Submitted:
01 October 2025
Posted:
02 October 2025
You are already at the latest version
Abstract
Keywords:
Introduction
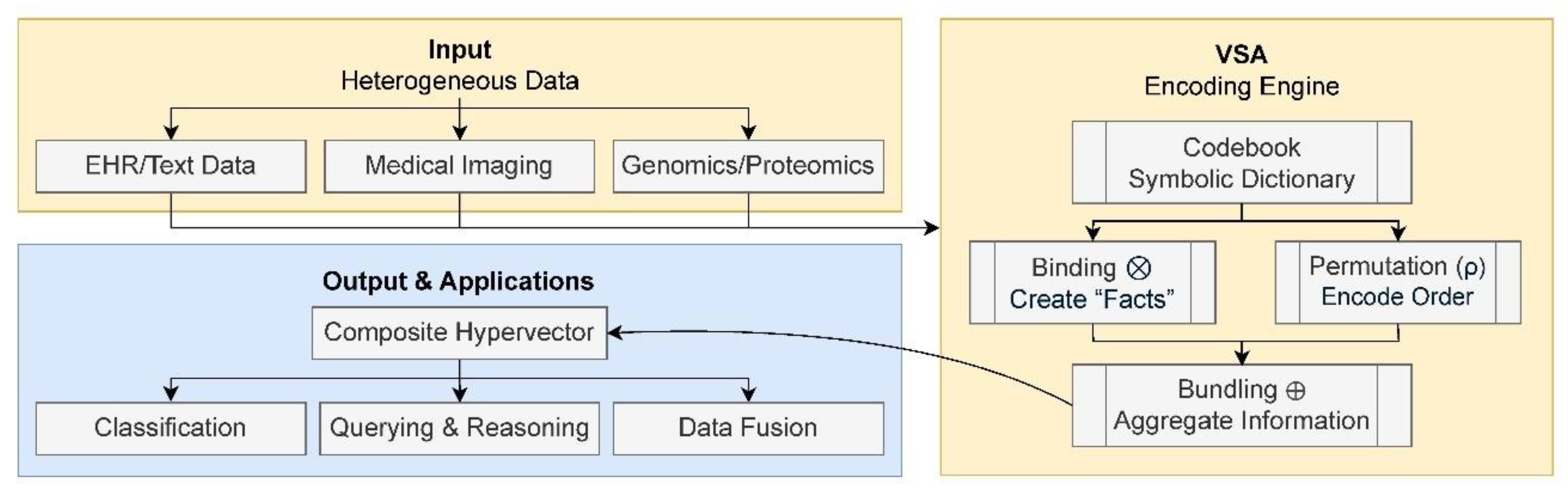
- Binding (element-wise multiplication, ⊗): links two concepts represented in the vectors together. For example, binding the vector representation of the concept serum level with the vector representation of the concept high creates a new, distinct hypervector representing the composite idea high serum level;
- Bundling (element-wise addition, ⊕): combines multiple concepts in vector format. One could bundle the vectors for high serum glucose, high blood pressure, and patient ID: 123 to create a single vector representing a patient’s state. This operation highlights a key property of HDC: the ability to easily integrate heterogeneous information, numerical values, categorical states, or identifiers, into one coherent data structure;
- Permutation (rotation, ρ): encodes order of sequences, which is critical for representing sequential data like DNA strands or time-series events in a patient’s record. In this operation, each hypervector is shifted or rotated according to its position in the sequence, ensuring that the same concept appearing at different positions is represented uniquely. This positional encoding preserves order while keeping the representations nearly orthogonal, thereby allowing clear distinction between sequences and their components.
Tip 1: For Electronic Health Records – Use Bundling to Create Holistic Patient Vectors
How to Create a Patient Vector
- Keys for data types: diagnosis, medication, lab_test, value;
- Specific diagnose: diabetes_type_2, hypertension;
- Specific medications: metformin, lisinopril;
- Specific lab tests: hba1c, creatinine;
- Discretized lab values: high, normal, low.


- A diagnosis of hypertension becomes: patient_diagnosis = diagnosis ⊗ hypertension
- A prescription for metformin becomes: patient_prescription = medication ⊗ metformin
- A HbA1c result of 8.1% (which we categorize as high) becomes: patient_hba1c = (lab_test ⊗ hba1c) ⊕ (value ⊗ high)
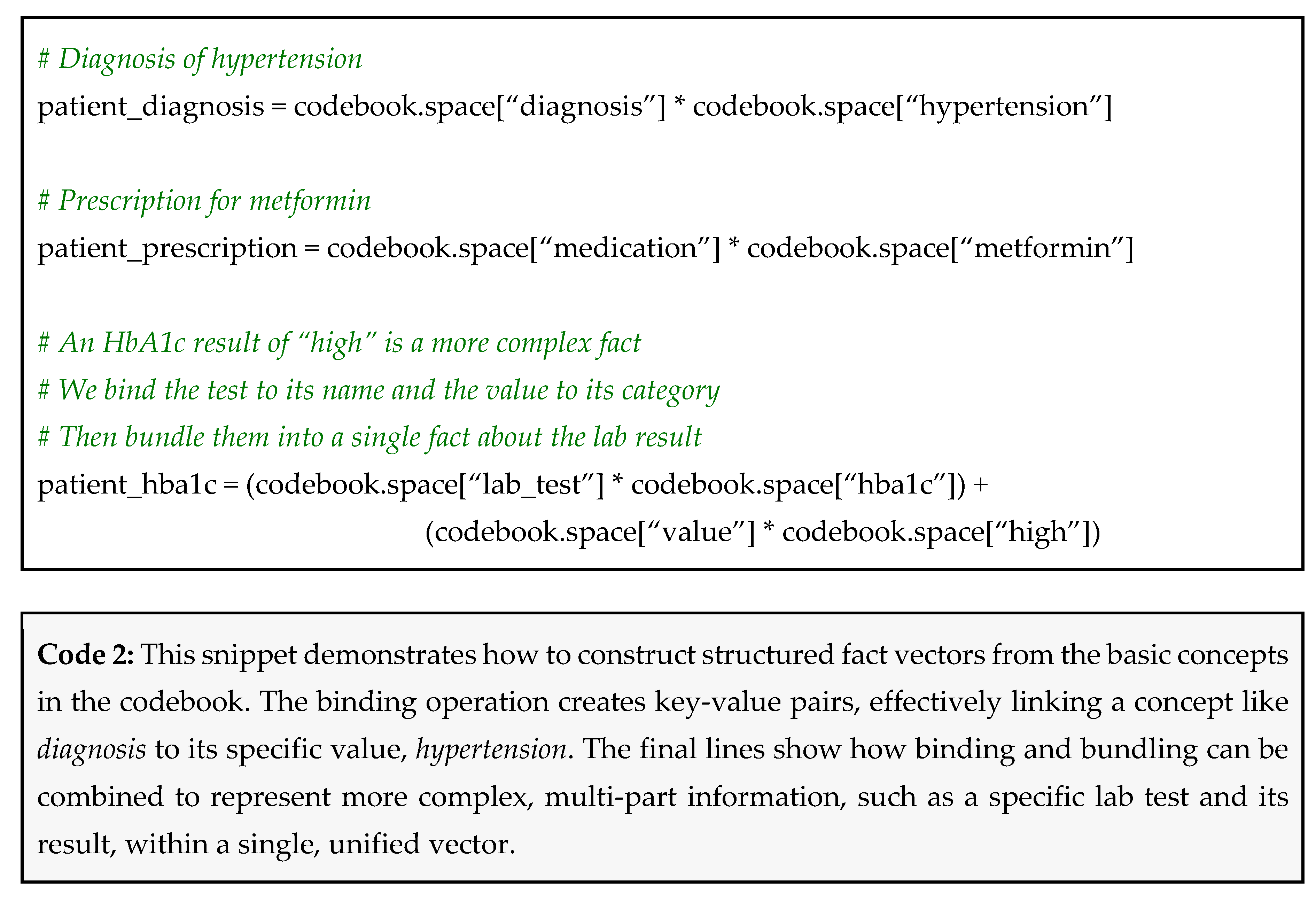

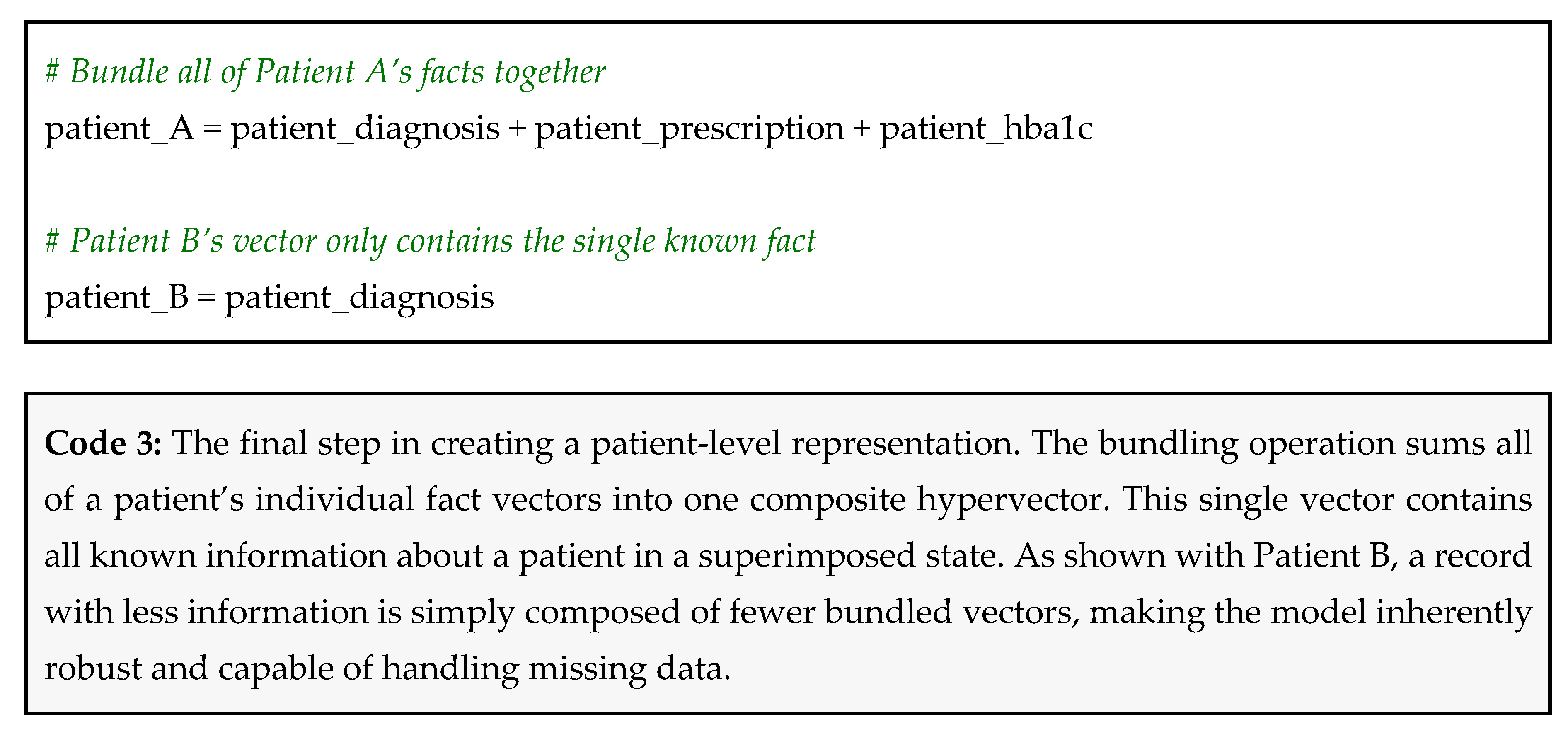
Why This Works
- Robustness to missing data: Patient A’s vector was created without knowing their creatinine level. Patient B’s vector was created with only one piece of information. The model does not break or require modification. It works with whatever data is available [4];
- Similarity-based reasoning: the magic happens when you compare patient vectors. The vector for Patient B is a component of the vector for Patient A. Mathematically, this means that their two vectors have a high degree of similarity. Thus, in general, you can find patients with similar clinical profiles by simply looking for vectors having high cosine similarity. Note that hdlib provides a dist function as Vector’s instance method to perform the cosine distance between the specific instance vector and a different Vector object, defined as 1 minus the cosine similarity. Following the previous example shown in Code 3, we can compute the cosine distance between patient_A and patient_B vectors using hdlib as: patient_A.dist(patient_B, method=“cosine”) [24].
Tip 2: For Genomics and Proteomics – Leverage Permutations to Encode Sequences
Encoding Positions
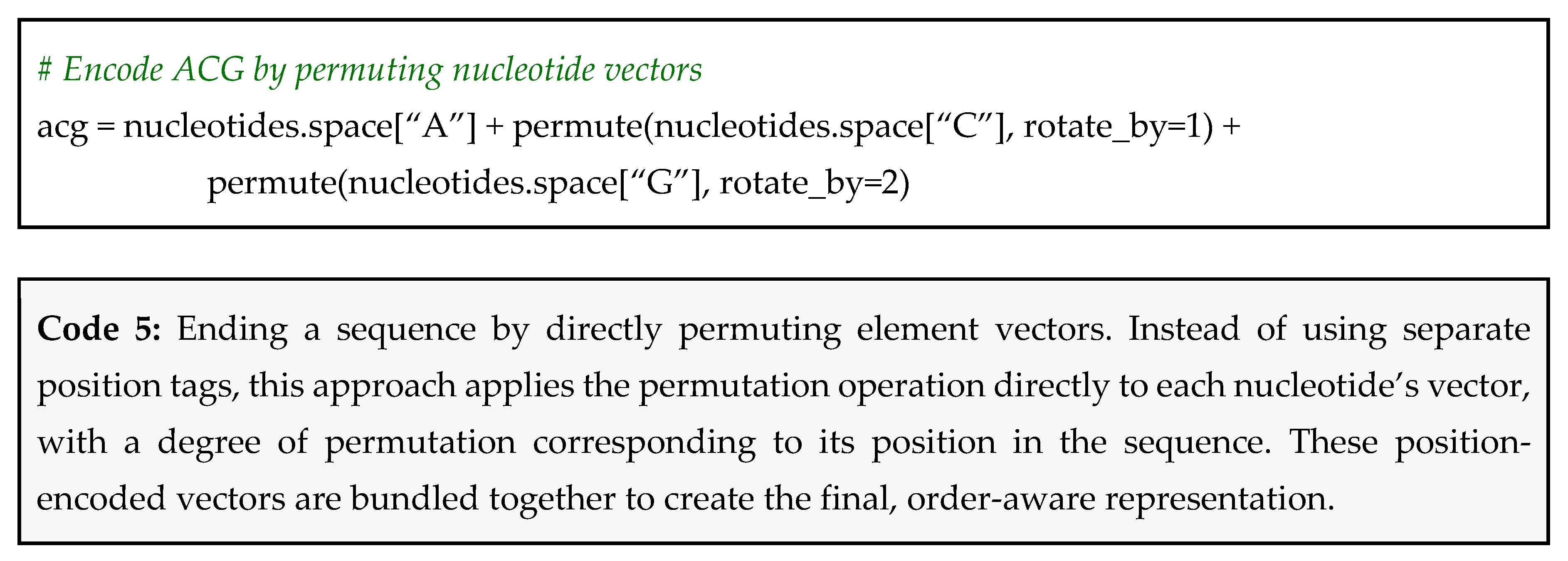


Which Method Should You Choose?
- Choose method 1 if your primary task is querying by position. For instance, if you frequently need to ask questions like “what nucleotide is at position 257?”, method 1 excels because you can directly query the full sequence vector with the position_257 vector to retrieve the answer;
- Choose method 2 if your goal is general sequence comparison, alignment, or if you need to ask questions like “at what positions does Adenine (“A”) appear?”. This approach is more elegant and memory efficient and requires a smaller codebook.
Tip 3: For Medical Imaging – Combine Vector-Symbolic Architectures with Convolutional Neural Networks for Explicable Artificial Intelligence
Building a Queryable Scene Description
- CNN acts as a junior technician. It has incredible eyes and can spot thousands of tiny, low-level patterns (textures, edges, shapes) in an image but cannot articulate what they mean;
- VSA acts as a senior pathologist. It takes the technician’s findings, gives them symbolic names (e.g., “spindle-shaped cells”, “high nuclear density”), notes their locations, and assembles everything into a structured, queryable report.
- Discover feature groups: we first run a clustering algorithm (like K-Means) on all the feature vectors produced by the CNN from a training set of images. This groups the thousands of slightly different numerical feature vectors into a small number of distinct clusters. Each cluster represents a recurring visual pattern, like a certain texture or cell shape;
- Assign symbolic names: we then treat each cluster as a single, unified concept. We assign a symbolic name to each one and generate a unique, random hypervector for each of these symbolic names in our codebook.
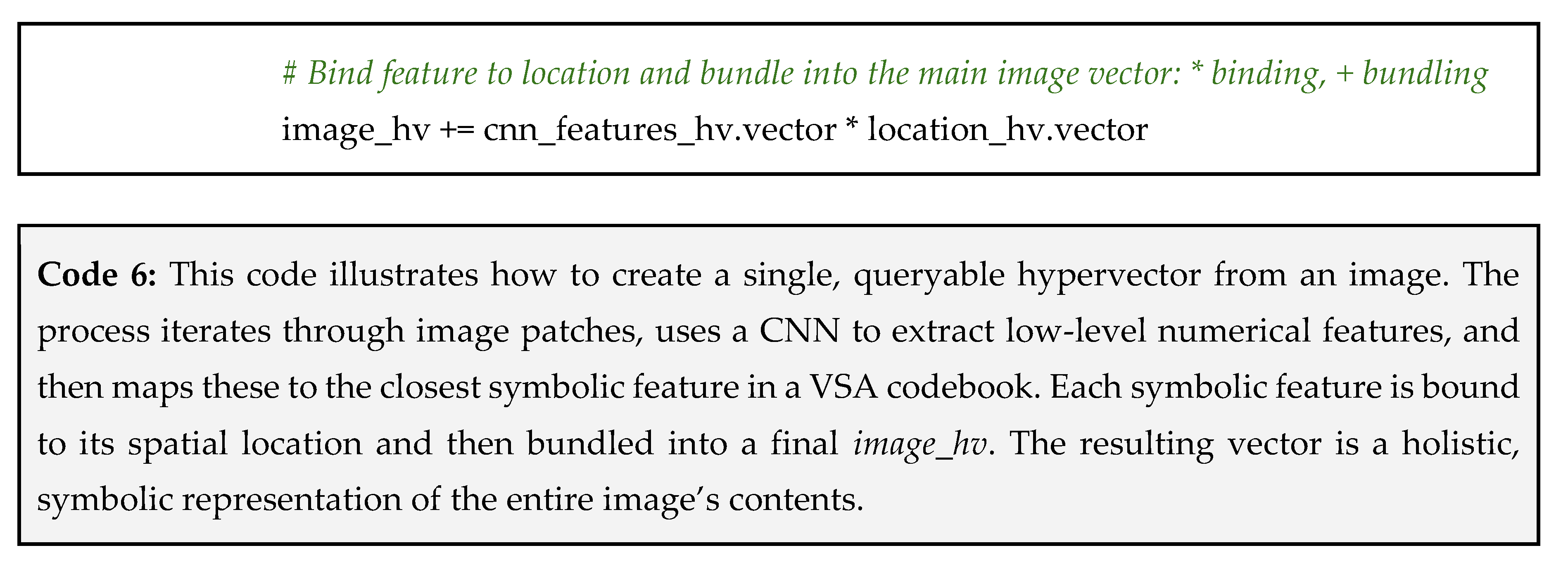


Why This Works

Tip 4: For Biosignal Processing (EEG/ECG) – Encode Time-Frequency Data Symbolically
A Symbolic Snapshot of the Signal’s State


Why This Works
- Create a prototype: collect many brain_state_hv samples while a subject is focused and bundle them together to create a single prototype_focused vector. Do the same for the relaxed state to create prototype_relaxed;
- Classify new data: to classify a new, unseen brain_state_hv after following the same encoding steps of a single test sample, simply check whether it is more similar to the focused or relaxed prototype (see Code 9 for a pseudocode using hdlib).

Tip 5: For Molecular Structures – Decompose Molecules into Atomic Fragments
A Sum of Its Parts
- o oxygen fragment: one central oxygen atom connected to two hydrogens via single bonds;
- o hydrogen fragment: each hydrogen is connected to one oxygen via a single bond.


Why This Works
Tip 6: For Biomedical Knowledge Graphs – Use Binding to Represent Relational Triples
Facts as Vectors


Why This Works

Tip 7: For Classification Tasks – Build and Refine Prototype Vectors
Learning the Average Case
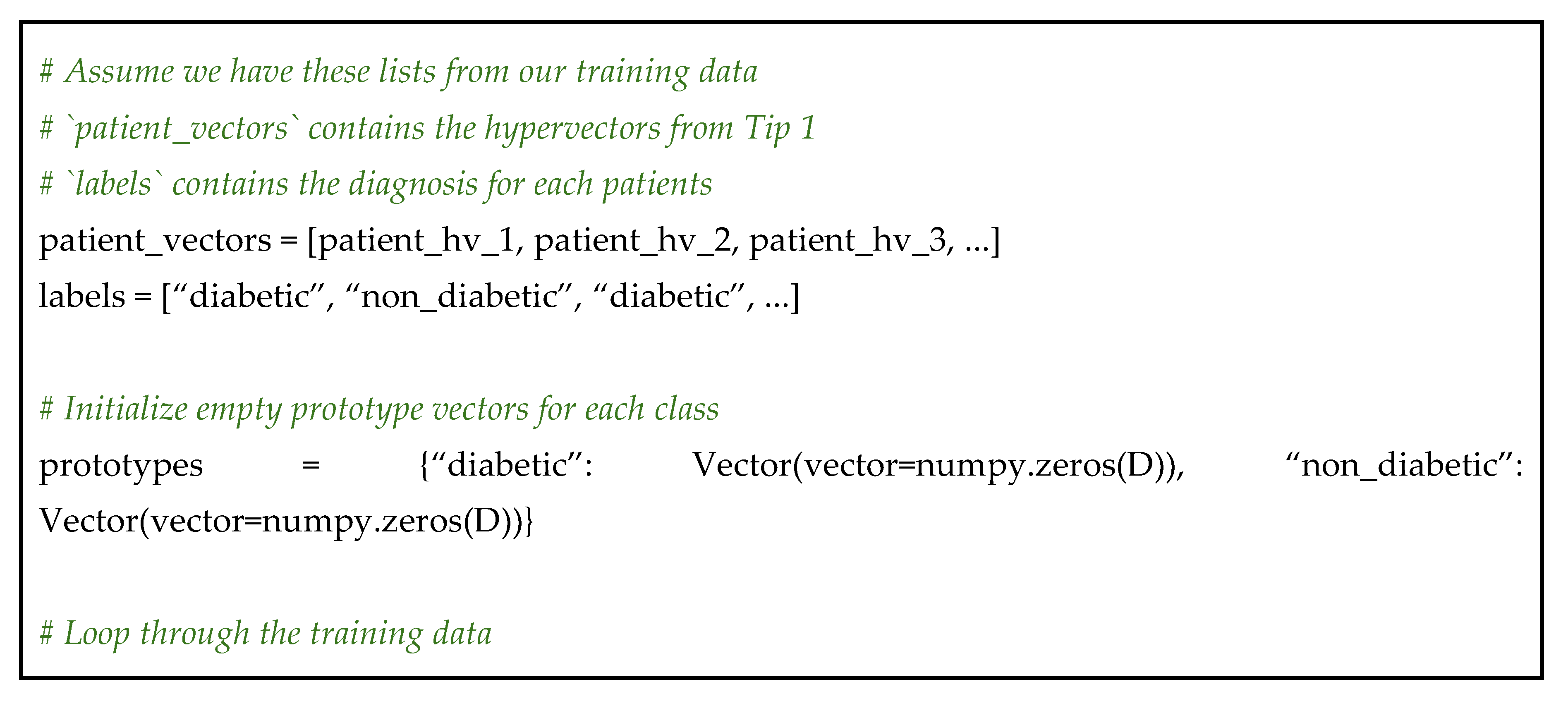


Why This Works
- Simplicity and speed: the training phase is just one pass of addition. The prediction phase is just a few similarity calculations. This is orders of magnitude faster than training a deep neural network;
- Excellent for few-shot learning: this method creates a reasonable prototype even with just a few samples per class;
- Incremental learning/unlearning: if you get new labeled data, you do not have to retrain your model from scratch. You can simply update the existing prototype sums with the new vectors, allowing your model to learn continuously. Conversely, if data needs to be removed, the update is just a subtraction from the prototype.
Tip 8: For Data Fusion – Map All Modalities into a Common Vector Space
A Universal Language
- Encode each modality separately into its own hypervector using the most appropriate technique (e.g., methods from Tip 1 for text/EHRs, Tip 2 for genetic sequences, Tip 3 for images);
- Bundle the resulting vectors together to create a single, multimodal hypervector that represents the complete picture.


Why This Works
Tip 9: For Interpreting Results – Probe Composite Vectors with Clean Pointers
Asking Your Vector Questions

Tip 10: For Reproducibility and Impact – Practice Open Science
- Ensure others can easily discover your work: assign a Digital Object Identifier (DOI) to your code and data by using a repository like Zenodo or Figshare. This makes them easily citable. Also, use rich metadata and keywords when you upload your assets. Describe what the data contains, the VSA parameters used, and the context of the study;
- Make your research available to everyone: publish your code in a public repository like GitHub or GitLab. Submit your manuscript to a fully Open Access Journal to remove paywalls. The protocol for accessing the data should be open and free;
- Ensure your data and models can be combined with other tools and datasets: use common, standard file formats for your input data instead of proprietary formats. Clearly document your VSA parameters, especially the dimensionality and the vector type. This allows others to integrate your hypervectors with their own VSA-based tools;
- Enable others to effectively build upon your work: provide clear and comprehensive documentation. A README.md file in your code repository should explain what the project does and how to run the analysis, including listing all necessary software libraries and their version. Choose a permissive open-source license (e.g., MIT or Apache 2.0) that explicitly tells others how they are allowed to reuse your code.
Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | Artificial Intelligence |
| CNN | Convolutional Neural Network |
| DOI | Digital Object Identifier |
| ECG | Electrocardiography |
| EEG | Electroencephalography |
| EHR | Electronic Health Record |
| EMG | Electromyography |
| FAIR | Findability, Accessibility, Interoperability, and Reusability |
| HDC | Hyperdimensional Computing |
| MAP | Multiply-Add-Permute |
| MRI | Magnetic Resonance Imaging |
| VSA | Vector-Symbolic Architecture |
References
- Kanerva, P. Hyperdimensional computing: An introduction to computing in distributed representation with high-dimensional random vectors. Cognit Comput. 2009, 1, 139–159. [Google Scholar] [CrossRef]
- Kanerva, P. Hyperdimensional computing: An algebra for computing with vectors. Advances in Semiconductor Technologies. Wiley; 2022. pp. 25–42. [CrossRef]
- Zhang S, Wang R, Zhang JJ, Rahimi A, Jiao X. Assessing robustness of hyperdimensional computing against errors in associative memory : (invited paper). 2021 IEEE 32nd International Conference on Application-specific Systems, Architectures and Processors (ASAP). IEEE; 2021. [CrossRef]
- Zhang S, Juretus K, Jiao X. Exploring hyperdimensional computing robustness against hardware errors. IEEE Trans Comput. 2025, 74, 1963–1977. [Google Scholar] [CrossRef]
- Burrello A, Schindler K, Benini L, Rahimi A. Hyperdimensional Computing With Local Binary Patterns: One-Shot Learning of Seizure Onset and Identification of Ictogenic Brain Regions Using Short-Time iEEG Recordings. IEEE Trans Biomed Eng. 2020, 67, 601–613. [Google Scholar] [CrossRef]
- Burrello A, Schindler K, Benini L, Rahimi A. One-shot learning for iEEG seizure detection using end-to-end binary operations: Local binary patterns with hyperdimensional computing. 2018 IEEE Biomedical Circuits and Systems Conference (BioCAS). IEEE; 2018. [CrossRef]
- Rahimi A, Tchouprina A, Kanerva P, Millán J del R, Rabaey JM. Hyperdimensional Computing for Blind and One-Shot Classification of EEG Error-Related Potentials. Mobile Networks and Applications. 2017, 25, 1958–1969. [Google Scholar] [CrossRef]
- Nair DR, Purushothaman A. Brain Inspired One Shot Learning Method for HD Computing. VLSI Design and Test. 2019, 286–297. [Google Scholar] [CrossRef]
- Cumbo F, Chicco D. Hyperdimensional computing in biomedical sciences: a brief review. PeerJ Comput Sci. 2025, 11, e2885. [Google Scholar] [CrossRef]
- Stock M, Van Criekinge W, Boeckaerts D, Taelman S, Van Haeverbeke M, Dewulf P, et al. Hyperdimensional computing: A fast, robust, and interpretable paradigm for biological data. PLOS Computational Biology. 2024, 20, e1012426. [Google Scholar] [CrossRef]
- Cumbo F, Weitschek E, Blankenberg D. hdlib: A Python library for designing Vector-Symbolic Architectures. J Open Source Softw. 2023, 8, 5704. [Google Scholar] [CrossRef]
- Sæthre E, Osborg Ose S, Krokstad S, Østgård Gismervik S. “Terrible Stuff. We’ve been had“: hospital staff reactions to a new electronic health record and implications for employee well-being – A qualitative study. International Journal of Medical Informatics. 2025, 204, 106039. [Google Scholar] [CrossRef]
- Tavabi N, Singh M, Pruneski J, Kiapour AM. Systematic evaluation of common natural language processing techniques to codify clinical notes. PLoS One. 2024, 19, e0298892. [Google Scholar] [CrossRef]
- Sheikhalishahi S, Miotto R, Dudley JT, Lavelli A, Rinaldi F, Osmani V. Natural Language Processing of Clinical Notes on Chronic Diseases: Systematic Review. JMIR Med Inform. 2019, 7, e12239. [Google Scholar] [CrossRef]
- Kuo T-T, Rao P, Maehara C, Doan S, Chaparro JD, Day ME, et al. Ensembles of NLP Tools for Data Element Extraction from Clinical Notes. AMIA Annu Symp Proc. 2016, 2016, 1880–1889. [Google Scholar]
- Wells BJ, Chagin KM, Nowacki AS, Kattan MW. Strategies for handling missing data in electronic health record derived data. EGEMS (Wash DC). 2013, 1, 1035. [Google Scholar] [CrossRef]
- Jones JA, Farnell B. Missing and Incomplete Data Reduces the Value of General Practice Electronic Medical Records as Data Sources in Research. Aust J Prim Health. 2007, 13, 74–80. [Google Scholar] [CrossRef]
- Joel LO, Doorsamy W, Paul BS. A Review of Missing Data Handling Techniques for Machine Learning. IJITIS. 2022, 5, 971–1005. [Google Scholar] [CrossRef]
- Emmanuel T, Maupong T, Mpoeleng D, Semong T, Mphago B, Tabona O. A survey on missing data in machine learning. Journal of Big Data. 2021, 8, 1–37. [Google Scholar] [CrossRef]
- Rizvi STH, Latif MY, Amin MS, Telmoudi AJ, Shah NA. Analysis of Machine Learning Based Imputation of Missing Data. Cybernetics and Systems. 2023, 818–832. [Google Scholar] [CrossRef]
- Bai T, Chanda AK, Egleston BL, Vucetic S. EHR phenotyping via jointly embedding medical concepts and words into a unified vector space. BMC Medical Informatics and Decision Making. 2018, 18, 15–25. [Google Scholar] [CrossRef]
- Chang E-J, Rahimi A, Benini L, Wu A-YA. Hyperdimensional computing-based multimodality emotion recognition with physiological signals. 2019 IEEE International Conference on Artificial Intelligence Circuits and Systems (AICAS). IEEE; 2019. [CrossRef]
- Zhao Q, Yu X, Hu S, Rosing T. MultimodalHD: Federated learning over heterogeneous sensor modalities using hyperdimensional computing. 2024 Design, Automation & Test in Europe Conference & Exhibition (DATE). IEEE; 2024. pp. 1–6. [CrossRef]
- Fanizzi N, d’Amato C. The blessing of dimensionality. Neurosymbolic Artificial Intelligence. 2025. [Google Scholar] [CrossRef]
- Imani M, Nassar T, Rahimi A, Rosing T. HDNA: Energy-efficient DNA sequencing using hyperdimensional computing. 2018 IEEE EMBS International Conference on Biomedical & Health Informatics (BHI). IEEE; 2018. [CrossRef]
- Zou Z, Chen H, Poduval P, Kim Y, Imani M, Sadredini E, et al. BioHD. Proceedings of the 49th Annual International Symposium on Computer Architecture. New York, NY, USA: ACM; 2022. [CrossRef]
- Xu W, Hsu P-K, Moshiri N, Yu S, Rosing T. HyperGen: compact and efficient genome sketching using hyperdimensional vectors. Bioinformatics. 2024, 40, btae452. [Google Scholar] [CrossRef]
- Salehi AW, Khan S, Gupta G, Alabduallah BI, Almjally A, Alsolai H, et al. A Study of CNN and Transfer Learning in Medical Imaging: Advantages, Challenges, Future Scope. Sustainability. 2023, 15, 5930. [Google Scholar] [CrossRef]
- Yao W, Bai J, Liao W, Chen Y, Liu M, Xie Y. From CNN to Transformer: A Review of Medical Image Segmentation Models. Journal of Imaging Informatics in Medicine. 2024, 37, 1529–1547. [Google Scholar] [CrossRef]
- Li Q, Cai W, Wang X, Zhou Y, Feng DD, Chen M. Medical image classification with convolutional neural network. 2014 13th International Conference on Control Automation Robotics & Vision (ICARCV). IEEE; 2014. [CrossRef]
- Xie Y, Zhang J, Shen C, Xia Y. CoTr: Efficiently Bridging CNN and Transformer for 3D Medical Image Segmentation. Medical Image Computing and Computer Assisted Intervention – MICCAI 2021. 2021; 171–180. [CrossRef]
- Yuan F, Zhang Z, Fang Z. An effective CNN and Transformer complementary network for medical image segmentation. Pattern Recognition. 2023, 136, 109228. [Google Scholar] [CrossRef]
- Hassan E, Halawani Y, Mohammad B, Saleh H. Hyper-dimensional computing challenges and opportunities for AI applications. IEEE Access. 2022, 10, 97651–97664. [Google Scholar] [CrossRef]
- Luczak P, Slot K, Kucharski J. Combining deep convolutional feature extraction with hyperdimensional computing for visual object recognition. 2022 International Joint Conference on Neural Networks (IJCNN). IEEE; 2022. pp. 1–8. [CrossRef]
- Lee H, Kim J, Chen H, Zeira A, Srinivasa N, Imani M, et al. Comprehensive integration of hyperdimensional computing with deep learning towards neuro-symbolic AI. 2023 60th ACM/IEEE Design Automation Conference (DAC). IEEE; 2023. pp. 1–6. [CrossRef]
- Alotaiby TN, Alshebeili SA, Alshawi T, Ahmad I, Abd El-Samie FE. EEG seizure detection and prediction algorithms: a survey. EURASIP Journal on Advances in Signal Processing. 2014, 2014, 1–21. [Google Scholar] [CrossRef]
- Ein Shoka AA, Dessouky MM, El-Sayed A, Hemdan EE-D. EEG seizure detection: concepts, techniques, challenges, and future trends. Multimedia Tools and Applications. 2023, 82, 42021–42051. [Google Scholar] [CrossRef] [PubMed]
- Rizal A, Priharti W, Hadiyoso S. Seizure detection in epileptic EEG using Short-Time Fourier Transform and support vector machine. Int J Onl Eng. 2021, 17, 65–78. [Google Scholar] [CrossRef]
- Pale U, Teijeiro T, Atienza D. Systematic Assessment of Hyperdimensional Computing for Epileptic Seizure Detection. Annu Int Conf IEEE Eng Med Biol Soc. 2021, 2021, 6361–6367. [Google Scholar] [CrossRef]
- Ge L, Parhi KK. Applicability of hyperdimensional computing to seizure detection. IEEE Open J Circuits Syst. 2022, 3, 59–71. [Google Scholar] [CrossRef]
- Schindler KA, Rahimi A. A Primer on Hyperdimensional Computing for iEEG Seizure Detection. Front Neurol. 2021, 12, 701791. [Google Scholar] [CrossRef]
- Asgarinejad F, Thomas A, Rosing T. Detection of Epileptic Seizures from Surface EEG Using Hyperdimensional Computing. Annu Int Conf IEEE Eng Med Biol Soc. 2020, 2020, 536–540. [Google Scholar] [CrossRef]
- Du Y, Ren Y, Wong N, Ngai ECH. Hyperdimensional computing with multiscale local binary patterns for scalp EEG-based epileptic seizure detection. IEEE Internet Things J. 2024, 11, 26046–26061. [Google Scholar] [CrossRef]
- Virtanen P, Gommers R, Oliphant TE, Haberland M, Reddy T, Cournapeau D, et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat Methods. 2020, 17, 261–272. [Google Scholar] [CrossRef]
- Gramfort A, Luessi M, Larson E, Engemann DA, Strohmeier D, Brodbeck C, et al. MEG and EEG data analysis with MNE-Python. Front Neurosci. 2013, 7, 267. [Google Scholar] [CrossRef]
- Ma D, Thapa R, Jiao X. MoleHD: Efficient drug discovery using brain inspired hyperdimensional computing. 2022 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE; 2022. [CrossRef]
- Jones D, Zhang X, Bennion BJ, Pinge S, Xu W, Kang J, et al. HDBind: encoding of molecular structure with hyperdimensional binary representations. Scientific Reports. 2024, 14, 1–16. [Google Scholar] [CrossRef]
- Cumbo F, Dhillon K, Joshi J, Raubenolt B, Chicco D, Aygun S, et al. Predicting the toxicity of chemical compounds via Hyperdimensional Computing. bioRxiv. 2025. [Google Scholar] [CrossRef]
- Nicholson DN, Greene CS. Constructing knowledge graphs and their biomedical applications. Computational and Structural Biotechnology Journal. 2020, 18, 1414–1428. [Google Scholar] [CrossRef] [PubMed]
- Bonner S, Barrett IP, Ye C, Swiers R, Engkvist O, Bender A, et al. A review of biomedical datasets relating to drug discovery: a knowledge graph perspective. Brief Bioinform. 2022, 23. [Google Scholar] [CrossRef]
- Harnoune A, Rhanoui M, Mikram M, Yousfi S, Elkaimbillah Z, El Asri B. BERT based clinical knowledge extraction for biomedical knowledge graph construction and analysis. Computer Methods and Programs in Biomedicine Update. 2021, 1, 100042. [Google Scholar] [CrossRef]
- Bugnon LA, Yones C, Milone DH, Stegmayer G. Deep Neural Architectures for Highly Imbalanced Data in Bioinformatics. IEEE Trans Neural Netw Learn Syst. 2020, 31, 2857–2867. [Google Scholar] [CrossRef]
- Pes, B. Learning from high-dimensional biomedical datasets: The issue of class imbalance. IEEE Access. 2020, 8, 13527–13540. [Google Scholar] [CrossRef]
- Cumbo F, Cappelli E, Weitschek E. A Brain-Inspired Hyperdimensional Computing Approach for Classifying Massive DNA Methylation Data of Cancer. Algorithms. 2020, 13, 233. [Google Scholar] [CrossRef]
- Cumbo F, Truglia S, Weitschek E, Blankenberg D. Feature selection with vector-symbolic architectures: a case study on microbial profiles of shotgun metagenomic samples of colorectal cancer. Brief Bioinform. 2025, 26, bbaf177. [Google Scholar] [CrossRef]
- Cumbo F, Dhillon K, Joshi J, Chicco D, Aygun S, Blankenberg D. A novel Vector-Symbolic Architecture for graph encoding and its application to viral pangenome-based species classification. bioRxiv. 2025. [CrossRef]
- Chicco D, Cumbo F, Angione C. Ten quick tips for avoiding pitfalls in multi-omics data integration analyses. PLoS Comput Biol. 2023, 19, e1011224. [Google Scholar] [CrossRef]
- Barkam HE, Yun S, Chen H, Gensler P, Mema A, Ding A, et al. Reliable hyperdimensional reasoning on unreliable emerging technologies. 2023 IEEE/ACM International Conference on Computer Aided Design (ICCAD). IEEE; 2023. [CrossRef]
- Mitrokhin A, Sutor P, Summers-Stay D, Fermüller C, Aloimonos Y. Symbolic Representation and Learning With Hyperdimensional Computing. Front Robot AI. 2020, 7, 535245. [Google Scholar] [CrossRef]
- Chang C-Y, Chuang Y-C, Huang C-T, Wu A-Y. Recent progress and development of hyperdimensional computing (HDC) for edge intelligence. IEEE J Emerg Sel Top Circuits Syst. 2023, 13, 119–136. [Google Scholar] [CrossRef]
- Amrouch H, Imani M, Jiao X, Aloimonos Y, Fermuller C, Yuan D, et al. Brain-inspired hyperdimensional computing for ultra-efficient edge AI. 2022 International Conference on Hardware/Software Codesign and System Synthesis (CODES+ISSS). IEEE; 2022. [CrossRef]
- Sim, I. Data Sharing and Reuse. Principles and Practice of Clinical Trials. 2022, 2137–2158. [Google Scholar] [CrossRef]
- Wilkinson MD, Dumontier M, Aalbersberg IJ, Appleton G, Axton M, Baak A, et al. The FAIR Guiding Principles for scientific data management and stewardship. Scientific Data. 2016, 3, 1–9. [Google Scholar] [CrossRef]
- Heddes M, Nunes I, Vergés P, Kleyko D, Abraham D, Givargis T, et al. Torchhd: An Open Source Python Library to Support Research on Hyperdimensional Computing and Vector Symbolic Architectures. Journal of Machine Learning Research. 2023, 24, 1–10. [Google Scholar]
- Wang J, Huang S, Imani M. DistHD: A learner-aware dynamic encoding method for hyperdimensional classification. 2023 60th ACM/IEEE Design Automation Conference (DAC). IEEE; 2023. pp. 1–6. [CrossRef]
- Kang J, Khaleghi B, Rosing T, Kim Y. OpenHD: A GPU-Powered Framework for Hyperdimensional Computing. IEEE Trans Comput. 2022, 71, 2753–2765. [Google Scholar] [CrossRef]
- Simon WA, Pale U, Teijeiro T, Atienza D. HDTorch: Accelerating Hyperdimensional Computing with GP-GPUs for Design Space Exploration. ICCAD ’22: Proceedings of the 41st IEEE/ACM International Conference on Computer-Aided Design. [CrossRef]
- Aygun S, Moghadam MS, Najafi MH. UHD: Unary processing for lightweight and dynamic hyperdimensional computing. 2024 Design, Automation & Test in Europe Conference & Exhibition (DATE). IEEE; 2024. pp. 1–6. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



