Submitted:
25 August 2025
Posted:
26 August 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Hypothesis
2.1. Supporting Evidence: Examples of How Ectopic Expression of Genes Controlling Differentiation Contribute to Carcinogenesis.
2.1.1. erbB
2.1.2. Anaplastic Lymphoma Kinase (ALK)
2.1.3. Caudal Type Homeobox 2 (CDX2)
2.1.3.1. Gastrointestinal Epithelial Metaplasia and Gastric Cancer
2.1.3.2. Acute Myeloid Leukemia (AML)
2.1.3.3. Relationship with the CCC Hypothesis
2.1.4. NKX2-1
2.1.5. VAV1
2.1.6. RE1-Silencing Transcription Factor (REST)
2.1.7. Sonic Hedgehog (Shh)
2.1.8. OCT4
2.1.9. PAX8
2.1.10. Twist
3. Genes with Carcinogenic Potential Exist Even Without Mutations
3.1. Four Criteria of the CCC Hypothesis
3.1.2. Tal1 Is Ectopically Expressed in T Cell Acute Lymphoblastic Leukemia
4. Implication of the Hypothesis
4.1. Explanation of Atypia
4.2. Explanation of Cancer Malignancy
5. Non-Mutated Drivers of Carcinogenesis
6. Pathological Diversity and Progression of Cancer
7. Explanation of the CCC Hypothesis Regarding Glycosylation Defect and Invasion/Metastasis
8. Relationship Between Various Carcinogenesis Theories and the CCC Hypothesis
8.1. Relationship Between the Mutation Theory and the CCC Hypothesis
8.2. Relationship Between the Multistage Carcinogenesis Hypothesis and CCC Hypothesis
8.3. Relationship Between the Epigenetic Hypothesis and CCC Hypothesis
8.4. Relationship Between the Aberrant Differentiation Hypothesis and CCC Hypothesis
8.5. Relationship Between the Cancer Stem Cell Hypothesis and CCC Hypothesis
8.6. Relationship Between the Clonal Evolution Hypothesis and CCC Hypothesis
9. Applications of the CCC Hypothesis
10. Differences Between Benign and Malignant Tumors and the Inevitability of Carcinogenesis in Multicellular Organisms
11. Metastasis and Organ Specificity
12. Methods for Verifying the CCC Hypothesis
13. Relationship Between Cancer Plasticity and the CCC Hypothesis
14. Repositioning Tumor Suppressor Genes in the CCC Hypothesis
15. Conclusion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CCC | Carcinogenic Chimera Cell |
| EMT | Epithelial–Mesenchymal Transition |
| CSC | Cancer Stem Cell |
| HPA | Human Protein Atlas |
| EGFR | Epidermal Growth Factor Receptor |
| AML | Acute Myeloid Leukemia |
| T-ALL | T-cell Acute Lymphoblastic Leukemia |
| iPS | Induced Pluripotent Stem Cell |
| MET | Mesenchymal–Epithelial Transition |
| DN | Double Negative (T cell developmental stage) |
| DP | Double Positive (T cell developmental stage) |
References
- Hida, T.; Ueda, R.; Sekido, Y.; Hibi, K.; Matsuda, R.; Ariyoshi, Y.; Sugiura, T.; Takahashi, T.; Takahashi, T. Ectopic expression of c-kit in small-cell lung cancer. Int. J. Cancer Suppl. 1994, 8, 108–109. [Google Scholar] [CrossRef]
- Graham, D.K.; Salzberg, D.B.; Kurtzberg, J.; Sather, S.; Matsushima, G.K.; Keating, A.K.; Liang, X.; Lovell, M.A.; Williams, S.A.; Dawson, T.L.; Schell, L.J.; Anwar, A.A.; Snodgrass, H.R.; Earp, H.S. Ectopic expression of the proto-oncogene Mer in pediatric T-cell acute lymphoblastic leukemia. Clin. Cancer Res. 2006, 12, 2662–269. [Google Scholar] [CrossRef]
- Yamamoto, H.; Hatano, M.; Iitsuka, Y.; Mahyar, N.S.; Yamamoto, M.; Tokuhisa, T. Two forms of Hox11 a T cell leukemia oncogene, are expressed in fetal spleen but not in primary lymphocytes. Mol. Immunol. 1995, 32, 1177–1182. [Google Scholar] [CrossRef]
- Choi, K.Y.; Chang, K.; Pickel, J.M.; Badger, J.D. 2nd; Roche, K.W. 2nd; Roche, K.W. Expression of metabotropic glutamate receptor 5 (mGluR5) induces melanoma in transgenic mice. Proc. Natl. Acad. Sci. USA. 2011, 108, 15219–15224. [Google Scholar] [CrossRef] [PubMed]
- Keilholz, U.; Menssen, H.D.; Gaiger, A.; Menke, A.; Oji, Y.; Oka, Y. Scheibenbogen, C.; Stauss, H.; Thiel, E.; Sugiyama, H. Wilms tumor gene 1 (WT1) in human neoplasia. Leukemia. 2005, 19, 1318–1323. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, T.; Toyoshima, K. The ErbB-related proto-oncogene genes encoding growth factor receptors. Gan To Kagaku Ryoho. 1987, 14, 2075–2082. [Google Scholar] [PubMed]
- Heldin, C.H.; Westermark, B. Mechanism of action and in vivo role of platelet-derived growth factor. Physiol Rev. 1999, 79, 1283–1316. [Google Scholar] [CrossRef]
- The Human Protein Atlas. ALK [Internet]. Available from: https://www.proteinatlas.org/ENSG00000171094-ALK.
- Tsuyama, N.; Sakamoto, K.; Sakata, S.; et al. . Anaplastic large cell lymphoma: pathology,genetics, and clinical aspects. J. Clin. Exp. Hematop. 2017, 57, 120―142. [Google Scholar] [CrossRef]
- Du, X.; Shao, Y.; Qin, H-F.; Tai, Y-H.; Gao, H-J. ALK rearrangements in non-small-cell lung cancer (NSCLC). Transl. Lung Cancer Res. 2018, 9, 423–430. [Google Scholar] [CrossRef]
- The Human Protein Atlas. CDX2 [Internet]. Available from: https://www.proteinatlas.org/ENSG00000165556-CDX2.
- Mutoh, H.; Hakamata, Y.; Sato, K.; Eda, A.; Yanaka, I.; Honda, S.; Osawa, H.; Kaneko, Y.; Sugano, K. Conversion of gastric mucosa to intestinal metaplasia in Cdx2-expressing transgenic mice. Biochem. Biophys. Res. Commun. 2002, 294, 470–479. [Google Scholar] [CrossRef]
- Mutoh, H.; Sakurai, S.; Satoh, K.; Tamada, K.; Kita, H.; Osawa, H.; Tomiyama, T.; Sato, Y.; Yamamoto, H.; Isoda, N.; Yoshida, T.; Ido, K.; Sugano, K. Development of gastric carcinoma from intestinal metaplasia in Cdx2-transgenic mice. Cancer Res. 2004, 64, 7740–7747. [Google Scholar] [CrossRef]
- Voutsadakis, I.A. Gastric adenocarcinomas with CDX2 induction show a higher frequency of TP53 and KMT2B mutations and MYC amplification but similar survival compared to cancers without CDX2 induction. J. Clin. Med. 2024, 13, 7635. [Google Scholar] [CrossRef] [PubMed]
- Khayyat, A.; Esmaeil Pour, M.A.; Poursina, O.; Zohouri, S.A.; Jian, P.V.; Patel, N.; Amin, A. Evaluation of the biomarkers CDX1 and CDX2 in gastric cancer prognosis: A meta-analysis. Int. J. Mol. Cell Med. 2024, 13, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Scholl, C.; Bansal, D.; Döhner, K.; Eiwen, K.; Huntly, B.J.P.; Lee, B.H.; Rücker, F.G.; Schlenk, R.F.; Bullinger, L.; Döhner, H.; Gilliland, D.G.; Fröhling, S. The homeobox gene, CDX2, is aberrantly expressed in most AML cases of acute myeloid leukemia and promotes leukemogenesis. J. Clin. Invest. 2007, 117, 1037–1048. [Google Scholar] [CrossRef]
- Rawat, V.P.S.; Cusan, M.; Deshpande, A.; Hiddemann, W.; Quintanilla-Martinez, L.; Humphries, R.K.; Bohlander, S.K.; Feuring-Buske, M.; Buske, C. Ectopic expression of the homeobox gene Cdx2 is the transforming event in a mouse model of t(12;13)(p13;q12) acute myeloid leukemia. Proc. Natl. Acad. Sci. USA. 2004, 101, 817–822. [Google Scholar] [CrossRef]
- The Human Protein Atlas. NKX2-1 [Internet]. Available from: https://www.proteinatlas.org/ENSG00000136352-NKX2-1.
- Fukagawa, K.; Takahashi, Y.; Yamamichi, N.; Kageyama-Yahara, N.; Sakaguchi, Y.; Obata, M.; Cho, R.; Sakuma, N.; Nagao, S.; Miura, Y.; Tamura, N.; Ohki, D.; Mizutani, H.; Yakabi, S.; Minatsuki, C.; Niimi, K.; Tsuji, Y.; Yamamichi, M.; Shigi, N.; Tomida, S.; Abe, H.; Ushiku, T.; Koike, K.; Fujishiro, M. Transcriptome analysis has revealed the essential role of NK2 homeobox 1/thyroid transcription factor 1 (NKX2-1/TTF-1) in fundic gland-type gastric adenocarcinoma. Gastric Cancer. 2023, 26, 44–54. [Google Scholar] [CrossRef]
- Homminga, I.; Pieters, R.; Langerak, A.W.; de Rooi, J.J.; Stubbs, A.; Verstegen, M.; Vuerhard, M.; Buijs-Gladdiness, J.; Kooi, C.; Klous, P.; van Vlierberghe, V.; Ferrando, A.A.; Cayuela, J.M.; Verhaaf, B.; Berna Beverloo, H.; Horstmann, M.; de Haas, V.; Weikmeijer, A.-S.; Pike-Overzet, K.; Staal, F.J.T.; de Laat, W.; Soulier, J.; Sigaux, F.; Meijerink, J.P.P. Integrated transcript and genome analyses have revealed that NKX2-1 and MEF2C are potential oncogenes in T-cell acute lymphoblastic leukemia. Cancer Cell. 2011, 19, 484–497. [Google Scholar] [CrossRef]
- The Human Protein Atlas. VAV1 [Internet]. Available from: https://www.proteinatlas.org/ENSG00000141968-VAV1.
- Fernandez-Zapico, M.E.; Gonzalez-Paz, N.C.; Weiss, E.; Savoy, D.N.; Molina, J.R.; Fonseca, R.; Smyrk, T.C.; Chari, S.T.; Urrutia, R.; Billadeau, B.B. The ectopic expression of VAV1 plays an unexpected role in pancreatic cancer tumorigenesis. Cancer Cell. 2005, 7, 39–49. [Google Scholar] [CrossRef]
- Coulson, J.M. Transcriptional regulation: Cancer, neurons, and REST. Review Curr. Biol. 2005, 15, R665–R668. [Google Scholar] [CrossRef]
- The Human Protein Atlas. SHH [Internet]. Available from: https://www.proteinatlas.org/ENSG00000164690-SHH.
- Sari, I.N.; Phi, L.T.H.; Jun, N.; Lee, S.; Kwon, H.Y. Hedgehog signaling in cancer: A prospective therapeutic target for eradicating cancer stem cells. Cells. 2018, 7, 208. [Google Scholar] [CrossRef]
- The Human Protein Atlas. POU5F1 (OCT4 [Internet]. Available at: https://www.proteinatlas.org/ENSG00000204531-POU5F1.
- von Eyben, F.E.; Kristiansen, K.; Kapp, D.S.; Hu, R.; Preda, O.; Nogales, F.F. Epigenetic regulation of driver genes involved in testicular tumorigenesis. Int. J. Mol. Sci. 2023, 24, 4148. [Google Scholar] [CrossRef]
- Herreros-Villanueva, M.; Bujanda, L.; Billadeau, D.D.; Zhang, J.-S. Embryonic stem cell factors and pancreatic cancer. World J. Gastroenterol. 2014, 20, 2247–2254. [Google Scholar] [CrossRef] [PubMed]
- Pádua, D.; Figueira, P.; Ribeiro, I.; Almeida, R.; Mesquita, P. Relevance of transcription factors in gastric and colorectal cancer stem cell identification and eradication. Front. Cell Dev. Biol. 2020, 8, 442. [Google Scholar] [CrossRef] [PubMed]
- The Human Protein Atlas. PAX8 [Internet]. Available from: https://www.proteinatlas.org/ENSG00000125618-PAX8.
- Di Palma, T.; Zannini, M. PAX8 as a potential target for ovarian cancer: what we know so far. Onco Targets Ther. 2022, 15, 1273–1280. [Google Scholar] [CrossRef] [PubMed]
- Lamouille, S.; Xu, J.; Derynck, R. Molecular mechanisms of epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2014, 15, 178–196. [Google Scholar] [CrossRef]
- Zhao, Z.; Rahman, M.A.; Chen, Z.G.; Shin, D.M. Multiple biological functions of Twist1 in various cancers. Oncotarget. 2017, 8, 20380–20393. [Google Scholar] [CrossRef]
- The Human Protein Atlas. PAX8 [Internet]. Available from: https://www.proteinatlas.org/ENSG00000125618-PAX8. [CrossRef]
- Takahashi,K.;Yamanaka,S.;Induction of pluripotent stem cells from mouse embryonic and adult fibroblast culyures by defined factors. Cell. 2006, 126, 663-676. [CrossRef]
- Silberg, D.G.; Sullivan, J.; Kang, E.; Swain, G.P.; Moffett, J.; Sund, N.J.; Sackett, S.D.; Kaestner, K.H. Cdx2 ectopic expression induces gastric intestinal metaplasia in transgenic mice. Gastroenterology. 2002, 122, 689–696. [Google Scholar] [CrossRef]
- Graf, T. Historical origins of transdifferentiation and reprogramming. Cell Stem Cell. 2011, 9504–516. [Google Scholar] [CrossRef]
- Horb, M.E.; Shen, C.N.; Tosh, D.; Slack, J.M. Experimental conversion of liver to pancreas. Curr. Biol. 2003, 13, 105–115. [Google Scholar] [CrossRef]
- Liu, Z.; Fan, H.; Li, Y.; Zheng, S.G. Experimental studies on the differentiation of fibroblasts into myoblasts induced by MyoD genes in vitro. Int. J. Biomed. Sci. 2008, 4, 14–19. [Google Scholar] [CrossRef]
- Liu, F.; Zhang, Y.; Chen, F.; Yuan, J.; Li, S.; Han, S.; Lu, D.; Geng, J.; Rao, Z.; Sun, L.; Xu, J.; Shi, Y.; Wang, X.; Liu, Y. Neurog2 directly converts astrocytes into functional neurons in midbrain and spinal cord. Cell Death Dis. 2021, 12, 225. [Google Scholar] [CrossRef]
- Cui, M.; Huang, J.; Zhang, S.; Liu, Q.; Liao, Q.; Qiu, X. Immunoglobulin expression in cancer cells and its critical roles in tumorigenesis. Front. Immunol. 2021, 12, 613530. [Google Scholar] [CrossRef]
- Ho, I.C.; Hodge, M.R.; Rooney, J.W.; Glimcher, L.H. The proto-oncogene c-maf is responsible for tissue-specific expression of interleukin-4. Cell. 1996, 85, 973–983. [Google Scholar] [CrossRef] [PubMed]
- DeRocco, S.E.; Iozzo, R.; Ma, X.P.; Schwarting, R.; Peterson, D.; Calabretta, B. Ectopic expression of A-myb in transgenic mice causes follicular hyperplasia and enhanced B lymphocyte proliferation. Proc. Natl. Acad. Sci. USA. 1997, 94, 3240–3244. [Google Scholar] [CrossRef] [PubMed]
- Neale, G.A.; Rehg, J.E.; Goorha, R.M. Ectopic expression of rhombotin-2 causes selective expansion of CD4-CD8- lymphocytes in the thymus and T-cell tumors in transgenic mice. Blood. 1995, 86, 3060–3071. [Google Scholar] [CrossRef]
- Neale, G.A.; Rehg, J.E.; Goorha, R.M. Disruption of T-cell differentiation precedes T-cell tumor formation in LMO-2 (rhombotin-2) transgenic mice. Leukemia. 1997, 11 Suppl 3, 289–290. [Google Scholar]
- Fu, X.; Kamps, M.P. E2a-Pbx1 induces aberrant expression of tissue-specific and developmentally regulated genes when expressed in NIH 3T3 fibroblasts. Mol. Cell Biol. 1997, 17, 1503–1512. [Google Scholar] [CrossRef]
- Souabni, A.; Jochum, W.; Busslinger, M. Oncogenic role of Pax5 in the T-lymphoid lineage upon ectopic expression from the immunoglobulin heavy-chain locus. Blood. 2007, 109, 281–289. [Google Scholar] [CrossRef]
- Hofmann, T.J.; Cole, M.D. The TAL1/Scl basic helix-loop-helix protein blocks myogenic differentiation and E-box dependent transactivation. Oncogene. 1996, 13, 617–624. [Google Scholar]
- Wang, X.T.; Wei, W.Y.; Kong, F.B.; Lian, C.; Luo, W.; Xiao, Q.; Xie, Y-B. Prognostic significance of Cdx2 immunohistochemical expression in gastric cancer: a meta-analysis of published literatures. J. Exp. Clin. Cancer Res. 2012, 31, 98. [Google Scholar] [CrossRef]
- Basati, G.; Mohammadpour, H.; Razavi, A.E. Association of high expression levels of SOX2, NANOG, and OCT4 in gastric cancer tumor tissues with progression and poor prognosis. J. Gastrointest. Cancer. 2020, 51, 41–47. [Google Scholar] [CrossRef]
- Scholl, C.; Bansal, D.; Döhner, K.; Eiwen, K.; Huntly, B.J.P.; Lee, B.H.; Rucker, F.G.; Schlenk, R.F.; Bullinger, L.; Dohner, H.; Gilliland, D.G.; Frohling, S. The homeobox gene CDX2 is aberrantly expressed in most cases of acute myeloid leukemia and promotes leukemogenesis. J. Clin. Invest. 2007, 117, 1037–1048. [Google Scholar] [CrossRef] [PubMed]
- Tan, T.K.; Zhang, C.; Sanda, T. Oncogenic transcriptional program driven by TAL1 in T-cell acute lymphoblastic leukemia. Int. J. Hematol. 2019, 109, 5–17. [Google Scholar] [CrossRef]
- Fearon, E.R.; Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell. 1990, 61, 759–767. [Google Scholar] [CrossRef]
- Feinberg, A.P.; Tycko, B. The history of cancer epigenetics. Nat. Rev.Cancer. 2004, 4, 143–153. [Google Scholar] [CrossRef] [PubMed]
- Clarke, M.F. Clinical and Therapeutic Implications of Cancer Stem Cells. N.Engl.J.Med. 2019, 380, 2237–2245. [Google Scholar] [CrossRef]
- Nguyen, L.V.; Vanner, R; Dirks, P.; Eaves, C.J. Cancer stem cells:an evolving concept. Nat. Rev. Cancer. 2012, 12, 133–143. [Google Scholar] [CrossRef]
- Swanton, C.; Bernard, E.; Abbosh, C.; André, F.; Auwerx, J.; Balmain, A.; Bar-Sagi, D.; Bernards, R.; Bullman, S.; DeGregori, J.; Elliott, C.; Erez, A.; Evan, G.; Febbraio, M.A.; Hidalgo, A.; Jamal-Hanjani, M.; Joyce, JA.; Kaiser, M.; Lamia, K.; Locasale, J.W.; Loi, S.; Malanchi, I.; Merad, M.; Musgrave, K.; Patel, K.J.; Quezada, S.; Wargo, J.A.; Weeraratna, A.; White, E.; Winkler, F.; Wood, J.N.; Vousden, K.H.; Hanahan, D. Embracing cancer complexity:Hallmarks of systemic disease. Cell. 2024, 187, 1589–1616. [Google Scholar] [CrossRef]
- Wang, M.L.; Chiou, S.H.; Wu, C.W. Targeting cancer stem cells: emerging role of Nanog transcription factor. Onco Targets Ther. 2013, 6, 1207–1220. [Google Scholar] [CrossRef]
- Zhang, Q.; Han, Z.; Zhu, Y.; Chen, J.; Li, W. The role and specific mechanism of OCT4 in cancer stem cells: a review. Int. J. Stem Cells. 2020, 13, 312–325. [Google Scholar] [CrossRef]
- Swain, N.; Thakur, M.; Pathak, J.; Swain, B. SOX2, OCT4 and NANOG: The core embryonic stem cell pluripotency regulators in oral carcinogenesis. J. Oral Maxillofac. Pathol. 2020, 24, 368–373. [Google Scholar] [CrossRef] [PubMed]
- Nelson, T.A.; Wang, N. Targeting lung cancer brain metastases: a narrative review of emerging insights for anaplastic lymphoma kinase (ALK)-positive disease. Transl. Lung Cancer Res. 2023, 12, 379–392. [Google Scholar] [CrossRef] [PubMed]
- Li, X.Q.; Lu, J.T.; Tan, C.C.; Wang, Q.S.; Feng, Y.M. RUNX2 promotes breast cancer bone metastasis by increasing integrin α5-mediated colonization. Cancer Lett. 2016, 380, 78–86. [Google Scholar] [CrossRef]
- Nagai, E.; Ueyama, T.; Yao, T.; Tsuneyoshi, M. Hepatoid adenocarcinoma of the stomach. A clinicopathologic and immunohistochemical analysis. Cancer. 1993, 72, 1827–1835. [Google Scholar] [CrossRef] [PubMed]
- Laisne,M,;lupien,M,;vallot.C,;Epigenomic heterogeneity as a source of tumour evolution.Nat.Rev.cancer. 2025, 25, 7-26. [CrossRef]
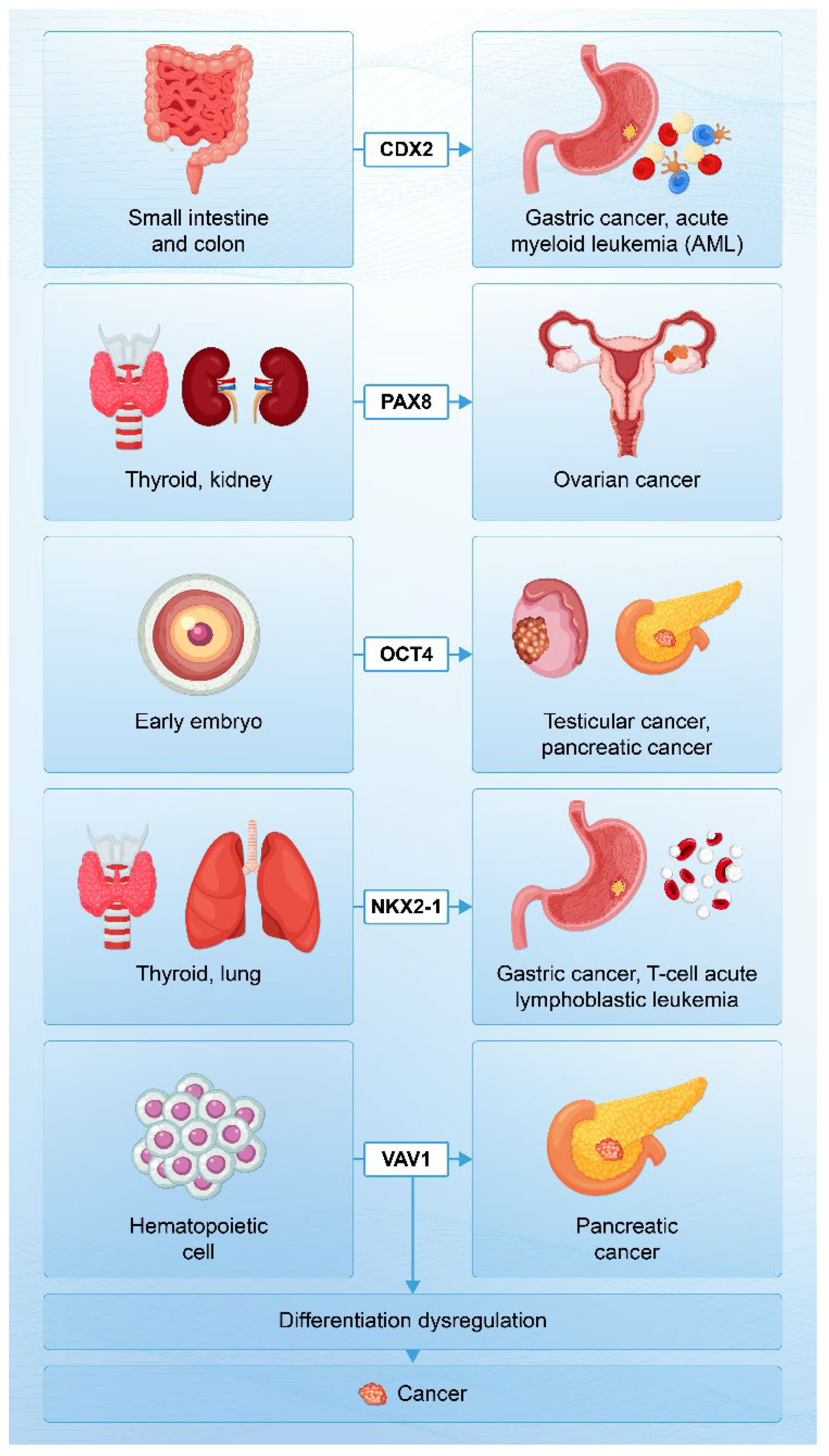
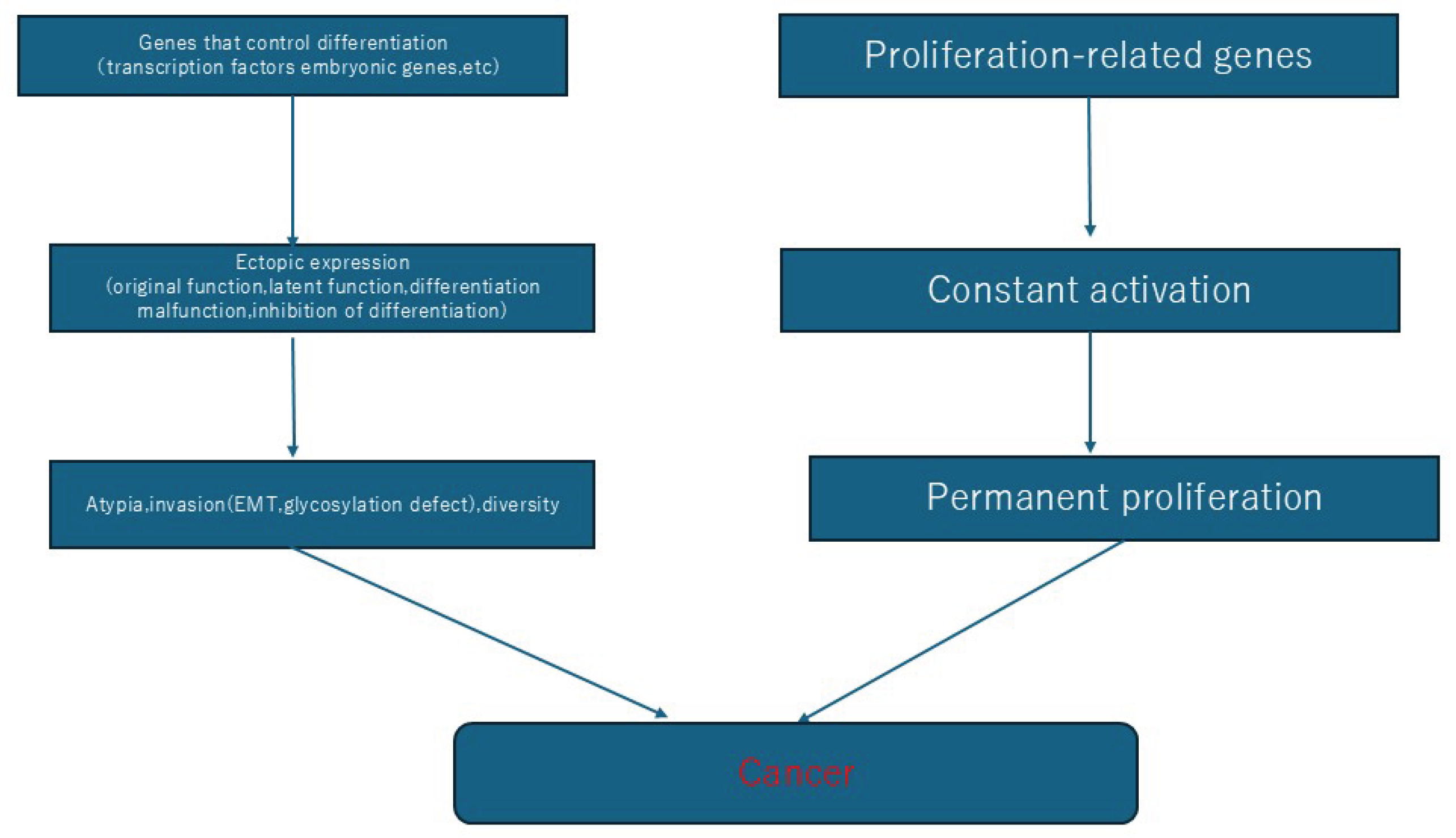
| Gene name | Normal expression site | Related cancers | Notes |
| erbB | Epithelial |
Erythroblastosis |
Not expressed in erythroblasts |
| ALK | Specific neurons in the brain and nervous system during development |
Anaplastic large cell lymphoma, small cell lung carcinoma | Small cell lung carcinoma expressing ALK are prone to metastasizing to the brain. |
| CDX2 | Small intestine, colon |
Gastric cancer, acute myeloid leukemia |
Not expressed in bone marrow and gastric mucosa |
| NKX2-1 | Thyroid, lung |
Gastric adenocarcinoma of fundic gland, T-cell acute lymphoblastic leukemia | Gastric adenocarcinoma of fundic gland also induces lung-specific surfactant proteins. |
| VAV1 | Expressed in hematopoietic cells |
Pancreatic cancer |
Expressed as wild type due to low methylation |
| REST | A transcription repressor that suppresses the expression of neuro-specific genes in non-neural cells |
Small cell lung carcinoma | Tumor suppressor gene |
| Shh | Expressed during embryogenesis |
Basal cell carcinoma, pancreatic cancer, lung cancer |
Not expressed in somatic cells |
| OCT4 | Embryonic |
Testicular germ cell tumors, pancreatic cancer | Not expressed in somatic cells |
| PAX8 | Thyroid, kidney, ureter, etc. | Ovarian cancer | Not expressed in the surface epithelium of the ovary |
| Twist | Embryonic | Breast cancer, ovarian cancer, prostate cancer, pancreatic cancer, etc. | Induces epithelial-mesenchymal transition |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




