Submitted:
03 September 2025
Posted:
03 September 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
2.2. Figures, Tables and Schemes
| Accuracy Dermatophyte detection in Nail samples | Accuracy Dermatophyte detection in Skin samples | Accuracy Dermatophyte detection in Hair samples | ||||
| Spike | Count of Correct | Percent of Correct | Count of Correct | Percent of Correct | Count of Correct | Percent of Correct |
| T. interdigitale | 27 | 100.0% | 27 | 100% | - | - |
| T. rubrum | 27 | 100.0% | - | - | - | - |
| C.albicans | 27 | 100.0% | - | - | - | - |
| C.parapsilosis | 27 | 100.0% | - | - | - | - |
| C.guillermondii | 27 | 100.0% | - | - | - | - |
| E.floccosum | 27 | 100.0% | 27 | 100% | - | - |
| S.brevicaulis | 27 | 100.0% | - | - | - | - |
| T.menta inter | - | - | 21 | 91.3% | - | - |
| T.equinum | - | - | 21 | 91.3% | - | - |
| T.violaceum | - | - | 27 | 100.0% | 24 | 96.0% |
| T. menta quinkeanum | - | - | 24 | 96.0% | - | - |
| M.canis | - | - | 27 | 100.0% | 18 | 100.0% |
| N.persicolor | - | - | 27 | 100.0% | 9 | 100.0% |
| T.schoenleinii | - | - | 27 | 100.0% | - | - |
| T.tonsurans | - | - | - | - | 27 | 100.0% |
| M.audouinii | - | - | - | - | 18 | 85.7% |
| Sensitivity | 189 | 100% | 228 | 97.9% | 96 | 96.3% |
| Sample Name | Hair | |||
| Expected | Detected with Dermatophytes | |||
| Not detetced with Y/M | ||||
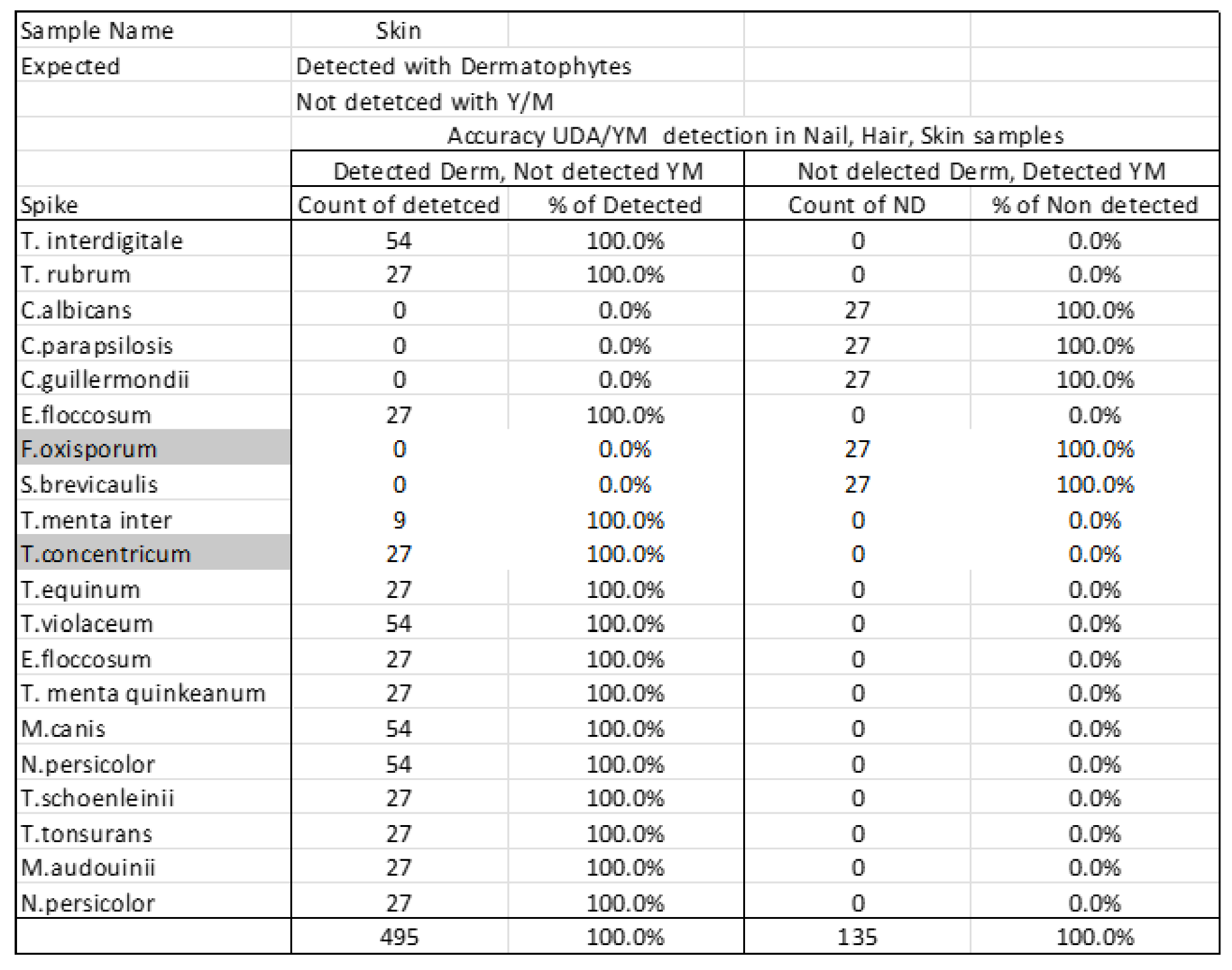
| Accuracy UDA/YM detection in Nail, Hair, Skin samples | ||||
| Detected Derm, Not detected YM | Not delected Derm, Detected YM | |||
| Spike | Count of detetced | % of Detected | Count of ND | % of Non detected |
| T. interdigitale | 54 | 100.0% | 0 | 0.0% |
| T. rubrum | 54 | 100.0% | 0 | 0.0% |
| C.albicans | 0 | 0.0% | 27 | 100.0% |
| C.parapsilosis | 0 | 0.0% | 27 | 100.0% |
| C.guillermondii | 0 | 0.0% | 27 | 100.0% |
| E.floccosum | 27 | 100.0% | 0 | 0.0% |
| F.oxisporum | 0 | 0.0% | 27 | 100.0% |
| S.brevicaulis | 0 | 0.0% | 27 | 100.0% |
| T.menta inter | 27 | 100.0% | 0 | 0.0% |
| T.concentricum | 27 | 100.0% | 0 | 0.0% |
| T.equinum | 27 | 100.0% | 0 | 0.0% |
| T.violaceum | 54 | 100.0% | 0 | 0.0% |
| E.floccosum | 27 | 100.0% | 0 | 0.0% |
| T. menta quinkeanum | 27 | 100.0% | 0 | 0.0% |
| M.canis | 54 | 100.0% | 0 | 0.0% |
| N.persicolor | 54 | 100.0% | 0 | 0.0% |
| T.schoenleinii | 27 | 100.0% | 0 | 0.0% |
| T.tonsurans | 27 | 100.0% | 0 | 0.0% |
| M.audouinii | 27 | 100.0% | 0 | 0.0% |
| N.persicolor | 27 | 100.0% | 0 | 0.0% |
| 540 | 100.0% | 135 | 100.0% | |
| Target Organism | Total Concordant | Total Discordant | Reproducibility | |
| Nail | T. interdigitale | 207 | 0 | 100.0% |
| T. rubrum | 207 | 0 | 100.0% | |
| C.albicans | 207 | 0 | 100.0% | |
| C.parapsilosis | 207 | 0 | 100.0% | |
| C.guillermondii | 207 | 0 | 100.0% | |
| E.floccosum | 207 | 0 | 100.0% | |
| F.oxisporum | 23 | 184 | 11.1% | |
| S.brevicaulis | 207 | 0 | 100.0% | |
| Skin | T.interdigitale | 207 | 0 | 100.0% |
| T.menta inter | 161 | 46 | 77.8% | |
| T.concentricum | 0 | 207 | 0.0% | |
| T.equinum | 161 | 46 | 77.8% | |
| T.violaceum | 207 | 0 | 100.0% | |
| E.floccosum | 207 | 0 | 100.0% | |
| T. menta quinkeanum | 184 | 23 | 88.9% | |
| M.canis | 207 | 0 | 100.0% | |
| N.persicolor | 207 | 0 | 100.0% | |
| T.schoenleinii | 207 | 0 | 100.0% | |
| Hair | T.tonsurans | 207 | 0 | 100.0% |
| T.violaceum | 184 | 23 | 88.9% | |
| M.audouinii | 138 | 69 | 66.7% | |
| M.canis | 207 | 0 | 100.0% | |
| N.persicolor | 207 | 0 | 100.0% |
| LOD Dermatophyte detection in Nail samples | ||||||||||
| 1,000 | 750 | 100 | 10 | 1 | 1,000 | 750 | 100 | 10 | ||
| 1.00E+03 | 7.50E+02 | 1.00E+02 | 1.00E+01 | 1.00E+00 | Average Intensity |
1.00E+03 | 7.50E+02 | 1.00E+02 | 1.00E+01 | |
| Spike | Count of Detected | Percent of Detected | ||||||||
| T. interdigitale | 3 | 3 | 3 | 3 | 3 | 9,482 | 100.0% | 100.0% | 100.0% | 100.0% |
| T. rubrum | 3 | 3 | 3 | 3 | 3 | 57,330 | 100.0% | 100.0% | 100.0% | 100.0% |
| C.albicans | 3 | 3 | 1 | 0 | 0 | 3,690 | 100.0% | 100.0% | 33.3% | 0.0% |
| C.parapsilosis | 3 | 3 | 3 | 1 | 1 | 48,884 | 100.0% | 100.0% | 100.0% | 33.3% |
| C.guillermondii | 3 | 3 | 3 | 3 | 3 | 14,915 | 100.0% | 100.0% | 100.0% | 100.0% |
| E.floccosum | 3 | 3 | 3 | 3 | 3 | 29,880 | 100.0% | 100.0% | 100.0% | 100.0% |
| F.oxisporum | 3 | 3 | 3 | 3 | 3 | 6,471 | 100.0% | 100.0% | 100.0% | 100.0% |
| S.brevicaulis | 3 | 3 | 3 | 3 | 3 | 28,481 | 100.0% | 100.0% | 100.0% | 100.0% |
| Universal Dermatophyte | 3 | 3 | 3 | 3 | 3 | 100.0% | 100.0% | 100.0% | 100.0% | |
| LOD Dermatophyte detection in Skin samples | ||||||||||
| 1,000 | 750 | 100 | 10 | 1 | 1,000 | 750 | 100 | 10 | ||
| 1.00E+03 | 7.50E+02 | 1.00E+02 | 1.00E+01 | 1.00E+00 | Average Intensity |
1.00E+03 | 7.50E+02 | 1.00E+02 | 1.00E+01 | |
| Spike | Count of Detected | Percent of Detected | ||||||||
| T.interdigitale | 4 | 4 | 4 | 4 | 2 | 5,010 | 100.0% | 100.0% | 100.0% | 100.0% |
| T.menta inter | 1 | 1 | 1 | 1 | 1 | 0 | 25.0% | 25.0% | 25.0% | 25.0% |
| T.concentricum | 0 | 0 | 0 | 0 | 0 | 0 | 0.0% | 0.0% | 0.0% | 0.0% |
| T.equinum | 2 | 1 | 0 | 0 | 0 | 1,378 | 50.0% | 25.0% | 0.0% | 0.0% |
| T.violaceum | 4 | 4 | 3 | 0 | 0 | 1,794 | 100.0% | 100.0% | 75.0% | 0.0% |
| E.floccosum | 4 | 4 | 4 | 3 | 3 | 7,544 | 100.0% | 100.0% | 100.0% | 75.0% |
| T. menta quinkeanum | 4 | 4 | 4 | 1 | 0 | 10,373 | 100.0% | 100.0% | 100.0% | 25.0% |
| M.canis | 3 | 2 | 2 | 2 | 0 | 2,714 | 75.0% | 50.0% | 50.0% | 50.0% |
| N.persicolor | 4 | 4 | 4 | 4 | 4 | 33,662 | 100.0% | 100.0% | 100.0% | 100.0% |
| T.schoenleinii | 4 | 4 | 4 | 3 | 0 | 18,366 | 100.0% | 100.0% | 100.0% | 75.0% |
| Universal Dermatophyte | 4 | 4 | 4 | 4 | 4 | 100.0% | 100.0% | 100.0% | 100.0% | |
| LOD Dermatophyte detection in Hair samples | ||||||||||
| 1,000 | 750 | 100 | 10 | 1 | 1,000 | 750 | 100 | 10 | ||
| 1.00E+03 | 7.50E+02 | 1.00E+02 | 1.00E+01 | 1.00E+00 | Average Intensity |
1.00E+03 | 7.50E+02 | 1.00E+02 | 1.00E+01 | |
| Spike | Count of Detected | Percent of Detected | ||||||||
| T.tonsurans | 4 | 4 | 4 | 4 | 0 | 5,010 | 100.0% | 100.0% | 100.0% | 100.0% |
| T.violaceum | 4 | 4 | 4 | 4 | 2 | 0 | 100.0% | 100.0% | 100.0% | 100.0% |
| M.audouinii | 4 | 4 | 4 | 4 | 0 | 0 | 100.0% | 100.0% | 100.0% | 100.0% |
| M.canis | 3 | 2 | 0 | 0 | 0 | 1,378 | 75.0% | 50.0% | 0.0% | 0.0% |
| N.persicolor | 4 | 4 | 4 | 4 | 4 | 1,794 | 100.0% | 100.0% | 100.0% | 100.0% |
| Universal Dermatophyte | 4 | 4 | 4 | 4 | 4 | 100.0% | 100.0% | 100.0% | 100.0% | |
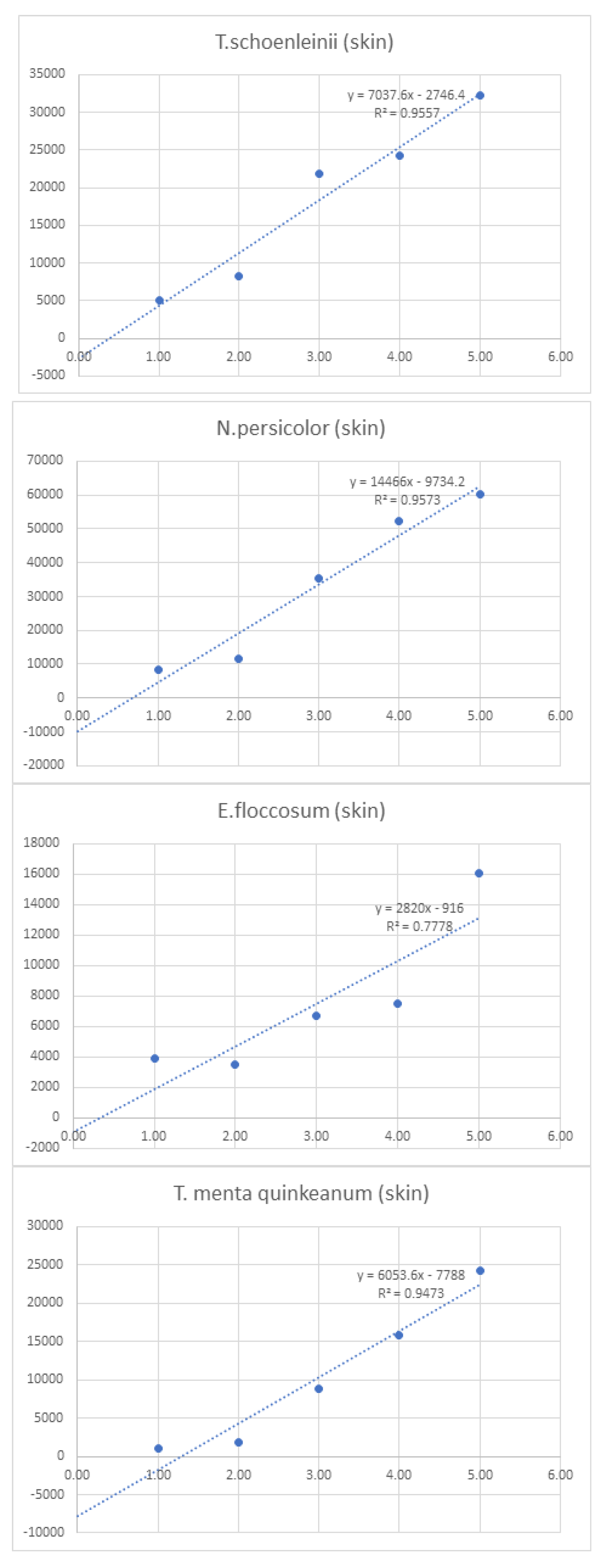
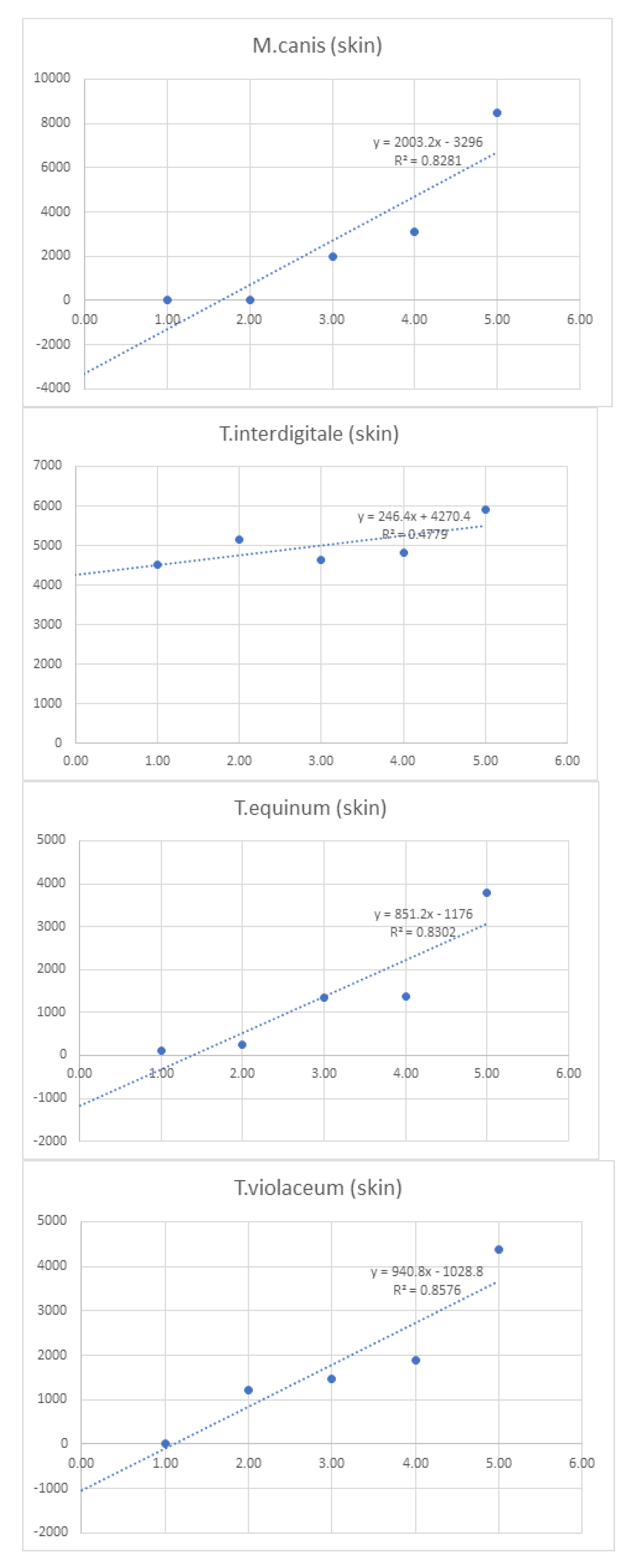
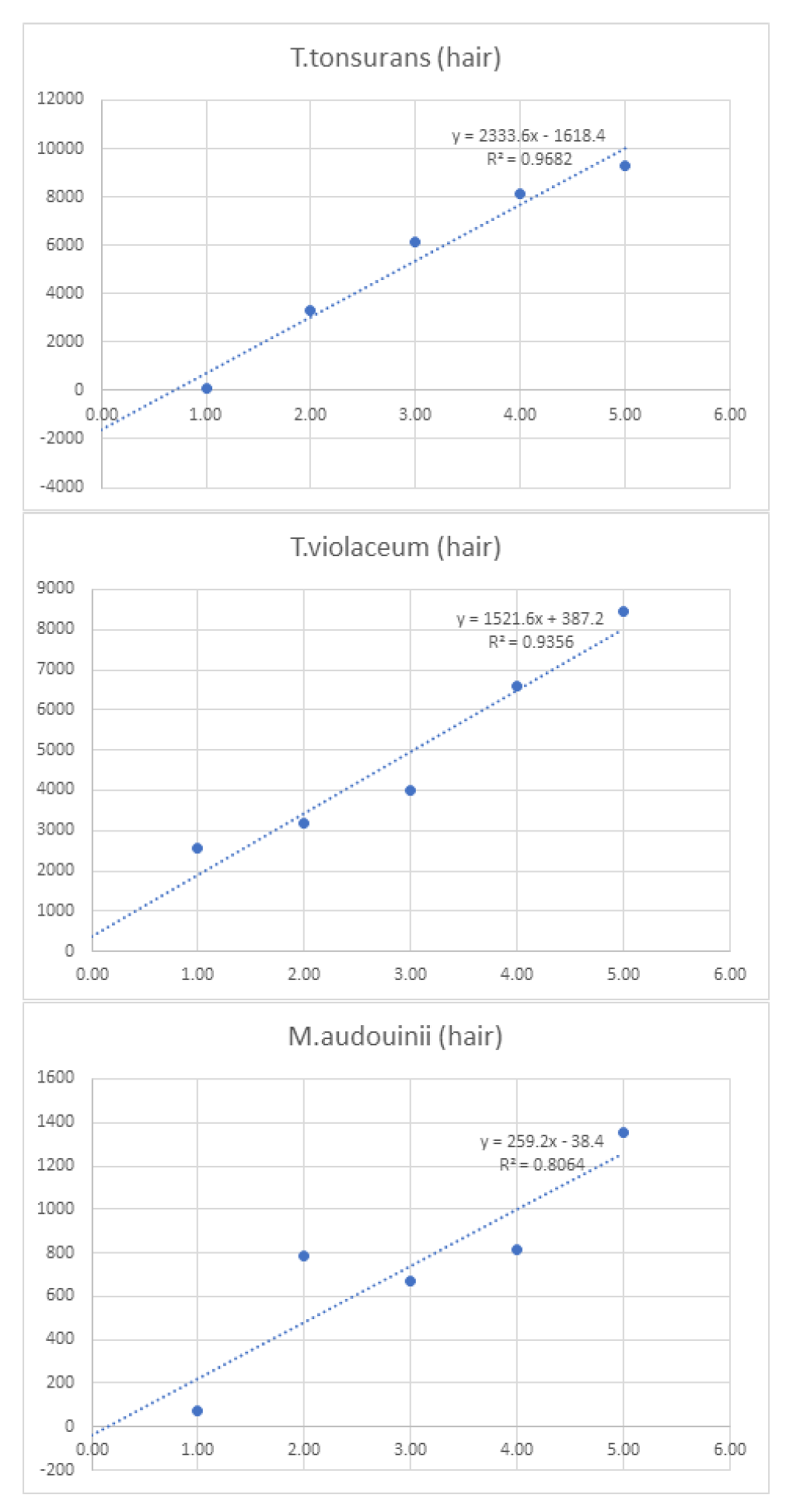
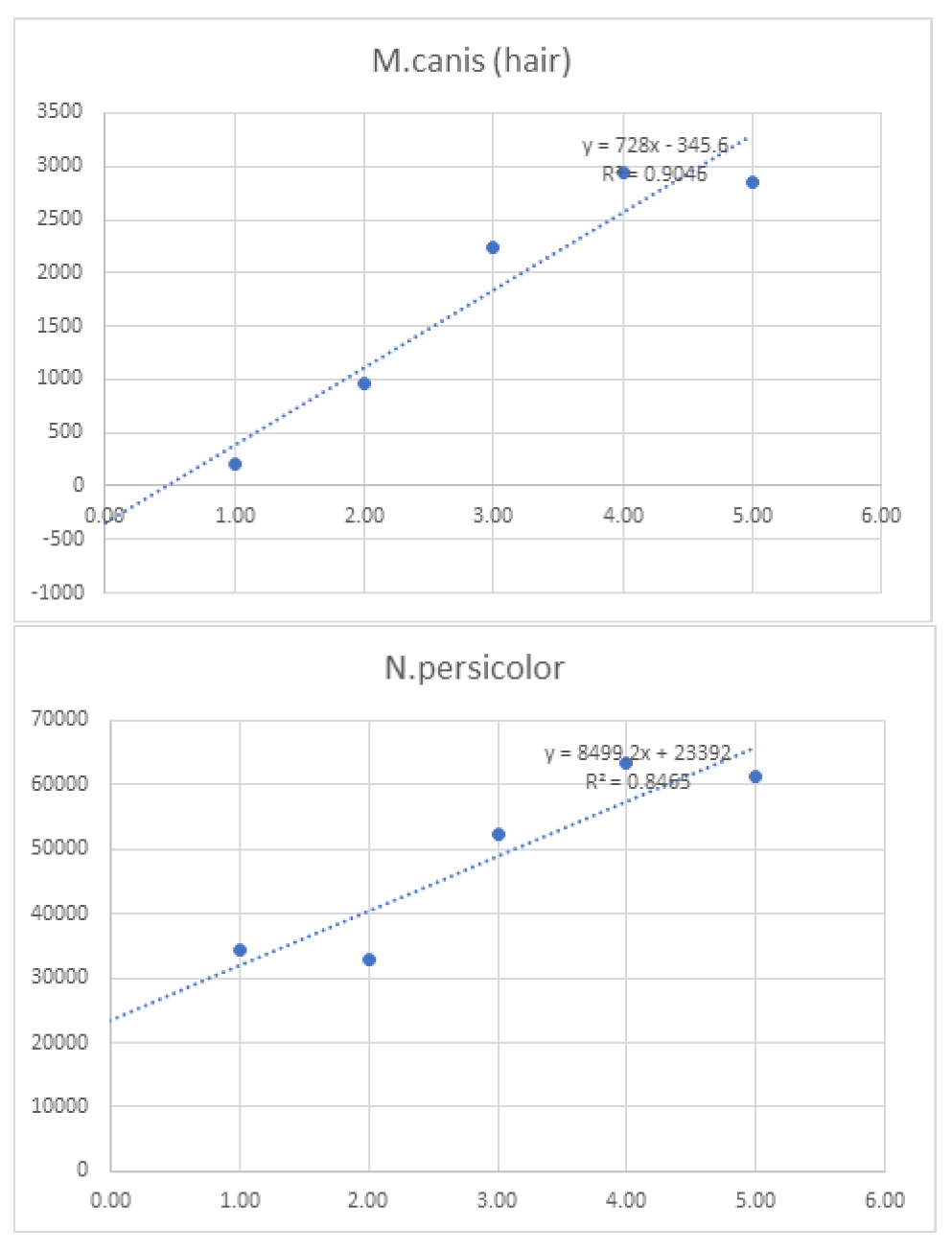
| Linearity | |||||||||
| 1 | 10 | 100 | 750 | 1,000 | 10,000 | ||||
| Log10 Concentration | 0.00 | 1.00 | 2.00 | 3.00 | 4.00 | 5.00 | Sum | ||
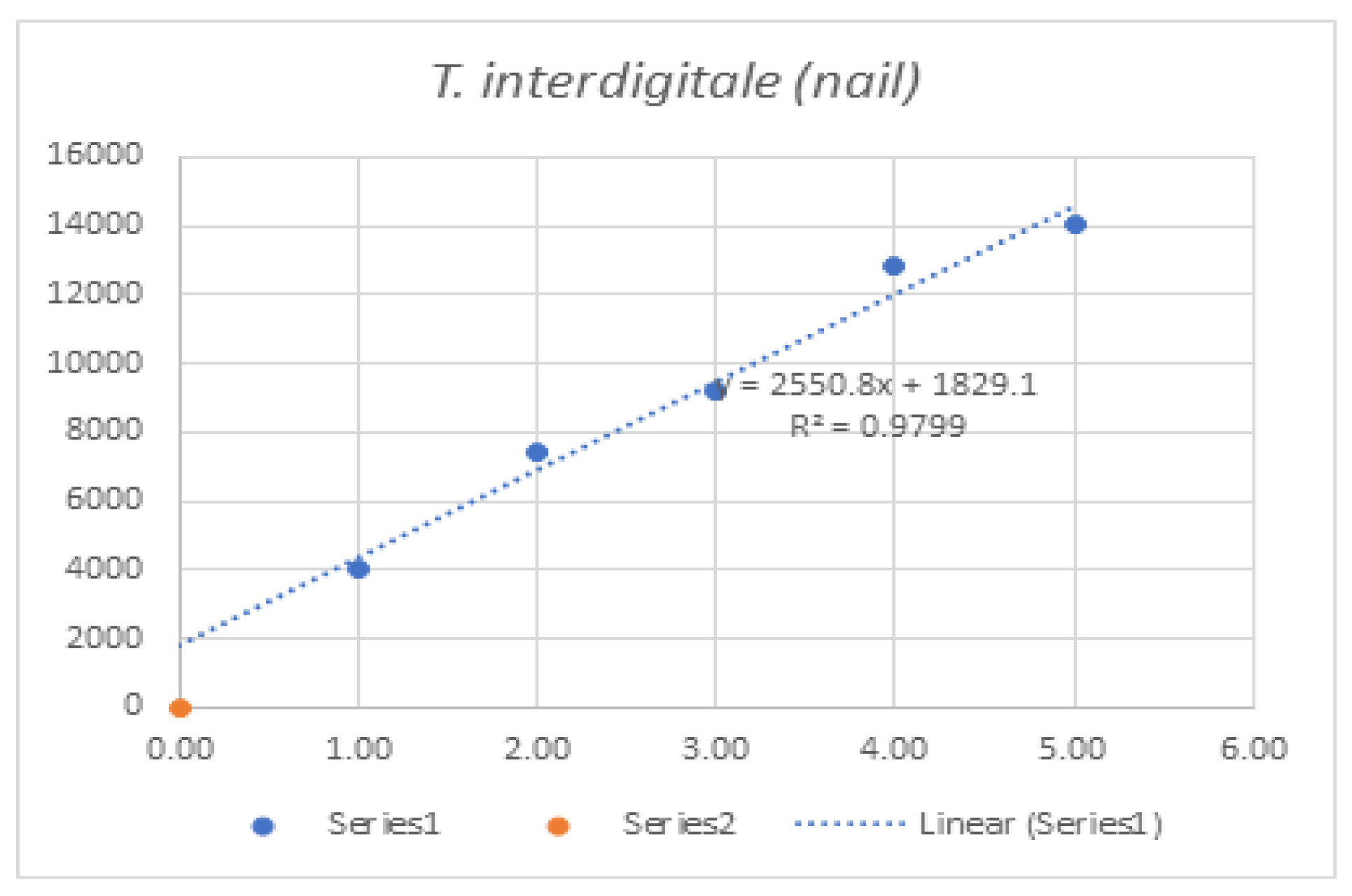
| Mean of Intensity | Nail | T. interdigitale (nail) | - | 3988 | 7383.75 | 9200 | 12804 | 14032 | 47,407.8 |
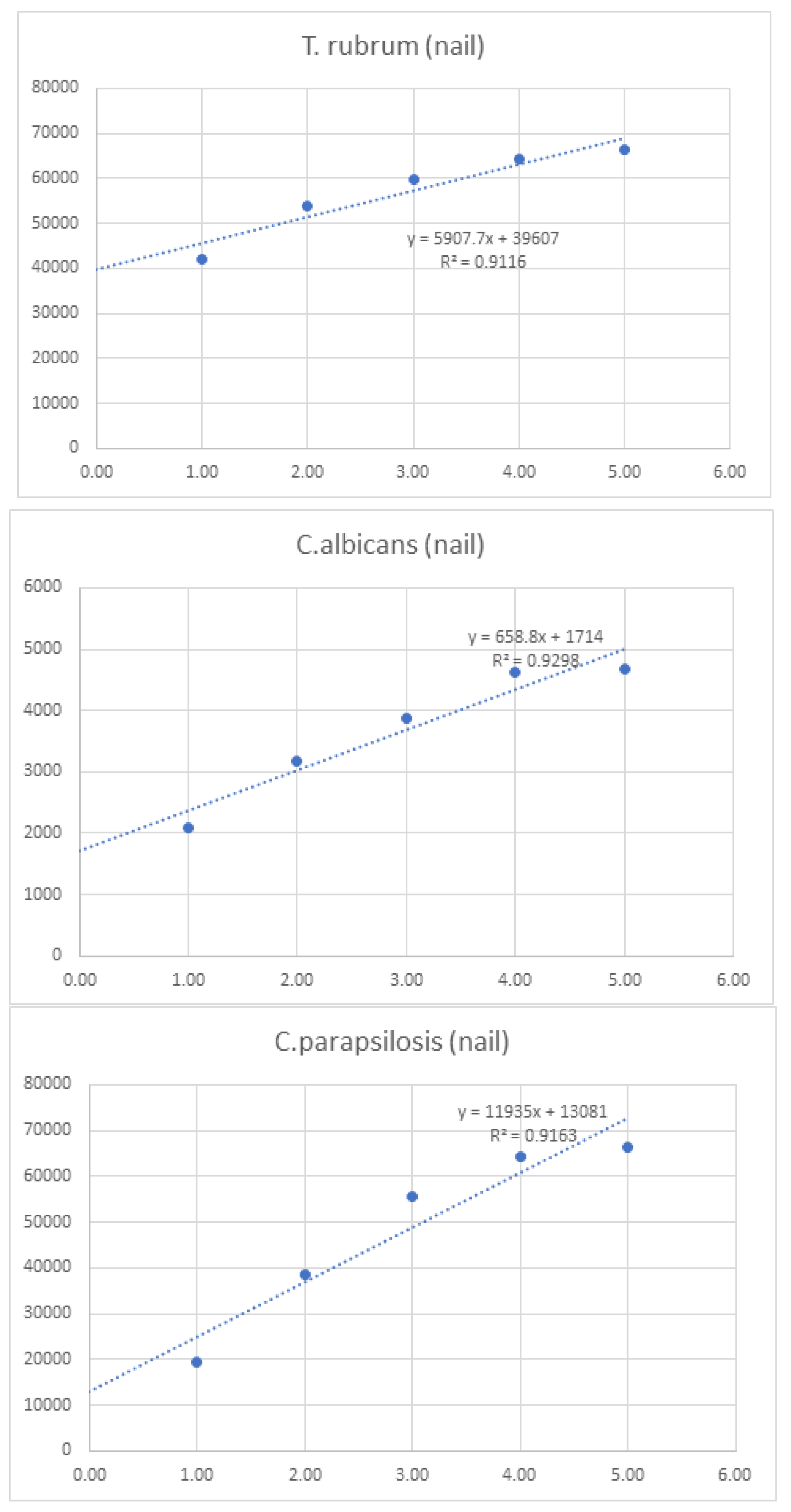
| T. rubrum (nail) | - | 42030 | 54028 | 59848 | 64319 | 66423 | 286,648.0 | ||
| C.albicans (nail) | - | 2096 | 3184 | 3876 | 4628 | 4668 | 18,452.0 | ||
| C.parapsilosis (nail) | - | 19520 | 38668 | 55579.75 | 64255 | 66399 | 244,421.8 | ||
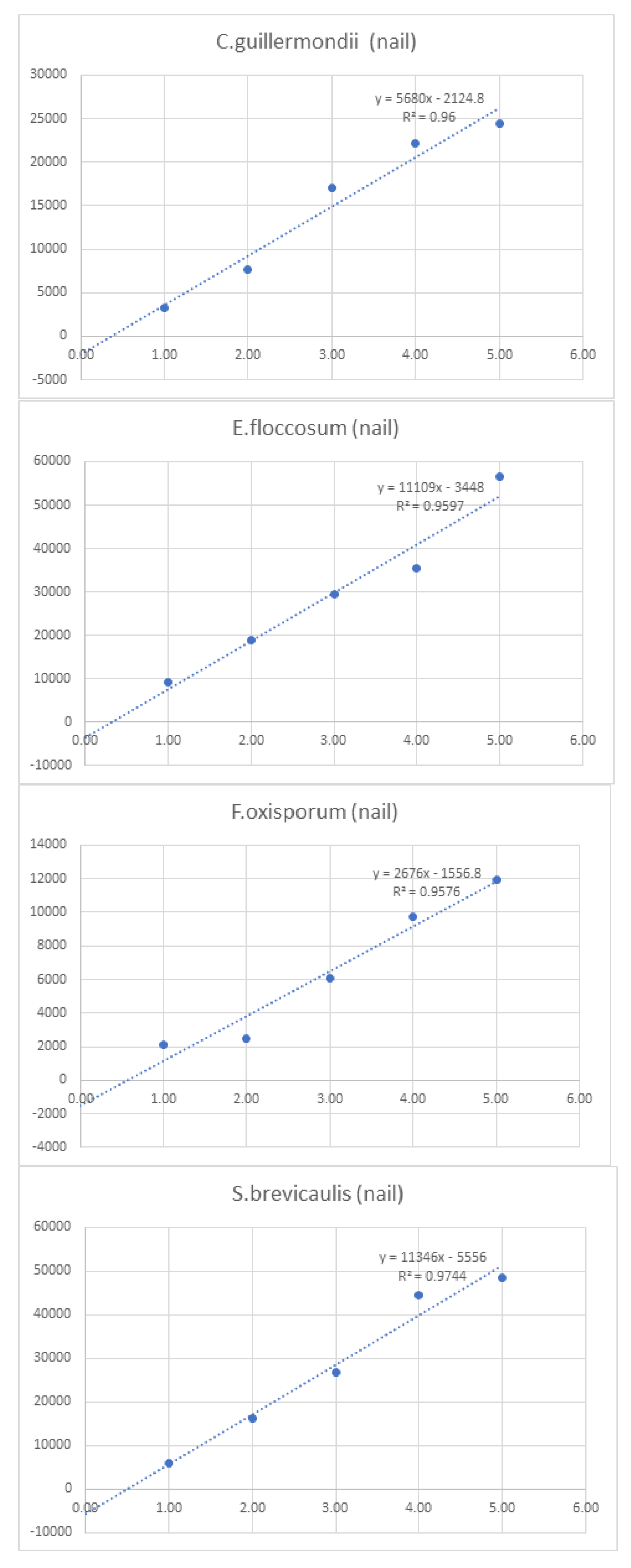
| C.guillermondii (nail) | - | 3308 | 7600 | 17052 | 22216 | 24400 | 74,576.0 | ||
| E.floccosum (nail) | - | 9272 | 18776 | 29388 | 35512 | 56450 | 149,398.0 | ||
| F.oxisporum (nail) | - | 2136 | 2508 | 6084 | 9716 | 11912 | 32,356.0 | ||
| S.brevicaulis (nail) | - | 6048 | 16340 | 26732 | 44676 | 48608 | 142,404.0 | ||
| Skin | T.interdigitale (skin) | - | 4512 | 5160 | 4648 | 4808 | 5920 | 25,048.0 | |
| T.menta inter (skin) | - | 0 | 0 | 0 | 0 | 0 | 0.0 | ||
| T.concentricum (skin) | - | 0 | 0 | 0 | 0 | 0 | 0.0 | ||
| T.equinum (skin) | - | 104 | 248 | 1360 | 1384 | 3792 | 6,888.0 | ||
| T.violaceum (skin) | - | 16 | 1216 | 1464 | 1888 | 4384 | 8,968.0 | ||
| E.floccosum (skin) | - | 3944 | 3536 | 6672 | 7512 | 16056 | 37,720.0 | ||
| T. menta quinkeanum (skin) | - | 1080 | 1776 | 8848 | 15848 | 24312 | 51,864.0 | ||
| M.canis (skin) | - | 8 | 0 | 1976 | 3120 | 8464 | 13,568.0 | ||
| N.persicolor (skin) | - | 8344 | 11768 | 35520 | 52248 | 60431.5 | 168,311.5 | ||
| T.schoenleinii (skin) | - | 5104 | 8304 | 21824 | 24312 | 32288 | 91,832.0 | ||
| Hair | T.tonsurans (hair) | - | 56 | 3280 | 6160 | 8104 | 9312 | 26,912.0 | |
| T.violaceum (hair) | - | 2544 | 3184 | 4000 | 6576 | 8456 | 24,760.0 | ||
| M.audouinii (hair) | - | 72 | 784 | 672 | 816 | 1352 | 3,696.0 | ||
| M.canis (hair) | - | 200 | 960 | 2240 | 2944 | 2848 | 9,192.0 | ||
| N.persicolor (hair) | - | 17128 | 16448 | 26168 | 31760 | 30720 | 122,224.0 | ||
3. Discussion
4. Materials and Methods
4.1. Sample Collection and DNA Extraction
4.2. PCR Amplification
4.3. Microarray Hybridization and Detection
4.4. Data Analysis
4.5. gDNA
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Weitzman, I., & Summerbell, R. C. (1995). The dermatophytes. Clinical microbiology reviews, 8(2), 240-259.
- Goldstein, A. O., & Goldstein, B. G. (2017). Dermatophyte (tinea) infections. Walthman, MA: UpToDate.
- Li, H. C., Bouchara, J. P., Hsu, M. M. L., Barton, R., Su, S., & Chang, T. C. (2008). Identification of dermatophytes by sequence analysis of the rRNA gene internal transcribed spacer regions. Journal of medical microbiology, 57(5), 592-600. [CrossRef]
- Pires, C. A. A., Cruz, N. F. S. D., Lobato, A. M., Sousa, P. O. D., Carneiro, F. R. O., & Mendes, A. M. D. (2014). Clinical, epidemiological, and therapeutic profile of dermatophytosis. Anais brasileiros de dermatologia, 89, 259-264. [CrossRef]
- Cortez, A. C. A., de Souza, J. V. B., Sadahiro, A., & de Oliveira, J. A. A. (2012). Frequency and aetiology of dermatophytosis in children age 12 and under in the state of Amazonas, Brazil. Revista iberoamericana de micologia, 29(4), 223-226.
- Microsporum, T. (2004). Ringworm, Tinea.
- Ghannoum, M. A., Hajjeh, R. A., Scher, R., Konnikov, N., Gupta, A. K., Summerbell, R., ... & Elewski, B. (2000). A large-scale North American study of fungal isolates from nails: the frequency of onychomycosis, fungal distribution, and antifungal susceptibility patterns. Journal of the American Academy of Dermatology, 43(4), 641-648. [CrossRef]
- Gupta, A. K., Jain, H. C., Lynde, C. W., MacDonald, P., Cooper, E. A., & Summerbell, R. C. (2000). Prevalence and epidemiology of onychomycosis in patients visiting physicians’ offices: a multicenter Canadian survey of 15,000 patients. Journal of the American Academy of Dermatology, 43(2), 244-248. [CrossRef]
- Liu, G., He, C., & Zhang, H. (2014). Identification and characterization of dermatophyte species and strains with PCR amplification. Experimental and therapeutic medicine, 8(2), 545-550. [CrossRef]
- Faggi, E., Pini, G., Campisi, E., Bertellini, C., Difonzo, E., & Mancianti, F. (2001). Application of PCR to distinguish common species of dermatophytes. Journal of clinical microbiology, 39(9), 3382-3385. [CrossRef]
- Kim, J. Y., Choe, Y. B., Ahn, K. J., & Lee, Y. W. (2011). Identification of dermatophytes using multiplex polymerase chain reaction. Annals of dermatology, 23(3), 304-312. [CrossRef]
- [EUROIMMUN Medizinische Labordiagnostika AG]. (2021, August 4). EUROArray Dermatomycosis: Multiplex PCR combined with a microarray [Video]. Youtube. https://www.youtube.com/watch?v=k5xoIuR7AgQ.
- Nasrin, S., Saeed, Z. B., Sina, M., & Sahar, N. (2017). Genotyping and molecular characterization of dermatophytes isolates collected from clinical samples. Archives of Pulmonology and Respiratory Care, 3(1), 052-057. [CrossRef]
- Shehata, A. S., Mukherjee, P. K., Aboulatta, H. N., El Akhras, A. I., Abbadi, S. H., & Ghannoum, M. A. (2008). Single-step PCR using (GACA) 4 primer: utility for rapid identification of dermatophyte species and strains. Journal of clinical microbiology, 46(8), 2641-2645. [CrossRef]
- Garg, J., Tilak, R., Garg, A., Prakash, P., Gulati, A. K., & Nath, G. (2009). Rapid detection of dermatophytes from skin and hair. BMC research notes, 2, 1-6. [CrossRef]
- Kano, R., Hirai, A., Muramatsu, M., Watari, T., & Hasegawa, A. (2003). Direct detection of dermatophytes in skin samples based on sequences of the chitin synthase 1 (CHS1) gene. Journal of veterinary medical science, 65(2), 267-270. [CrossRef]
- Robert, R., & Pihet, M. (2008). Conventional methods for the diagnosis of dermatophytosis. Mycopathologia, 166, 295-306. [CrossRef]
- Pihet, M., & Le Govic, Y. (2017). Reappraisal of conventional diagnosis for dermatophytes. Mycopathologia, 182(1), 169-180. [CrossRef]
- Nenoff, P., Krüger, C., Schaller, J., Ginter-Hanselmayer, G., Schulte-Beerbühl, R., & Tietz, H. J. (2014). Mycology–an update part 2: dermatomycoses: clinical picture and diagnostics. JDDG: Journal der Deutschen Dermatologischen Gesellschaft, 12(9), 749-777. [CrossRef]
- Lockhart, S. R., Smith, D. J., & Gold, J. A. (2023). Trichophyton indotineae and other terbinafine-resistant dermatophytes in North America. Journal of Clinical Microbiology, 61(12), e00903-23. [CrossRef]
- Gupta, A. K., Wang, T., Mann, A., Piguet, V., Chowdhary, A., & Bakotic, W. L. (2025). Mechanisms of resistance against allylamine and azole antifungals in Trichophyton: A renewed call for innovative molecular diagnostics in susceptibility testing. PLoS pathogens, 21(2), e1012913. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




