Submitted:
02 September 2025
Posted:
04 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
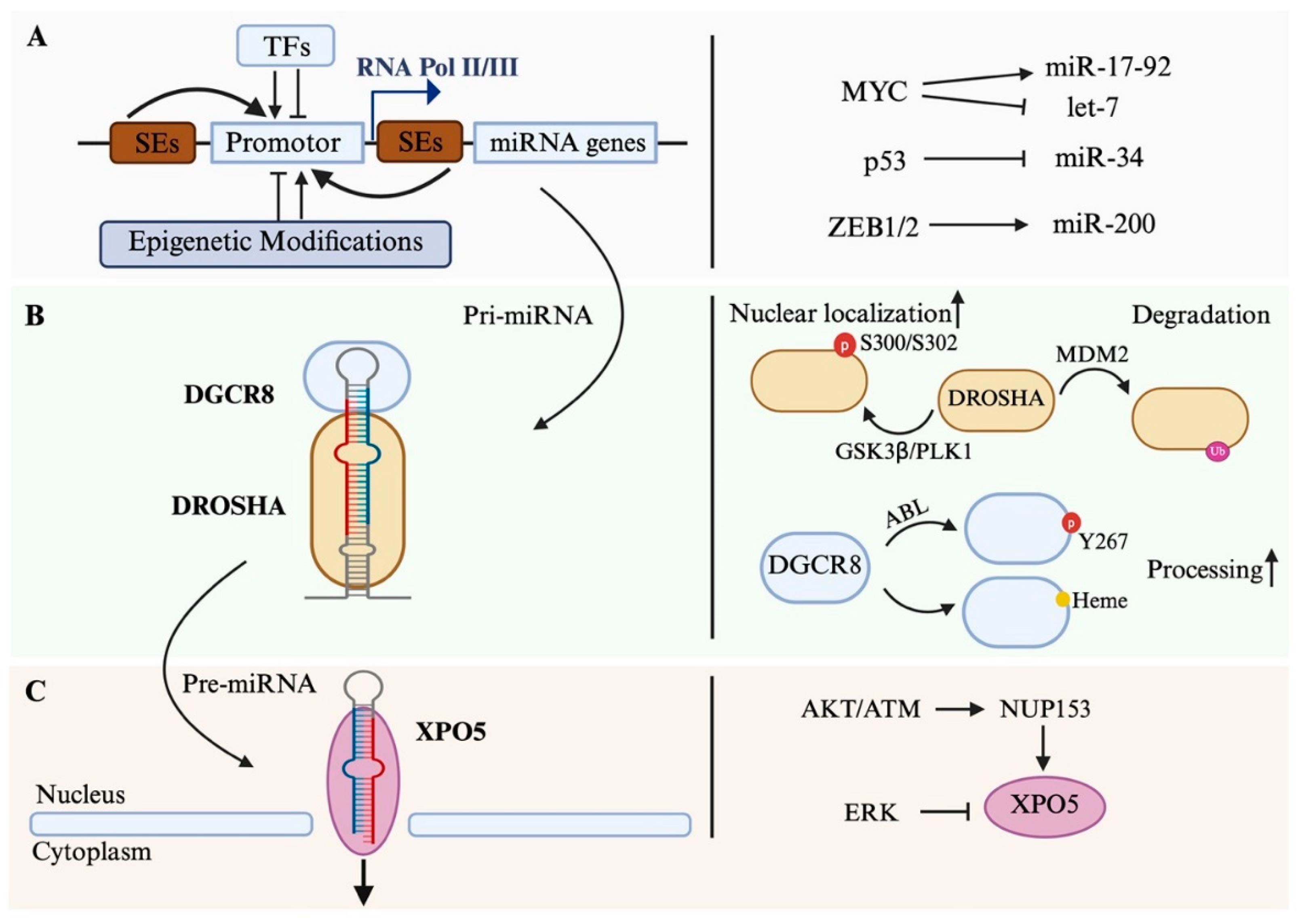
2. Transcription of MiRNA
2.1. Transcriptional Regulation
2.2. Mutations of Transcriptional Regulators
3. Processing in the Nucleus
3.1. The Microprocessor Complex
3.2. Regulation of Nuclear Processing
3.3. Nuclear Export of Pre-miRNA
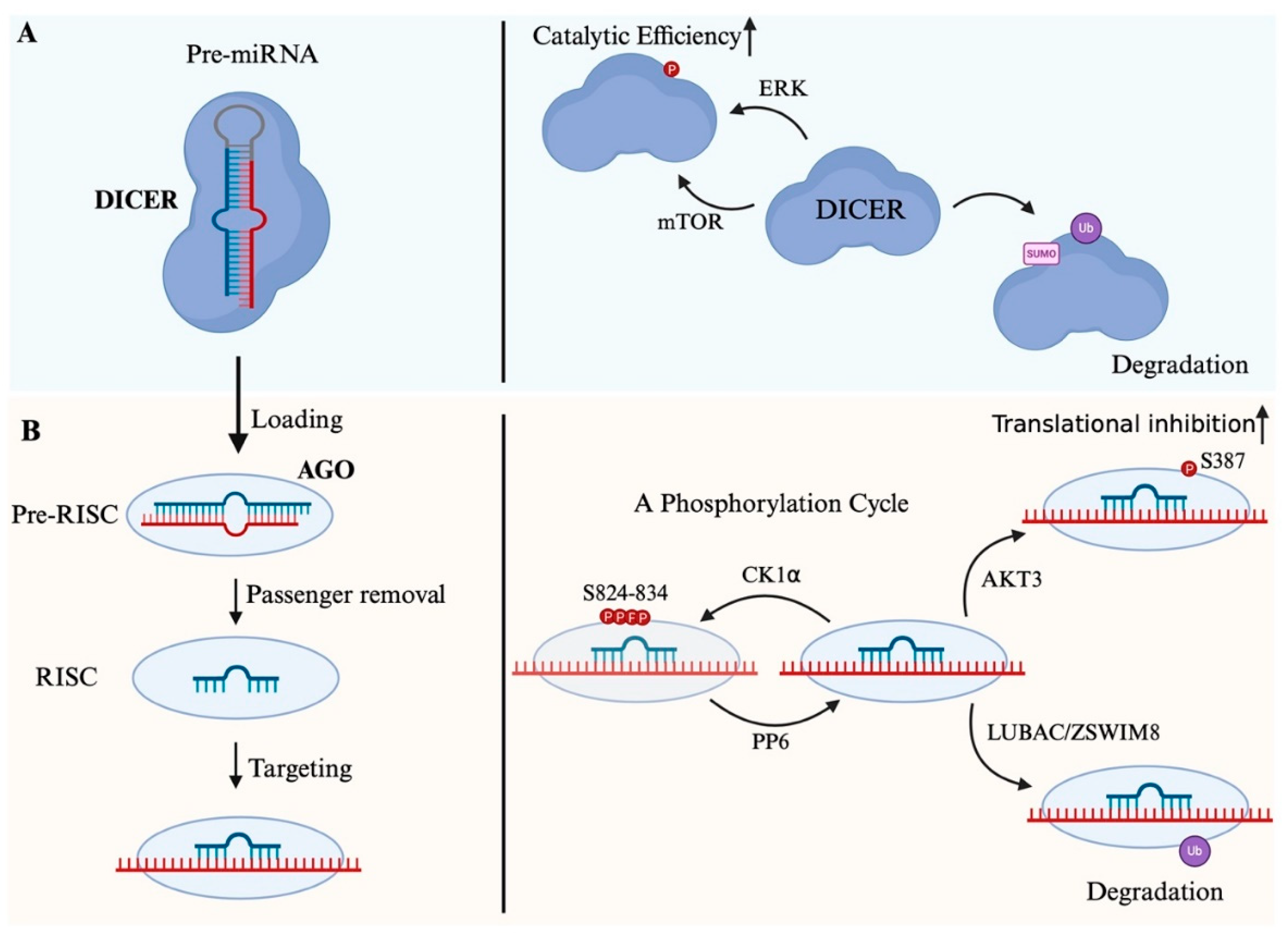
4. Processing in the Cytoplasm
4.1. DICER-Mediated Cleavage
4.2. Regulation of Cytoplasmic Processing by TRBP and PACT
4.3. MiRNA Loading and Strand Selection
5. Function of MiRNA in RISC
5.1. Canonical Function: Translational Inhibition and mRNA Destabilization
5.2. Functions in the Nucleus, Mitochondria and Condensate
5.3. Post-Translational Regulation of RISC in Diseases
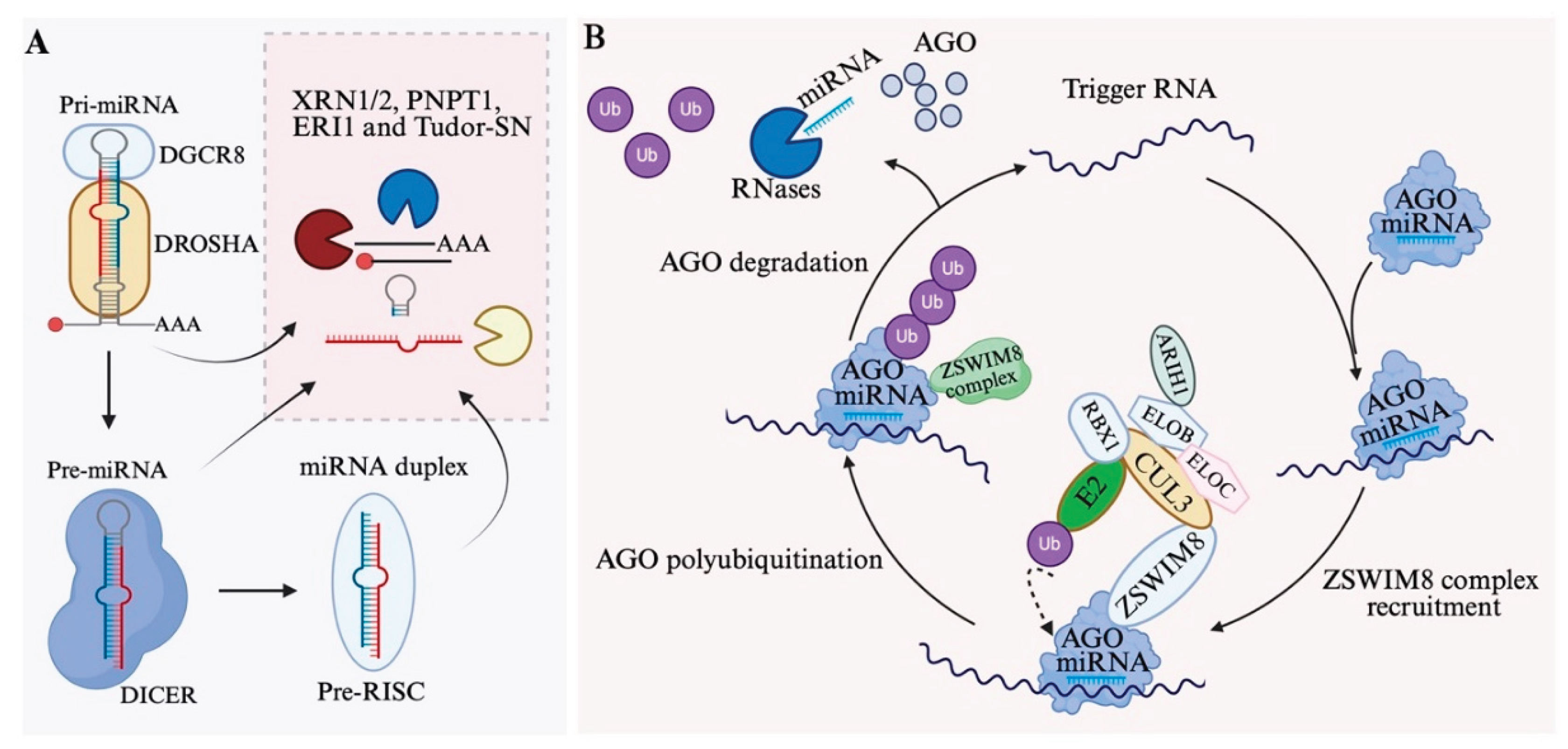
6. MiRNA Decay
6.1. Degradation of MiRNA Intermediates and Mature MiRNAs
6.2. Target-Directed miRNA Degradation
7. RNA Modifications Add a New Regulatory Layer to MiRNA Biogenesis and Function
7.1. RNA Modifications During MiRNA Biogenesis
7.2. Modifications Impacting miRNA function and stability
8. Advances in MiRNA Research and Challenges in MiRNA Therapeutics
8.1. Expansion of the Small RNA Landscape
8.2. AGO-Independent MiRNA Functions
8.3. Clinical Outlook and Challenges
9. Conclusions and Outlook

Acknowledgments
Conflicts of Interest
Abbreviations
| RNAi | RNA interference |
| TF | Transcription factors |
| ESC | Embryonic stem cell |
| SE | Super enhancers |
| EMT | Epithelial-mesenchymal transition |
| dsRBD | Double strand RNA-binding domain |
| RBP | RNA-binding protein |
| PTM | Post-translational modification |
| RISC | RNA-induced silencing complex |
| UTR | Untranslated region |
| MitomiR | Mitochondrial microRNA |
| PP6 | Protein Phosphatase 6 complex |
| TENT | Terminal nucleotidyl transferase |
| TDMD | Target-Directed miRNA Degradation |
| ROS | Reactive oxygen species |
| tsRNA | tRNA-derived small RNA |
| rsRNA | rRNA-derived small RNA |
| EV | Extracellular vesicle |
References
- Lee, R.C.; Feinbaum, R.L.; Ambros, V. The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense complementarity to lin-14. Cell 1993, 75, 843–854. [Google Scholar] [CrossRef]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000, 403, 901–906. [Google Scholar] [CrossRef]
- Almeida, M.I.; Reis, R.M.; Calin, G.A. MicroRNA history: discovery, recent applications, and next frontiers. Mutat Res 2011, 717, 1–8. [Google Scholar] [CrossRef]
- Callaway, E.; Sanderson, K. Medicine Nobel awarded for gene-regulating 'microRNAs'. Nature 2024, 634, 524–525. [Google Scholar] [CrossRef]
- Kilikevicius, A.; Meister, G.; Corey, D.R. Reexamining assumptions about miRNA-guided gene silencing. Nucleic Acids Res 2022, 50, 617–634. [Google Scholar] [CrossRef]
- Park, C.Y.; Choi, Y.S.; McManus, M.T. Analysis of microRNA knockouts in mice. Hum Mol Genet 2010, 19, R169–175. [Google Scholar] [CrossRef]
- Park, C.Y.; Jeker, L.T.; Carver-Moore, K.; Oh, A.; Liu, H.J.; Cameron, R.; Richards, H.; Li, Z.; Adler, D.; Yoshinaga, Y.; et al. A resource for the conditional ablation of microRNAs in the mouse. Cell Rep 2012, 1, 385–391. [Google Scholar] [CrossRef] [PubMed]
- Lin, S.; Gregory, R.I. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer 2015, 15, 321–333. [Google Scholar] [CrossRef] [PubMed]
- Eacker, S.M.; Dawson, T.M.; Dawson, V.L. Understanding microRNAs in neurodegeneration. Nat Rev Neurosci 2009, 10, 837–841. [Google Scholar] [CrossRef] [PubMed]
- Pauley, K.M.; Cha, S.; Chan, E.K. MicroRNA in autoimmunity and autoimmune diseases. J Autoimmun 2009, 32, 189–194. [Google Scholar] [CrossRef] [PubMed]
- Marsico, A.; Huska, M.R.; Lasserre, J.; Hu, H.; Vucicevic, D.; Musahl, A.; Orom, U.; Vingron, M. PROmiRNA: a new miRNA promoter recognition method uncovers the complex regulation of intronic miRNAs. Genome Biol 2013, 14, R84. [Google Scholar] [CrossRef] [PubMed]
- Ghorai, A.; Ghosh, U. miRNA gene counts in chromosomes vary widely in a species and biogenesis of miRNA largely depends on transcription or post-transcriptional processing of coding genes. Front Genet 2014, 5, 100. [Google Scholar] [CrossRef]
- Altuvia, Y.; Landgraf, P.; Lithwick, G.; Elefant, N.; Pfeffer, S.; Aravin, A.; Brownstein, M.J.; Tuschl, T.; Margalit, H. Clustering and conservation patterns of human microRNAs. Nucleic Acids Res 2005, 33, 2697–2706. [Google Scholar] [CrossRef] [PubMed]
- Mogilyansky, E.; Rigoutsos, I. The miR-17/92 cluster: a comprehensive update on its genomics, genetics, functions and increasingly important and numerous roles in health and disease. Cell Death Differ 2013, 20, 1603–1614. [Google Scholar] [CrossRef]
- Lee, H.; Han, S.; Kwon, C.S.; Lee, D. Biogenesis and regulation of the let-7 miRNAs and their functional implications. Protein Cell 2016, 7, 100–113. [Google Scholar] [CrossRef]
- Kim, H.; Lee, Y.Y.; Kim, V.N. The biogenesis and regulation of animal microRNAs. Nat Rev Mol Cell Biol 2025, 26, 276–296. [Google Scholar] [CrossRef]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA genes are transcribed by RNA polymerase II. EMBO J 2004, 23, 4051–4060. [Google Scholar] [CrossRef]
- Borchert, G.M.; Lanier, W.; Davidson, B.L. RNA polymerase III transcribes human microRNAs. Nat Struct Mol Biol 2006, 13, 1097–1101. [Google Scholar] [CrossRef]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Rupaimoole, R.; Calin, G.A.; Lopez-Berestein, G.; Sood, A.K. miRNA Deregulation in Cancer Cells and the Tumor Microenvironment. Cancer Discov 2016, 6, 235–246. [Google Scholar] [CrossRef]
- Kumar, M.S.; Lu, J.; Mercer, K.L.; Golub, T.R.; Jacks, T. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet 2007, 39, 673–677. [Google Scholar] [CrossRef]
- Keller, A.; Groger, L.; Tschernig, T.; Solomon, J.; Laham, O.; Schaum, N.; Wagner, V.; Kern, F.; Schmartz, G.P.; Li, Y.; et al. miRNATissueAtlas2: an update to the human miRNA tissue atlas. Nucleic Acids Res 2022, 50, D211–D221. [Google Scholar] [CrossRef]
- Ludwig, N.; Leidinger, P.; Becker, K.; Backes, C.; Fehlmann, T.; Pallasch, C.; Rheinheimer, S.; Meder, B.; Stahler, C.; Meese, E.; et al. Distribution of miRNA expression across human tissues. Nucleic Acids Res 2016, 44, 3865–3877. [Google Scholar] [CrossRef] [PubMed]
- Lagos-Quintana, M.; Rauhut, R.; Yalcin, A.; Meyer, J.; Lendeckel, W.; Tuschl, T. Identification of tissue-specific microRNAs from mouse. Curr Biol 2002, 12, 735–739. [Google Scholar] [CrossRef]
- Wang, Y.; Baskerville, S.; Shenoy, A.; Babiarz, J.E.; Baehner, L.; Blelloch, R. Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat Genet 2008, 40, 1478–1483. [Google Scholar] [CrossRef]
- Wu, D.R.; Gu, K.L.; Yu, J.C.; Fu, X.; Wang, X.W.; Guo, W.T.; Liao, L.Q.; Zhu, H.; Zhang, X.S.; Hui, J.; et al. Opposing roles of miR-294 and MBNL1/2 in shaping the gene regulatory network of embryonic stem cells. EMBO Rep 2018, 19. [Google Scholar] [CrossRef] [PubMed]
- Valdmanis, P.N.; Kim, H.K.; Chu, K.; Zhang, F.; Xu, J.; Munding, E.M.; Shen, J.; Kay, M.A. miR-122 removal in the liver activates imprinted microRNAs and enables more effective microRNA-mediated gene repression. Nat Commun 2018, 9, 5321. [Google Scholar] [CrossRef]
- Zhelankin, A.V.; Iulmetova, L.N.; Ahmetov, II; Generozov, E.V.; Sharova, E.I. Diversity and Differential Expression of MicroRNAs in the Human Skeletal Muscle with Distinct Fiber Type Composition. Life (Basel) 2023, 13. [CrossRef]
- Hobert, O. Common logic of transcription factor and microRNA action. Trends Biochem Sci 2004, 29, 462–468. [Google Scholar] [CrossRef]
- Guo, Z.; Maki, M.; Ding, R.; Yang, Y.; Zhang, B.; Xiong, L. Genome-wide survey of tissue-specific microRNA and transcription factor regulatory networks in 12 tissues. Sci Rep 2014, 4, 5150. [Google Scholar] [CrossRef]
- Chen, J.F.; Mandel, E.M.; Thomson, J.M.; Wu, Q.; Callis, T.E.; Hammond, S.M.; Conlon, F.L.; Wang, D.Z. The role of microRNA-1 and microRNA-133 in skeletal muscle proliferation and differentiation. Nat Genet 2006, 38, 228–233. [Google Scholar] [CrossRef]
- Rosenberg, M.I.; Georges, S.A.; Asawachaicharn, A.; Analau, E.; Tapscott, S.J. MyoD inhibits Fstl1 and Utrn expression by inducing transcription of miR-206. J Cell Biol 2006, 175, 77–85. [Google Scholar] [CrossRef] [PubMed]
- Wanet, A.; Tacheny, A.; Arnould, T.; Renard, P. miR-212/132 expression and functions: within and beyond the neuronal compartment. Nucleic Acids Res 2012, 40, 4742–4753. [Google Scholar] [CrossRef]
- Conaco, C.; Otto, S.; Han, J.J.; Mandel, G. Reciprocal actions of REST and a microRNA promote neuronal identity. Proc Natl Acad Sci U S A 2006, 103, 2422–2427. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Papagiannakopoulos, T.; Pan, G.; Thomson, J.A.; Kosik, K.S. MicroRNA-145 regulates OCT4, SOX2, and KLF4 and represses pluripotency in human embryonic stem cells. Cell 2009, 137, 647–658. [Google Scholar] [CrossRef]
- Divisato, G.; Passaro, F.; Russo, T.; Parisi, S. The Key Role of MicroRNAs in Self-Renewal and Differentiation of Embryonic Stem Cells. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef]
- Suzuki, H.; Maruyama, R.; Yamamoto, E.; Kai, M. Epigenetic alteration and microRNA dysregulation in cancer. Front Genet 2013, 4, 258. [Google Scholar] [CrossRef]
- Suzuki, H.I.; Young, R.A.; Sharp, P.A. Super-Enhancer-Mediated RNA Processing Revealed by Integrative MicroRNA Network Analysis. Cell 2017, 168, 1000–1014 e1015. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Rai, S.N. Predicting the Effect of miRNA on Gene Regulation to Foster Translational Multi-Omics Research-A Review on the Role of Super-Enhancers. Noncoding RNA 2024, 10. [Google Scholar] [CrossRef]
- Tan, H.; Huang, S.; Zhang, Z.; Qian, X.; Sun, P.; Zhou, X. Pan-cancer analysis on microRNA-associated gene activation. EBioMedicine 2019, 43, 82–97. [Google Scholar] [CrossRef]
- Urbanek-Trzeciak, M.O.; Galka-Marciniak, P.; Nawrocka, P.M.; Kowal, E.; Szwec, S.; Giefing, M.; Kozlowski, P. Pan-cancer analysis of somatic mutations in miRNA genes. EBioMedicine 2020, 61, 103051. [Google Scholar] [CrossRef]
- Bonci, D.; Coppola, V.; Musumeci, M.; Addario, A.; Giuffrida, R.; Memeo, L.; D'Urso, L.; Pagliuca, A.; Biffoni, M.; Labbaye, C.; et al. The miR-15a-miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nat Med 2008, 14, 1271–1277. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Choi, P.S.; Casey, S.C.; Dill, D.L.; Felsher, D.W. MYC through miR-17-92 suppresses specific target genes to maintain survival, autonomous proliferation, and a neoplastic state. Cancer Cell 2014, 26, 262–272. [Google Scholar] [CrossRef] [PubMed]
- Chang, T.C.; Yu, D.; Lee, Y.S.; Wentzel, E.A.; Arking, D.E.; West, K.M.; Dang, C.V.; Thomas-Tikhonenko, A.; Mendell, J.T. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet 2008, 40, 43–50. [Google Scholar] [CrossRef]
- Hermeking, H. p53 enters the microRNA world. Cancer Cell 2007, 12, 414–418. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Chai, B.; Li, L.; Lu, Z.; Ma, Z. p53/MicroRNA-34 axis in cancer and beyond. Heliyon 2023, 9, e15155. [Google Scholar] [CrossRef]
- Jo, H.; Shim, K.; Jeoung, D. Potential of the miR-200 Family as a Target for Developing Anti-Cancer Therapeutics. Int J Mol Sci 2022, 23. [Google Scholar] [CrossRef]
- Parreno, V.; Loubiere, V.; Schuettengruber, B.; Fritsch, L.; Rawal, C.C.; Erokhin, M.; Gyorffy, B.; Normanno, D.; Di Stefano, M.; Moreaux, J.; et al. Transient loss of Polycomb components induces an epigenetic cancer fate. Nature 2024, 629, 688–696. [Google Scholar] [CrossRef]
- Agirre, X.; Vilas-Zornoza, A.; Jimenez-Velasco, A.; Martin-Subero, J.I.; Cordeu, L.; Garate, L.; San Jose-Eneriz, E.; Abizanda, G.; Rodriguez-Otero, P.; Fortes, P.; et al. Epigenetic silencing of the tumor suppressor microRNA Hsa-miR-124a regulates CDK6 expression and confers a poor prognosis in acute lymphoblastic leukemia. Cancer Res 2009, 69, 4443–4453. [Google Scholar] [CrossRef]
- Strmsek, Z.; Kunej, T. MicroRNA Silencing by DNA Methylation in Human Cancer: a Literature Analysis. Noncoding RNA 2015, 1, 44–52. [Google Scholar] [CrossRef]
- Feinberg, A.P.; Koldobskiy, M.A.; Gondor, A. Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat Rev Genet 2016, 17, 284–299. [Google Scholar] [CrossRef] [PubMed]
- Sampath, D.; Liu, C.; Vasan, K.; Sulda, M.; Puduvalli, V.K.; Wierda, W.G.; Keating, M.J. Histone deacetylases mediate the silencing of miR-15a, miR-16, and miR-29b in chronic lymphocytic leukemia. Blood 2012, 119, 1162–1172. [Google Scholar] [CrossRef] [PubMed]
- Bollati, V.; Marinelli, B.; Apostoli, P.; Bonzini, M.; Nordio, F.; Hoxha, M.; Pegoraro, V.; Motta, V.; Tarantini, L.; Cantone, L.; et al. Exposure to metal-rich particulate matter modifies the expression of candidate microRNAs in peripheral blood leukocytes. Environ Health Perspect 2010, 118, 763–768. [Google Scholar] [CrossRef]
- Berezikov, E.; Chung, W.J.; Willis, J.; Cuppen, E.; Lai, E.C. Mammalian mirtron genes. Mol Cell 2007, 28, 328–336. [Google Scholar] [CrossRef]
- Salim, U.; Kumar, A.; Kulshreshtha, R.; Vivekanandan, P. Biogenesis, characterization, and functions of mirtrons. Wiley Interdiscip Rev RNA 2022, 13, e1680. [Google Scholar] [CrossRef]
- Havens, M.A.; Reich, A.A.; Duelli, D.M.; Hastings, M.L. Biogenesis of mammalian microRNAs by a non-canonical processing pathway. Nucleic Acids Res 2012, 40, 4626–4640. [Google Scholar] [CrossRef]
- Yi, R.; Pasolli, H.A.; Landthaler, M.; Hafner, M.; Ojo, T.; Sheridan, R.; Sander, C.; O'Carroll, D.; Stoffel, M.; Tuschl, T.; et al. DGCR8-dependent microRNA biogenesis is essential for skin development. Proc Natl Acad Sci U S A 2009, 106, 498–502. [Google Scholar] [CrossRef]
- Kwon, S.C.; Nguyen, T.A.; Choi, Y.G.; Jo, M.H.; Hohng, S.; Kim, V.N.; Woo, J.S. Structure of Human DROSHA. Cell 2016, 164, 81–90. [Google Scholar] [CrossRef]
- Partin, A.C.; Zhang, K.; Jeong, B.C.; Herrell, E.; Li, S.; Chiu, W.; Nam, Y. Cryo-EM Structures of Human Drosha and DGCR8 in Complex with Primary MicroRNA. Mol Cell 2020, 78, 411–422 e414. [Google Scholar] [CrossRef] [PubMed]
- Guil, S.; Caceres, J.F. The multifunctional RNA-binding protein hnRNP A1 is required for processing of miR-18a. Nat Struct Mol Biol 2007, 14, 591–596. [Google Scholar] [CrossRef]
- Garg, A.; Shang, R.; Cvetanovic, T.; Lai, E.C.; Joshua-Tor, L. The structural landscape of Microprocessor-mediated processing of pri-let-7 miRNAs. Mol Cell 2024, 84, 4175–4190 e4176. [Google Scholar] [CrossRef]
- Hynes, C.; Kakumani, P.K. Regulatory role of RNA-binding proteins in microRNA biogenesis. Front Mol Biosci 2024, 11, 1374843. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Pedersen, J.S.; Kwon, S.C.; Belair, C.D.; Kim, Y.K.; Yeom, K.H.; Yang, W.Y.; Haussler, D.; Blelloch, R.; Kim, V.N. Posttranscriptional crossregulation between Drosha and DGCR8. Cell 2009, 136, 75–84. [Google Scholar] [CrossRef]
- Grasso, G.; Higuchi, T.; Mac, V.; Barbier, J.; Helsmoortel, M.; Lorenzi, C.; Sanchez, G.; Bello, M.; Ritchie, W.; Sakamoto, S.; et al. NF90 modulates processing of a subset of human pri-miRNAs. Nucleic Acids Res 2020, 48, 6874–6888. [Google Scholar] [CrossRef]
- Balzeau, J.; Menezes, M.R.; Cao, S.; Hagan, J.P. The LIN28/let-7 Pathway in Cancer. Front Genet 2017, 8, 31. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Li, M.; Tucker, L.; Ramratnam, B. Glycogen synthase kinase 3 beta (GSK3beta) phosphorylates the RNAase III enzyme Drosha at S300 and S302. PLoS One 2011, 6, e20391. [Google Scholar] [CrossRef]
- Fletcher, C.E.; Taylor, M.A.; Bevan, C.L. PLK1 Regulates MicroRNA Biogenesis through Drosha Phosphorylation. Int J Mol Sci 2023, 24. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Wen, S.; Zheng, D.; Tucker, L.; Cao, L.; Pantazatos, D.; Moss, S.F.; Ramratnam, B. Acetylation of drosha on the N-terminus inhibits its degradation by ubiquitination. PLoS One 2013, 8, e72503. [Google Scholar] [CrossRef]
- Ye, P.; Liu, Y.; Chen, C.; Tang, F.; Wu, Q.; Wang, X.; Liu, C.G.; Liu, X.; Liu, R.; Liu, Y.; et al. An mTORC1-Mdm2-Drosha axis for miRNA biogenesis in response to glucose- and amino acid-deprivation. Mol Cell 2015, 57, 708–720. [Google Scholar] [CrossRef]
- Tu, C.C.; Zhong, Y.; Nguyen, L.; Tsai, A.; Sridevi, P.; Tarn, W.Y.; Wang, J.Y. The kinase ABL phosphorylates the microprocessor subunit DGCR8 to stimulate primary microRNA processing in response to DNA damage. Sci Signal 2015, 8, ra64. [Google Scholar] [CrossRef]
- Zhu, C.; Chen, C.; Huang, J.; Zhang, H.; Zhao, X.; Deng, R.; Dou, J.; Jin, H.; Chen, R.; Xu, M.; et al. SUMOylation at K707 of DGCR8 controls direct function of primary microRNA. Nucleic Acids Res 2015, 43, 7945–7960. [Google Scholar] [CrossRef]
- Quick-Cleveland, J.; Jacob, J.P.; Weitz, S.H.; Shoffner, G.; Senturia, R.; Guo, F. The DGCR8 RNA-binding heme domain recognizes primary microRNAs by clamping the hairpin. Cell Rep 2014, 7, 1994–2005. [Google Scholar] [CrossRef]
- Partin, A.C.; Ngo, T.D.; Herrell, E.; Jeong, B.C.; Hon, G.; Nam, Y. Heme enables proper positioning of Drosha and DGCR8 on primary microRNAs. Nat Commun 2017, 8, 1737. [Google Scholar] [CrossRef]
- Galka-Marciniak, P.; Urbanek-Trzeciak, M.O.; Nawrocka, P.M.; Kozlowski, P. A pan-cancer atlas of somatic mutations in miRNA biogenesis genes. Nucleic Acids Res 2021, 49, 601–620. [Google Scholar] [CrossRef] [PubMed]
- Torrezan, G.T.; Ferreira, E.N.; Nakahata, A.M.; Barros, B.D.; Castro, M.T.; Correa, B.R.; Krepischi, A.C.; Olivieri, E.H.; Cunha, I.W.; Tabori, U.; et al. Recurrent somatic mutation in DROSHA induces microRNA profile changes in Wilms tumour. Nat Commun 2014, 5, 4039. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Lee, J.E.; Riemondy, K.; Yu, Y.; Marquez, S.M.; Lai, E.C.; Yi, R. XPO5 promotes primary miRNA processing independently of RanGTP. Nat Commun 2020, 11, 1845. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Liu, F.; Yang, F.; Meng, Z.; Zeng, Y. Selectivity of Exportin 5 binding to human precursor microRNAs. RNA Biol 2021, 18, 730–737. [Google Scholar] [CrossRef]
- Zeng, Y.; Cullen, B.R. Structural requirements for pre-microRNA binding and nuclear export by Exportin 5. Nucleic Acids Res 2004, 32, 4776–4785. [Google Scholar] [CrossRef]
- Kim, Y.K.; Kim, B.; Kim, V.N. Re-evaluation of the roles of DROSHA, Export in 5, and DICER in microRNA biogenesis. Proc Natl Acad Sci U S A 2016, 113, E1881–1889. [Google Scholar] [CrossRef]
- Wan, G.; Zhang, X.; Langley, R.R.; Liu, Y.; Hu, X.; Han, C.; Peng, G.; Ellis, L.M.; Jones, S.N.; Lu, X. DNA-damage-induced nuclear export of precursor microRNAs is regulated by the ATM-AKT pathway. Cell Rep 2013, 3, 2100–2112. [Google Scholar] [CrossRef]
- Sun, H.L.; Cui, R.; Zhou, J.; Teng, K.Y.; Hsiao, Y.H.; Nakanishi, K.; Fassan, M.; Luo, Z.; Shi, G.; Tili, E.; et al. ERK Activation Globally Downregulates miRNAs through Phosphorylating Exportin-5. Cancer Cell 2016, 30, 723–736. [Google Scholar] [CrossRef]
- Wu, K.; He, J.; Pu, W.; Peng, Y. The Role of Exportin-5 in MicroRNA Biogenesis and Cancer. Genomics Proteomics Bioinformatics 2018, 16, 120–126. [Google Scholar] [CrossRef]
- Lee, Y.Y.; Lee, H.; Kim, H.; Kim, V.N.; Roh, S.H. Structure of the human DICER-pre-miRNA complex in a dicing state. Nature 2023, 615, 331–338. [Google Scholar] [CrossRef]
- Liu, Z.; Wang, J.; Cheng, H.; Ke, X.; Sun, L.; Zhang, Q.C.; Wang, H.W. Cryo-EM Structure of Human Dicer and Its Complexes with a Pre-miRNA Substrate. Cell 2018, 173, 1191–1203 e1112. [Google Scholar] [CrossRef]
- Nguyen, T.D.; Trinh, T.A.; Bao, S.; Nguyen, T.A. Secondary structure RNA elements control the cleavage activity of DICER. Nat Commun 2022, 13, 2138. [Google Scholar] [CrossRef]
- Bernstein, E.; Kim, S.Y.; Carmell, M.A.; Murchison, E.P.; Alcorn, H.; Li, M.Z.; Mills, A.A.; Elledge, S.J.; Anderson, K.V.; Hannon, G.J. Dicer is essential for mouse development. Nat Genet 2003, 35, 215–217. [Google Scholar] [CrossRef]
- Chen, J.F.; Murchison, E.P.; Tang, R.; Callis, T.E.; Tatsuguchi, M.; Deng, Z.; Rojas, M.; Hammond, S.M.; Schneider, M.D.; Selzman, C.H.; et al. Targeted deletion of Dicer in the heart leads to dilated cardiomyopathy and heart failure. Proc Natl Acad Sci U S A 2008, 105, 2111–2116. [Google Scholar] [CrossRef] [PubMed]
- Oikawa, S.; Lee, M.; Akimoto, T. Conditional Deletion of Dicer in Adult Mice Impairs Skeletal Muscle Regeneration. Int J Mol Sci 2019, 20. [Google Scholar] [CrossRef]
- De Cauwer, A.; Loustau, T.; Erne, W.; Pichot, A.; Molitor, A.; Stemmelen, T.; Carapito, R.; Orend, G.; Bahram, S.; Georgel, P. Dicer1 deficient mice exhibit premature aging and metabolic perturbations in adipocytes. iScience 2022, 25, 105149. [Google Scholar] [CrossRef] [PubMed]
- Robertson, J.C.; Jorcyk, C.L.; Oxford, J.T. DICER1 Syndrome: DICER1 Mutations in Rare Cancers. Cancers (Basel) 2018, 10. [Google Scholar] [CrossRef] [PubMed]
- Theotoki, E.I.; Pantazopoulou, V.I.; Georgiou, S.; Kakoulidis, P.; Filippa, V.; Stravopodis, D.J.; Anastasiadou, E. Dicing the Disease with Dicer: The Implications of Dicer Ribonuclease in Human Pathologies. Int J Mol Sci 2020, 21. [Google Scholar] [CrossRef] [PubMed]
- Chou, P.C.; Rajput, S.; Zhao, X.; Patel, C.; Albaciete, D.; Oh, W.J.; Daguplo, H.Q.; Patel, N.; Su, B.; Werlen, G.; et al. mTORC2 Is Involved in the Induction of RSK Phosphorylation by Serum or Nutrient Starvation. Cells 2020, 9. [Google Scholar] [CrossRef]
- Reyes-Castro, R.A.; Chen, S.Y.; Seemann, J.; Kundu, S.T.; Gibbons, D.L.; Arur, S. Phosphorylated nuclear DICER1 promotes open chromatin state and lineage plasticity of AT2 tumor cells in lung adenocarcinomas. Sci Adv 2023, 9, eadf6210. [Google Scholar] [CrossRef]
- Lai, H.H.; Li, J.N.; Wang, M.Y.; Huang, H.Y.; Croce, C.M.; Sun, H.L.; Lyu, Y.J.; Kang, J.W.; Chiu, C.F.; Hung, M.C.; et al. HIF-1alpha promotes autophagic proteolysis of Dicer and enhances tumor metastasis. J Clin Invest 2018, 128, 625–643. [Google Scholar] [CrossRef]
- Kurzynska-Kokorniak, A.; Koralewska, N.; Pokornowska, M.; Urbanowicz, A.; Tworak, A.; Mickiewicz, A.; Figlerowicz, M. The many faces of Dicer: the complexity of the mechanisms regulating Dicer gene expression and enzyme activities. Nucleic Acids Res 2015, 43, 4365–4380. [Google Scholar] [CrossRef] [PubMed]
- Wilson, R.C.; Tambe, A.; Kidwell, M.A.; Noland, C.L.; Schneider, C.P.; Doudna, J.A. Dicer-TRBP complex formation ensures accurate mammalian microRNA biogenesis. Mol Cell 2015, 57, 397–407. [Google Scholar] [CrossRef]
- Lee, Y.; Hur, I.; Park, S.Y.; Kim, Y.K.; Suh, M.R.; Kim, V.N. The role of PACT in the RNA silencing pathway. EMBO J 2006, 25, 522–532. [Google Scholar] [CrossRef]
- Lee, H.Y.; Zhou, K.; Smith, A.M.; Noland, C.L.; Doudna, J.A. Differential roles of human Dicer-binding proteins TRBP and PACT in small RNA processing. Nucleic Acids Res 2013, 41, 6568–6576. [Google Scholar] [CrossRef]
- Chendrimada, T.P.; Gregory, R.I.; Kumaraswamy, E.; Norman, J.; Cooch, N.; Nishikura, K.; Shiekhattar, R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005, 436, 740–744. [Google Scholar] [CrossRef]
- Fukunaga, R.; Han, B.W.; Hung, J.H.; Xu, J.; Weng, Z.; Zamore, P.D. Dicer partner proteins tune the length of mature miRNAs in flies and mammals. Cell 2012, 151, 533–546. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.Y.; Doudna, J.A. TRBP alters human precursor microRNA processing in vitro. RNA 2012, 18, 2012–2019. [Google Scholar] [CrossRef]
- Haase, A.D.; Jaskiewicz, L.; Zhang, H.; Laine, S.; Sack, R.; Gatignol, A.; Filipowicz, W. TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep 2005, 6, 961–967. [Google Scholar] [CrossRef]
- Heyam, A.; Lagos, D.; Plevin, M. Dissecting the roles of TRBP and PACT in double-stranded RNA recognition and processing of noncoding RNAs. Wiley Interdiscip Rev RNA 2015, 6, 271–289. [Google Scholar] [CrossRef] [PubMed]
- Rybak, A.; Fuchs, H.; Smirnova, L.; Brandt, C.; Pohl, E.E.; Nitsch, R.; Wulczyn, F.G. A feedback loop comprising lin-28 and let-7 controls pre-let-7 maturation during neural stem-cell commitment. Nat Cell Biol 2008, 10, 987–993. [Google Scholar] [CrossRef] [PubMed]
- Heo, I.; Joo, C.; Cho, J.; Ha, M.; Han, J.; Kim, V.N. Lin28 mediates the terminal uridylation of let-7 precursor MicroRNA. Mol Cell 2008, 32, 276–284. [Google Scholar] [CrossRef]
- Wu, S.L.; Fu, X.; Huang, J.; Jia, T.T.; Zong, F.Y.; Mu, S.R.; Zhu, H.; Yan, Y.; Qiu, S.; Wu, Q.; et al. Genome-wide analysis of YB-1-RNA interactions reveals a novel role of YB-1 in miRNA processing in glioblastoma multiforme. Nucleic Acids Res 2015, 43, 8516–8528. [Google Scholar] [CrossRef]
- Dueck, A.; Ziegler, C.; Eichner, A.; Berezikov, E.; Meister, G. microRNAs associated with the different human Argonaute proteins. Nucleic Acids Res 2012, 40, 9850–9862. [Google Scholar] [CrossRef]
- Nakanishi, K. Anatomy of four human Argonaute proteins. Nucleic Acids Res 2022, 50, 6618–6638. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Carmell, M.A.; Rivas, F.V.; Marsden, C.G.; Thomson, J.M.; Song, J.J.; Hammond, S.M.; Joshua-Tor, L.; Hannon, G.J. Argonaute2 is the catalytic engine of mammalian RNAi. Science 2004, 305, 1437–1441. [Google Scholar] [CrossRef]
- Medley, J.C.; Panzade, G.; Zinovyeva, A.Y. microRNA strand selection: Unwinding the rules. Wiley Interdiscip Rev RNA 2021, 12, e1627. [Google Scholar] [CrossRef]
- Schwarz, D.S.; Hutvagner, G.; Du, T.; Xu, Z.; Aronin, N.; Zamore, P.D. Asymmetry in the assembly of the RNAi enzyme complex. Cell 2003, 115, 199–208. [Google Scholar] [CrossRef]
- Frank, F.; Sonenberg, N.; Nagar, B. Structural basis for 5'-nucleotide base-specific recognition of guide RNA by human AGO2. Nature 2010, 465, 818–822. [Google Scholar] [CrossRef]
- Pinhal, D.; Goncalves, L.B.; Campos, V.F.; Patton, J.G. Decoding microRNA arm switching: a key to evolutionary innovation and gene regulation. Cell Mol Life Sci 2025, 82, 197. [Google Scholar] [CrossRef]
- Kim, H.; Kim, J.; Yu, S.; Lee, Y.Y.; Park, J.; Choi, R.J.; Yoon, S.J.; Kang, S.G.; Kim, V.N. A Mechanism for microRNA Arm Switching Regulated by Uridylation. Mol Cell 2020, 78, 1224–1236 e1225. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, K. Anatomy of RISC: how do small RNAs and chaperones activate Argonaute proteins? Wiley Interdiscip Rev RNA 2016, 7, 637–660. [Google Scholar] [CrossRef]
- Iwasaki, S.; Kobayashi, M.; Yoda, M.; Sakaguchi, Y.; Katsuma, S.; Suzuki, T.; Tomari, Y. Hsc70/Hsp90 chaperone machinery mediates ATP-dependent RISC loading of small RNA duplexes. Mol Cell 2010, 39, 292–299. [Google Scholar] [CrossRef] [PubMed]
- Ohanian, M.; Humphreys, D.T.; Anderson, E.; Preiss, T.; Fatkin, D. A heterozygous variant in the human cardiac miR-133 gene, MIR133A2, alters miRNA duplex processing and strand abundance. BMC Genet 2013, 14, 18. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Pi, J.; Zou, D.; Wang, X.; Xu, J.; Yu, S.; Zhang, T.; Li, F.; Zhang, X.; Zhao, H.; et al. microRNA arm-imbalance in part from complementary targets mediated decay promotes gastric cancer progression. Nat Commun 2019, 10, 4397. [Google Scholar] [CrossRef]
- Friedman, R.C.; Farh, K.K.; Burge, C.B.; Bartel, D.P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res 2009, 19, 92–105. [Google Scholar] [CrossRef]
- Tian, S.; Wang, J.; Zhang, F.; Wang, D. Comparative Analysis of microRNA Binding Site Distribution and microRNA-Mediated Gene Expression Repression of Oncogenes and Tumor Suppressor Genes. Genes (Basel) 2022, 13. [Google Scholar] [CrossRef]
- Vidigal, J.A. A catalytic function for mammalian Argonautes. Nat Rev Mol Cell Biol 2024, 25, 339. [Google Scholar] [CrossRef]
- Hutvagner, G.; Zamore, P.D. A microRNA in a multiple-turnover RNAi enzyme complex. Science 2002, 297, 2056–2060. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Becker, W.R.; Ober-Reynolds, B.; Jouravleva, K.; Jolly, S.M.; Zamore, P.D.; Greenleaf, W.J. High-Throughput Analysis Reveals Rules for Target RNA Binding and Cleavage by AGO2. Mol Cell 2019, 75, 741–755 e711. [Google Scholar] [CrossRef]
- Mohamed, A.A.; Wang, P.Y.; Bartel, D.P.; Vos, S.M. The structural basis for RNA slicing by human Argonaute2. Cell Rep 2025, 44, 115166. [Google Scholar] [CrossRef] [PubMed]
- Iwakawa, H.O.; Tomari, Y. Life of RISC: Formation, action, and degradation of RNA-induced silencing complex. Mol Cell 2022, 82, 30–43. [Google Scholar] [CrossRef] [PubMed]
- Fukaya, T.; Iwakawa, H.O.; Tomari, Y. MicroRNAs block assembly of eIF4F translation initiation complex in Drosophila. Mol Cell 2014, 56, 67–78. [Google Scholar] [CrossRef]
- Chen, C.Y.; Zheng, D.; Xia, Z.; Shyu, A.B. Ago-TNRC6 triggers microRNA-mediated decay by promoting two deadenylation steps. Nat Struct Mol Biol 2009, 16, 1160–1166. [Google Scholar] [CrossRef]
- Johnson, S.M.; Grosshans, H.; Shingara, J.; Byrom, M.; Jarvis, R.; Cheng, A.; Labourier, E.; Reinert, K.L.; Brown, D.; Slack, F.J. RAS is regulated by the let-7 microRNA family. Cell 2005, 120, 635–647. [Google Scholar] [CrossRef]
- Kim, H.H.; Kuwano, Y.; Srikantan, S.; Lee, E.K.; Martindale, J.L.; Gorospe, M. HuR recruits let-7/RISC to repress c-Myc expression. Genes Dev 2009, 23, 1743–1748. [Google Scholar] [CrossRef]
- Meng, F.; Henson, R.; Wehbe-Janek, H.; Ghoshal, K.; Jacob, S.T.; Patel, T. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 2007, 133, 647–658. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Wu, H.; Wu, F.; Nie, D.; Sheng, S.; Mo, Y.Y. MicroRNA-21 targets tumor suppressor genes in invasion and metastasis. Cell Res 2008, 18, 350–359. [Google Scholar] [CrossRef]
- Zheng, L.; Chen, Y.; Ye, L.; Jiao, W.; Song, H.; Mei, H.; Li, D.; Yang, F.; Li, H.; Huang, K.; et al. miRNA-584-3p inhibits gastric cancer progression by repressing Yin Yang 1- facilitated MMP-14 expression. Sci Rep 2017, 7, 8967. [Google Scholar] [CrossRef]
- Matsui, M.; Chu, Y.; Zhang, H.; Gagnon, K.T.; Shaikh, S.; Kuchimanchi, S.; Manoharan, M.; Corey, D.R.; Janowski, B.A. Promoter RNA links transcriptional regulation of inflammatory pathway genes. Nucleic Acids Res 2013, 41, 10086–10109. [Google Scholar] [CrossRef]
- Xiao, M.; Li, J.; Li, W.; Wang, Y.; Wu, F.; Xi, Y.; Zhang, L.; Ding, C.; Luo, H.; Li, Y.; et al. MicroRNAs activate gene transcription epigenetically as an enhancer trigger. RNA Biol 2017, 14, 1326–1334. [Google Scholar] [CrossRef]
- Chen, D.; Fu, L.Y.; Zhang, Z.; Li, G.; Zhang, H.; Jiang, L.; Harrison, A.P.; Shanahan, H.P.; Klukas, C.; Zhang, H.Y.; et al. Dissecting the chromatin interactome of microRNA genes. Nucleic Acids Res 2014, 42, 3028–3043. [Google Scholar] [CrossRef]
- Chu, Y.; Yokota, S.; Liu, J.; Kilikevicius, A.; Johnson, K.C.; Corey, D.R. Argonaute binding within human nuclear RNA and its impact on alternative splicing. RNA 2021, 27, 991–1003. [Google Scholar] [CrossRef] [PubMed]
- Tarallo, R.; Giurato, G.; Bruno, G.; Ravo, M.; Rizzo, F.; Salvati, A.; Ricciardi, L.; Marchese, G.; Cordella, A.; Rocco, T.; et al. The nuclear receptor ERbeta engages AGO2 in regulation of gene transcription, RNA splicing and RISC loading. Genome Biol 2017, 18, 189. [Google Scholar] [CrossRef]
- Ameyar-Zazoua, M.; Rachez, C.; Souidi, M.; Robin, P.; Fritsch, L.; Young, R.; Morozova, N.; Fenouil, R.; Descostes, N.; Andrau, J.C.; et al. Argonaute proteins couple chromatin silencing to alternative splicing. Nat Struct Mol Biol 2012, 19, 998–1004. [Google Scholar] [CrossRef]
- Guidi, R.; Wedeles, C.; Xu, D.; Kolmus, K.; Headland, S.E.; Teng, G.; Guillory, J.; Zeng, Y.J.; Cheung, T.K.; Chaudhuri, S.; et al. Argonaute3-SF3B3 complex controls pre-mRNA splicing to restrain type 2 immunity. Cell Rep 2023, 42, 113515. [Google Scholar] [CrossRef] [PubMed]
- El Fatimy, R.; Zhang, Y.; Deforzh, E.; Ramadas, M.; Saravanan, H.; Wei, Z.; Rabinovsky, R.; Teplyuk, N.M.; Uhlmann, E.J.; Krichevsky, A.M. A nuclear function for an oncogenic microRNA as a modulator of snRNA and splicing. Mol Cancer 2022, 21, 17. [Google Scholar] [CrossRef] [PubMed]
- Bandiera, S.; Ruberg, S.; Girard, M.; Cagnard, N.; Hanein, S.; Chretien, D.; Munnich, A.; Lyonnet, S.; Henrion-Caude, A. Nuclear outsourcing of RNA interference components to human mitochondria. PLoS One 2011, 6, e20746. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Song, C.; Zhou, X.; Han, X.; Li, J.; Wang, Z.; Shang, H.; Liu, Y.; Cao, H. Mitochondria Associated MicroRNA Expression Profiling of Heart Failure. Biomed Res Int 2017, 2017, 4042509. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Dai, B.; Fan, J.; Chen, C.; Nie, X.; Yin, Z.; Zhao, Y.; Zhang, X.; Wang, D.W. The Different Roles of miRNA-92a-2-5p and let-7b-5p in Mitochondrial Translation in db/db Mice. Mol Ther Nucleic Acids 2019, 17, 424–435. [Google Scholar] [CrossRef]
- Karginov, F.V.; Hannon, G.J. Remodeling of Ago2-mRNA interactions upon cellular stress reflects miRNA complementarity and correlates with altered translation rates. Genes Dev 2013, 27, 1624–1632. [Google Scholar] [CrossRef]
- Sheu-Gruttadauria, J.; MacRae, I.J. Phase Transitions in the Assembly and Function of Human miRISC. Cell 2018, 173, 946–957 e916. [Google Scholar] [CrossRef]
- Zhou, Y.; Panhale, A.; Shvedunova, M.; Balan, M.; Gomez-Auli, A.; Holz, H.; Seyfferth, J.; Helmstadter, M.; Kayser, S.; Zhao, Y.; et al. RNA damage compartmentalization by DHX9 stress granules. Cell 2024, 187, 1701–1718 e1728. [Google Scholar] [CrossRef]
- Cai, C.; Yu, J.; Zhang, X.; Zhou, T.; Chen, Q. A model for propagation of RNA structural memory through biomolecular condensates. Nat Cell Biol 2025. [Google Scholar] [CrossRef]
- Duan, Y.; Li, L.; Panzade, G.P.; Piton, A.; Zinovyeva, A.; Ambros, V. Modeling neurodevelopmental disorder-associated human AGO1 mutations in Caenorhabditis elegans Argonaute alg-1. Proc Natl Acad Sci U S A 2024, 121, e2308255121. [Google Scholar] [CrossRef]
- Schalk, A.; Cousin, M.A.; Dsouza, N.R.; Challman, T.D.; Wain, K.E.; Powis, Z.; Minks, K.; Trimouille, A.; Lasseaux, E.; Lacombe, D.; et al. De novo coding variants in the AGO1 gene cause a neurodevelopmental disorder with intellectual disability. J Med Genet 2022, 59, 965–975. [Google Scholar] [CrossRef]
- Lessel, D.; Zeitler, D.M.; Reijnders, M.R.F.; Kazantsev, A.; Hassani Nia, F.; Bartholomaus, A.; Martens, V.; Bruckmann, A.; Graus, V.; McConkie-Rosell, A.; et al. Germline AGO2 mutations impair RNA interference and human neurological development. Nat Commun 2020, 11, 5797. [Google Scholar] [CrossRef]
- Fawzy, M.S.; Toraih, E.A.; Alelwani, W.; Kattan, S.W.; Alnajeebi, A.M.; Hassan, R. The prognostic value of microRNA-biogenesis genes Argonaute 1 and 2 variants in breast cancer patients. Am J Transl Res 2020, 12, 1994–2006. [Google Scholar]
- Nowak, I.; Sarshad, A.A. Argonaute Proteins Take Center Stage in Cancers. Cancers (Basel) 2021, 13. [Google Scholar] [CrossRef] [PubMed]
- Golden, R.J.; Chen, B.; Li, T.; Braun, J.; Manjunath, H.; Chen, X.; Wu, J.; Schmid, V.; Chang, T.C.; Kopp, F.; et al. An Argonaute phosphorylation cycle promotes microRNA-mediated silencing. Nature 2017, 542, 197–202. [Google Scholar] [CrossRef] [PubMed]
- Paradis-Isler, N.; Boehm, J. NMDA receptor-dependent dephosphorylation of serine 387 in Argonaute 2 increases its degradation and affects dendritic spine density and maturation. J Biol Chem 2018, 293, 9311–9325. [Google Scholar] [CrossRef]
- Kingston, E.R.; Bartel, D.P. Global analyses of the dynamics of mammalian microRNA metabolism. Genome Res 2019, 29, 1777–1790. [Google Scholar] [CrossRef] [PubMed]
- Zangari, J.; Ilie, M.; Rouaud, F.; Signetti, L.; Ohanna, M.; Didier, R.; Romeo, B.; Goldoni, D.; Nottet, N.; Staedel, C.; et al. Rapid decay of engulfed extracellular miRNA by XRN1 exonuclease promotes transient epithelial-mesenchymal transition. Nucleic Acids Res 2017, 45, 4131–4141. [Google Scholar] [CrossRef]
- Zhang, H.; Lu, Y.; Chen, E.; Li, X.; Lv, B.; Vikis, H.G.; Liu, P. XRN2 promotes EMT and metastasis through regulating maturation of miR-10a. Oncogene 2017, 36, 3925–3933. [Google Scholar] [CrossRef]
- Das, S.K.; Sokhi, U.K.; Bhutia, S.K.; Azab, B.; Su, Z.Z.; Sarkar, D.; Fisher, P.B. Human polynucleotide phosphorylase selectively and preferentially degrades microRNA-221 in human melanoma cells. Proc Natl Acad Sci U S A 2010, 107, 11948–11953. [Google Scholar] [CrossRef]
- Thomas, M.F.; Abdul-Wajid, S.; Panduro, M.; Babiarz, J.E.; Rajaram, M.; Woodruff, P.; Lanier, L.L.; Heissmeyer, V.; Ansel, K.M. Eri1 regulates microRNA homeostasis and mouse lymphocyte development and antiviral function. Blood 2012, 120, 130–142. [Google Scholar] [CrossRef]
- Elbarbary, R.A.; Miyoshi, K.; Myers, J.R.; Du, P.; Ashton, J.M.; Tian, B.; Maquat, L.E. Tudor-SN-mediated endonucleolytic decay of human cell microRNAs promotes G(1)/S phase transition. Science 2017, 356, 859–862. [Google Scholar] [CrossRef]
- Reichholf, B.; Herzog, V.A.; Fasching, N.; Manzenreither, R.A.; Sowemimo, I.; Ameres, S.L. Time-Resolved Small RNA Sequencing Unravels the Molecular Principles of MicroRNA Homeostasis. Mol Cell 2019, 75, 756–768 e757. [Google Scholar] [CrossRef]
- Yuan, Y.R.; Pei, Y.; Ma, J.B.; Kuryavyi, V.; Zhadina, M.; Meister, G.; Chen, H.Y.; Dauter, Z.; Tuschl, T.; Patel, D.J. Crystal structure of A. aeolicus argonaute, a site-specific DNA-guided endoribonuclease, provides insights into RISC-mediated mRNA cleavage. Mol Cell 2005, 19, 405–419. [Google Scholar] [CrossRef]
- Ameres, S.L.; Horwich, M.D.; Hung, J.H.; Xu, J.; Ghildiyal, M.; Weng, Z.; Zamore, P.D. Target RNA-directed trimming and tailing of small silencing RNAs. Science 2010, 328, 1534–1539. [Google Scholar] [CrossRef] [PubMed]
- Yang, A.; Shao, T.J.; Bofill-De Ros, X.; Lian, C.; Villanueva, P.; Dai, L.; Gu, S. AGO-bound mature miRNAs are oligouridylated by TUTs and subsequently degraded by DIS3L2. Nat Commun 2020, 11, 2765. [Google Scholar] [CrossRef]
- Cazalla, D.; Yario, T.; Steitz, J.A. Down-regulation of a host microRNA by a Herpesvirus saimiri noncoding RNA. Science 2010, 328, 1563–1566. [Google Scholar] [CrossRef]
- Sheu-Gruttadauria, J.; Pawlica, P.; Klum, S.M.; Wang, S.; Yario, T.A.; Schirle Oakdale, N.T.; Steitz, J.A.; MacRae, I.J. Structural Basis for Target-Directed MicroRNA Degradation. Mol Cell 2019, 75, 1243–1255 e1247. [Google Scholar] [CrossRef] [PubMed]
- Kleaveland, B. SnapShot: Target-directed miRNA degradation. Cell 2023, 186, 5674–5674 e5671. [Google Scholar] [CrossRef]
- Shi, C.Y.; Kingston, E.R.; Kleaveland, B.; Lin, D.H.; Stubna, M.W.; Bartel, D.P. The ZSWIM8 ubiquitin ligase mediates target-directed microRNA degradation. Science 2020, 370. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; LaVigne, C.A.; Jones, B.T.; Zhang, H.; Gillett, F.; Mendell, J.T. A ubiquitin ligase mediates target-directed microRNA decay independently of tailing and trimming. Science 2020, 370. [Google Scholar] [CrossRef]
- Kleaveland, B.; Shi, C.Y.; Stefano, J.; Bartel, D.P. A Network of Noncoding Regulatory RNAs Acts in the Mammalian Brain. Cell 2018, 174, 350–362 e317. [Google Scholar] [CrossRef]
- Ghini, F.; Rubolino, C.; Climent, M.; Simeone, I.; Marzi, M.J.; Nicassio, F. Endogenous transcripts control miRNA levels and activity in mammalian cells by target-directed miRNA degradation. Nat Commun 2018, 9, 3119. [Google Scholar] [CrossRef]
- Li, L.; Sheng, P.; Li, T.; Fields, C.J.; Hiers, N.M.; Wang, Y.; Li, J.; Guardia, C.M.; Licht, J.D.; Xie, M. Widespread microRNA degradation elements in target mRNAs can assist the encoded proteins. Genes Dev 2021, 35, 1595–1609. [Google Scholar] [CrossRef]
- Shi, C.Y.; Elcavage, L.E.; Chivukula, R.R.; Stefano, J.; Kleaveland, B.; Bartel, D.P. ZSWIM8 destabilizes many murine microRNAs and is required for proper embryonic growth and development. Genome Res 2023, 33, 1482–1496. [Google Scholar] [CrossRef]
- Jones, B.T.; Han, J.; Zhang, H.; Hammer, R.E.; Evers, B.M.; Rakheja, D.; Acharya, A.; Mendell, J.T. Target-directed microRNA degradation regulates developmental microRNA expression and embryonic growth in mammals. Genes Dev 2023, 37, 661–674. [Google Scholar] [CrossRef]
- Kingston, E.R.; Blodgett, L.W.; Bartel, D.P. Endogenous transcripts direct microRNA degradation in Drosophila, and this targeted degradation is required for proper embryonic development. Mol Cell 2022, 82, 3872–3884 e3879. [Google Scholar] [CrossRef]
- Kawahara, Y.; Megraw, M.; Kreider, E.; Iizasa, H.; Valente, L.; Hatzigeorgiou, A.G.; Nishikura, K. Frequency and fate of microRNA editing in human brain. Nucleic Acids Res 2008, 36, 5270–5280. [Google Scholar] [CrossRef]
- Alarcon, C.R.; Lee, H.; Goodarzi, H.; Halberg, N.; Tavazoie, S.F. N6-methyladenosine marks primary microRNAs for processing. Nature 2015, 519, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Cheray, M.; Etcheverry, A.; Jacques, C.; Pacaud, R.; Bougras-Cartron, G.; Aubry, M.; Denoual, F.; Peterlongo, P.; Nadaradjane, A.; Briand, J.; et al. Cytosine methylation of mature microRNAs inhibits their functions and is associated with poor prognosis in glioblastoma multiforme. Mol Cancer 2020, 19, 36. [Google Scholar] [CrossRef] [PubMed]
- Pandolfini, L.; Barbieri, I.; Bannister, A.J.; Hendrick, A.; Andrews, B.; Webster, N.; Murat, P.; Mach, P.; Brandi, R.; Robson, S.C.; et al. METTL1 Promotes let-7 MicroRNA Processing via m7G Methylation. Mol Cell 2019, 74, 1278–1290 e1279. [Google Scholar] [CrossRef] [PubMed]
- Liang, H.; Jiao, Z.; Rong, W.; Qu, S.; Liao, Z.; Sun, X.; Wei, Y.; Zhao, Q.; Wang, J.; Liu, Y.; et al. 3'-Terminal 2'-O-methylation of lung cancer miR-21-5p enhances its stability and association with Argonaute 2. Nucleic Acids Res 2020, 48, 7027–7040. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Lu, R.; Huang, J.; Li, L.; Cao, Y.; Huang, C.; Chen, R.; Wang, Y.; Huang, J.; Zhao, X.; et al. N4-acetylcytidine modifies primary microRNAs for processing in cancer cells. Cell Mol Life Sci 2024, 81, 73. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.X.; Gao, J.; Ding, S.L.; Wang, K.; Jiao, J.Q.; Wang, Y.; Sun, T.; Zhou, L.Y.; Long, B.; Zhang, X.J.; et al. Oxidative Modification of miR-184 Enables It to Target Bcl-xL and Bcl-w. Mol Cell 2015, 59, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Xhemalce, B.; Robson, S.C.; Kouzarides, T. Human RNA methyltransferase BCDIN3D regulates microRNA processing. Cell 2012, 151, 278–288. [Google Scholar] [CrossRef]
- Yuan, X.; Su, Y.; Johnson, B.; Kirchner, M.; Zhang, X.; Xu, S.; Jiang, S.; Wu, J.; Shi, S.; Russo, J.J.; et al. Mass Spectrometry-Based Direct Sequencing of tRNAs De Novo and Quantitative Mapping of Multiple RNA Modifications. J Am Chem Soc 2024, 146, 25600–25613. [Google Scholar] [CrossRef]
- Zhang, X.; Cozen, A.E.; Liu, Y.; Chen, Q.; Lowe, T.M. Small RNA Modifications: Integral to Function and Disease. Trends Mol Med 2016, 22, 1025–1034. [Google Scholar] [CrossRef]
- Blow, M.J.; Grocock, R.J.; van Dongen, S.; Enright, A.J.; Dicks, E.; Futreal, P.A.; Wooster, R.; Stratton, M.R. RNA editing of human microRNAs. Genome Biol 2006, 7, R27. [Google Scholar] [CrossRef]
- Yang, W.; Chendrimada, T.P.; Wang, Q.; Higuchi, M.; Seeburg, P.H.; Shiekhattar, R.; Nishikura, K. Modulation of microRNA processing and expression through RNA editing by ADAR deaminases. Nat Struct Mol Biol 2006, 13, 13–21. [Google Scholar] [CrossRef]
- Kawahara, Y.; Zinshteyn, B.; Chendrimada, T.P.; Shiekhattar, R.; Nishikura, K. RNA editing of the microRNA-151 precursor blocks cleavage by the Dicer-TRBP complex. EMBO Rep 2007, 8, 763–769. [Google Scholar] [CrossRef]
- Ota, H.; Sakurai, M.; Gupta, R.; Valente, L.; Wulff, B.E.; Ariyoshi, K.; Iizasa, H.; Davuluri, R.V.; Nishikura, K. ADAR1 forms a complex with Dicer to promote microRNA processing and RNA-induced gene silencing. Cell 2013, 153, 575–589. [Google Scholar] [CrossRef]
- Park, Y.M.; Hwang, S.J.; Masuda, K.; Choi, K.M.; Jeong, M.R.; Nam, D.H.; Gorospe, M.; Kim, H.H. Heterogeneous nuclear ribonucleoprotein C1/C2 controls the metastatic potential of glioblastoma by regulating PDCD4. Mol Cell Biol 2012, 32, 4237–4244. [Google Scholar] [CrossRef]
- Alarcon, C.R.; Goodarzi, H.; Lee, H.; Liu, X.; Tavazoie, S.; Tavazoie, S.F. HNRNPA2B1 Is a Mediator of m(6)A-Dependent Nuclear RNA Processing Events. Cell 2015, 162, 1299–1308. [Google Scholar] [CrossRef]
- Heo, I.; Ha, M.; Lim, J.; Yoon, M.J.; Park, J.E.; Kwon, S.C.; Chang, H.; Kim, V.N. Mono-uridylation of pre-microRNA as a key step in the biogenesis of group II let-7 microRNAs. Cell 2012, 151, 521–532. [Google Scholar] [CrossRef] [PubMed]
- Salzman, D.W.; Nakamura, K.; Nallur, S.; Dookwah, M.T.; Metheetrairut, C.; Slack, F.J.; Weidhaas, J.B. miR-34 activity is modulated through 5'-end phosphorylation in response to DNA damage. Nat Commun 2016, 7, 10954. [Google Scholar] [CrossRef]
- Wang, Y.; Xu, X.; Yu, S.; Jeong, K.J.; Zhou, Z.; Han, L.; Tsang, Y.H.; Li, J.; Chen, H.; Mangala, L.S.; et al. Systematic characterization of A-to-I RNA editing hotspots in microRNAs across human cancers. Genome Res 2017, 27, 1112–1125. [Google Scholar] [CrossRef]
- Paul, D.; Sinha, A.N.; Ray, A.; Lal, M.; Nayak, S.; Sharma, A.; Mehani, B.; Mukherjee, D.; Laddha, S.V.; Suri, A.; et al. A-to-I editing in human miRNAs is enriched in seed sequence, influenced by sequence contexts and significantly hypoedited in glioblastoma multiforme. Sci Rep 2017, 7, 2466. [Google Scholar] [CrossRef]
- Choudhury, Y.; Tay, F.C.; Lam, D.H.; Sandanaraj, E.; Tang, C.; Ang, B.T.; Wang, S. Attenuated adenosine-to-inosine editing of microRNA-376a* promotes invasiveness of glioblastoma cells. J Clin Invest 2012, 122, 4059–4076. [Google Scholar] [CrossRef] [PubMed]
- Heo, I.; Joo, C.; Kim, Y.K.; Ha, M.; Yoon, M.J.; Cho, J.; Yeom, K.H.; Han, J.; Kim, V.N. TUT4 in concert with Lin28 suppresses microRNA biogenesis through pre-microRNA uridylation. Cell 2009, 138, 696–708. [Google Scholar] [CrossRef]
- Thornton, J.E.; Chang, H.M.; Piskounova, E.; Gregory, R.I. Lin28-mediated control of let-7 microRNA expression by alternative TUTases Zcchc11 (TUT4) and Zcchc6 (TUT7). RNA 2012, 18, 1875–1885. [Google Scholar] [CrossRef]
- Seok, H.; Lee, H.; Lee, S.; Ahn, S.H.; Lee, H.S.; Kim, G.D.; Peak, J.; Park, J.; Cho, Y.K.; Jeong, Y.; et al. Position-specific oxidation of miR-1 encodes cardiac hypertrophy. Nature 2020, 584, 279–285. [Google Scholar] [CrossRef] [PubMed]
- Eom, S.; Peak, J.; Park, J.; Ahn, S.H.; Cho, Y.K.; Jeong, Y.; Lee, H.S.; Lee, J.; Ignatova, E.; Lee, S.E.; et al. Widespread 8-oxoguanine modifications of miRNA seeds differentially regulate redox-dependent cancer development. Nat Cell Biol 2023, 25, 1369–1383. [Google Scholar] [CrossRef]
- Shi, J.; Zhang, Y.; Zhou, T.; Chen, Q. tsRNAs: The Swiss Army Knife for Translational Regulation. Trends Biochem Sci 2019, 44, 185–189. [Google Scholar] [CrossRef]
- Shi, J.; Zhou, T.; Chen, Q. Exploring the expanding universe of small RNAs. Nat Cell Biol 2022, 24, 415–423. [Google Scholar] [CrossRef]
- Shi, J.; Zhang, Y.; Li, Y.; Zhang, L.; Zhang, X.; Yan, M.; Chen, Q.; Zhang, Y. Optimized identification and characterization of small RNAs with PANDORA-seq. Nat Protoc 2025. [Google Scholar] [CrossRef]
- Shi, J.; Zhang, Y.; Tan, D.; Zhang, X.; Yan, M.; Zhang, Y.; Franklin, R.; Shahbazi, M.; Mackinlay, K.; Liu, S.; et al. PANDORA-seq expands the repertoire of regulatory small RNAs by overcoming RNA modifications. Nat Cell Biol 2021, 23, 424–436. [Google Scholar] [CrossRef]
- Hernandez, R.; Shi, J.; Liu, J.; Li, X.; Wu, J.; Zhao, L.; Zhou, T.; Chen, Q.; Zhou, C. PANDORA-Seq unveils the hidden small noncoding RNA landscape in atherosclerosis of LDL receptor-deficient mice. J Lipid Res 2023, 64, 100352. [Google Scholar] [CrossRef] [PubMed]
- Lai, H.; Feng, N.; Zhai, Q. Discovery of the major 15-30 nt mammalian small RNAs, their biogenesis and function. Nat Commun 2023, 14, 5796. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Zhang, Z.; O'Loughlin, E.; Lee, T.; Houel, S.; O'Carroll, D.; Tarakhovsky, A.; Ahn, N.G.; Yi, R. Quantitative functions of Argonaute proteins in mammalian development. Genes Dev 2012, 26, 693–704. [Google Scholar] [CrossRef]
- Bissels, U.; Wild, S.; Tomiuk, S.; Holste, A.; Hafner, M.; Tuschl, T.; Bosio, A. Absolute quantification of microRNAs by using a universal reference. RNA 2009, 15, 2375–2384. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Shi, J.; Zhang, Y.; Zhang, H.; Liao, S.; Li, W.; Lei, L.; Han, C.; Ning, L.; Cao, Y.; et al. A novel class of tRNA-derived small RNAs extremely enriched in mature mouse sperm. Cell Res 2012, 22, 1609–1612. [Google Scholar] [CrossRef]
- Chen, Q.; Yan, M.; Cao, Z.; Li, X.; Zhang, Y.; Shi, J.; Feng, G.H.; Peng, H.; Zhang, X.; Zhang, Y.; et al. Sperm tsRNAs contribute to intergenerational inheritance of an acquired metabolic disorder. Science 2016, 351, 397–400. [Google Scholar] [CrossRef]
- Chen, Q.; Yan, W.; Duan, E. Epigenetic inheritance of acquired traits through sperm RNAs and sperm RNA modifications. Nat Rev Genet 2016, 17, 733–743. [Google Scholar] [CrossRef]
- Tomar, A.; Gomez-Velazquez, M.; Gerlini, R.; Comas-Armangue, G.; Makharadze, L.; Kolbe, T.; Boersma, A.; Dahlhoff, M.; Burgstaller, J.P.; Lassi, M.; et al. Epigenetic inheritance of diet-induced and sperm-borne mitochondrial RNAs. Nature 2024, 630, 720–727. [Google Scholar] [CrossRef]
- Cai, C.; Chen, Q. Father's diet influences son's metabolic health through sperm RNA. Nature 2024, 630, 571–573. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, X.; Shi, J.; Tuorto, F.; Li, X.; Liu, Y.; Liebers, R.; Zhang, L.; Qu, Y.; Qian, J.; et al. Dnmt2 mediates intergenerational transmission of paternally acquired metabolic disorders through sperm small non-coding RNAs. Nat Cell Biol 2018, 20, 535–540. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Zhang, X.; Shi, J.; Yan, M.; Zhou, T. Origins and evolving functionalities of tRNA-derived small RNAs. Trends Biochem Sci 2021, 46, 790–804. [Google Scholar] [CrossRef] [PubMed]
- Gu, W.; Shi, J.; Liu, H.; Zhang, X.; Zhou, J.J.; Li, M.; Zhou, D.; Li, R.; Lv, J.; Wen, G.; et al. Peripheral blood non-canonical small non-coding RNAs as novel biomarkers in lung cancer. Mol Cancer 2020, 19, 159. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Trebak, F.; Souza, L.A.C.; Shi, J.; Zhou, T.; Kehoe, P.G.; Chen, Q.; Feng Earley, Y. Small RNA modifications in Alzheimer's disease. Neurobiol Dis 2020, 145, 105058. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Y.; Shi, J.; Zhang, H.; Cao, Z.; Gao, X.; Ren, W.; Ning, Y.; Ning, L.; Cao, Y.; et al. Identification and characterization of an ancient class of small RNAs enriched in serum associating with active infection. J Mol Cell Biol 2014, 6, 172–174. [Google Scholar] [CrossRef]
- Zhang, Y.; Shi, J.; Chen, Q. tsRNAs: new players in mammalian retrotransposon control. Cell Res 2017, 27, 1307–1308. [Google Scholar] [CrossRef]
- Eiring, A.M.; Harb, J.G.; Neviani, P.; Garton, C.; Oaks, J.J.; Spizzo, R.; Liu, S.; Schwind, S.; Santhanam, R.; Hickey, C.J.; et al. miR-328 functions as an RNA decoy to modulate hnRNP E2 regulation of mRNA translation in leukemic blasts. Cell 2010, 140, 652–665. [Google Scholar] [CrossRef]
- Lehmann, S.M.; Kruger, C.; Park, B.; Derkow, K.; Rosenberger, K.; Baumgart, J.; Trimbuch, T.; Eom, G.; Hinz, M.; Kaul, D.; et al. An unconventional role for miRNA: let-7 activates Toll-like receptor 7 and causes neurodegeneration. Nat Neurosci 2012, 15, 827–835. [Google Scholar] [CrossRef]
- Wang, Y.; Liang, H.; Jin, F.; Yan, X.; Xu, G.; Hu, H.; Liang, G.; Zhan, S.; Hu, X.; Zhao, Q.; et al. Injured liver-released miRNA-122 elicits acute pulmonary inflammation via activating alveolar macrophage TLR7 signaling pathway. Proc Natl Acad Sci U S A 2019, 116, 6162–6171. [Google Scholar] [CrossRef]
- Park, C.K.; Xu, Z.Z.; Berta, T.; Han, Q.; Chen, G.; Liu, X.J.; Ji, R.R. Extracellular microRNAs activate nociceptor neurons to elicit pain via TLR7 and TRPA1. Neuron 2014, 82, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Zhang, X.; Cai, C.; Zhou, T.; Chen, Q. Small RNA and Toll-like receptor interactions: origins and disease mechanisms. Trends Biochem Sci 2025, 50, 385–401. [Google Scholar] [CrossRef]
- Han, Q.; Liu, D.; Convertino, M.; Wang, Z.; Jiang, C.; Kim, Y.H.; Luo, X.; Zhang, X.; Nackley, A.; Dokholyan, N.V.; et al. miRNA-711 Binds and Activates TRPA1 Extracellularly to Evoke Acute and Chronic Pruritus. Neuron 2018, 99, 449–463 e446. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Wan, X.; Dennis, A.T.; Bektik, E.; Wang, Z.; Costa, M.G.S.; Fagnen, C.; Venien-Bryan, C.; Xu, X.; Gratz, D.H.; et al. MicroRNA Biophysically Modulates Cardiac Action Potential by Direct Binding to Ion Channel. Circulation 2021, 143, 1597–1613. [Google Scholar] [CrossRef]
- Seyhan, A.A. Trials and Tribulations of MicroRNA Therapeutics. Int J Mol Sci 2024, 25. [Google Scholar] [CrossRef]
- What will it take to get miRNA therapies to market? Nat Biotechnol 2024, 42, 1623–1624. [CrossRef]
- Segal, M.; Slack, F.J. Challenges identifying efficacious miRNA therapeutics for cancer. Expert Opin Drug Discov 2020, 15, 987–992. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Zhu, L.; Wang, X.; Jin, H. RNA-based therapeutics: an overview and prospectus. Cell Death Dis 2022, 13, 644. [Google Scholar] [CrossRef] [PubMed]
- Lockhart, J.; Canfield, J.; Mong, E.F.; VanWye, J.; Totary-Jain, H. Nucleotide Modification Alters MicroRNA-Dependent Silencing of MicroRNA Switches. Mol Ther Nucleic Acids 2019, 14, 339–350. [Google Scholar] [CrossRef] [PubMed]
- Groot, M.; Lee, H. Sorting Mechanisms for MicroRNAs into Extracellular Vesicles and Their Associated Diseases. Cells 2020, 9. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Wartmann, T.; Accuffi, S.; Al-Madhi, S.; Perrakis, A.; Kahlert, C.; Link, A.; Venerito, M.; Keitel-Anselmino, V.; Bruns, C.; et al. Integrating a microRNA signature as a liquid biopsy-based tool for the early diagnosis and prediction of potential therapeutic targets in pancreatic cancer. Br J Cancer 2024, 130, 125–134. [Google Scholar] [CrossRef]
- Metcalf, G.A.D. MicroRNAs: circulating biomarkers for the early detection of imperceptible cancers via biosensor and machine-learning advances. Oncogene 2024, 43, 2135–2142. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).



