Submitted:
04 August 2025
Posted:
06 August 2025
You are already at the latest version
Abstract
Keywords:
Chapter One Introduction
1.1. Background Information
1.2. Statement of the Problem
1.3. Objective of the Project
1.3.1. General Objective
1.3.2. Specific Objective of the Project
- To study different deep learning models.
- To design and implement user management functionalities, allowing different roles (patients, medical professionals, administrators) to access the system securely.
- To develop modules for documenting patient histories and initial diagnoses, ensuring comprehensive and accurate record-keeping.
- To implement a referral system for MRI scans, enabling medical professionals to refer patients seamlessly and track the status of referrals.
- To integrate deep learning model for brain tumor classification and segmentation, enhancing diagnostic accuracy.
- To establish a specialist referral system, enabling medical professionals to seek expert opinions for complex cases and manage specialist responses efficiently.
- To conduct comprehensive testing and evaluation of Brain tumor Track to ensure usability, reliability, and effectiveness in real-world clinical settings.
- To enhance the accuracy of brain tumor detection using deep learning algorithms.
- To reduce the time required for diagnosing brain tumors by automating image analysis.
- To integrate patient follow-up features which facilitate timely communication between patients and healthcare providers.
- To provide educational resources to improve patient understanding and adherence to treatment plans.
1.4. Scope of the Project
- Brain tumor Detection: The project will focus on developing a deep learning- based brain tumor detection model that can accurately identify and classify brain tumors in MRI images.
- Patient Data Follow Up: The project will develop a patient data management system that can store and manage patient data, including medical history, treatment plans, and test results.
1.5. Significance of the Project
- Improve brain tumor detection accuracy and reduce false positives.
- Enhance patient outcomes and quality of life through timely and accurate diagnosis.
- Reduce the burden on radiologists and doctors through automation.
- Contribute to the development of innovative deep learning and deep learning techniques in medical imaging.
- Facilitate the communication of specialists with doctors or radiologists (technician) which found in rural areas.
1.6. Outline of the Project
Chapter Two Literature Review
- Introduction
2.1. Theoretical Review
2.2. Empirical Review
2.1.1. Patient Follow-up Web Applications Study on Patient Engagement and Usability:
- Research on Healthcare Utilization and Cost Reduction:
2.1.2. Tumor Detection Using Deep Learning Application of Deep Learning in Tumor Detection:
- Study on Image Segmentation and Classification:
2.3. Research Gap
Chapter Three Method and Design
3.1. Introduction
3.2. Data Collection
3.2.1. Methods Used to Collect Data
3.2.2. Data Sources
3.4. Software Development
3.4.1. Software Development Method
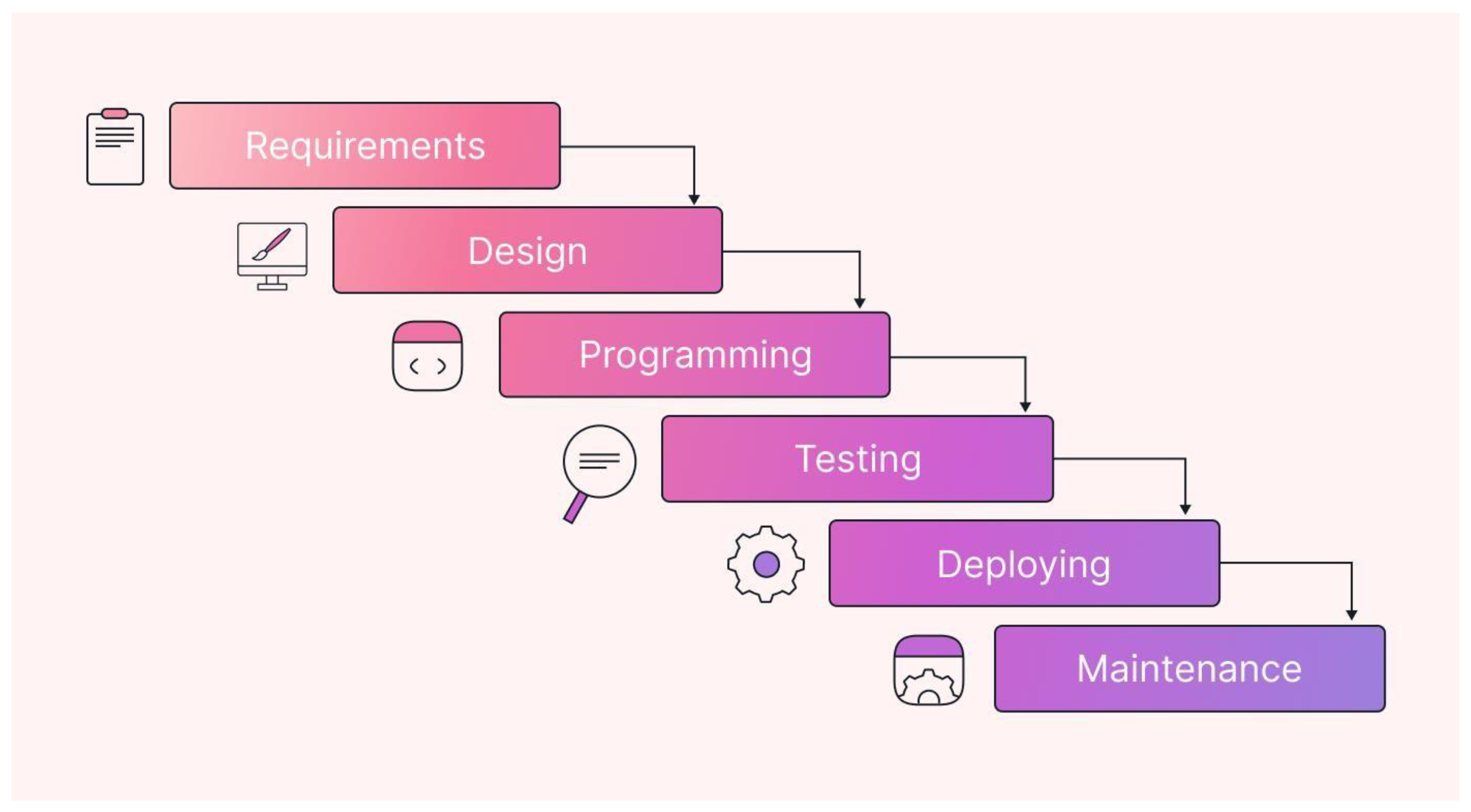
3.4.2. Requirements Analysis
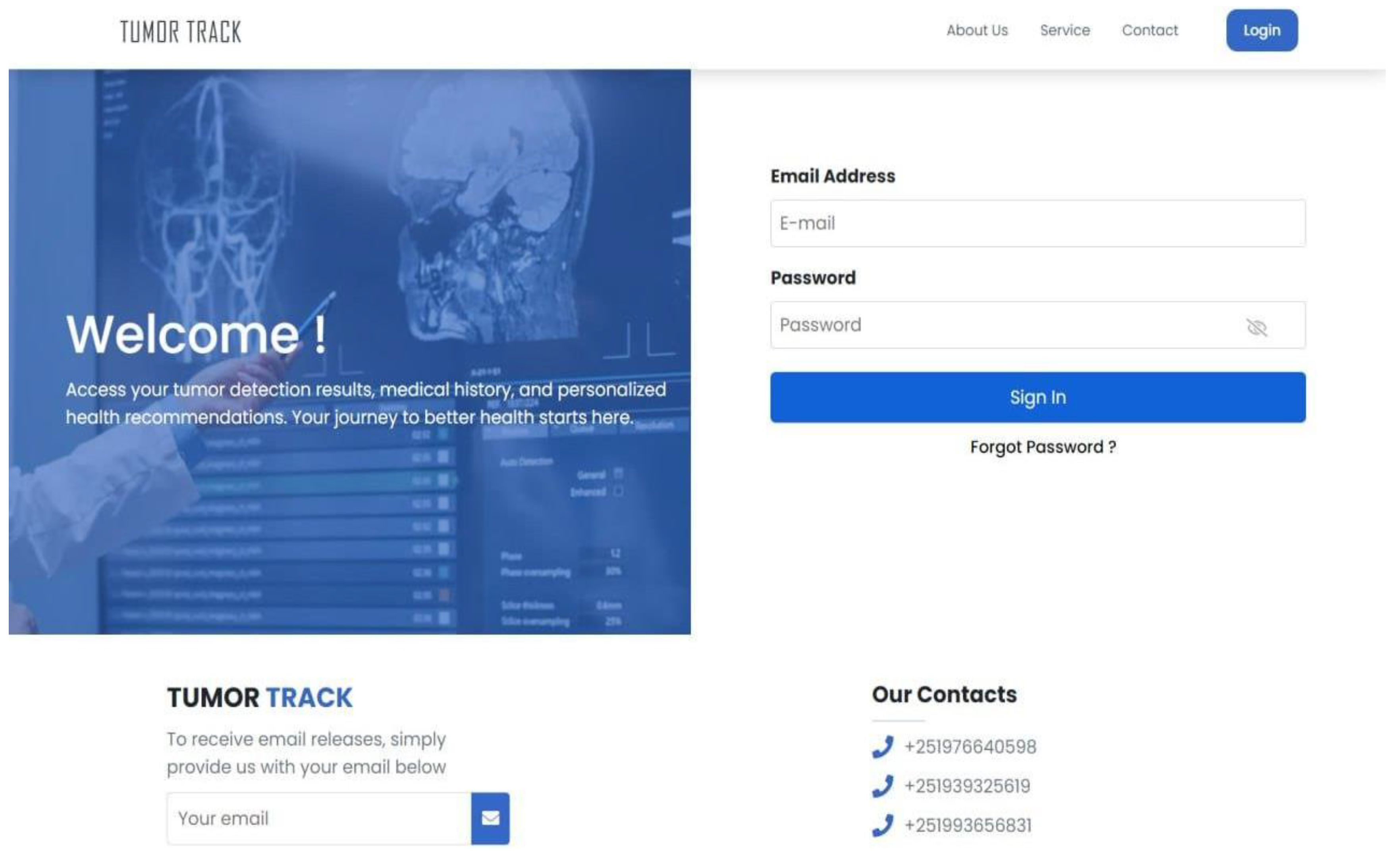
- User Authentication: The system must allow users to register, log in, and manage their accounts.
- Role-Based Access Control: Different roles (patient, medical receptionist, doctor, radiologist, specialist, and admin) must have appropriate access levels and functionalities.
- Patient History Documentation: Doctors must be able to document patient history and diagnosis.
- MRI Referral: Doctors must be able to refer patients to radiologists for MRI scans.
- MRI Result Upload: Radiologists must be able to upload MRI results and their diagnosis.
- Secure Communication: Communication between users must be compliant with relevant healthcare regulations and standards.
- Specialist Consultation: Doctors must be able to consult specialists or refer patients to them if needed.
- Notifications: The system must notify users about anything that might require their response or concern them.
- Security and Privacy: The system must ensure the confidentiality, integrity, and availability of patient data.
- Usability: The interface must be user-friendly and accessible.
- Performance: The system must be responsive and handle multiple user requests efficiently.
- Scalability: The system must be able to scale to accommodate a growing number of users and data.
3.4.2.1. UML Use Case Model UML Use Case Diagram
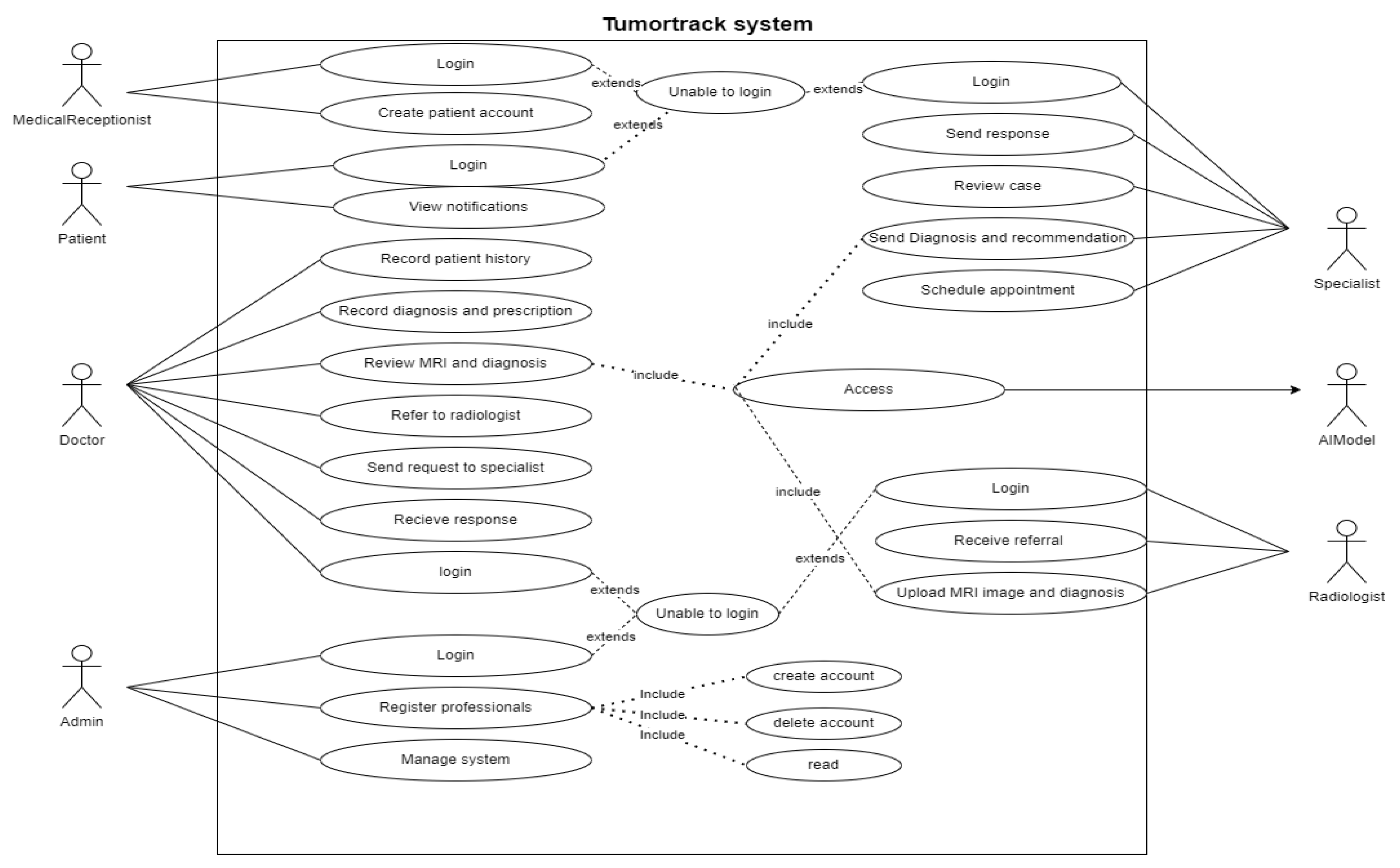
- UML Use Case Table
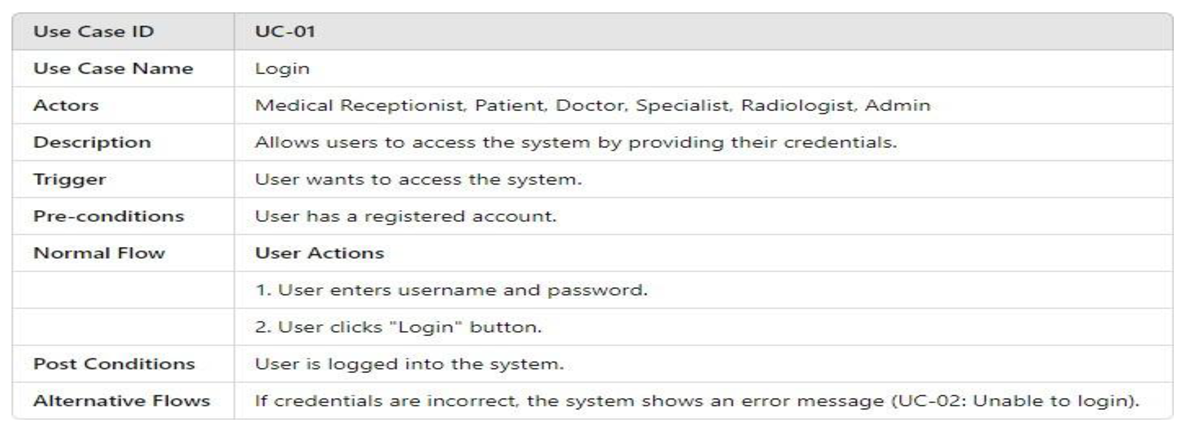
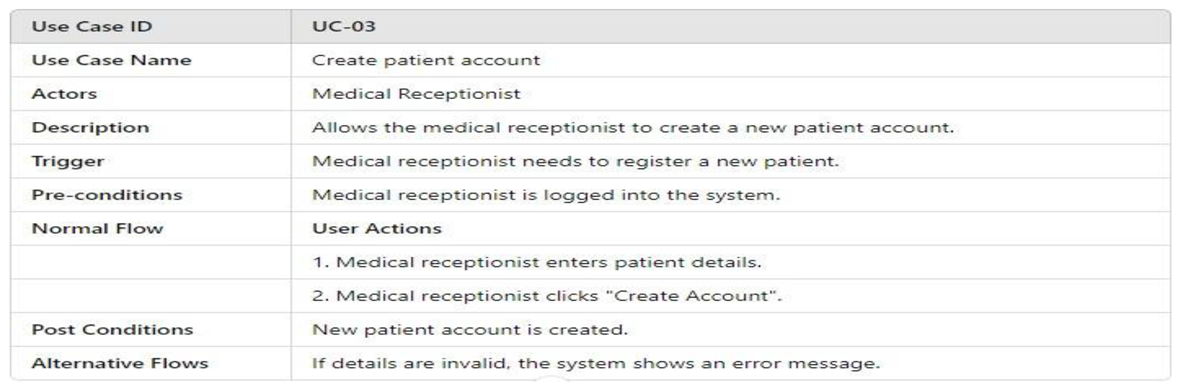
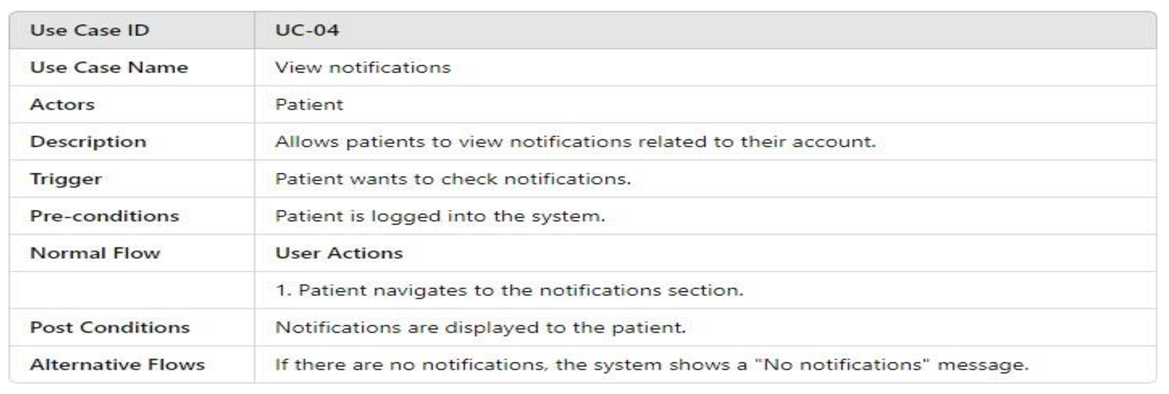
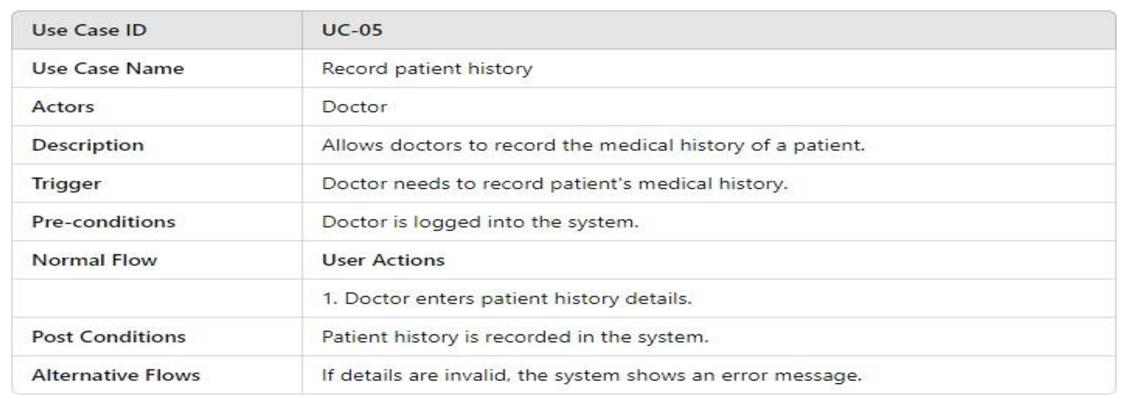

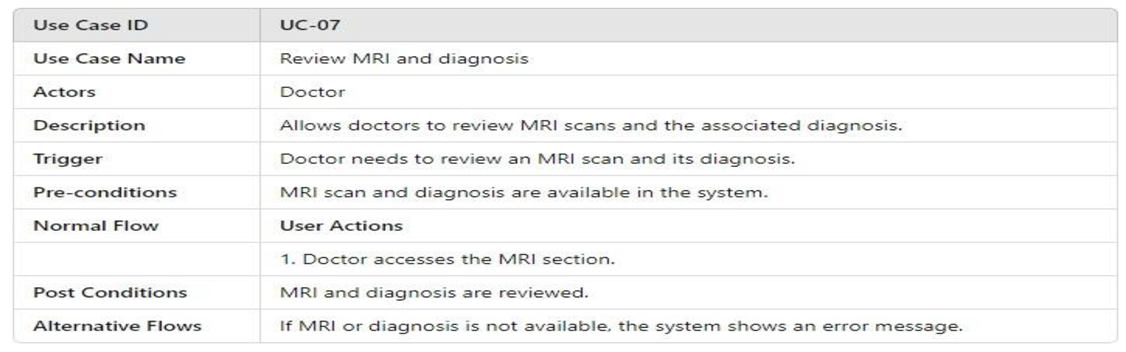
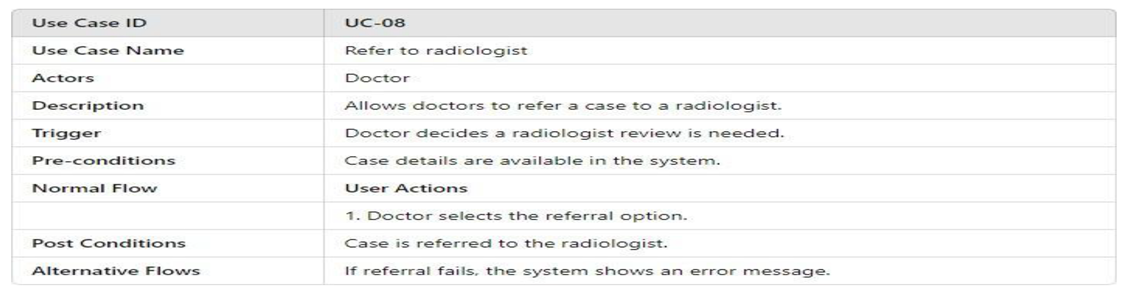
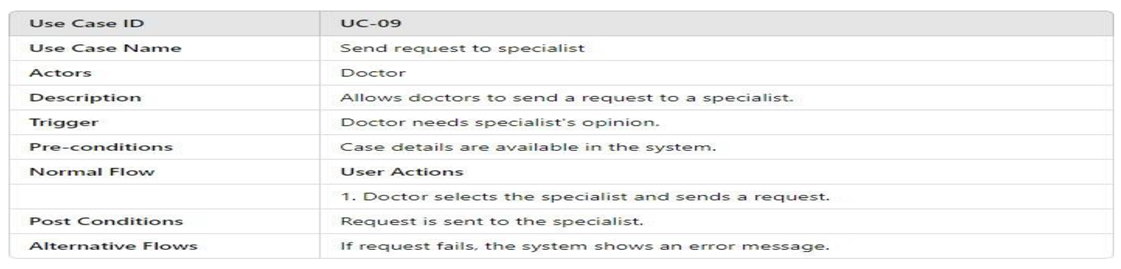
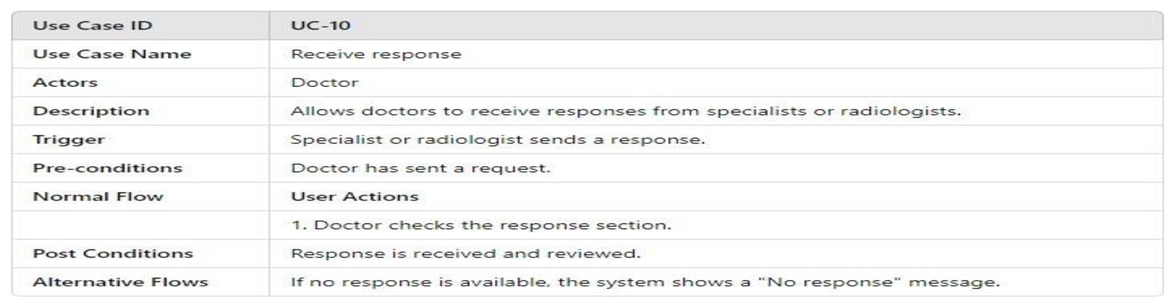
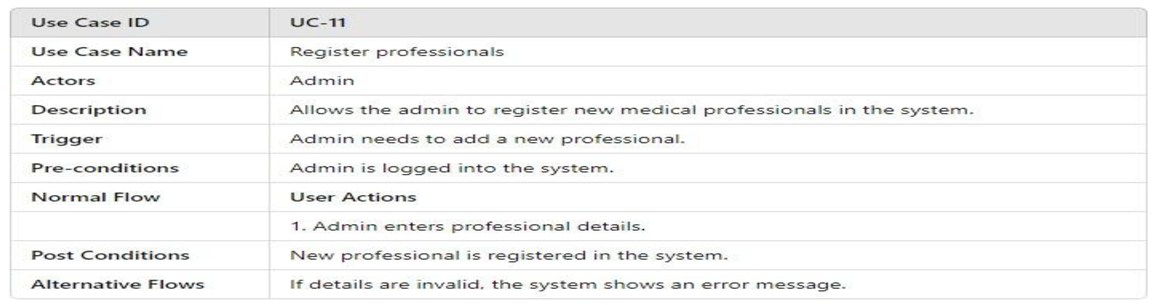
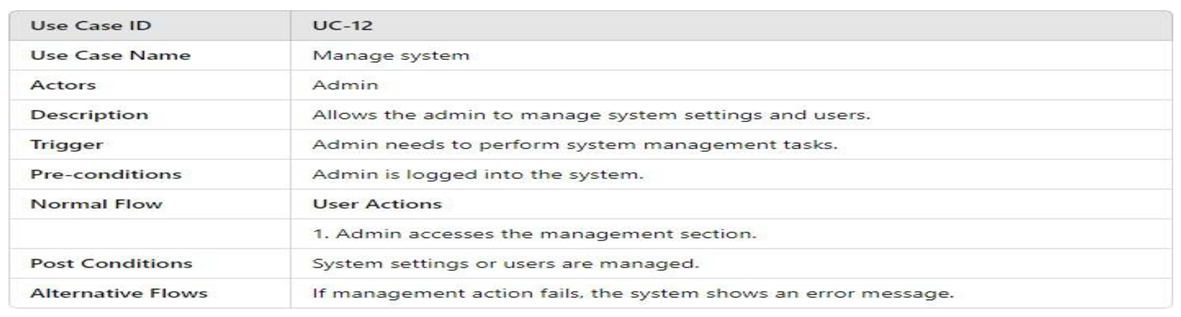
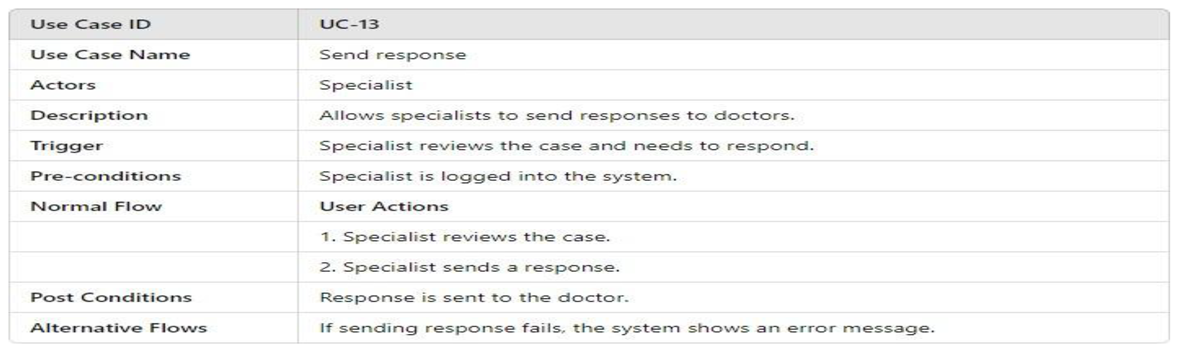
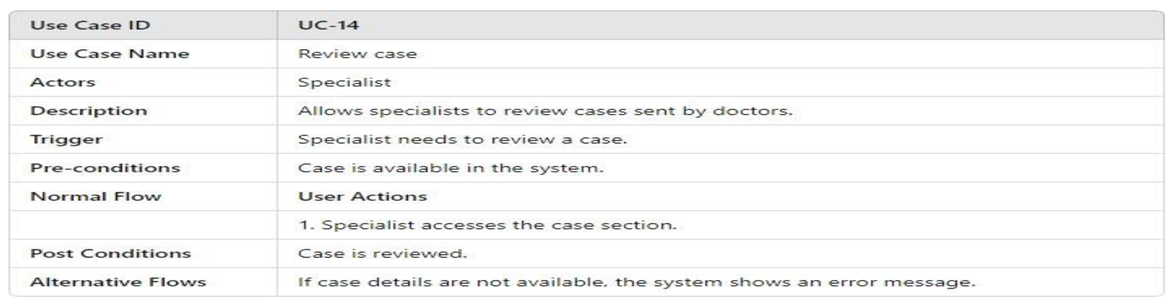

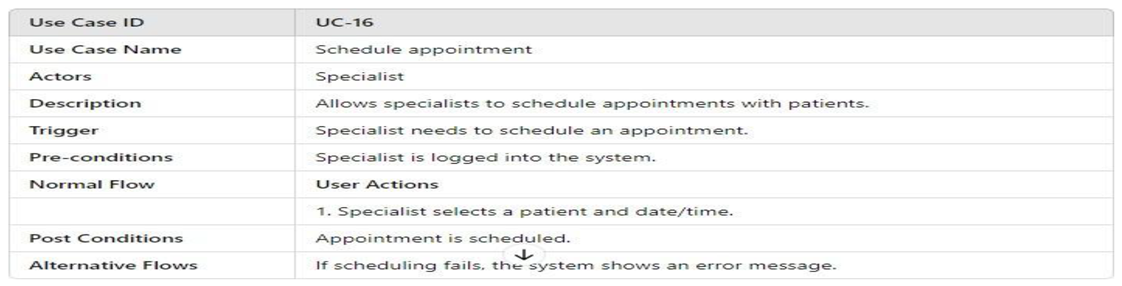
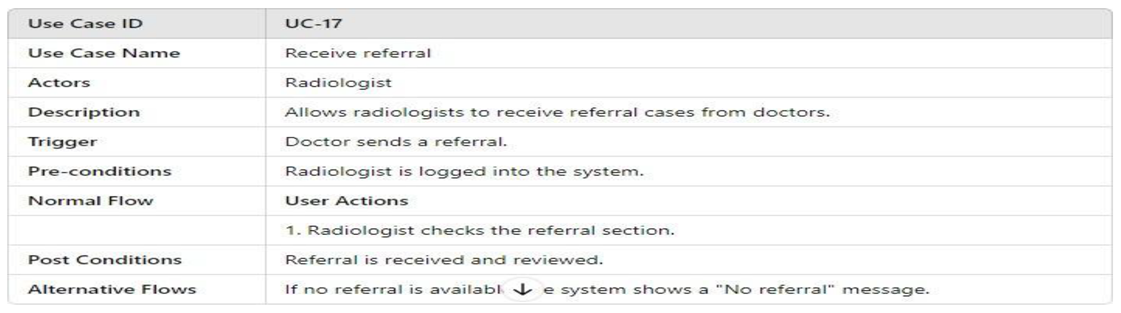
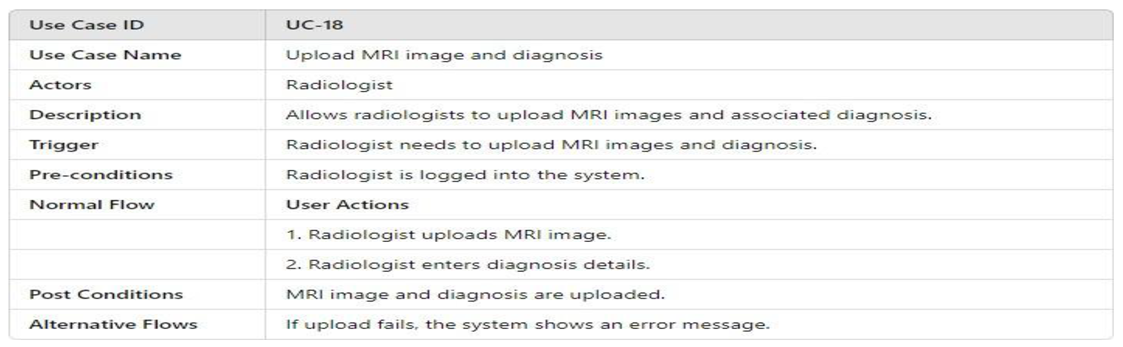
3.4.2.2. Activity Model
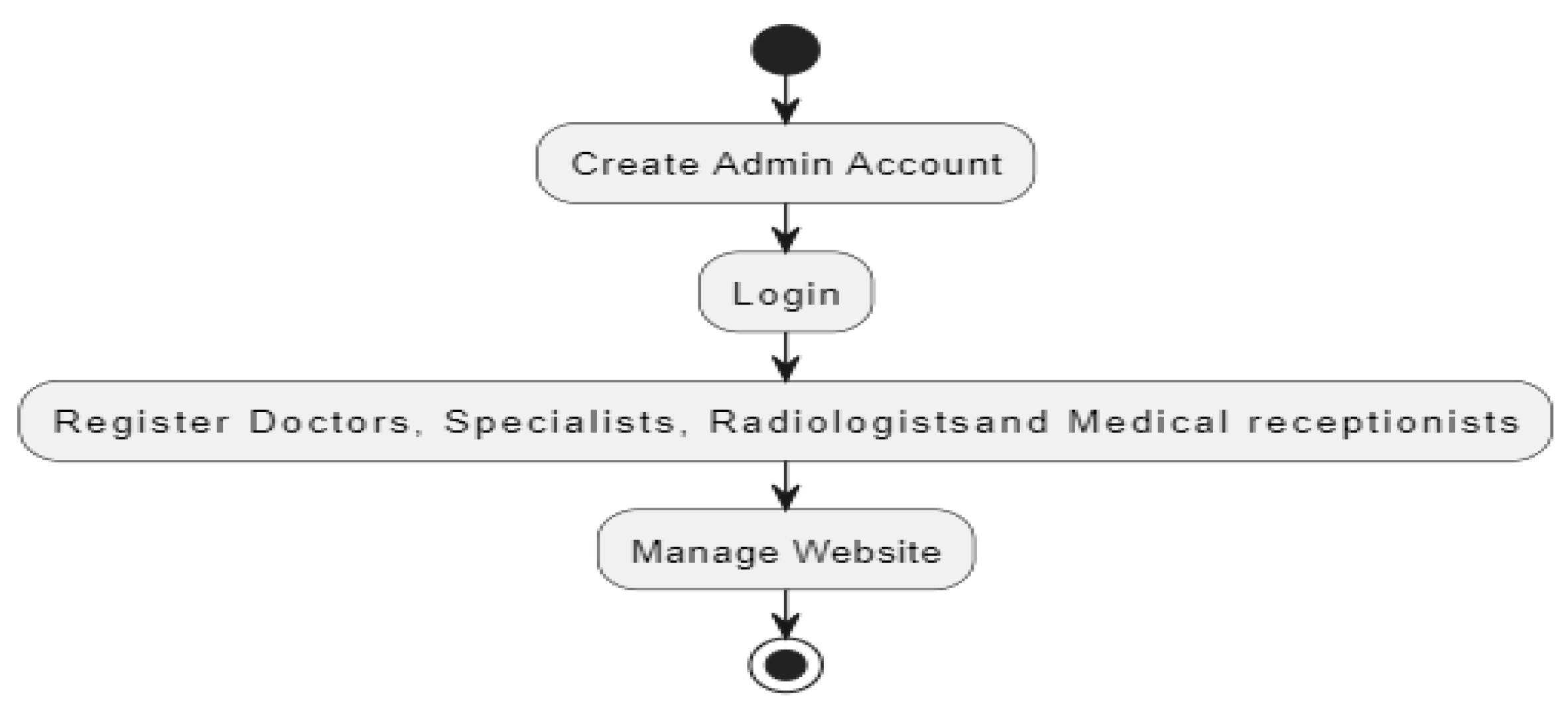
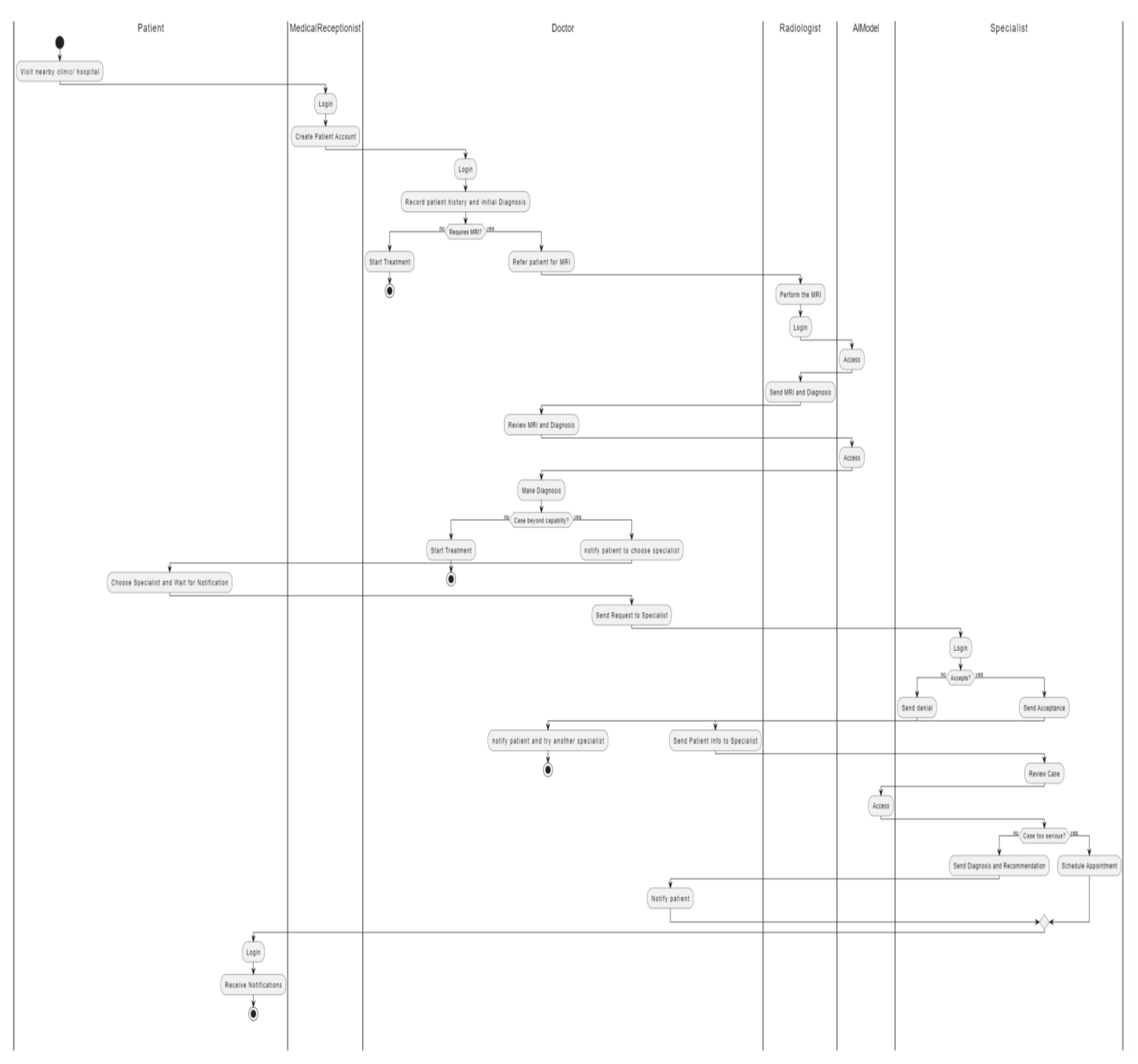
3.4.2.3. Sequence Diagram
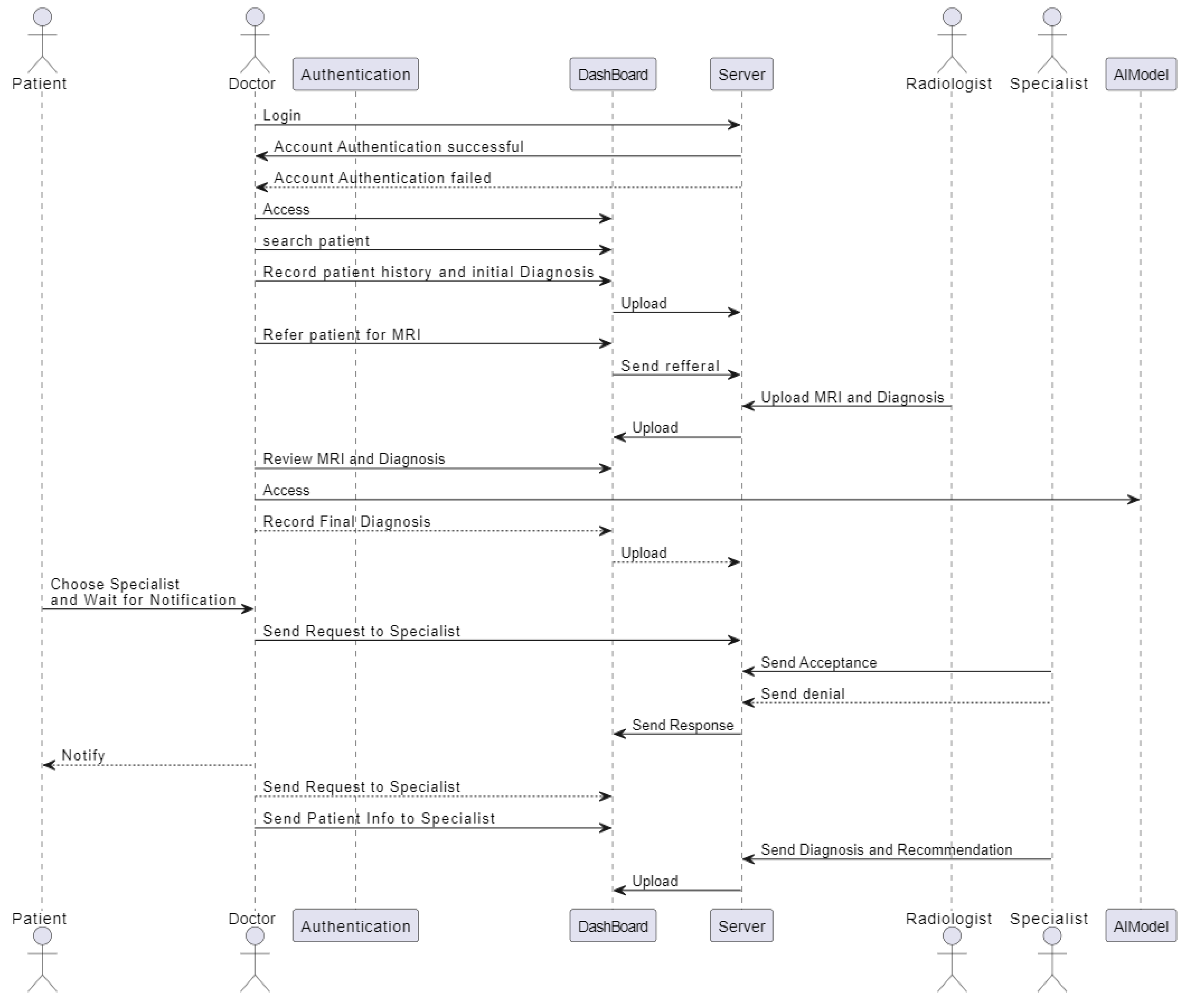
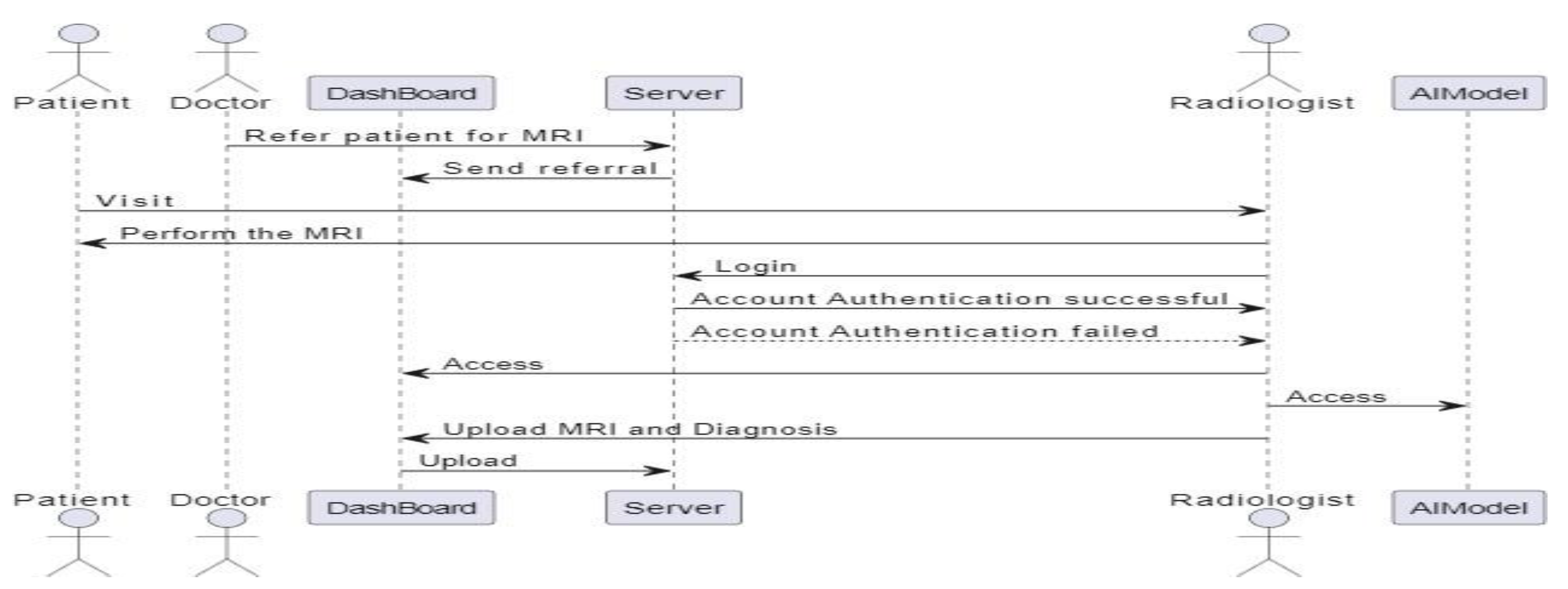
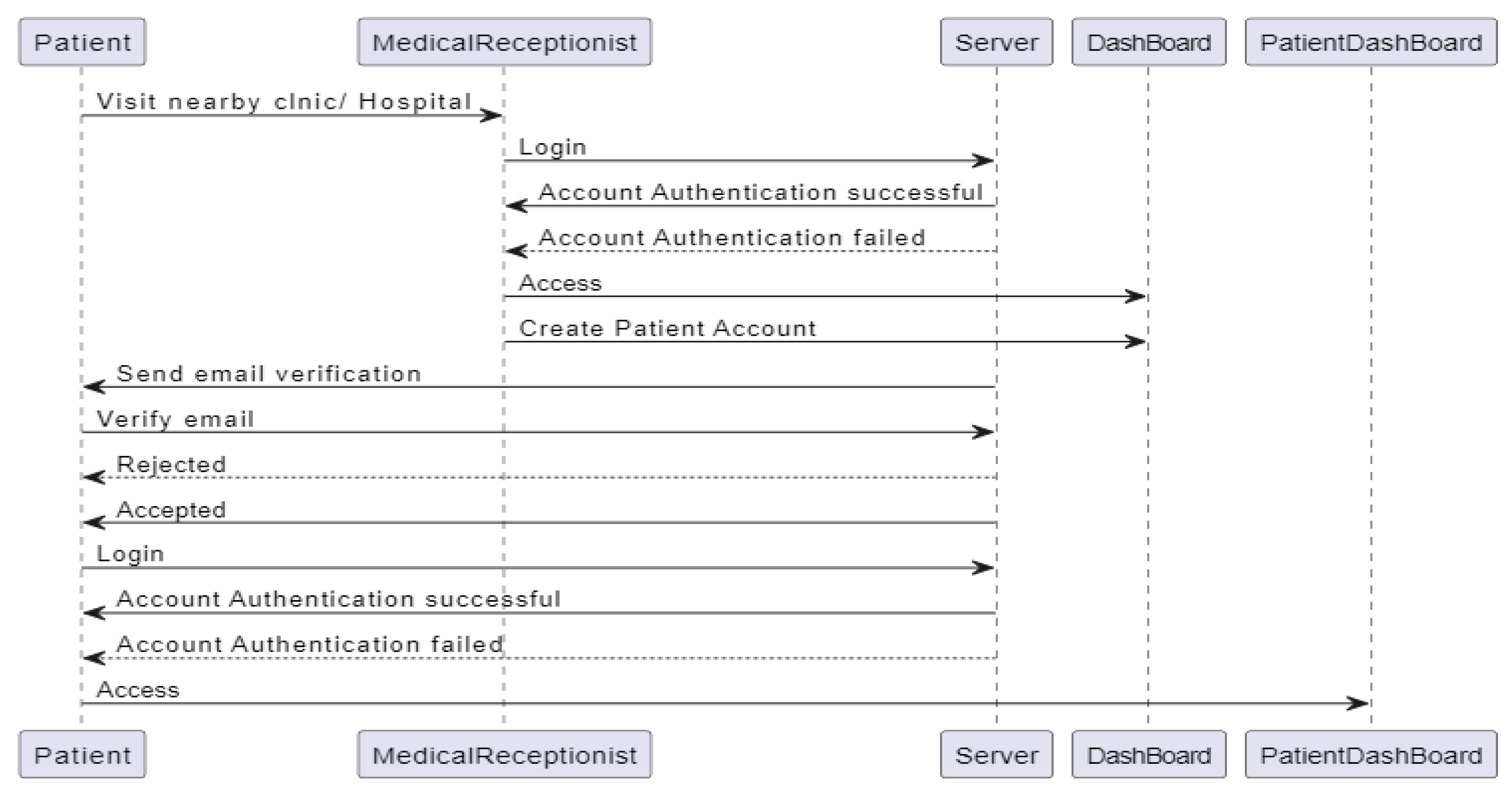
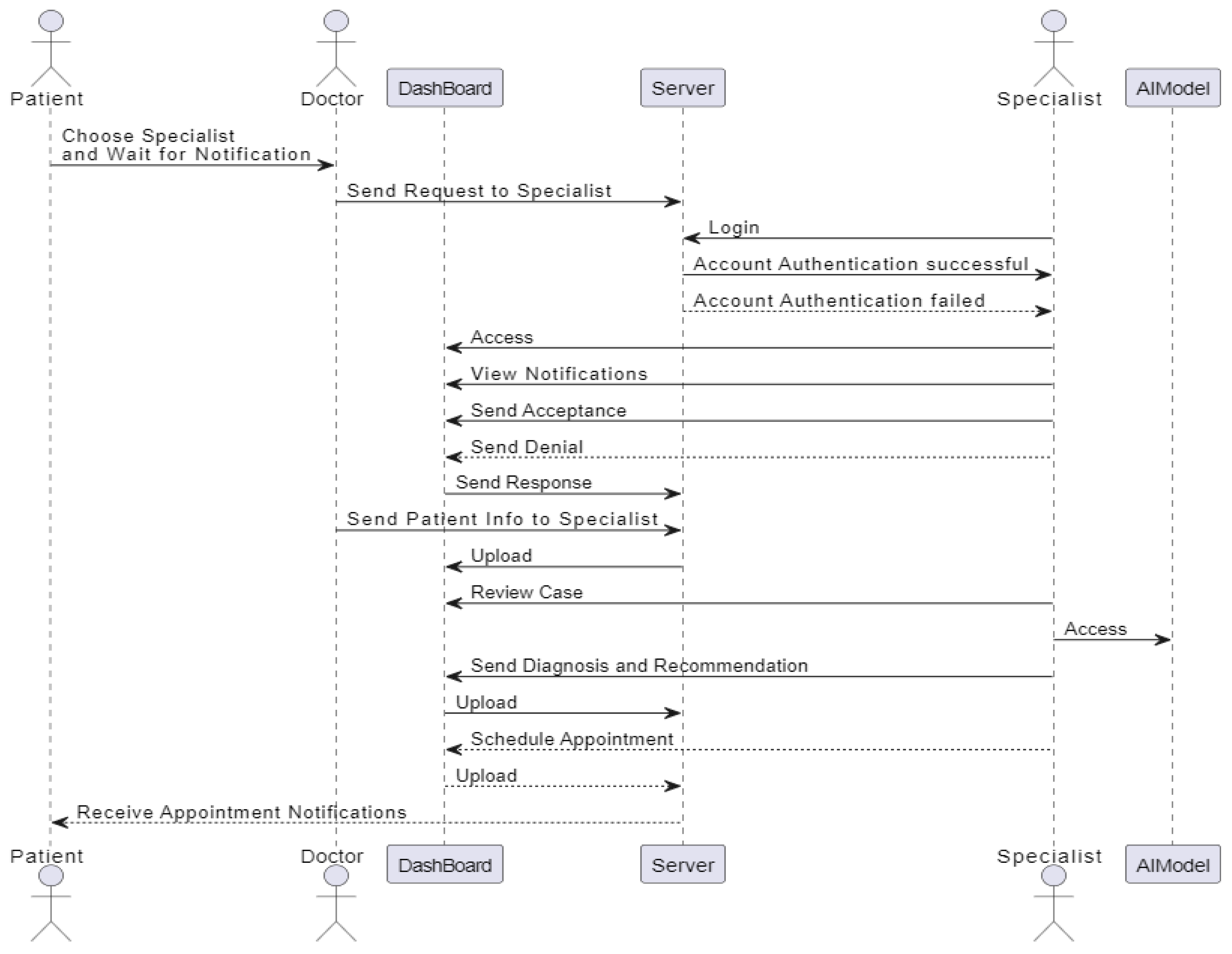
3.4.3. Software Design
3.4.3.1. System Architecture
- ➢
- Presentation Layer: This layer includes the user interface components for patients, doctors, radiologists, specialists, medical receptionists, and admins.
- ➢
- Business Logic Layer: This layer contains the core application logic, including patient management, diagnosis workflows, and communication between different actors.
- ➢
- Data Access Layer: This layer handles the interaction with the database, including CRUD operations for patient records, user accounts, and system configurations.
- ➢
- Database Layer: This layer consists of the database which stores all the persistent data required by the system.
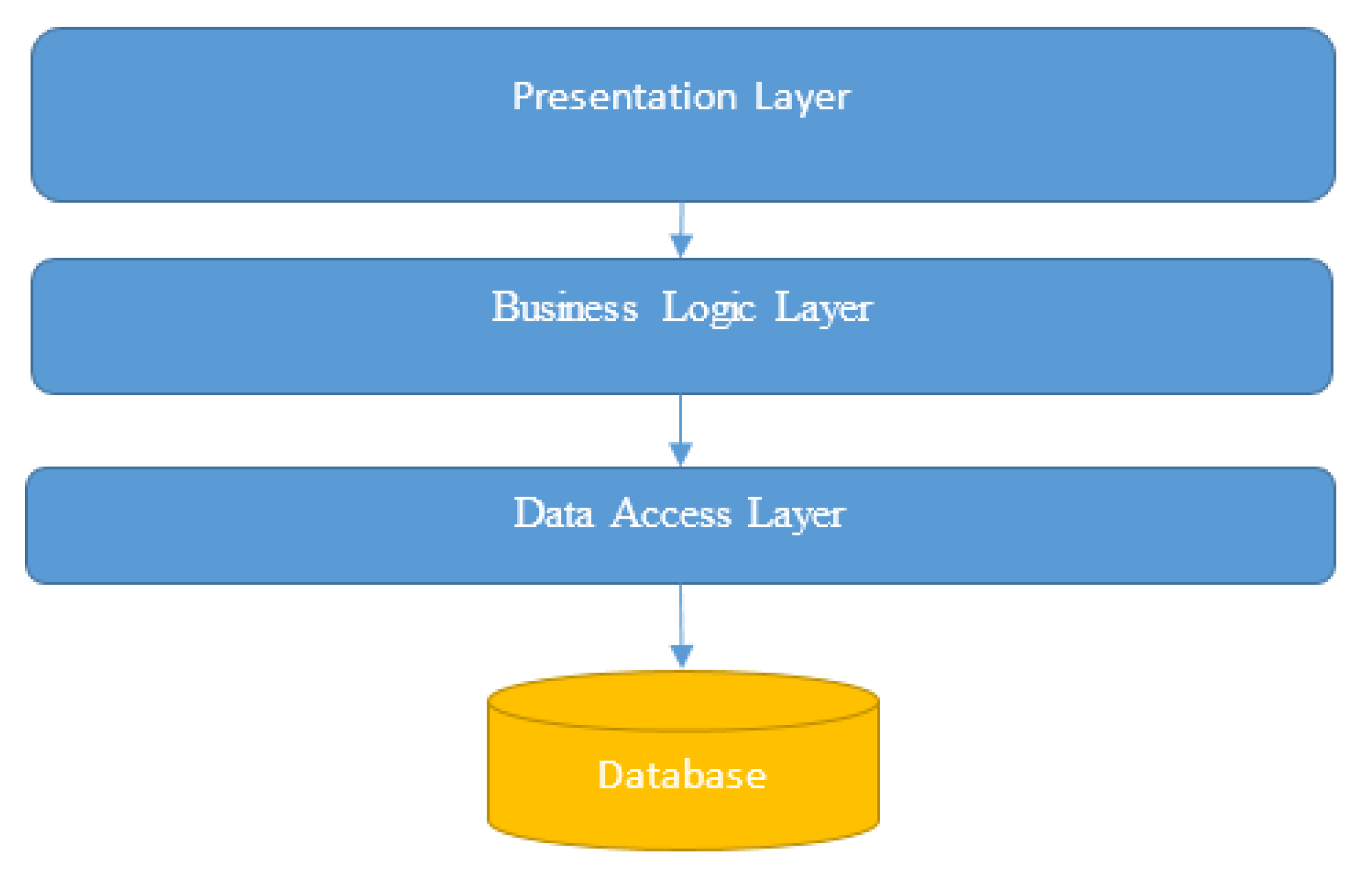
3.4.3.2. Database Design
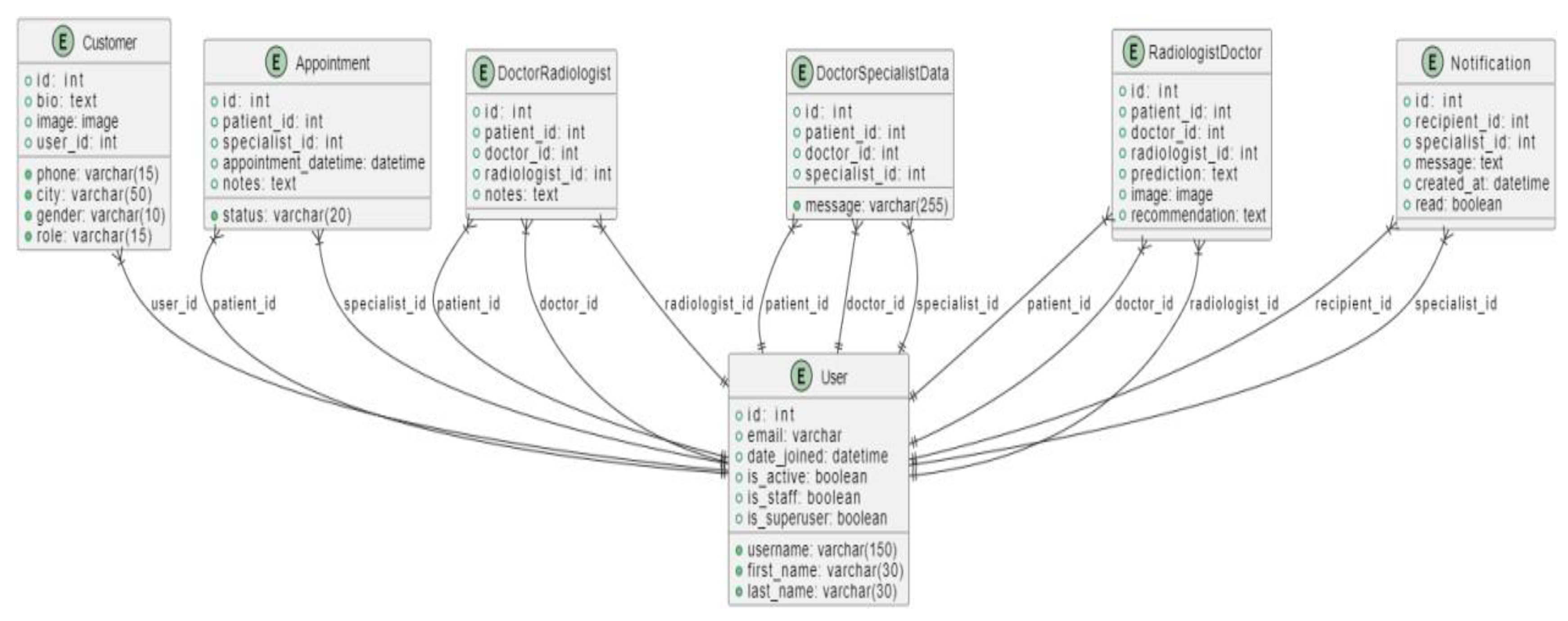
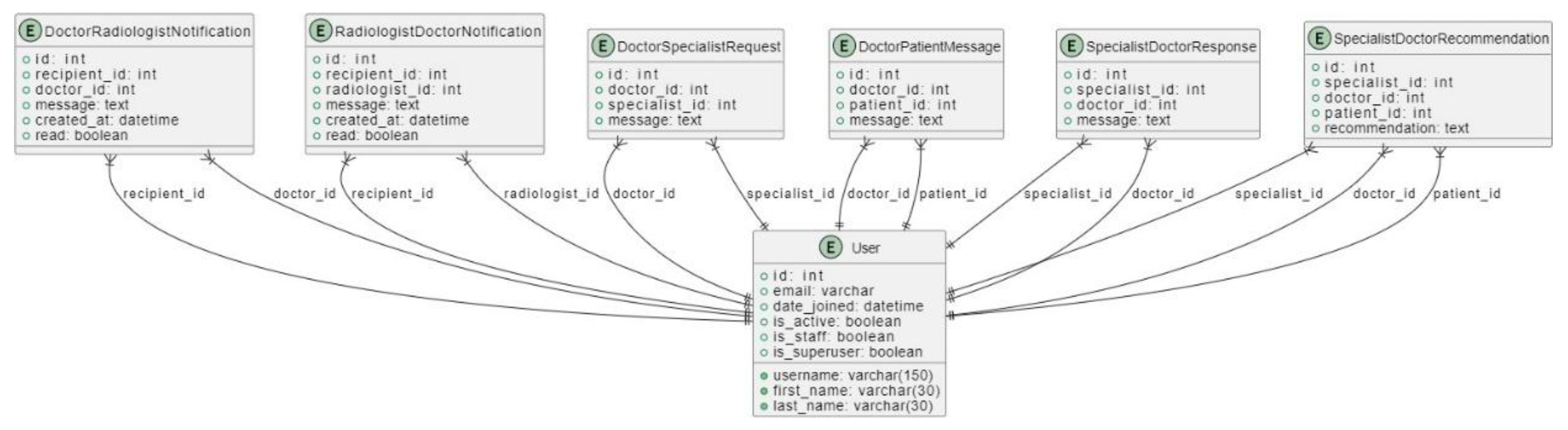
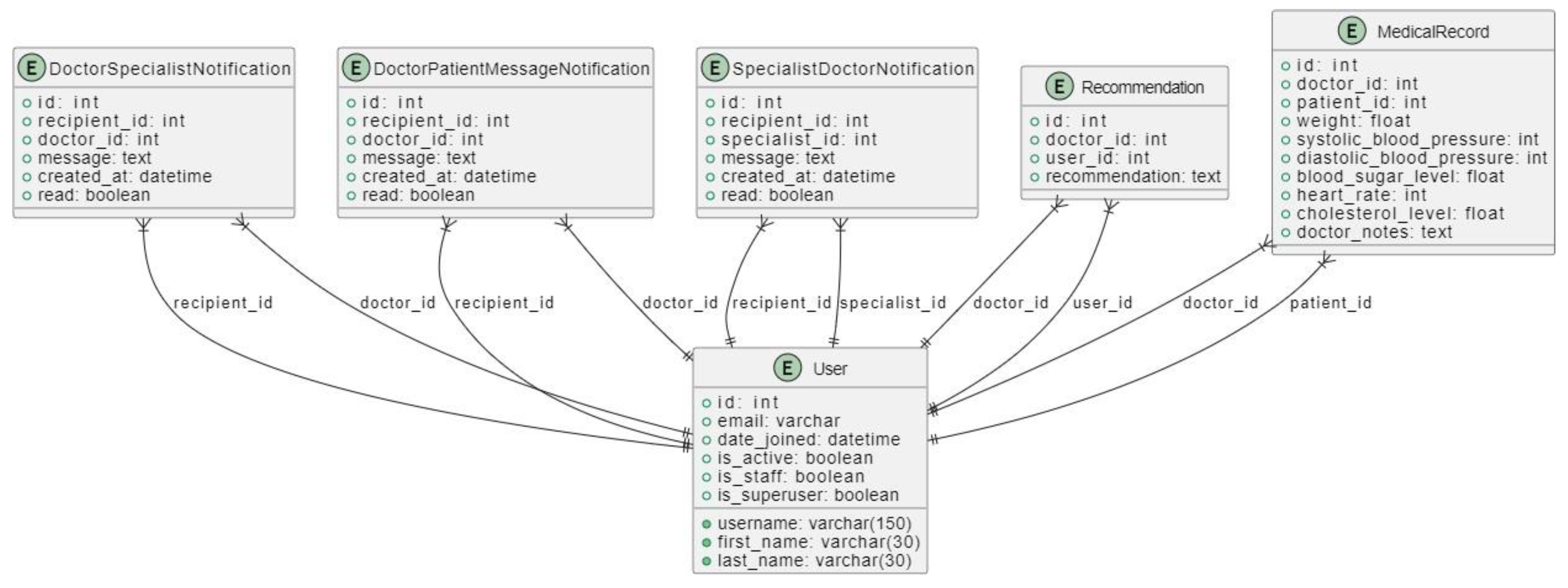
3.4.3.2. Class Diagram
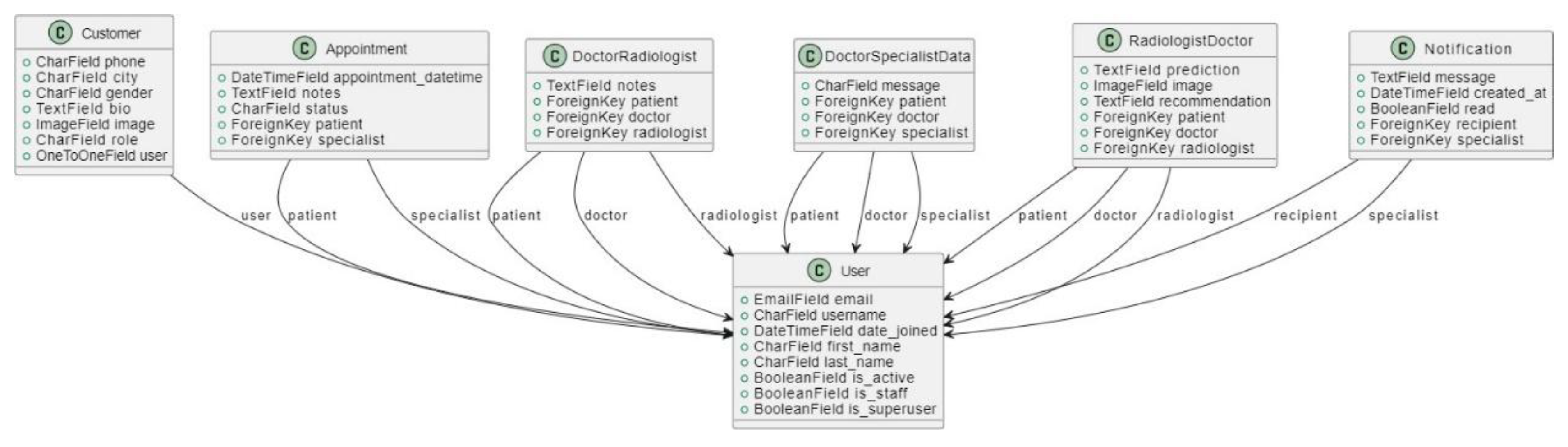
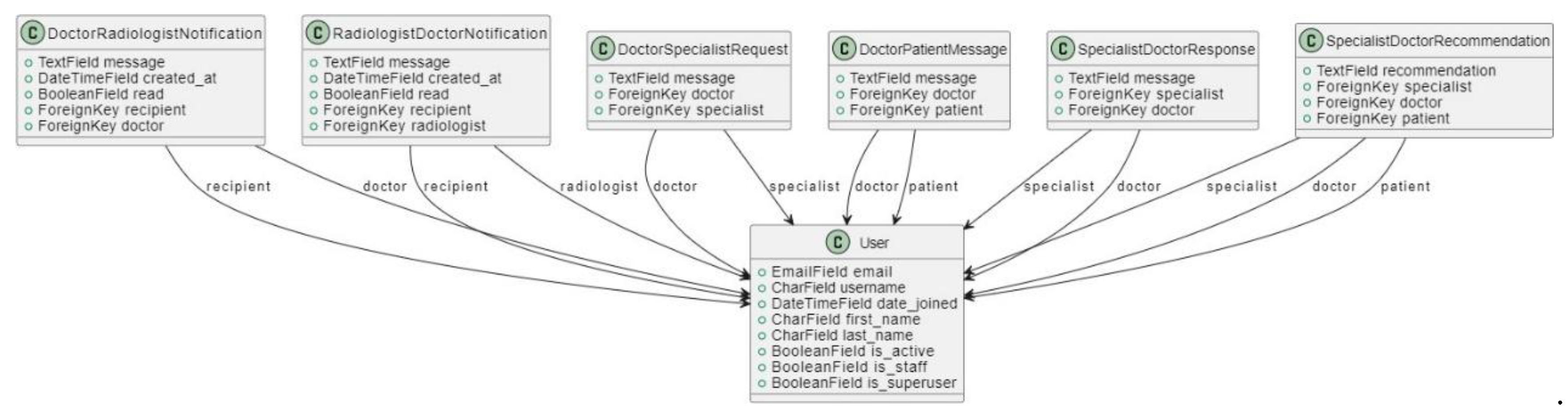
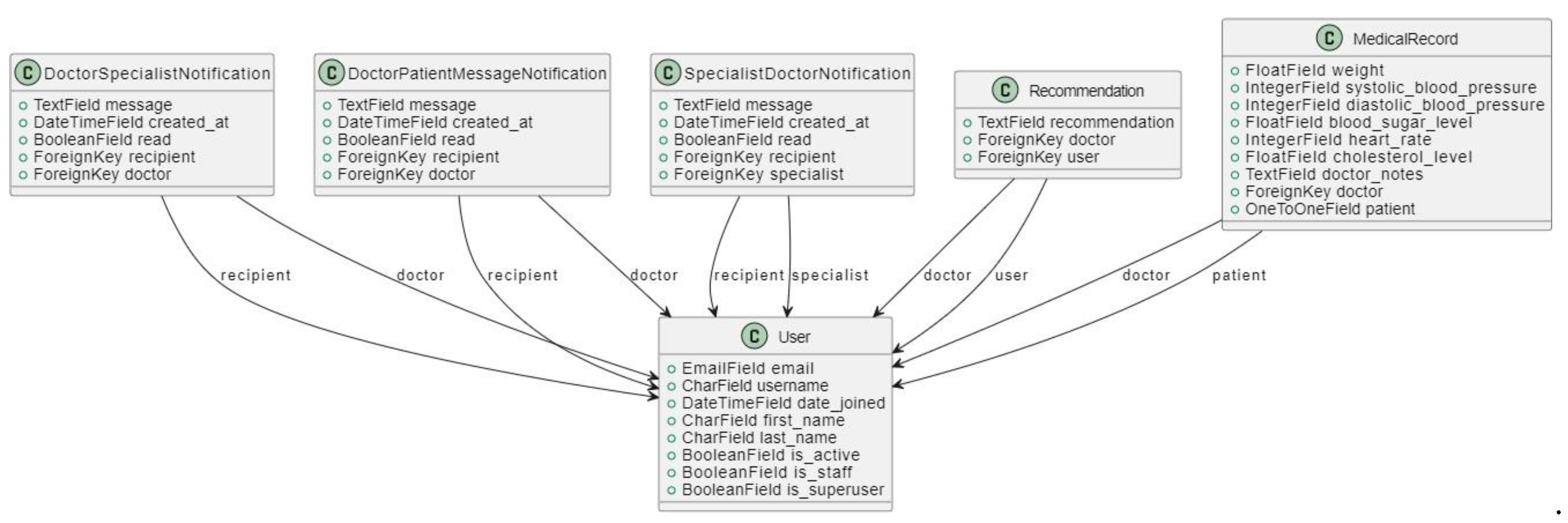
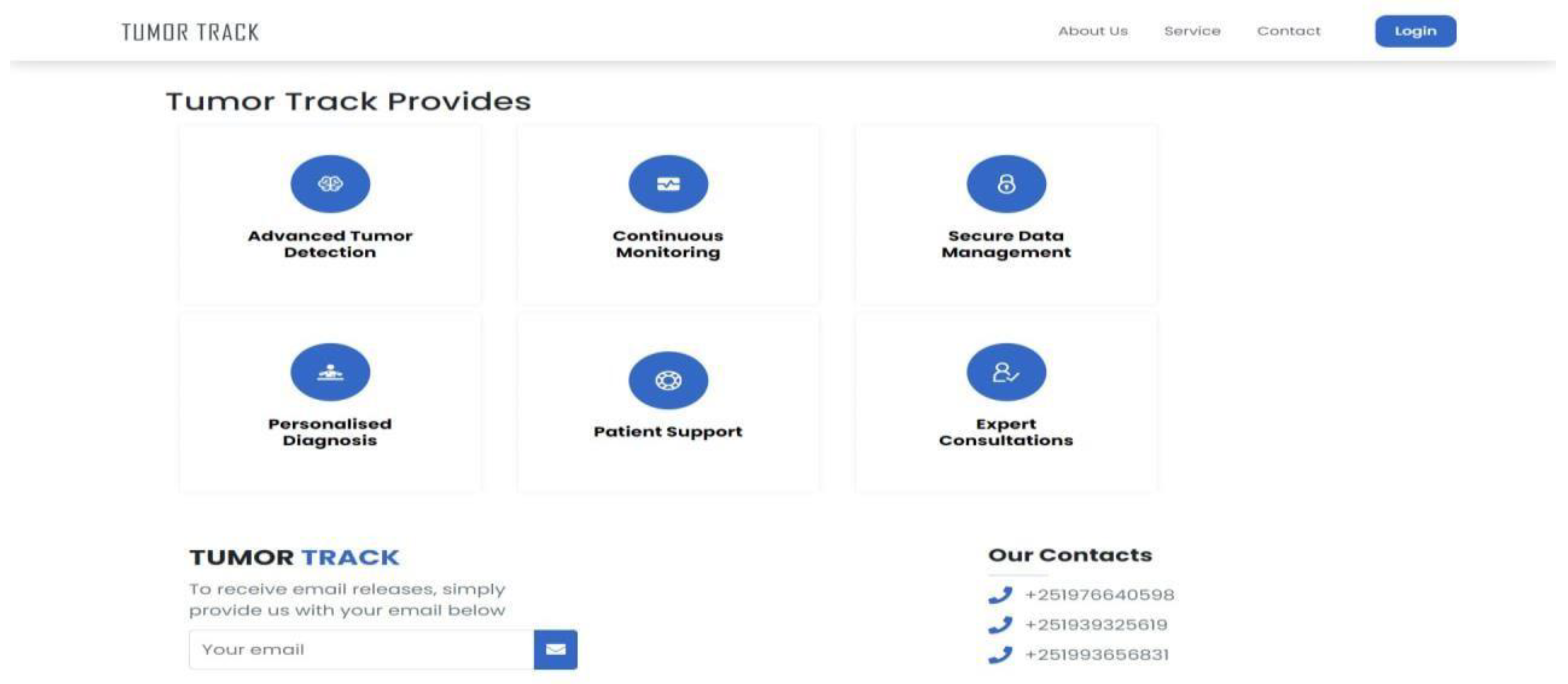
3.4.3.4. User Interface Design
- 1.
-
User-Centered Design:
- User Research: Conducted surveys and interviews with potential users to understand their needs, preferences, and pain points.
- Personas: Developed user personas to guide design decisions, ensuring the interface meets the specific needs of each user group.
- 2.
-
Simplicity and Consistency:
- Minimalistic Design: Emphasized a clean and simple layout to reduce cognitive load and make the interface intuitive.
- Consistency: Used consistent design elements, such as color schemes, fonts, and button styles, throughout the application to create a cohesive experience.
- 3.
-
Accessibility:
- Responsive Design: Ensured the interface is responsive and works well on various devices, including desktops, tablets, and smartphones.
- Accessibility Standards: Followed WCAG (Web Content Accessibility Guidelines) to make the interface accessible to users with disabilities.
- 4.
-
Visual Hierarchy:
- Prioritization of Information: Organized information and functionalities based on their importance and frequency of use.
- Highlighting Key Actions: Used visual cues, such as contrasting colors and icons, to draw attention to critical actions and information.
- 1.
-
Dashboard:
- Overview: Provides a summary of important information and quick access to common functionalities.
- User-Specific Data: Displays relevant data based on the user's role (e.g., upcoming appointments for specialists, patient records for doctors).
- 2.
-
Navigation:
- Top Navigation Bar: Includes links to different sections of the application, such as patient records, appointment scheduling, diagnostic tools, user profile settings, notifications, and
- 3.
-
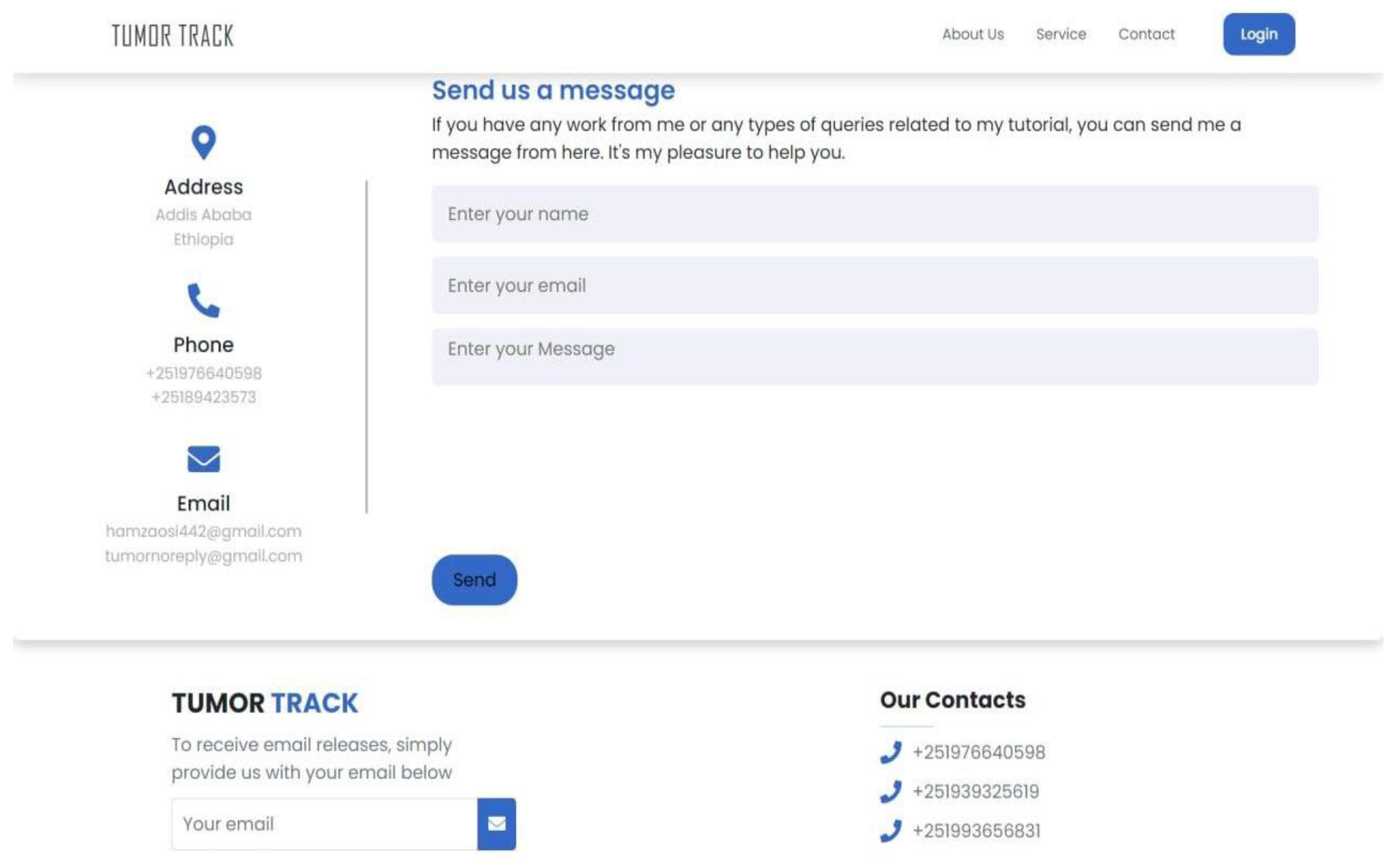
Forms and Data Entry:
- Intuitive Forms: Designed forms for tasks such as patient registration, diagnostic input, and appointment scheduling to be straightforward and easy to fill out.
- Validation and Feedback: Implemented real-time validation and feedback to guide users in providing correct and complete information.
- 4.
-
Patient Records:
- Detailed View: Allows doctors and medical staff to view comprehensive patient records, including medical history, diagnostic reports, and treatment plans.
- Edit and Update: Provides functionality for authorized users to edit and update patient information as needed.
- 5.
-
Appointment Scheduling:
- Calendar View: Displays a calendar for scheduling and managing appointments, with the ability to view by day, week, or month.
- Automated Reminders: Sends automated reminders to patients and staff about upcoming appointments.
- 6.
-
Diagnostic Tools:
- Image Viewer: Integrates with AI tools for viewing and analyzing medical images, such as MRI scans.
- Notes: Allows doctors to add notes for reference and collaboration.
- 7.
-
Notifications and Alerts:
- Real-Time Alerts: Provides real-time notifications for critical updates, such as new patient records, appointment changes, and diagnostic results.
- Notification Center: Centralizes all alerts and messages, allowing users to manage and review them efficiently.
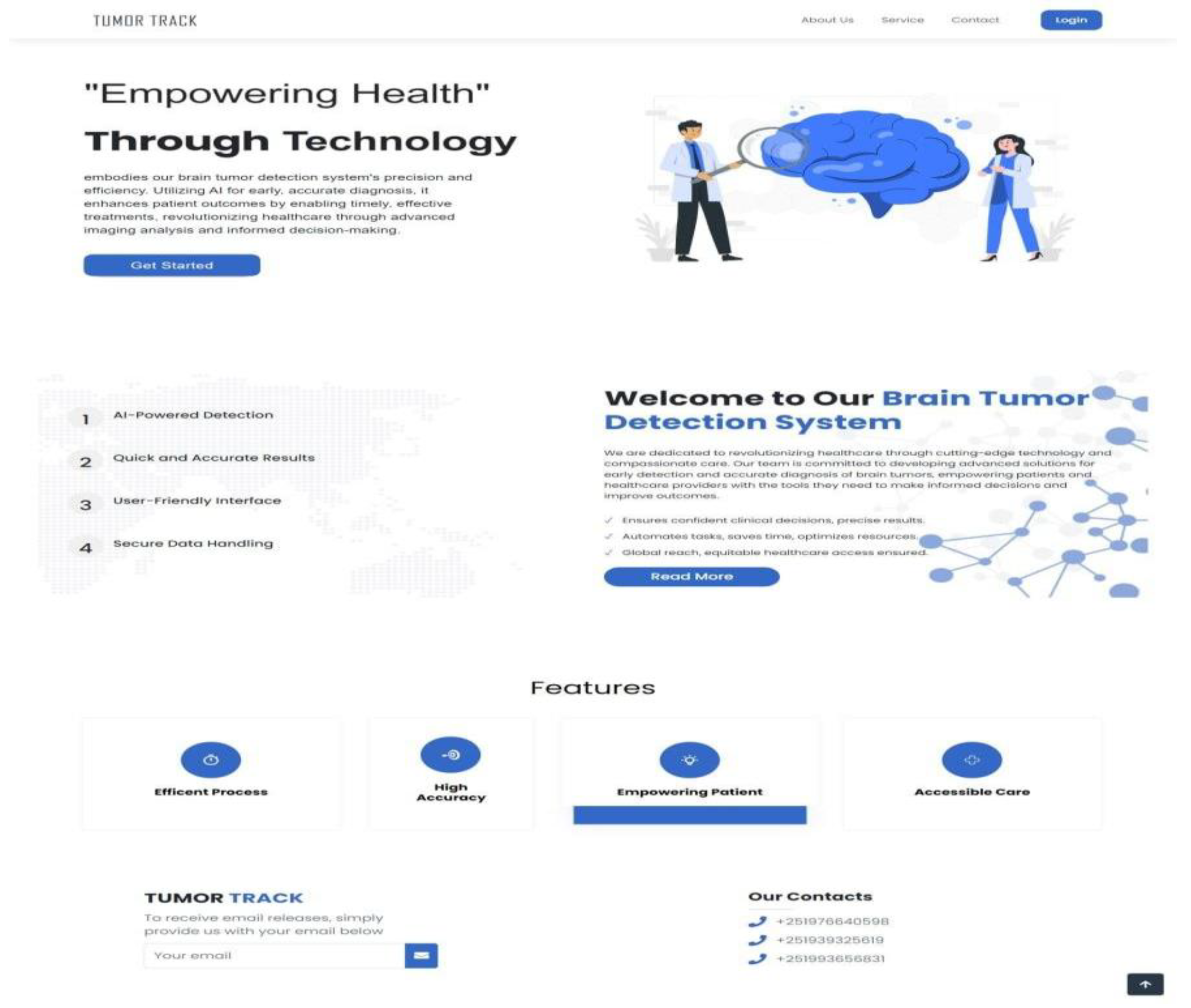
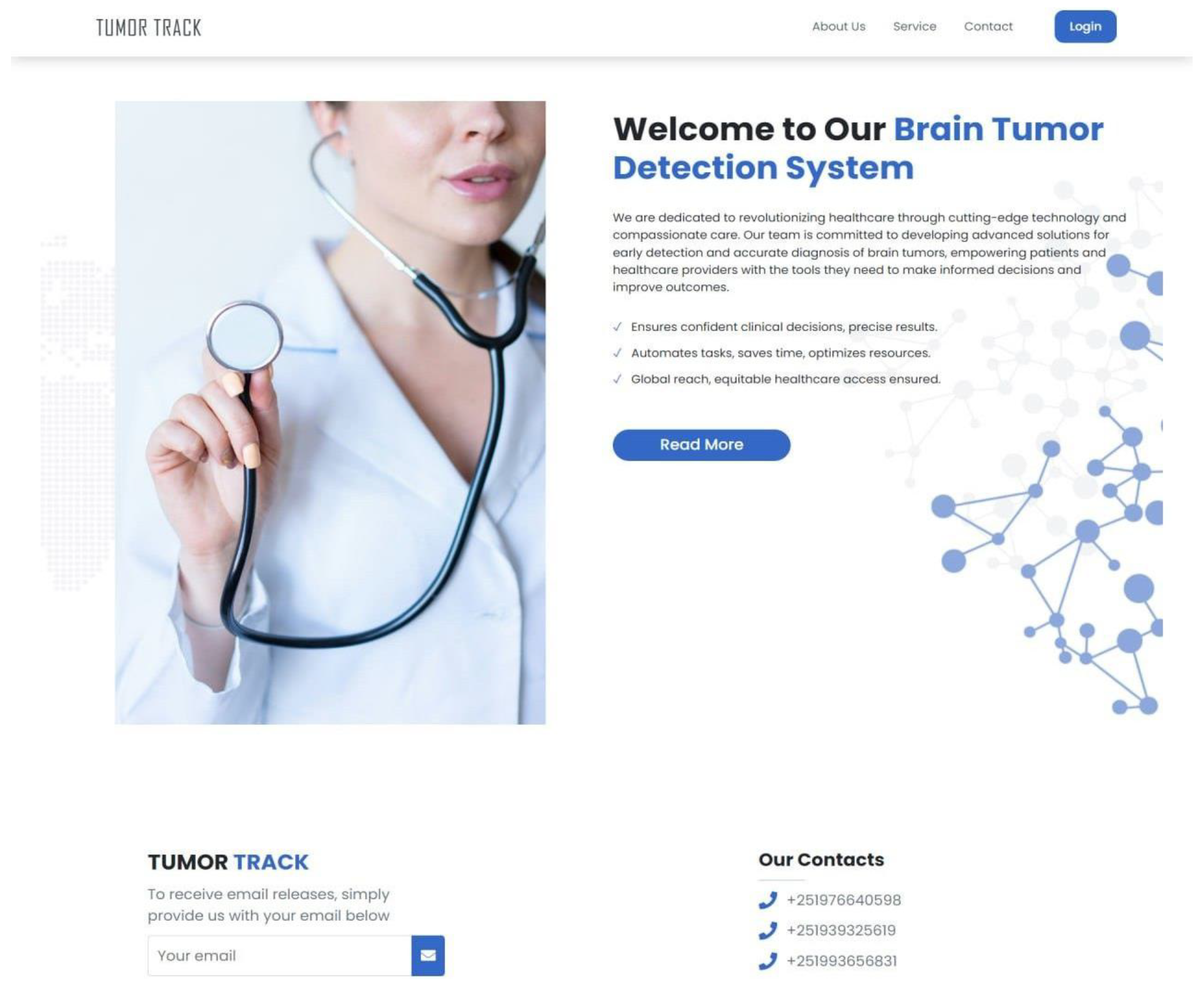
3.4.4. Implementation
3.4.4.1. Frontend Implementation
3.4.4.2. Backend Implementation:
3.5. Model Development
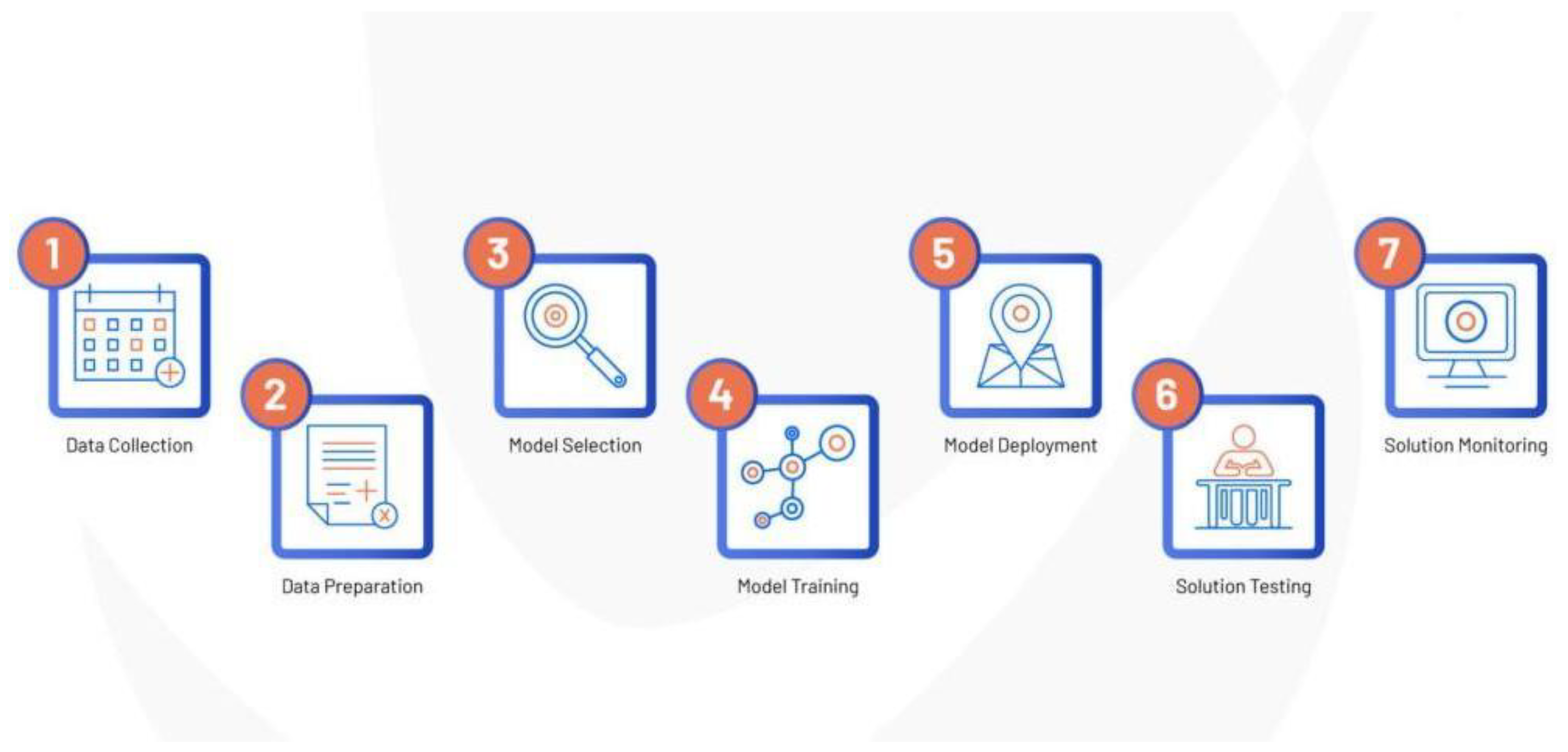
3.5.1. Brain Tumor Classification Model
3.5.1.1. Data Acquisition and Preparation
| Classes | Number of Images |
|---|---|
| Glioma | 1321 images |
| Meningioma | 1339 images |
| Pituitary | 1457 images |
| Healthy | 1595 images |
3.5.1.2. Model Selection
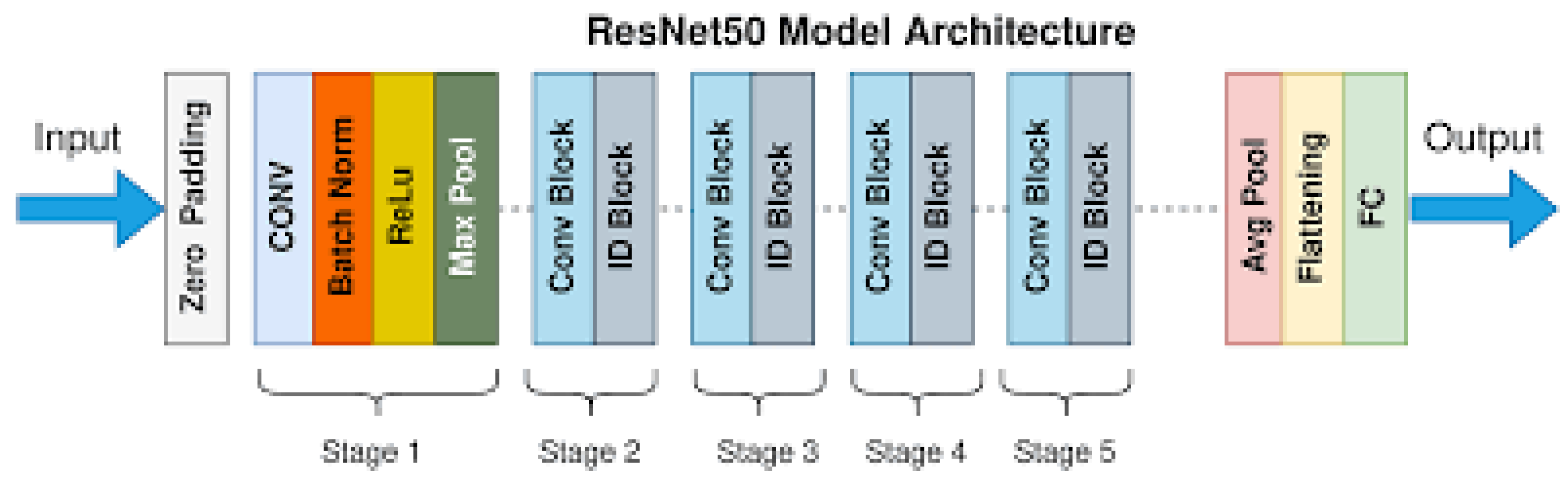
- Input Layer: Receives the input image.
- Convolutional Layers: Initial layers that perform feature extraction through convolution operations.
- Residual Blocks:
- Core building blocks of ResNet.
- Each block contains two convolutional layers with batch normalization and ReLU activation.
- Introduces skip connections to facilitate the learning of residual mappings.
- Pooling Layers: Subsequent layers that reduce the spatial dimensions of feature maps while preserving important information.
- Fully Connected Layers: Layers near the end of the network responsible for classification.
- Output Layer: Final layer that produces output predictions.
- Typically uses softmax activation for classification tasks.
3.5.1.3. Training Procedure
- The dataset was splitted into training, validation, and test sets (80% training, 20% testing).
- The model was trained for 50 epochs with early stopping based on validation loss to prevent overfitting.
- Batch normalization was used to stabilize and accelerate the training process.
3.5.2. Segmentation
3.5.2.1. Data Acquisition Data Set Selection:
| Parameter | Values |
|---|---|
| Labels | 3064 |
| Images | 3064 |
| Masks | 3064 |
| Augmented images | 4902 |
3.5.2.2. Data Pre Processing
3.5.2.3. Model Architecture
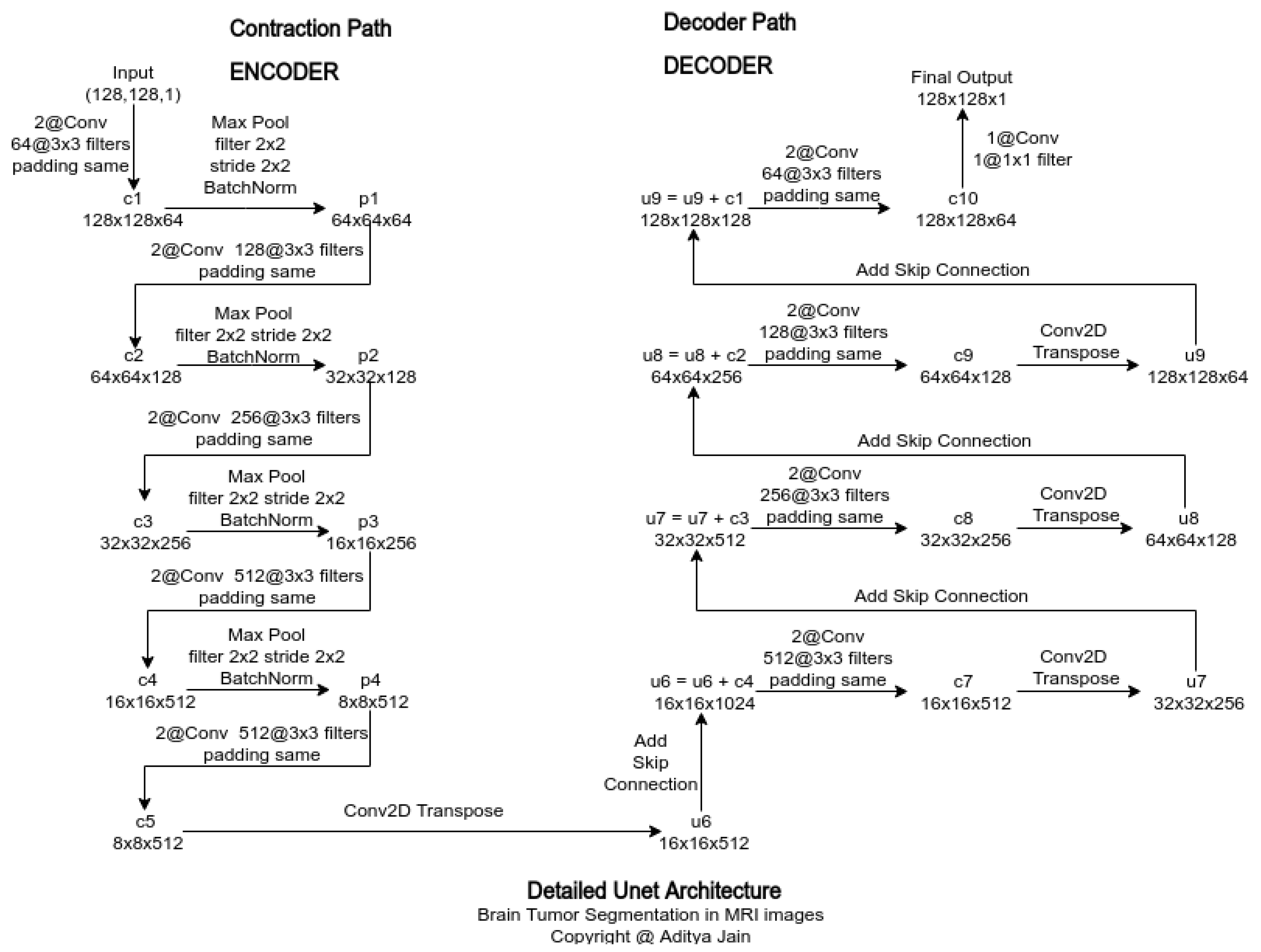
- Use 3x3 convolutional layers with padding to preserve spatial dimensions.
- Employ ReLU activation functions after each convolution.
- Use max-pooling layers with a pool size of 2x2 for down-sampling.
- Use transposed convolutions for up-sampling in the decoder part.
- Apply dropout layers where necessary to prevent over-fitting.
3.5.2.4. Training Procedure
- We split the data set into training, and test sets (80% training, 20% validation).
- We train the model for 100 epochs with early stopping based on validation loss to prevent over fitting.
- We use batch normalization to stabilize and accelerate the training process.
3.5.2.5. Data Post-Processing
- Improved Performance: Colab Pro provides faster GPUs and more memory, which improves the performance of resource-intensive tasks.
- Increased Flexibility: Colab Pro offers longer runtimes and background execution, which increases flexibility and control over resource allocation.
- Enhanced Debugging: Colab Pro's terminal access and continuous execution enhance debugging capabilities and ensure that users can resume their work seamlessly.
- Cost-Effective: Colab Pro is priced at $9.99 per month, making it a cost-effective option for users who require advanced features and resources.
- Community Support: Colab Pro has a large and active community of users and developers who contribute to its development and provide support through various resources.
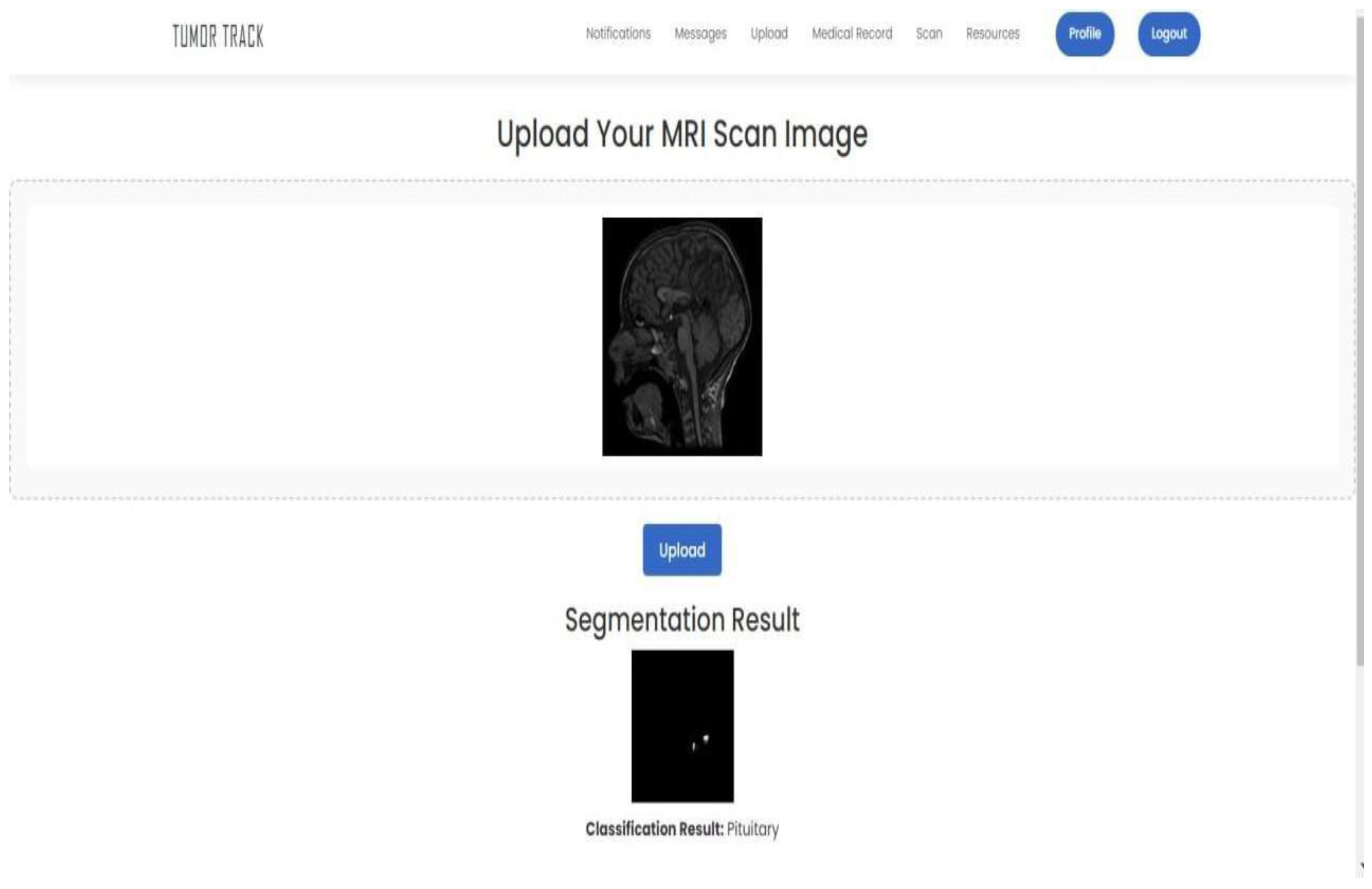
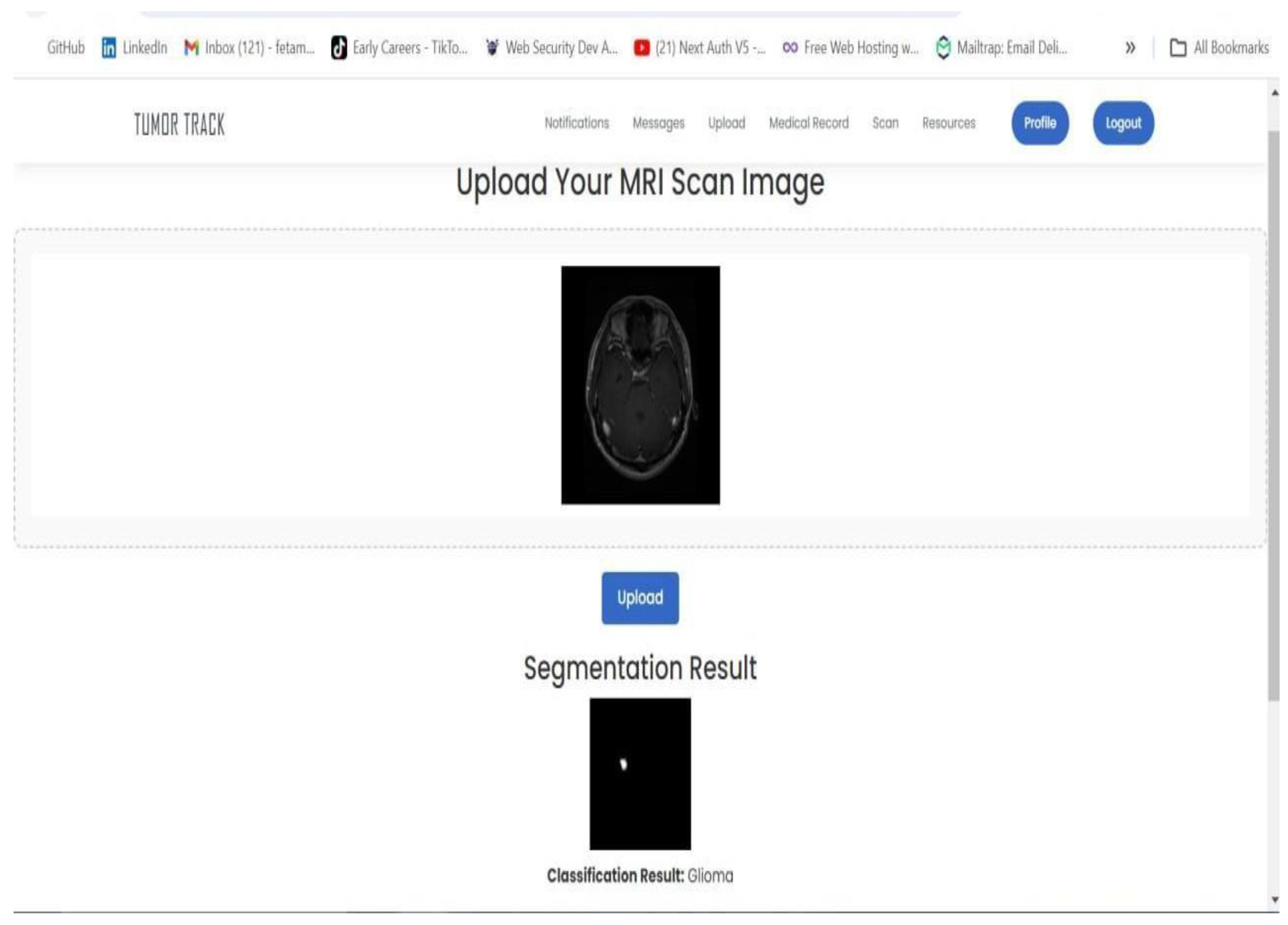
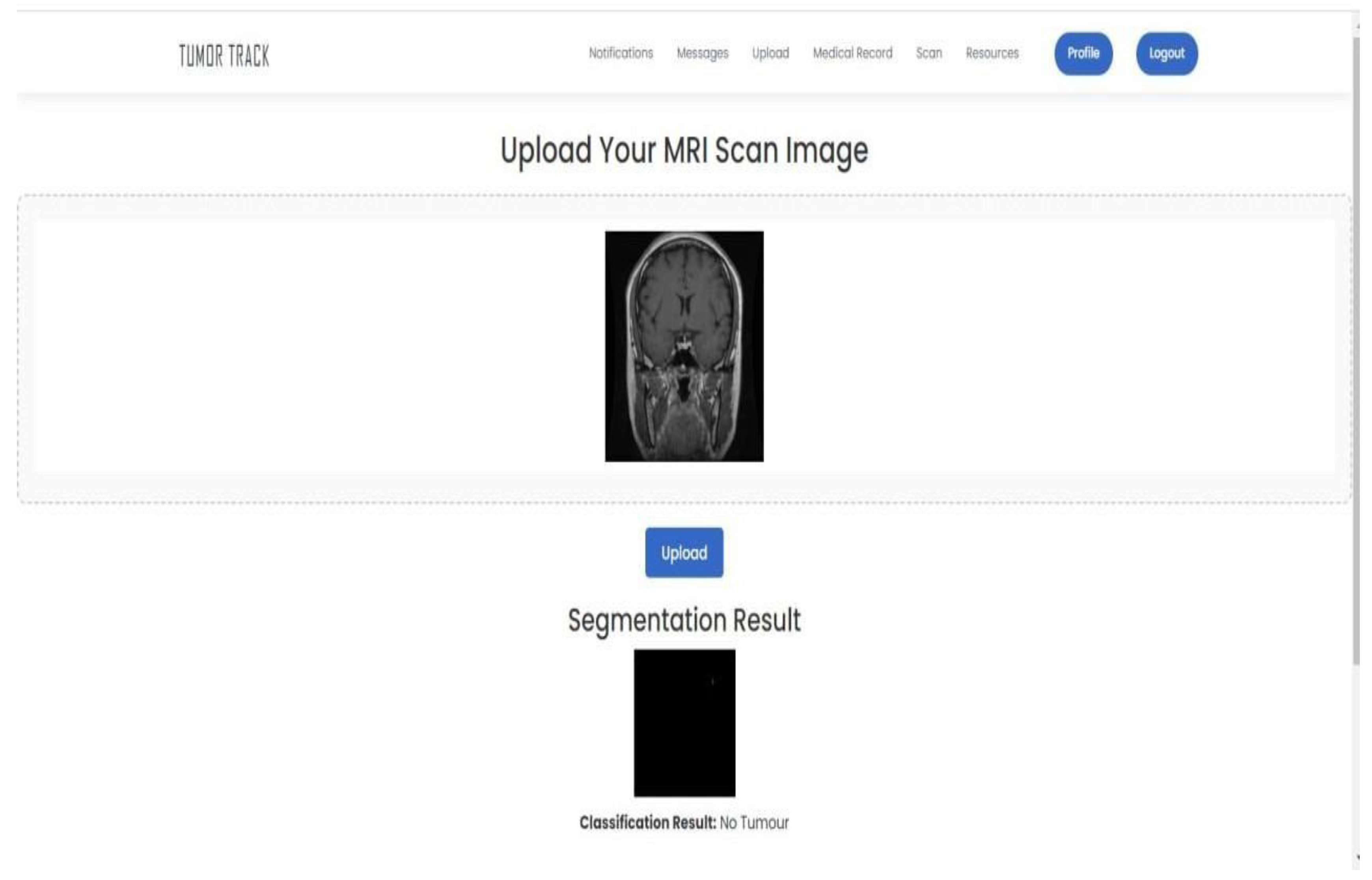
3.6. Integration
Chapter Four Result and Discussion
4.1. Introduction
4.2. Experimental Setup
4.2.1. Introduction
4.2.2. Materials and Equipment Hardware:
- Computers: For development, testing, and user access.
- 2.
- Development Tools: IDE (Visual Studio Code and Google Colab), version control (Git), web server (XAMPP).
- 3.
- Programming Languages: React for frontend and Django framework for backend development.
- 4.
- Database: SQL-based database (MySQL) for storing user data, patient records, MRI images, and other necessary information.
- 5.
- AI Model: Pre-trained deep learning models for brain tumor classification andsegmentation using ResNet 50 and U-Net respectively.
- 6.
- Testing Tools: Manual testing scripts.
4.2.3. Testing Strategy
- 1.
- Unit Testing: Test individual components of the system to ensure they function correctly.
- 2.
- Integration Testing: Verify that different modules of the system work together as expected.
- 3.
- System Testing: Conduct end-to-end testing to ensure the entire system operates correctly under various scenarios.
4.2.4. Test Procedure
- 1.
- Develop System Tests: Write test cases that cover all functional and non- functional requirements.
- 2.
- Execute Tests: Conduct the tests to validate overall system behavior.
- 3.
- Analyze Results: Review test results to ensure the system meets all specified requirements.
4.2.5. Safety Considerations
4.3. Result
4.3.1. Software Testing and Evaluation
| Scenario Id | 01 |
| Scenario Name | Patient with severe Tumor case |
| Participant Actor | Patient (Girma Worku), Medical receptionist (Sara Bilew), Doctor (Dr. Yonas Geda), Radiologist (Leole Masresha), Specialist (Dr. Hana Asfaw) |
| Flow of events |
|
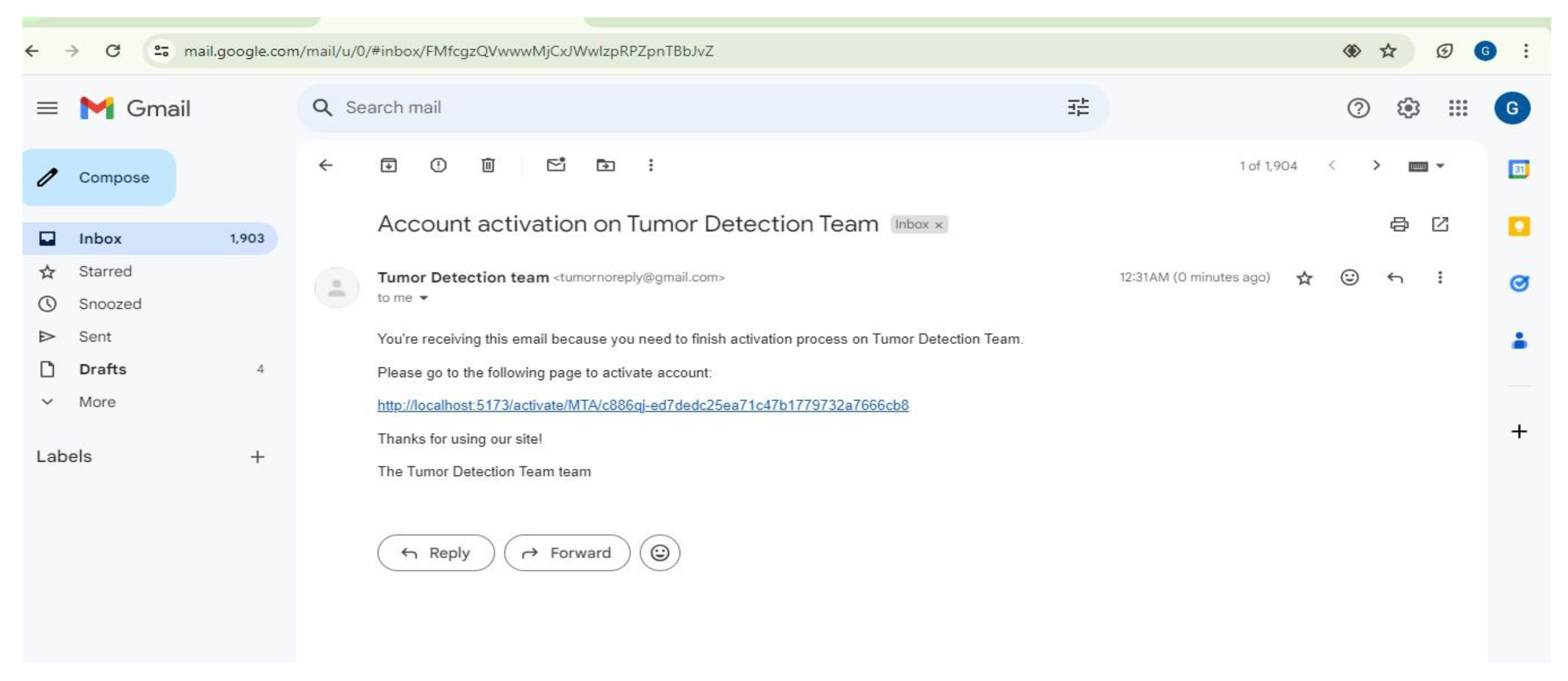
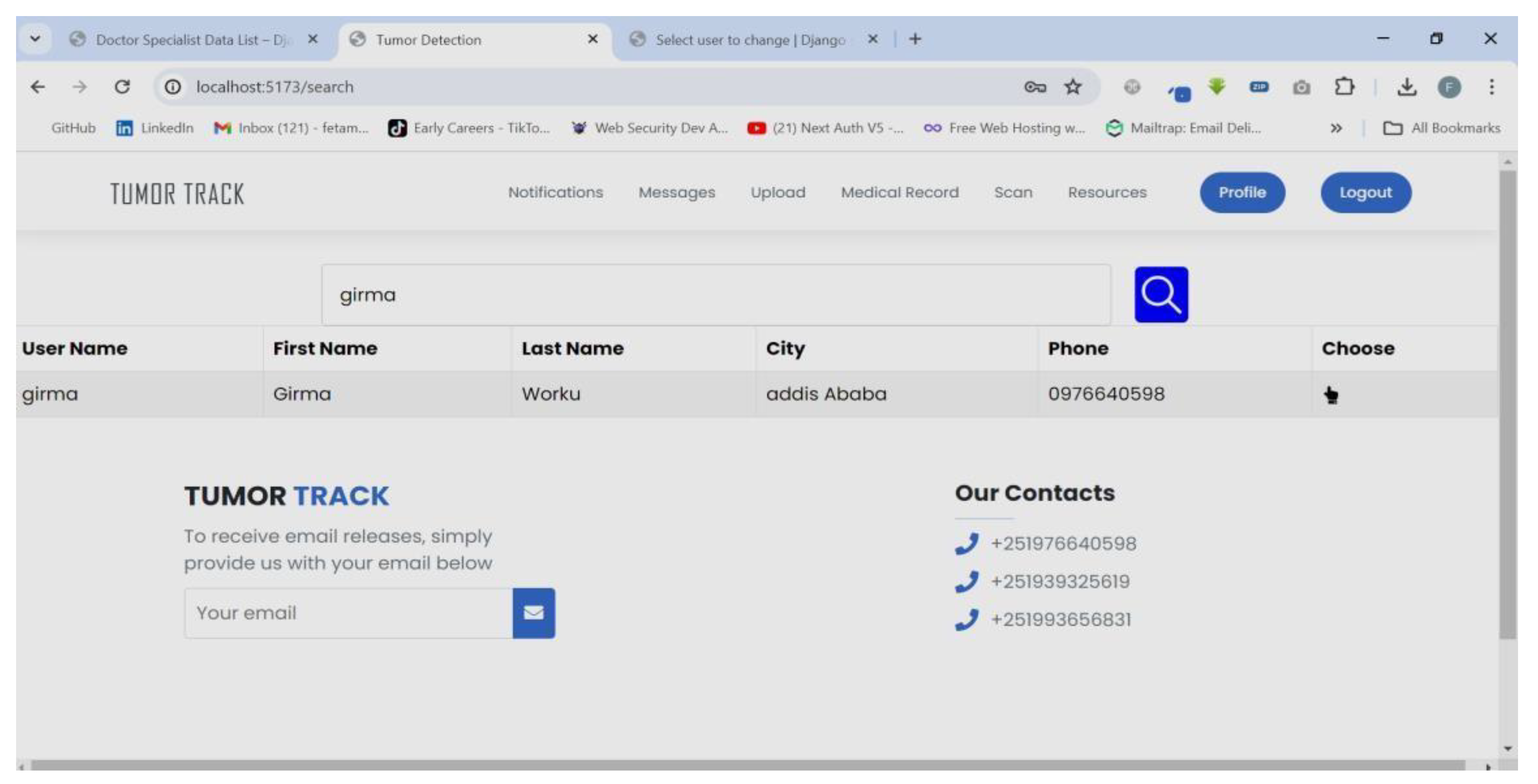
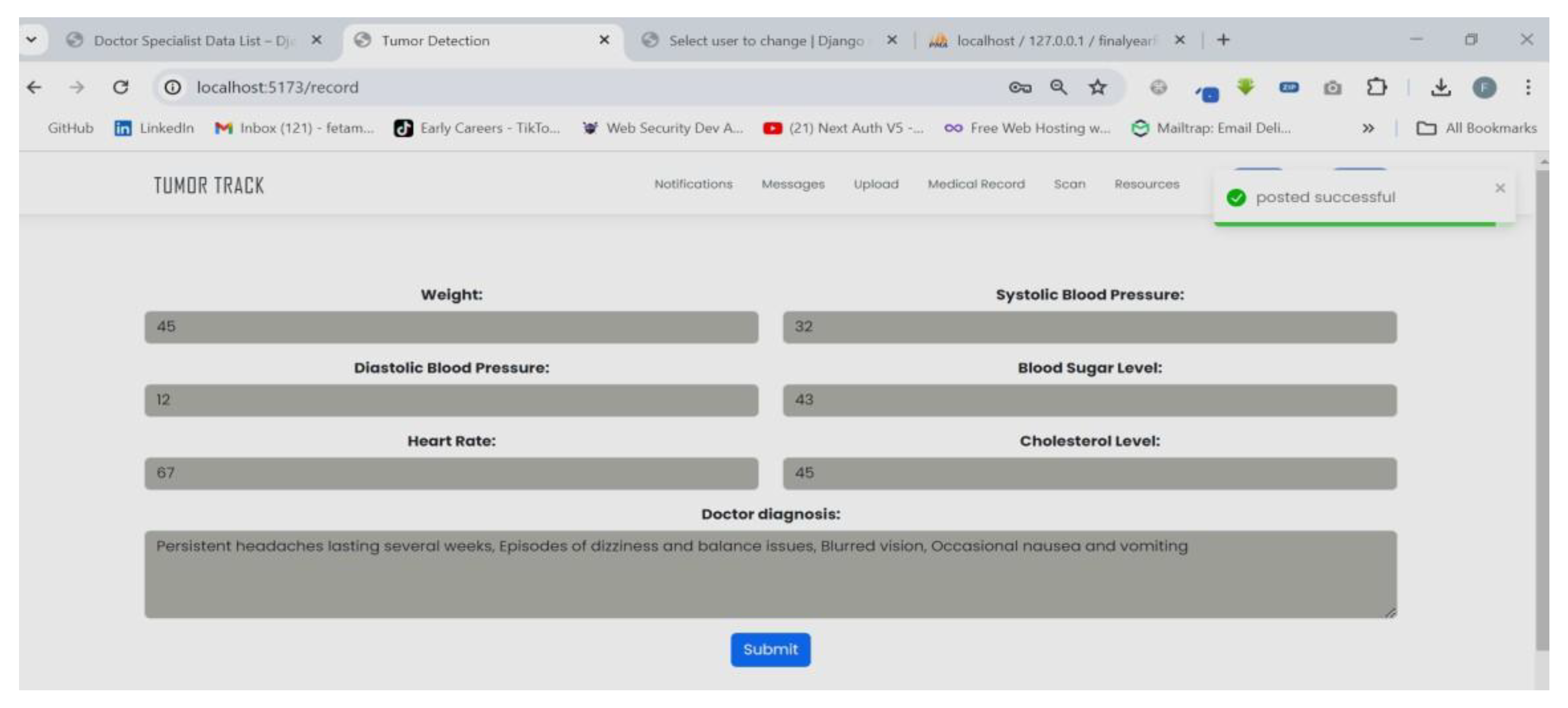
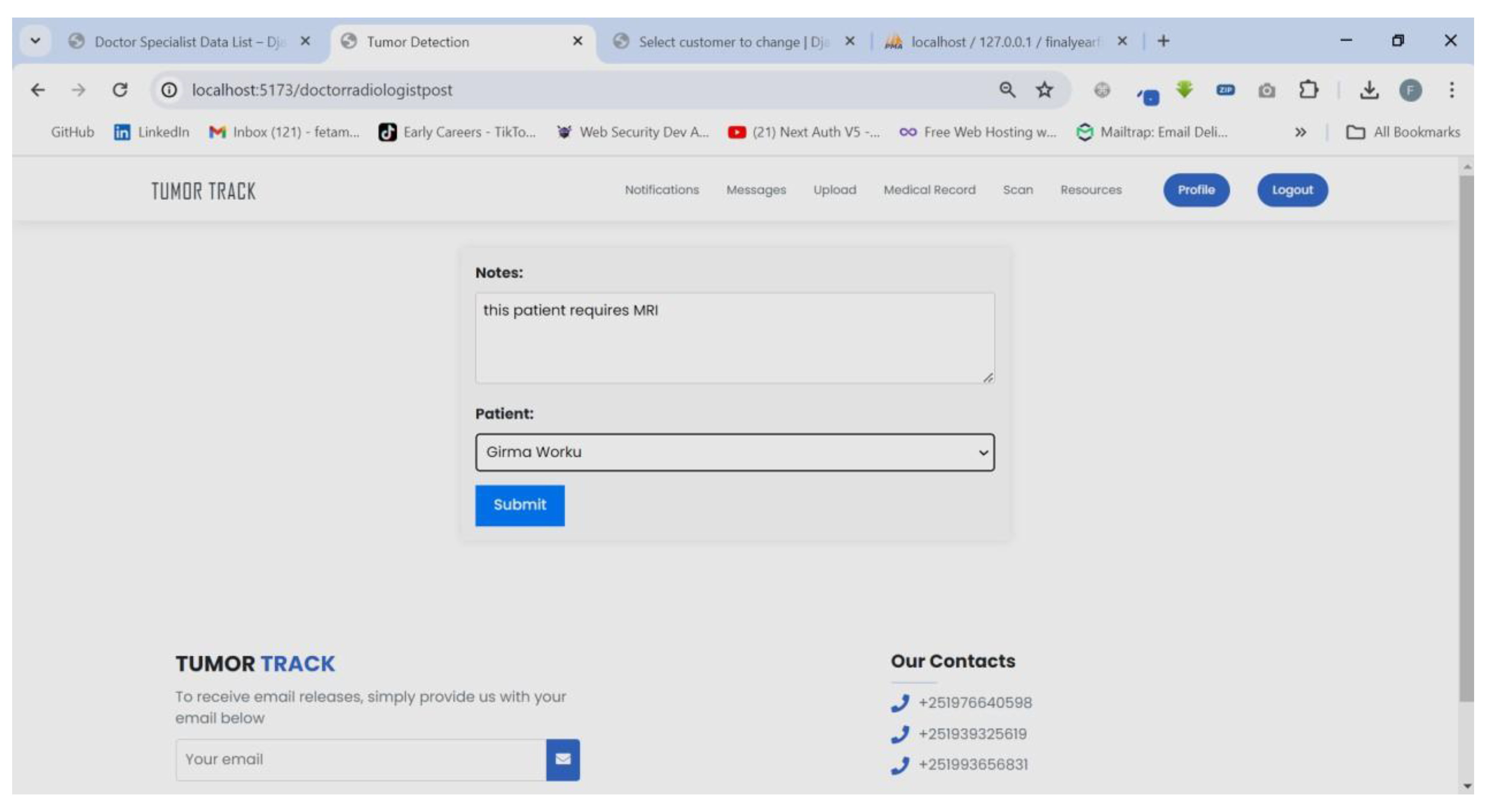
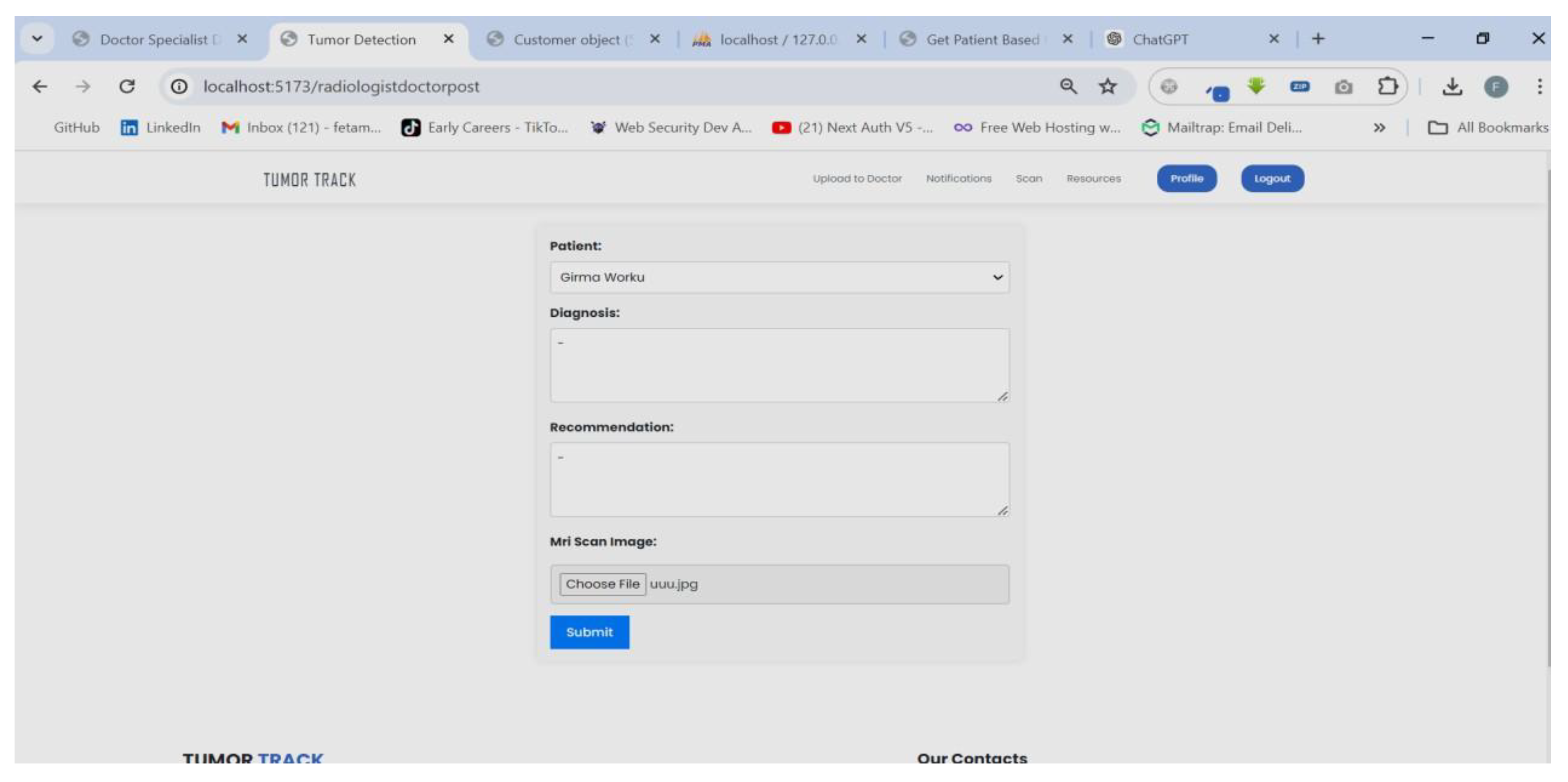

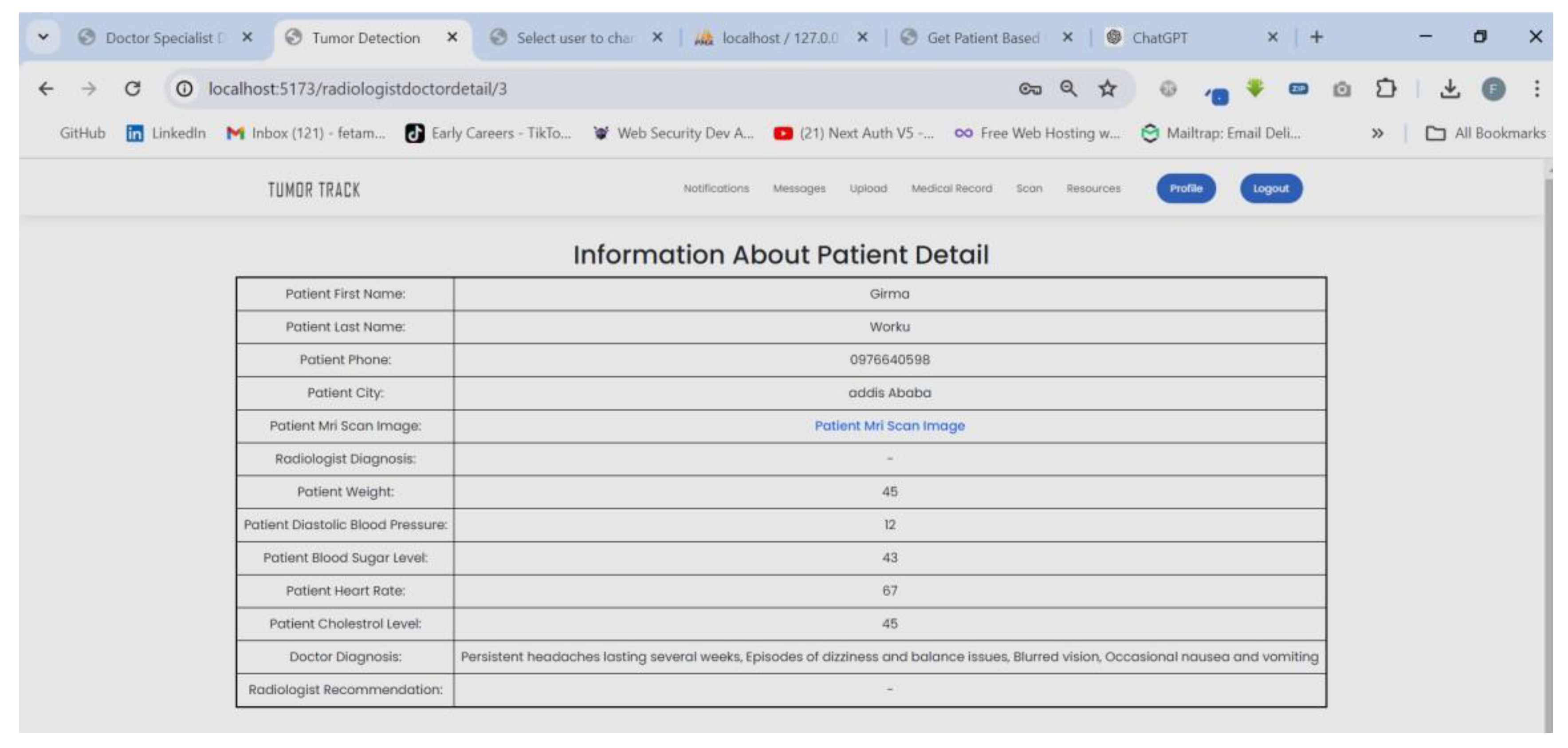
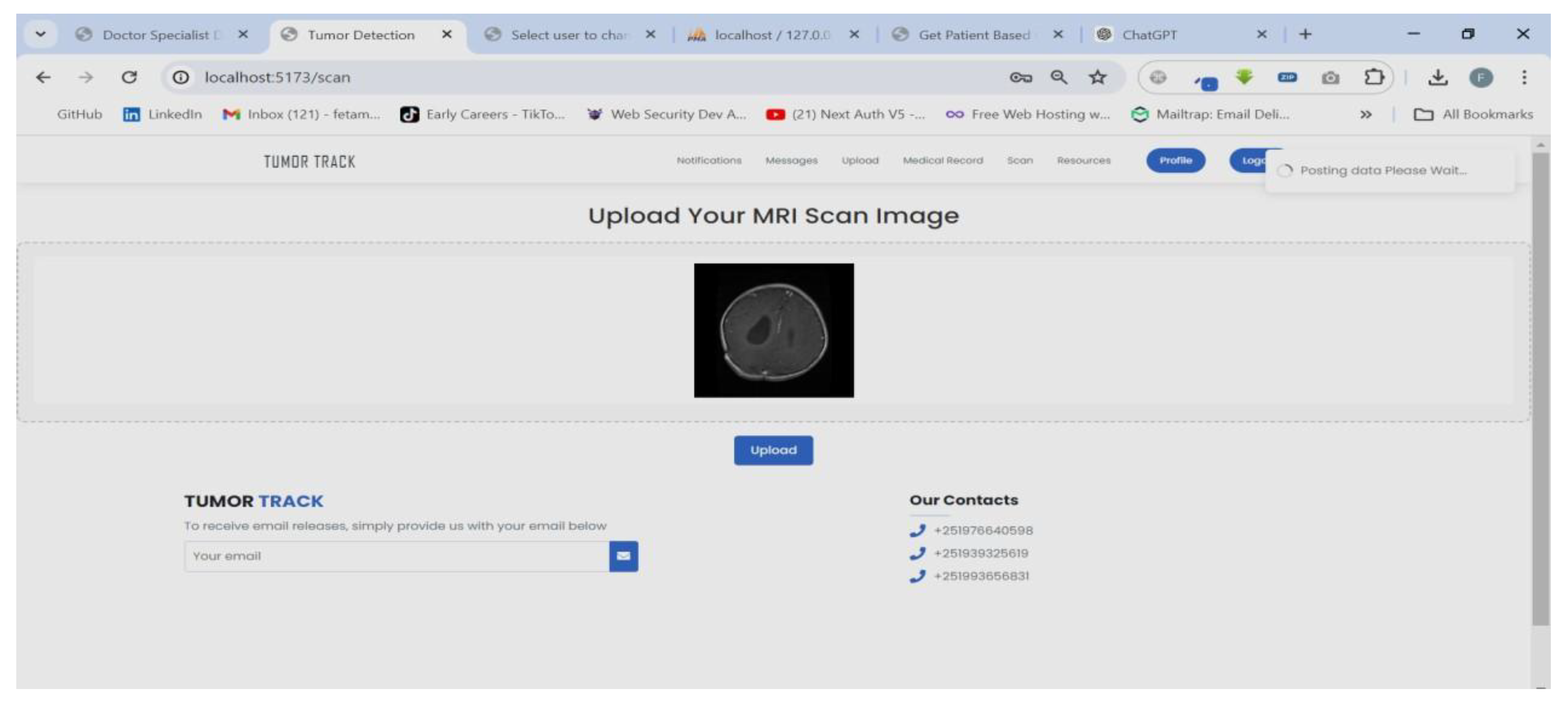
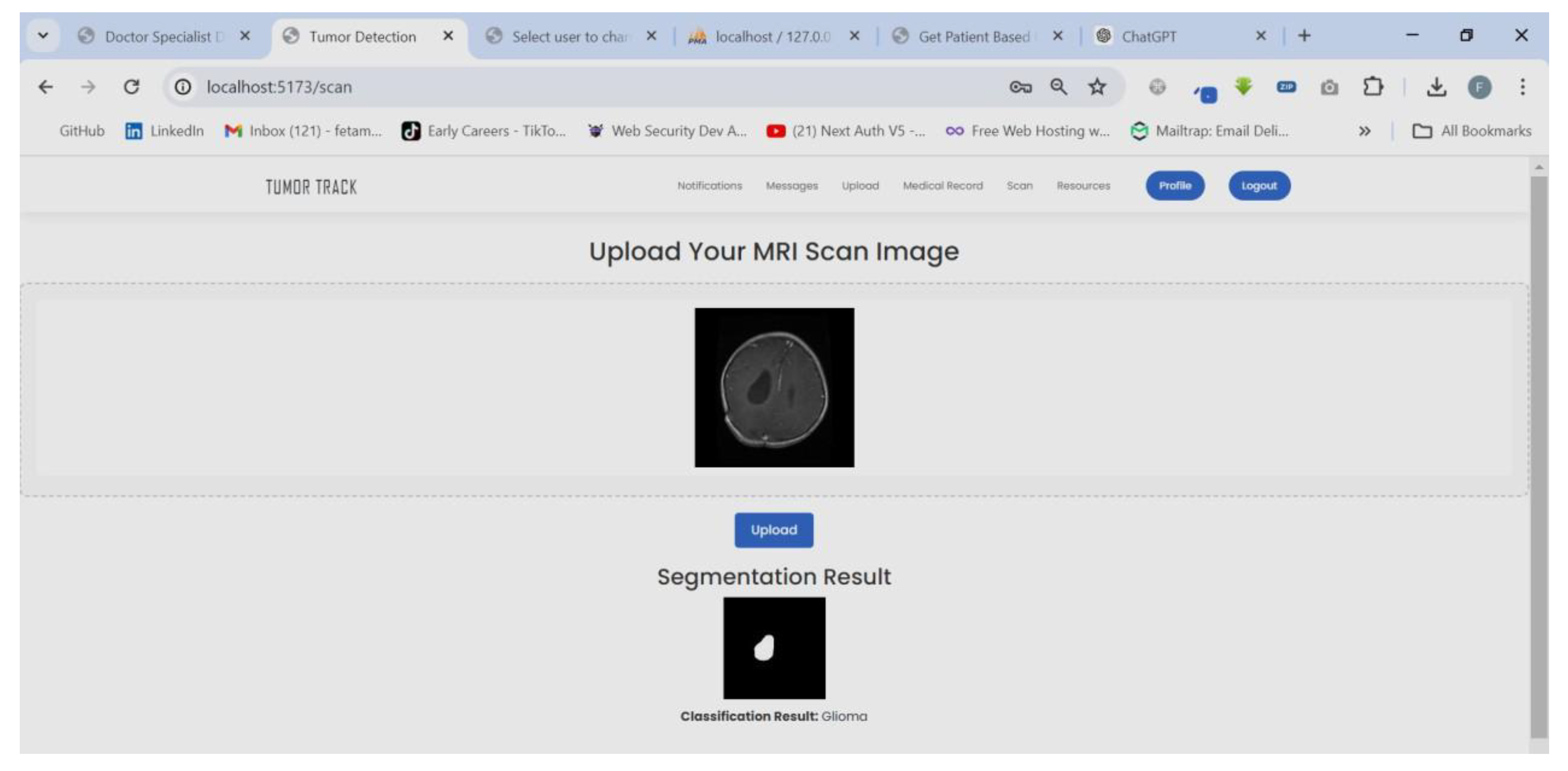
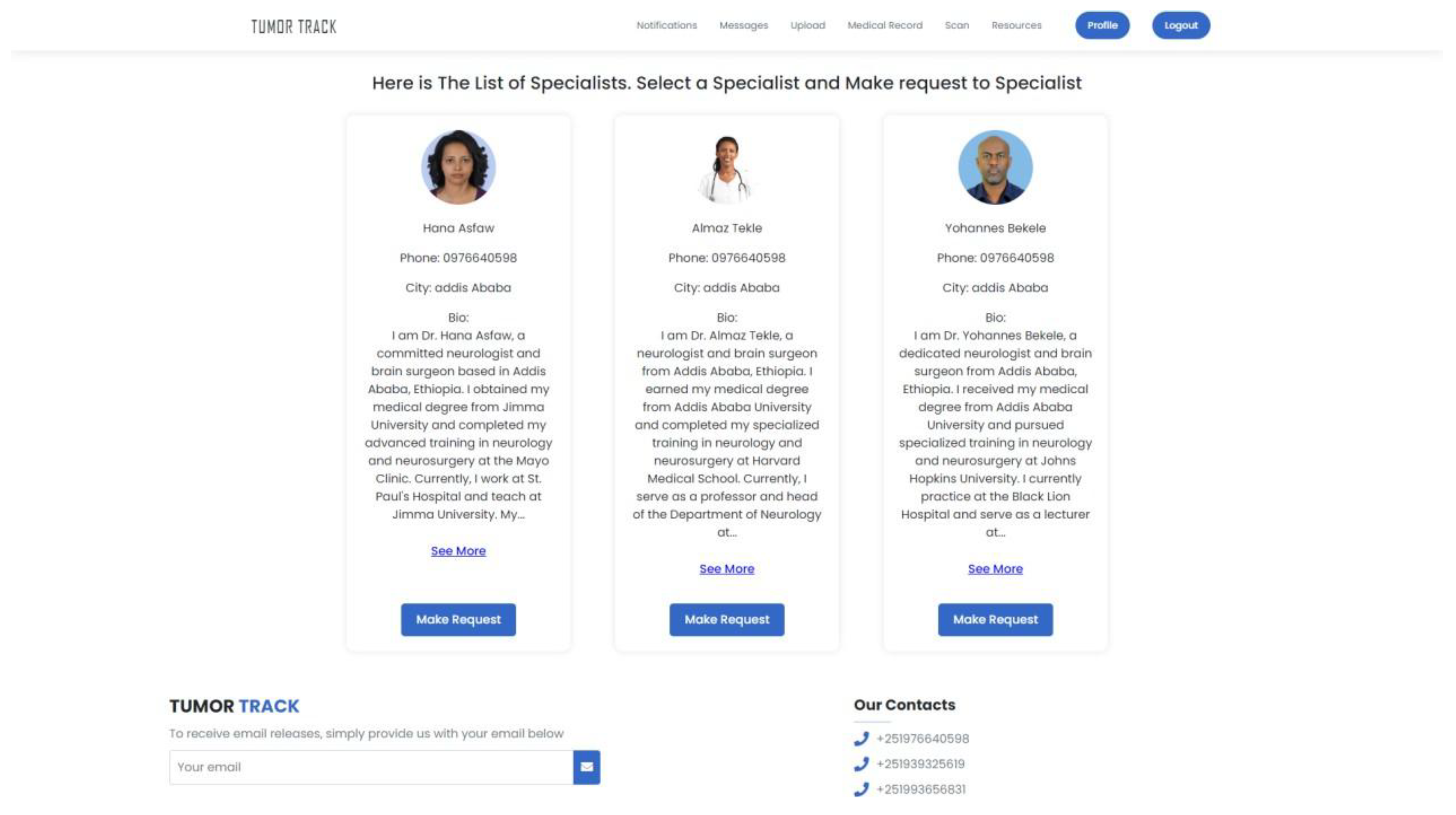
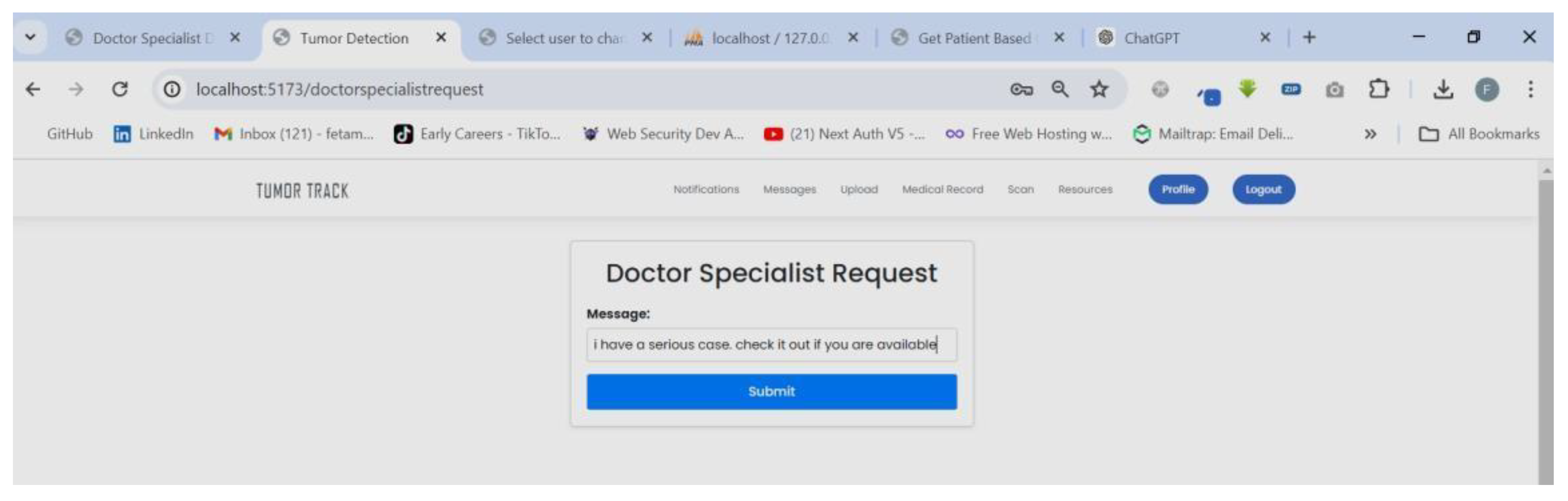


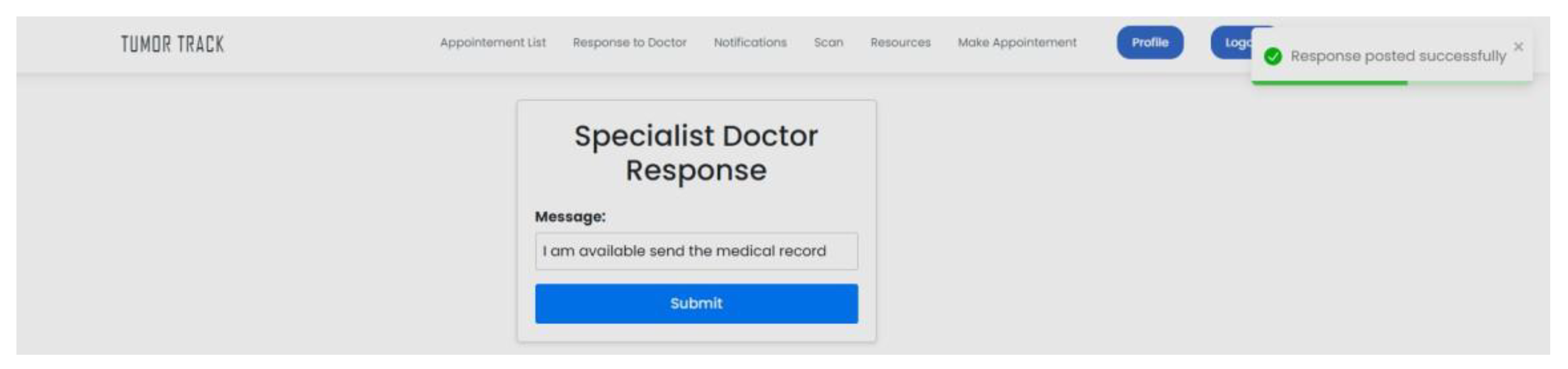

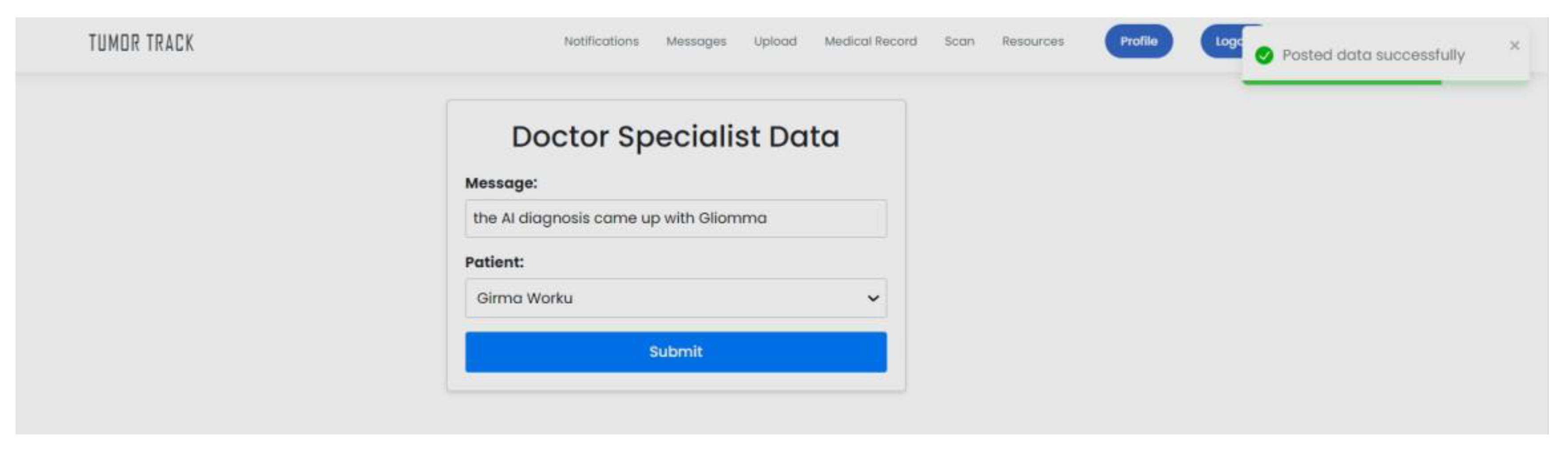
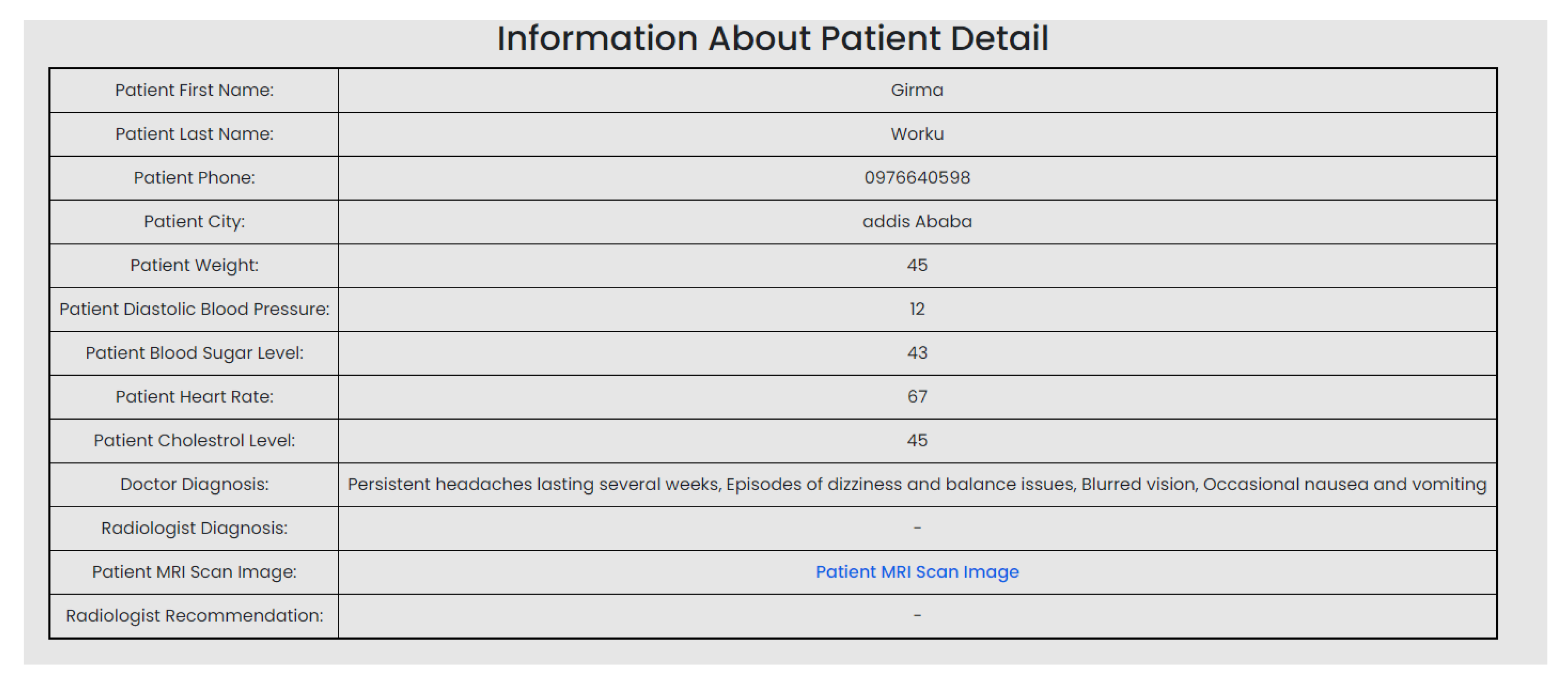
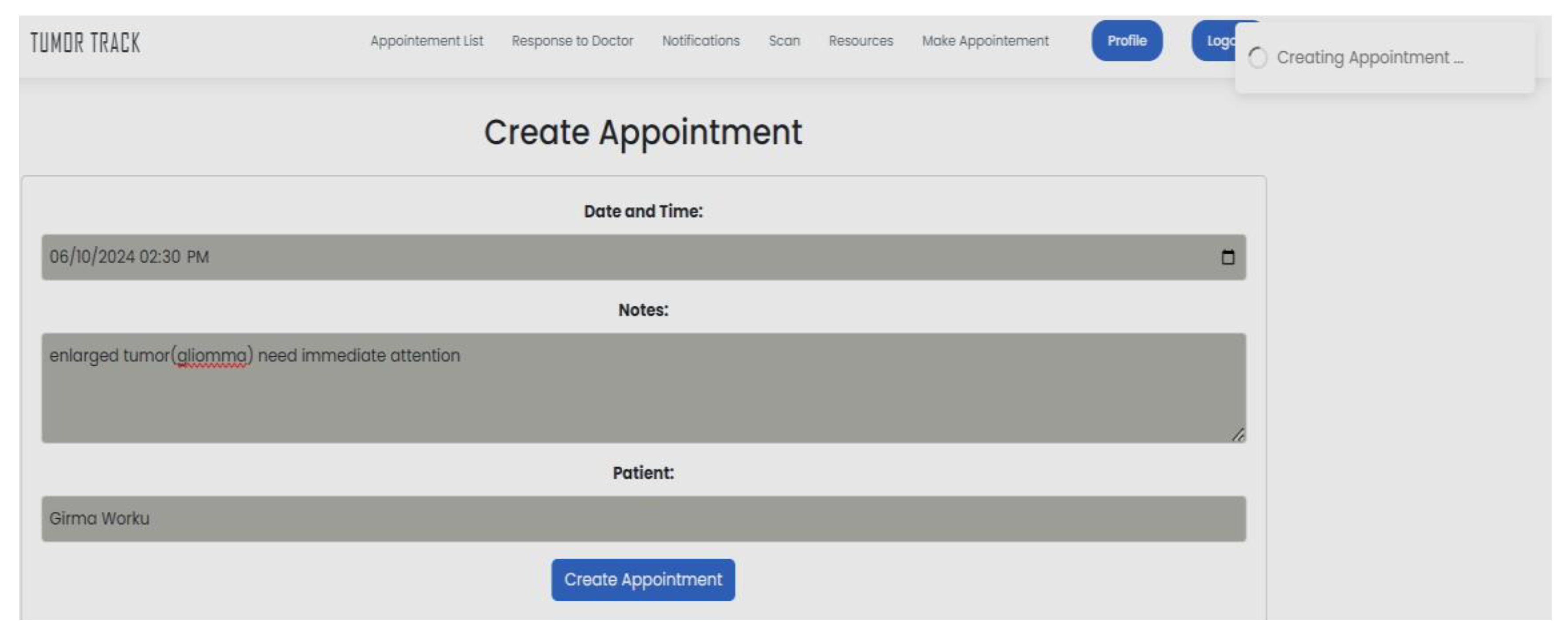

| Scenario Id | 02 |
| Scenario Name | Registration |
| Participant Actor | Admin |
| Flow of events | The admin logs into his account. Chooses add user. Chooses role for this case she is a specialist. Fills in the required form and save it. |
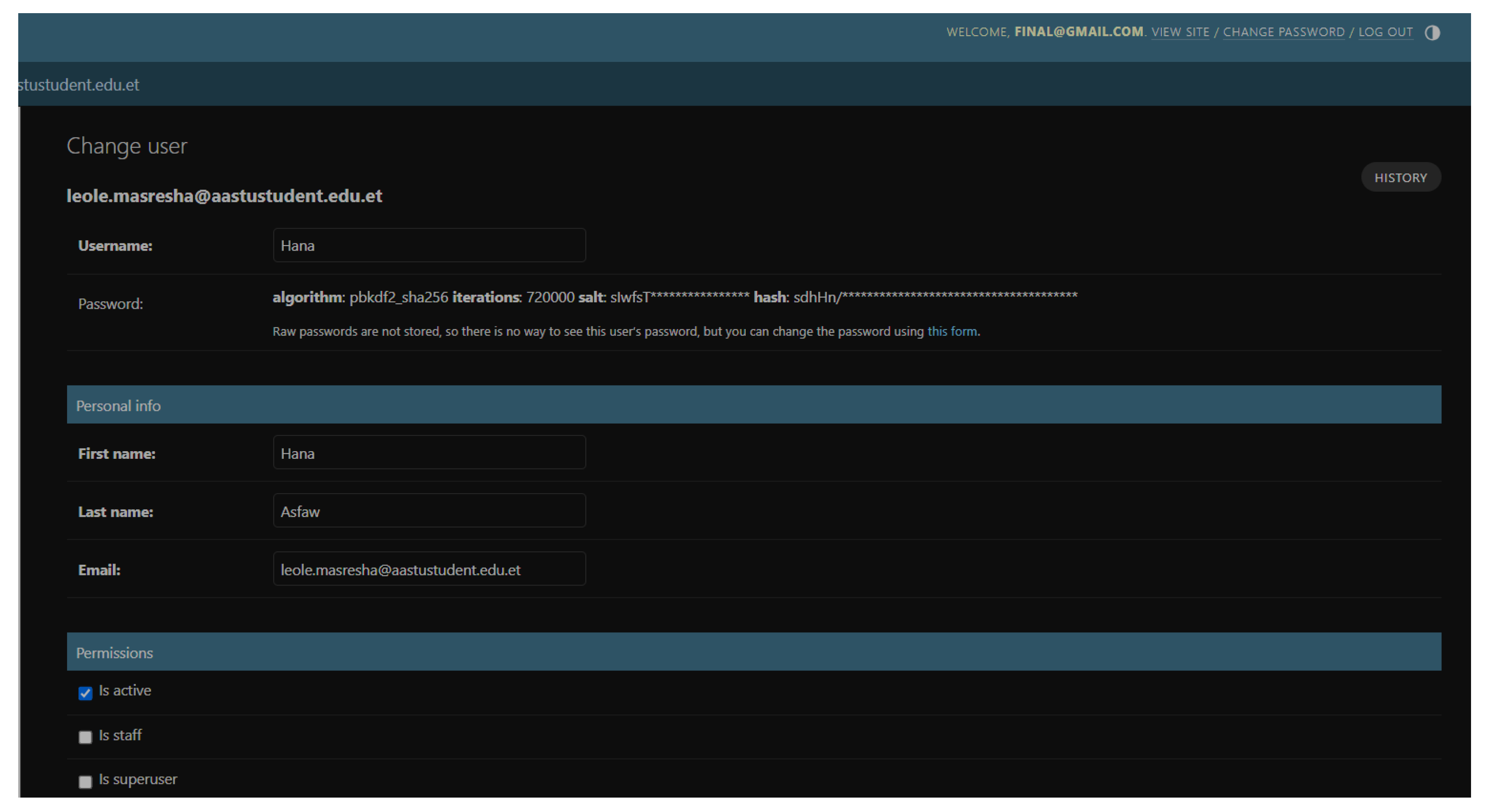
| Scenario Id | 03 |
| Scenario Name | Unable to login |
| Participant Actor | Patient (Feta M.) |
| Flow of events |
|
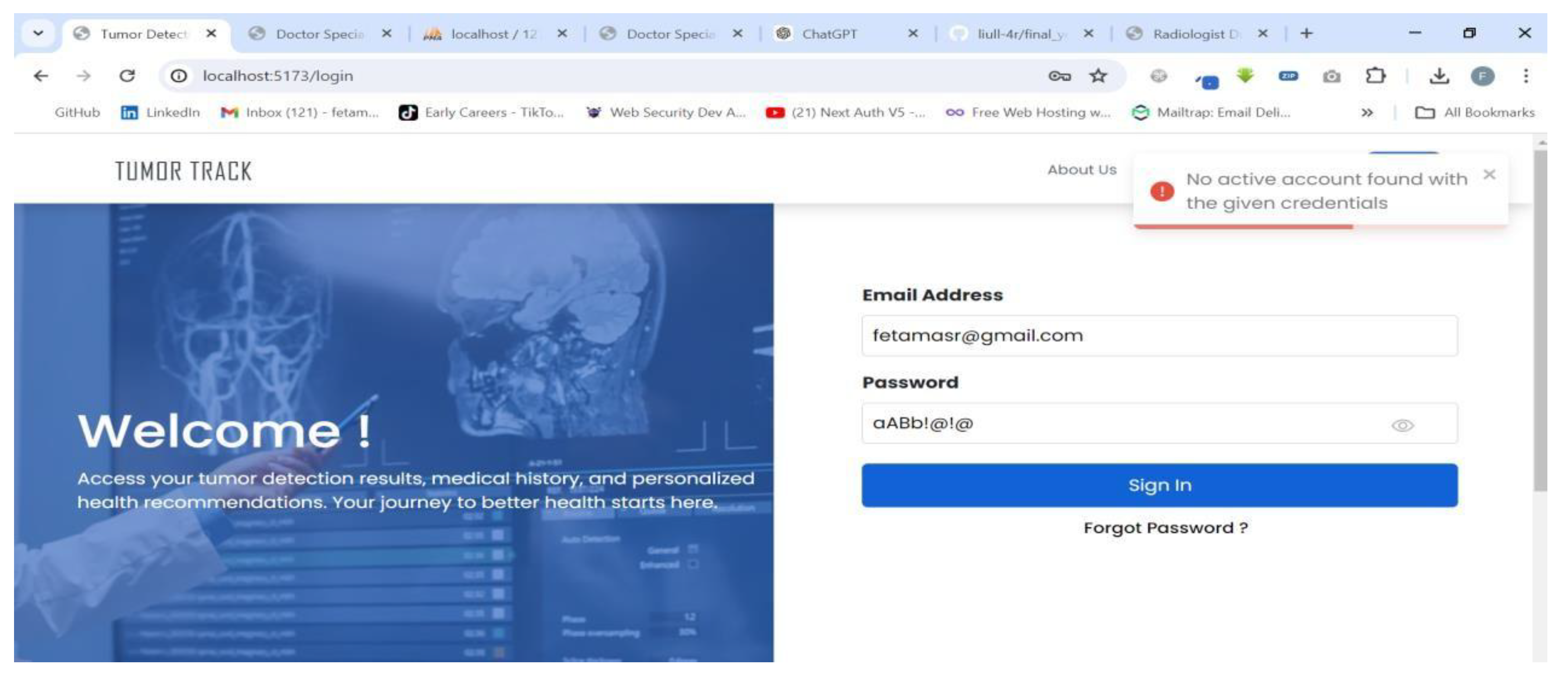
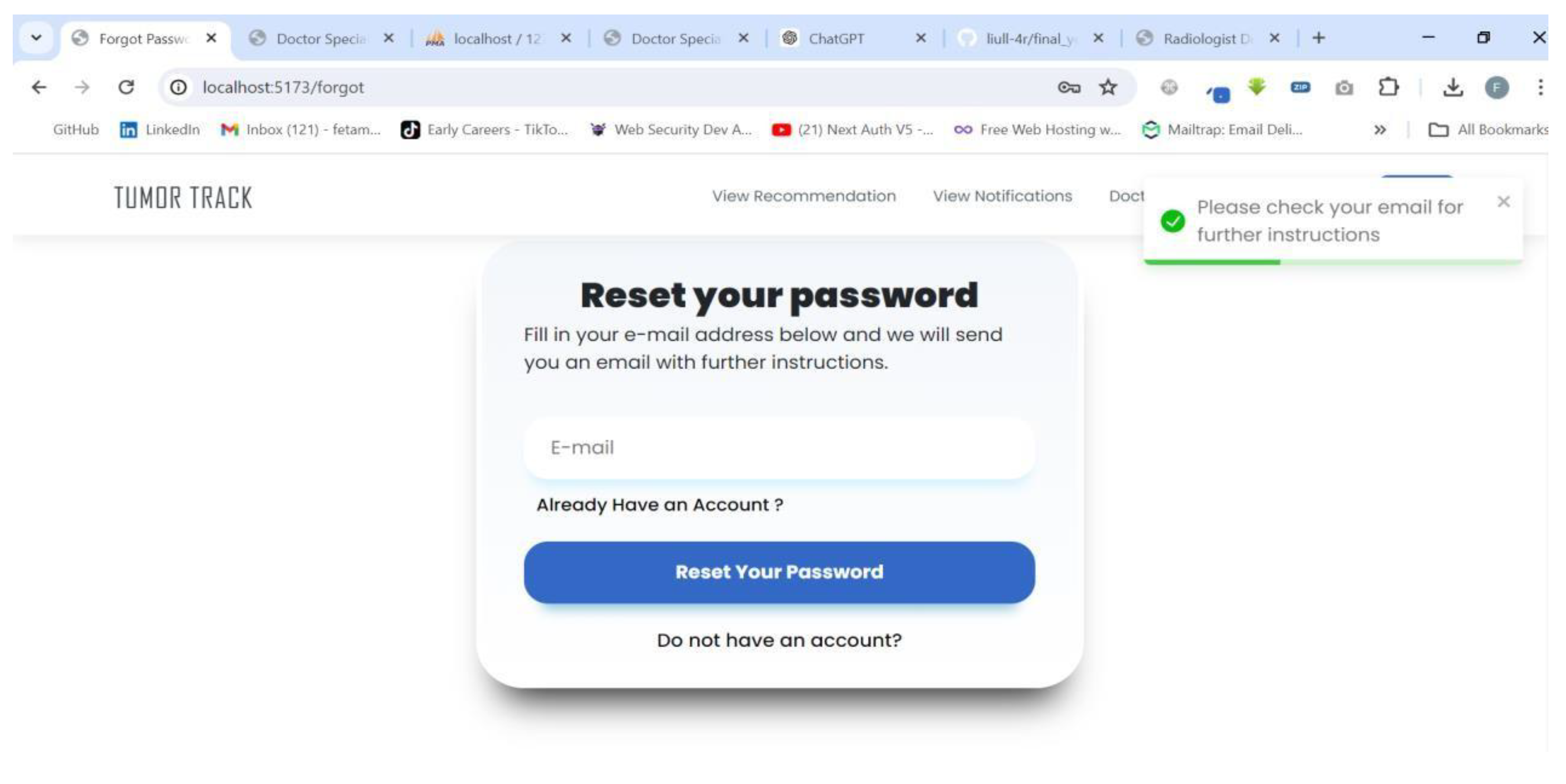
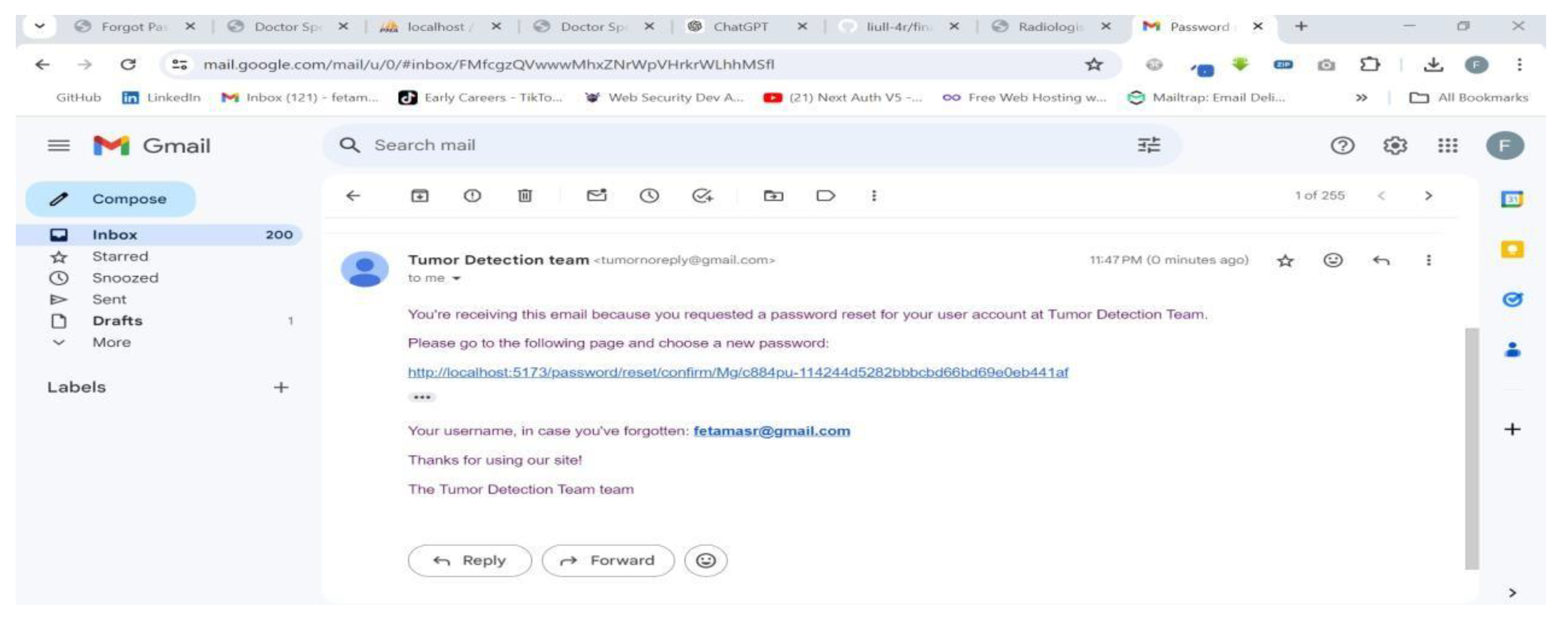
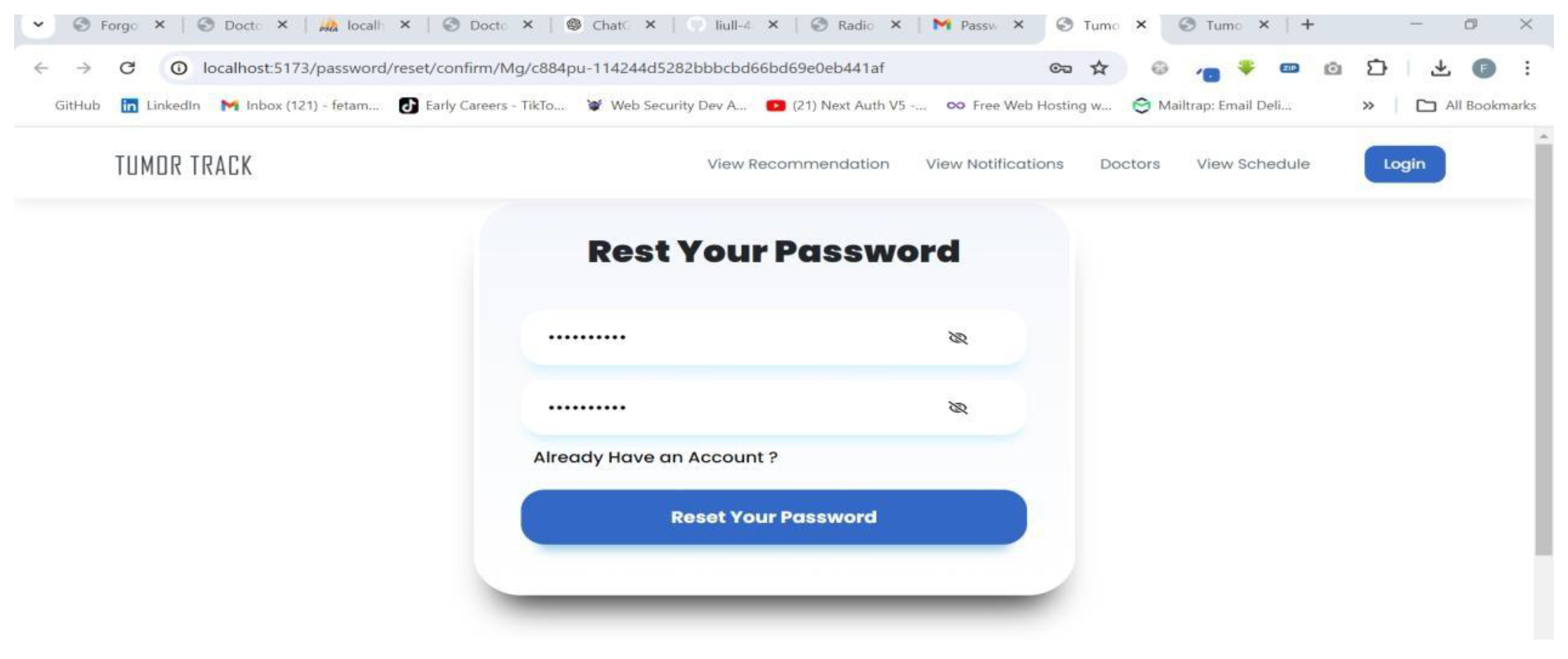
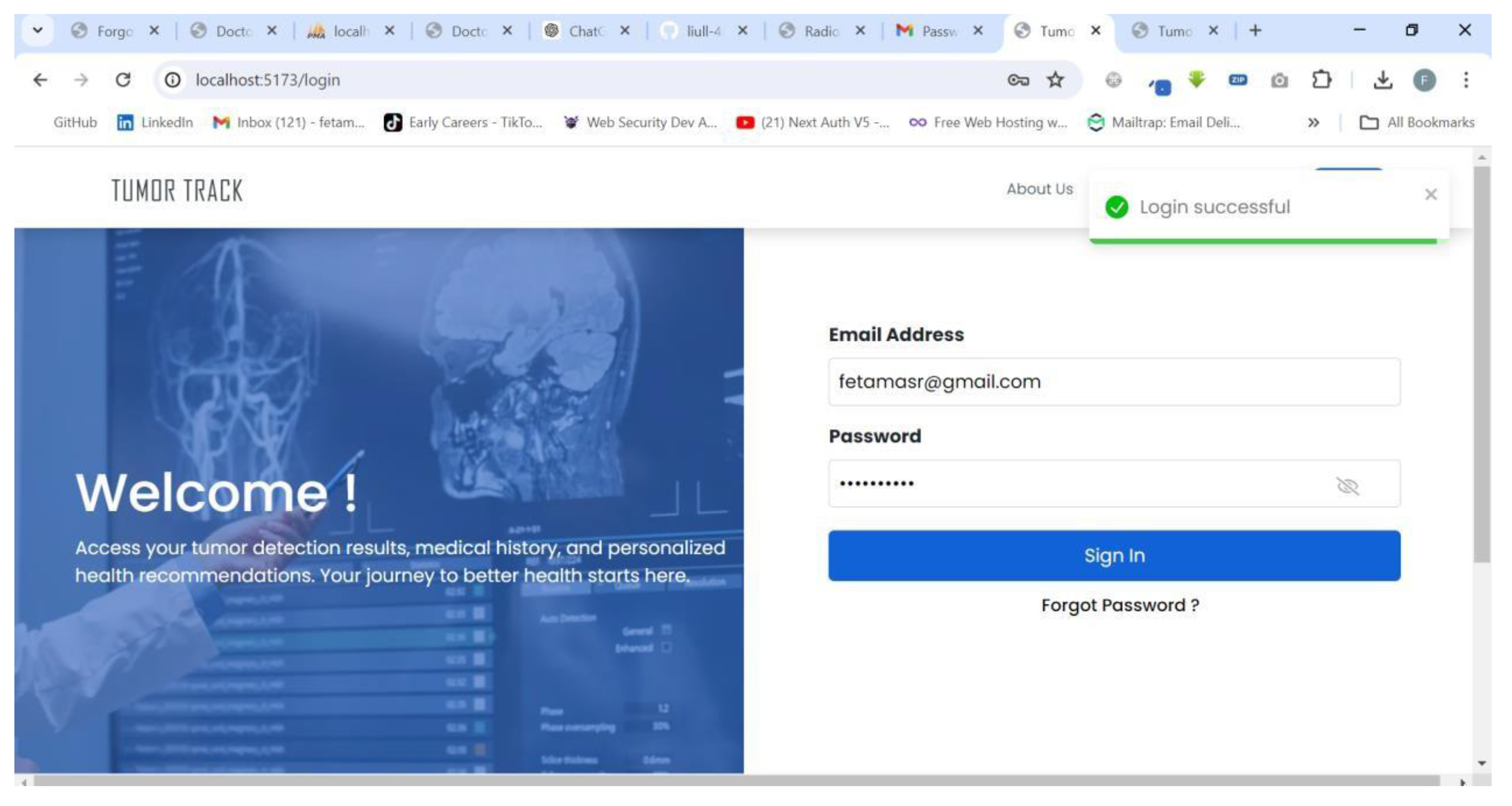
4.3.2. AI model Testing and Evaluation
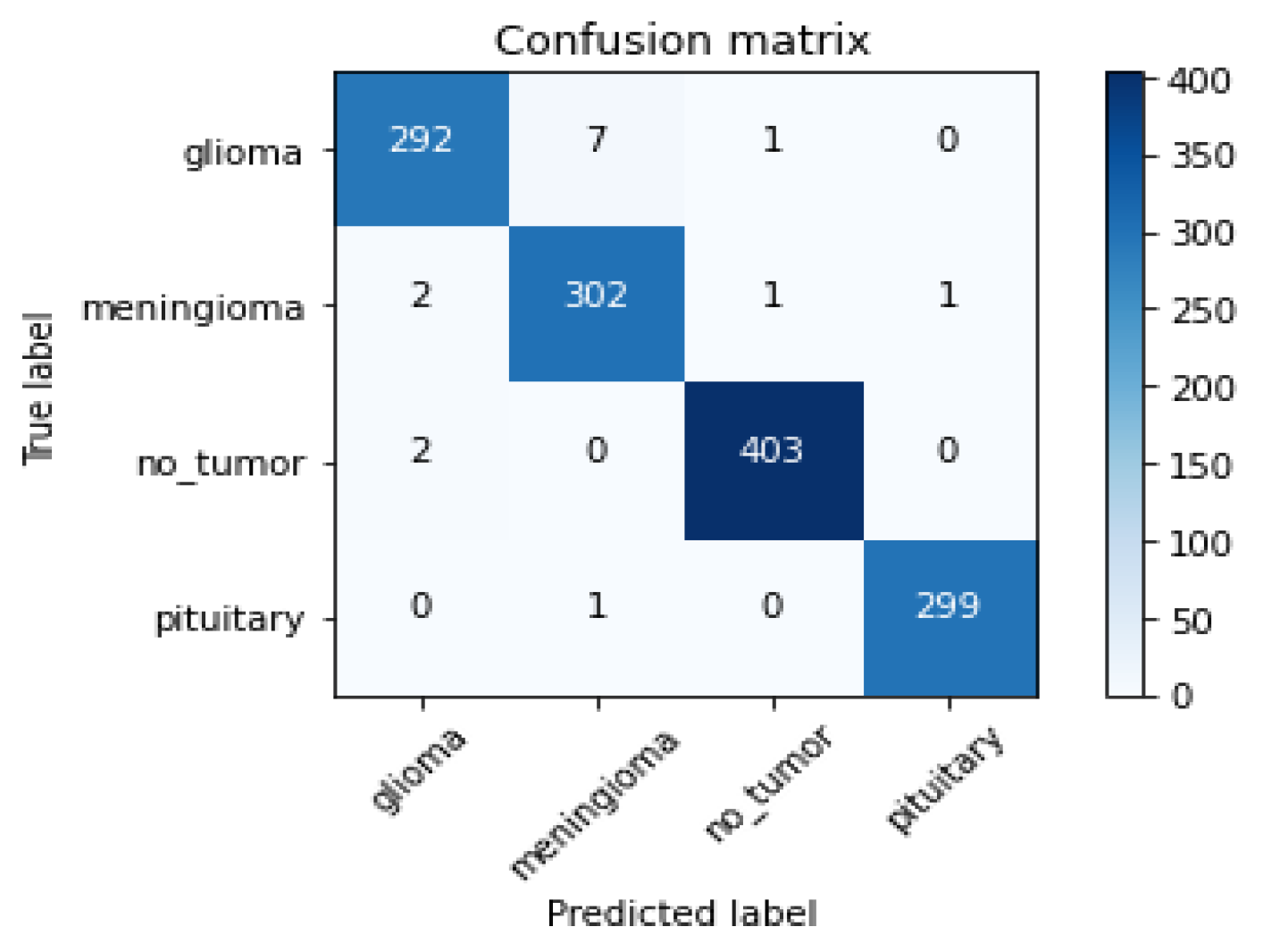
4.3.2.1. Evaluation Metrics for Classification Model (ResNet 50)
| Performance Metrics | Results |
|---|---|
| Test Accuracy | 98.86% |
| Precision | 99.00% |
| Sensitivity (Glioma) | 97% |
| Sensitivity (Meningioma) | 99% |
| Sensitivity (Pituitary) | 100% |
| Sensitivity (Normal) | 100% |
| F1-score | 99.00% |
| AUC | 1.0 |
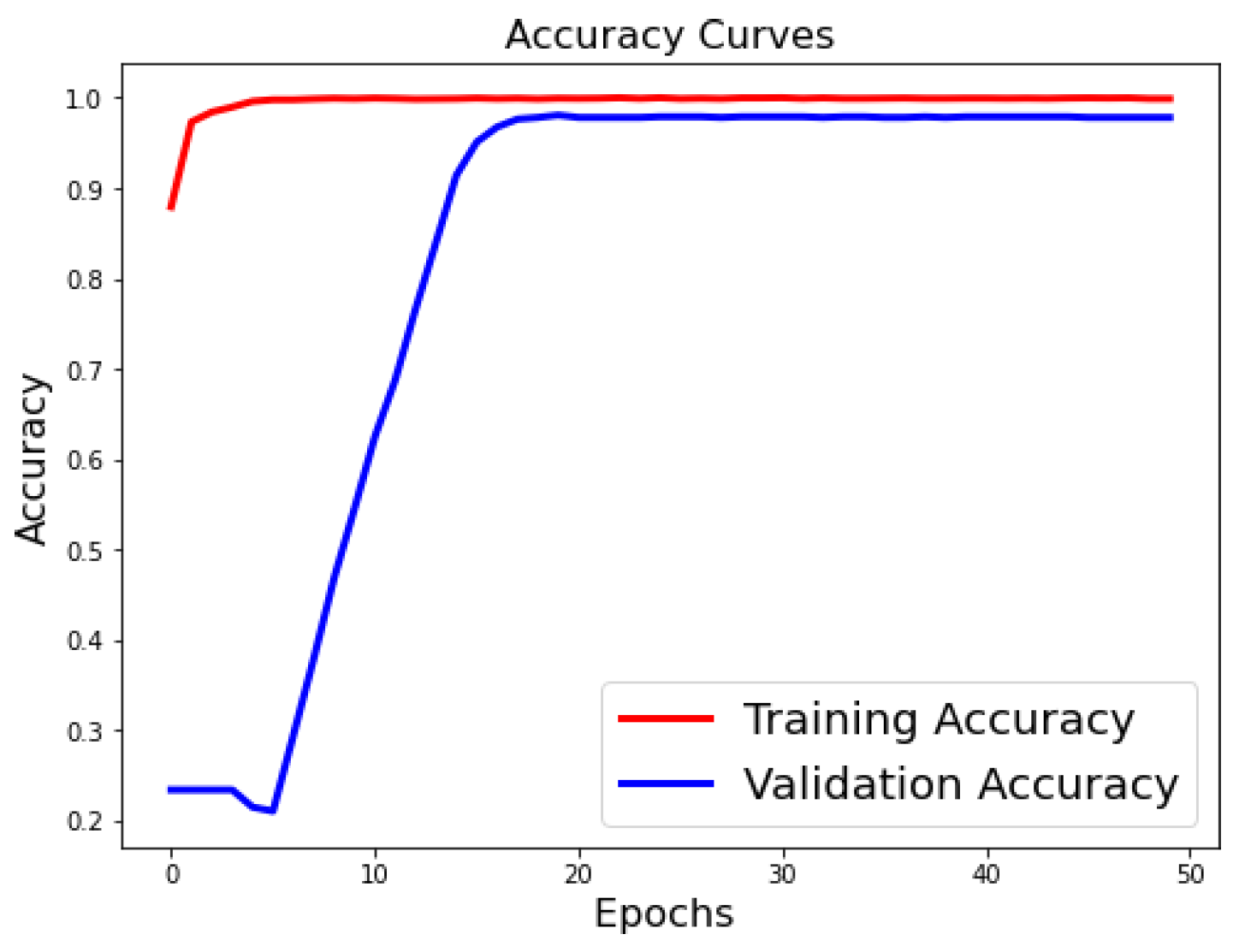
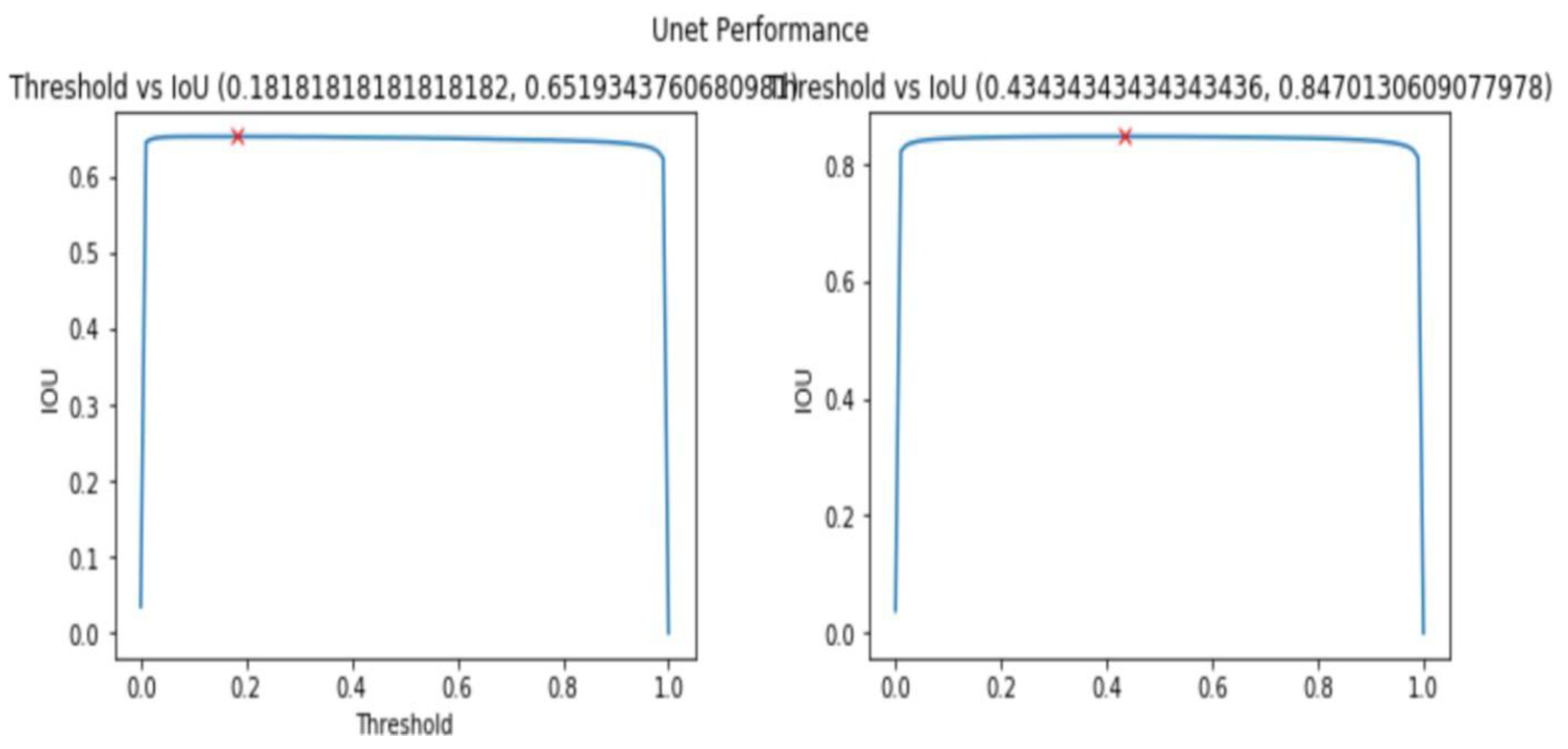
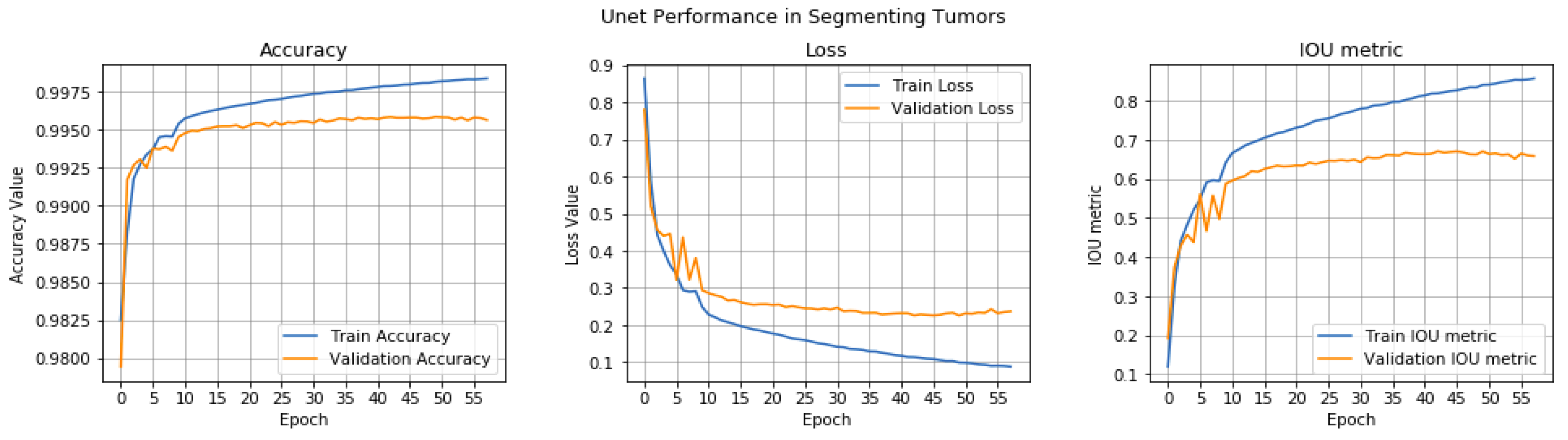
4.3.2.2. Evaluation Metrics for Segmentation Model (UNet)
| Metric | Training Value | Validation Value |
|---|---|---|
| Accuracy | 0.9975 | 0.9950 |
| Loss | 0.1 | 0.2 |
| IoU Metric | 0.8 | 0.6 |
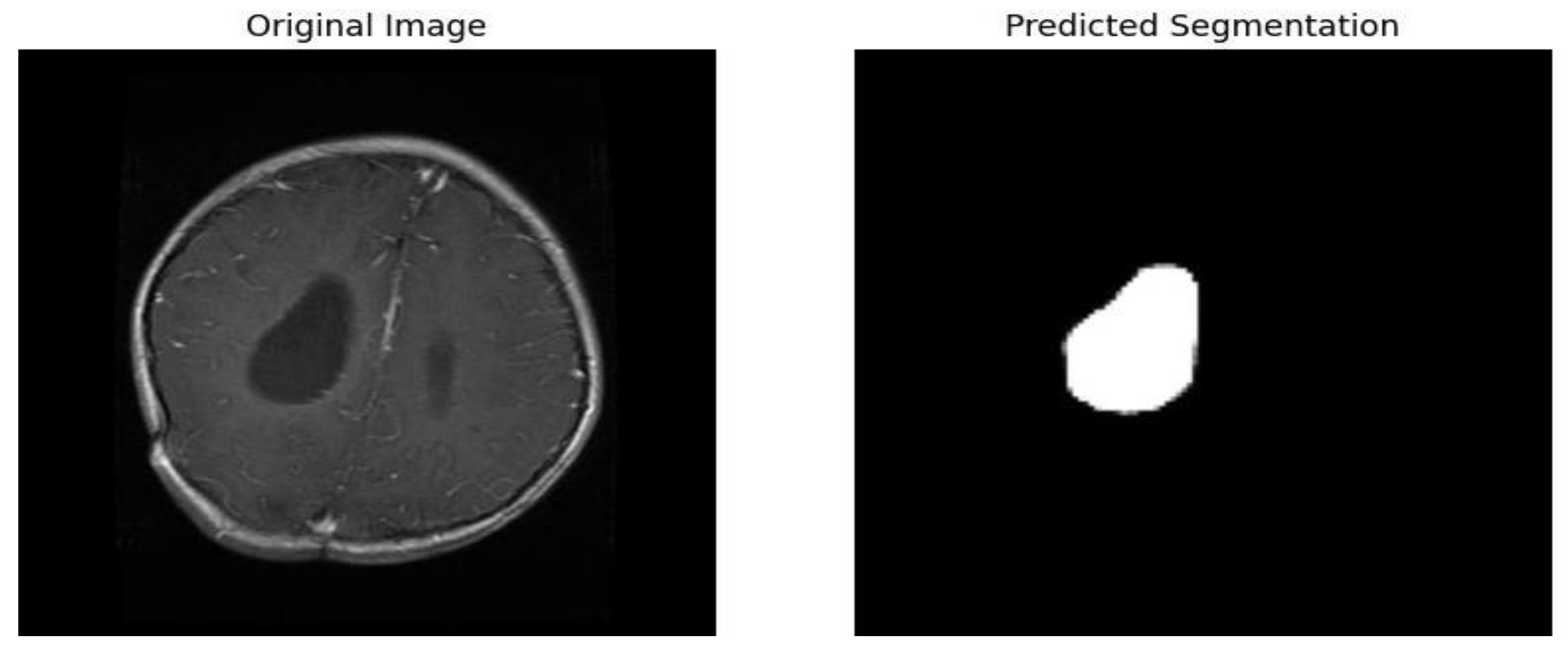
4.3.3. Integration
4.4. Discussion
4.4.1. Challenges Faced
4.4.2. Expenses
4.4.3. Contributions
4.4.4. Limitations
Chapter Five Conclusion and Recommendation
5.1. Conclusion
5.2. Recommendation
5.3. Future Directions
Acknowledgment
Appendix A
- Backend sample code:
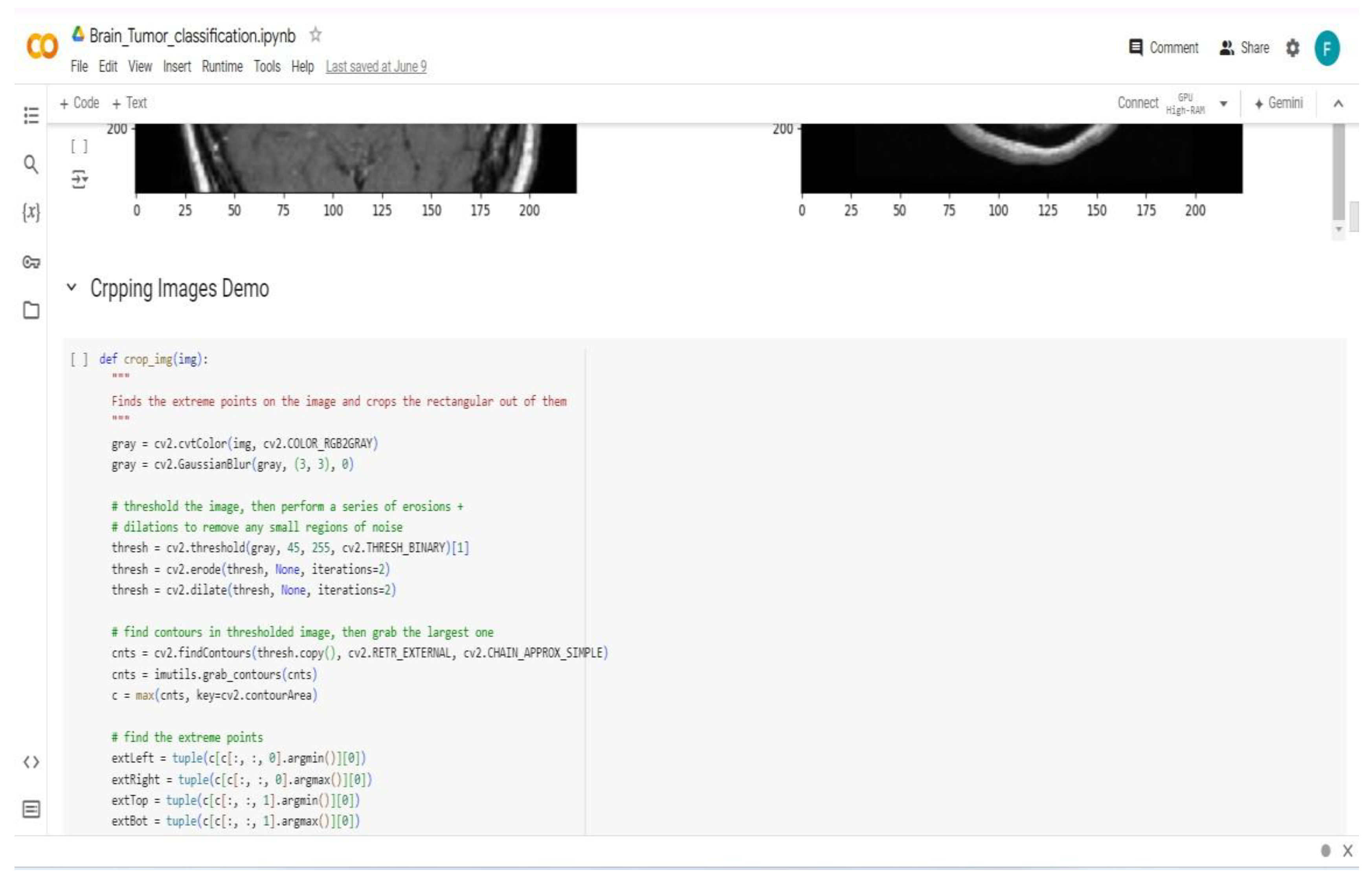
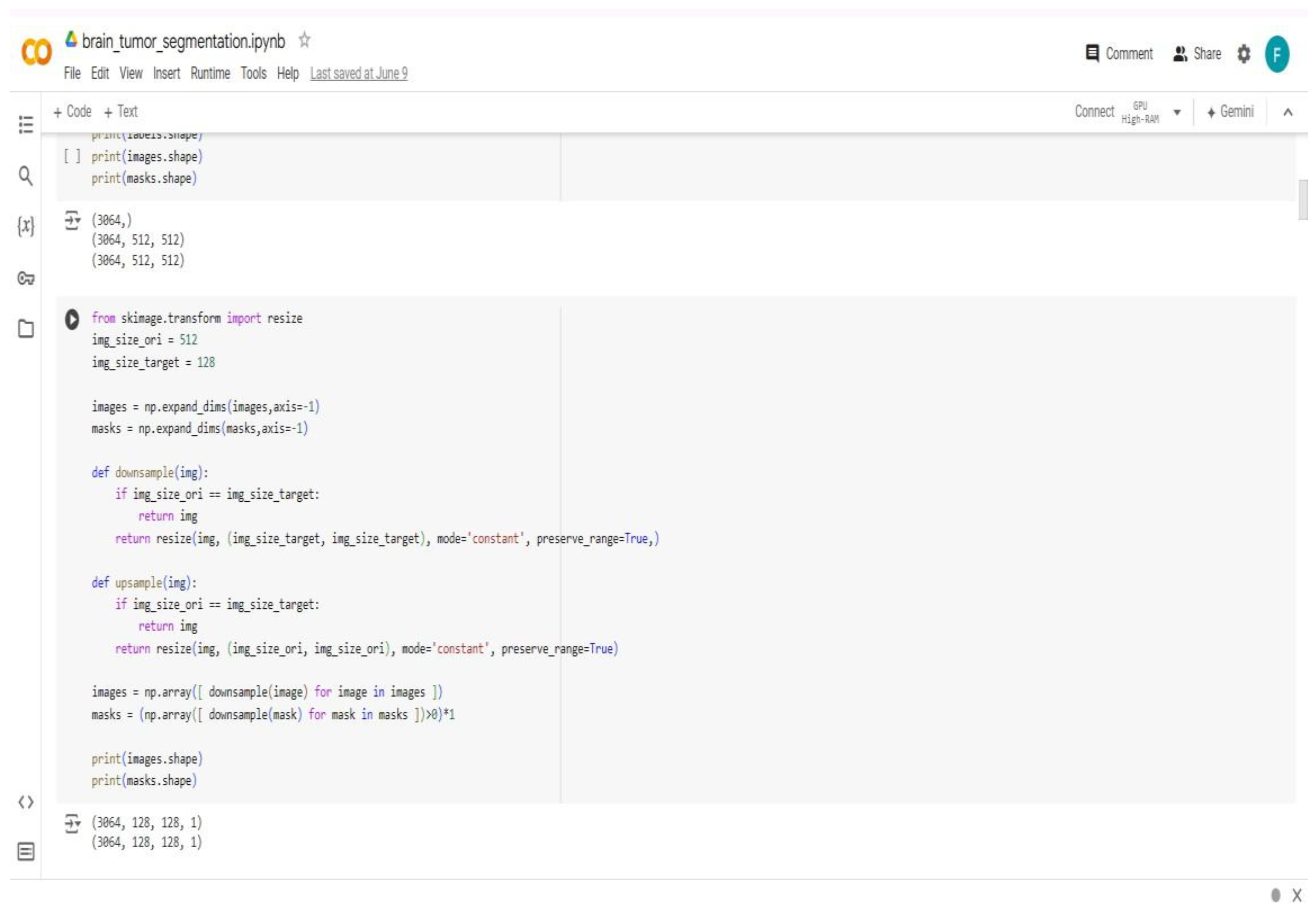
Appendix B
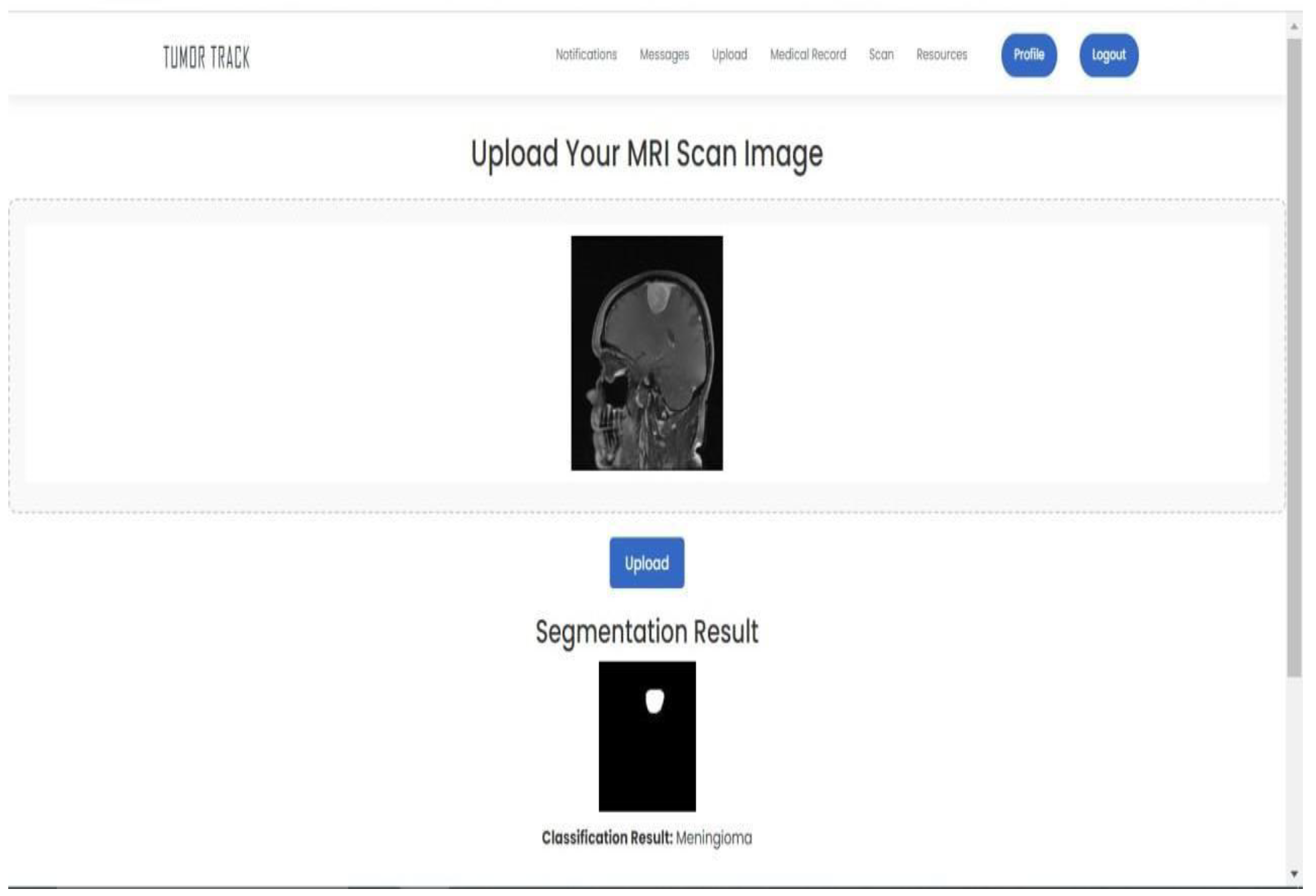
References
- Buser, A.A. The Training and Practice of Radiology in Ethiopia: Challenges and prospect. Ethiop J Health Sci. 2022, 32, 1–2. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kaifi, R. A Review of Recent Advances in Brain Tumor Diagnosis Based on AI- Based Classification. Diagnostics (Basel). 2023, 13, 3007. [Google Scholar] [CrossRef] [PubMed]
- Sagaro, G.G.; Battineni, G.; Amenta, F. Barriers to Sustainable Telemedicine Implementation in Ethiopia: A Systematic Review. Telemed Rep. 2020, 1, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Smith, J.; Johnson, K.; Lee, S.; Wang, X. Patient engagement and usability of a web-based patient follow-up application. Journal of Medical Informatics 2023, 123, 456–789. [Google Scholar]
- Johnson, M.; Lee, T.; Chen, Y. Impact of a patient follow-up web application on healthcare utilization and cost reduction. Journal of Healthcare Management 2022, 67, 123–135. [Google Scholar]
- Lee, J.; Wang, Z.; Chen, X. Application of deep learning in brain tumor detection from MRI images. Journal of Neuroradiology 2021, 48, 123–456. [Google Scholar]
- Wang, Y.; Chen, L. Image segmentation and classification algorithms for brain tumor detection. Journal of Imaging Science and Technology 2020, 64, 040402. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




