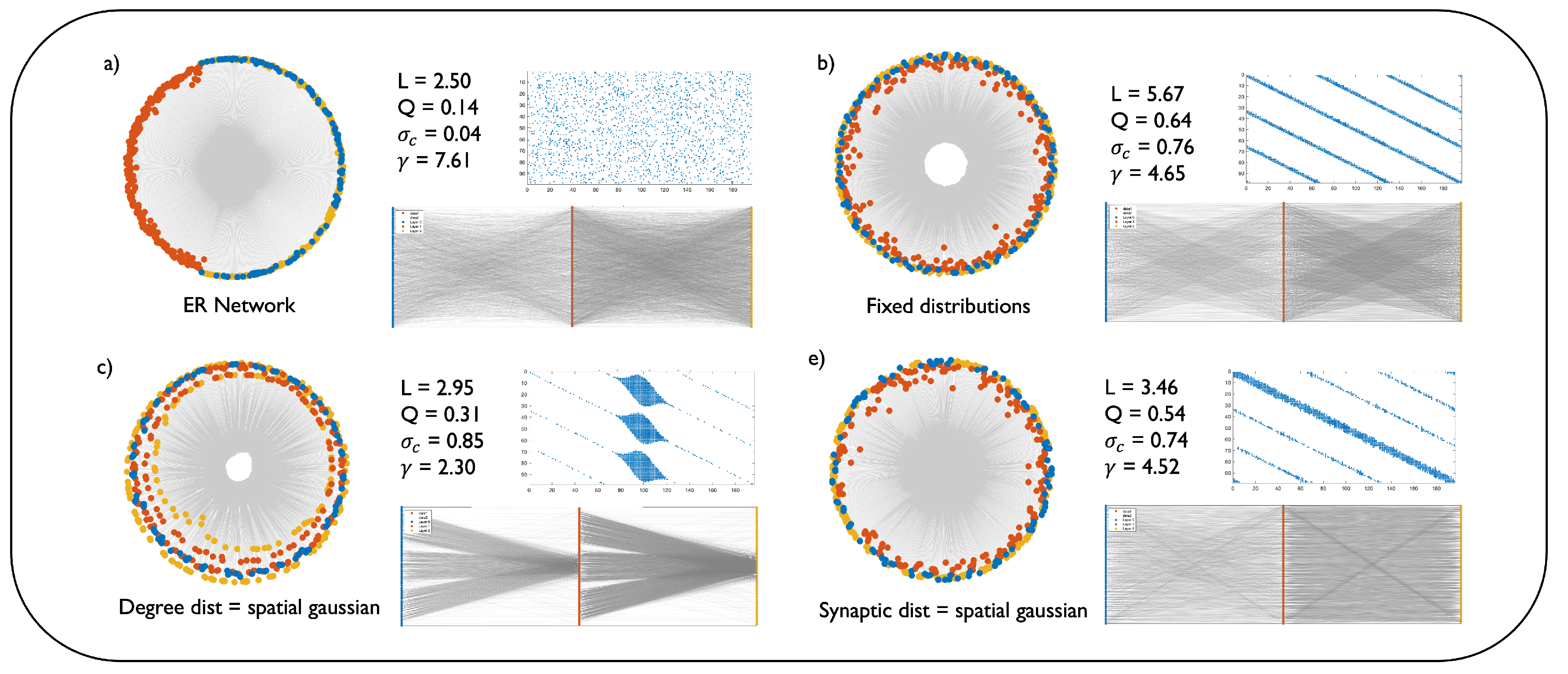
3.3. Network Topology and Geometric Characterization
4. Experiments
4.1. Experimental Setup
Baseline methods
4.2. MLP for Image Classification
Static sparse training
Dynamic sparse training
4.3. Transformer for Machine Translation
5. Results Analysis
6. Conclusions
Appendix A. Glossary of Network Science
Scale-Free Network
Watts-Strogatz Model and Small-World Network
Structural Consistency
Modularity
Characteristic path length
Coalescent Embedding
Appendix B. Hyperparameter Settings and Implementation Details
Appendix B.1. MLP for Image Classification
Appendix B.2. Transformer for Machine Translation
- Multi30k: Trained for 5,000 steps with a learning rate of 0.25, 1000 warmup steps, and a batch size of 1024.
- IWSLT14: Trained for 20,000 steps with a learning rate of 2.0, 6000 warmup steps, and a batch size of 10240.
- WMT17: Trained for 80,000 steps with a learning rate of 2.0, 8000 warmup steps, and a batch size of 12000.
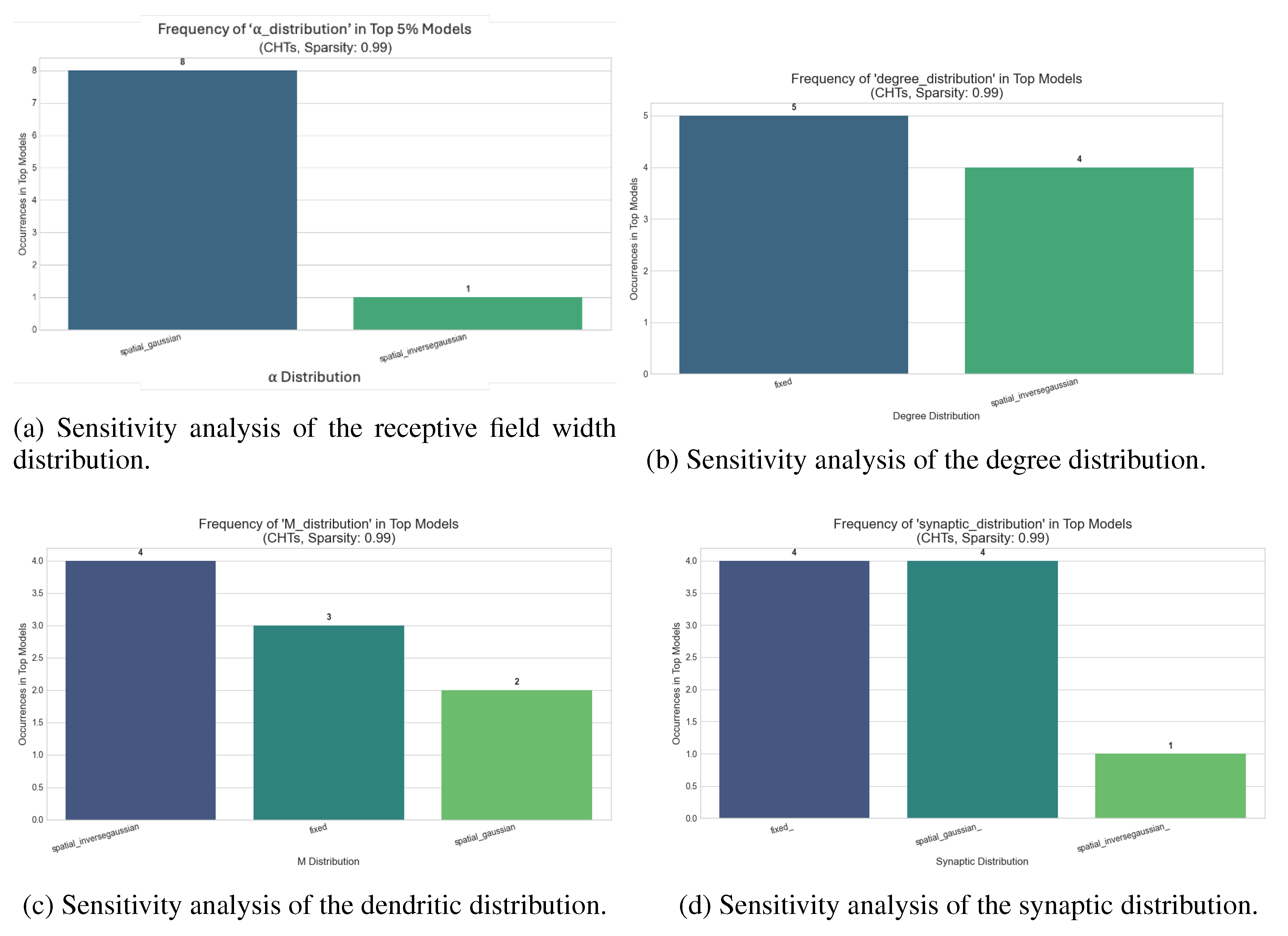
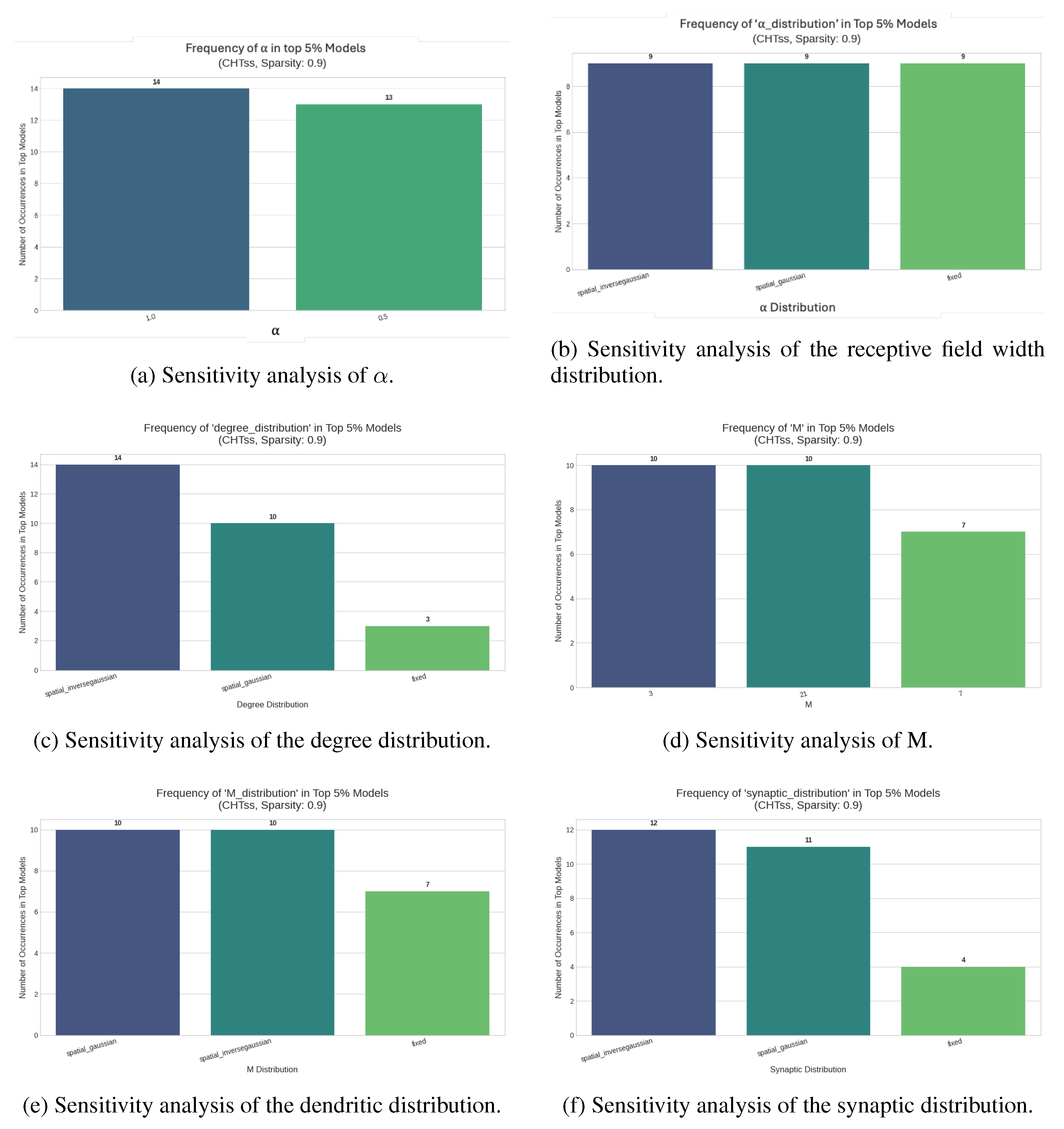
Appendix C. Sensitivity Tests
Dynamic Sparse Training for Image Classification
| MNIST | EMNIST | Fashion MNIST | CIFAR-10 | |||||
|---|---|---|---|---|---|---|---|---|
| Parameter | CHTs | CHTss | CHTs | CHTss | CHTs | CHTss | CHTs | CHTss |
| Degree Dist | 0.001538 | 0.001538 | 0.001419 | 0.009172 | 0.003379 | 0.017639 | 0.017639 | 0.017639 |
| Rec Field Width Dist | 0.000506 | 0.000506 | 0.001585 | 0.006154 | 0.001205 | 0.005381 | 0.005381 | 0.005381 |
| Dendritic Dist | 0.000525 | 0.000525 | 0.001265 | 0.003620 | 0.001136 | 0.004983 | 0.004983 | 0.004983 |
| Synaptic Dist | 0.000464 | 0.000464 | 0.000989 | 0.002538 | 0.000964 | 0.004306 | 0.004306 | 0.004306 |
Dynamic Sparse Training for Machine Translation
| Parameter | Average Coefficient of Variation (CV) | |
|---|---|---|
| 90% | 95% | |
| Dendritic distribution | 0.010643 | 0.009340 |
| Degree distribution | 0.010270 | 0.008436 |
| Receptive Field Width Distribution | 0.009830 | 0.009103 |
| Synaptic distribution | 0.009663 | 0.009310 |
| M | 0.008959 | 0.007806 |
| 0.008210 | 0.007461 | |
| Note: Higher CV indicates greater impact on performance. | ||
Appendix D. Modelling Dendritic Networks
- Determine the degree of each output neuron in the layer based on one of the three distribution strategies (fixed, non-spatial, spatial). A probabilistic rounding and adjustment mechanism ensures that no output neuron is disconnected and the sampled degrees sum precisely to the target total number of connections of the layer.
- Next, for each output neuron j, define a receptive field. This is done by topologically mapping the output neuron’s position at a central point in the input layer and establishing a receptive window around this center. The size of this window is controlled by the parameter , which determines the fraction of the input layer that the neuron can connect to. itself can be fixed or sampled from a spatial or non-spatial distribution.
- For each output neuron, determine the number of dendritic branches, to be used to connect it to the input layer. Again, is determined based on one of the three configurations (fixed for all neurons, or sampled from a distribution that could depend on the neuron’s position in the layer).
- Place the dendrites as dendritic centers within the neuron’s receptive window, spacing them evenly across the window.
- The neuron’s total degree, obtained from step 1, is distributed across its dendrites according to a synaptic distribution (fixed, spatial, or non-spatial). For each dendrite, connections are formed with the input neurons that are spatially closest to its center.
- Finally, the process ensures connection uniqueness and adherence to the precise degree constraints.
Appendix E. Dynamic Sparse Training (DST)
Appendix E.1. Dynamic Sparse Training
Appendix F. Baseline Methods
Appendix F.1. Sparse Network Initialization
Bipartite Scale-Free (BSF)
Bipartite Small-World (BSW)
SNIP
Ramanujan Graphs
- Generate d random permutation matrices, where d is the desired degree of the graph. Each permutation matrix represents a perfect matching, which is a set of edges that connects each node in one layer to exactly one node in the other layer without any overlaps.
- Iteratively combine these matchings. In each step, deterministically decide whether to add or subtract the successive matching to the current adjacency matrix. This decision is made by minimizing a barrier function that ensures that the eigenvalues remain within the Ramanujan bounds.
Bipartite Receptive Field (BRF)
Appendix F.2. Dynamic Sparse Training (DST)
SET [28]
RigL [29]
CHTs and CHTss [26]
Appendix G. Impact of initial sparsity on DNM performance
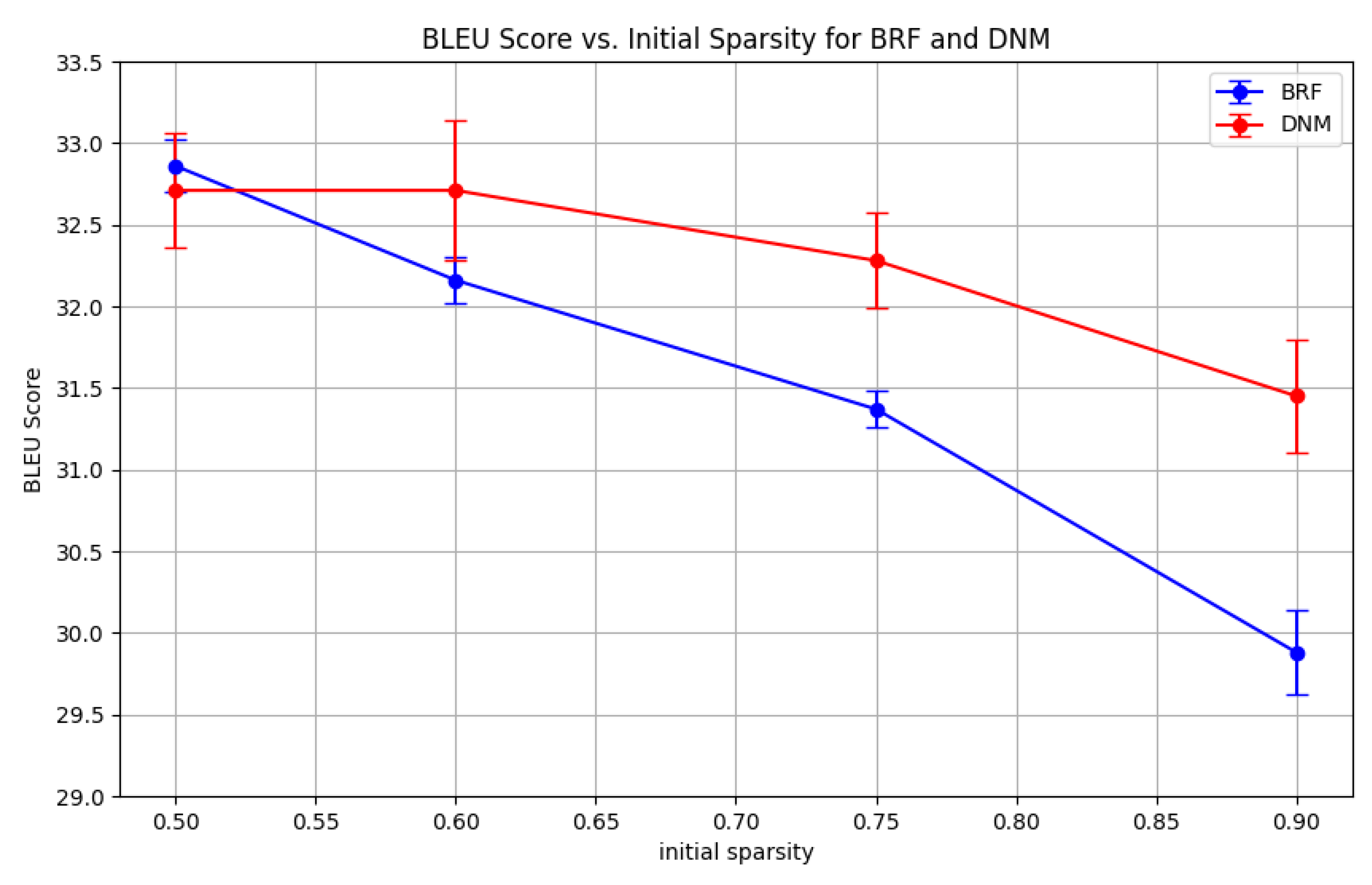
| With parameter search | ||||
|---|---|---|---|---|
| 0.50 | 0.60 | 0.75 | 0.90 | |
| DNM | 32.71±0.35 | 32.71±0.43 | 32.28±0.29 | 31.45±0.35 |
| BRF | 32.86±0.16 | 32.16±0.14 | 31.20±0.36 | 29.81±0.37 |
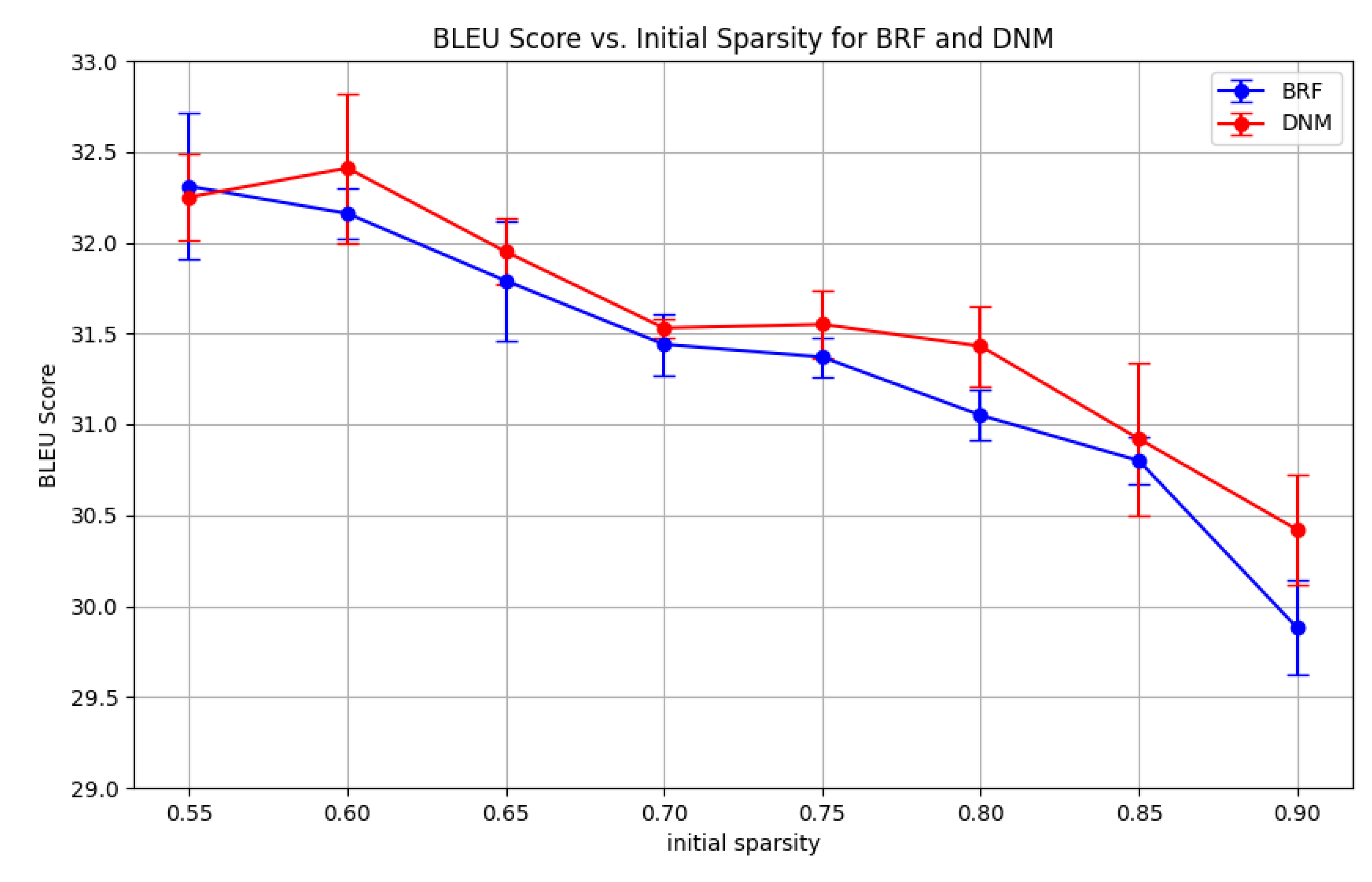
| No DNM parameter search | ||||||||
|---|---|---|---|---|---|---|---|---|
| 0.55 | 0.60 | 0.65 | 0.70 | 0.75 | 0.80 | 0.85 | 0.90 | |
| DNM | 32.25 | 32.41 | 31.95 | 31.53 | 31.55 | 31.43 | 30.92 | 30.42 |
| BRF | 32.31 | 32.16 | 31.79 | 31.44 | 31.37 | 31.05 | 30.80 | 29.88 |
Appendix H. Analysis of the Bounded Layer Border Wiring Pattern
| Static Sparse Training | ||||
|---|---|---|---|---|
| MNIST | EMNIST | Fashion_MNIST | CIFAR10 | |
| Bounded | 97.64±0.10 | 84.00±0.06 | 89.19±0.01 | 61.63±0.18 |
| Wrap-around | 97.82±0.03 | 84.76±0.13 | 88.47±0.03 | 59.04±0.17 |
| SET | ||||
|---|---|---|---|---|
| MNIST | EMNIST | Fashion_MNIST | CIFAR10 | |
| Bounded | 98.40±0.02 | 86.52±0.02 | 89.78±0.09 | 65.67±0.18 |
| Wrap-around | 98.36±0.05 | 86.50±0.06 | 89.75±0.04 | 64.81±0.01 |
| CHTs | ||||
|---|---|---|---|---|
| MNIST | EMNIST | Fashion_MNIST | CIFAR10 | |
| Bounded | 98.66±0.03 | 87.35±0.00 | 90.68±0.09 | 68.03±0.14 |
| Wrap-around | 98.62±0.01 | 87.40±0.04 | 90.62±0.16 | 68.76±0.11 |
| CHTss | ||||
|---|---|---|---|---|
| MNIST | EMNIST | Fashion_MNIST | CIFAR10 | |
| Bounded | 98.66±0.03 | 87.37±0.04 | 90.68±0.09 | 68.03±0.14 |
| Wrap-around | 98.90±0.01 | 87.55±0.01 | 90.88±0.03 | 68.50±0.21 |
Appendix I. Transferability of Optimal Topologies
Experimental Design
Results
| Static Sparse Training | |||
|---|---|---|---|
| Fashion MNIST | EMNIST | CIFAR10 | |
| FC | 90.88±0.02 | 87.13±0.04 | 62.85±0.16 |
| CSTI | 88.52±0.14 | 84.66±0.13 | 52.64±0.30 |
| SNIP | 87.85±0.22 | 84.08±0.08 | 61.81±0.58 |
| Random | 87.34±0.11 | 82.66±0.08 | 55.28±0.09 |
| BSW | 88.18±0.18 | 82.94±0.06 | 56.54±0.15 |
| BRF | 87.41±0.13 | 82.98±0.02 | 54.73±0.07 |
| Ramanujan | 86.45±0.15 | 81.80±0.13 | 55.05±0.40 |
| DNM (Fine-tuned) | 89.19±0.01 | 84.76±0.13 | 61.63±0.18 |
| DNM (Transferred) | 87.98±0.06 | 83.67±0.06 | 58.70±0.42 |
Discussion
Appendix J. Reproduction statement
Appendix K. Claim of LLM Usage
References
- Drachman, D.A. Do we have brain to spare?, 2005.
- Walsh, C.A. Peter Huttenlocher (1931–2013). Nature 2013, 502, 172–172.
- Cuntz, H.; Forstner, F.; Borst, A.; Häusser, M. One rule to grow them all: a general theory of neuronal branching and its practical application. PLoS computational biology 2010, 6, e1000877. [CrossRef]
- London, M.; Häusser, M. Dendritic computation. Annu. Rev. Neurosci. 2005, 28, 503–532. [CrossRef]
- Larkum, M. A cellular mechanism for cortical associations: an organizing principle for the cerebral cortex. Trends in neurosciences 2013, 36, 141–151. [CrossRef]
- Poirazi, P.; Brannon, T.; Mel, B.W. Pyramidal neuron as two-layer neural network. Neuron 2003, 37, 989–999. [CrossRef]
- Lauditi, C.; Malatesta, E.M.; Pittorino, F.; Baldassi, C.; Brunel, N.; Zecchina, R. Impact of Dendritic Nonlinearities on the Computational Capabilities of Neurons. PRX Life 2025, 3, 033003.
- Baek, E.; Song, S.; Baek, C.K.; Rong, Z.; Shi, L.; Cannistraci, C.V. Neuromorphic dendritic network computation with silent synapses for visual motion perception. Nature Electronics 2024, 7, 454–465. [CrossRef]
- Jones, I.S.; Kording, K.P. Might a single neuron solve interesting machine learning problems through successive computations on its dendritic tree? Neural Computation 2021, 33, 1554–1571. [CrossRef]
- Jones, I.S.; Kording, K.P. Do biological constraints impair dendritic computation? Neuroscience 2022, 489, 262–274. [CrossRef]
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proceedings of the IEEE 2002, 86, 2278–2324. [CrossRef]
- Cohen, G.; Afshar, S.; Tapson, J.; Van Schaik, A. EMNIST: Extending MNIST to handwritten letters. In Proceedings of the 2017 international joint conference on neural networks (IJCNN). IEEE, 2017, pp. 2921–2926.
- Xiao, H.; Rasul, K.; Vollgraf, R. Fashion-mnist: a novel image dataset for benchmarking machine learning algorithms. arXiv preprint arXiv:1708.07747 2017.
- Krizhevsky, A. Learning Multiple Layers of Features from Tiny Images 2009. pp. 32–33.
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, .; Polosukhin, I. Attention is all you need. Advances in neural information processing systems 2017, 30.
- Elliott, D.; Frank, S.; Sima’an, K.; Specia, L. Multi30K: Multilingual English-German Image Descriptions. In Proceedings of the Proceedings of the 5th Workshop on Vision and Language. Association for Computational Linguistics, 2016, pp. 70–74. https://doi.org/10.18653/v1/W16-3210.
- Cettolo, M.; Niehues, J.; Stüker, S.; Bentivogli, L.; Federico, M. Report on the 11th IWSLT evaluation campaign. In Proceedings of the Proceedings of the 11th International Workshop on Spoken Language Translation: Evaluation Campaign; Federico, M.; Stüker, S.; Yvon, F., Eds., Lake Tahoe, California, 4-5 2014; pp. 2–17.
- Bojar, O.; Chatterjee, R.; Federmann, C.; Graham, Y.; Haddow, B.; Huang, S.; Huck, M.; Koehn, P.; Liu, Q.; Logacheva, V.; et al. Findings of the 2017 conference on machine translation (wmt17). Association for Computational Linguistics, 2017.
- Erdős, P.; Rényi, A. On the evolution of random graphs. Publication of the Mathematical Institute of the Hungarian Academy of Sciences 1960, 5, 17–60.
- Watts, D.J.; Strogatz, S.H. Collective dynamics of ’small-world’ networks. Nature 1998, 393, 440–442. [CrossRef]
- Barabási, A.L.; Albert, R. Emergence of scaling in random networks. science 1999, 286, 509–512. [CrossRef]
- Zhang, Y.; Zhao, J.; Wu, W.; Muscoloni, A.; Cannistraci, C.V. Epitopological learning and Cannistraci-Hebb network shape intelligence brain-inspired theory for ultra-sparse advantage in deep learning. In Proceedings of the The Twelfth International Conference on Learning Representations, 2024.
- Zhang, Y.; Zhao, J.; Wu, W.; Muscoloni, A. Epitopological learning and cannistraci-hebb network shape intelligence brain-inspired theory for ultra-sparse advantage in deep learning. In Proceedings of the The Twelfth International Conference on Learning Representations, 2024.
- Lee, N.; Ajanthan, T.; Torr, P.H. Snip: Single-shot network pruning based on connection sensitivity. arXiv preprint arXiv:1810.02340 2018.
- Kalra, S.A.; Biswas, A.; Mitra, P.; Basu, B. Sparse Neural Architectures via Deterministic Ramanujan Graphs. Transactions on Machine Learning Research.
- Zhang, Y.; Cerretti, D.; Zhao, J.; Wu, W.; Liao, Z.; Michieli, U.; Cannistraci, C.V. Brain network science modelling of sparse neural networks enables Transformers and LLMs to perform as fully connected. arXiv preprint arXiv:2501.19107 2025.
- Cacciola, A.; Muscoloni, A.; Narula, V.; Calamuneri, A.; Nigro, S.; Mayer, E.A.; Labus, J.S.; Anastasi, G.; Quattrone, A.; Quartarone, A.; et al. Coalescent embedding in the hyperbolic space unsupervisedly discloses the hidden geometry of the brain. arXiv preprint arXiv:1705.04192 2017.
- Mocanu, D.C.; Mocanu, E.; Stone, P.; Nguyen, P.H.; Gibescu, M.; Liotta, A. Scalable training of artificial neural networks with adaptive sparse connectivity inspired by network science. Nature communications 2018, 9, 2383. [CrossRef]
- Evci, U.; Gale, T.; Menick, J.; Castro, P.S.; Elsen, E. Rigging the lottery: Making all tickets winners. In Proceedings of the International conference on machine learning. PMLR, 2020, pp. 2943–2952.
- Cettolo, M.; Girardi, C.; Federico, M. Wit3: Web inventory of transcribed and translated talks. Proceedings of EAMT 2012, pp. 261–268.
- Lü, L.; Pan, L.; Zhou, T.; Zhang, Y.C.; Stanley, H.E. Toward link predictability of complex networks. Proceedings of the National Academy of Sciences 2015, 112, 2325–2330. [CrossRef]
- Newman, M.E. Modularity and community structure in networks. Proceedings of the national academy of sciences 2006, 103, 8577–8582.
- Cannistraci, C.V.; Muscoloni, A. Geometrical congruence, greedy navigability and myopic transfer in complex networks and brain connectomes. Nature Communications 2022, 13, 7308. [CrossRef]
- Muscoloni, A.; Thomas, J.M.; Ciucci, S.; Bianconi, G.; Cannistraci, C.V. Machine learning meets complex networks via coalescent embedding in the hyperbolic space. Nature communications 2017, 8, 1615. [CrossRef]
- Balasubramanian, M.; Schwartz, E.L. The isomap algorithm and topological stability. Science 2002, 295, 7–7. [CrossRef]
- Muscoloni, A.; Abdelhamid, I.; Cannistraci, C.V. Local-community network automata modelling based on length-three-paths for prediction of complex network structures in protein interactomes, food webs and more. BioRxiv 2018, p. 346916.
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 2014.
- Bellec, G.; Kappel, D.; Maass, W.; Legenstein, R. Deep rewiring: Training very sparse deep networks. arXiv preprint arXiv:1711.05136 2017.
- Yuan, G.; Ma, X.; Niu, W.; Li, Z.; Kong, Z.; Liu, N.; Gong, Y.; Zhan, Z.; He, C.; Jin, Q.; et al. Mest: Accurate and fast memory-economic sparse training framework on the edge. Advances in Neural Information Processing Systems 2021, 34, 20838–20850.
- Jayakumar, S.; Pascanu, R.; Rae, J.; Osindero, S.; Elsen, E. Top-kast: Top-k always sparse training. Advances in Neural Information Processing Systems 2020, 33, 20744–20754.
- Lasby, M.; Golubeva, A.; Evci, U.; Nica, M.; Ioannou, Y. Dynamic sparse training with structured sparsity. arXiv preprint arXiv:2305.02299 2023.
- Lubotzky, A.; Phillips, R.; Sarnak, P. Ramanujan graphs. Combinatorica 1988, 8, 261–277.
- Marcus, A.; Spielman, D.A.; Srivastava, N. Interlacing families I: Bipartite Ramanujan graphs of all degrees. In Proceedings of the 2013 IEEE 54th Annual Symposium on Foundations of computer science. IEEE, 2013, pp. 529–537.
- Muscoloni, A.; Michieli, U.; Zhang, Y.; Cannistraci, C.V. Adaptive network automata modelling of complex networks 2022.
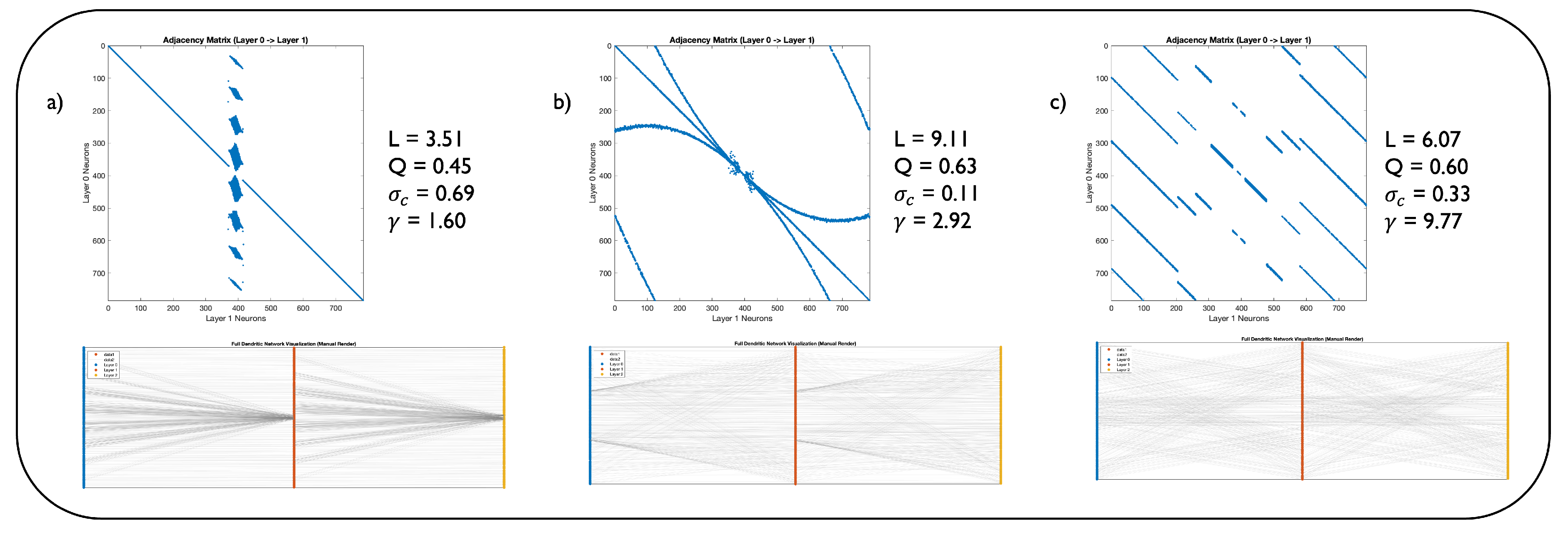
| 1 | To facilitate an intuitive exploration of this landscape, we have developed an interactive web application where readers can adjust the model’s parameters and visualize the resulting network structures. The application is available at: https://dendritic-network-model.streamlit.app/
|
| Static Sparse Training | ||||
|---|---|---|---|---|
| MNIST | Fashion MNIST | EMNIST | CIFAR10 | |
| FC | 98.78±0.02 | 90.88±0.02 | 87.13±0.04 | 62.85±0.16 |
| CSTI | 98.07±0.02 | 88.52±0.14 | 84.66±0.13 | 52.64±0.30 |
| SNIP | 97.59±0.08 | 87.85±0.22 | 84.08±0.08 | 61.81±0.58 |
| Random | 96.72±0.04 | 87.34±0.11 | 82.66±0.08 | 55.28±0.09 |
| BSW | 97.32±0.02 | 88.18±0.18 | 82.94±0.06 | 56.54±0.15 |
| BRF | 96.85±0.01 | 87.41±0.13 | 82.98±0.02 | 54.73±0.07 |
| Ramanujan | 96.51±0.17 | 86.45±0.15 | 81.80±0.13 | 55.05±0.40 |
| DNM | 97.82±0.03 | 89.19±0.01 | 84.76±0.13 | 61.63±0.18 |
| SET | ||||
|---|---|---|---|---|
| MNIST | Fashion MNIST | EMNIST | CIFAR10 | |
| FC | 98.78±0.02 | 90.88±0.02 | 87.13±0.04 | 62.85±0.16 |
| SNIP | 98.31±0.05 | 89.49±0.05 | 86.44±0.09 | 64.26±0.14 |
| CSTI | 98.47±0.02 | 89.91±0.11 | 86.66±0.06 | 65.05±0.14 |
| Random | 98.14±0.02 | 89.00±0.09 | 86.31±0.08 | 62.70±0.11 |
| BSW | 98.25±0.02 | 89.25±0.03 | 86.26±0.08 | 63.66±0.07* |
| BRF | 98.25±0.01 | 89.44±0.01 | 86.20±0.04 | 62.64±0.10 |
| Ramanujan | 97.96±0.05 | 89.16±0.07 | 85.80±0.03 | 62.58±0.20 |
| DNM | 98.40±0.02 | 89.78±0.09 | 86.52±0.02 | 65.67±0.18* |
| RigL | ||||
|---|---|---|---|---|
| MNIST | Fashion MNIST | EMNIST | CIFAR10 | |
| FC | 98.78±0.02 | 90.88±0.02 | 87.13±0.04 | 62.85±0.16 |
| SNIP | 98.75±0.04 | 89.89±0.01 | 87.34±0.03 | 63.79±0.23 |
| CSTI | 98.76±0.02 | 90.14±0.10 | 87.41±0.01 | 60.52±0.31 |
| Random | 98.61±0.01 | 89.53±0.04 | 86.99±0.09 | 64.24±0.07* |
| BSW | 98.71±0.06 | 89.85±0.07 | 87.18±0.07* | 65.06±0.19* |
| BRF | 98.65±0.03 | 89.76±0.09 | 87.08±0.05 | 64.33±0.16* |
| Ramanujan | 98.42±0.03 | 89.80±0.07 | 87.09±0.04 | 64.31±0.08* |
| DNM | 98.75±0.04 | 90.08±0.09 | 87.28±0.28* | 65.68±0.01* |
| CSTI | Random | BSW | BRF | Ramanujan | DNM | |
|---|---|---|---|---|---|---|
| FC | 62.85±0.16 | |||||
| CHTs | 69.77±0.06 | 66.96±0.24 | 66.96±0.24 | 64.96±0.17 | 67.19±0.17 | 68.76±0.11 |
| CHTss | 71.29±0.14 | 66.89±0.23 | 66.89±0.23 | 64.96±0.17 | 67.37±0.12 | 68.50±0.21 |
| MNIST | Fashion MNIST | EMNIST | ||||
|---|---|---|---|---|---|---|
| CHTs | CHTss | CHTs | CHTss | CHTs | CHTss | |
| FC | 98.78±0.02 | 90.88±0.02 | 87.13±0.04 | |||
| CSTI | 98.81±0.04 | 98.83±0.02 | 90.93±0.03 | 90.81±0.11 | 87.82±0.04 | 87.52±0.04 |
| Random | 98.57±0.04 | 98.61±0.03 | 90.42±0.03 | 90.30±0.10 | 87.12±0.13 | 87.20±0.09 |
| BSW | 98.57±0.04 | 98.61±0.04 | 90.46±0.06 | 90.46±0.06 | 87.12±0.13 | 87.20±0.09 |
| BRF | 98.47±0.03 | 98.47±0.03 | 90.04±0.12 | 90.04±0.12 | 87.03±0.07 | 87.03±0.07 |
| Ramnujan | 98.57±0.03 | 98.57±0.03 | 89.82±0.14 | 98.78±0.08 | 87.24±0.08 | 87.24±0.08 |
| DNM | 98.66±0.03 | 98.90±0.01* | 90.68±0.09 | 90.88±0.03* | 87.40±0.04 | 87.55±0.01* |
| Multi30k | IWSLT | |||
| 0.95 | 0.90 | 0.95 | 0.90 | |
| FC | 31.38±0.38 | 24.48±0.30 | ||
| CHTsB | 28.94±0.57 | 29.81±0.37 | 21.15±0.10 | 21.92±0.17 |
| CHTsD | 30.54±0.42 | 31.45±0.35* | 22.09±0.14 | 23.52±0.24 |
| CHTssB | 32.03±0.29* | 32.86±0.16* | 24.51±0.02* | 24.31±0.04 |
| CHTssD | 32.62±0.28* | 33.00±0.31* | 24.43±0.14 | 24.20±0.07 |
| Model | WMT | |
| 0.95 | 0.90 | |
| FC | 25.52 | |
| CHTsB | 20.94±0.63 | 22.40±0.06 |
| CHTsD | 21.34±0.20 | 22.56±0.14 |
| CHTssB | 23.73±0.43 | 24.61±0.14 |
| CHTssD | 24.52±0.12 | 24.40±0.15 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 1996 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).




