Submitted:
27 September 2025
Posted:
29 September 2025
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Related Work
2.1. Deep Learning for Breast Ultrasound Image Segmentation
2.2. Lightweight Architectures for Medical Image Segmentation
2.3. Attention Mechanisms in Medical Image Analysis
2.4. Boundary-Aware and Robust Segmentation
3. Methodology
3.1. Overall Architecture
3.2. Lightweight Design of LBA-Net
3.3. Lightweight Encoder
3.4. Lightweight Attention Block (LBA-Block) Design
3.5. Dual-Head Supervision
3.6. Loss Function
3.6.1. Segmentation Loss:
3.6.2. Boundary Loss:
3.7. Training Strategy
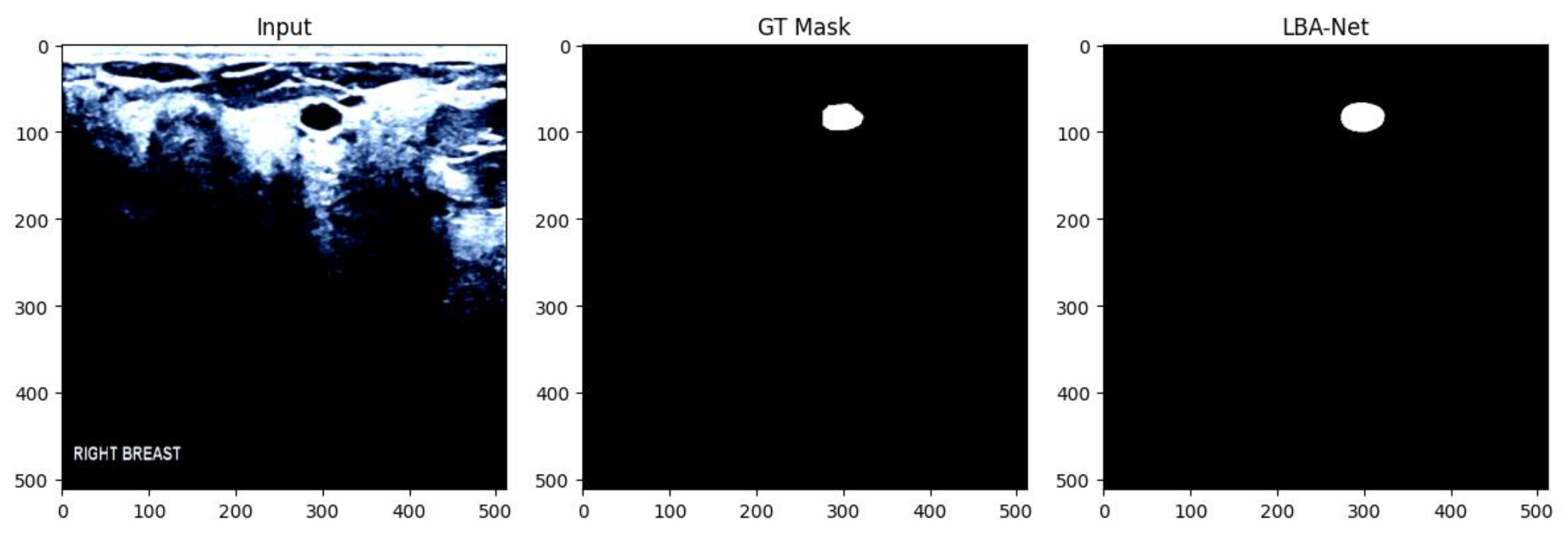
4. Experiments
4.1. The Impact of Input Resolution on Performance
| Input size | Validation Dice(%) | IoU (%) | HD95 (mm) | FPS (GPU) |
|---|---|---|---|---|
| 512×512 | 81.4 | 68.3 | 6.1 | 122 |
| 256×256 | 71.8 | 56.0 | 7.4 | 213 |
4.2. Datasets and Evaluation Metrics
4.3. Implementation Details
4.4. Comparison with State-of-the-Art
4.5. Ablation Studies
- (1)
- V1 (w/o LBA-Block): all local-boundary attention blocks in the decoder were replaced with 1×1 convolutions.
- (2)
- V2 (w/o ASPP): the ASPP bottleneck was removed and substituted by a 1×1 convolution, discarding multi-scale context.
- (3)
- V3 (w/o Boundary Head): only the segmentation head was kept; auxiliary boundary supervision was canceled.
- (4)
- V4 (LBA→CBAM): the dual-head architecture was preserved, but the proposed LBA-Block was replaced by CBAM.
| Model Variant | Val Dice (%)↑ | Val IoU (%) ↑ | Params (M) ↓ | FLOPs (G) ↓ |
|---|---|---|---|---|
| Baseline (Full LBA-Net) | 81.42 | 68.62 | 2.14 | 4.83 |
| V1 (w/o LBA-Block) | 78.91 | 65.21 | 2.03 | 4.75 |
| V2 (w/o ASPP) | 79.53 | 66.88 | 1.98 | 4.62 |
| V3 (w/o Boundary Head) | 80.10 | 66.25 | 2.08 | 4.79 |
| V4 (LBA→CBAM) | 77.65 | 63.95 | 2.15 | 4.86 |
5. Discussion
5.1. Advantages of LBA-Net
5.2. Clinical Implications and Future Work
5.3. Summary
6. Conclusion
Author Contributions
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bizuayehu, H.M.; Ahmed, K.Y.; Kibret, G.D.; Dadi, A.F.; Belachew, S.A.; Bagade, T.; Tegegne, T.K.; Venchiarutti, R.L.; Kibret, K.T.; Hailegebireal, A.H.; Assefa, Y.; Khan, M.N.; Abajobir, A.; Alene, K.A.; Mengesha, Z.; Erku, D.; Enquobahrie, D.A.; Minas, T.Z.; Misgan, E.; Ross, A.G. Global Disparities of Cancer and Its Projected Burden in 2050. JAMA Netw Open. 2024, 7, e2443198. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chen, F.; Chen, Y.J.; Hu, Y.Z. Multimodal Ultrasound Imaging in the Diagnosis of Primary Giant Cell Tumor of the Breast: A Case Report and Literature Review. J Clin Ultrasound. 2025, 53, 885–892. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Liu, S.; Cao, Z.; Zhang, J.; Pu, X.; Yu, J. Accurate classification of benign and malignant breast tumors in ultrasound imaging with an enhanced deep learning model. Front Bioeng Biotechnol. 2025, 13, 1526260. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yu, L.; Gou, B.; Xia, X.; Yang, Y.; Yi, Z.; Min, X.; He, T. BUS-M2AE: Multi-scale Masked Autoencoder for Breast Ultrasound Image Analysis. Comput Biol Med. 2025, 191, 110159. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, M.R.; Torres, H.R.; Oliveira, B.; de Araujo, A.R.V.F.; Morais, P.; Novais, P.; Vilaca, J.L. Deep Learning Networks for Breast Lesion Classification in Ultrasound Images: A Comparative Study. Annu Int Conf IEEE Eng Med Biol Soc. 2023, 2023, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Guo, Z.; Xie, J.; Wan, Y.; Zhang, M.; Qiao, L.; Yu, J.; Chen, S.; Li, B.; Yao, Y. A review of the current state of the computer-aided diagnosis (CAD) systems for breast cancer diagnosis. Open Life Sci. 2022, 17, 1600–1611. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Abdelrahman, L.; Al Ghamdi, M.; Collado-Mesa, F.; Abdel-Mottaleb, M. Convolutional neural networks for breast cancer detection in mammography: A survey. Comput Biol Med. 2021, 131, 104248. [Google Scholar] [CrossRef] [PubMed]
- Derakhshandeh, S.; Mahloojifar, A. Modifying the U-Net's Encoder-Decoder Architecture for Segmentation of Tumors in Breast Ultrasound Images. J Imaging Inform Med. 2025. [Google Scholar] [CrossRef] [PubMed]
- [9] Khaled, R.; Vidal, J.; Vilanova, J.C.; Martí, R. A U-Net Ensemble for breast lesion segmentation in DCE MRI. Comput Biol Med. 2022, 140, 105093. [Google Scholar] [CrossRef] [PubMed]
- Pramanik, P.; Roy, A.; Cuevas, E.; Perez-Cisneros, M.; Sarkar, R. DAU-Net: Dual attention-aided U-Net for segmenting tumor in breast ultrasound images. PLoS One. 2024, 19, e0303670. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wu, Y.; Huang, L.; Yang, T. Breast Ultrasound Image Segmentation Using Multi-branch Skip Connection Search. J Imaging Inform Med. 2025. [Google Scholar] [CrossRef] [PubMed]
- Sadeghi-Goughari, M.; Rajabzadeh, H.; Han, J.W.; Kwon, H.J. Artificial intelligence-assisted ultrasound-guided focused ultrasound therapy: a feasibility study. Int J Hyperthermia. 2023, 40, 2260127. [Google Scholar] [CrossRef] [PubMed]
- Anandha Praba, R.; Suganthi, L. Human activity recognition utilizing optimized attention induced Multihead Convolutional Neural Network with Mobile Net V1 from Mobile health data. Network. 2025, 36, 294–321. [Google Scholar] [CrossRef] [PubMed]
- Kanchana, K.; et al. "Enhancing skin cancer classification using efficient net b0-b7 through convolutional neural networks and transfer learning with patient-specific data. " Asian Pacific Journal of Cancer Prevention: APJCP 2024, 25, 1795. [Google Scholar]
- Sara Koshy, S.; Anbarasi, L.J. HMA-Net: a hybrid mixer framework with multihead attention for breast ultrasound image segmentation. Front Artif Intell. 2025, 8, 1572433. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Lou, Q.; Li, Y.; Qian, Y.; Lu, F.; Ma, J. Mammogram classification based on a novel convolutional neural network with efficient channel attention. Comput Biol Med. 2022, 150, 106082. [Google Scholar] [CrossRef] [PubMed]
- Nissar, I.; Alam, S.; Masood, S.; Kashif, M. MOB-CBAM: A dual-channel attention-based deep learning generalizable model for breast cancer molecular subtypes prediction using mammograms. Comput Methods Programs Biomed. 2024, 248, 108121. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Q.; Wang, Q.; Bao, Y.; et al. LAEDNet: a lightweight attention encoder–decoder network for ultrasound medical image segmentation. Computers and Electrical Engineering, 2022, 99, 107777. [Google Scholar] [CrossRef]
- Wang, R.; Chen, S.; Ji, C.; Fan, J.; Li, Y. Boundary-aware context neural network for medical image segmentation. Med Image Anal. 2022, 78, 102395. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Min, W.; Wang, S. Boundary-Aware Gradient Operator Network for Medical Image Segmentation. IEEE J Biomed Health Inform. 2024, 28, 4711–4723. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Liu, H.; Zeng, Z.; et al. BES-Net: Boundary enhancing semantic context network for high-resolution image semantic segmentation. Remote Sensing, 2022, 14, 1638. [Google Scholar] [CrossRef]
- Xiong, Y.; Shu, X.; Liu, Q.; et al. HCMNet: A Hybrid CNN-Mamba Network for Breast Ultrasound Segmentation for Consumer Assisted Diagnosis. IEEE Transactions on Consumer Electronics 2025. [Google Scholar] [CrossRef]
- Zhou, J.; Kuang, H.; Wang, J. HCM-Net: Hybrid CNN and Mamba Network with Multi-scale Awareness Feature Fusion for Lung Cancer Pathological Complete Response Prediction[C]//International Symposium on Bioinformatics Research and Applications. Singapore: Springer Nature Singapore, 2025, 38-48.
- DeVoe, K.; Takahashi, G.; Tarshizi, E.; Sacker, A. Evaluation of the precision and accuracy in the classification of breast histopathology images using the MobileNetV3 model. J Pathol Inform. 2024, 15, 100377. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Lou, Q.; Li, Y.; Qian, Y.; et al. Mammogram classification based on a novel convolutional neural network with efficient channel attention. Computers in Biology and Medicine, 2022, 150, 106082. [Google Scholar] [CrossRef] [PubMed]
- Sulaiman, A.; Anand, V.; Gupta, S.; et al. Attention based UNet model for breast cancer segmentation using BUSI dataset. Scientific Reports, 2024, 14, 22422. [Google Scholar] [CrossRef]
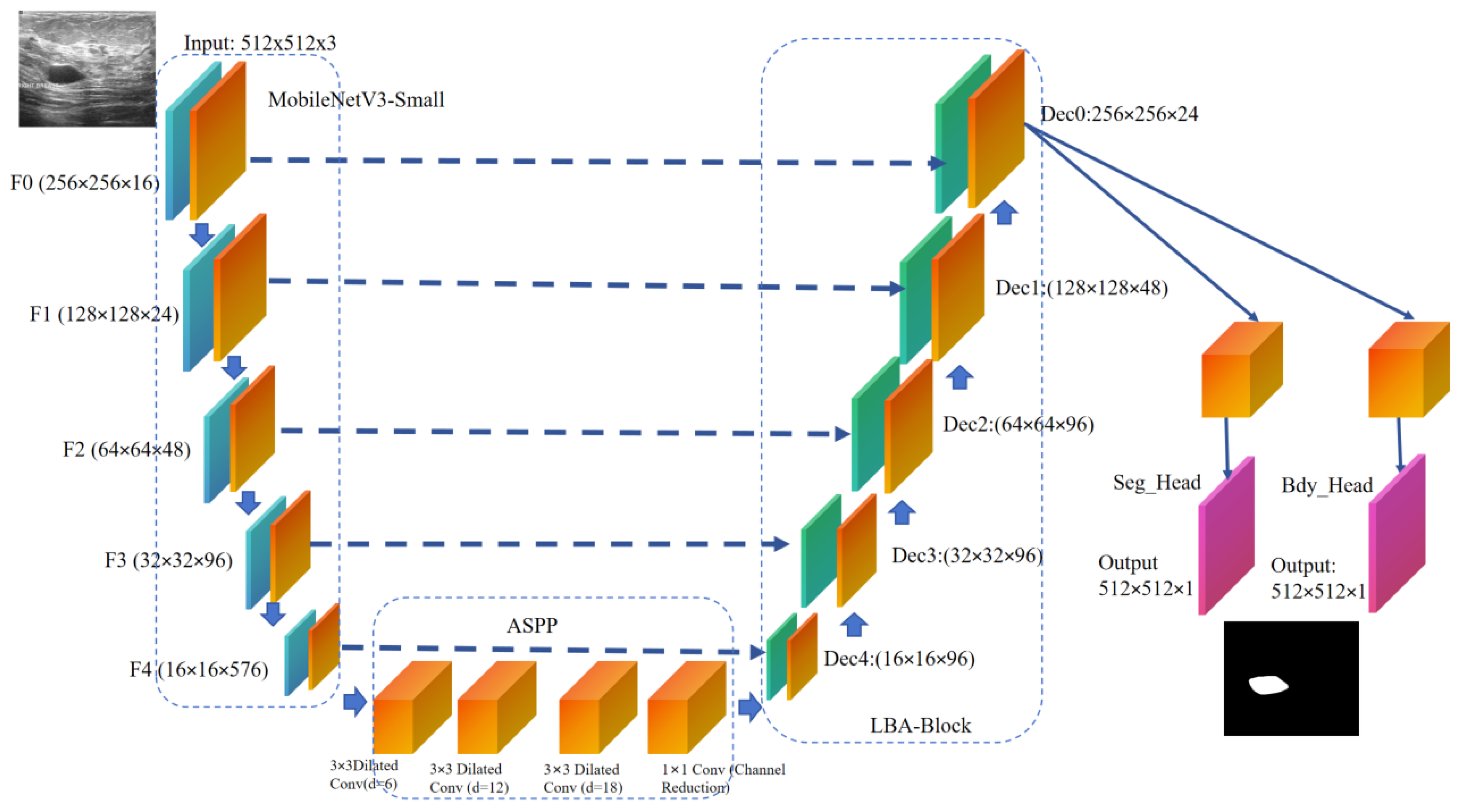
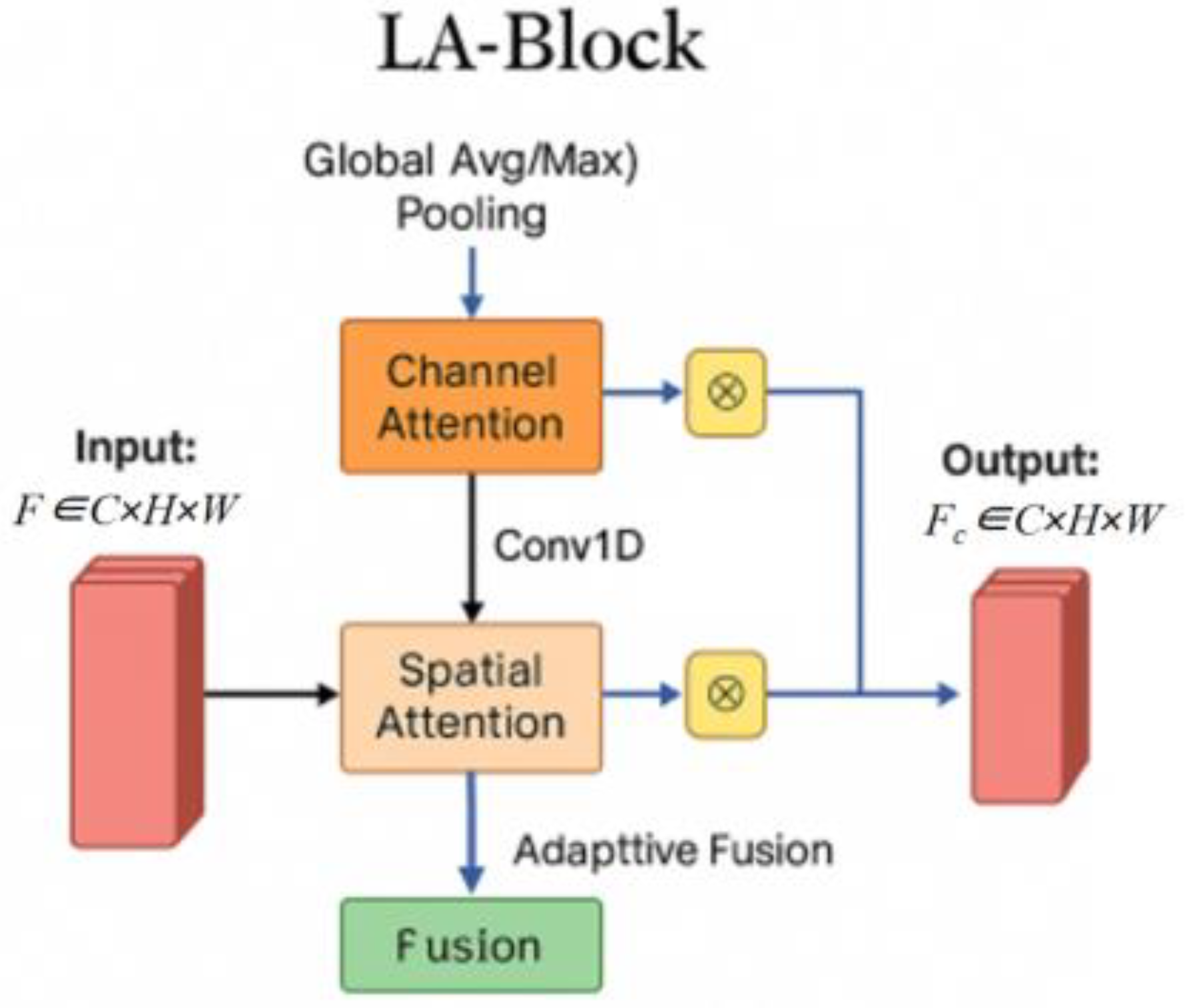
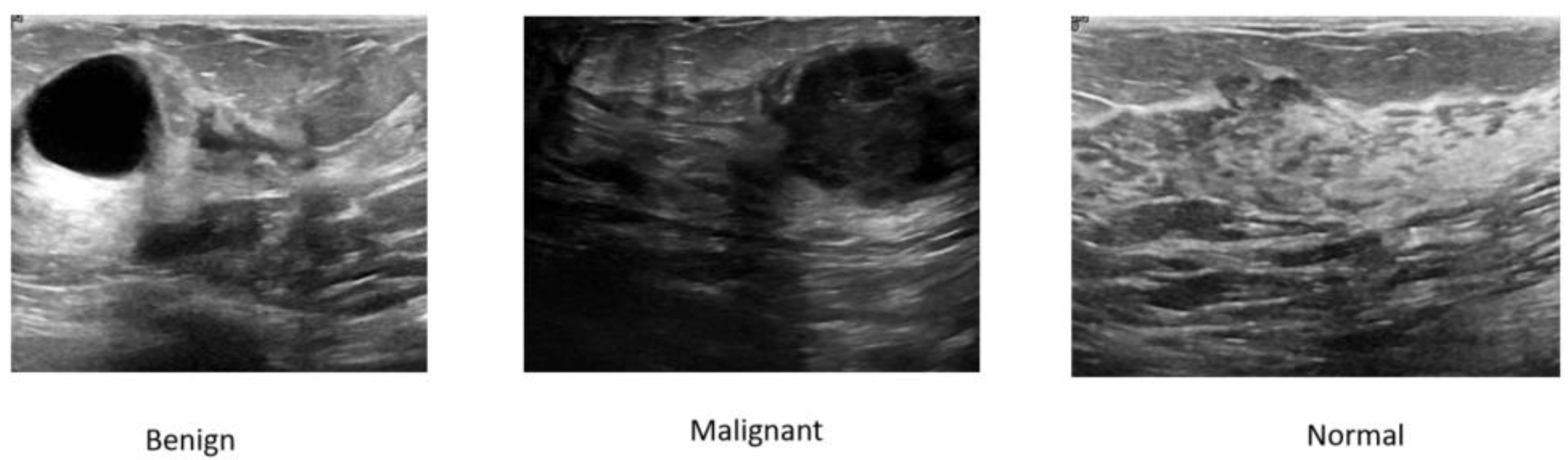
| Model Category | Model Name | Optimal Val Dice (%) | Optimal Val IoU (%) | Params (M) | FLOPs (G) | Minimum Training Loss | Minimum Validation Loss |
|---|---|---|---|---|---|---|---|
| CNN-based | U-Net | 75.66 | 60.10 | 7.83 | 15.63 | 0.2575 | 0.4848 |
| CNN-based | UNet++ | 75.68 | 60.48 | 9.04 | 16.23 | 0.2464 | 0.5065 |
| CNN-based | DeepLabV3+ | 75.53 | 60.68 | 20.56 | 18.72 | 0.2412 | 0.4963 |
| CNN-based | FPN | 76.41 | 61.54 | 1.83 | 8.34 | 0.2548 | 0.4961 |
| Proposed | LBA-Net | 81.42 | 68.62 | 2.14 | 4.86 | 0.2410 | 0.4902 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




