Submitted:
22 August 2025
Posted:
27 August 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
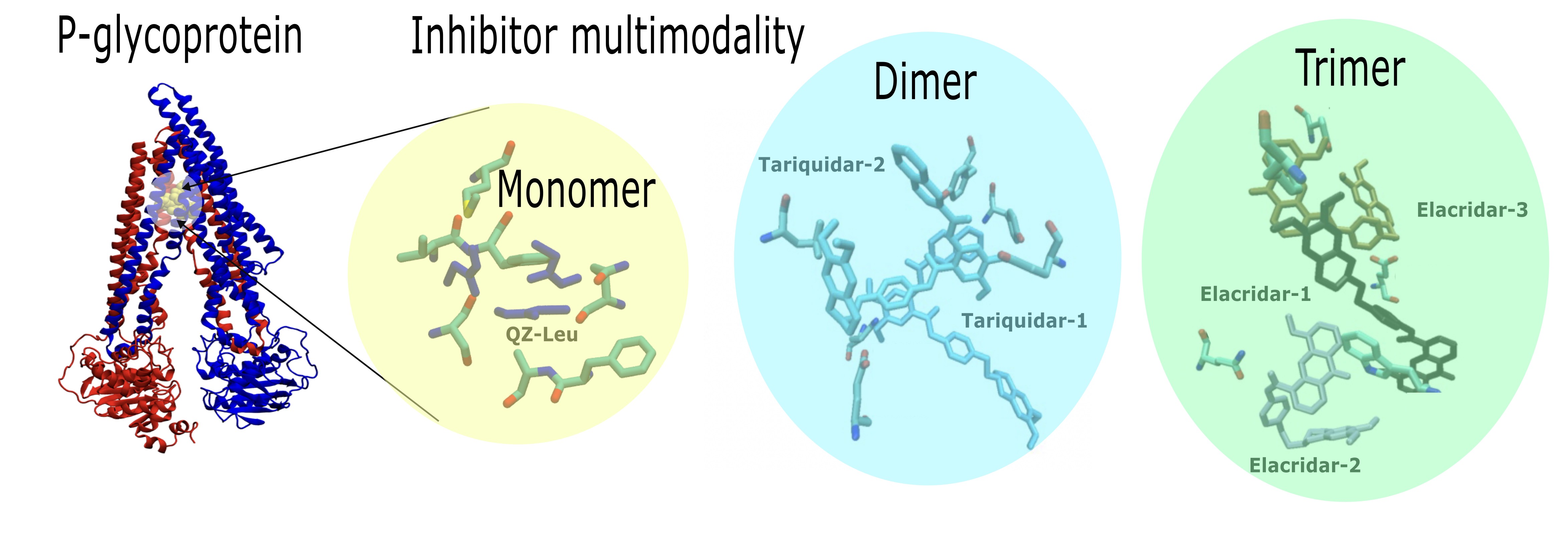
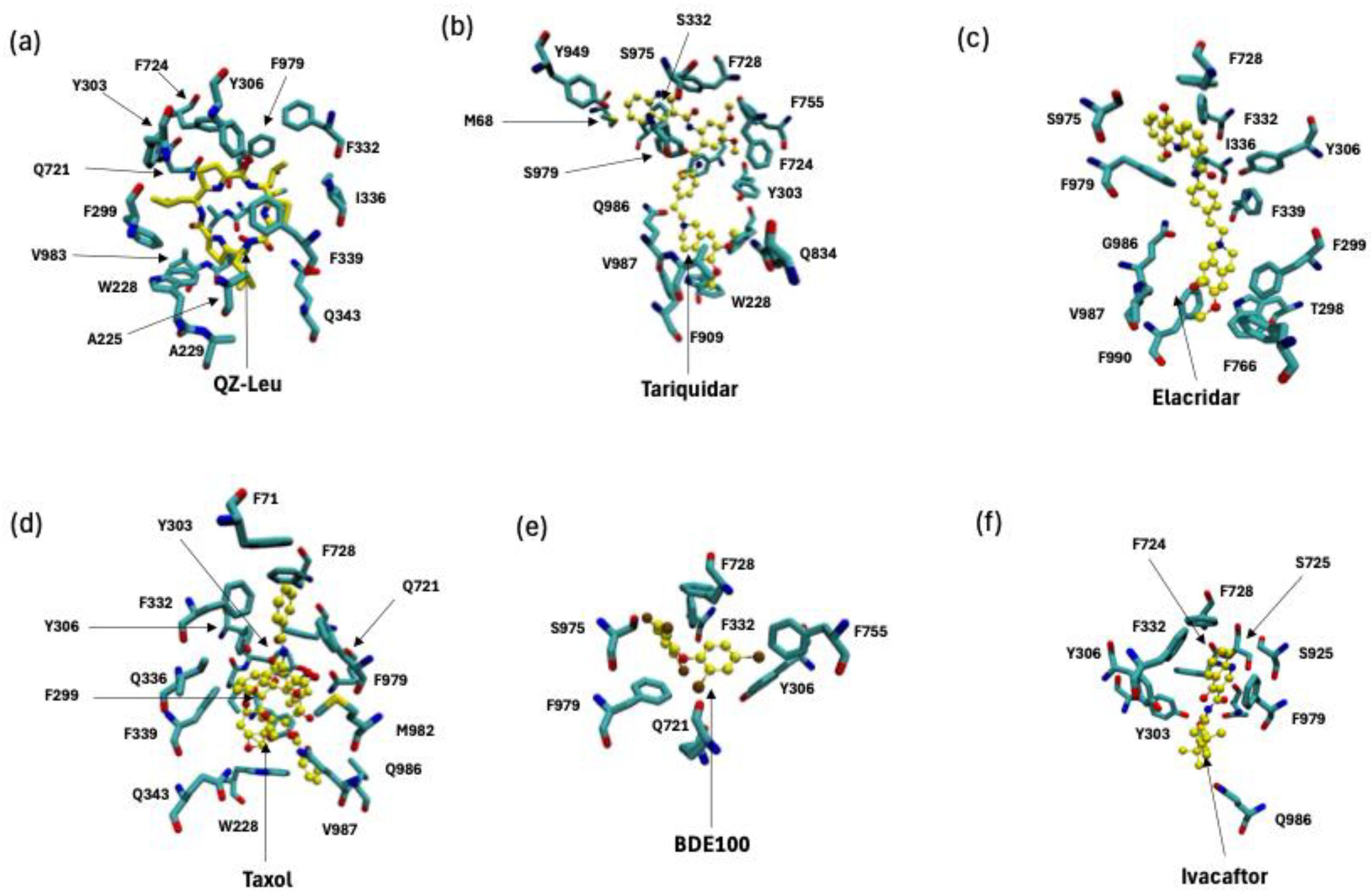
2.1. P-Gp Inhibitor Pharmacology: Multimodality of Inhibition
2.2. P-Glycoprotein Substrate Pharmacology: The Handling of Chemotherapeutics and Environmental Pollutants from PDB Database Analysis
2.3. Assessing IC50 of P-Gp Inhibitors and Substrates with AI
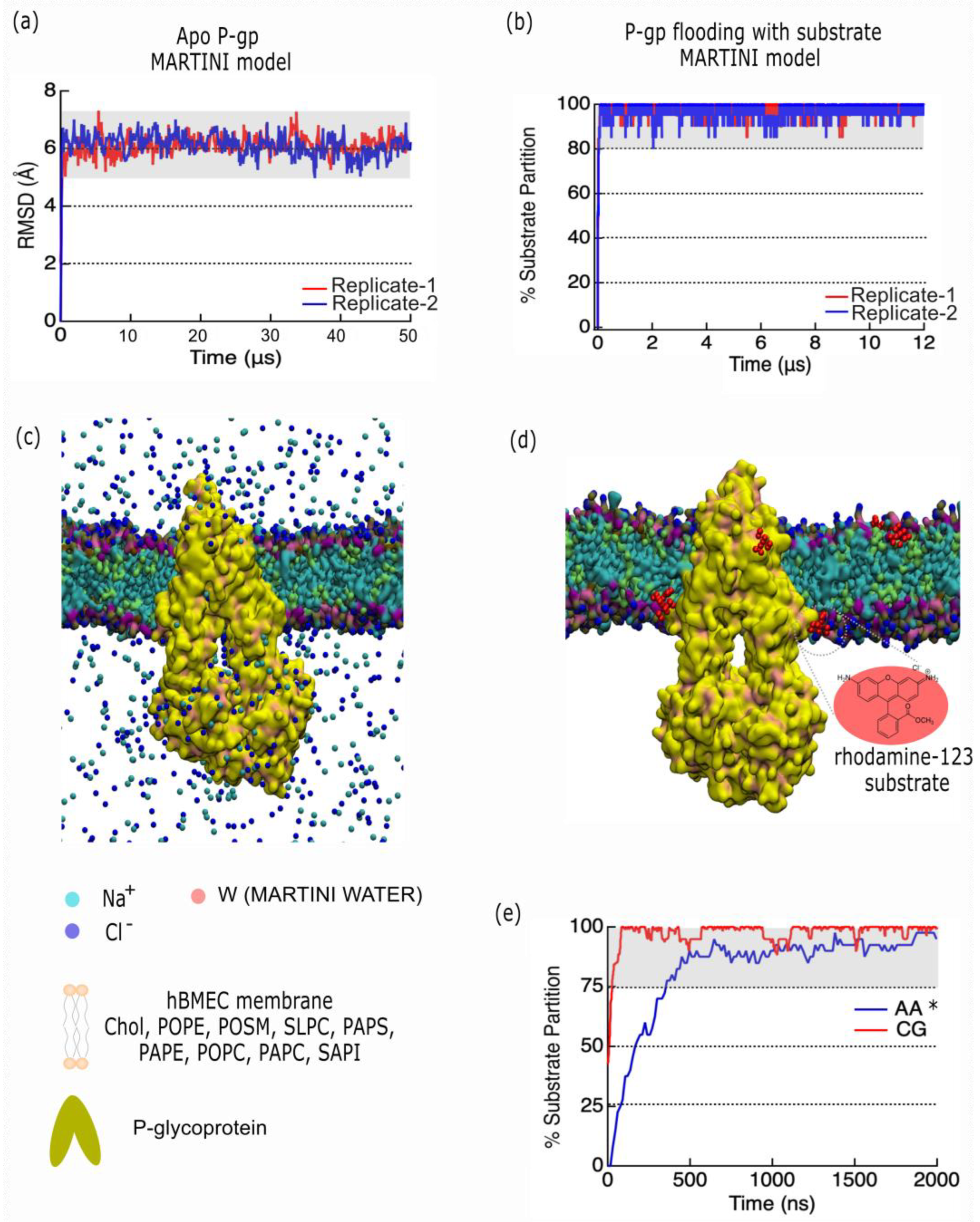
2.4. Developing a P-Glycoprotein Coarse-Grain Model to Explore Substrate Interactions.
3. Discussion
3.1. P-Gp Inhibition Exhibits Multimodal Character
3.2. P-Gp Substrate Pharmacology: The Handling of Chemotherapeutics and Environmental Pollutants from PDB Database Analysis Shows Diversity in Character
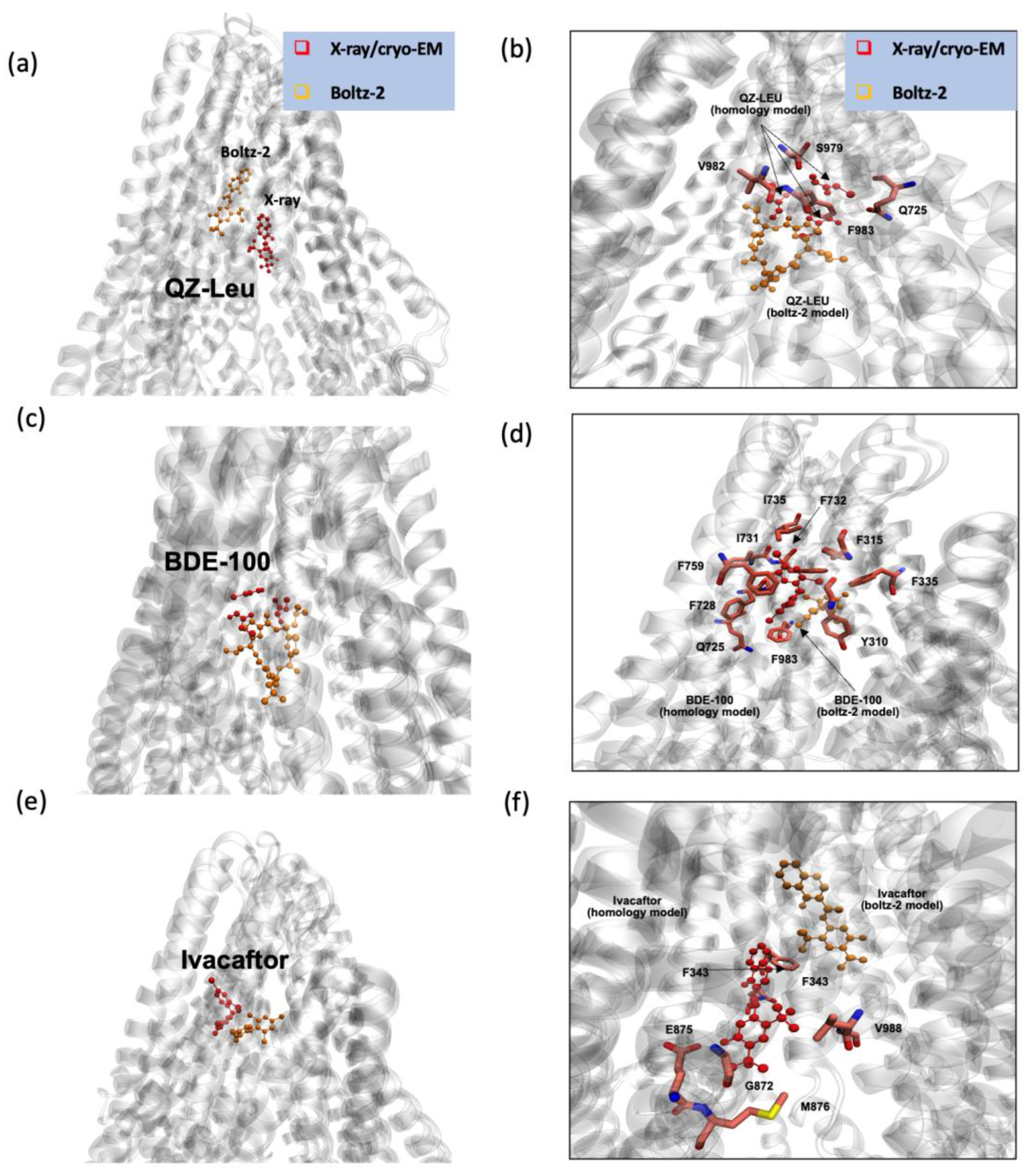
3.3. Boltz-2 AI Suggests That Later Generation Inhibitors Bind More Efficiently.
3.4. P-Gp Coarse-Grained Model
3.5. Conclusion
4. Materials and Methods
4.1. P-Gp Human Homology Models.
4.2. Protein-Ligand Co-Folding Using Boltz-2
4.3. Coarse-Grain Protein-Membrane Model Setup
4.4. Coarse-Grained Molecular Dynamics
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| P-gp | P-glycoprotein |
| ABC | ATP-Binding Cassette |
| BMEC | Brain microvascular endothelial cell |
| BBB | Blood-brain barrier |
| TMD | Transmembrane domain |
| NBD | Nucleotide binding domain |
| MD | Molecular Dynamics |
| CG | Coarse-grain |
| POPC | 1-Palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine |
| POPE | 1-Palmitoyl-2-oleoyl-sn-glycero-3-phosphoethanolamine |
| POSM | Palmitoyl Sphingomyelin |
| SLPC | 1-Stearoyl-2-linoleoyl-sn-glycero-3-phosphocholine |
| PAPS | 1-Palmitoyl-2-arachidonoyl-sn-glycero-3-phosphoserine |
| PAPE | 1-Palmitoyl-2-arachidonoyl-sn-glycero-3-phosphoethanolamine |
| PAPC | 1-Palmitoyl-2-arachidonoyl-sn-glycero-3-phosphocholine |
| SAPI | Stearoyl Arachidonoyl Phosphatidylinositol |
| SMILES | Simplified Molecular Input Line Entry System |
| PDB | Protein Data Bank |
| COM | Center of Mass |
| CFTR | Cystic Fibrosis Transmembrane Conductance Regulator |
| pTM | Predicted TM-score |
| ipTM | Interface Predicted TM-score |
| pLDDT | Predicted Local Distance Difference Test |
| IC50 | Half Maximal Inhibitory Concentration |
| pIC50 | Negative Log of IC50 |
| QMEAN | Qualitative Model Energy Analysis |
| MSA | Multiple Sequence Alignment |
| PME | Particle Mesh Ewald |
| LINCS | Linear Constraint Solver |
| GROMACS | GROningen MAchine for Chemical Simulations |
References
- Wong, A.D.; Ye, M.; Levy, A.F.; Rothstein, J.D.; Bergles, D.E.; Searson, P.C. The Blood-Brain Barrier: An Engineering Perspective. Front Neuroeng 2013, 6, 7. [CrossRef]
- Gottesman, M.M.; Ling, V. The Molecular Basis of Multidrug Resistance in Cancer: The Early Years of P-Glycoprotein Research. FEBS Lett 2006, 580, 998–1009. [CrossRef]
- van Veen, H.W.; Konings, W.N. Structure and Function of Multidrug Transporters. Resolving the Antibiotic Paradox: Progress in Understanding Drug Resistance and Development of New Antibiotics 1998, 145–158.
- Kim, Y.; Chen, J. Molecular Structure of Human P-Glycoprotein in the ATP-Bound, Outward-Facing Conformation. Science (1979) 2018, 359, 915–919. [CrossRef]
- Condic-Jurkic, K.; Subramanian, N.; Mark, A.E.; O’Mara, M.L. The Reliability of Molecular Dynamics Simulations of the Multidrug Transporter P-Glycoprotein in a Membrane Environment. PLoS One 2018, 13, e0191882. [CrossRef]
- Jorgensen, C.; Ulmschneider, M.B.; Searson, P.C. Modeling Substrate Entry into the P-Glycoprotein Efflux Pump at the Blood–Brain Barrier. J Med Chem 2023, 66, 16615–16627. [CrossRef]
- Szewczyk, P.; Tao, H.; McGrath, A.P.; Villaluz, M.; Rees, S.D.; Lee, S.C.; Doshi, R.; Urbatsch, I.L.; Zhang, Q.; Chang, G. Snapshots of Ligand Entry, Malleable Binding and Induced Helical Movement in P-Glycoprotein. Biological Crystallography 2015, 71, 732–741. [CrossRef]
- Jin, M.S.; Oldham, M.L.; Zhang, Q.; Chen, J. Crystal Structure of the Multidrug Transporter P-Glycoprotein from Caenorhabditis Elegans. Nature 2012, 490, 566–569. [CrossRef]
- Aller, S.G.; Yu, J.; Ward, A.; Weng, Y.; Chittaboina, S.; Zhuo, R.; Harrell, P.M.; Trinh, Y.T.; Zhang, Q.; Urbatsch, I.L. Structure of P-Glycoprotein Reveals a Molecular Basis for Poly-Specific Drug Binding. Science (1979) 2009, 323, 1718–1722. [CrossRef]
- Alam, A.; Kowal, J.; Broude, E.; Roninson, I.; Locher, K.P. Structural Insight into Substrate and Inhibitor Discrimination by Human P-Glycoprotein. Science (1979) 2019, 363, 753–756. [CrossRef]
- Xing, J.; Huang, S.; Heng, Y.; Mei, H.; Pan, X. Computational Insights into Allosteric Conformational Modulation of P-Glycoprotein by Substrate and Inhibitor Binding. Molecules 2020, 25, 6006. [CrossRef]
- Loo, T.W.; Bartlett, M.C.; Clarke, D.M. Drug Binding in Human P-Glycoprotein Causes Conformational Changes in Both Nucleotide-Binding Domains. Journal of Biological Chemistry 2003, 278, 1575–1578. [CrossRef]
- McCormick, J.W.; Vogel, P.D.; Wise, J.G. Multiple Drug Transport Pathways through Human P-Glycoprotein. Biochemistry 2015, 54, 4374–4390. [CrossRef]
- Ward, A.B.; Szewczyk, P.; Grimard, V.; Lee, C.-W.; Martinez, L.; Doshi, R.; Caya, A.; Villaluz, M.; Pardon, E.; Cregger, C. Structures of P-Glycoprotein Reveal Its Conformational Flexibility and an Epitope on the Nucleotide-Binding Domain. Proceedings of the National Academy of Sciences 2013, 110, 13386–13391. [CrossRef]
- Callaghan, R.; Ford, R.C.; Kerr, I.D. The Translocation Mechanism of P-Glycoprotein. FEBS Lett 2006, 580, 1056–1063. [CrossRef]
- Prajapati, R.; Sangamwar, A.T. Translocation Mechanism of P-Glycoprotein and Conformational Changes Occurring at Drug-Binding Site: Insights from Multi-Targeted Molecular Dynamics. Biochimica et Biophysica Acta (BBA)-Biomembranes 2014, 1838, 2882–2898. [CrossRef]
- Li, H.; Gong, W. Study of Allosteric Transitions of Human P-Glycoprotein by Using the Two-State Anisotropic Network Model. Front Med (Lausanne) 2022, 9, 815355. [CrossRef]
- De Vecchis, D.; Schäfer, L. V Coupling the Role of Lipids to the Conformational Dynamics of the ABC Transporter P-Glycoprotein. Biophys J 2024, 123, 2522–2536.
- Venable, R.M.; Kramer, A.; Pastor, R.W. Molecular Dynamics Simulations of Membrane Permeability. Chem Rev 2019, 119, 5954–5997. [CrossRef]
- Van Breemen, R.B.; Li, Y. Caco-2 Cell Permeability Assays to Measure Drug Absorption. Expert Opin Drug Metab Toxicol 2005, 1, 175–185.
- Bednarek, R. In Vitro Methods for Measuring the Permeability of Cell Monolayers. Methods Protoc 2022, 5, 17. [CrossRef]
- Jorgensen, C.; Linville, R.M.; Galea, I.; Lambden, E.; Vögele, M.; Chen, C.; Troendle, E.P.; Ruggiu, F.; Ulmschneider, M.B.; Schiøtt, B. Permeability Benchmarking: Guidelines for Comparing in Silico, in Vitro, and in Vivo Measurements. J Chem Inf Model 2025. [CrossRef]
- Nosol, K.; Romane, K.; Irobalieva, R.N.; Alam, A.; Kowal, J.; Fujita, N.; Locher, K.P. Cryo-EM Structures Reveal Distinct Mechanisms of Inhibition of the Human Multidrug Transporter ABCB1. Proceedings of the National Academy of Sciences 2020, 117, 26245–26253. [CrossRef]
- Hamaguchi-Suzuki, N.; Adachi, N.; Moriya, T.; Yasuda, S.; Kawasaki, M.; Suzuki, K.; Ogasawara, S.; Anzai, N.; Senda, T.; Murata, T. Cryo-EM Structure of P-Glycoprotein Bound to Triple Elacridar Inhibitor Molecules. Biochem Biophys Res Commun 2024, 709, 149855. [CrossRef]
- Le, C.A.; Harvey, D.S.; Aller, S.G. Structural Definition of Polyspecific Compensatory Ligand Recognition by P-Glycoprotein. IUCrJ 2020, 7, 663–672. [CrossRef]
- Barbieri, A.; Thonghin, N.; Shafi, T.; Prince, S.M.; Collins, R.F.; Ford, R.C. Structure of ABCB1/P-Glycoprotein in the Presence of the CFTR Potentiator Ivacaftor. Membranes (Basel) 2021, 11, 923. [CrossRef]
- Passaro, S.; Corso, G.; Wohlwend, J.; Reveiz, M.; Thaler, S.; Ram Somnath, V.; Getz, N.; Portnoi, T.; Roy, J.; Stark, H. Boltz-2: Towards Accurate and Efficient Binding Affinity Prediction. BioRxiv 2025, 2025–2026.
- Jorgensen, C.; Ulmschneider, M.B.; Searson, P.C. Atomistic Model of Solute Transport across the Blood–Brain Barrier. ACS Omega 2021, 7, 1100–1112. [CrossRef]
- Jorgensen, C.; Troendle, E.P.; Ulmschneider, J.P.; Searson, P.C.; Ulmschneider, M.B. A Least-Squares-Fitting Procedure for an Efficient Preclinical Ranking of Passive Transport across the Blood–Brain Barrier Endothelium. J Comput Aided Mol Des 2023, 37, 537–549. [CrossRef]
- Wassenaar, T.A.; Ingólfsson, H.I.; Bockmann, R.A.; Tieleman, D.P.; Marrink, S.J. Computational Lipidomics with Insane: A Versatile Tool for Generating Custom Membranes for Molecular Simulations. J Chem Theory Comput 2015, 11, 2144–2155. [CrossRef]
- Souza, P.C.T.; Alessandri, R.; Barnoud, J.; Thallmair, S.; Faustino, I.; Grünewald, F.; Patmanidis, I.; Abdizadeh, H.; Bruininks, B.M.H.; Wassenaar, T.A. Martini 3: A General Purpose Force Field for Coarse-Grained Molecular Dynamics. Nat Methods 2021, 18, 382–388. [CrossRef]
- Periole, X.; Cavalli, M.; Marrink, S.-J.; Ceruso, M.A. Combining an Elastic Network with a Coarse-Grained Molecular Force Field: Structure, Dynamics, and Intermolecular Recognition. J Chem Theory Comput 2009, 5, 2531–2543. [CrossRef]
- Monticelli, L.; Kandasamy, S.K.; Periole, X.; Larson, R.G.; Tieleman, D.P.; Marrink, S.-J. The MARTINI Coarse-Grained Force Field: Extension to Proteins. J Chem Theory Comput 2008, 4, 819–834. [CrossRef]
- Vanommeslaeghe, K.; Hatcher, E.; Acharya, C.; Kundu, S.; Zhong, S.; Shim, J.; Darian, E.; Guvench, O.; Lopes, P.; Vorobyov, I. CHARMM General Force Field: A Force Field for Drug-like Molecules Compatible with the CHARMM All-atom Additive Biological Force Fields. J Comput Chem 2010, 31, 671–690. [CrossRef]
- Nørholm, M.H.H.; von Heijne, G.; Draheim, R.R. Forcing the Issue: Aromatic Tuning Facilitates Stimulus-Independent Modulation of a Two-Component Signaling Circuit. ACS Synth Biol 2015, 4, 474–481. [CrossRef]
- Yusuf, R.; Yau, J.J.Y.; Baker-Valla, G.W.; Lawrence, R.J.; Draheim, R.R. EnvZ Signal Output Correlates with Capacity for Transmembrane Water Encapsulation. bioRxiv 2025, 2023–2025.
- Ma, J.; Biggin, P.C. Substrate versus Inhibitor Dynamics of P-glycoprotein. Proteins: Structure, Function, and Bioinformatics 2013, 81, 1653–1668. [CrossRef]
- Klepsch, F.; Vasanthanathan, P.; Ecker, G.F. Ligand and Structure-Based Classification Models for Prediction of P-Glycoprotein Inhibitors. J Chem Inf Model 2014, 54, 218–229. [CrossRef]
- Weidner, L.D.; Fung, K.L.; Kannan, P.; Moen, J.K.; Kumar, J.S.; Mulder, J.; Innis, R.B.; Gottesman, M.M.; Hall, M.D. Tariquidar Is an Inhibitor and Not a Substrate of Human and Mouse P-Glycoprotein. Drug Metabolism and Disposition 2016, 44, 275–282. [CrossRef]
- Domicevica, L.; Biggin, P.C. Homology Modelling of Human P-Glycoprotein. Biochem Soc Trans 2015, 43, 952–958. [CrossRef]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L. SWISS-MODEL: Homology Modelling of Protein Structures and Complexes. Nucleic Acids Res 2018, 46, W296–W303. [CrossRef]
- Kiefer, F.; Arnold, K.; Künzli, M.; Bordoli, L.; Schwede, T. The SWISS-MODEL Repository and Associated Resources. Nucleic Acids Res 2009, 37, D387–D392. [CrossRef]
- UniProt: The Universal Protein Knowledgebase in 2021. Nucleic Acids Res 2021, 49, D480–D489.
- Kim, S.; Thiessen, P.A.; Bolton, E.E.; Chen, J.; Fu, G.; Gindulyte, A.; Han, L.; He, J.; He, S.; Shoemaker, B.A. PubChem Substance and Compound Databases. Nucleic Acids Res 2016, 44, D1202–D1213. [CrossRef]
- Van Der Spoel, D.; Lindahl, E.; Hess, B.; Groenhof, G.; Mark, A.E.; Berendsen, H.J.C. GROMACS: Fast, Flexible, and Free. J Comput Chem 2005, 26, 1701–1718. [CrossRef]
- Berendsen, H.J.C.; Postma, J.P.M. van; Van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular Dynamics with Coupling to an External Bath. J Chem Phys 1984, 81, 3684–3690. [CrossRef]
- Bussi, G.; Donadio, D.; Parrinello, M. Canonical Sampling through Velocity Rescaling. J Chem Phys 2007, 126. [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A Linear Constraint Solver for Molecular Simulations. J Comput Chem 1997, 18, 1463–1472.
- Consortium, U. UniProt: A Hub for Protein Information. Nucleic Acids Res 2015, 43, D204–D212.
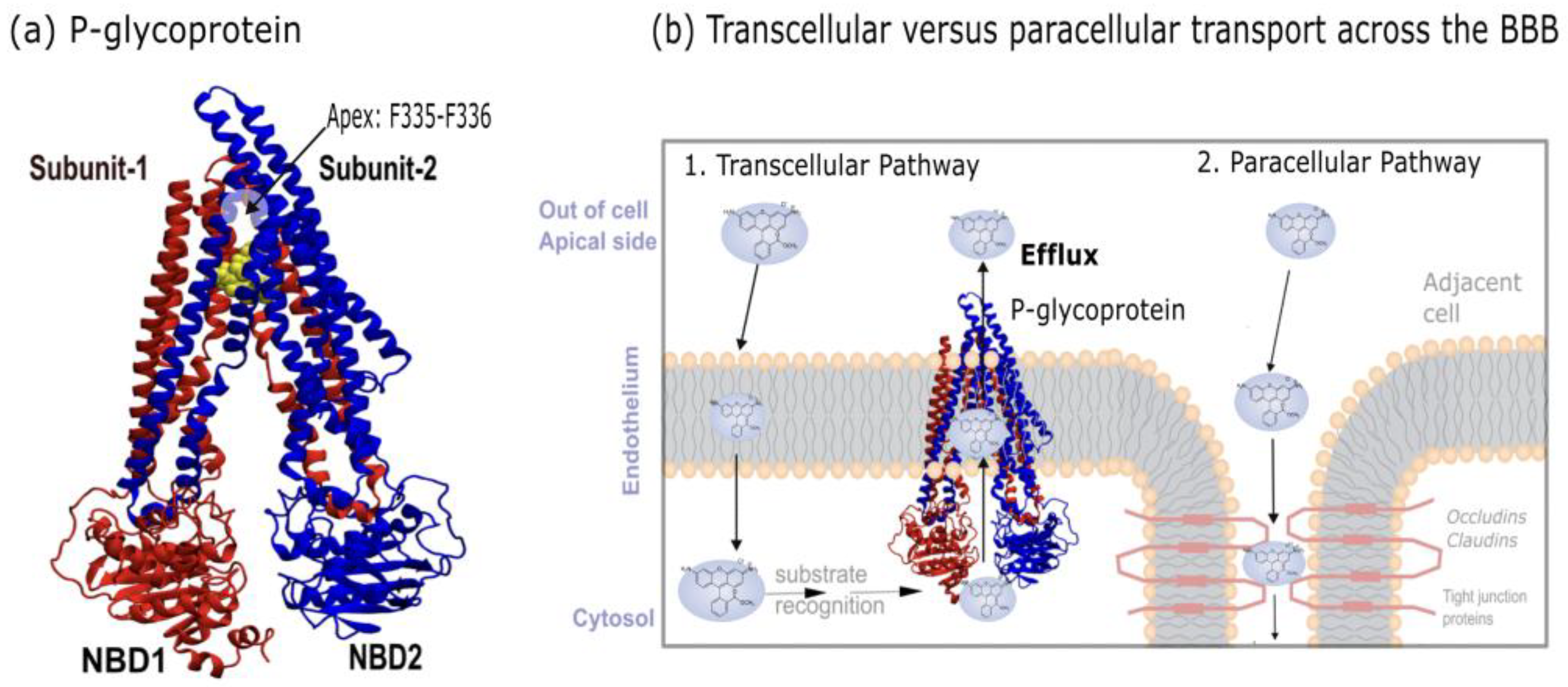
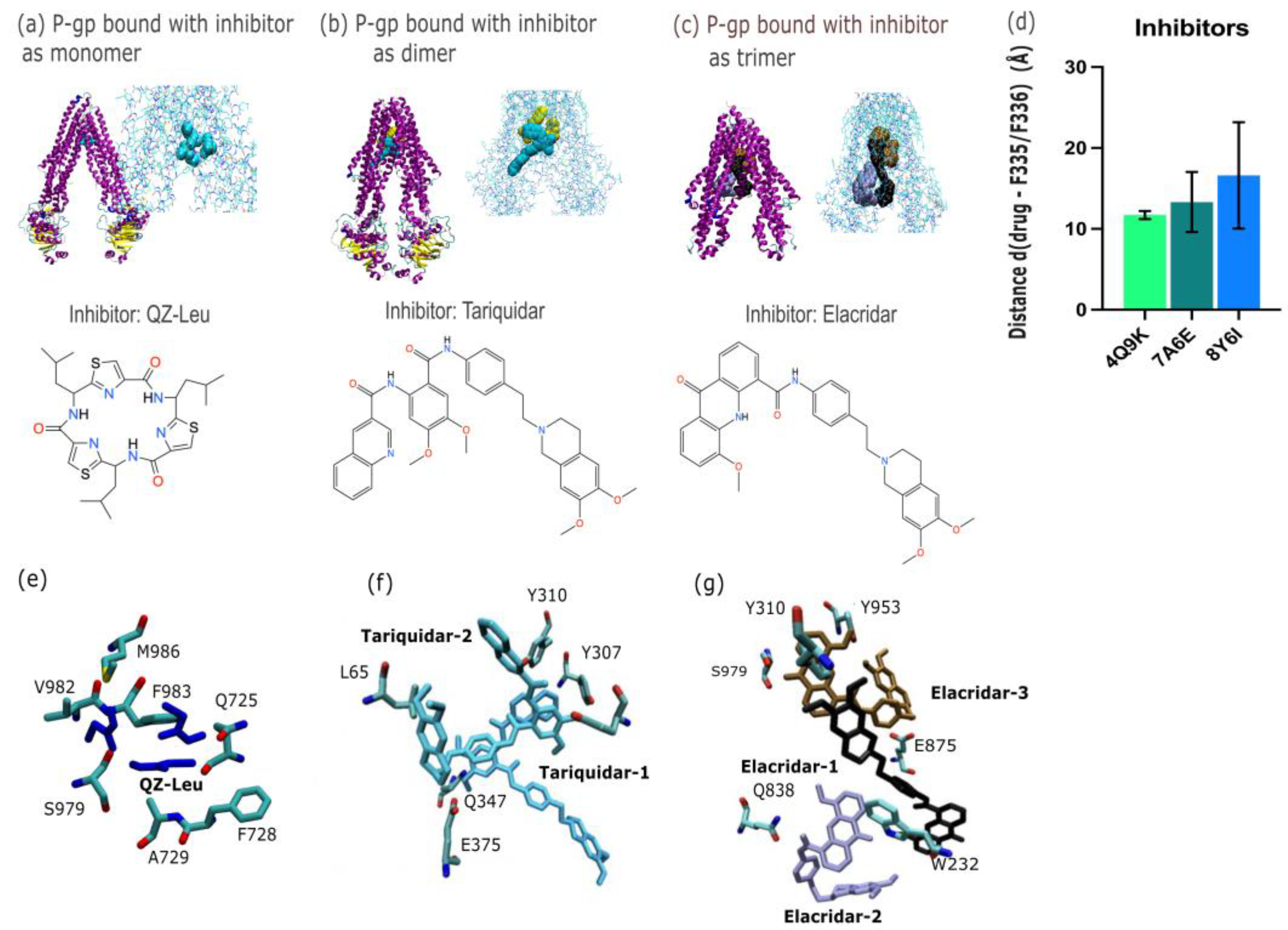
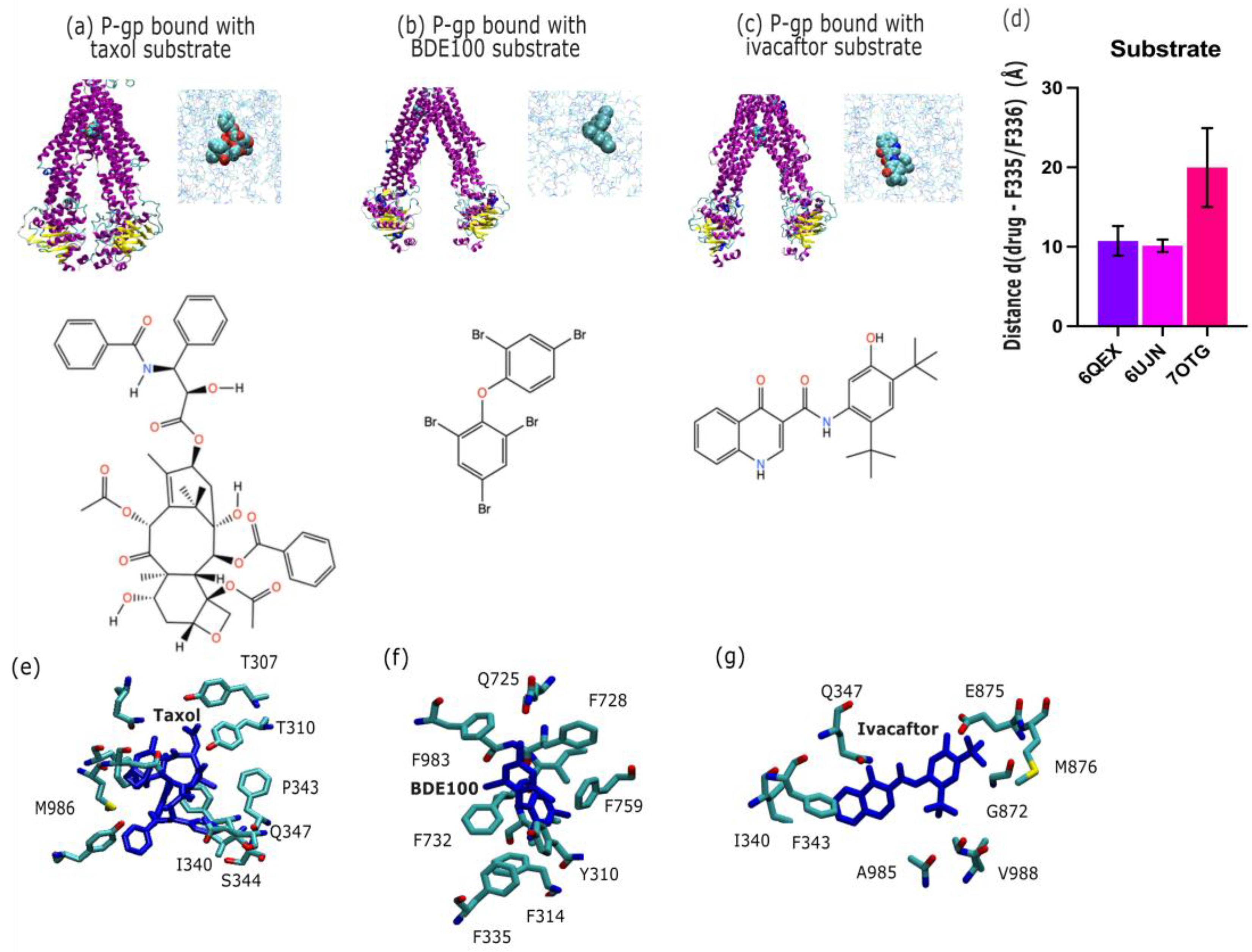
| Inhibitors | Substrates | |||||
|---|---|---|---|---|---|---|
| QZ-Leu | Tariquidar | Elacridar | Taxol | BDE100 | Ivacaftor | |
| Predicted TM-Score (pTM) | 0.763 | 0.772 | 0.781 | 0.780 | 0.792 | 0.774 |
| Interface Predicted TM-Score (ipTM) | 0.890 | 0.911 | 0.933 | 0.892 | 0.779 | 0.915 |
| Confidence Score | 0.743 | 0.785 | 0.784 | 0.755 | 0.781 | 0.777 |
| Average predicted local distance difference test (pLDDT) | 0.706 | 0.753 | 0.746 | 0.721 | 0.782 | 0.743 |
| Affinity Probability | 0.402 | 0.667 | 0.672 | 0.545 | 0.503 | 0.663 |
| Predicted pIC50 | 6.565 | 7.243 | 6.447 | 6.928 | 5.589 | 6.237 |
| Predicted IC50 (nM) | 272.3 | 57.1 | 357.3 | 118.0 | 2580 | 579.4 |
| Lipid | Total bilayer (%) |
|---|---|
| CHOL | 30 (96) |
| POPE | 6 (20) |
| POSM | 19 (61) |
| SLPC | 8 (27) |
| PAPS | 8 (25) |
| PAPE | 15 (47) |
| POPC | 4 (13) |
| PAPC | 8 (27) |
| SAPI | 2 (6) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




