Submitted:
19 July 2025
Posted:
21 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- We utilized advanced data augmentation techniques to tackle the issues of limited and imbalanced datasets, resulting in enhanced model robustness and generalization.
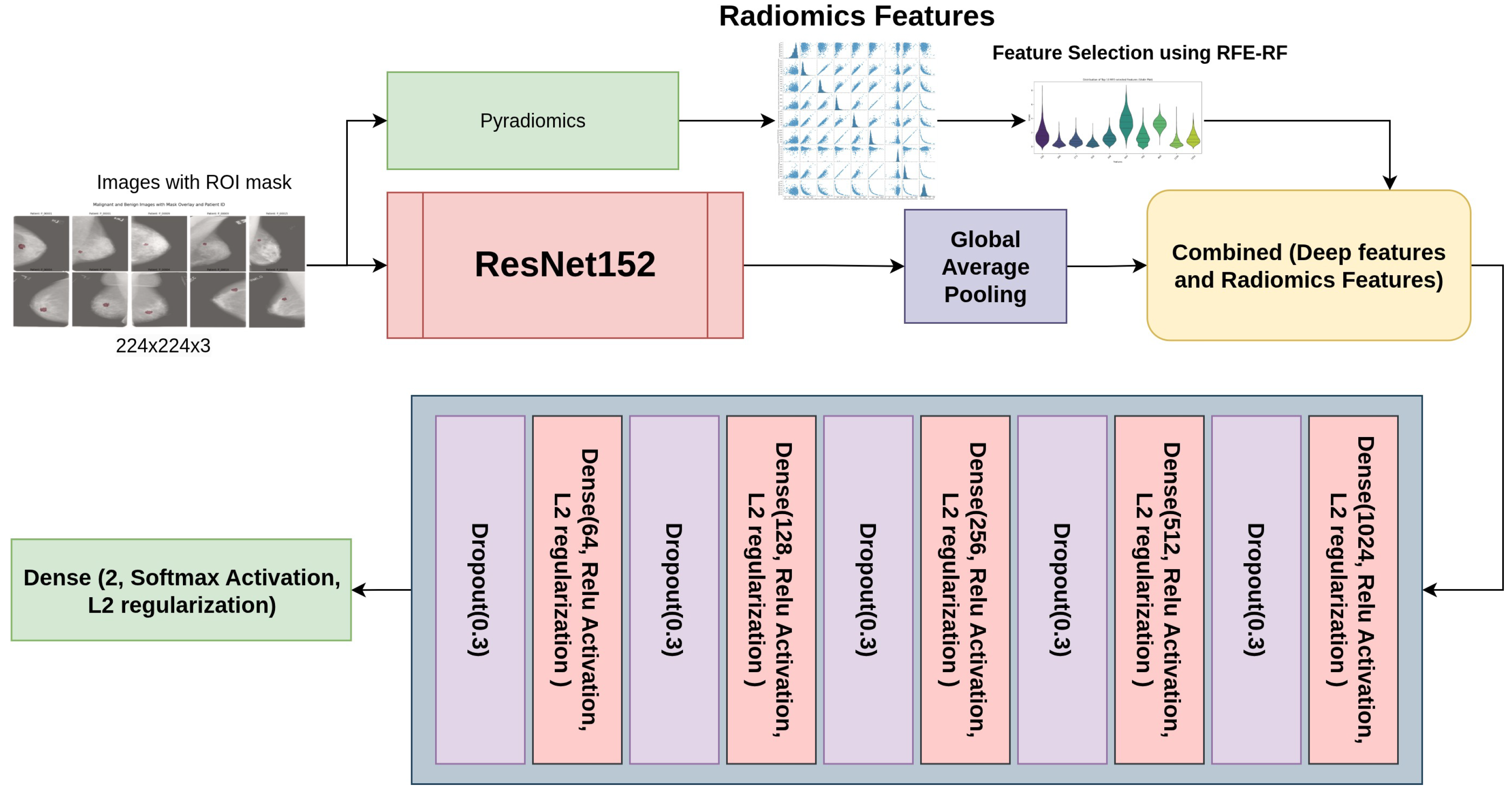
- Perform feature engineering by integrating multimodal features (radiomics and deep features) from the augmented training dataset. Radiomics feature selection was conducted using RFE (with Random Forest and Logistic Regression), ANOVA F-test, LASSO, and mutual information, alongside embedded approaches leveraging XGBoost, LightGBM, and CatBoost with GPU acceleration. The most informative features were subsequently integrated with deep features to build a unified multimodal feature space for final model development.
- Conducted a comprehensive analysis of 13 transfer learning models, highlighting their comparative performance and demonstrating significant improvements in the accuracy of breast cancer detection.
2. Literature Review
2.1. Research Gap
3. Materials and Methods
3.1. Dataset
3.2. Image Pre-Processing and Data Augmentation
| Algorithm 1:Image augmentation pipeline |
|
3.3. Feature Engineering
3.3.1. Radiomics Features
| Algorithm 2:Radiomics Feature Extraction |
|
3.3.2. Radiomics Feature Selection
3.3.3. Deep Learning Features
- = activation value at position in the c-th feature map.
- = resultant scalar for the c-th channel after applying the global average pooling.
3.4. Multimodal Feature Preparation
| Algorithm 3:Multimodal Feature Preparation |
|
3.5. Transfer Learning Models
3.6. Model Evaluation
3.7. Experiment Setup
4. Results
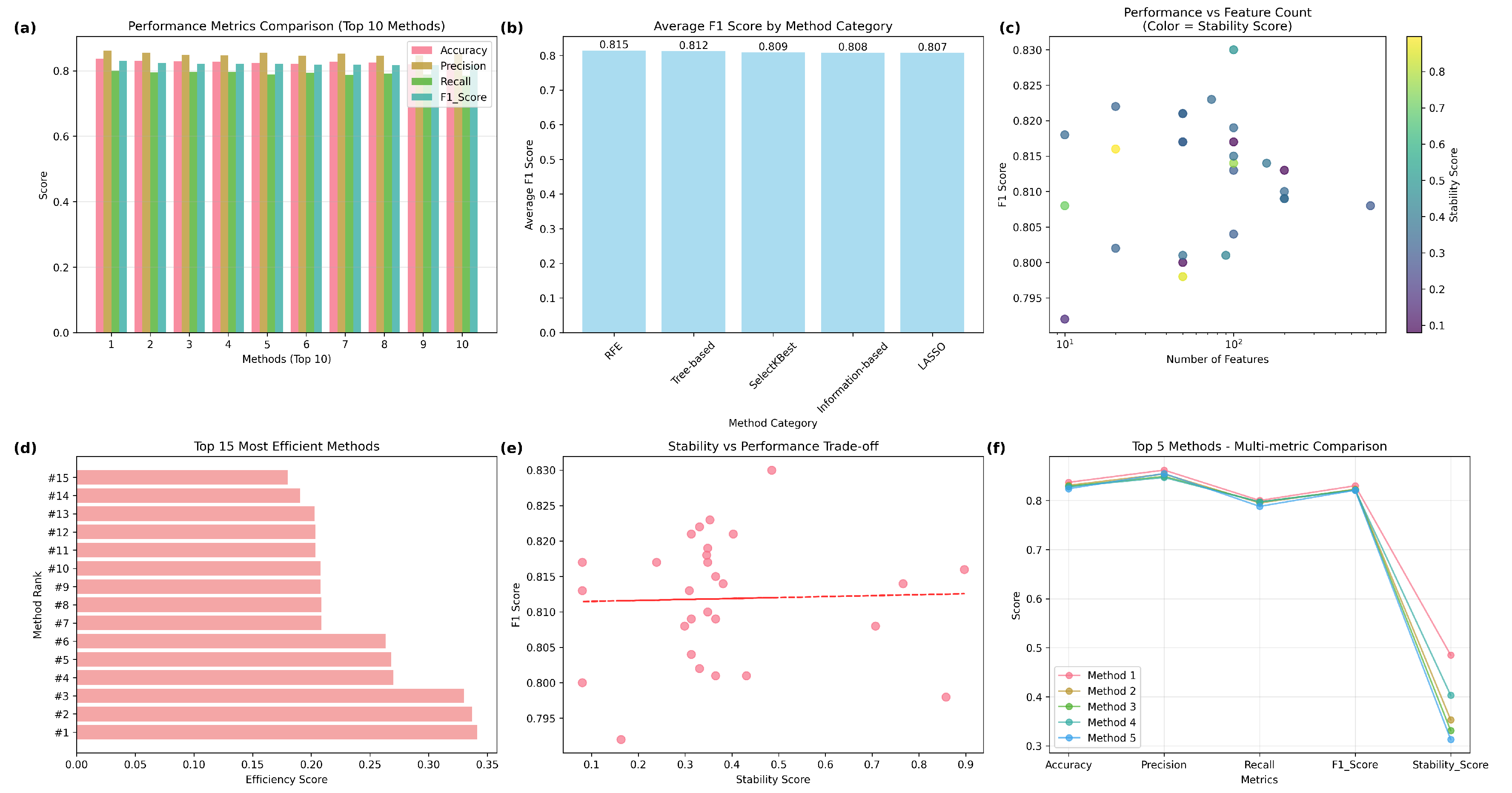
4.1. Radiomics Analysis
Radiomics Feature selection
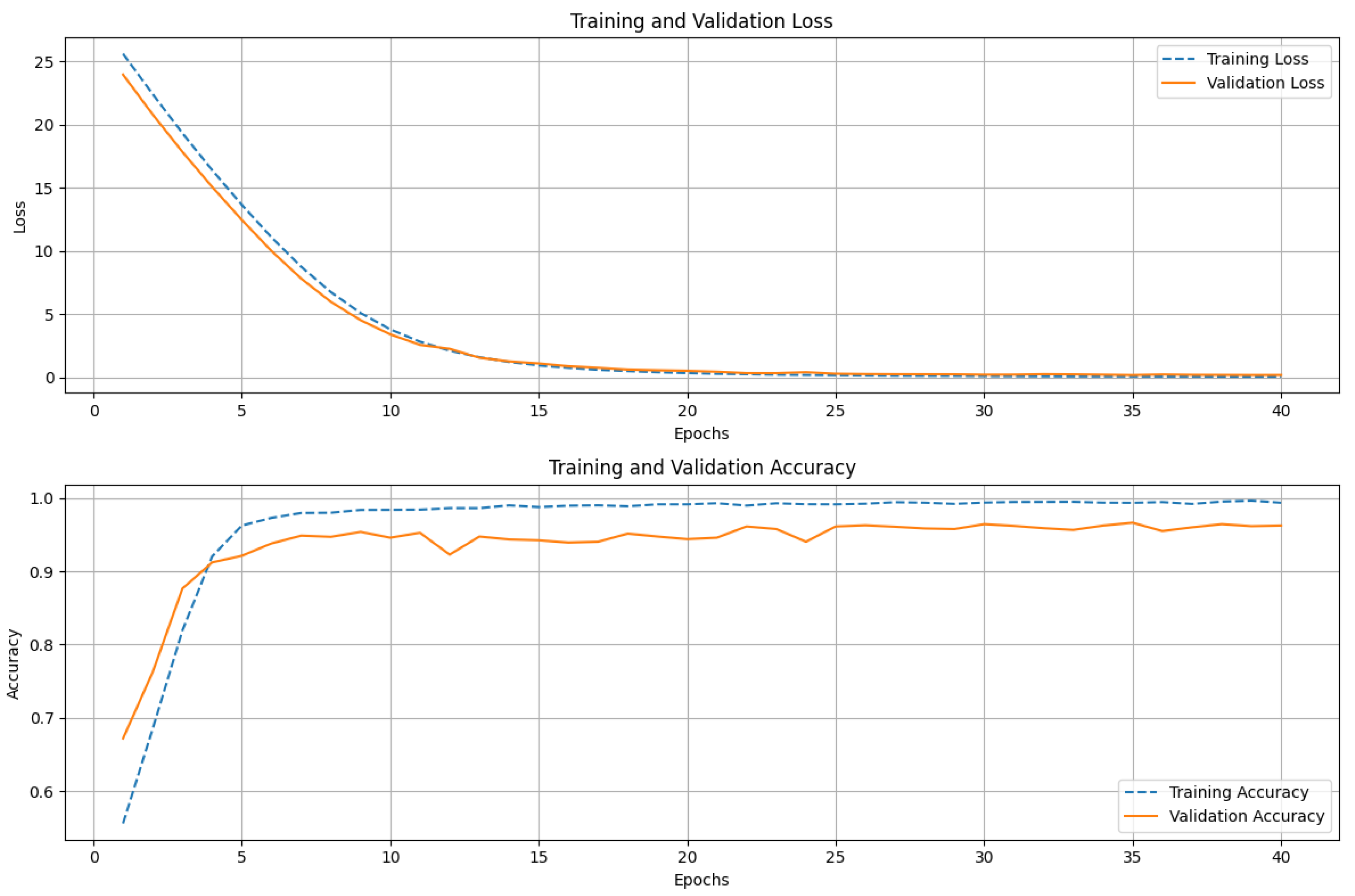
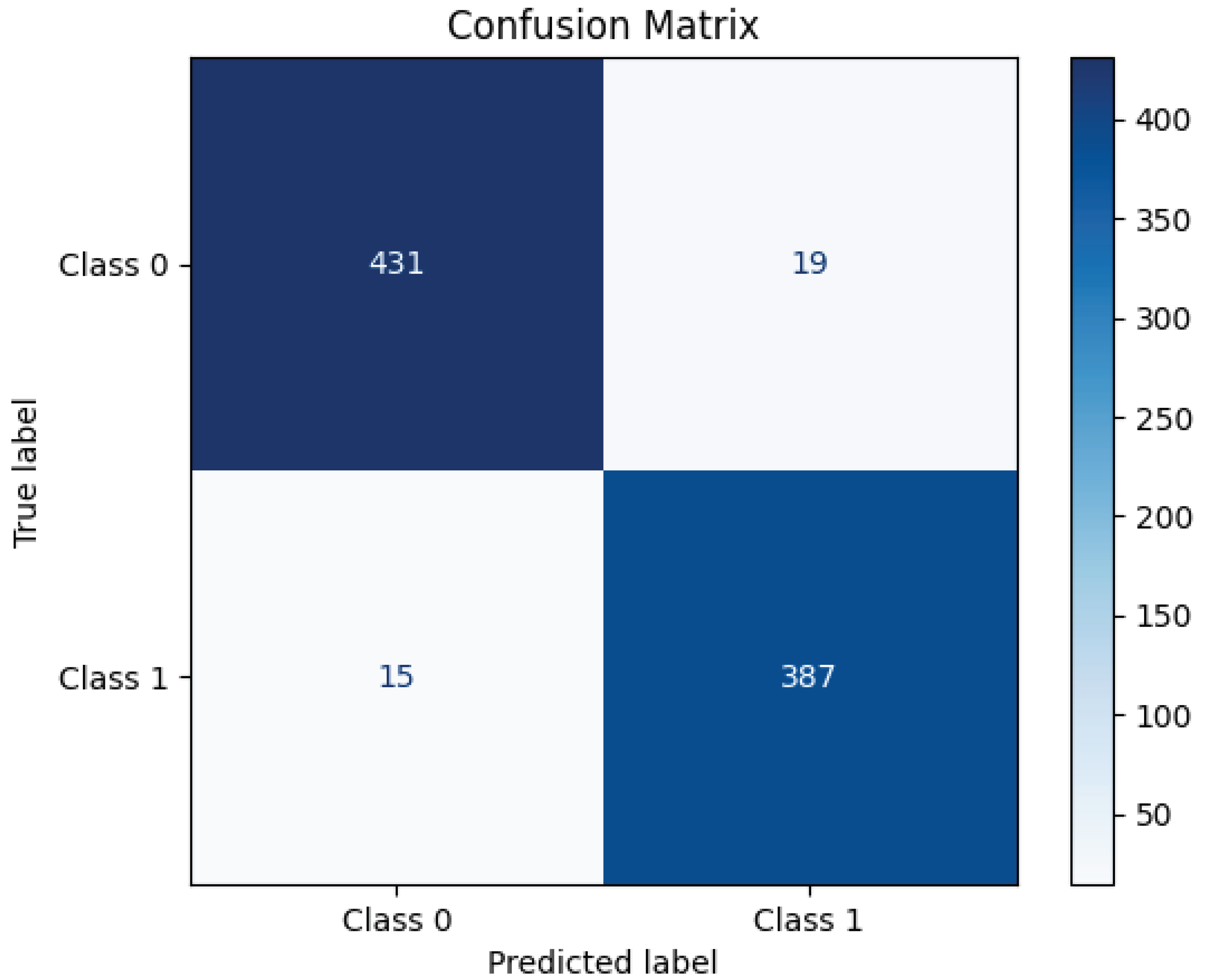
4.2. Models Performance
5. Discussion
6. Conclusion
References
- Arnold, M.; Morgan, E.; Rumgay, H.; Mafra, A.; Singh, D.; Laversanne, M.; Vignat, J.; Gralow, J.R.; Cardoso, F.; Siesling, S.; et al. Current and future burden of breast cancer: Global statistics for 2020 and 2040. Breast (Edinburgh, Scotland) 2022, 66, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Bhagat, J.; Lobo, R.; Kumar, N.; Mathew, J.E.; Pai, A. Cytotoxic potential of Anisochilus carnosus (L.f.) wall and estimation of luteolin content by HPLC 2014.
- Balkenende, L.; Teuwen, J.; Mann, R.M. Application of Deep Learning in Breast Cancer Imaging. Seminars in Nuclear Medicine 2022, 52, 584–596. [Google Scholar] [CrossRef] [PubMed]
- Maruf, N.A.; Basuhail, A. Breast cancer diagnosis using radiomics-guided DL/ML model-systematic review and meta-analysis. Frontiers in Computer Science 2025, 7, 1446270. [Google Scholar] [CrossRef]
- Zheng, D.; He, X.; Jing, J. Overview of Artificial Intelligence in Breast Cancer Medical Imaging. Journal of Clinical Medicine 2023, 12. [Google Scholar] [CrossRef] [PubMed]
- Karthiga, R.; Narasimhan, K. Medical imaging technique using curvelet transform and machine learning for the automated diagnosis of breast cancer from thermal image. Pattern Analysis and Applications 2021, 24, 981–991. [Google Scholar] [CrossRef]
- Kora, P.; Ooi, C.P.; Faust, O.; Raghavendra, U.; Gudigar, A.; Chan, W.Y.; Meenakshi, K.; Swaraja, K.; Plawiak, P.; Rajendra Acharya, U. Transfer learning techniques for medical image analysis: A review. Biocybernetics and Biomedical Engineering 2022, 42, 79–107. [Google Scholar] [CrossRef]
- Oladimeji, O.O.; Ayaz, H.; McLoughlin, I.; Unnikrishnan, S. Mutual information-based radiomic feature selection with SHAP explainability for breast cancer diagnosis. Results in Engineering 2024, 24, 103071. [Google Scholar] [CrossRef]
- Gamal, A.; Sharafeldeen, A.; Alnaghy, E.; Alghandour, R.; Saleh Alghamdi, N.; Ali, K.M.; Shamaa, S.; Aboueleneen, A.; Elsaid Tolba, A.; Elmougy, S.; et al. A Novel Machine Learning Approach for Predicting Neoadjuvant Chemotherapy Response in Breast Cancer: Integration of Multimodal Radiomics With Clinical and Molecular Subtype Markers. IEEE Access 2024, 12, 104983–105003. [Google Scholar] [CrossRef]
- Jiang, Y.; Zeng, Y.; Zuo, Z.; Yang, X.; Liu, H.; Zhou, Y.; Fan, X. Leveraging multimodal MRI-based radiomics analysis with diverse machine learning models to evaluate lymphovascular invasion in clinically node-negative breast cancer. Heliyon 2024, 10, e23916. [Google Scholar] [CrossRef] [PubMed]
- Del Corso, G.; Germanese, D.; Caudai, C.; Anastasi, G.; Belli, P.; Formica, A.; Nicolucci, A.; Palma, S.; Pascali, M.A.; Pieroni, S.; et al. Adaptive Machine Learning Approach for Importance Evaluation of Multimodal Breast Cancer Radiomic Features. Journal of Imaging Informatics in Medicine 2024, 37, 1642–1651. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, A.; Primakov, S.; Beuque, M.; Woodruff, H.C.; Halilaj, I.; Wu, G.; Refaee, T.; Granzier, R.; Widaatalla, Y.; Hustinx, R.; et al. Radiomics for precision medicine: Current challenges, future prospects, and the proposal of a new framework. Methods 2021, 188, 20–29. [Google Scholar] [CrossRef] [PubMed]
- Haghshomar, M.; Rodrigues, D.; Kalyan, A.; Velichko, Y.; Borhani, A. Leveraging radiomics and AI for precision diagnosis and prognostication of liver malignancies. Frontiers in Oncology 2024, Volume 14 - 2024. [CrossRef]
- Khourdifi, Y.; Alami, A.E.; Zaydi, M.; Maleh, Y. Early Breast Cancer Detection Based on Deep Learning: An Ensemble Approach Applied to Mammograms. MDPI 2024.
- Luong, H.H.; Vo, M.D.; Phan, H.P.; Dinh, T.A. Improving Breast Cancer Prediction via Progressive Ensemble and Image Enhancement. Multimedia Tools and Applications 2024.
- Chugh, G.; Kumar, S.; Singh, N. TransNet: A Comparative Study on Breast Carcinoma Diagnosis with Classical Machine Learning and Transfer Learning Paradigm. Multimedia Tools and Applications 2024.
- Subha, T.D.; Charan, G.S.; Reddy, C.C. An Efficient Image-Based Mammogram Classification Framework Using Depth-Wise Convolutional Neural Network. IEEE Conference on Artificial Intelligence 2023.
- Gujar, S.A.; Lei, X.; Cen, S.Y.; Hwang, D.H. Breast Lesion Classification using Radiomics-derived Regions of Interest on Mammograms. IEEE Symposium on Image Processing 2023.
- Alkhaleefah, M.; Tan, T.H.; Chang, C.H.; Wang, T.C.; Ma, S.C. Connected-segNets: A Deep Learning Model for Breast Tumor Segmentation from X-ray Images. Cancers 2022.
- Lee, R.S.; Gimenez, F.; Hoogi, A.; Miyake, K.K.; Gorovoy, M.; Rubin, D.L. A curated mammography data set for use in computer-aided detection and diagnosis research. Scientific Data 2017 4:1 2017, 4, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Marufaiub. CBIS-DDSM-Radiomics-And-13-Transfer-Learning-Models, 2024. Accessed: 2025-02-02.
- Van Griethuysen, J.J.; Fedorov, A.; Parmar, C.; Hosny, A.; Aucoin, N.; Narayan, V.; Beets-Tan, R.G.; Fillion-Robin, J.C.; Pieper, S.; Aerts, H.J. Computational radiomics system to decode the radiographic phenotype. Cancer Research 2017, 77, e104–e107. [Google Scholar] [CrossRef] [PubMed]
- Yu, F.H.; Miao, S.M.; Li, C.Y.; Hang, J.; Deng, J.; Ye, X.H.; Liu, Y. Pretreatment ultrasound-based deep learning radiomics model for the early prediction of pathologic response to neoadjuvant chemotherapy in breast cancer. European Radiology 2023, 33, 5634–5644. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Zhong, X.; Li, W.; Li, Q.; Shao, H.; Wang, Z.; Dai, Y.; Ma, H.; Shi, Y.; Zhang, H.; et al. Attention-based Deep Learning for the Preoperative Differentiation of Axillary Lymph Node Metastasis in Breast Cancer on DCE-MRI. Journal of Magnetic Resonance Imaging 2023, 57, 1842–1853. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Ma, Q.; Feng, H.; Wei, T.; Jiang, F.; Fan, L.; Zhang, W.; Xu, J.; Zhang, X. Deep learning radiomics for prediction of axillary lymph node metastasis in patients with clinical stage T1–2 breast cancer. Quantitative Imaging in Medicine and Surgery 2023, 13, 4995–5011. [Google Scholar] [CrossRef] [PubMed]
- Sharmin, S.; Ahammad, T.; Talukder, M.A.; Ghose, P. A Hybrid Dependable Deep Feature Extraction and Ensemble-Based Machine Learning Approach for Breast Cancer Detection. IEEE Access 2023, 11, 87694–87708. [Google Scholar] [CrossRef]
- Yang, X.; Fan, X.; Lin, S.; Zhou, Y.; Liu, H.; Wang, X.; Zuo, Z.; Zeng, Y. Assessment of Lymphovascular Invasion in Breast Cancer Using a Combined <scp>MRI</scp> Morphological Features, Radiomics, and Deep Learning Approach Based on Dynamic Contrast-Enhanced <scp>MRI</scp>. Journal of Magnetic Resonance Imaging 2023. [Google Scholar] [CrossRef]
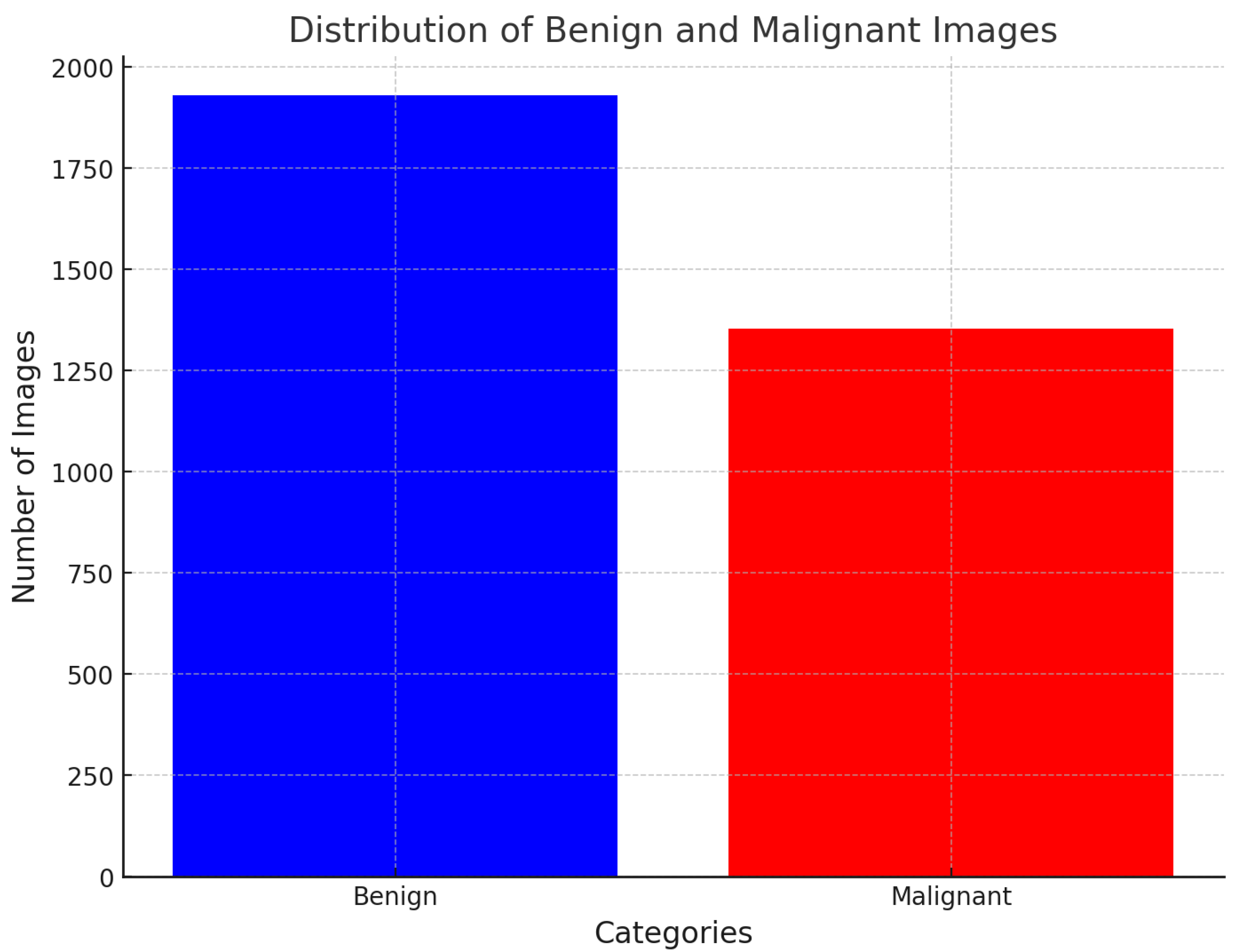
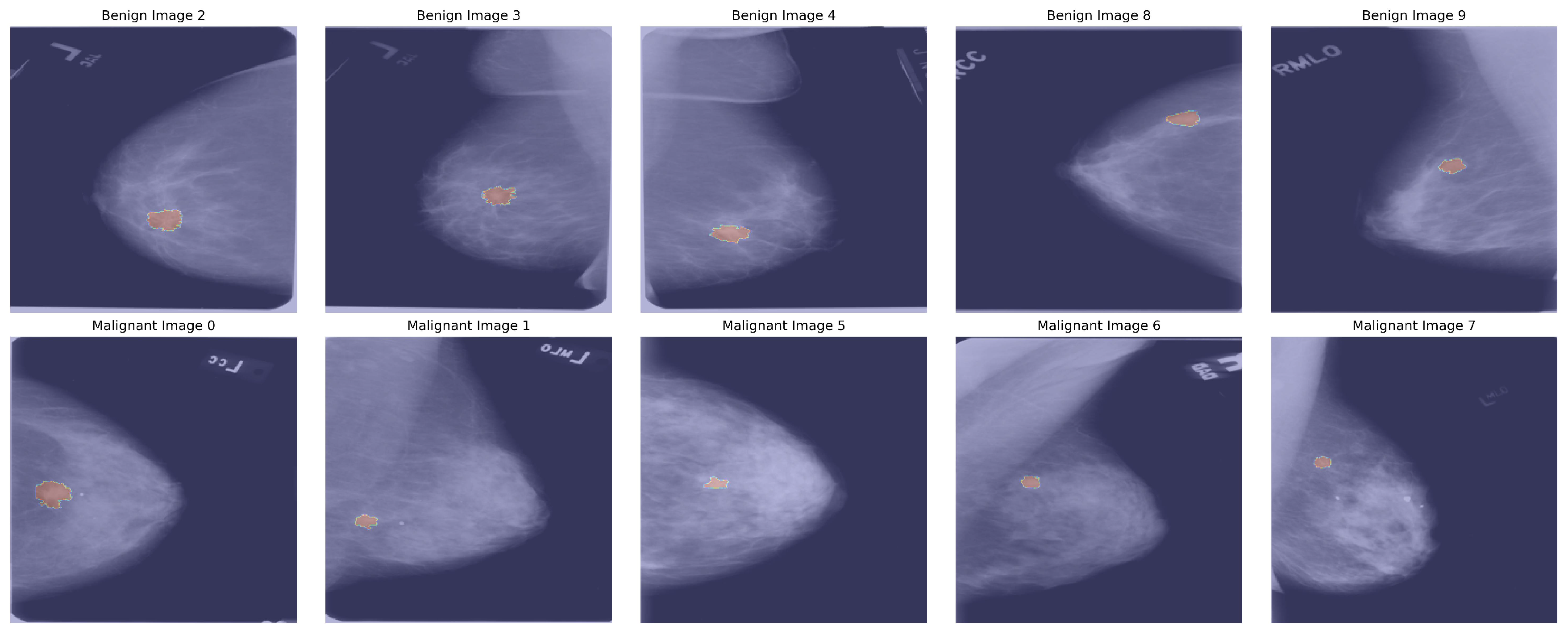
| Labels | Original Images | Augmented Images |
|---|---|---|
| Benign | 1930 | 8498 |
| Malignant | 1354 | 8498 |
| Total | 3284 | 16996 |
| Method | Num_Features |
|---|---|
| RFE (Random Forest) | 10, 20, 50, 100 |
| RFE (Logistic Regression) | 10, 20, 50, 100 |
| RFECV (Random Forest) | 74 |
| RFECV (Logistic Regression) | 647 |
| SelectKBest (ANOVA) | 10, 20, 50, 100 |
| LASSO (LassoCV) | 90 |
| LASSO (LassoCV) | 157 |
| XGBoost GPU | 50, 100, 200 |
| LightGBM GPU | 50, 100, 200 |
| CatBoost GPU | 50, 100, 200 |
| Mutual Information | 50, 100, 200 |
| Category | Number of Features |
|---|---|
| GLCM | 264 |
| GLSZM | 176 |
| GLRLM | 176 |
| First-order | 198 |
| GLDM | 154 |
| NGTDM | 55 |
| Shape | 9 |
| Diagnostics | 5 |
| Other | 3 |
| Method | Features | Ac | Pre | Rec | F1s | St |
|---|---|---|---|---|---|---|
| RFE (RF) | 100 | 0.837 | 0.862 | 0.800 | 0.830 | 0.485 |
| RFECV (RF) | 74 | 0.831 | 0.854 | 0.795 | 0.823 | 0.353 |
| RFE (RF) | 20 | 0.829 | 0.849 | 0.797 | 0.822 | 0.331 |
| RFE (Random Forest) | 50 | 0.828 | 0.847 | 0.797 | 0.821 | 0.403 |
| RFE (Random Forest) | 10 | 0.827 | 0.852 | 0.787 | 0.818 | 0.346 |
| RFE (LR) | 50 | 0.825 | 0.846 | 0.791 | 0.817 | 0.239 |
| CatBoost GPU | 50 | 0.824 | 0.855 | 0.788 | 0.821 | 0.313 |
| SelectKBest (ANOVA) | 100 | 0.824 | 0.856 | 0.775 | 0.814 | 0.766 |
| SelectKBest (ANOVA) | 20 | 0.822 | 0.835 | 0.798 | 0.816 | 0.897 |
| LightGBM GPU | 100 | 0.822 | 0.846 | 0.794 | 0.819 | 0.348 |
| Model | Prec | Reca | F1s | Ac | Ep |
|---|---|---|---|---|---|
| DenseNet169 | 0.94 | 0.94 | 0.95 | 0.95 | 50 |
| DenseNet201 | 0.93 | 0.94 | 0.94 | 0.94 | 50 |
| DenseNet121 | 0.93 | 0.94 | 0.94 | 0.93 | 50 |
| ResNet152 | 0.97 | 0.98 | 0.97 | 0.97 | 40 |
| ResNet101V2 | 0.96 | 0.96 | 0.96 | 0.96 | 45 |
| ResNet101 | 0.96 | 0.96 | 0.96 | 0.96 | 48 |
| ResNet50 | 0.94 | 0.94 | 0.94 | 0.94 | 50 |
| ResNet152V2 | 0.94 | 0.94 | 0.94 | 0.94 | 50 |
| ResNet50V2 | 0.93 | 0.93 | 0.93 | 0.93 | 50 |
| InceptionV3 | 0.89 | 0.88 | 0.90 | 0.89 | 50 |
| MobileNet | 0.87 | 0.86 | 0.88 | 0.88 | 50 |
| VGG19 | 0.96 | 0.96 | 0.96 | 0.96 | 50 |
| VGG16 | 0.94 | 0.94 | 0.94 | 0.94 | 29 |
| Author(s) | Methods | Accuracy (%) |
|---|---|---|
| Yu et al. [23] | ResNet34 | 72 |
| ResNet50 | 82 | |
| VGG16 | 71 | |
| Gao et al. [24] | ResNet | 82 |
| Wei et al. [25] | ResNet50 | 72 |
| ResNet101 | 76 | |
| VGG19 | 83 | |
| Inception_v3 | 72 | |
| Sharmin et al. [26] | ResNet50V2 | 95 |
| Yang et al. [27] | 3DResNet | 74 |
| Our Study | ResNet50 | 94 |
| ResNet50V2 | 93 | |
| ResNet101 | 96 | |
| ResNet101V2 | 96 | |
| ResNet152 | 97 | |
| ResNet152V2 | 94 | |
| VGG19 | 96 | |
| VGG16 | 94 | |
| Inception_v3 | 89 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




