Submitted:
31 August 2025
Posted:
01 September 2025
You are already at the latest version
Abstract
Keywords:
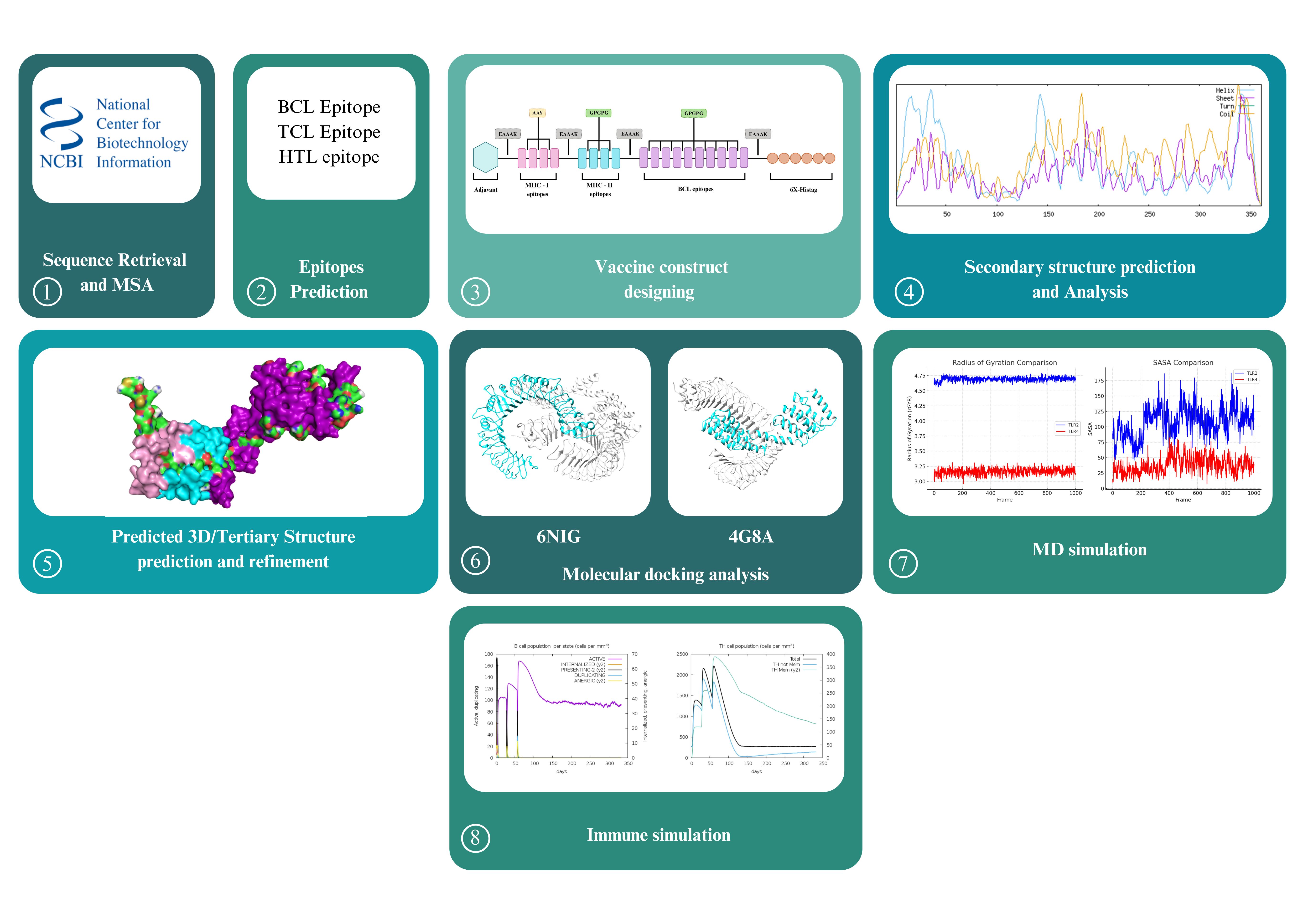
1. Introduction
2. Material and Methods
Protein Sequence Retrieval
BCL Epitope Prediction
CTL and HTL Based Epitopes Prediction
Prediction of CD8+T Cell Epitopes
Prediction of CD4+T Cell Epitopes
Prediction of Discontinuous Epitopes
Design of the SRLV Based Vaccine
Physiochemical Properties
Secondary Structure Prediction
Tertiary Structure Prediction
Molecular Docking Analysis of the SRLV Based Vaccine Construct with TLR-2 and TLR-4 Receptors
Molecular Dynamic Simulations
Immune Simulation Studies
Codon Optimization and In-Silico Cloning
3. Results
Sequence Retrieval
Linear BCL Epitopes Prediction
CTL Epitopes
| Allele | Epitope Sequence | Score | Percentile Rank | AllerTOP | VaxiJen | ToxinPred |
|---|---|---|---|---|---|---|
| BoLA-T2c | SMMHQQMLL | 0.933478 | 0.04 | Non-Allergen | Non-Immunogen (100%) | Non-Toxin |
| BoLA-HD6 | RQQEQEKLL | 0.945614 | 0.03 | Non-Allergen | Immunogen (100%) | Non-Toxin |
| BoLA-1:02301 | RQQEQEKLL | 0.768478 | 0.07 | Non-Allergen | Immunogen (100%) | Non-Toxin |
| BoLA-2:00801 | YIAGGKQFW | 0.859677 | 0.02 | Non-Allergen | Immunogen (66%) | Non-Toxin |
| This first column shows the respective allele used to predict the CTL epitope sequence for the vaccine construct whereas the epitope sequence length, score and percentile length are listed in the second, third and fourth column respecttively. Furthermore, the results of allergenicity analysis, antigenicity, toxicity are depicted in column 4-7 respectively. | ||||||
HTL Epitopes
Prediction of Discontinuous Epitopes
Epitopes Selection Criteria
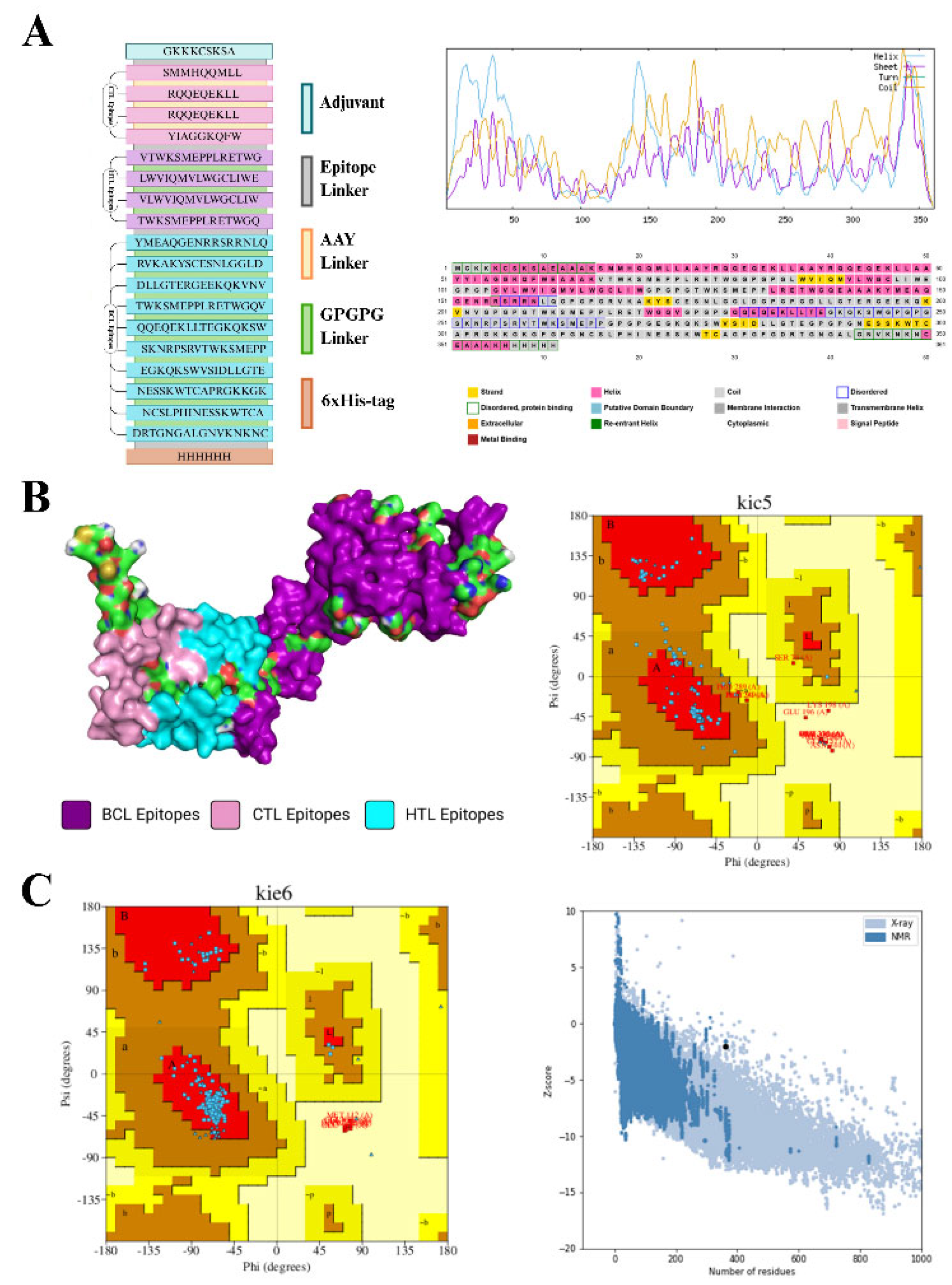
Design of the SRLV Based Vaccine
Physiochemical Properties
Secondary Structure Prediction
Tertiary Structure Prediction
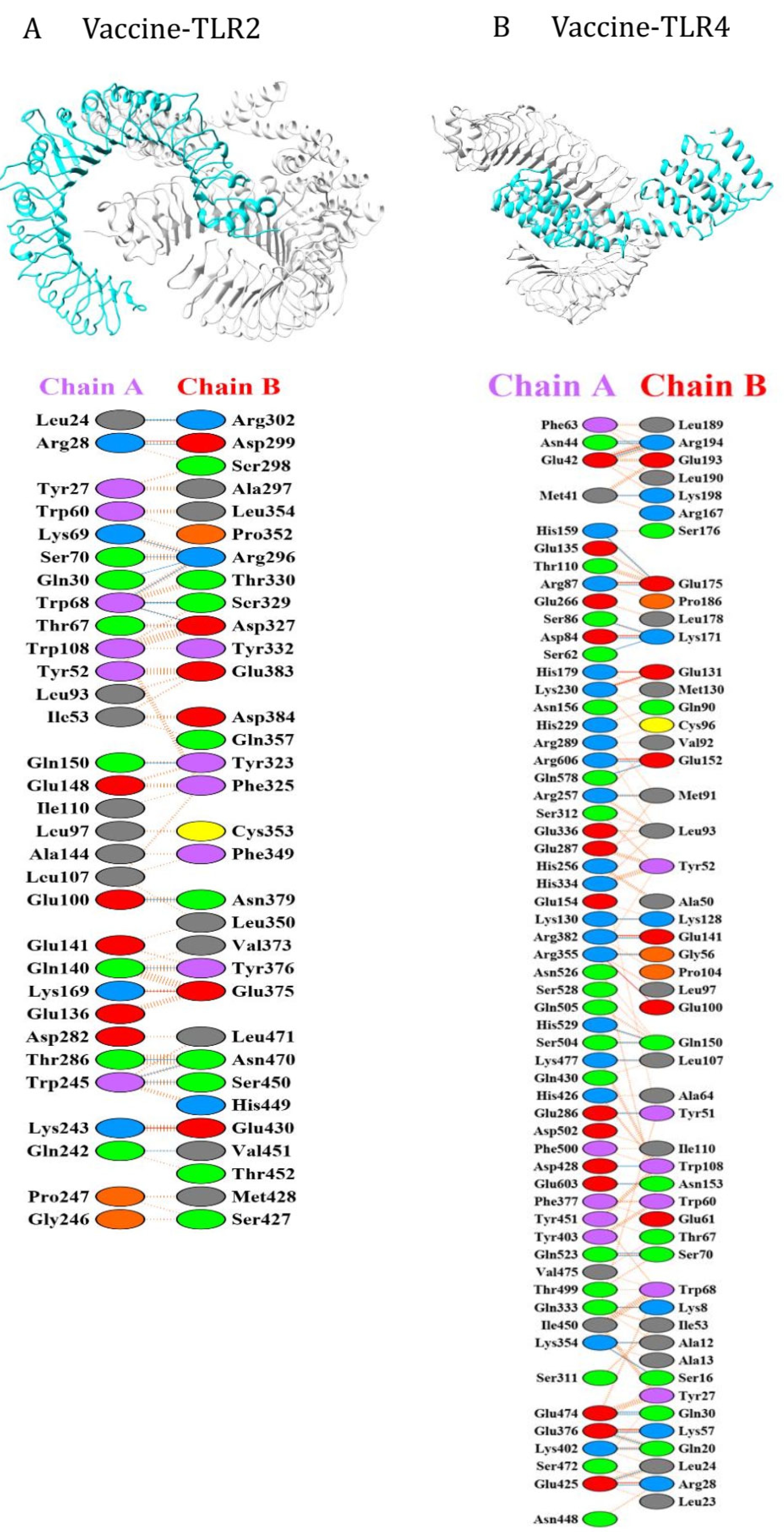
Molecular Docking Analysis of the SRLV Based Vaccine Construct with TLR-2 and TLR-4 Receptors
| Protein-Protein Complex | ΔG (kcal mol-1) | Kd (M) at °C |
ICs charged-charged | ICs charge-polar | ICs charged-apolar | ICs polar-polar | ICs polar-apolar | ICs apolar-apolar | NIS charged | NIS apolar |
|---|---|---|---|---|---|---|---|---|---|---|
| 6NIG_ Construct |
-17.4 | 1.7e-13 | 22 | 21 | 50 | 16 | 33 | 39 | 27.14 | 33.27 |
| The above table shows the outcome of the PRODIGY analysis for the docked vaccine construct for the TLR-2 based ligand 6NIG. Various affinity binding parameters are listed in the respective columns. | ||||||||||
| Protein-Protein Complex | ΔG (kcal mol-1) | Kd (M) at °C |
ICs charged-charged | ICs charged-polar | ICs charged-apolar | ICs polar-polar | ICs polar-apolar | ICs apolar-apolar | NIS charged | NIS apolar | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 4G8A-Construct | -22.5 | 3e-17 | 30 | 38 | 96 | 13 | 31 | 20 | 23.69 | 37.85 | |
| The above table shows the outcome of the PRODIGY analysis for the docked vaccine construct for the TLR-4 based ligand 4G8A. Various affinity binding parameters are listed in the respective columns. | |||||||||||
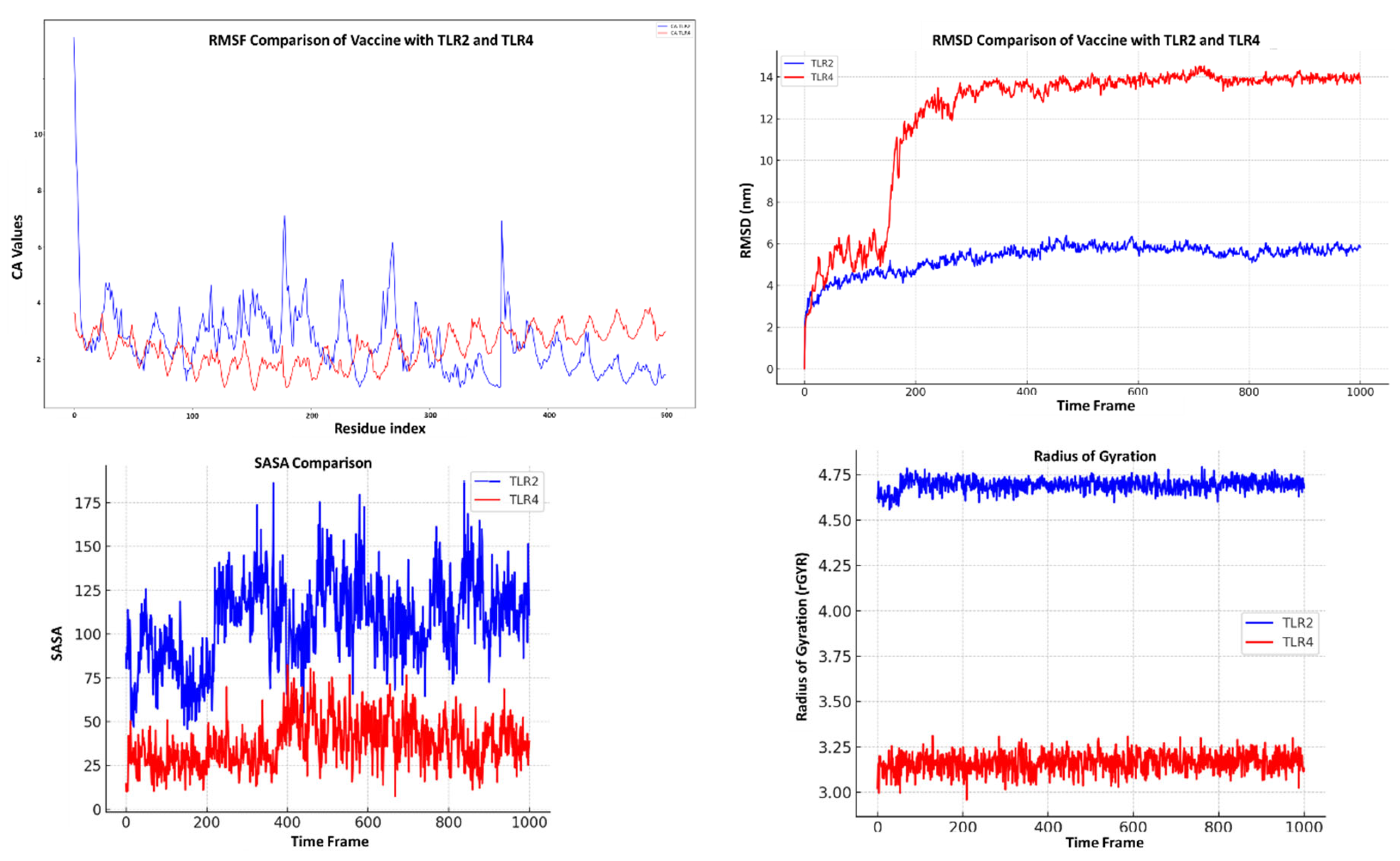
MD Simulations
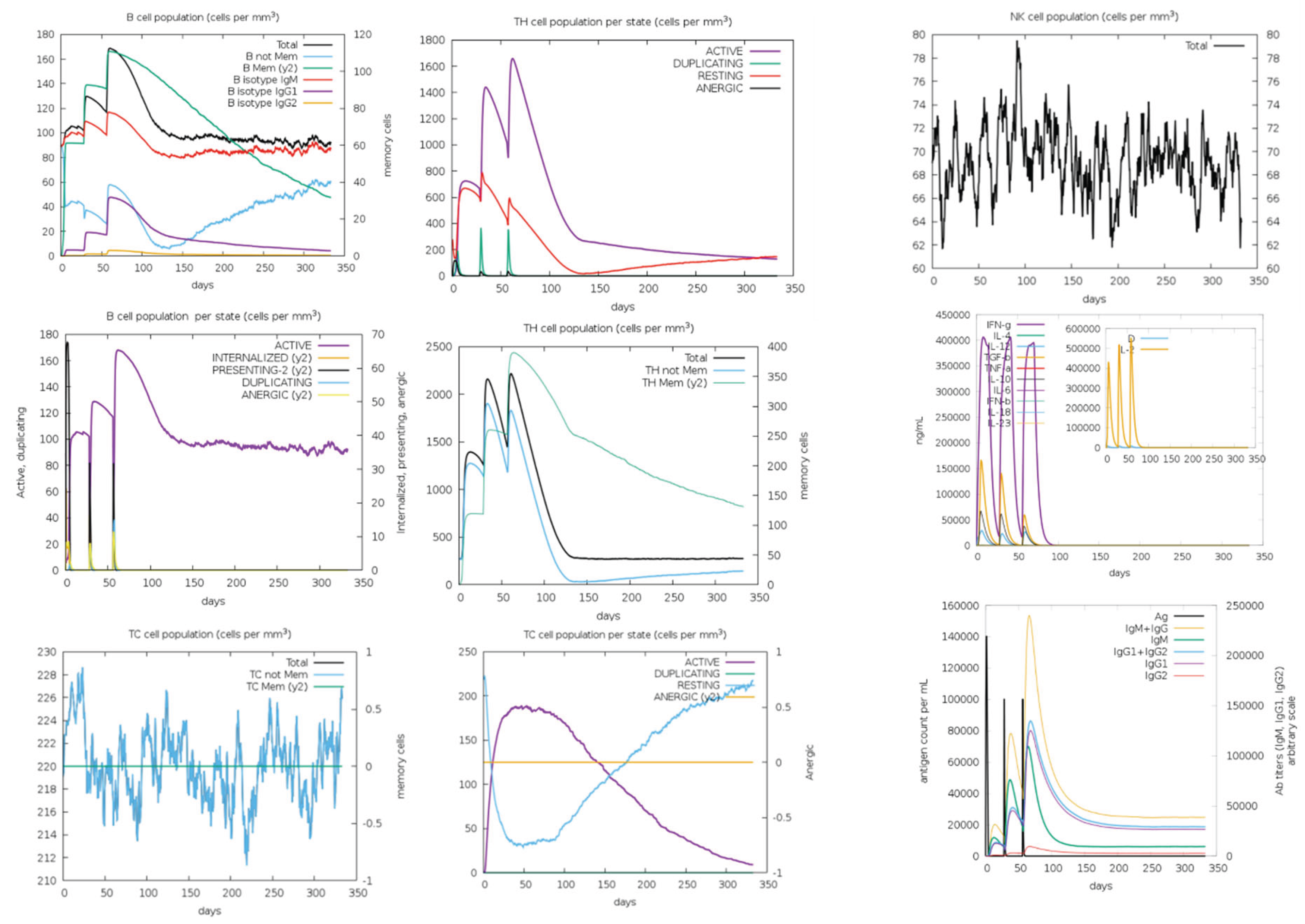
Immune Simulations
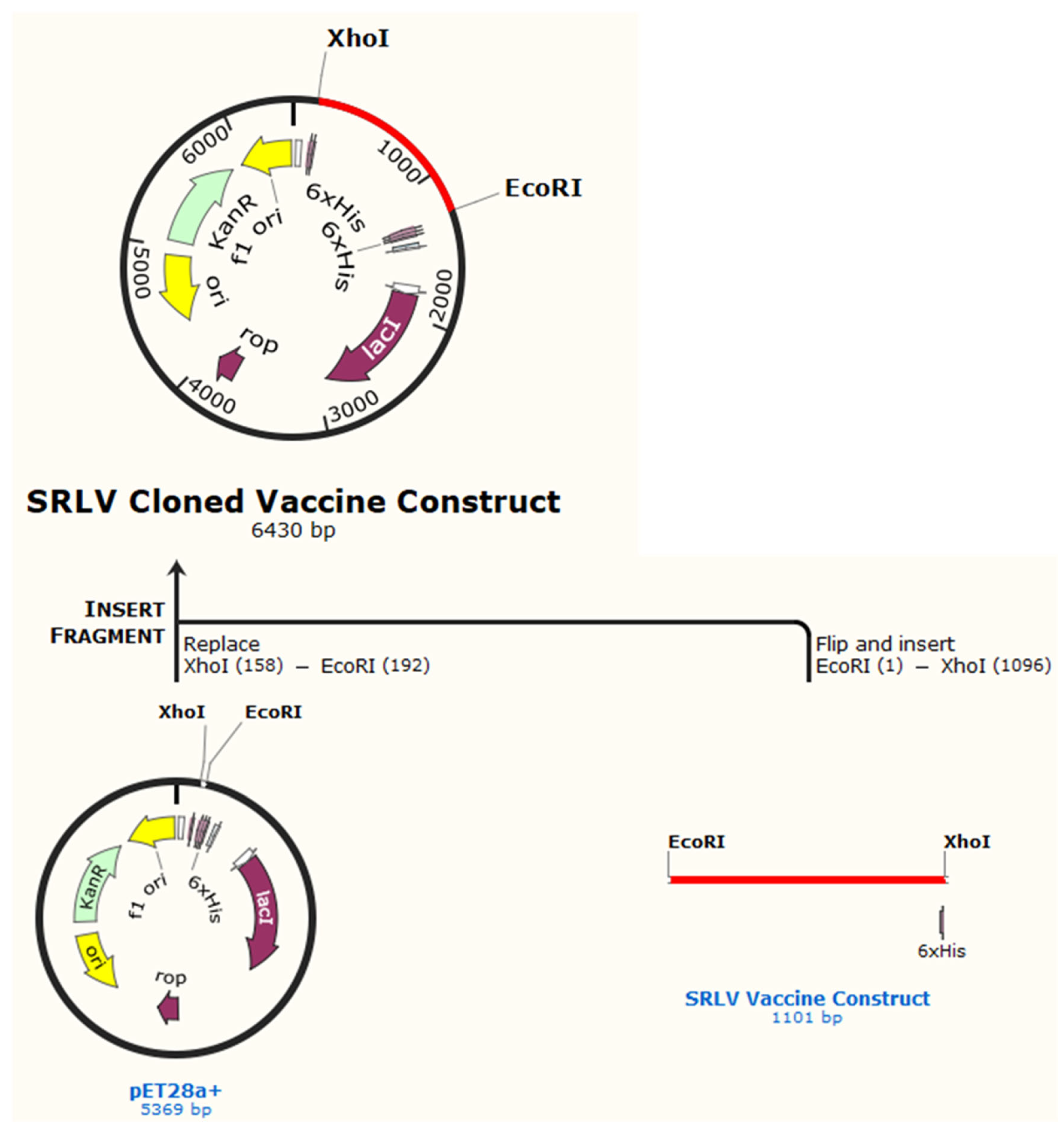
SRLV Vaccine Molecular Cloning and Codon-Optimization
4. Discussion
5. Limitations
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Ethics Statement/Consent Information
Conflicts of Interest
References
- Arcangeli, C., M. Torricelli, C. Sebastiani, D. Lucarelli, M. Ciullo, F. Passamonti, M. Giammarioli and M. J. V. Biagetti (2022). "Genetic characterization of small ruminant lentiviruses (SRLVs) circulating in naturally infected sheep in Central Italy." 14(4): 686.
- Blacklaws, B., E. Berriatua, S. Torsteinsdottir, N. Watt, D. De Andres, D. Klein and G. J. V. M. Harkiss (2004). "Transmission of small ruminant lentiviruses." 101(3): 199-208.
- Blacklaws, B. A. J. C. i., microbiology and i. diseases (2012). "Small ruminant lentiviruses: Immunopathogenesis of visna-maedi and caprine arthritis and encephalitis virus." 35(3): 259-269.
- Cheevers, W., D. Knowles, T. McGuire, T. Baszler, G. J. V. i. Hullinger and immunopathology (1994). "Caprine arthritis-encephalitis lentivirus (CAEV) challenge of goats immunized with recombinant vaccinia virus expressing CAEV surface and transmembrane envelope glycoproteins." 42(3-4): 237-251.
- Davaasuren, N., V. Molaee, T.-O. Erdene-Ochir, G. Nyamdavaa, S. Ganzorig, M. Mazzei, Y. Sakoda, G. Lühken and S. J. V. R. C. Tumenjargal (2024). "Phylogenetic analysis of small ruminant lentiviruses in Mongolian sheep supports an ancient east-west split for the genotype A." 48(3): 1955-1962.
- De Andres, X., R. Reina, J. Ciriza, H. Crespo, I. Glaria, H. Ramírez, M. J. Grilló, M. M. Pérez, V. Andresdottir and S. J. V. Rosati (2009). "Use of B7 costimulatory molecules as adjuvants in a prime-boost vaccination against Visna/Maedi ovine lentivirus." 27(34): 4591-4600.
- Dimitrov, I., I. Bangov, D. R. Flower and I. J. J. o. m. m. Doytchinova (2014). "AllerTOP v. 2—a server for in silico prediction of allergens." 20: 1-6.
- Doytchinova, I. A. and D. R. J. B. b. Flower (2007). "VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines." 8: 1-7.
- Doytchinova, I. A., P. Guan and D. R. J. B. b. Flower (2006). "EpiJen: a server for multistep T cell epitope prediction." 7: 1-11.
- ENACHE, D. A., S. BARAITAREANU, M. DAN, M. R. GURAU, F. OTELEA, A. DOBRE and D. J. S. W. S. C. DANES, Veterinary Medicine (2017). "Preliminary results of MVV and CAEV seroprevalence in Romanian sheep and goats." 63(1).
- Gasteiger, E., C. Hoogland, A. Gattiker, S. e. Duvaud, M. R. Wilkins, R. D. Appel and A. J. T. p. p. h. Bairoch (2005). "Protein identification and analysis tools on the ExPASy server." 571-607.
- Geourjon, C. and G. J. B. Deleage (1995). "SOPMA: significant improvements in protein secondary structure prediction by consensus prediction from multiple alignments." 11(6): 681-684.
- Gomez-Lucia, E., N. Barquero, A. J. V. M. R. Domenech and Reports (2018). "Maedi-Visna virus: current perspectives." 11-21.
- González, B., R. Reina, I. García, S. Andrés, I. Glaria, M. Alzueta, M. I. Mora, B. M. Jugo, I. Arrieta-Aguirre and J. M. P. J. V. de la Lastra (2005). "Mucosal immunization of sheep with a Maedi-Visna virus (MVV) env DNA vaccine protects against early MVV productive infection." 23(34): 4342-4352.
- Grote, A., K. Hiller, M. Scheer, R. Münch, B. Nörtemann, D. C. Hempel and D. J. N. a. r. Jahn (2005). "JCat: a novel tool to adapt codon usage of a target gene to its potential expression host." 33(suppl_2): W526-W531.
- Hall, T., I. Biosciences and C. J. G. b. b. Carlsbad (2011). "BioEdit: an important software for molecular biology." 2(1): 60-61.
- Heo, L., H. Park and C. J. N. a. r. Seok (2013). "GalaxyRefine: Protein structure refinement driven by side-chain repacking." 41(W1): W384-W388.
- Jespersen, M. C., B. Peters, M. Nielsen and P. J. N. a. r. Marcatili (2017). "BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes." 45(W1): W24-W29.
- Koçkaya, E. S., H. Can, Y. Yaman and C. J. B. Ün (2023). "In silico discovery of epitopes of gag and env proteins for the development of a multi-epitope vaccine candidate against maedi visna virus using reverse vaccinology approach." 84: 101715.
- Kosloff, D. and R. J. J. o. C. P. Kosloff (1983). "A Fourier method solution for the time dependent Schrödinger equation as a tool in molecular dynamics." 52(1): 35-53.
- Kozakov, D., D. R. Hall, B. Xia, K. A. Porter, D. Padhorny, C. Yueh, D. Beglov and S. J. N. p. Vajda (2017). "The ClusPro web server for protein–protein docking." 12(2): 255-278.
- Krissinel, E. and K. J. J. o. m. b. Henrick (2007). "Inference of macromolecular assemblies from crystalline state." 372(3): 774-797.
- Laskowski, R. A., J. Jabłońska, L. Pravda, R. S. Vařeková and J. M. J. P. s. Thornton (2018). "PDBsum: Structural summaries of PDB entries." 27(1): 129-134.
- Laskowski, R. A., M. W. MacArthur, D. S. Moss and J. M. J. A. C. Thornton (1993). "PROCHECK: a program to check the stereochemical quality of protein structures." 26(2): 283-291.
- Magnan, C. N., A. Randall and P. J. B. Baldi (2009). "SOLpro: accurate sequence-based prediction of protein solubility." 25(17): 2200-2207.
- Martin, D., J. Procter, B. Soares, A. Waterhouse, S. Shehata, N. Giang, M. Carstairs, C. Ofoegbu, K. Mourão, S. J. M. Duce and I. Tutorial (2020). "Jalview 2.11.".
- McGuffin, L. J., K. Bryson and D. T. J. B. Jones (2000). "The PSIPRED protein structure prediction server." 16(4): 404-405.
- Michiels, R., E. Van Mael, C. Quinet, N. R. Adjadj, A. B. Cay and N. J. V. De Regge (2018). "Comparative analysis of different serological and molecular tests for the detection of small ruminant lentiviruses (SRLVs) in Belgian sheep and goats." 10(12): 696.
- Minardi da Cruz, J. C., D. K. Singh, A. Lamara and Y. J. V. Chebloune (2013). "Small ruminant lentiviruses (SRLVs) break the species barrier to acquire new host range." 5(7): 1867-1884.
- Molaee, V., M. Bazzucchi, G. M. De Mia, V. Otarod, D. Abdollahi, S. Rosati and G. J. S. r. Lühken (2020). "Phylogenetic analysis of small ruminant lentiviruses in Germany and Iran suggests their expansion with domestic sheep." 10(1): 2243.
- Moroz, A., M. Czopowicz, M. Sobczak-Filipiak, I. Dolka, M. Rzewuska, M. Kizerwetter-Świda, D. Chrobak-Chmiel, M. Mickiewicz, L. Witkowski and O. J. P. Szaluś-Jordanow (2022). "The prevalence of histopathological features of pneumonia in goats with symptomatic caprine arthritis-encephalitis." 11(6): 629.
- Mosa, A. H. and M. M. Zenad (2020). "Serological and histopathological detection of Maedi-Visna virus in middle Iraq regions.".
- Olech, M., S. Valas and J. J. P. O. Kuźmak (2018). "Epidemiological survey in single-species flocks from Poland reveals expanded genetic and antigenic diversity of small ruminant lentiviruses." 13(3): e0193892.
- Pépin, M., C. Vitu, P. Russo, J.-F. Mornex and E. J. V. r. Peterhans (1998). "Maedi-visna virus infection in sheep: a review." 29(3-4): 341-367.
- Pettersen, E. F., T. D. Goddard, C. C. Huang, G. S. Couch, D. M. Greenblatt, E. C. Meng and T. E. J. J. o. c. c. Ferrin (2004). "UCSF Chimera—a visualization system for exploratory research and analysis." 25(13): 1605-1612.
- Pétursson, G., S. Matthíasdóttir, V. Svansson, V. Andrésdóttir, G. Georgsson, A. H. Martin, G. Agnarsdóttir, E. Gísladóttir, S. Árnadóttir and S. J. V. Högnadóttir (2005). "Mucosal vaccination with an attenuated maedi–visna virus clone." 23(24): 3223-3228.
- Ponomarenko, J., H.-H. Bui, W. Li, N. Fusseder, P. E. Bourne, A. Sette and B. J. B. b. Peters (2008). "ElliPro: a new structure-based tool for the prediction of antibody epitopes." 9: 1-8.
- Querat, G., G. Audoly, P. Sonigo and R. J. V. Vigne (1990). "Nucleotide sequence analysis of SA-OMVV, a visna-related ovine lentivirus: phylogenetic history of lentiviruses." 175(2): 434-447.
- Rahman, M. H., E. Ahmed, M. N. Haque, M. Z. Hassan and M. Z. J. B. J. o. L. R. Ali (2022). "Major respiratory diseases of goat and their epidemiology, prevention and control." 1-20.
- Rapin, N., O. Lund and F. J. B. Castiglione (2011). "Immune system simulation online." 27(14): 2013-2014.
- Saha, S. and G. P. S. Raghava (2004). BcePred: prediction of continuous B-cell epitopes in antigenic sequences using physico-chemical properties. International conference on artificial immune systems, Springer.
- Saha, S., G. P. S. J. P. S. Raghava, Function, and Bioinformatics (2006). "Prediction of continuous B-cell epitopes in an antigen using recurrent neural network." 65(1): 40-48.
- Santry, L. A., J. de Jong, A. C. Gold, S. R. Walsh, P. I. Menzies and S. K. J. V. R. Wootton (2013). "Genetic characterization of small ruminant lentiviruses circulating in naturally infected sheep and goats in Ontario, Canada." 175(1): 30-44.
- Seeliger, D. and B. L. J. J. o. c.-a. m. d. de Groot (2010). "Ligand docking and binding site analysis with PyMOL and Autodock/Vina." 24(5): 417-422.
- Sharma, N., L. D. Naorem, S. Jain and G. P. J. B. i. b. Raghava (2022). "ToxinPred2: an improved method for predicting toxicity of proteins." 23(5): bbac174.
- Sievers, F. and D. G. J. C. p. i. b. Higgins (2014). "Clustal omega." 48(1): 3.13. 11-13.13. 16.
- Singh, H. and G. J. B. Raghava (2001). "ProPred: prediction of HLA-DR binding sites." 17(12): 1236-1237.
- Torsteinsdóttir, S., H. M. Carlsdóttir, V. Svansson, S. Matthíasdóttir, A. H. Martin and G. J. V. Pétursson (2007). "Vaccination of sheep with Maedi-visna virus gag gene and protein, beneficial or harmful?" 25(37-38): 6713-6720.
- Trolle, T., I. G. Metushi, J. A. Greenbaum, Y. Kim, J. Sidney, O. Lund, A. Sette, B. Peters and M. J. B. Nielsen (2015). "Automated benchmarking of peptide-MHC class I binding predictions." 31(13): 2174-2181.
- Wiederstein, M. and M. J. J. N. a. r. Sippl (2007). "ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins." 35(suppl_2): W407-W410.
- Xue, L. C., J. P. Rodrigues, P. L. Kastritis, A. M. Bonvin and A. J. B. Vangone (2016). "PRODIGY: a web server for predicting the binding affinity of protein–protein complexes." 32(23): 3676-3678.
- Yang, Y., Z. Wei, G. Cia, X. Song, F. Pucci, M. Rooman, F. Xue and Q. J. F. i. i. Hou (2024). "MHCII-peptide presentation: an assessment of the state-of-the-art prediction methods." 15: 1293706.
- Zhang, Y. J. B. b. (2008). "I-TASSER server for protein 3D structure prediction." 9: 1-8.
| TYPE OF VACCINE | TARGETED VIRAL AGENT | TARGETED SPECIE | STUDY PARAMETERS | OUTCOMES | REFERENCES |
|---|---|---|---|---|---|
| Attenuated molecular clone of Maedi Visna virus (MVV) (2005) |
Maedi-Visna Virus (MVV) |
Sheep |
Four sheep were infected with attenuated molecular clone of maedi visna virus (MVV) (LV1-1KS1); while comparing them with control group (n=4) | Low viral titer was detected in the ELISA test indicating partial protection. | (Pétursson, Matthíasdóttir et al. 2005) |
| DNA based vaccine based on the envelope gene (2005) |
Maedi-Visna Virus (MVV) | Sheep | pcDNA plasmid (pcDNA-env) encoding the env glycoproteins of MVV, boosted with combined pcDNA-env and pCR3.1-IFN-γ plasmid inoculations were introduced in both test and control groups | Early protective effect was observed in the vaccinated sheeps that restricted the virus replication however it disappeared after two years. | (González, Reina et al. 2005) |
| DNA vaccine containing the Maedi-Visna virus (MVV) gag gene (2007) |
Maedi-Visna Virus (MVV) | Sheep |
Sheep were immunized eight times over a period of 2.5 years with the Maedi-Visna (MVV) gag gene, VR1012-gag-CTE and pcDNA3.1-gag-CTE. | No positive outcome was observed. In fact, the vaccine may have made them more susceptible to the virus. | (Torsteinsdóttir, Carlsdóttir et al. 2007) |
| Use of B7 costimulatory molecules as adjuvants in a prime-boost vaccination against Visna/Maedi ovine lentivirus (2009) |
Visna/Maedi ovine lentivirus |
Sheep (Llewyn sheep breed |
Sheep were primed with particle-mediated epidermal bombardment (PMED) using plasmids containing the VMV gag and env genes. | Enhanced early cellular immune response. | (De Andres, Reina et al. 2009) |
| In silico discovery of epitopes of gag and env proteins for the development of a multi-epitope vaccine candidate against Maedi Visna Virus (2023) | Maedi-Visna | Sheep | Retrieved the DNA sequence for gag and env gene of MVV KV1514 isolate from NCBI, and a multi-epitope vaccine was designed based on the 19 epitopes. | Gag protein was detected to be more conserved and a strong affinity was shown between the final multi-epitope vaccine and TLR-2/4. | (Koçkaya, Can et al. 2023) |
| Recombinant Vaccinia Virus rWR63 Expressing CAEV Envelope Glycoproteins | CEAV | Goat | Goats (n=6) were vaccinated thrice with rWR63, followed by different booster immunizations. Control group received rWRSC11 | All vaccinated goats developed antibodies to CAEV glycoproteins, but no neutralizing antibodies were detected. | (Cheevers, Knowles et al. 1994) |
| The table depicts the various types of vaccine candidates that have been reported over the years against MVV and CAEV infections. The type of vaccine candidate is shown in the first column, whereas the targeted viral agent and specie are mentioned in the second and third column. Furthermore, the parameters chosen for the specific study and the outcomes are depicted in the fourth and fifth column respectively. Moreover, the reference articles of the study are mentioned in the sixth column. | |||||
| Serotype | NCBI Accession Number | Region | Gene Type | Accession Links |
|---|---|---|---|---|
| A1 | M60609 | Iceland | Envelope | https://www.ncbi.nlm.nih.gov/nuccore/M60609 |
| A1 | OL436272 | Poland | Envelope | https://www.ncbi.nlm.nih.gov/nuccore/OL436272 |
| A2 | AY101611 | USA | Envelope | https://www.ncbi.nlm.nih.gov/nuccore/AY101611 |
| B1 | OL436260 | Poland | Envelope | https://www.ncbi.nlm.nih.gov/nuccore/OL436260 |
| B2 | OL436262 | Poland | Envelope | https://www.ncbi.nlm.nih.gov/nuccore/OL436262 |
| This first column shows the serotype of the selected isolates whereas its accession number is given in the second column. The third column contains the information about the region and followed by its targeted gene type in fourth column. All the information is retrieved through NCBI for which the link is provided in the last column. | ||||
| Sequence | Sequence Length | Score | AllerTop | VexiJen | ToxinPred | SVM Score | Molecular Weight |
|---|---|---|---|---|---|---|---|
| YMEAQGENRRSRRNLQ | 16 | 0.80 | Non-Allergen | Antigen (0.6151) |
Non-Toxin | -1.04 | 1880.28 |
| RVKAKYSCESNLGGLD | 16 | 0.77 | Non-Allergen | Antigen (1.1465) |
Non-Toxin | -0.59 | 1625.09 |
| DLLGTERGEEKQKVNV | 16 | 0.89 | Non-Allergen | Antigen (0.7215) |
Non-Toxin | -1.40 | 1716.11 |
| TWKSMEPPLRETWGQV | 16 | 0.80 | Non-Allergen | Antigen (0.5101) |
Non-Toxin | -0.80 | 1846.31 |
| QQEQEKLLTEGKQKSW |
16 | 0.79 | Non-Allergen | Antigen (0.5243) |
Non-Toxin | -1.29 | 1744.20 |
| SKNRPSRVTWKSMEPP |
16 | 0.70 | Non-Allergen | Antigen (0.8433) |
Non-Toxin | -0.99 | 1803.27 |
| EGKQKSWVSIDLLGTE |
16 | 0.60 | Non-Allergen | Antigen (1.0230) |
Non-Toxin | -1.01 | 1661.12 |
| NESSKWTCAPRGKKGK | 16 | 0.89 | Non-Allergen | Antigen (0.9891) |
Non-Toxin | -0.57 | 1649.06 |
| NCSLPHINESSKWTCA | 16 | 0.70 | Non-Allergen | Antigen (0.5145) |
Non-Toxin | -0.57 | 1719.13 |
| DRTGNGALGNVKNKNC | 16 | 0.82 | Non-Allergen | Antigen (1.2684) |
Non-Toxin | 0.73 | 1557.91 |
| The table shows the selected BCL epitopes for vaccine development. The first column shows the sequence of the selected BCL epitope sequence whereas the sequence length and scores are listed in the second and third column respectively. Furthermore, the results of allergenicity analysis, antigenicity, toxicity are depicted in column 4-6 respectively. SMV score and molecular weight information is mentioned in the seventh and eighth column. | |||||||
| Sequence | Length | Allele | Affinity nM | Percentile Rank | Binding Level | AllerTOP | VaxiJen | Toxin Pred (SVM, Molecular Weight) |
|---|---|---|---|---|---|---|---|---|
| VTWKSMEPPLRETWG |
16 | BoLA-DRB3*0301 |
0.644 |
30.63 | Strong Binder | Non-Allergen | Immunogen (100%) |
Non-Toxin (-0.89, 1817.31) |
| LWVIQMVLWGCLIWE |
16 | DRB1*0102 |
4.41 |
7.00 | Strong Binder | Non-Allergen | Immunogen (66%) |
Non-Toxin (-0.14, 1889.64) |
|
VLWVIQMVLWGCLIW |
16 | DRB1*0104 |
0.761 |
10.00 | Strong Binder | Non-Allergen | Immunogen (66%) |
Non-Toxin (-0.30, 1859.62) |
| TWKSMEPPLRETWGQ |
16 | DRB1*0120 |
0.648 |
29.46 | Strong Binder | Non-Allergen | Immunogen (66%) |
Non-Toxin (-0.80, 1846.31) |
| The predicted HTL epitope sequence for the vaccine construct is shown in the first column whereas the epitope sequence length, respective allele, binding affinity and percentile rank are depicted in the column and percentile length are listed in the second, third and fourth column respecttively. Furthermore, the results of allergenicity analysis, antigenicity, toxicity are depicted in column 4-7 respectively. | ||||||||
| Tool Used | Link |
|---|---|
| Clustal Omega | https://www.ebi.ac.uk/jdispatcher/msa/clustalo |
| JalView | https://www.jalview.org/ |
| BioEdit | https://bioedit.software.informer.com/7.2/ |
| ABCPRED | https://webs.iiitd.edu.in/raghava/abcpred/ |
| BCEPRED | https://webs.iiitd.edu.in/raghava/bcepred/bcepred_submission.html |
| BEPIPRED | https://services.healthtech.dtu.dk/services/BepiPred-2.0/ |
| VAXIJEN 2.0 | https://www.ddgpharmfac.net/vaxijen/VaxiJen/VaxiJen.html |
| AllerTOP | https://www.ddg-0pharmfac.net/allertop_test/ |
| ToxinPred | https://webs.iiitd.edu.in/raghava/toxinpred/ |
| ProPred-I | http://crdd.osdd.net/raghava/propred1/ |
| EpiJen | https://www.ddg-pharmfac.net/epijen/ |
| ElliPro | http://tools.iedb.org/ellipro/ |
| IEDB MHC-I Binding Prediction tools | http://tools.iedb.org/mhci/ |
| NetMHCIIpan 2.1 server | https://services.healthtech.dtu.dk/services/NetMHCIIpan-2.1/ |
| Expasy ProtParam | https://web.expasy.org/cgi-bin/protparam/protparam |
| SolPro | https://scratch.proteomics.ics.uci.edu/ |
| SOPMA | https://npsa.lyon.inserm.fr/cgi-bin/npsa_automat.pl?page=/NPSA/npsa_sopma.html |
| PSIPRED | https://bioinf.cs.ucl.ac.uk/psipred/ |
| I-TASSER | https://zhanggroup.org/I-TASSER/ |
| ProCheck server | https://www.ebi.ac.uk/thornton-srv/software/PROCHECK/ |
| Galaxy Refine server | https://bio.tools/galaxyrefine |
| ProSa webserver | https://prosa.services.came.sbg.ac.at/prosa.php |
| ClusPro | https://cluspro.bu.edu/ |
| Protein Data Bank | https://www.rcsb.org/ |
| PRODIGY | https://rascar.science.uu.nl/prodigy/ |
| PDBePISA | https://www.ebi.ac.uk/pdbe/pisa/ |
| PDBSum | https://www.ebi.ac.uk/thornton-srv/databases/pdbsum/ |
| Schrodinger Maestro software package | https://www.schrodinger.com/platform/products/maestro/ |
| C-ImmSim server | https://wwwold.iac.rm.cnr.it/ |
| The table describes the tools utilized in the current study. The first column depicts the name of the tool whereas the respective link is mentioned in the second column. | |
| Physiochemical Property | Result |
|---|---|
| No. of amino acids | 361 |
| Theoretical pI | 9.38 |
| Molecular Weight | 39077.45 |
| Total number of negatively charged residues (Asp + Glu) | 33 |
| Total number of positively charged residues (Arg + Lys) | 47 |
| Total no. of atoms | 5426 |
| Formula | C1723H2679N505O500S19 |
| Extinction coefficients | Ext. coefficient 108440 M-1 cm-1 Abs 0.1% (=1 g/l) 2.775, assuming all pairs of Cys residues form cystines |
| Estimated half-life | 30 hours (mammalian reticulocytes, in vitro) >20 hours (yeast, in vivo) >10 hours (Escherichia coli, in vivo) |
| Instability Index | 35.15 (Stable) |
| Aliphatic index |
53.02 |
| Grand average of hydropathicity (GRAVY) | -0.840 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




