Submitted:
01 August 2025
Posted:
04 August 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
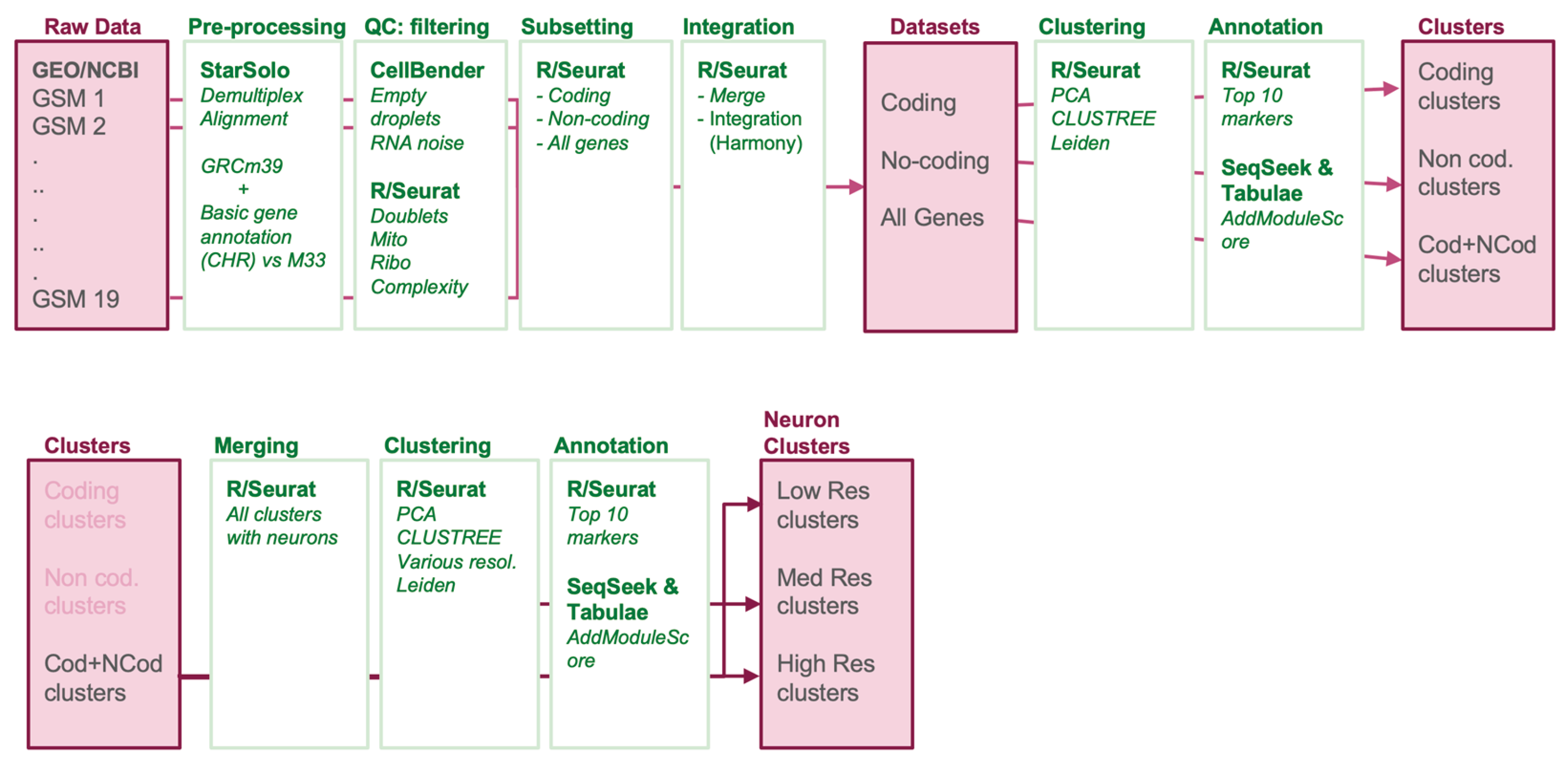
2. Materials and Methods
2.1. Data Selection and Preprocessing
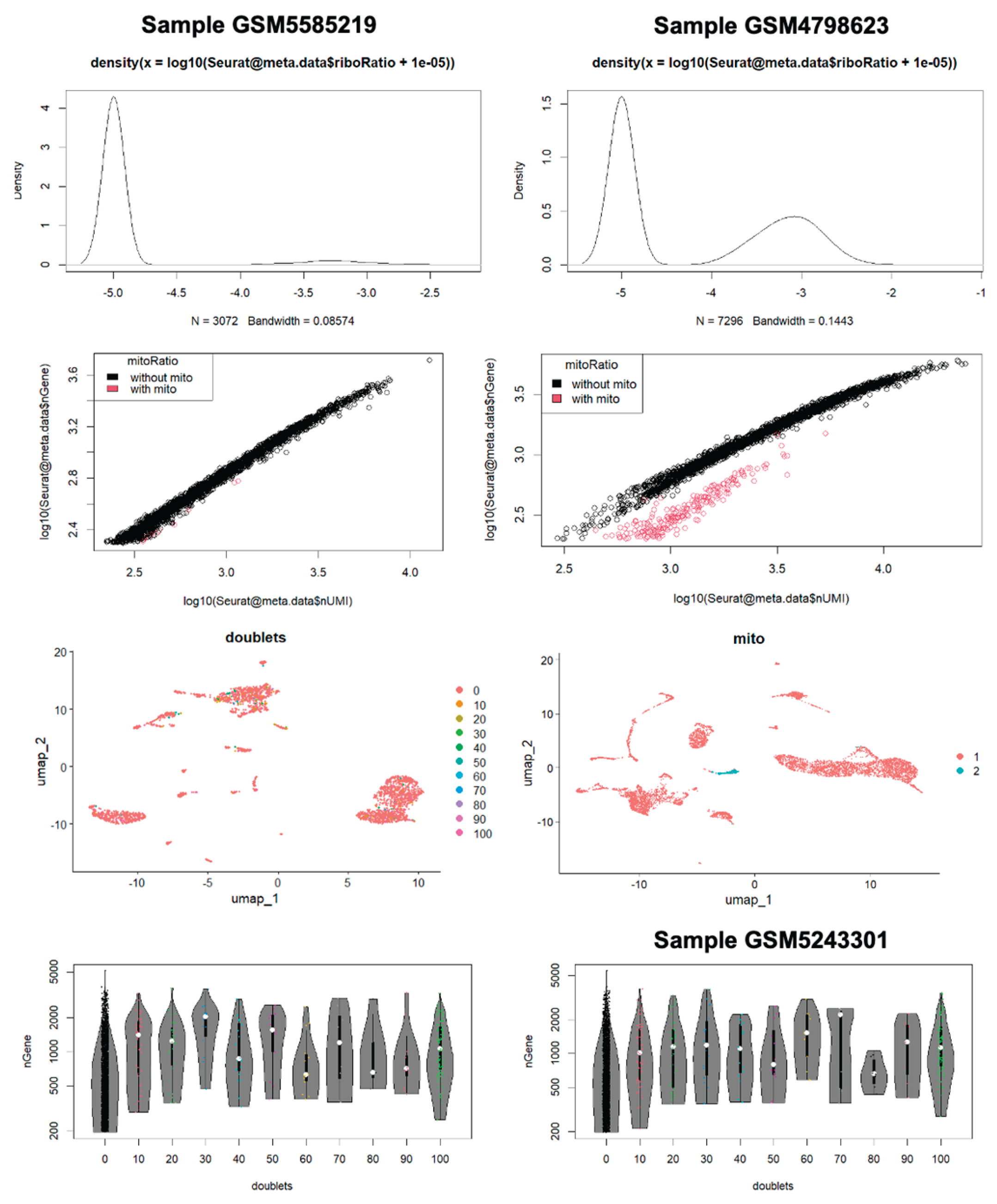
2.2. Quality Control and Cell Filtering
2.3. Gene Set Definition: Coding, Non-Coding, and Combined
- Combined (ALL): Dataset with all transcripts included in the M33 annotation of the genome;
- Coding genes (CG): a subset of the transcripts annotated as “protein_coding”;
- Non-coding genes (NCG): a subset of the transcripts annotated as “lncRNAs”, “antisense”, “pseudogenes”, “TECs”, and “non-coding isoforms”.
2.4. Normalization and Integration
2.5. Clustering
2.6. Cell Type Annotation
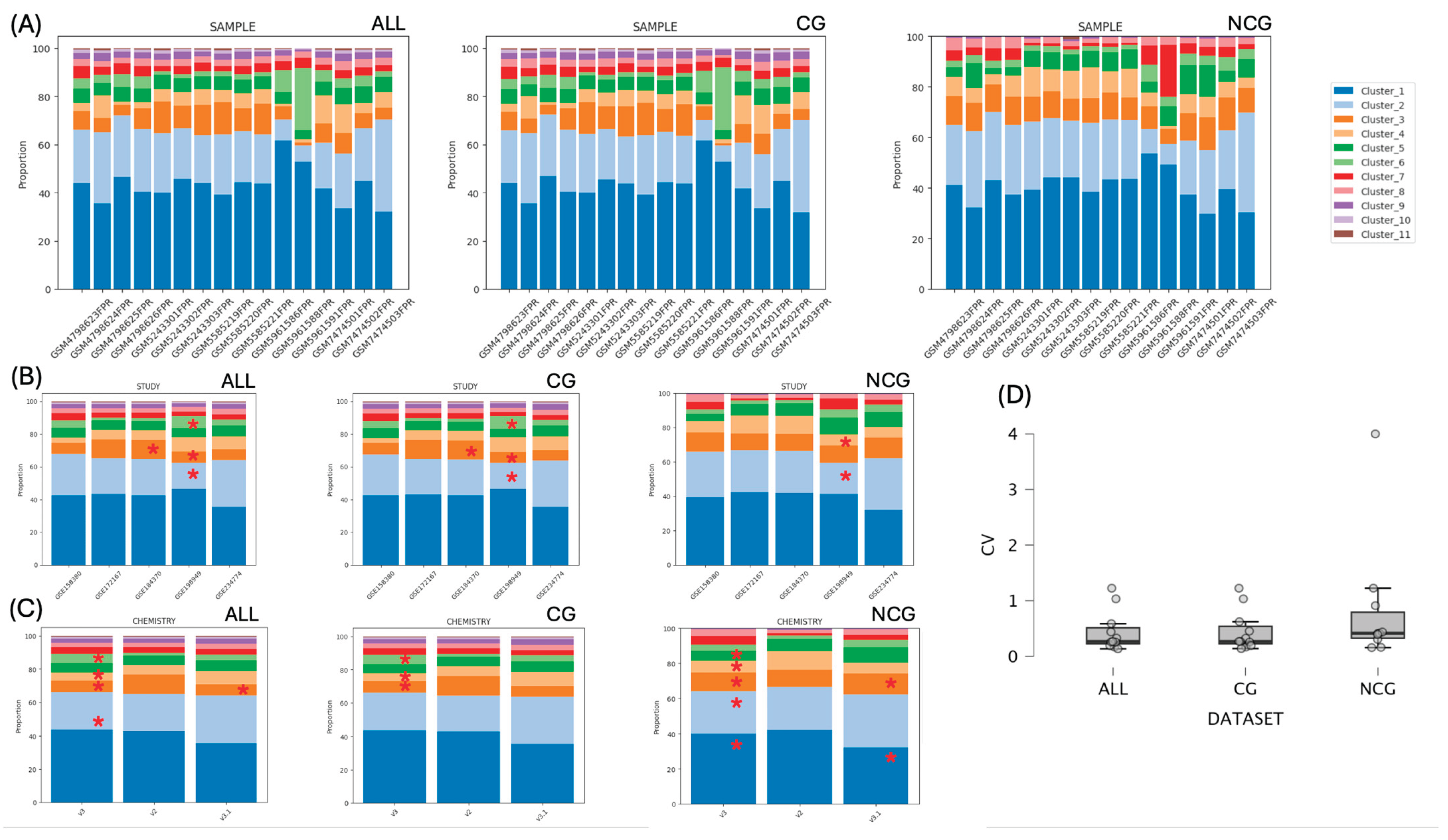
2.7. Compositional Analysis
2.8. Data Analysis
3. Results
3.1. Quality Control
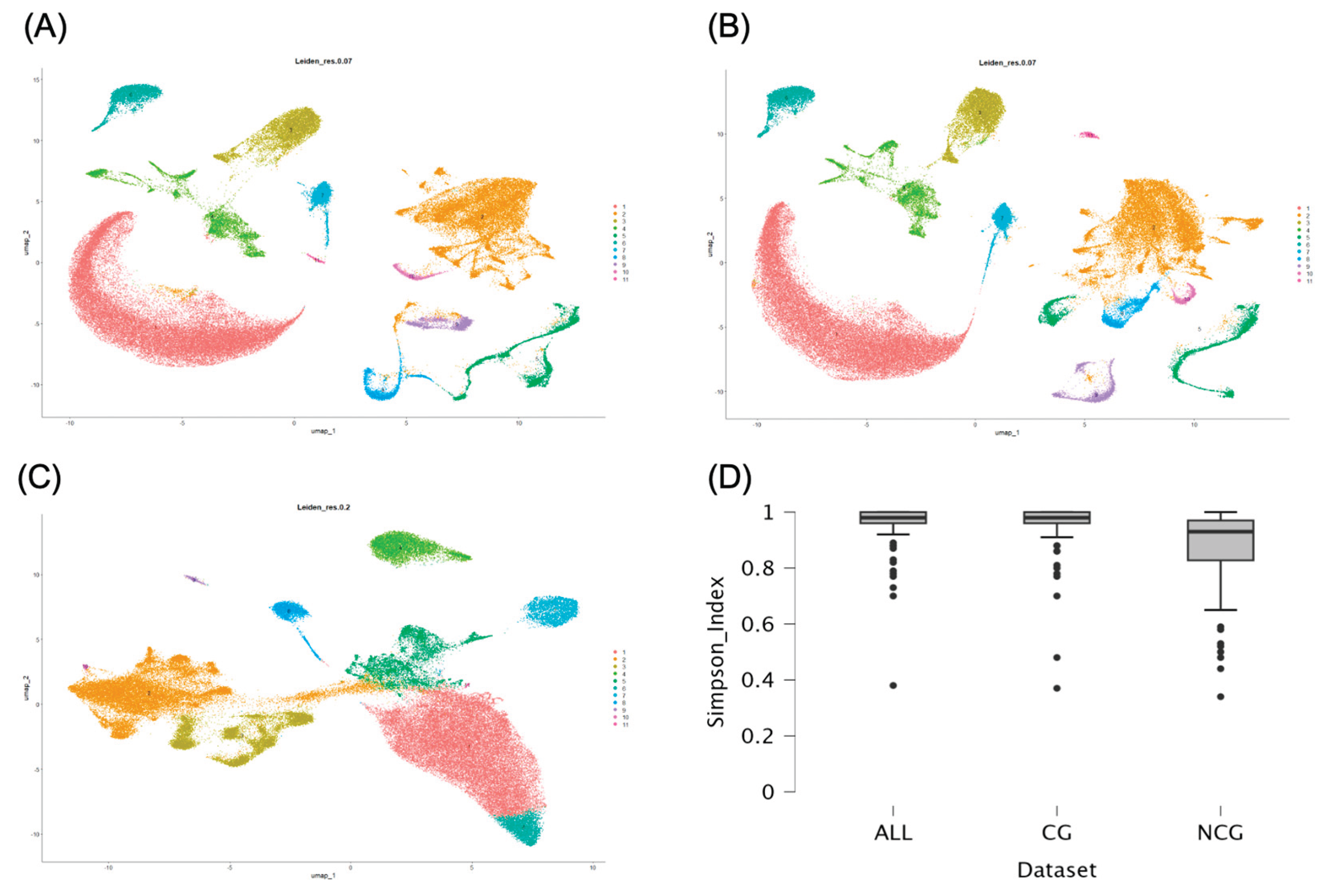
3.2. Clustering of Spinal Cell Types
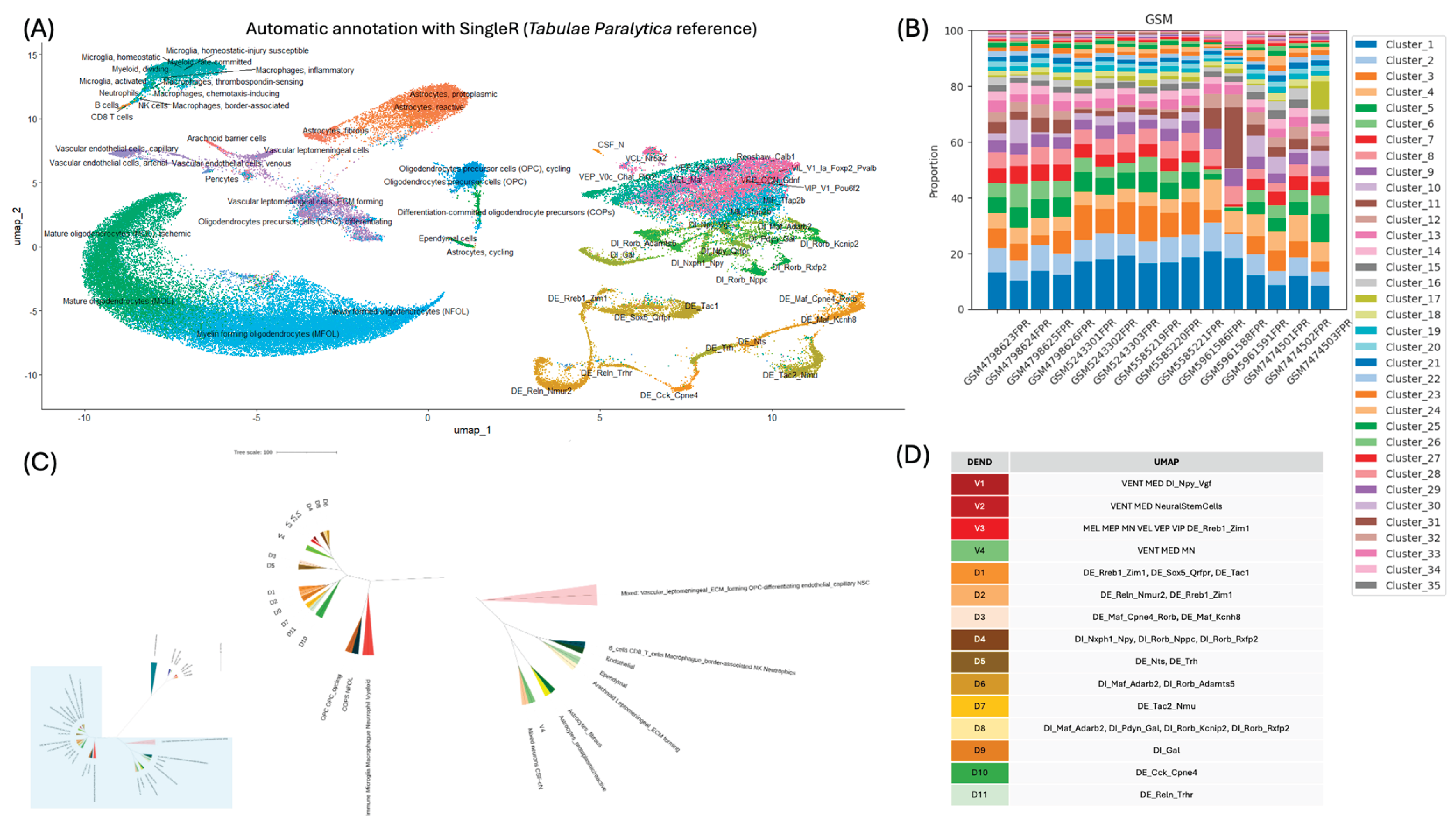
3.3. Annotation of Major Clusters
3.4. Comparison of Clusterings Among ALL, CG, and NCG Datasets
3.5. Cluster Composition is Retained Among Samples
3.6. Fine Clustering and Neuronal Populations
3.7. Non-Coding RNA Markers of Spinal Populations
4. Discussion
Limitations and Caveats
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ALL | Coding + non-coding genes dataset |
| CG | Coding genes dataset |
| CNS | Central nervous system |
| CV | Coefficient of variation |
| DE | Dorsal excitatory neurons |
| DI | Dorsal inhibitory neurons |
| GNR | Glial neuronal ratio |
| IF | Isotropic Fractionator |
| k-NN | k-nearest neighbor |
| lincRNAs | Long intervening non-coding RNAs |
| lncRNAs | Long non-coding RNAs |
| NCG | Non-coding genes dataset |
| ncRNAs | Non-coding RNAs |
| nNNR | Non-neuronal to neuronal ratio |
| OPCs | Oligodendrocyte precursor cells |
| PC | Principal Component |
| PCA | Principal Component Analysis |
| QC | Quality control |
| SCI | Spinal cord injury |
| snRNA-seq | Single-nucleus RNA sequencing |
| snT | Single nucleus transcriptomics |
| STE | Stereology |
| UMAP | Uniform Manifold Approximation and Projection |
| UMI | Unique molecular identifier |
References
- Andrews, S. FastQC: A quality control tool for high throughput sequence data [Software]. 2010. https://www.bioinformatics.babraham.ac.uk/projects/fastqc/.
- Aran, D.; Looney, A.P.; Liu, L.; Wu, E.; Fong, V.; Hsu, A.; Chak, S.; Naikawadi, R.P.; Wolters, P.J.; Abate, A.R.; Butte, A.J.; Bhattacharya, M. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat Immunol. 2019, 20, 163–172. [Google Scholar] [CrossRef] [PubMed]
- Bjugn, R. The use of the optical disector to estimate the number of neurons, glial and endothelial cells in the spinal cord of the mouse--with a comparative note on the rat spinal cord. Brain Res. 1993, 627, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Büttner, M.; Ostner, J.; Müller, C.L.; Theis, F.J.; Schubert, B. scCODA is a Bayesian model for compositional single-cell data analysis. Nat Commun. 2021, 12, 6876. [Google Scholar] [CrossRef] [PubMed]
- DeJong, T.M. A comparison of three diversity indices based on their components of richness and evenness. Oikos 1975, 222–227. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Dobrott, C.I.; Sathyamurthy, A.; Levine, A.J. Decoding Cell Type Diversity Within the Spinal Cord. Curr Opin Physiol. 2019, 8, 1–6. [Google Scholar] [CrossRef]
- Fleming, S.J.; Chaffin, M.D.; Arduini, A.; Akkad, A.D.; Banks, E.; Marioni, J.C.; Philippakis, A.A.; Ellinor, P.T.; Babadi, M. Unsupervised removal of systematic background noise from droplet-based single-cell experiments using CellBender. Nat Methods. 2023, 20, 1323–1335. [Google Scholar] [CrossRef]
- Fu, Y.; Rusznák, Z.; Herculano-Houzel, S.; Watson, C.; Paxinos, G. Cellular composition characterizing postnatal development and maturation of the mouse brain and spinal cord. Brain Struct Funct. 2013, 218, 1337–54. [Google Scholar] [CrossRef]
- Fu, Y.; Yu, Y.; Paxinos, G.; Watson, C.; Rusznák, Z. Aging-dependent changes in the cellular composition of the mouse brain and spinal cord. Neuroscience 2015, 290, 406–20. [Google Scholar] [CrossRef]
- Germain, P.L.; Lun, A.; Garcia Meixide, C.; Macnair, W.; Robinson, M.D. Doublet identification in single-cell sequencing data using scDblFinder. F1000Res. 2022, 10, 979. [Google Scholar] [CrossRef]
- Hao, Y.; Stuart, T.; Kowalski, M.H.; Choudhary, S.; Hoffman, P.; Hartman, A.; Srivastava, A.; Molla, G.; Madad, S.; Fernandez-Granda, C.; Satija, R. Dictionary learning for integrative, multimodal and scalable single-cell analysis. Nat Biotechnol. 2024, 42, 293–304. [Google Scholar] [CrossRef]
- JASP team. JASP (Version 0.19.3) Computer software. 2024.
- Kaminow, B.; Yusunov, D.; Dobin, A. STARsolo: accurate, fast and versatile mapping/quantification of single-cell and single-nucleus RNA-seq data. bioRxiv 2024. [CrossRef]
- Kathe, C.; Skinnider, M.A.; Hutson, T.H.; Regazzi, N.; Gautier, M.; Demesmaeker, R.; Komi, S.; Ceto, S.; James, N.D.; Cho, N.; Baud, L.; Galan, K.; Matson, K.J.E.; Rowald, A.; Kim, K.; Wang, R.; Minassian, K.; Prior, J.O.; Asboth, L.; Barraud, Q.; Lacour, S.P.; Levine, A.J.; Wagner, F.; Bloch, J.; Squair, J.W.; Courtine, G. The neurons that restore walking after paralysis. Nature 2022, 611, 540–547. [Google Scholar] [CrossRef]
- Korsunsky, I.; Millard, N.; Fan, J.; Slowikowski, K.; Zhang, F.; Wei, K.; Baglaenko, Y.; Brenner, M.; Loh, P.R.; Raychaudhuri, S. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat Methods. 2019, 16, 1289–1296. [Google Scholar] [CrossRef]
- Matson, K.J.E.; Russ, D.E.; Kathe, C.; Hua, I.; Maric, D.; Ding, Y.; Krynitsky, J.; Pursley, R.; Sathyamurthy, A.; Squair, J.W.; Levi, B.P.; Courtine, G.; Levine, A.J. Single cell atlas of spinal cord injury in mice reveals a pro-regenerative signature in spinocerebellar neurons. Nat Commun. 2022, 13, 5628. [Google Scholar] [CrossRef] [PubMed]
- Open, AI. ChatGPT. Large language model. 2025. https://chat.openai.com/.
- R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. 2024. https://www.R-project.org/.
- Russ, D.E.; Cross, R.B.P.; Li, L.; Koch, S.C.; Matson, K.J.E.; Yadav, A.; Alkaslasi, M.R.; Lee, D.I.; Le Pichon, C.E.; Menon, V.; Levine, A.J. Author Correction: A harmonized atlas of mouse spinal cord cell types and their spatial organization. Nat Commun. 2021, 13, 1033–10.1038, Erratum for: Nat Commun. 2021, 12, 5722.. [Google Scholar] [CrossRef] [PubMed]
- Sathyamurthy, A.; Johnson, K.R.; Matson, K.J.E.; Dobrott, C.I.; Li, L.; Ryba, A.R.; Bergman, T.B.; Kelly, M.C.; Kelley, M.W.; Levine, A.J. Massively Parallel Single Nucleus Transcriptional Profiling Defines Spinal Cord Neurons and Their Activity during Behavior. Cell Rep. 2018, 22, 2216–2225. [Google Scholar] [CrossRef] [PubMed]
- Skinnider, M.A.; Gautier, M.; Teo, A.Y.Y.; Kathe, C.; Hutson, T.H.; Laskaratos, A.; de Coucy, A.; Regazzi, N.; Aureli, V.; James, N.D.; Schneider, B.; Sofroniew, M.V.; Barraud, Q.; Bloch, J.; Anderson, M.A.; Squair, J.W.; Courtine, G. Single-cell and spatial atlases of spinal cord injury in the Tabulae Paralytica. Nature 2024, 631, 150–163. [Google Scholar] [CrossRef]
- Squair, J.W.; Gautier, M.; Kathe, C.; Anderson, M.A.; James, N.D.; Hutson, T.H.; Hudelle, R.; Qaiser, T.; Matson, K.J.E.; Barraud, Q.; Levine, A.J.; La Manno, G.; Skinnider, M.A.; Courtine, G. Confronting false discoveries in single-cell differential expression. Nat Commun. 2021, 12, 5692. [Google Scholar] [CrossRef]
- Squair, J.W.; Milano, M.; de Coucy, A.; Gautier, M.; Skinnider, M.A.; James, N.D.; Cho, N.; Lasne, A.; Kathe, C.; Hutson, T.H.; Ceto, S.; Baud, L.; Galan, K.; Aureli, V.; Laskaratos, A.; Barraud, Q.; Deming, T.J.; Kohman, R.E.; Schneider, B.L.; He, Z.; Bloch, J.; Sofroniew, M.V.; Courtine, G.; Anderson, M.A. Recovery of walking after paralysis by regenerating characterized neurons to their natural target region. Science 2023, 381, 1338–1345. [Google Scholar] [CrossRef]
- Traag, V.A.; Waltman, L.; van Eck, N.J. From Louvain to Leiden: guaranteeing well-connected communities. Sci Rep. 2019, 9, 5233. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Wen, Z.J.; Xu, H.M.; Zhang, Y.; Zhang, Y.F. Exosomal noncoding RNAs in central nervous system diseases: biological functions and potential clinical applications. Front Mol Neurosci. 2022, 15, 1004221. [Google Scholar] [CrossRef]
- Zappia, L.; Oshlack, A. Clustering trees: a visualization for evaluating clusterings at multiple resolutions. Gigascience 2018, 7, giy083. [Google Scholar] [CrossRef]
- Zeisel, A.; Hochgerner, H.; Lönnerberg, P.; Johnsson, A.; Memic, F.; van der Zwan, J.; Häring, M.; Braun, E.; Borm, L.E.; La Manno, G.; Codeluppi, S.; Furlan, A.; Lee, K.; Skene, N.; Harris, K.D.; Hjerling-Leffler, J.; Arenas, E.; Ernfors, P.; Marklund, U.; Linnarsson, S. Molecular Architecture of the Mouse Nervous System. Cell 2018, 174, 999–1014.e22. [Google Scholar] [CrossRef]

| GEO series ID |
GEO samples ID (number of runs in SRA) |
10X Genomics Chromium version |
Reads (M) |
Sex |
Age (weeks) |
Publication |
| GSE158380 | GSM4798623 (6) GSM4798624 (6) GSM4798625 (6) GSM4798626 (6) |
Single Cell 3’ Kit Version 3 | 172.4 166.0 174.2 178.7 |
M M F F |
9 | Russ et al. (2022) |
| GSE165003 | GSM5024317 (8)* GSM5024318 (8)* GSM5024319 (8)* |
Single Cell Kit Version 2 | 267.2 282.8 274.5 |
F F F |
12-30 | Squair et al. (2021) |
| GSE198949 | GSM5961586 (1) GSM5961588 (2) GSM5961591 (8) |
Single Cell 3’ Kit Version 3 | 84.1 100.6 903.2 |
F F F |
8-15 | Squair et al. (2023) |
| GSE172167 | GSM5243301 (15) GSM5243302 (15) GSM5243303 (15) |
Single Cell Kit Version 2 | 505.6 593.9 648 |
F F F |
12-30 | Matson et al. (2022) |
| GSE184370 | GSM5585219 (8) GSM5585220 (8) GSM5585221 (8) |
Single Cell Kit Version 2 | 267.2 282.8 274.5 |
F F F |
12-30 | Kathe et al. (2022) |
| GSE234774 | GSM7474501 (8) GSM7474502 (8) GSM7474503 (8) |
Single Cell Kit Version 3.1 | 698.2 465 480.2 |
F F F |
8 | Skinnider et al. (2024) |
| Lineage | Coding | Non-coding | Cod + Non Cod |
| Oligodendrocytes | 1 (34,836) 98.9% | 2 (32,302/3,021) 97.6% | 1 (34,834) 98.9% |
| Oligodendrocyte precursor cells | 1 (2,968) 89.2% | 1 (2,697) 98.2% | 1 (2,986) 88.7% |
| Astrocytes | 1 (6,635) 93.9% | 1 (6,313) 98.7% | 1 (6,728) 92.6% |
| Ependymal | 1 (491) 93.7% | 1 (492) 93.5% | 1 (494) 93.1% |
| Vascular | 1 (5,603) 89.7%* | 2 (6,158/2,893) 89.2% | 1 (5,485) 90.8% |
| Microglia/immune | 1 (3,405) | 1 (3,407) | |
| Neurons | 5 (32,440) 98.7% | 3 (32,464) 98.6% | 5 (32,444) 98.7% |
| VE, VI, DI | 1 (21,168) | 1 (22,903) | 1 (21,396) |
| DE | 3 (5,141/2,601/2,542) | 1 (9,455) | 3 (5,179/2,573/2,307) |
| DI-Gal | 1 (988) | 1 (989) | |
| VE-VI | 1 (106) | ||
| Other | 1 (38**) | ||
| Total clusters | 11 (86,378) | ||
| SOURCE | OLIGO | NEUR | ASTRO | VASC | IMM | OPC |
EPEND |
OTHER | nN/N | GNR | TECH |
| ALL Dataset (Median) | 43.0 | 35.5 | 9.0 | 6.0 | 2.5 | 4.0 | 1.0 | 1.8 | 1.6 | snT | |
| ALL Dataset (Mean) | 41.1 | 36.5 | 8.9 | 6.1 | 3.1 | 3.4 | 0.6 | 1.7 | 1.5 | snT | |
| Fu et al., 2015 | 3.7 | IF | |||||||||
| Fu et al., 2013 – 4w | 3.2 | IF | |||||||||
| Fu et al., 2013 – 40w | 4.1 | IF | |||||||||
| Bjugn, 1993 | 39.3 | 33.6 | 19.1 | 8.0 | 2.0 | 1.2 | STE | ||||
| Zeisel (Russ) | 40.4 | 9.2 | 18.0 | 18.6 | 5.2 | 6.4 | 2.0 | 0.1 | 9.9 | 7.3 | snT |
| Skinnider et al., 2024 | 49.7 | 35.3 | 3.4 | 4.2 | 3.1 | 3.3 | 0.7 | 1.8 | 1.6 | snT | |
| Sathyamurthy (Russ) | 24.8 | 28.0 | 13.6 | 7.9 | 2.4 | 1.6 | 2.4 | 19.3 | 2.6 | 1.5 | snT |
| Sathyamurthy paper | 16.0 | 52.0 | 9.0 | 5.0 | 1.0 | 1.0 | 14.0 | 0.9 | 0.5 | snT |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




