Submitted:
05 September 2025
Posted:
05 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Ethics and Regulatory Statement
2.2. RNA-Seq and Bioinformatics
2.3. RT-qPCR of Nasopharyngeal Samples
2.4. Phylogenetic Tree Generation
2.5. Statistics
3. Results
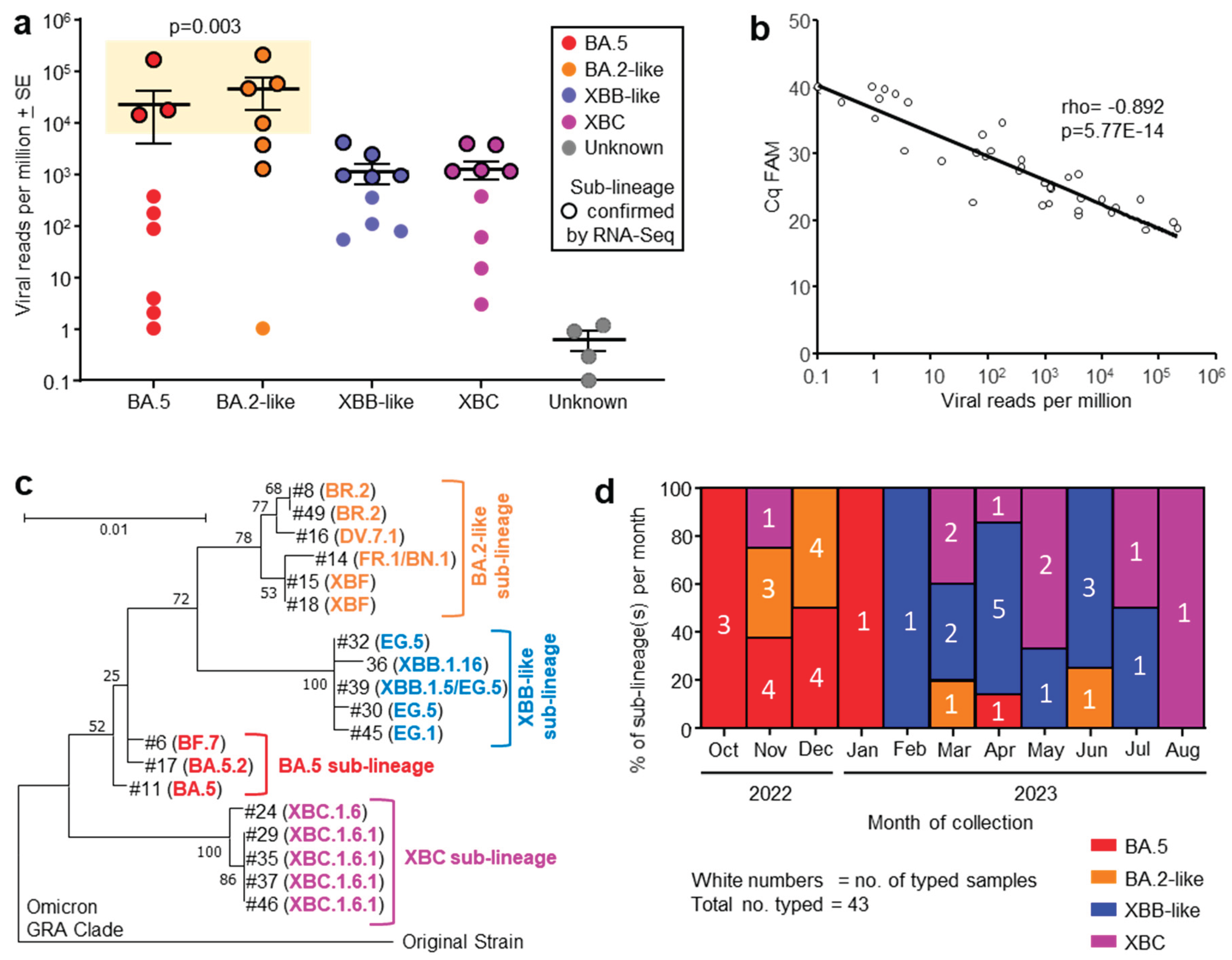
3.1. Omicron Sublineages and Viral Read Counts from COVID-19 ICU Patients
3.2. Quantitation by RT-qPCR Correlated Well with Viral read Counts
3.3. Phylogenetic Tree of Omicron Sublineages
3.4. Progression of Omicron Sublineages from Oct 2022 to Aug 2023
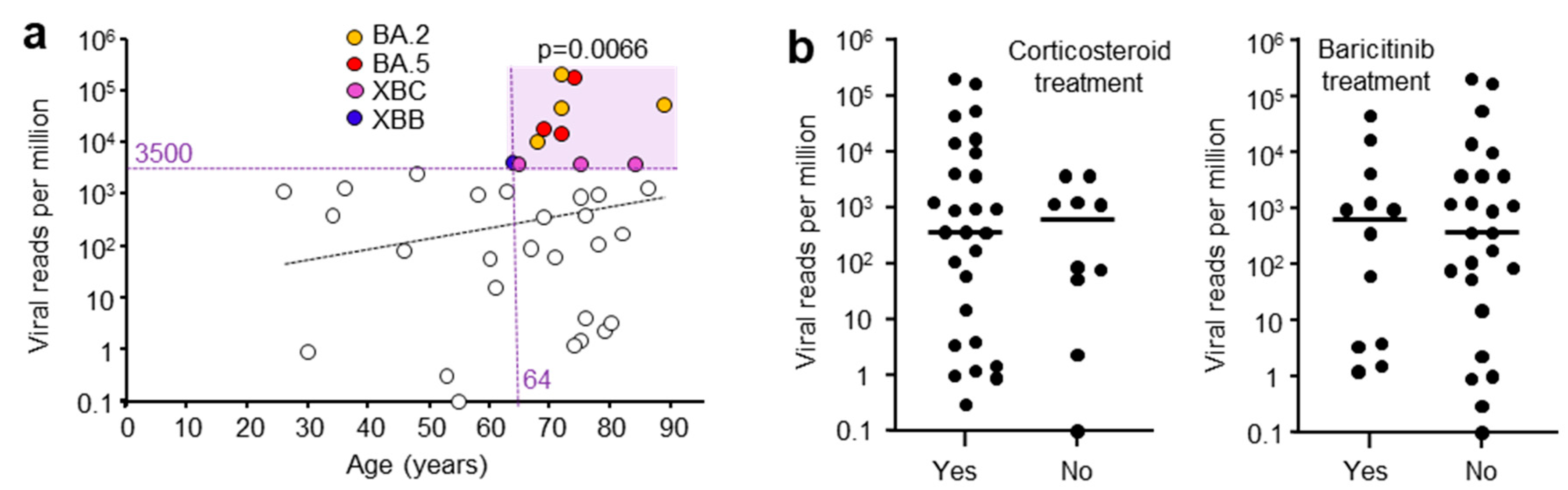
3.5. Correlations of Viral Read Counts with Age and Other Patient Data
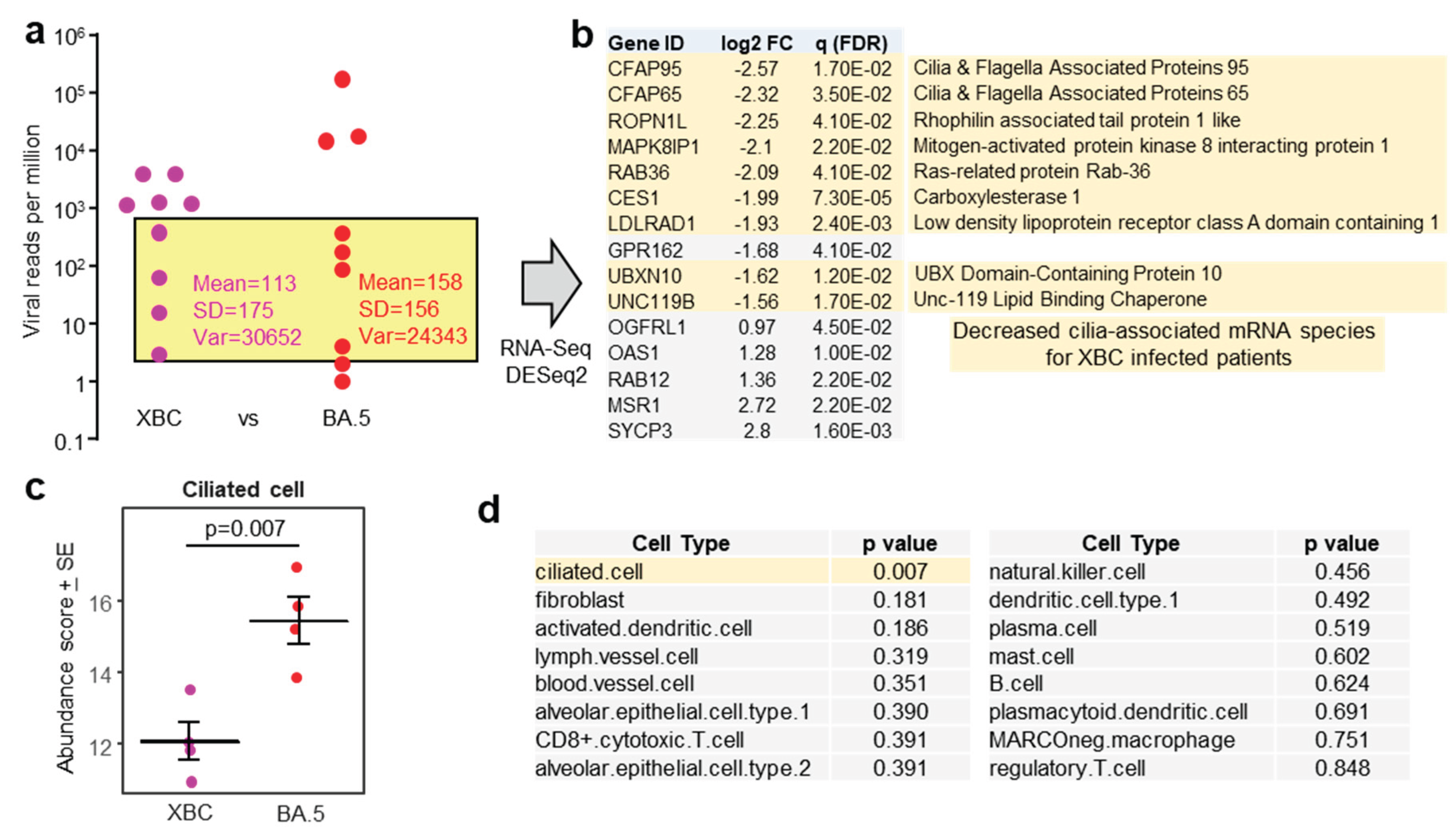
3.5. RNA-Seq Analysis of Human Gene Expression
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| COVID-19 | Coronavirus Disease of 2019 |
| SARS-CoV-2 | Severe acute respiratory syndrome coronavirus 2 |
| RT-qPCR | Reverse transcriptase quantitative polymerase chain reaction |
| RNA-Seq | Ribonucleic acid sequencing |
| mRNA | Messenger RNA |
| RIN | RNA integrity number |
| DEGs | Differentially expressed genes |
| RPM | Reads per million |
| FDR | False discovery rate (q) |
| FC | Fold change |
References
- Markov, P. V.; Ghafari, M.; Beer, M.; Lythgoe, K.; Simmonds, P.; Stilianakis, N. I.; Katzourakis, A., The evolution of SARS-CoV-2. Nature Reviews Microbiology 2023, 21, 361-379. [CrossRef]
- Roemer, C.; Sheward, D. J.; Hisner, R.; Gueli, F.; Sakaguchi, H.; Frohberg, N.; Schoenmakers, J.; Sato, K.; O’Toole, Á.; Rambaut, A., et al., SARS-CoV-2 evolution in the Omicron era. Nature Microbiology 2023, 8, 1952-1959. [CrossRef]
- Hu, B.; Chan, J. F.-W.; Liu, Y.; Liu, H.; Chen, Y.-X.; Shuai, H.; Hu, Y.-F.; Hartnoll, M.; Chen, L.; Xia, Y., et al., Divergent trajectory of replication and intrinsic pathogenicity of SARS-CoV-2 Omicron post-BA.2/5 subvariants in the upper and lower respiratory tract. eBioMedicine 2024, 99, 104916. [CrossRef]
- Velavan, T. P.; Ntoumi, F.; Kremsner, P. G.; Lee, S. S.; Meyer, C. G., Emergence and geographic dominance of Omicron subvariants XBB/XBB.1.5 and BF.7 - the public health challenges. Int J Infect Dis 2023, 128, 307-309. [CrossRef]
- Wang, Q.; Guo, Y.; Zhang, R. M.; Ho, J.; Mohri, H.; Valdez, R.; Manthei, D. M.; Gordon, A.; Liu, L.; Ho, D. D., Antibody neutralisation of emerging SARS-CoV-2 subvariants: EG.5.1 and XBC.1.6. Lancet Infect Dis 2023, 23, e397-e398. [CrossRef]
- Carabelli, A. M.; Peacock, T. P.; Thorne, L. G.; Harvey, W. T.; Hughes, J.; Peacock, S. J.; Barclay, W. S.; de Silva, T. I.; Towers, G. J.; Robertson, D. L., SARS-CoV-2 variant biology: immune escape, transmission and fitness. Nat Rev Microbiol 2023, 21, 162-177. [CrossRef]
- Dadonaite, B.; Brown, J.; McMahon, T. E.; Farrell, A. G.; Figgins, M. D.; Asarnow, D.; Stewart, C.; Lee, J.; Logue, J.; Bedford, T., et al., Spike deep mutational scanning helps predict success of SARS-CoV-2 clades. Nature 2024, 631, 617-626. [CrossRef]
- Steiner, S.; Kratzel, A.; Barut, G. T.; Lang, R. M.; Aguiar Moreira, E.; Thomann, L.; Kelly, J. N.; Thiel, V., SARS-CoV-2 biology and host interactions. Nature Reviews Microbiology 2024, 22, 206-225. [CrossRef]
- Gupta, S.; Gupta, D.; Bhatnagar, S., Analysis of SARS-CoV-2 genome evolutionary patterns. Microbiology Spectrum 2024, 12, e02654-23. [CrossRef]
- Ivanov, K. I.; Yang, H.; Sun, R.; Li, C.; Guo, D., The emerging role of SARS-CoV-2 nonstructural protein 1 (nsp1) in epigenetic regulation of host gene expression. FEMS Microbiology Reviews 2024, 48, fuae023. [CrossRef]
- Garcia Lopez, V.; Plate, L., Comparative Interactome Profiling of Nonstructural Protein 3 Across SARS-CoV-2 Variants Emerged During the COVID-19 Pandemic. Viruses 2025, 17, 447. [CrossRef]
- Takada, K.; Orba, Y.; Kida, Y.; Wu, J.; Ono, C.; Matsuura, Y.; Nakagawa, S.; Sawa, H.; Watanabe, T., Genes involved in the limited spread of SARS-CoV-2 in the lower respiratory airways of hamsters may be associated with adaptive evolution. Journal of Virology 2024, 98, e01784-23. [CrossRef]
- Wu, C.-T.; Lidsky, P. V.; Xiao, Y.; Cheng, R.; Lee, I. T.; Nakayama, T.; Jiang, S.; He, W.; Demeter, J.; Knight, M. G., et al., SARS-CoV-2 replication in airway epithelia requires motile cilia and microvillar reprogramming. Cell 2023, 186, 112-130.e20. [CrossRef]
- Fonseca, B. F.; Robinot, R.; Michel, V.; Mendez, A.; Lebourgeois, S.; Chivé, C.; Jeger-Madiot, R.; Vaid, R.; Bondet, V.; Maloney, E., et al., Stealth replication of SARS-CoV-2 Omicron in the nasal epithelium at physiological temperature. bioRxiv 2025, 2025.05.03.652024. [CrossRef]
- Dumenil, T.; Le, T. T.; Rawle, D. J.; Yan, K.; Tang, B.; Nguyen, W.; Bishop, C.; Suhrbier, A., Warmer ambient air temperatures reduce nasal turbinate and brain infection, but increase lung inflammation in the K18-hACE2 mouse model of COVID-19. Sci Total Environ 2023, 859, 160163. [CrossRef]
- Dai, X.; Xu, R.; Li, N., The Interplay between Airway Cilia and Coronavirus Infection, Implications for Prevention and Control of Airway Viral Infections. Cells 2024, 13, 1353. [CrossRef]
- El-Daly, M. M., Advances and Challenges in SARS-CoV-2 Detection: A Review of Molecular and Serological Technologies. Diagnostics 2024, 14, 519. [CrossRef]
- Pollak, N. M.; Rawle, D. J.; Yan, K.; Buckley, C.; Le, T. T.; Wang, C. Y. T.; Ertl, N. G.; van Huyssteen, K.; Crkvencic, N.; Hashmi, M., et al., Rapid inactivation and sample preparation for SARS-CoV-2 PCR-based diagnostics using TNA-Cifer Reagent E. Frontiers in Microbiology 2023, 14, 1238542. [CrossRef]
- Wu, Y.; Guo, Z.; Yuan, J.; Cao, G.; Wang, Y.; Gao, P.; Liu, J.; Liu, M., Duration of viable virus shedding and polymerase chain reaction positivity of the SARS-CoV-2 Omicron variant in the upper respiratory tract: a systematic review and meta-analysis. International Journal of Infectious Diseases 2023, 129, 228-235. [CrossRef]
- Stewart, R.; Yan, K.; Ellis, S. A.; Bishop, C. R.; Dumenil, T.; Tang, B.; Nguyen, W.; Larcher, T.; Parry, R.; Sng, J. J., et al., SARS-CoV-2 omicron BA.5 and XBB variants have increased neurotropic potential over BA.1 in K18-hACE2 mice and human brain organoids. Front Microbiol 2023, 14, 1320856. [CrossRef]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J.; Homer, N.; Marth, G.; Abecasis, G.; Durbin, R.; Genome Project Data Processing, S., The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009, 25, 2078-2079. [CrossRef]
- Love, M. I.; Huber, W.; Anders, S., Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome biology 2014, 15, 550. [CrossRef]
- Bishop, C. R.; Yan, K.; Nguyen, W.; Rawle, D. J.; Tang, B.; Larcher, T.; Suhrbier, A., Microplastics dysregulate innate immunity in the SARS-CoV-2 infected lung. Frontiers in Immunology 2024, 15, 1382655. [CrossRef]
- Danaher, P.; Kim, Y.; Nelson, B.; Griswold, M.; Yang, Z.; Piazza, E.; Beechem, J. M., Advances in mixed cell deconvolution enable quantification of cell types in spatial transcriptomic data. Nature Communications 2022, 13, 385. [CrossRef]
- Madissoon, E.; Wilbrey-Clark, A.; Miragaia, R. J.; Saeb-Parsy, K.; Mahbubani, K. T.; Georgakopoulos, N.; Harding, P.; Polanski, K.; Huang, N.; Nowicki-Osuch, K., scRNA-seq assessment of the human lung, spleen, and esophagus tissue stability after cold preservation. Genome biology 2019, 21, 1. [CrossRef]
- Vogels, C. B. F.; Brito, A. F.; Wyllie, A. L.; Fauver, J. R.; Ott, I. M.; Kalinich, C. C.; Petrone, M. E.; Casanovas-Massana, A.; Catherine Muenker, M.; Moore, A. J., et al., Analytical sensitivity and efficiency comparisons of SARS-CoV-2 RT–qPCR primer–probe sets. Nature Microbiology 2020, 5, 1299-1305. [CrossRef]
- Robinson, J. T.; Thorvaldsdottir, H.; Turner, D.; Mesirov, J. P., igv.js: an embeddable JavaScript implementation of the Integrative Genomics Viewer (IGV). Bioinformatics 2023, 39, btac830. [CrossRef]
- Scarpa, F.; Locci, C.; Azzena, I.; Casu, M.; Fiori, P. L.; Ciccozzi, A.; Giovanetti, M.; Quaranta, M.; Ceccarelli, G.; Pascarella, S., et al., SARS-CoV-2 recombinants: Genomic comparison between XBF and its parental lineages. Microorganisms 2023, 11, 1824. [CrossRef]
- Chhoung, C.; Ko, K.; Ouoba, S.; Phyo, Z.; Akuffo, G. A.; Sugiyama, A.; Akita, T.; Sasaki, H.; Yamamoto, T.; Takahashi, K., et al., Sustained applicability of SARS-CoV-2 variants identification by Sanger Sequencing Strategy on emerging various SARS-CoV-2 Omicron variants in Hiroshima, Japan. BMC Genomics 2024, 25, 1063. [CrossRef]
- To, K. K.-W.; Tsang, O. T.-Y.; Leung, W.-S.; Tam, A. R.; Wu, T.-C.; Lung, D. C.; Yip, C. C.-Y.; Cai, J.-P.; Chan, J. M.-C.; Chik, T. S.-H., et al., Temporal profiles of viral load in posterior oropharyngeal saliva samples and serum antibody responses during infection by SARS-CoV-2: an observational cohort study. The Lancet Infectious Diseases 2020, 20, 565-574. [CrossRef]
- Zhong, W.; Yang, X.; Jiang, X.; Duan, Z.; Wang, W.; Sun, Z.; Chen, W.; Zhang, W.; Xu, J.; Cheng, J., et al., Factors associated with prolonged viral shedding in older patients infected with Omicron BA.2.2. Frontiers in Public Health 2023, 10, 1087800. [CrossRef]
- Johns, M.; George, S.; Taburyanskaya, M.; Poon, Y. K., A Review of the Evidence for Corticosteroids in COVID-19. Journal of Pharmacy Practice 2021, 35, 626-637. [CrossRef]
- Huang, R.; Zhu, C.; Jian, W.; Xue, L.; Li, C.; Yan, X.; Huang, S.; Zhang, B.; Zhu, L.; Xu, T., et al., Corticosteroid therapy is associated with the delay of SARS-CoV-2 clearance in COVID-19 patients. European Journal of Pharmacology 2020, 889, 173556. [CrossRef]
- Spagnuolo, V.; Guffanti, M.; Galli, L.; Poli, A.; Querini, P. R.; Ripa, M.; Clementi, M.; Scarpellini, P.; Lazzarin, A.; Tresoldi, M., et al., Viral clearance after early corticosteroid treatment in patients with moderate or severe covid-19. Scientific Reports 2020, 10, 21291. [CrossRef]
- Patanwala, A. E.; Xiao, X.; Hills, T. E.; Higgins, A. M.; McArthur, C. J.; Alexander, G. C.; Mehta, H. B.; on behalf of National Covid Cohort Collaborative, C., Comparative Effectiveness of Baricitinib Versus Tocilizumab in Hospitalized Patients With COVID-19: A Retrospective Cohort Study of the National Covid Collaborative. Critical Care Medicine 2025, 53, e29-41. [CrossRef]
- Viermyr, H.-K.; Tonby, K.; Ponzi, E.; Trouillet-Assant, S.; Poissy, J.; Arribas, J. R.; Dyon-Tafani, V.; Bouscambert-Duchamp, M.; Assoumou, L.; Halvorsen, B., et al., Safety of baricitinib in vaccinated patients with severe and critical COVID-19 sub study of the randomised Bari-SolidAct trial. eBioMedicine 2025, 111, 105511. [CrossRef]
- Maltezou, H. C.; Raftopoulos, V.; Vorou, R.; Papadima, K.; Mellou, K.; Spanakis, N.; Kossyvakis, A.; Gioula, G.; Exindari, M.; Froukala, E., et al., Association Between Upper Respiratory Tract Viral Load, Comorbidities, Disease Severity, and Outcome of Patients With SARS-CoV-2 Infection. The Journal of Infectious Diseases 2021, 223, 1132-1138. [CrossRef]
- Cocconcelli, E.; Castelli, G.; Onelia, F.; Lavezzo, E.; Giraudo, C.; Bernardinello, N.; Fichera, G.; Leoni, D.; Trevenzoli, M.; Saetta, M., et al., Disease Severity and Prognosis of SARS-CoV-2 Infection in Hospitalized Patients Is Not Associated With Viral Load in Nasopharyngeal Swab. Frontiers in Medicine 2021, 8, 714221. [CrossRef]
- Liu, J.; Wen, R.; Wang, N.; Li, G.; Xu, P.; Li, X.; Zeng, X.; Liu, C., A retrospective study on COVID-19 infections caused by omicron variant with clinical, epidemiological, and viral load evaluations in breakthrough infections. International Journal of Medical Sciences 2024, 21, 454. [CrossRef]
- Ura, H.; Niida, Y., Comparison of RNA-Sequencing Methods for Degraded RNA. Int J Mol Sci 2024, 25, 6143. [CrossRef]
- Lu, W.; Zhou, Q.; Chen, Y., Impact of RNA degradation on next-generation sequencing transcriptome data. Genomics 2022, 114, 110429. [CrossRef]
- Carolin, A.; Frazer, D.; Yan, K.; Bishop, C. R.; Tang, B.; Nguyen, W.; Helman, S. L.; Horvat, J.; Larcher, T.; Rawle, D. J., et al., The effects of iron deficient and high iron diets on SARS-CoV-2 lung infection and disease. Frontiers in Microbiology 2024, 15, 1441495. [CrossRef]
- Carolin, A.; Yan, K.; Bishop, C. R.; Tang, B.; Nguyen, W.; Rawle, D. J.; Suhrbier, A., Tracking inflammation resolution signatures in lungs after SARS-CoV-2 omicron BA.1 infection of K18-hACE2 mice. PLOS ONE 2024, 19, e0302344. [CrossRef]
- Rochette, L.; Zeller, M.; Cottin, Y.; Vergely, C., GDF15: an emerging modulator of immunity and a strategy in COVID-19 in association with iron metabolism. Trends Endocrinol Metab 2021, 32, 875-889. [CrossRef]
- Andersen, J. S.; Vijayakumaran, A.; Godbehere, C.; Lorentzen, E.; Mennella, V.; Schou, K. B., Uncovering structural themes across cilia microtubule inner proteins with implications for human cilia function. Nature Communications 2024, 15, 2687. [CrossRef]
- Wang, W.; Tu, C.; Nie, H.; Meng, L.; Li, Y.; Yuan, S.; Zhang, Q.; Du, J.; Wang, J.; Gong, F., et al., Biallelic mutations in CFAP65 lead to severe asthenoteratospermia due to acrosome hypoplasia and flagellum malformations. J Med Genet 2019, 56, 750-757. [CrossRef]
- Liu, Y.; Liu, T.; Ruan, L.; Zhu, D.; He, Y.; Jia, J.; Chen, Y., Cilia Plays a Pivotal Role in the Hypersecretion of Airway Mucus in Mice. Current Molecular Pharmacology 2024, 17, E18761429368288. [CrossRef]
- Verhey, K. J.; Hammond, J. W., Traffic control: regulation of kinesin motors. Nature Reviews Molecular Cell Biology 2009, 10, 765-777. [CrossRef]
- Ganga, A. K.; Kennedy, M. C.; Oguchi, M. E.; Gray, S.; Oliver, K. E.; Knight, T. A.; De La Cruz, E. M.; Homma, Y.; Fukuda, M.; Breslow, D. K., Rab34 GTPase mediates ciliary membrane formation in the intracellular ciliogenesis pathway. Curr Biol 2021, 31, 2895-2905.e7. [CrossRef]
- Li, R.; Liclican, A.; Xu, Y.; Pitts, J.; Niu, C.; Zhang, J.; Kim, C.; Zhao, X.; Soohoo, D.; Babusis, D., et al., Key Metabolic Enzymes Involved in Remdesivir Activation in Human Lung Cells. Antimicrobial Agents and Chemotherapy 2021, 65, 10.1128/aac.00602-21. [CrossRef]
- Geremek, M.; Bruinenberg, M.; Ziętkiewicz, E.; Pogorzelski, A.; Witt, M.; Wijmenga, C., Gene expression studies in cells from primary ciliary dyskinesia patients identify 208 potential ciliary genes. Human Genetics 2011, 129, 283-293. [CrossRef]
- Raman, M.; Sergeev, M.; Garnaas, M.; Lydeard, J. R.; Huttlin, E. L.; Goessling, W.; Shah, J. V.; Harper, J. W., Systematic proteomics of the VCP–UBXD adaptor network identifies a role for UBXN10 in regulating ciliogenesis. Nature Cell Biology 2015, 17, 1356-1369. [CrossRef]
- Jean, F.; Pilgrim, D., Coordinating the uncoordinated: UNC119 trafficking in cilia. European Journal of Cell Biology 2017, 96, 643-652. [CrossRef]
- Tanneti, N. S.; Patel, A. K.; Tan, L. H.; Marques, A. D.; Perera, R. A. P. M.; Sherrill-Mix, S.; Kelly, B. J.; Renner, D. M.; Collman, R. G.; Rodino, K., et al., Comparison of SARS-CoV-2 variants of concern in primary human nasal cultures demonstrates Delta as most cytopathic and Omicron as fastest replicating. mBio 2024, 15, e03129-23. [CrossRef]
- Zaderer, V.; Abd El Halim, H.; Wyremblewsky, A.-L.; Lupoli, G.; Dächert, C.; Muenchhoff, M.; Graf, A.; Blum, H.; Lass-Flörl, C.; Keppler, O. T., et al., Omicron subvariants illustrate reduced respiratory tissue penetration, cell damage and inflammatory responses in human airway epithelia. Frontiers in Immunology 2023, 14, 1258268. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



