Submitted:
14 July 2025
Posted:
16 July 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Relevant Work
2.1. Machine Learning Techniques
2.2. Deep Learning Models for Binary Classification
2.3. Transfer Learning and Model Optimization
2.4. Deep Learning Models for Multi-Class Classification
3. Dataset and Preprocessing
3.1. BreaKHis Dataset
3.2. Preprocessing and Augmentation
3.2.1. Image Resizing
3.2.2. Normalization
3.3. Data Splitting Strategy
3.4. Data Augmentation
3.4.1. Random Horizontal Flipping
3.4.2. Random Rotation ()
3.5. Handling Class Imbalance
4. Methodology
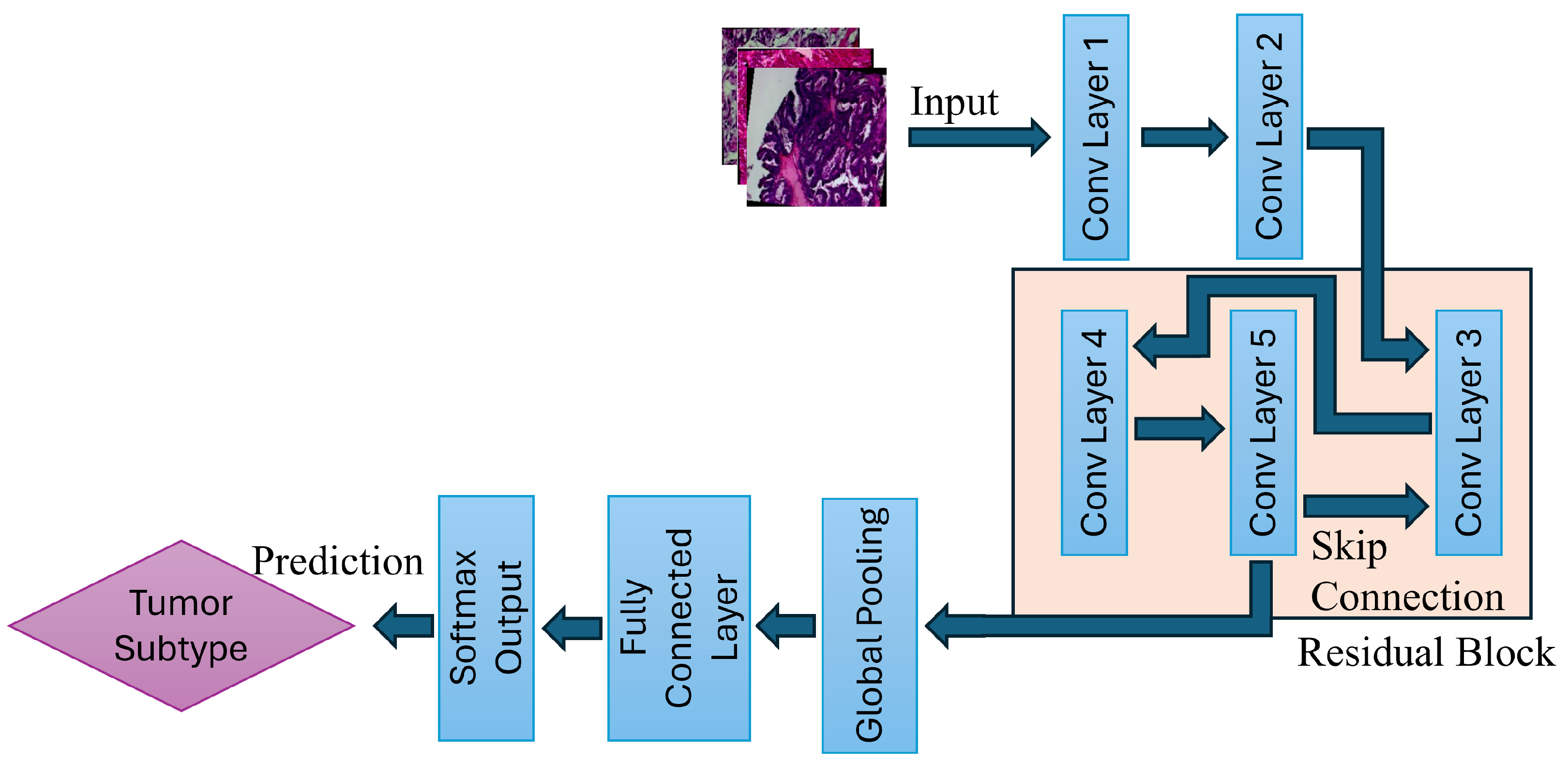
4.1. Overview of the Proposed Model
4.2. Deep Residual Networks (ResNet) for Tumor Classification
4.2.1. ResNet Architecture
4.3. Model Training Process
4.3.1. Forward Propagation
- 1.
- Input Image Processing:
- 2.
- Convolutional Feature Extraction:
- 3.
- Residual Learning via Skip Connections:
5. Results and Analysis
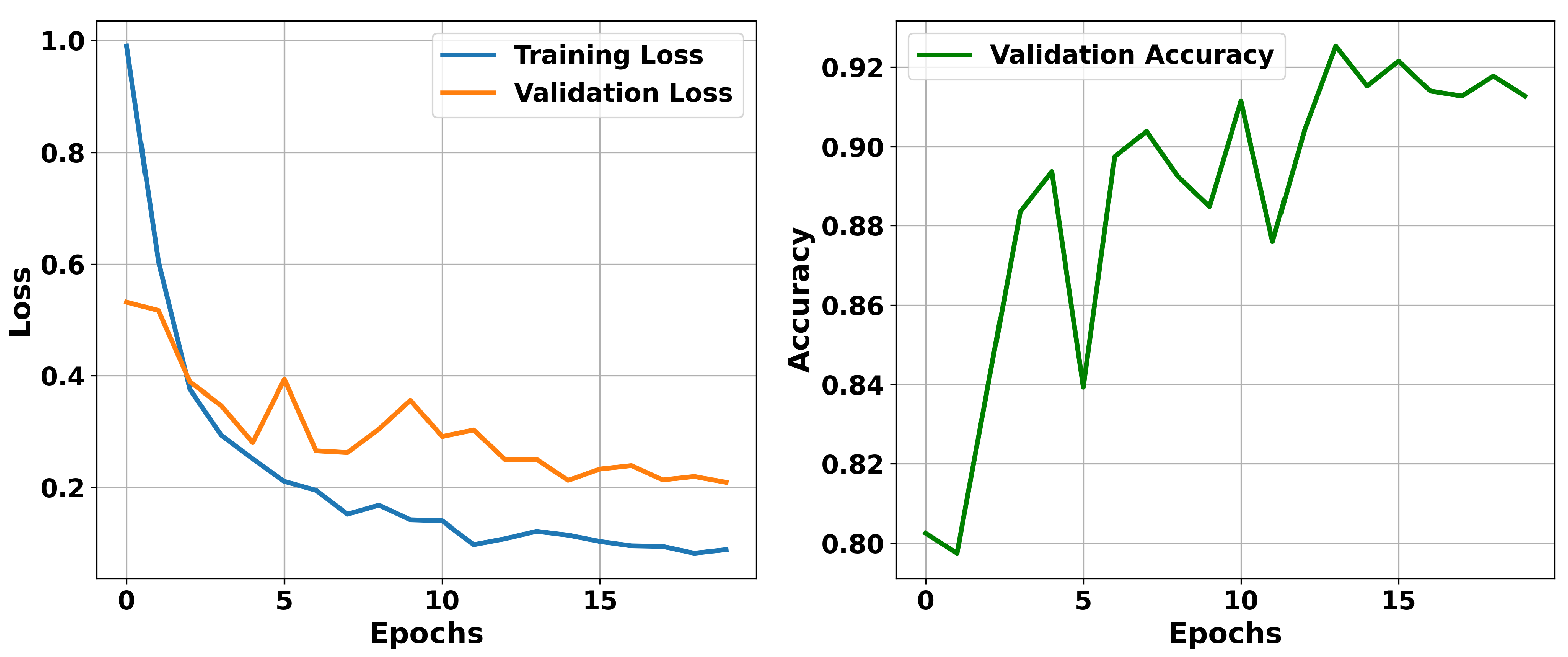
5.1. Model Training Dynamics and Convergence
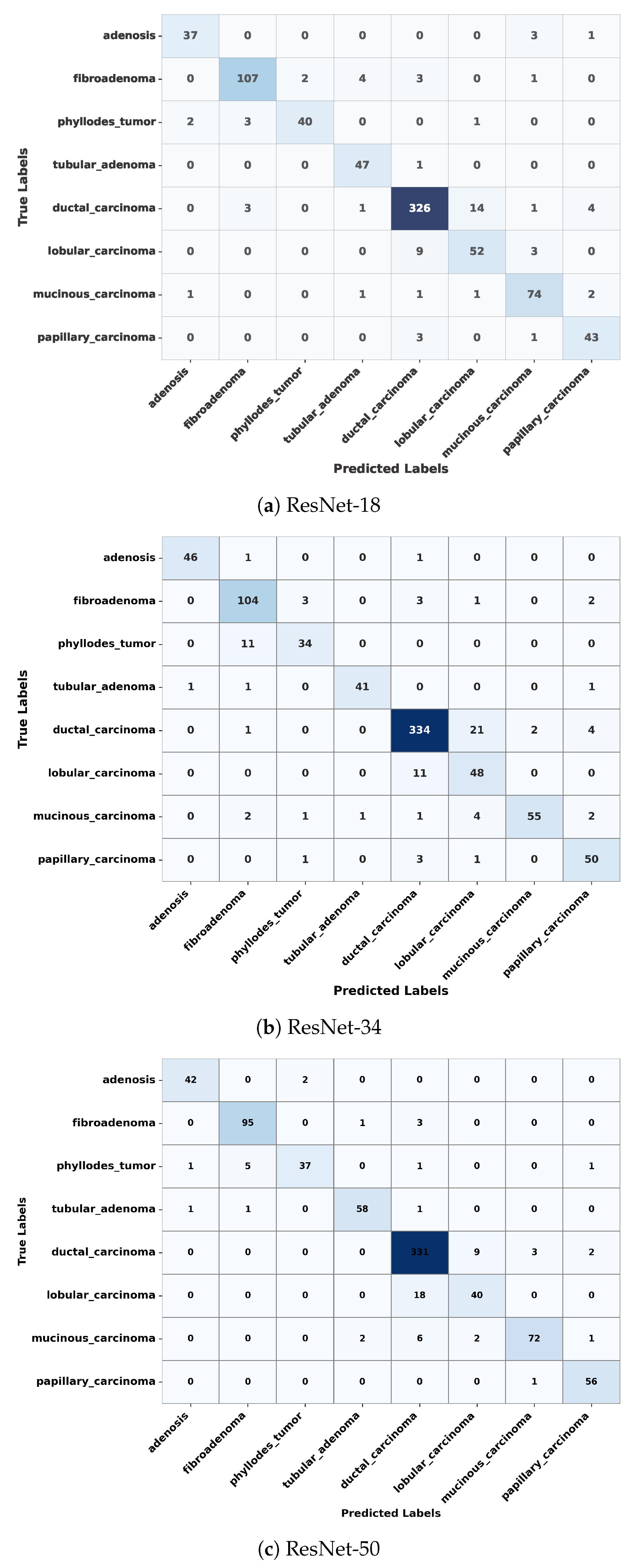
5.2. Confusion Matrix Analysis
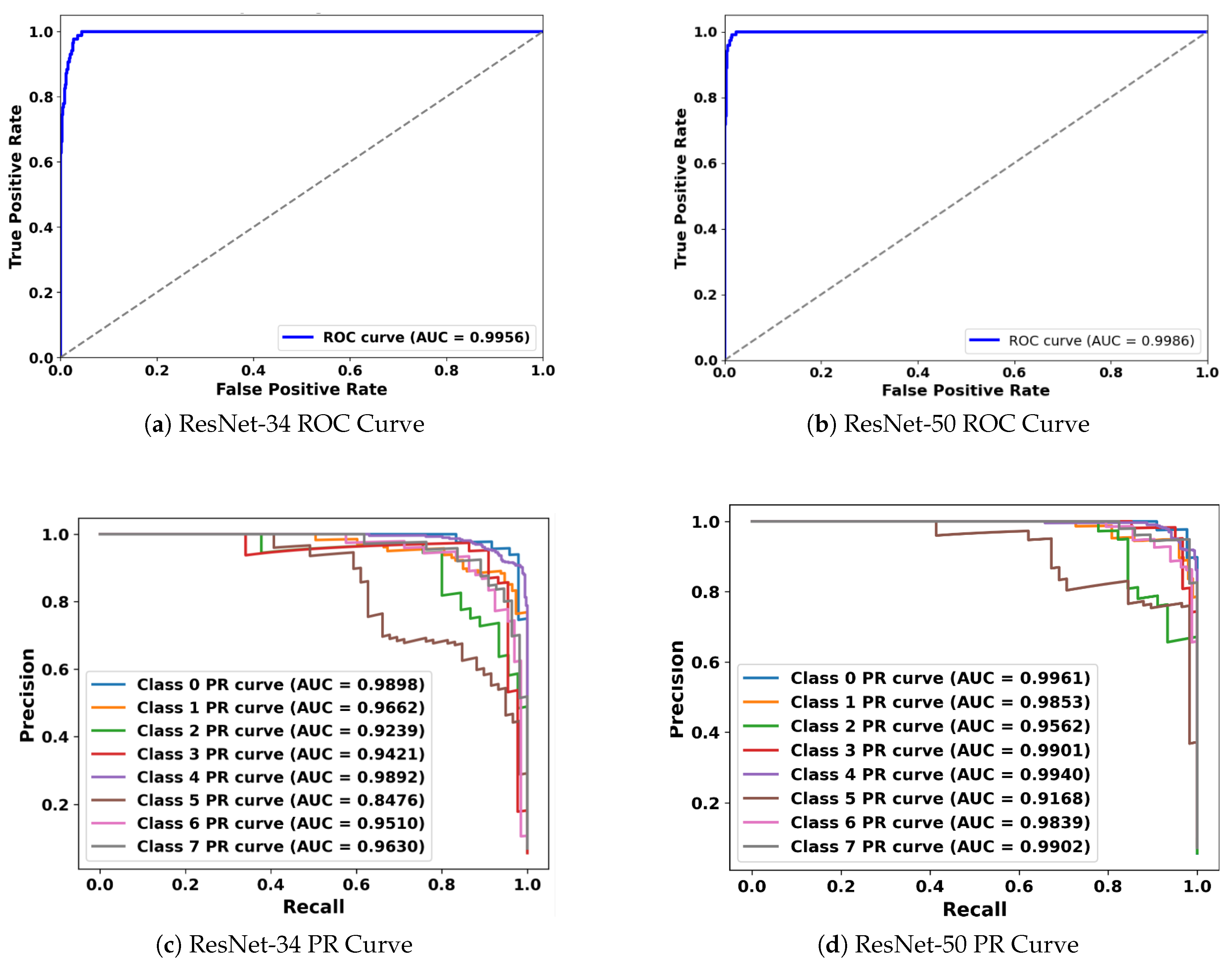
5.3. Receiver Operating Characteristic (ROC) and Precision-Recall (PR) Curve Analysis
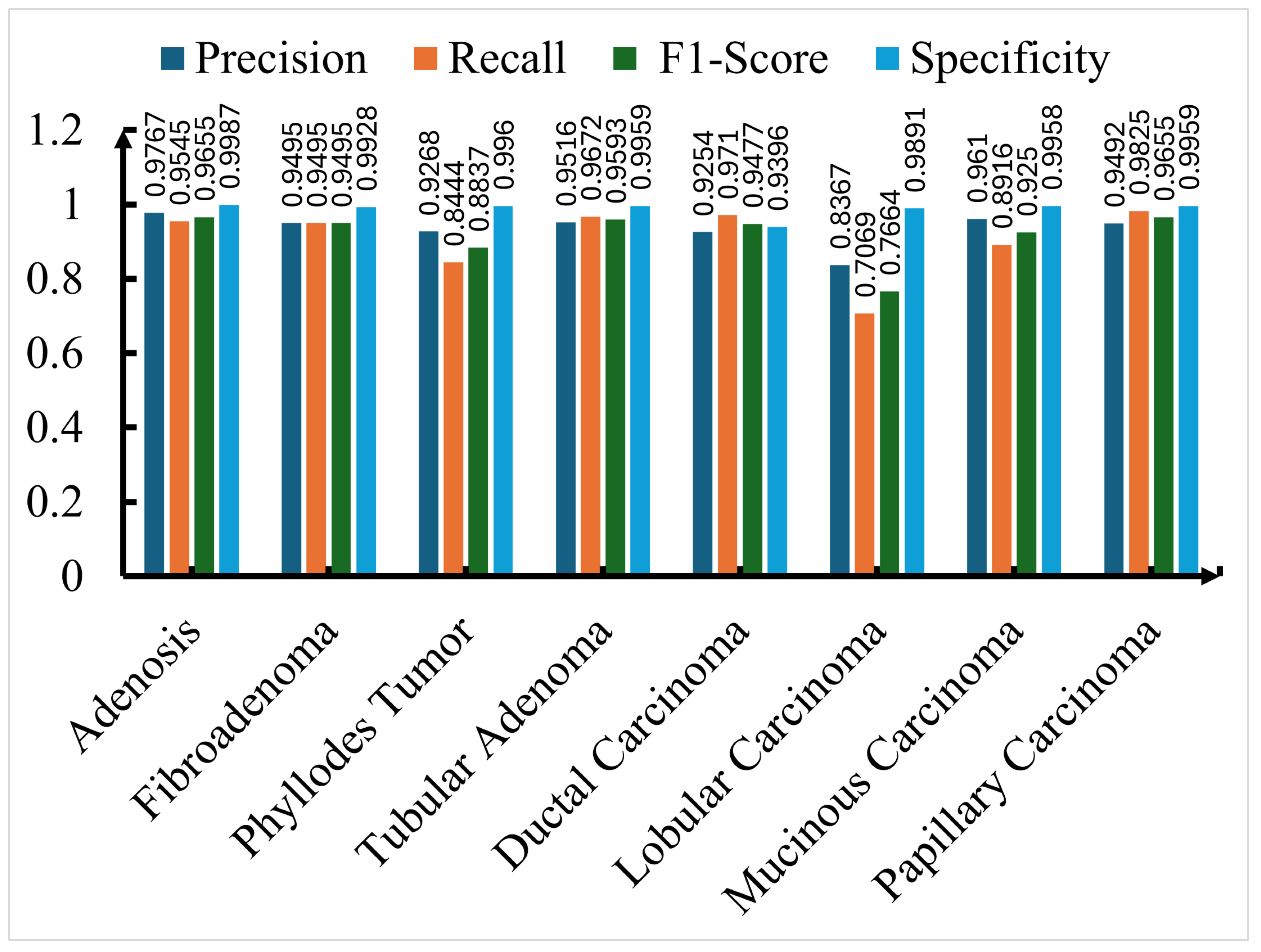
5.4. Class-wise Performance Metrics and Detailed Analysis
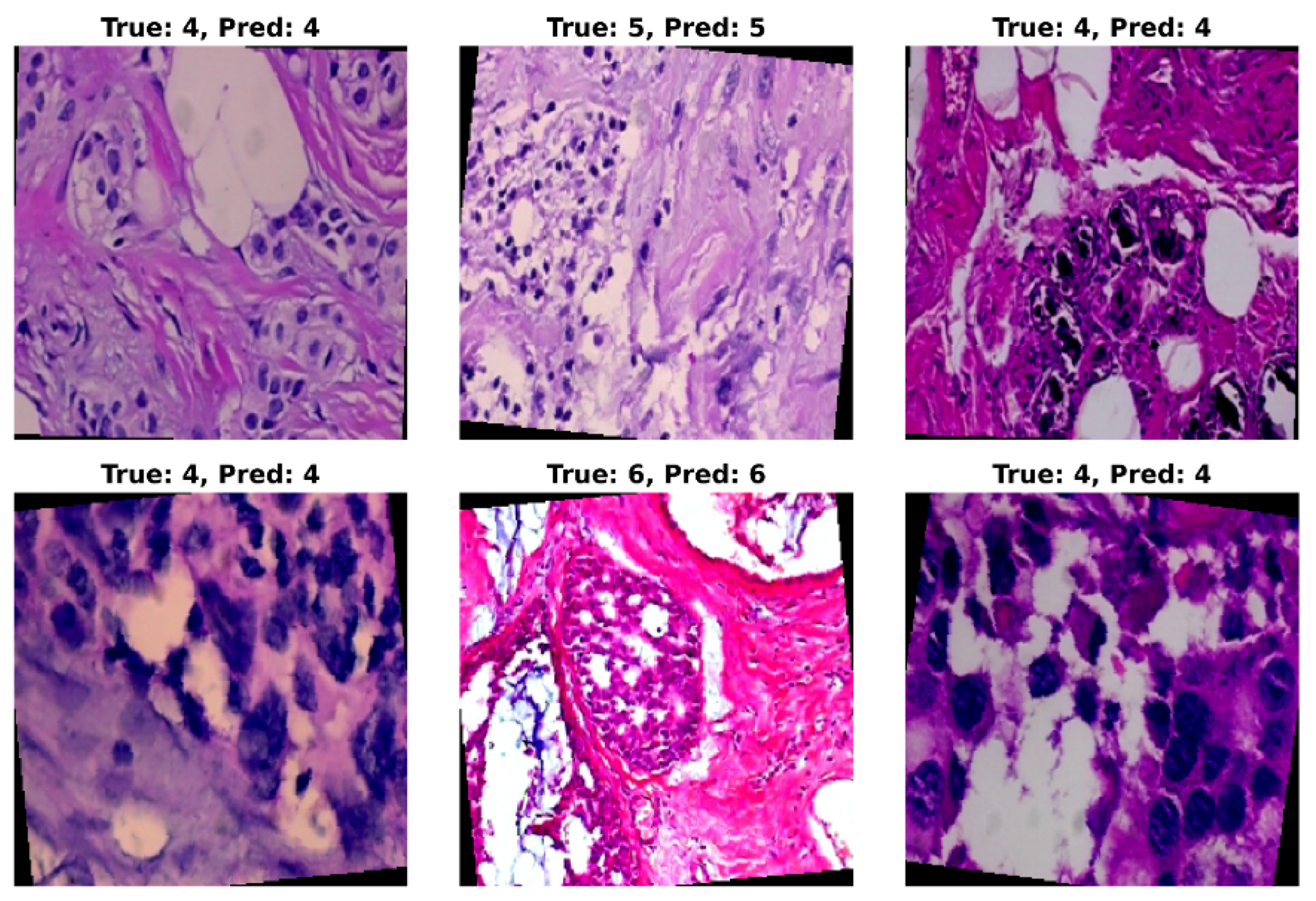
5.5. Visual Assessment of Model Predictions
6. Discussion
7. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Løyland, B.; Sandbekken, I.H.; Grov, E.K.; Utne, I. Causes and risk factors of breast cancer, what do we know for sure? An evidence synthesis of systematic reviews and meta-analyses. Cancers 2024, 16, 1583. [Google Scholar] [CrossRef] [PubMed]
- Veta, M.; Pluim, J.P.; Diest, P.J.V.; Viergever, M.A. Breast cancer histopathology image analysis: A review. IEEE Transactions on Biomedical Engineering 2014, 61, 1400–1411. [Google Scholar] [CrossRef] [PubMed]
- Aswathy, M.A.; Jagannath, M. Detection of breast cancer on digital histopathology images: Present status and future possibilities. Informatics in Medicine Unlocked 2017, 8, 74–79. [Google Scholar] [CrossRef]
- Li, C.I.; Uribe, D.J.; Daling, J.R. Clinical characteristics of different histologic types of breast cancer. British Journal of Cancer 2005, 93, 1046–1052. [Google Scholar] [CrossRef] [PubMed]
- Rakha, E.A.; et al. Breast cancer prognostic classification in the molecular era: the role of histological grade. Breast Cancer Research 2010, 12, 207. [Google Scholar] [CrossRef] [PubMed]
- Gøtzsche, P.C.; Olsen, O. Is screening for breast cancer with mammography justifiable? The Lancet 2000, 355, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Olsen, O.; Gøtzsche, P.C. Screening for breast cancer with mammography. The Cochrane Database of Systematic Reviews 2001, p. CD001877.
- Gøtzsche, P.C.; Jørgensen, K.J. Screening for breast cancer with mammography. Cochrane Database of Systematic Reviews 2013. Accessed: Mar. 01, 2025. [Online].
- Guo, R.; Lu, G.; Qin, B.; Fei, B. Ultrasound imaging technologies for breast cancer detection and management: a review. Ultrasound in Medicine & Biology 2018, 44, 37–70. [Google Scholar] [CrossRef] [PubMed]
- Gordon, P.B. Ultrasound for breast cancer screening and staging. Radiologic Clinics 2002, 40, 431–441. [Google Scholar] [CrossRef] [PubMed]
- Sood, R.; et al. Ultrasound for Breast Cancer Detection Globally: A Systematic Review and Meta-Analysis. JGO 2019, pp. 1–17. [CrossRef]
- Bluemke, D.A.; et al. Magnetic resonance imaging of the breast prior to biopsy. JAMA 2004, 292, 2735–2742. [Google Scholar] [CrossRef] [PubMed]
- Kwok, T.C.; et al. Histological grading of breast cancer on needle core biopsy: the role of immunohistochemical assessment of proliferation. Histopathology 2010, 57, 212–219. [Google Scholar] [CrossRef] [PubMed]
- Ellis, I.O.; Humphreys, S.; Michell, M.; Pinder, S.E.; Wells, C.A.; Zakhour, H. Best Practice No 179: Guidelines for breast needle core biopsy handling and reporting in breast screening assessment. Journal of Clinical Pathology 2004, 57, 897–902. [Google Scholar] [CrossRef] [PubMed]
- Chougrad, H.; Zouaki, H.; Alheyane, O. Deep Convolutional Neural Networks for breast cancer screening. Computer Methods and Programs in Biomedicine 2018, 157, 19–30. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Ge, Y.; Zhao, Y.; Guan, E.; Yan, W. Benign and malignant mammographic image classification based on Convolutional Neural Networks. In Proceedings of the Proceedings of the 2018 10th International Conference on Machine Learning and Computing, New York, NY, USA, 2018; ICMLC ’18, p. 247–251. [CrossRef]
- Saber, A.; Sakr, M.; Abo-Seida, O.M.; Keshk, A.; Chen, H. A novel deep-learning model for automatic detection and classification of breast cancer using the transfer-learning technique. IEEE Access 2021, 9, 71194–71209. [Google Scholar] [CrossRef]
- Saini, M.; Susan, S. Vggin-net: Deep transfer network for imbalanced breast cancer dataset. IEEE/ACM Transactions on Computational Biology and Bioinformatics 2022, 20, 752–762. [Google Scholar] [CrossRef] [PubMed]
- Shahidi, F.; Daud, S.M.; Abas, H.; Ahmad, N.A.; Maarop, N. Breast cancer classification using deep learning approaches and histopathology image: a comparison study. IEEE Access 2020, 8, 187531–187552. [Google Scholar] [CrossRef]
- Spanhol, F.A.; Oliveira, L.S.; Petitjean, C.; Heutte, L. A Dataset for Breast Cancer Histopathological Image Classification. IEEE Transactions on Biomedical Engineering 2016, 63, 1455–1462. [Google Scholar] [CrossRef] [PubMed]
- Alqudah, A.; Alqudah, A.M. Sliding Window Based Support Vector Machine System for Classification of Breast Cancer Using Histopathological Microscopic Images. IETE Journal of Research 2022, 68, 59–67. [Google Scholar] [CrossRef]
- Ariateja, D.; Aprilliyani, R.; Chaidir, M.D. Breast cancer histopathological images classification based on weighted K-nearest neighbor. AIP Conference Proceedings 2024, 3215, 120007. [Google Scholar] [CrossRef]
- Murtaza, G.; Abdul Wahab, A.W.; Raza, G.; Shuib, L. A tree-based multiclassification of breast tumor histopathology images through deep learning. Computerized Medical Imaging and Graphics 2021, 89, 101870. [Google Scholar] [CrossRef] [PubMed]
- Araújo, T. Others. Classification of breast cancer histopathological images using Convolutional Neural Networks. PLoS ONE 2017, 12, e0177544. [Google Scholar] [CrossRef] [PubMed]
- Bayramoglu, N.; Kannala, J.; Heikkilä, J. Deep learning for magnification independent breast cancer histopathology image classification. In Proceedings of the 2016 23rd International Conference on Pattern Recognition (ICPR); 2016; pp. 2440–2445. [Google Scholar] [CrossRef]
- Mehta, S.; Khurana, S. Enhanced Breast Tumor Detection with a CNN-LSTM Hybrid Approach: Advancing Accuracy and Precision. In Proceedings of the 2024 2nd International Conference on Recent Trends in Microelectronics, Automation, Computing and Communications Systems (ICMACC); 2024; pp. 14–18. [Google Scholar] [CrossRef]
- Kaddes, M.; Ayid, Y.M.; Elshewey, A.M.; Others. Breast cancer classification based on hybrid CNN with LSTM model. Scientific Reports 2025, 15, 4409. [Google Scholar] [CrossRef] [PubMed]
- Toma, T.A.; Biswas, S.; Miah, M.S.; Alibakhshikenari, M.; Virdee, B.S.; Fernando, S.; Rahman, M.H.; Ali, S.M.; Arpanaei, F.; Hossain, M.A.; et al. Breast Cancer Detection Based on Simplified Deep Learning Technique With Histopathological Image Using BreaKHis Database. Radio Science 2023, 58, e2023RS007761. [Google Scholar] [CrossRef]
- Benhammou, Y.; Achchab, B.; Herrera, F.; Tabik, S. BreakHis based breast cancer automatic diagnosis using deep learning: Taxonomy, survey and insights. Neurocomputing 2020, 375, 9–24. [Google Scholar] [CrossRef]
- Umer, M.J.; Sharif, M.; Kadry, S.; Alharbi, A. Multi-Class Classification of Breast Cancer Using 6B-Net with Deep Feature Fusion and Selection Method. Journal of Personalized Medicine 2022, 12. [Google Scholar] [CrossRef] [PubMed]
- Rafiq, A.; Jaffar, A.; Latif, G.; Masood, S.; Abdelhamid, S.E. Enhanced Multi-Class Breast Cancer Classification from Whole-Slide Histopathology Images Using a Proposed Deep Learning Model. Diagnostics 2025, 15. [Google Scholar] [CrossRef] [PubMed]
- Bardou, D.; Zhang, K.; Ahmad, S.M. Classification of Breast Cancer Based on Histology Images Using Convolutional Neural Networks. IEEE Access 2018, 6, 24680–24693. [Google Scholar] [CrossRef]
- Mi, W.; et al. . Deep Learning-Based Multi-Class Classification of Breast Digital Pathology Images. Cancer Management and Research 2021, 13, 4605–4617. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, P.T.; Nguyen, T.T.; Nguyen, N.C.; Le, T.T. Multiclass Breast Cancer Classification Using Convolutional Neural Network. In Proceedings of the 2019 International Symposium on Electrical and Electronics Engineering (ISEE), Oct. 2019; pp. 130–134. [Google Scholar] [CrossRef]
- Aldakhil, L.A.; Alhasson, H.F.; Alharbi, S.S. Attention-Based Deep Learning Approach for Breast Cancer Histopathological Image Multi-Classification. Diagnostics 2024, 14, 1402. [Google Scholar] [CrossRef] [PubMed]
- Tummala, S.; Kim, J.; Kadry, S. BreaST-Net: Multi-Class Classification of Breast Cancer from Histopathological Images Using Ensemble of Swin Transformers. Mathematics 2022, 10, 4109. [Google Scholar] [CrossRef]
- Joseph, A.A.; Abdullahi, M.; Junaidu, S.B.; Ibrahim, H.H.; Chiroma, H. Improved multi-classification of breast cancer histopathological images using handcrafted features and deep neural network (dense layer). Intelligent Systems with Applications 2022, 14, 200066. [Google Scholar] [CrossRef]
- Chikkala, R.B.; et al. Enhancing Breast Cancer Diagnosis With Bidirectional Recurrent Neural Networks: A Novel Approach for Histopathological Image Multi-Classification. IEEE Access 2025, 13, 41682–41707. [Google Scholar] [CrossRef]
- Sharma, S.; Mehra, R. Conventional Machine Learning and Deep Learning Approach for Multi-Classification of Breast Cancer Histopathology Images—a Comparative Insight. Journal of Digital Imaging 2020, 33, 632–654. [Google Scholar] [CrossRef] [PubMed]
| Magnification Level | Description | Application |
|---|---|---|
| 40X | Low-resolution overview of tissue structure | Identifying overall morphology |
| 100X | Balanced detail of cell structure and tissue morphology | Intermediate analysis |
| 200X | Detailed examination of cellular organization | Feature extraction for AI models |
| 400X | High-resolution visualization of individual cell structures | Fine-grained classification |
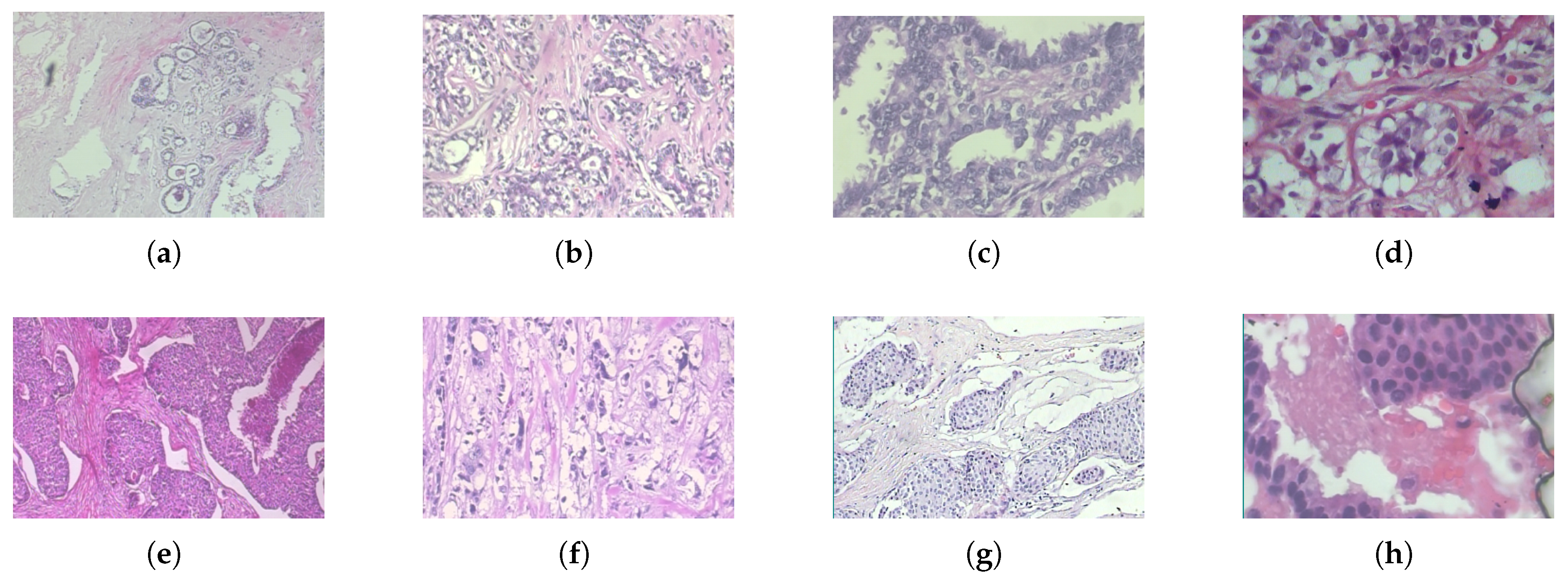
| Benign Tumors | Malignant Tumors |
|---|---|
| Adenosis: Non-cancerous overgrowth of glands within the lobules | Ductal Carcinoma: The most common malignant tumor, originating in the milk ducts |
| Fibroadenoma: Common benign tumor composed of fibrous and glandular tissues | Lobular Carcinoma: Cancer that begins in the lobules and tends to spread diffusely |
| Phyllodes Tumor: Rare fibroepithelial tumor with potential to recur | Mucinous Carcinoma: Malignant tumor characterized by mucin production |
| Tubular Adenoma: Well-circumscribed benign tumor of tightly packed tubules | Papillary Carcinoma: Malignant tumor with papillary structural patterns |
| ResNet Model | Depth | Residual Block Type | Parameters (millions) |
|---|---|---|---|
| ResNet-18 | 18 layers | Basic Block | 11.7 |
| ResNet-34 | 34 layers | Basic Block | 21.8 |
| ResNet-50 | 50 layers | Bottleneck Block | 25.6 |
| Work | Method | Dataset Split (Train:Test) | Classification Type | Accuracy |
|---|---|---|---|---|
| [32] | CNN | 70:30 | 8 Class | 88.23% |
| [33] | Inception V3 CNN | 80:20 | 8 Class | 88.16% |
| [34] | CNN | 90:10 | 8 Class | 73.68% |
| [35] | ECSAnet | 70:30 | 8 Class | 91.3% |
| Proposed | ResNet-18 | 80:20 | 8 Class | 91.41% |
| Proposed | ResNet-34 | 80:20 | 8 Class | 90.40% |
| Proposed | ResNet-50 | 80:20 | 8 Class | 92.30% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




