Submitted:
06 September 2025
Posted:
08 September 2025
You are already at the latest version
Abstract
Keywords:
Introduction
- YOLOv11 detection outcomes,
- vegetation indices (NDVI, EVI, NDWI),
- morphological characteristics (leaf perimeter, area),
- texture descriptors (GLCM, LBP).
- Design of a multimodal crop health monitoring system that integrates UAV images, IoT sensor readings, and deep learning analytics.
- Building of YOLOv11 with transformer-based attention mechanisms, which is specifically targeted at detecting small lesions in heavy field conditions in real time.
- Developing the Composite Health Index (CHI), an integrated measure that combines vegetation indices, morphological and texture characteristics and AI predictions.
- Implementation of a hybrid edge–cloud deployment paradigm, enabling real-time on-field usage and cloud-scale processing.
- Past field studies have referenced improvements such as reduced usage of pesticides, improved scouting effectiveness, and less yield loss when AI-IoT-UAV systems were used.
Related Work
- Early Image Processing Methods (2000–2012)
- 2.
- CNN-Based Disease Classification (2012–2017)
- Models were only trained on lab-captured or mobile photos, not UAV field imagery.
- High accuracy was dataset-specific and did not hold up under actual farm variation.
- No publicly available, India-specific datasets hindered generalization.
- While CNN-based systems gave the first proof that AI might revolutionize Indian crop monitoring, they identified the lab-to-field gap.
- 3.
- Real-Time Object Detection (YOLO Family, 2018–2022)

- 4.
- Hybrid CNN–Transformer Architectures (2022–Present)


- 5.
- IoT and UAV-Based Remote Sensing (2018–Present)
- 6.
- Comparative Insights (India vs Global)
- 7.
- Research Gaps (India-Specific)
- Gap in Transferability: All models perform well in constrained sets, but perform poorly in heterogeneous Indian fields where lighting, background clutter, and mixed cropping prevail.
- Gap in Affordability: Capable YOLO and Transformer models need costly GPUs, out of reach for Indian small farmers.
- Gap in Data: Large-scale labeled public crop disease datasets, particularly UAV-based ones, are not available in India.
- Integration Gap: There are very few Indian systems that integrate IoT + UAV + AI into a real-time deployable solution by farmers.
- Policy/Infrastructure Gap: UAV operation in India is restricted under DGCA norms, battery life in hot temperatures, and low rural connectivity.
- 8.
- Comparative Summary of Key Approaches
| Study / Model | Data Type | Methodology | Strengths | Limitations |
|---|---|---|---|---|
| Pooja et al. (2017) [31] | RGB leaf images | Image processing (color/shape) | Simple, low-cost | Poor generalization, noise-sensitive |
| Agarwal et al. (2019) [12] | RGB leaf images | CNN classifier | High accuracy (>95%) | Controlled datasets only |
| Qi et al. (2022) [26] | Tomato leaf images | Improved YOLOv5 + attention | Real-time detection, robust | Needs large annotated datasets |
| Islam et al. (2023) [6] | Web + mobile images | DeepCrop (CNN + Transformer) | Accessible, high precision | Limited multi-crop support |
| Ngugi et al. (2024) [13] | IoT + UAV multispectral | ML/DL fusion models | Early detection, multi-source data | Expensive sensors, rural adoption issues |
| Sharma et al. (2024) [18] | Multi-weed datasets | YOLOv8–YOLOv11 comparison | YOLOv11 best for small lesions | High training cost, limited trials |
Materials and Methods
System Architecture
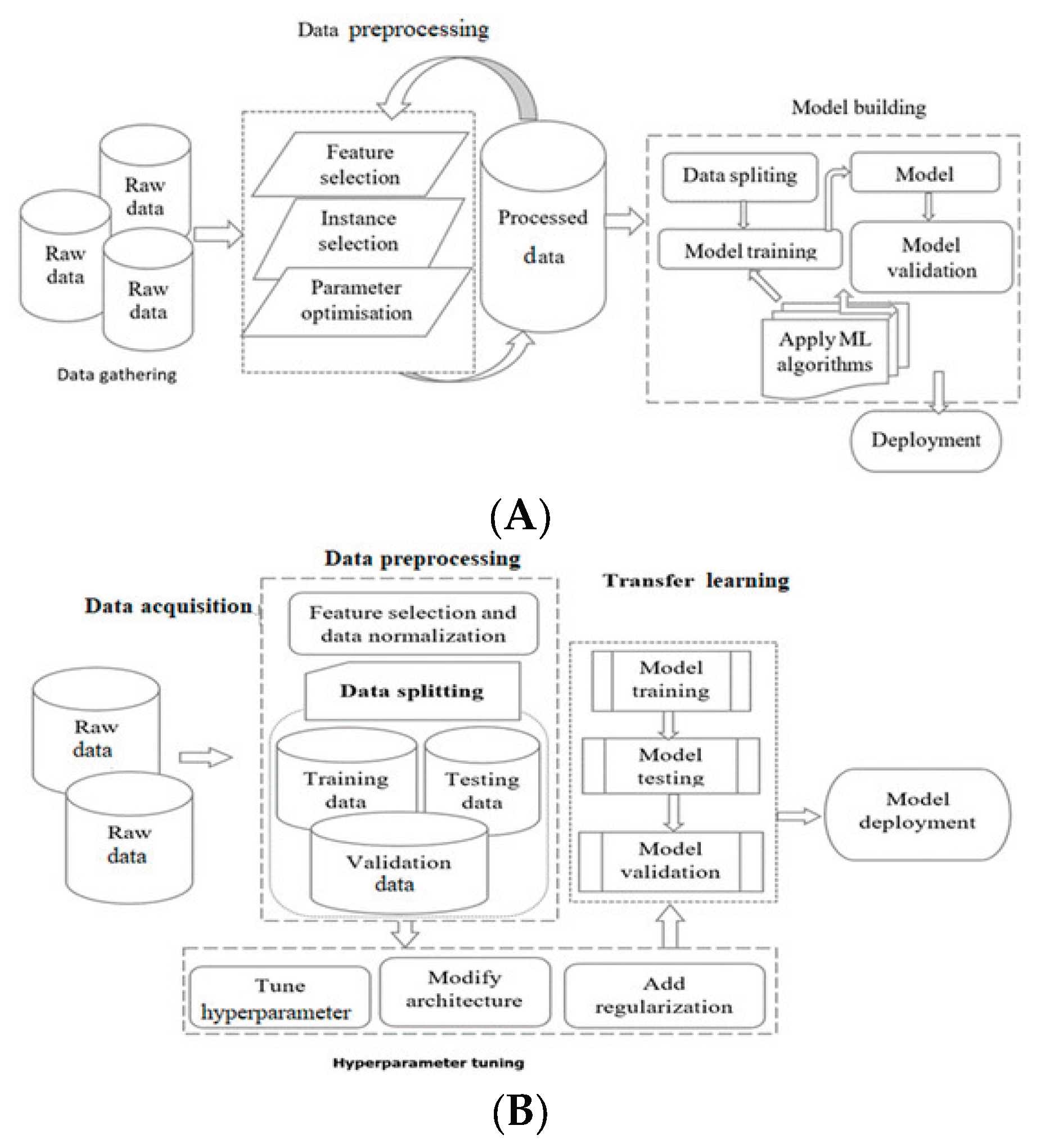
General Process Flow of Machine Learning and Deep Learning Systems
Data Acquisitions
Data Preprocessing
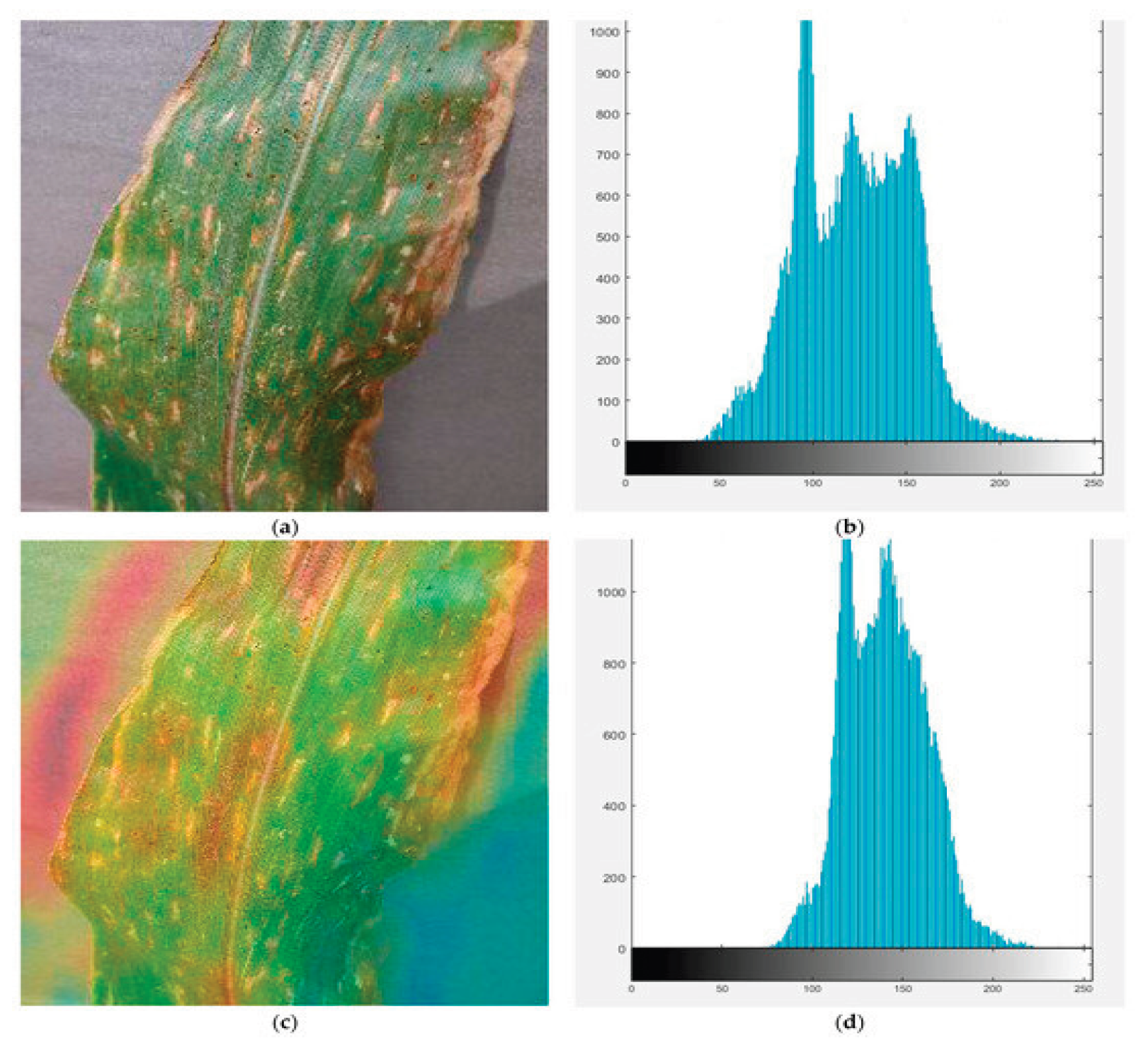
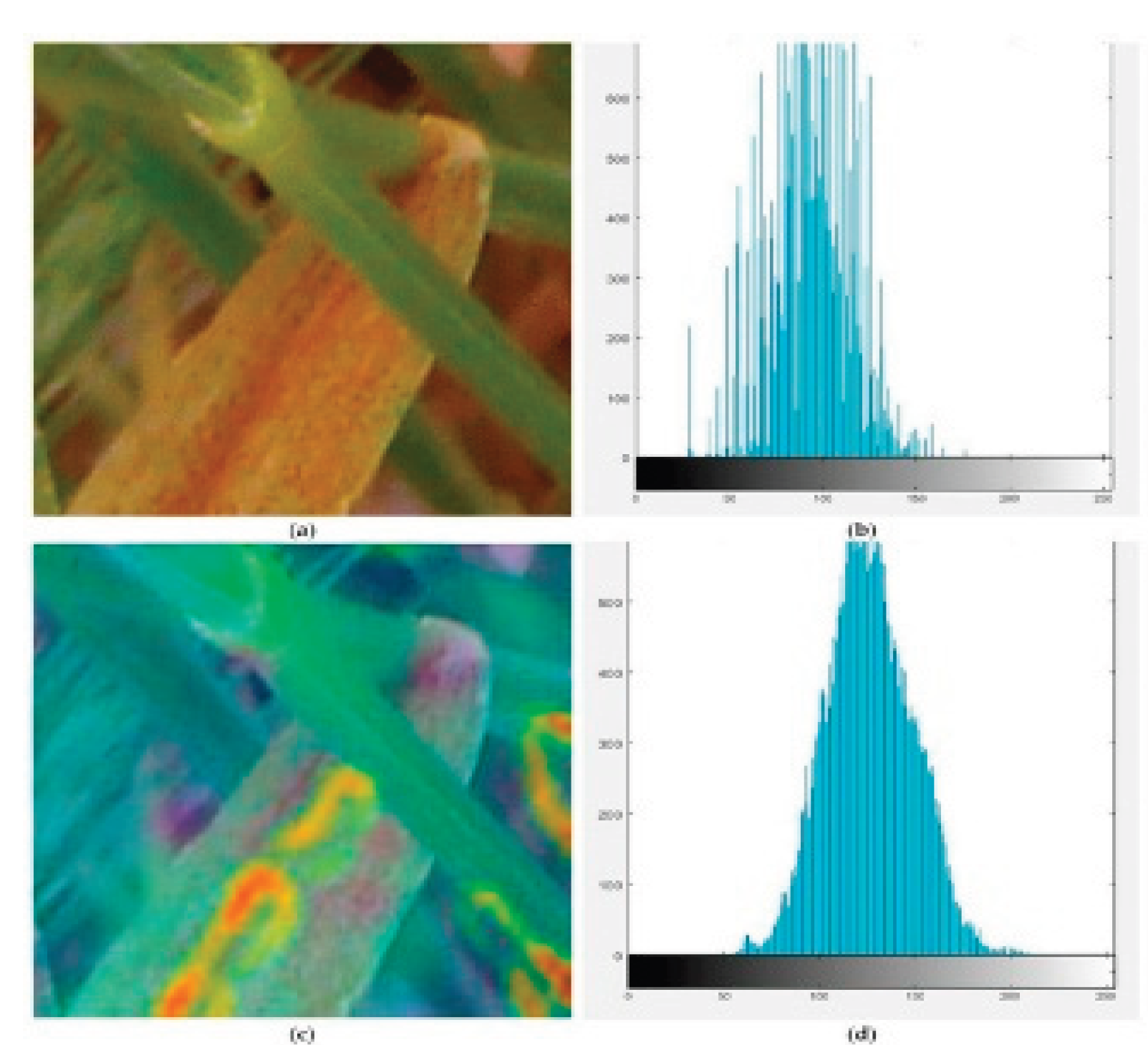
Feature Engineering
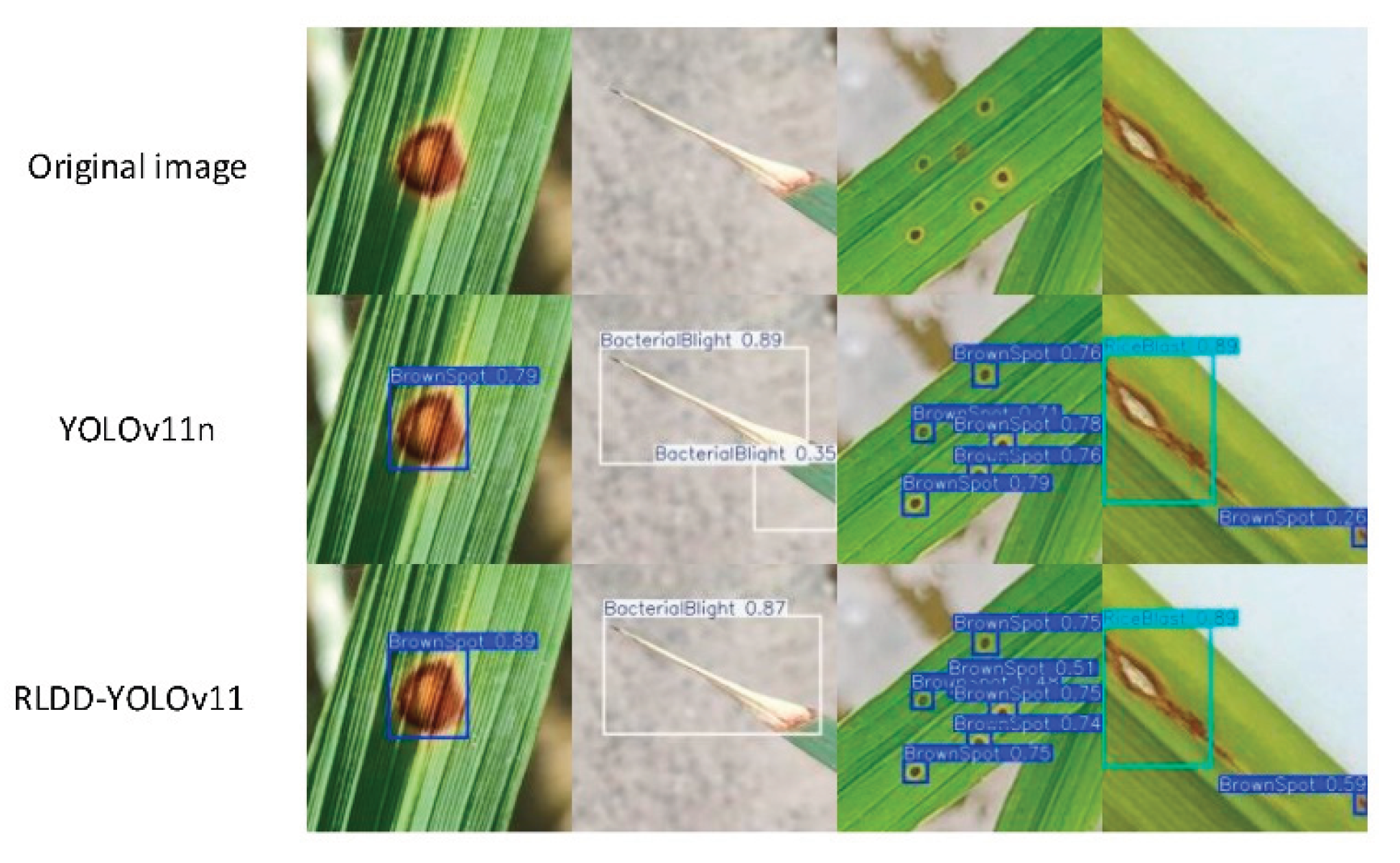
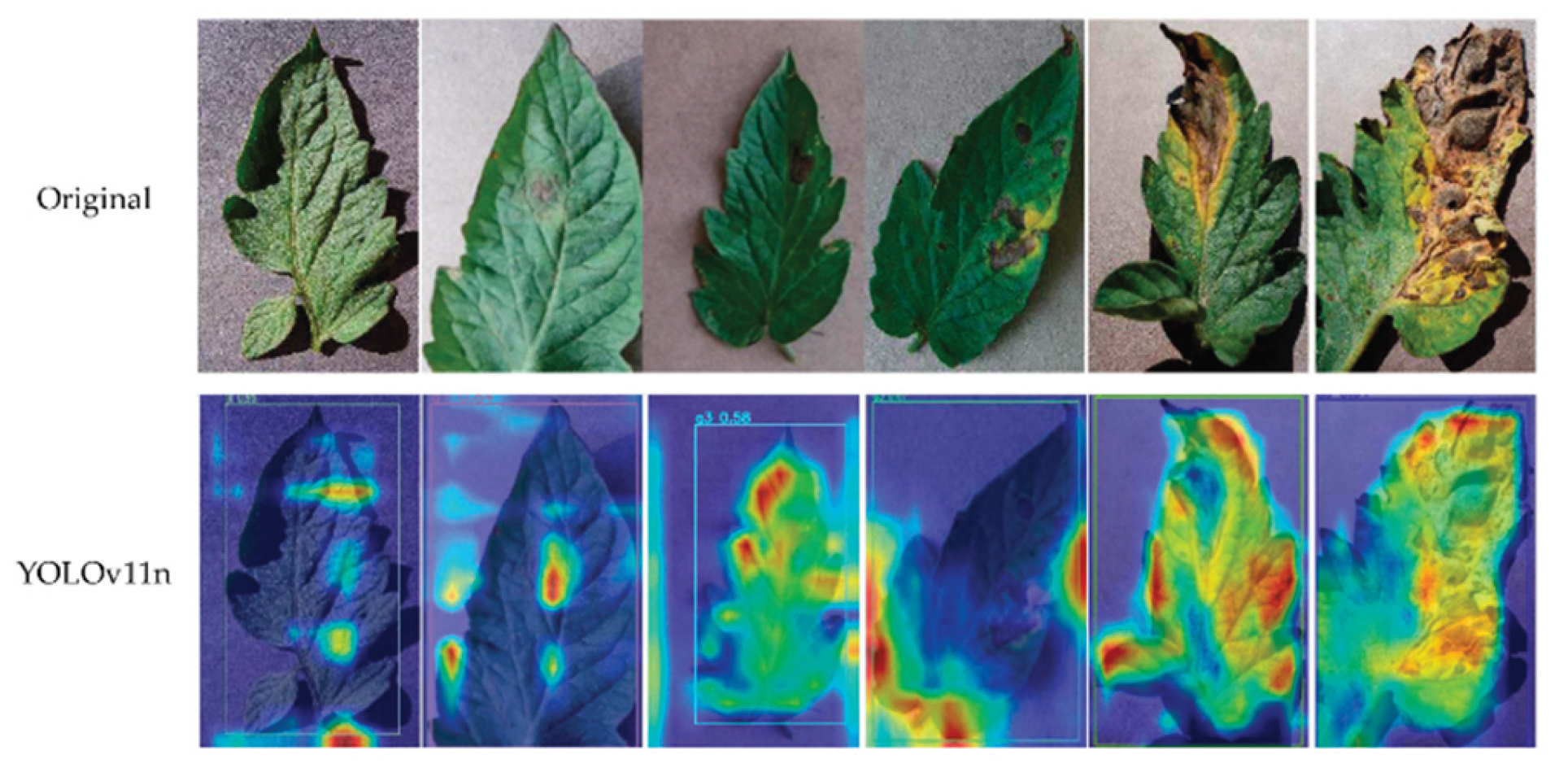
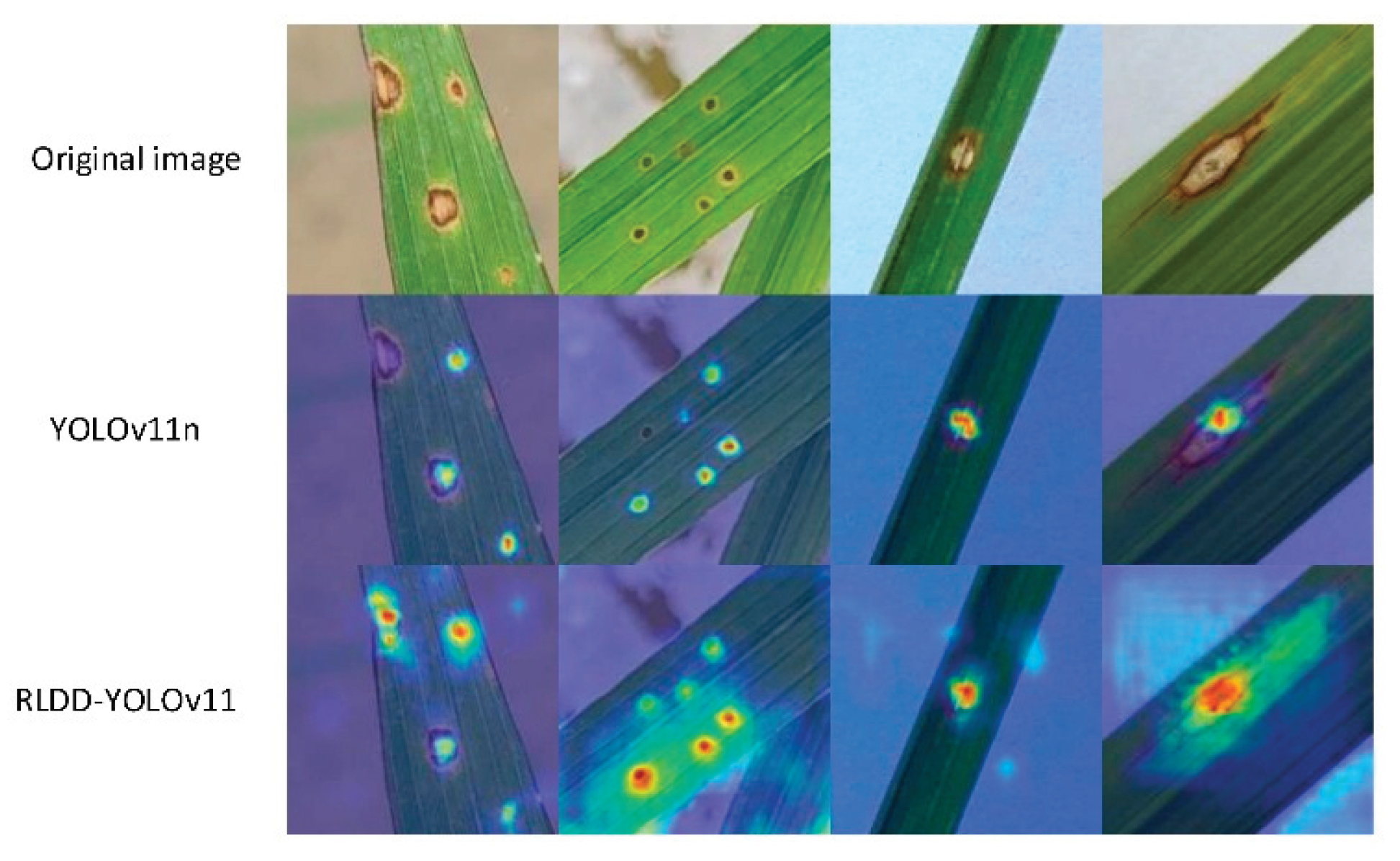
Disease Detection with YOLOv11
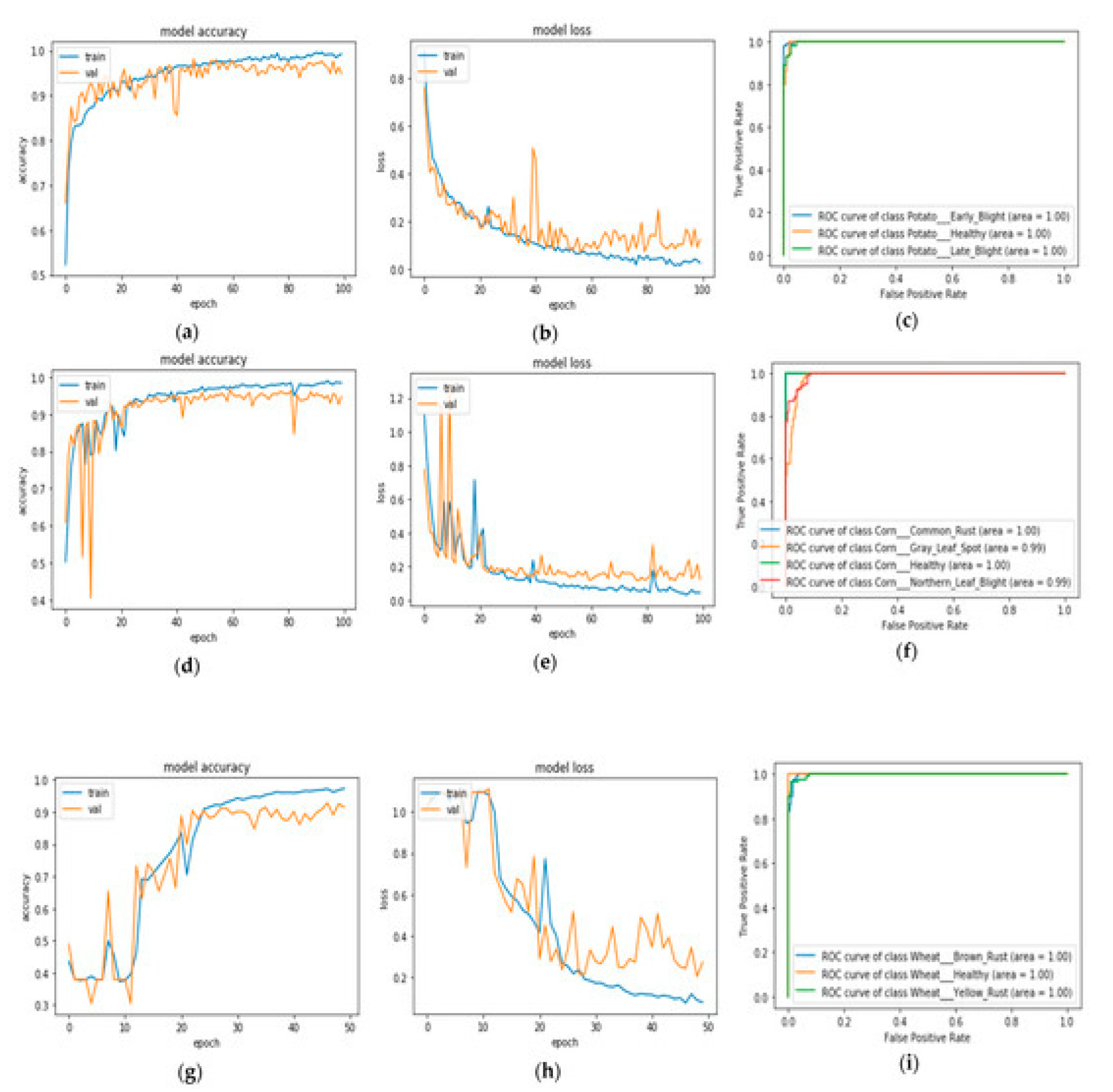
- [31]Accuracy: The proportion of test cases for which a correct prediction was made can be expressed as follows:
- [31] Precision: The ratio of correctly predicted disease-affected leaves to all positively predicted leaves by the model is known as precision and can be defined as follows:
- [31] Recall: recall is defined as the percentage of correctly predicted disease-affected leaves relative to the total positive instances of the test case:
- [31] F1-Score:
- Normalized Difference Vegetation Index (NDVI):
- NIR = Near Infrared band (plants strongly reflect NIR if healthy).
- VIS = Visible band, usually red light (plants absorb red for photosynthesis).
- Measures greenness and photosynthetic activity.
- Values range from -1 to +1:
- +0.6 to +0.9 → healthy, dense vegetation.
- 0 to 0.2 → bare soil/stressed vegetation.
- Negative → water, clouds, non-vegetated surfaces.
- Simple Ratio Index (SR):
- Ratio of reflectance in visible vs. NIR bands.
- Simpler than NDVI, but less normalized (more sensitive to illumination differences).
- Quick indicator of vegetation density.
- High SR → stressed or sparse vegetation.
- Low SR → healthy vegetation (since NIR >> VIS).
- Soil Adjusted Vegetation Index (SAVI):
- L = soil brightness correction factor (usually 0.5).
- Corrects for soil background noise in areas with low vegetation cover.
- Useful in semi-arid regions or early crop growth stages when soil is visible.
- Pasture Index (PI):
- Uses two red bands and NIR.
- Designed to measure pasture biomass and forage quality.
- Detects grassland/pasture conditions more precisely than NDVI.
- Helps in livestock management and grazing capacity estimation.
- Normalised Difference Water Index (NDWI):
- Uses NIR bands at 860 nm and 1240 nm.
- Detects water content in vegetation (leaf water content, canopy moisture).
- Useful for identifying drought stress and irrigation needs.
Confusion Matrix
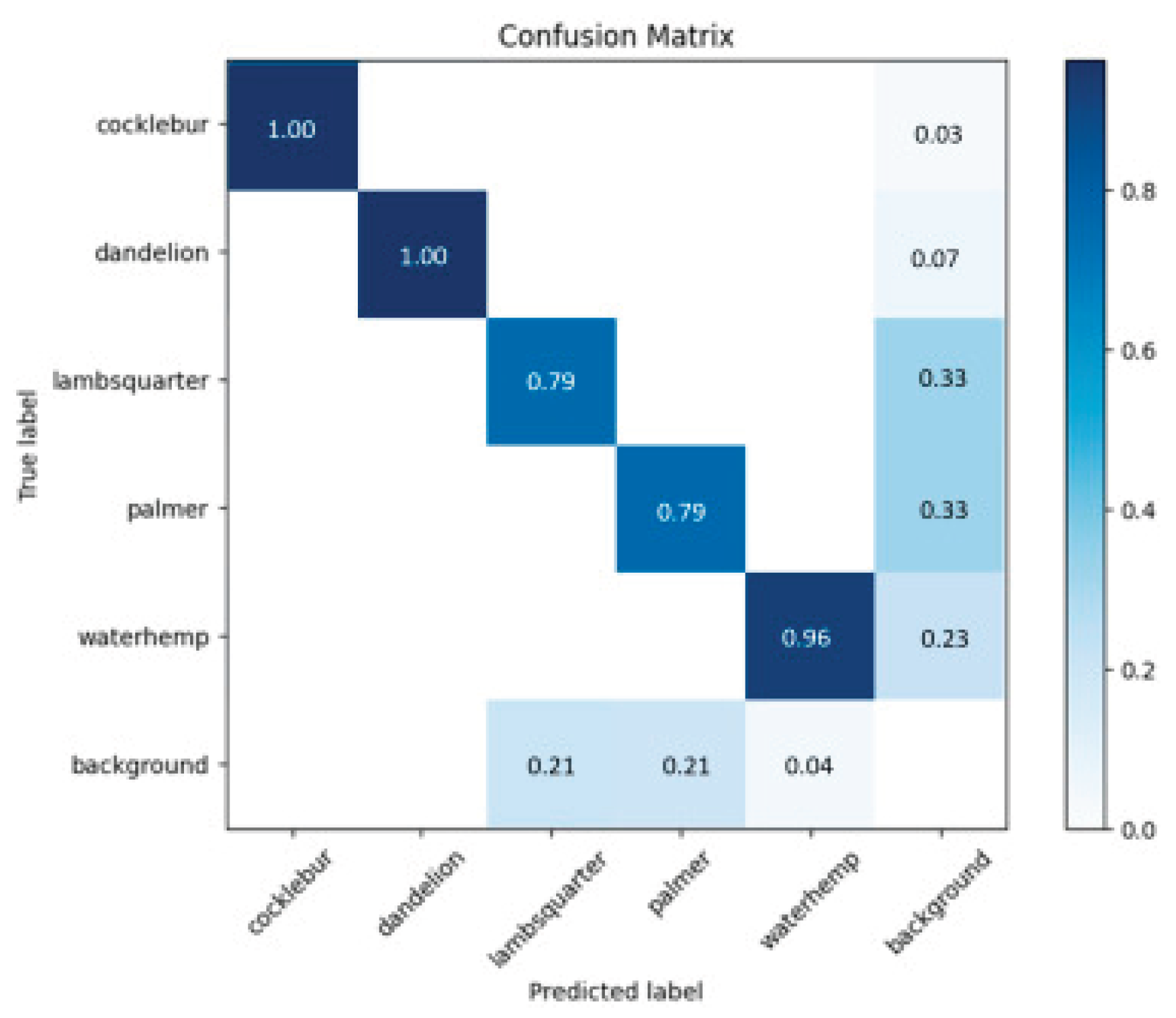
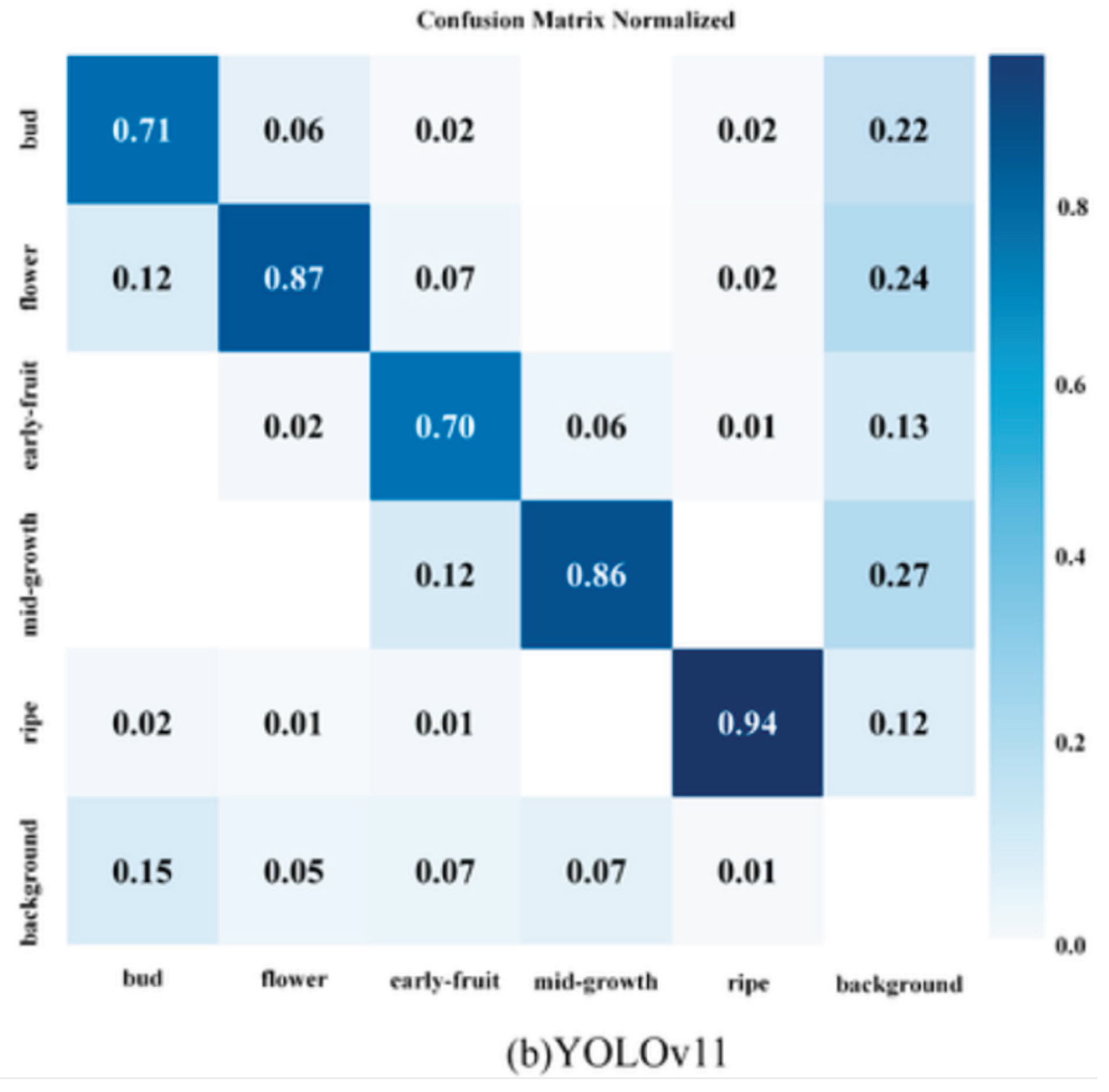
Precision Recall Curve
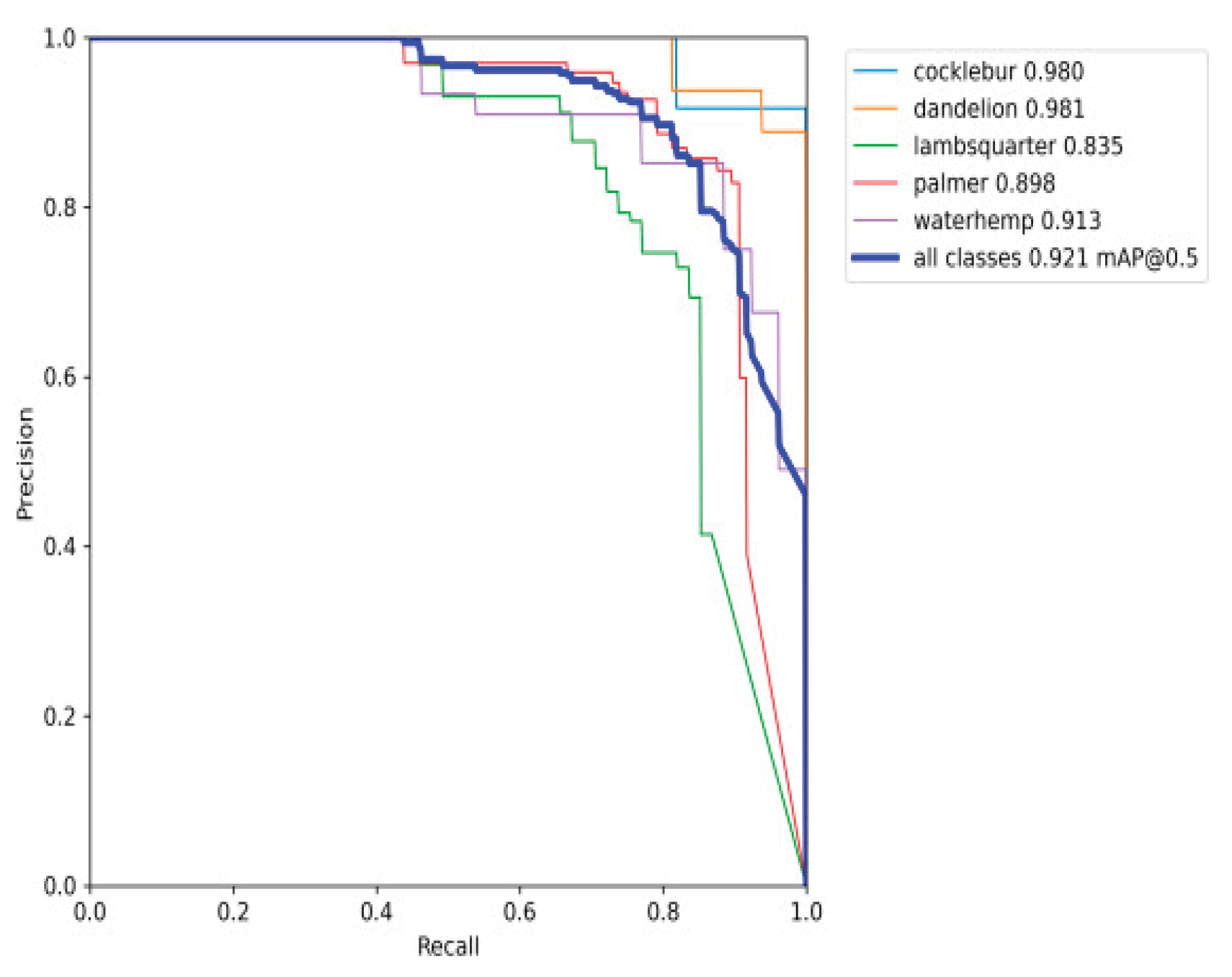
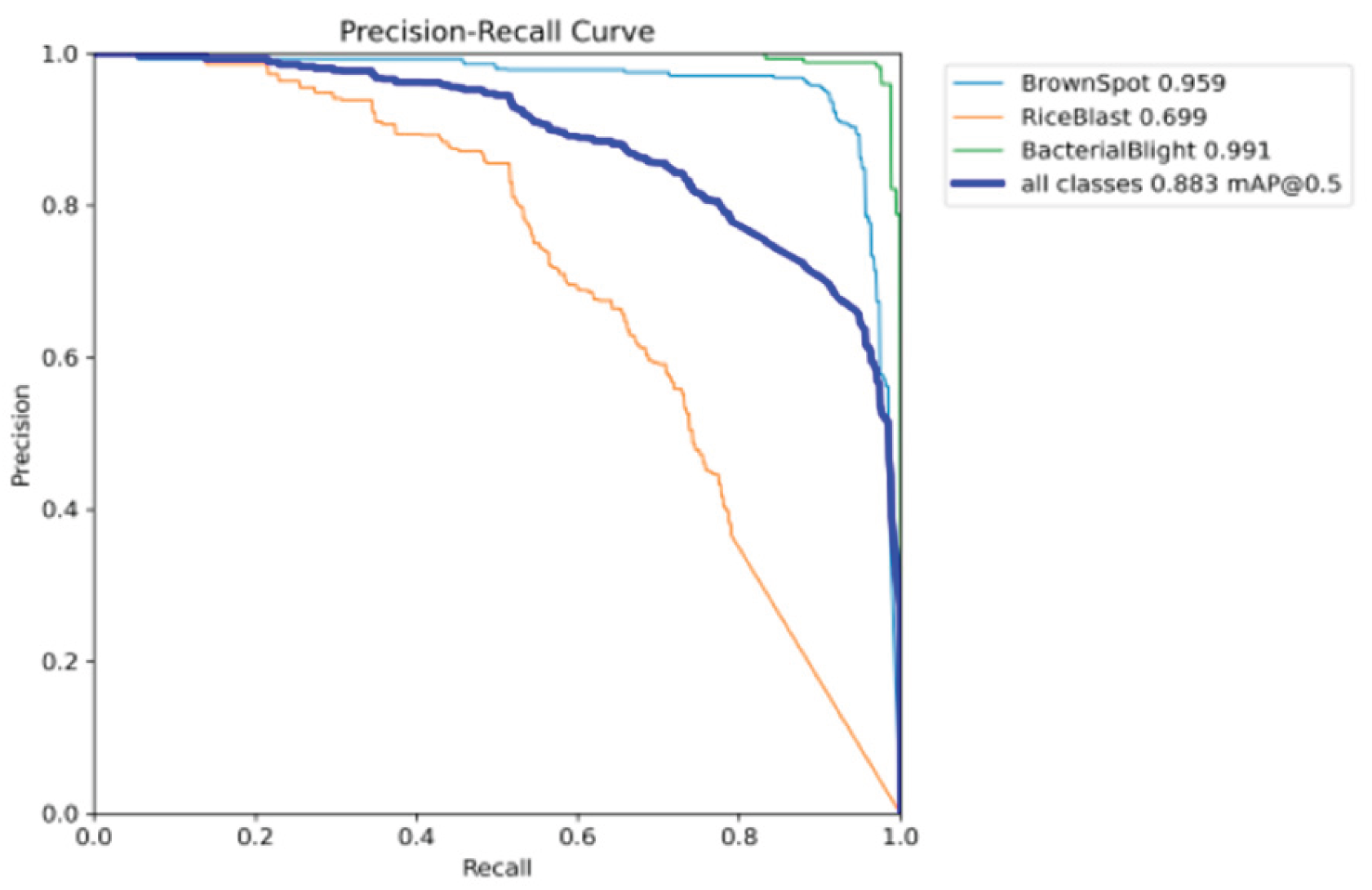
Visualization and Decision Support
Evaluation and Validation
Results and Discussion
Detection Accuracy and Model Performance
Comparison of Accuracy
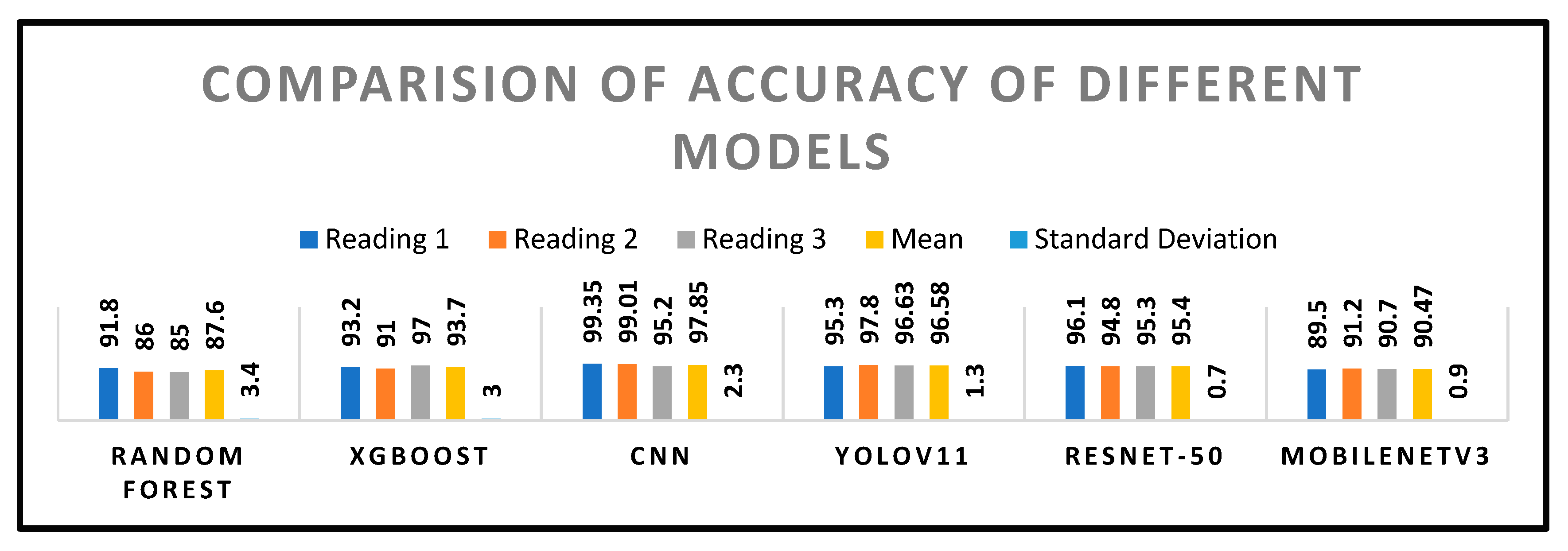
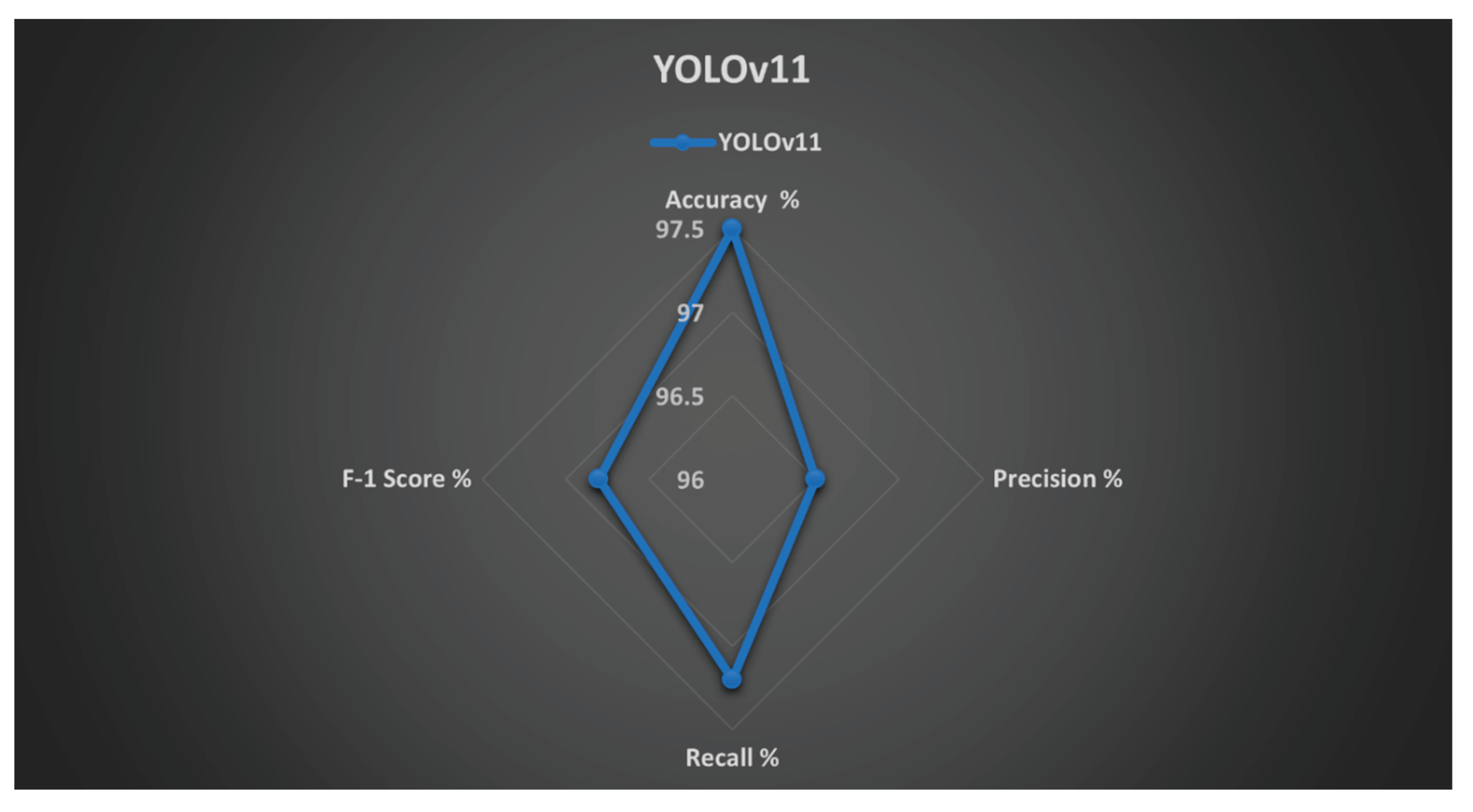
| Model | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) | mAP@0.5 |
|---|---|---|---|---|---|
| YOLOv11 | 97.5 | 96.5 | 97.2 | 96.8 | 0.975 |
| YOLOv10 | 95.1 | 94.2 | 94.8 | 94.2 | 0.951 |
| YOLOv8 | 93.7 | 92.5 | 93.1 | 92.5 | 0.937 |
| ResNet-50 | 92.1 | 90.7 | 91.5 | 90.7 | 0.921 |
| Random Forest | 75.8 | 74.1 | 73.5 | 77.9 | - |
| XGBoost | 79.3 | 78.1 | 77.1 | 77.6 | - |
| MobileNetV3 | 89.7 | 88.2 | 88.9 | 88.2 | 0.897 |
Composite Health Index (CHI) Assessment
- NDVI, EVI, NDWI: Computed values of these vegetation indices for the region.
- Morphological Features: A norm-scored measure obtained from morphological measurement (e.g., leaf area, perimeter, ratio of aspect).
- Texture Features: A score normalized from texture analysis procedures such as GLCM (Contrast, Homogeneity) or LBP.
- w₁, w₂, w₃, w₄, w₅, w₆: The weights that each factor is optimized to. They are optimized during the training of the model to achieve maximum correlation with true plant health responses. All weights typically sum to 1.
Field Trial Outcomes
- Pesticide use declined by 26% with targeted intervention, reducing costs and environmental impact.
- Efficiency of scouting was increased threefold, as UAVs and automated analysis replaced human inspection.
- Yield loss was reduced by 32%, attributed to early disease detection and timely treatment interventions, Yield increased by 12–15% in monitored plots.
- The system enabled a 20% reduction in chemical usage by providing precision alerts, thereby promoting more sustainable crop management practices.
| Metric | Conventional Scouting | System | Improvement |
|---|---|---|---|
| Pesticide Use | 100% (baseline) | 74% | -26% |
| Scouting Efficiency | 1× (baseline) | 3× | +200% |
| Yield Loss | 100% (baseline) | 68% | -32% |
| Yield Improvement | – | +12–15% | +12–15% |
Comparison with Other Models
| Model | Inference (ms) | Power (W) | FPS | RAM (GB) |
|---|---|---|---|---|
| YOLOv11 (Ours) | 142 | 1.8 | 5-7 FPS | 1.2 |
| YOLOv10 | 155 | 2.1 | 4-6 FPS | 1.4 |
| YOLOv8 | 175 | 2.3 | 4-5 FPS | 1.3 |
| ResNet-50 | 480 | 8.5 | <1 FPS | 3.8 |
| MobileNetV3 | 95 | 1.2 | 6-7 FPS | 0.9 |
| XGBoost | 22 | 4.1 | - | 0.4 |
| Random Forest | 18 | 4.0 | - | 0.6 |
Challenges and Limitations
Implications for Sustainable Agriculture
Future Prospects
- Federated learning techniques to facilitate model improvement without data centralization, maintaining farmer data confidentiality.
- Predictive analysis to forecast potential outbreaks based on historical and real-time environmental data.
- Automation by robots for precision spraying with a reduction in human labor force and chemical utilization.
- Dataset increase to cover diverse crops, climatic regions, and diseases for enhanced model generalization.
Overall Discussion
Conclusions
References
- S. P. Mohanty, D. P. Hughes, and M. Salathé, “Using deep learning for image-based plant disease detection,” Front. Plant Sci., vol. 7, Sep. 2016, Art. no. 1419. [CrossRef]
- M. Abu John, I. Bankole, O. Ajayi-Moses, T. Ijila, T. Jeje, P. Lalit, and O. Jeje, “Relevance of advanced plant disease detection techniques in disease and pest management for ensuring food security and their implication: A review,” Amer. J. Plant Sci., vol. 14, no. 11, pp. 1260– 1295, 2023. [CrossRef]
- Kaggle, “Plant village dataset.” [Online]. Available: https://www.kaggle.com/emmarex/plantdisease.
- M. Redowan and A. H. Kanan, “Potentials and limitations of NDVI and other vegetation indices (VIS) for monitoring vegetation parameters from remotely sensed data,” Bangladesh Res. Publ. J, vol. 7, no. 3, pp. 291–299, Sept.–Oct. 2012.
- S. S. Harakannanavar, J. M. Rudagi, V. I. Puranikmath, A. Siddiqua, and R. Pramodhini, “Plant leaf disease detection using computer vision and machine learning algorithms,” Glob. Transitions Proc., vol. 3, pp. 305–310, 2022. [CrossRef]
- M. Islam, et al., “DeepCrop: Deep learning-based crop disease prediction with web application,” J. Agric. Food Res., vol. 14, Art. no. 100764, 2023. [CrossRef]
- DroneDeploy, “Agriculture report 2023.” [Online]. Available: https://www.dronedeploy.com/resources/reports/agriculture-2023.
- M. B. Khan, S. Tamkin, J. Ara, M. Alam, and H. Bhuiyan, “CropsDisNet: An AI-based platform for disease detection,” Comput. Electron. Agric., early access, Feb. 2025. [CrossRef]
- Md. M. Islam, “DeepCrop: Deep learning-based crop disease prediction with web application,” J. Agric. Food Res., vol. 14, Dec. 2023, published Aug. 28, 2023.
- Anjna, M. Sood, and P. K. Singh, “Hybrid system for detection and classification of plant disease using qualitative texture features analysis,” Procedia Comput. Sci., vol. 167, pp. 1056–1065, 2020. [CrossRef]
- R. Gajjar, N. Gajjar, V. J. Thakor, and N. P. Patel, “Real-time detection and identification of plant leaf diseases using convolutional neural networks on an embedded platform,” Vis. Comput., vol. 38, pp. 2923–2938, 2021. [CrossRef]
- M. Agarwal, V. K. Bohat, M. D. Ansari, A. Sinha, S. K. Gupta, and D. Garg, “A convolution neural network-based approach to detect the disease in corn crop,” in Proc. IEEE 9th Int. Conf. Adv. Comput. (IACC), Tiruchirappalli, India, Dec. 2019, pp. 176–181.
- H. N. Ngugi, A. A. Akinyelu, and A. E. Ezugwu, “Machine learning and deep learning for crop disease diagnosis,” Comput. Electron. Agric., published Dec. 2024. [CrossRef]
- K. R. B. Leelavathy, “Plant disease classification using VGG-16: Performance evaluation and comparative analysis,” Int. J. Environ. Sci. Technol., vol. 23, pp. 219–229, 2023.
- D. Tirkey, K. K. Singh, and S. Tripathi, “Performance analysis of AI-based solutions for crop disease identification, detection, and classification,” in Smart Agric. Technol., vol. 5, Oct. 2023. [CrossRef]
- T. Kavzoglu and A. Teke, “Predictive performances of ensemble machine learning algorithms in landslide susceptibility mapping using random forest, extreme gradient boosting (XGBoost) and natural gradient boosting (NGBoost),” Arab. J. Sci. Eng., vol. 47, pp. 7367–7385, 2022. [CrossRef]
- L. Sosa, A. Justel, and Í. Molina, “Detection of crop hail damage with a machine learning algorithm using time series of remote sensing data,” Agronomy, vol. 11, Art. no. 2078, 2021. [CrossRef]
- Sharma, V. Kumar, and L. Longchamps, “Comparative performance of YOLOv8, YOLOv9, YOLOv10, YOLOv11 and Faster R-CNN models for detection of multiple weed species,” in Smart Agric. Technol., vol. 9, Dec. 2024.
- M. A. Talukder, et al., “An efficient deep learning model to categorize brain tumor using reconstruction and fine-tuning,” Expert Syst. Appl., 2023, Art. no. 120534.
- N. Ahmed, et al., “Machine learning based diabetes prediction and development of smart web application,” Int. J. Cogn. Comput. Eng., vol. 2, pp. 229–241, 2021.
- M. D. Bah, A. Hafiane, and R. Canals, “Deep learning with unsupervised data labeling for weed detection in line crops in UAV images,” Remote Sens., vol. 10, no. 11, p. 1690, 2018.
- M. A. Talukder, M. M. Islam, M. A. Uddin, A. Akhter, K. F. Hasan, and M. A. Moni, “Machine learning-based lung and colon cancer detection using deep feature extraction and ensemble learning,” Expert Syst. Appl., vol. 205, Art. no. 117695, 2022.
- Z.-Q. Zhao, P. Zheng, S.-T. Xu, and X. Wu, “Object detection with deep learning: A review,” IEEE Trans. Neural Netw. Learn. Syst., vol. 30, no. 11, pp. 3212–3232, Nov. 2019.
- V. N. T. Le, G. Truong, and K. Alameh, “Detecting weeds from crops under complex field environments based on Faster R-CNN,” in Proc. IEEE Int. Conf., 2021, pp. 350–355.
- N. Jmour, S. Zayen, and A. Abdelkrim, “Convolutional neural networks for image classification,” in Proc. Int. Conf. Adv. Syst. Elect. Technol. (IC_ASET), Hammamet, Tunisia, 2018, pp. 397–402.
- J. Qi, et al., “An improved YOLOv5 model based on visual attention mechanism: Application to recognition of tomato virus disease,” Comput. Electron. Agric., vol. 194, Art. no. 106780, 2022. [CrossRef]
- E. Hirani, V. Magotra, J. Jain, and P. Bide, “Plant disease detection using deep learning,” in Proc. Int. Conf. Convergence Technol. (I2CT), Maharashtra, India, 2021.
- F. Ertam and G. Aydın, “Data classification with deep learning using TensorFlow,” in Proc. Int. Conf. Comput. Sci. Eng. (UBMK), 2017, pp. 755–758. [CrossRef]
- S. Pawar, S. Shedge, N. Panigrahi, A. Jyoti, P. Thorave, and S. Sayyad, “Leaf disease detection of multiple plants using deep learning,” in Proc. Int. Conf. Mach. Learn., Big Data, Cloud Parallel Comput. (COM-IT-CON), 2022, pp. 241–245.
- D. Tiwari, M. Ashish, N. Gangwar, A. Sharma, S. Patel, and S. Bhardwaj, “Potato leaf diseases detection using deep learning,” in Proc. 4th Int. Conf. Intell. Comput. Control Syst. (ICICCS), 2020, pp. 461–466.
- V. Pooja, R. Das, and V. Kanchana, “Identification of plant leaf diseases using image processing techniques,” in Proc. Int. Conf. Tech. Innov. ICT Agric. Rural Develop., 2017, pp. 130–133. [CrossRef]
- W. Shafik, A. Tufail, C. D. S. Liyanage, and R. A. A. H. M. Apong, “Using transfer learning-based plant disease classification and detection for sustainable agriculture,” BMC Plant Biol., vol. 24, pp. 1–19, 2024. [CrossRef]
- T. Hayit, H. Erbay, F. Varçın, F. Hayit, and N. Akci, “Determination of the severity level of yellow rust disease in wheat by using convolutional neural networks,” J. Plant Pathol., vol. 103, pp. 923–934, 2021. [CrossRef]
- J. Yule and R. R. Pullanagari, Optical Sensors to Assist Agricultural Crop and Pasture Management. Palmerston North, New Zealand: Massey Univ., Jan. 2012.
- H. Zhu, “Intelligent agriculture: Deep learning in UAV-based remote sensing imagery for crop diseases and pests detection,” Front. Plant Sci., vol. 15, 2024, Art. no. e1435016.
- H. M. Faisal, “Detection of cotton crops diseases using customized deep learning models including VGG16, ResNet152, EfficientNet, InceptionV3, MobileNet, and others: ResNet152 outperforms in cotton disease recognition,” Sci. Rep., 2025.
- B. V. Baiju, “Robust CRW crops leaf disease detection and classification in corn, rice, and wheat using Slender-CNN,” Plant Methods, 2025.
- Y. Ashurov, “Enhancing plant disease detection through deep learning: A modified depthwise CNN with squeeze-and-excitation blocks and improved residual skip connections,” Front. Plant Sci., 2025.
- S. Wang, et al., “Advances in deep learning applications for plant disease and pest detection: A review,” Remote Sens., vol. 17, no. 4, Art. no. 698, 2025. [CrossRef]
- R. Sujatha, S. Krishnan, J. M. Chatterjee, and A. H. Gandomi, “Advancing plant leaf disease detection integrating machine learning and deep learning,” Sci. Rep., vol. 15, Art. no. 11552, 2025. [CrossRef]
- K. Fang, R. Zhou, N. Deng, C. Li, and X. Zhu, “RLDD-YOLOv11n: Research on rice leaf disease detection based on YOLOv11,” Agronomy, vol. 15, no. 6, p. 1266, May 2025. [CrossRef]
- X. Tang, Z. Sun, L. Yang, Q. Chen, Z. Liu, P. Wang, and Y. Zhang, “YOLOv11-AIU: A lightweight detection model for the grading detection of early blight disease in tomatoes,” Plant Methods, vol. 21, Article 118, 2025. [CrossRef]
- L. Xu, B. Li, X. Fu, Z. Lu, Z. Li, B. Jiang, and S. Jia, “YOLO-MSNet: Real-time detection algorithm for pomegranate fruit improved by YOLOv11n,” Agriculture, vol. 15, no. 10, p. 1028, May 2025. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).