Submitted:
29 June 2025
Posted:
01 July 2025
You are already at the latest version
Abstract
Keywords:
Background Introduction
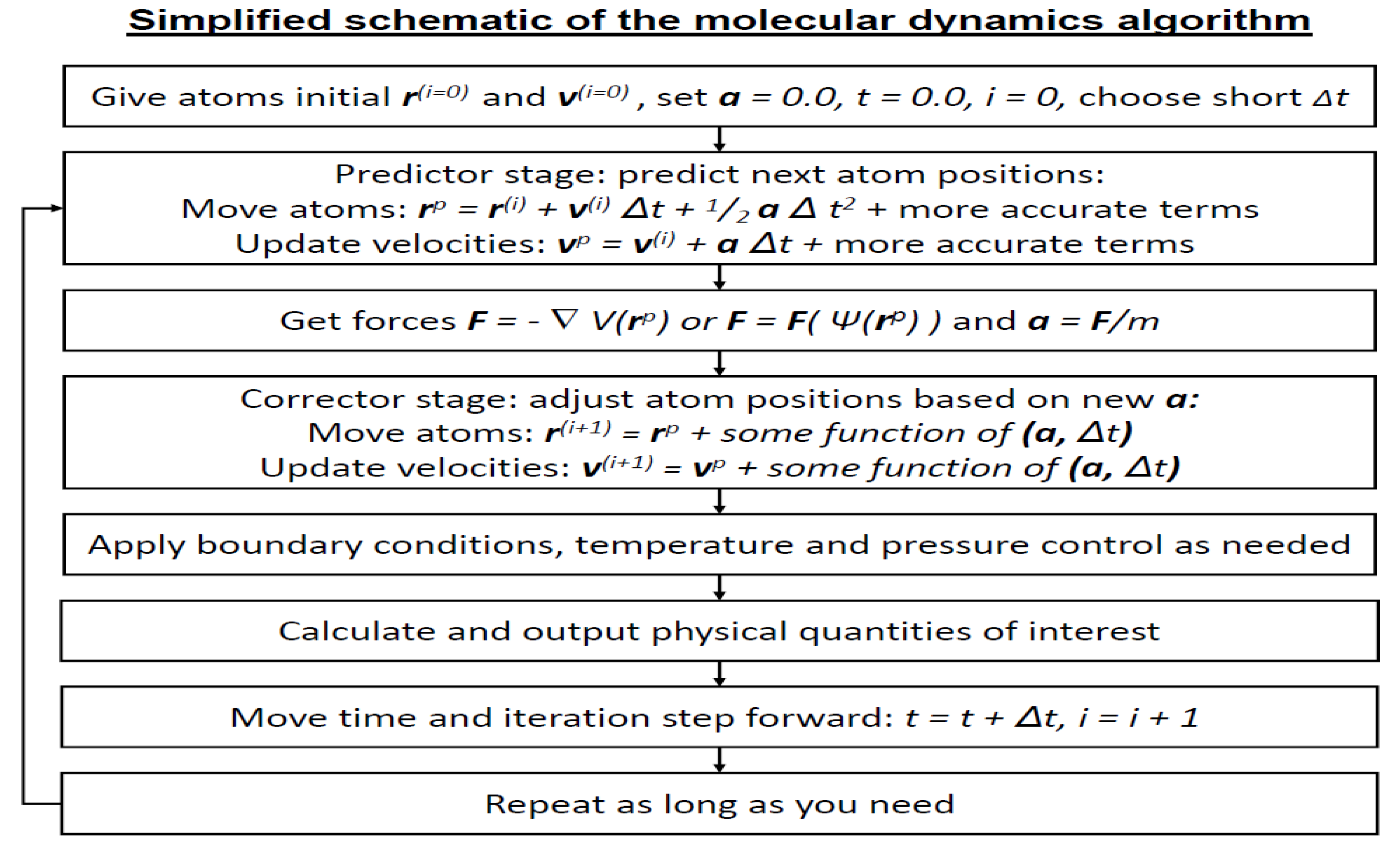
- Initializing atomic positions and velocities.
- Computing forces via classical or quantum-based potential energy calculations.
- Predicting new positions and velocities.
- Correcting these predictions using computed forces.
- Iterating this process over time steps, which may be fixed or adaptive depending on the integrator used.
- Selective synthesis – producing only the desired enantiomer directly through stereoselective chemical methods.
- Resolution of racemic mixtures – separating the two enantiomers after they’ve been formed as an achiral mixture.
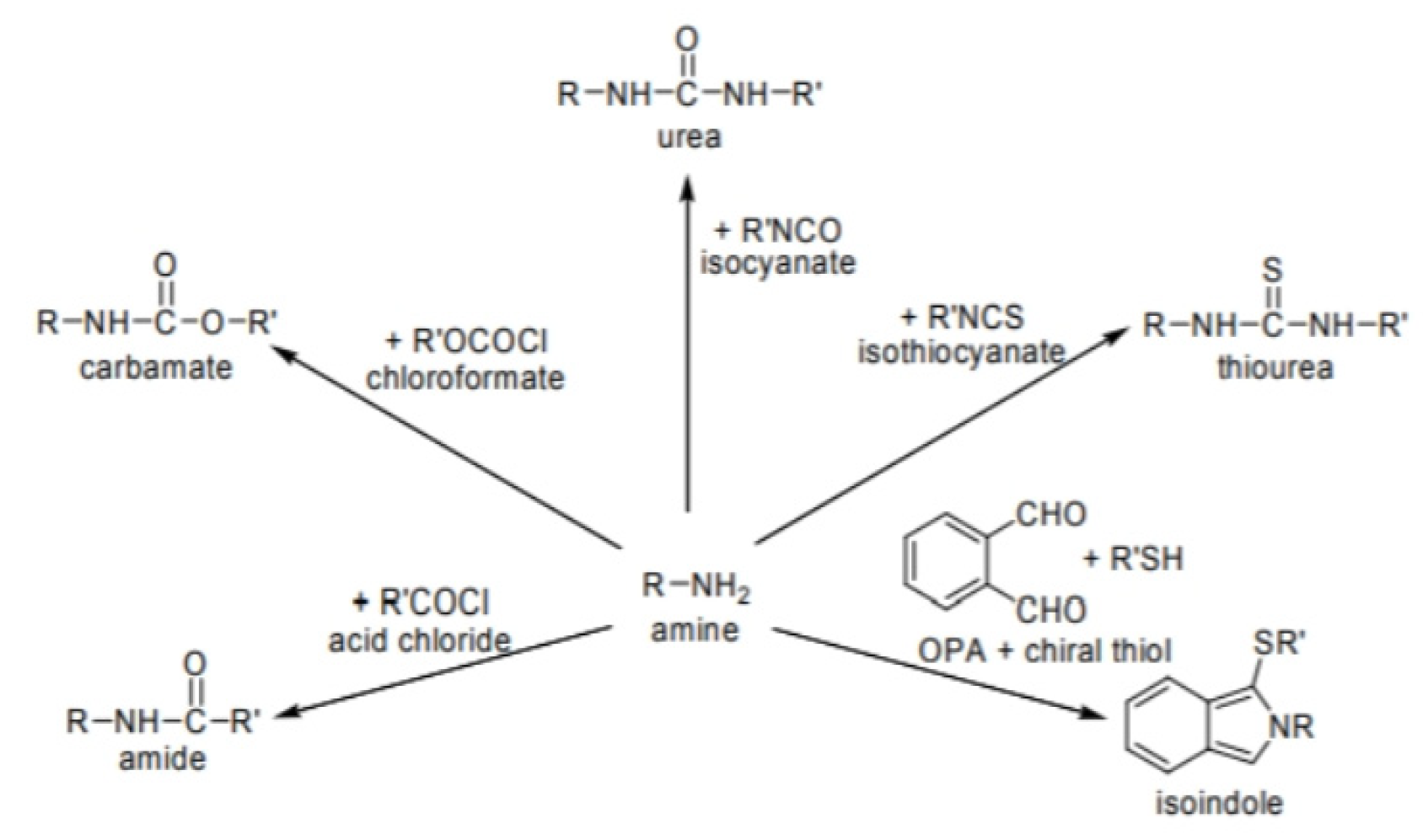
- Indirect resolution involves converting enantiomers into covalent diastereomers, which have different physicochemical properties. These differences allow separation using techniques such as crystallization, non-chiral chromatography, or distillation.
- Direct resolution relies on the formation of noncovalent diastereomeric complexes between the racemate and a chiral selector. This can be achieved using a chiral stationary phase (CSP) in chromatographic systems or by adding a chiral mobile phase additive (CMA) to interact selectively with one enantiomer over the other.
Indirect Chromatographic Methods
Direct Chromatographic Methods
Chiral Mobile Phase Additives (CMAs)
Chiral Stationary Phases (CSPs)
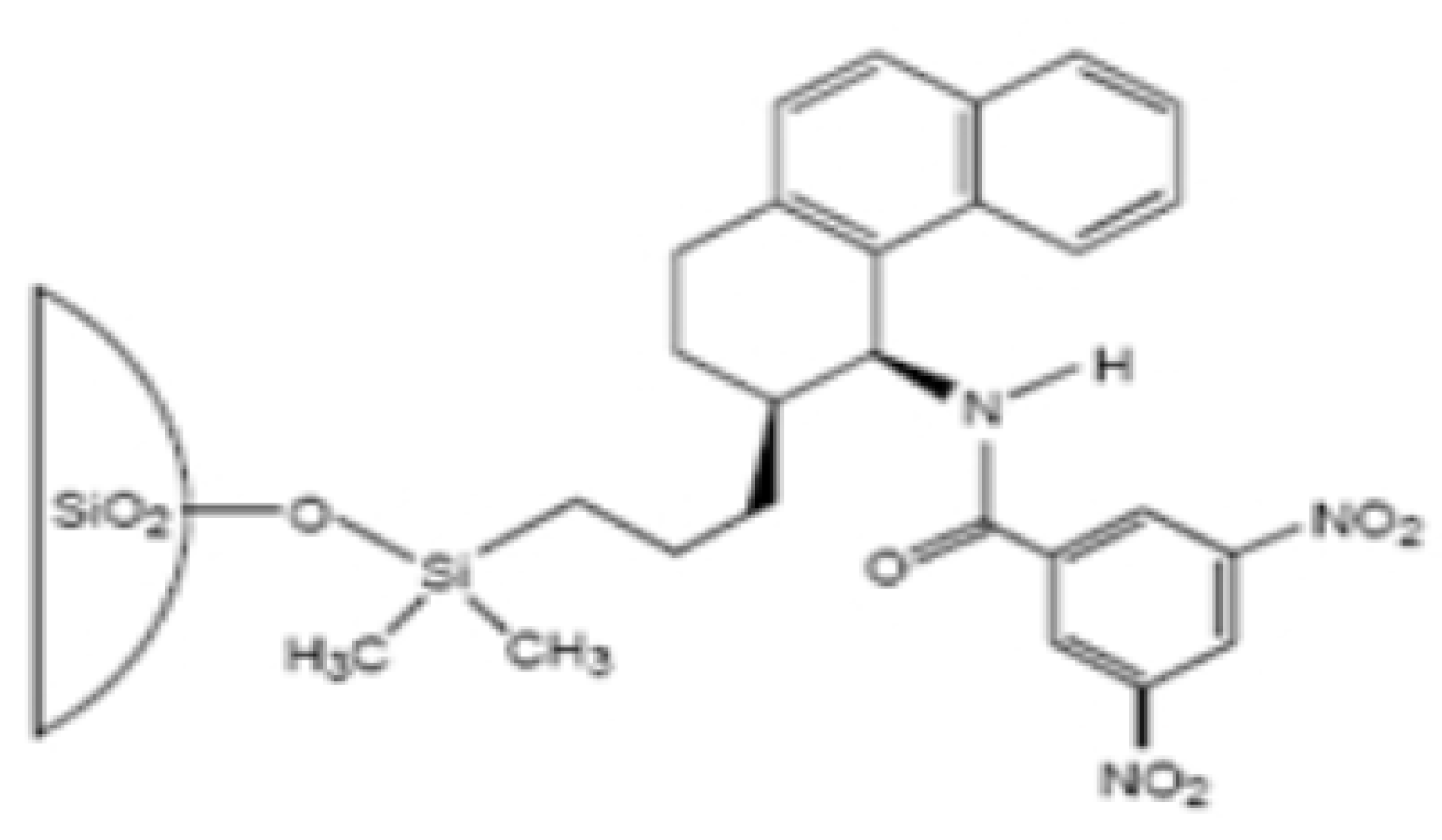
- Pirkle-type phases
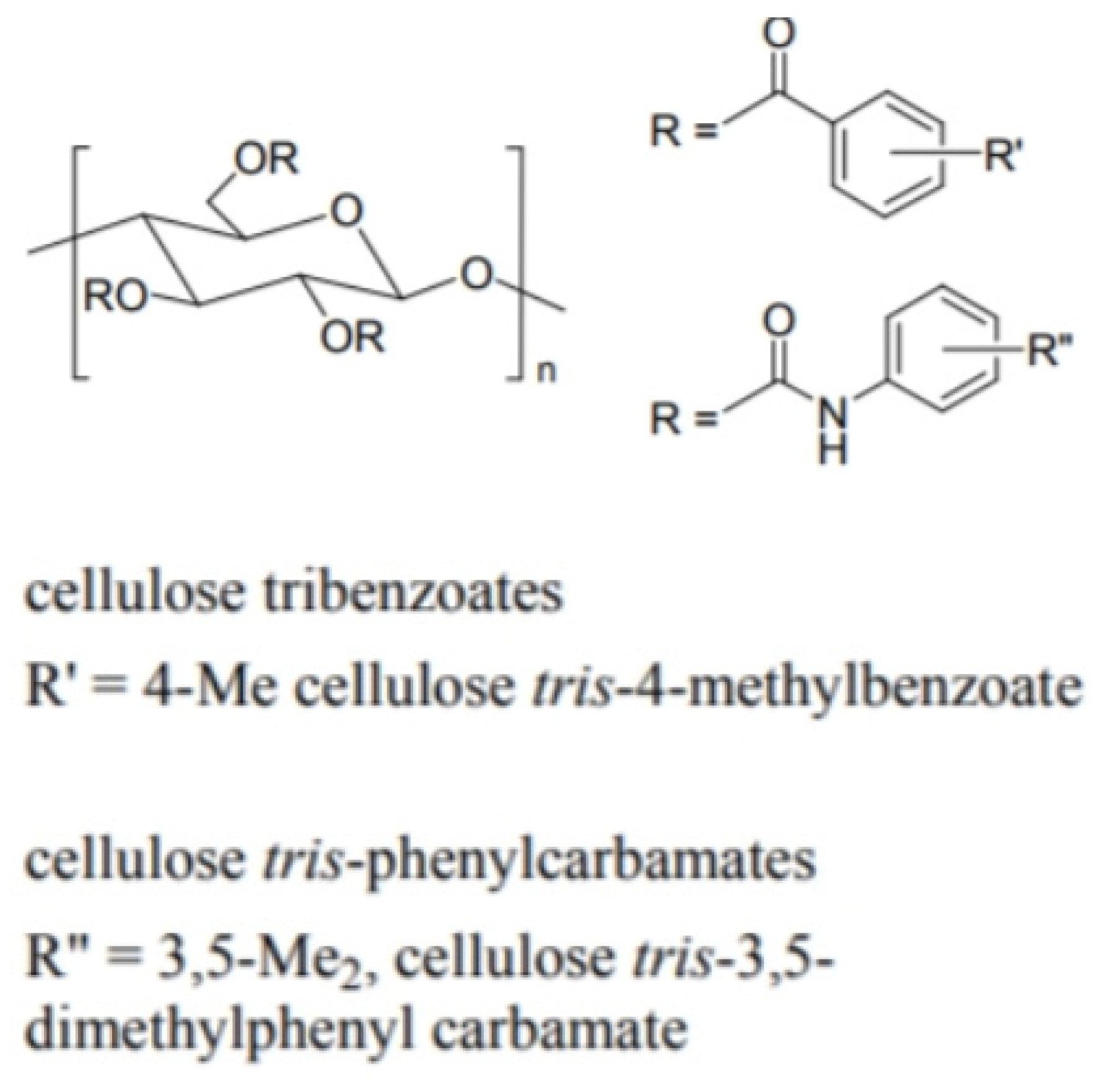
- Helical polysaccharides, such as cellulose and amylose derivatives
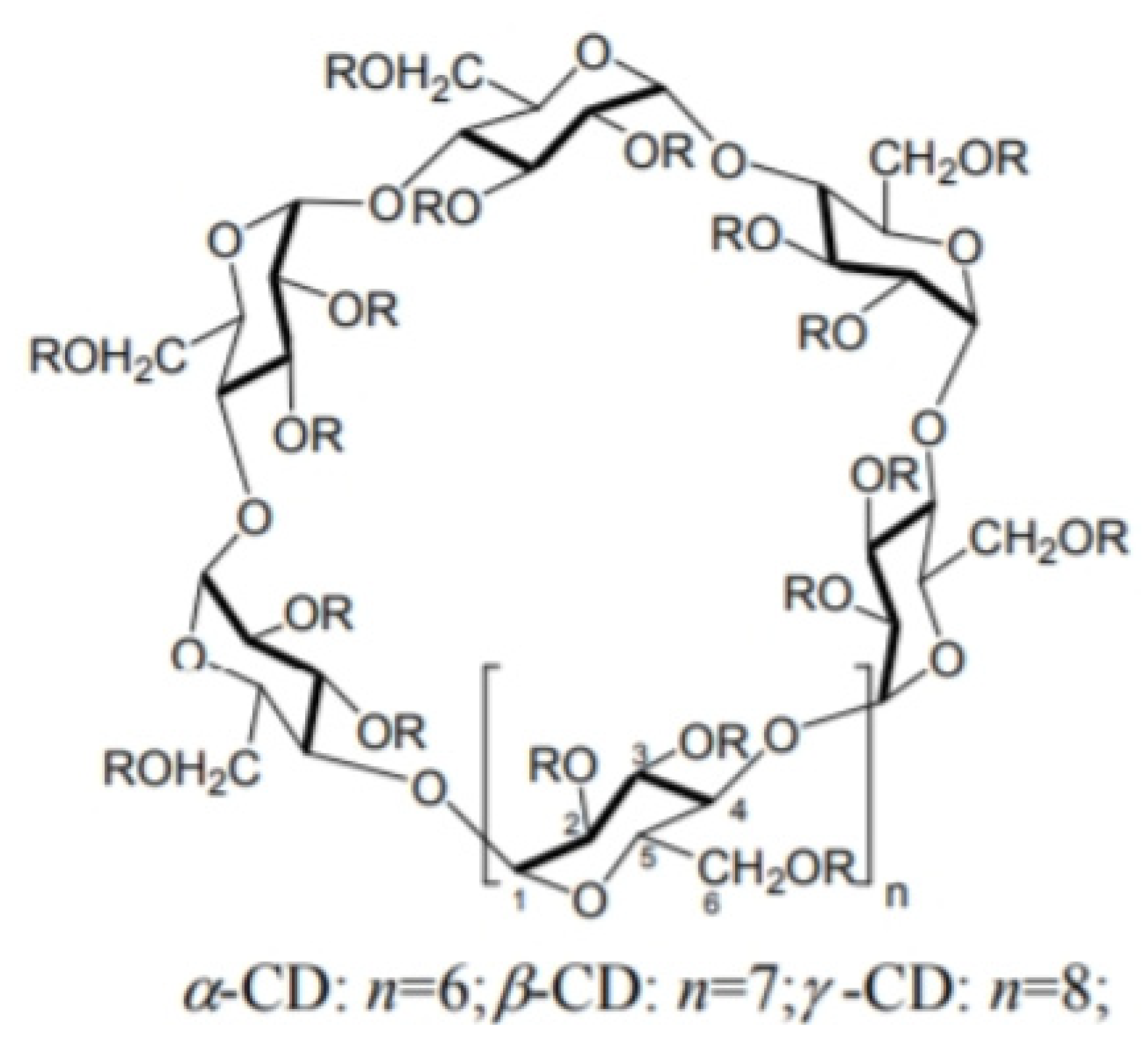
- Cavity-based materials, including cyclodextrin-derivatives
- Macrocyclic antibiotics
- Protein- and ligand-exchange phases
- Ansamycins (such as rifamycins)
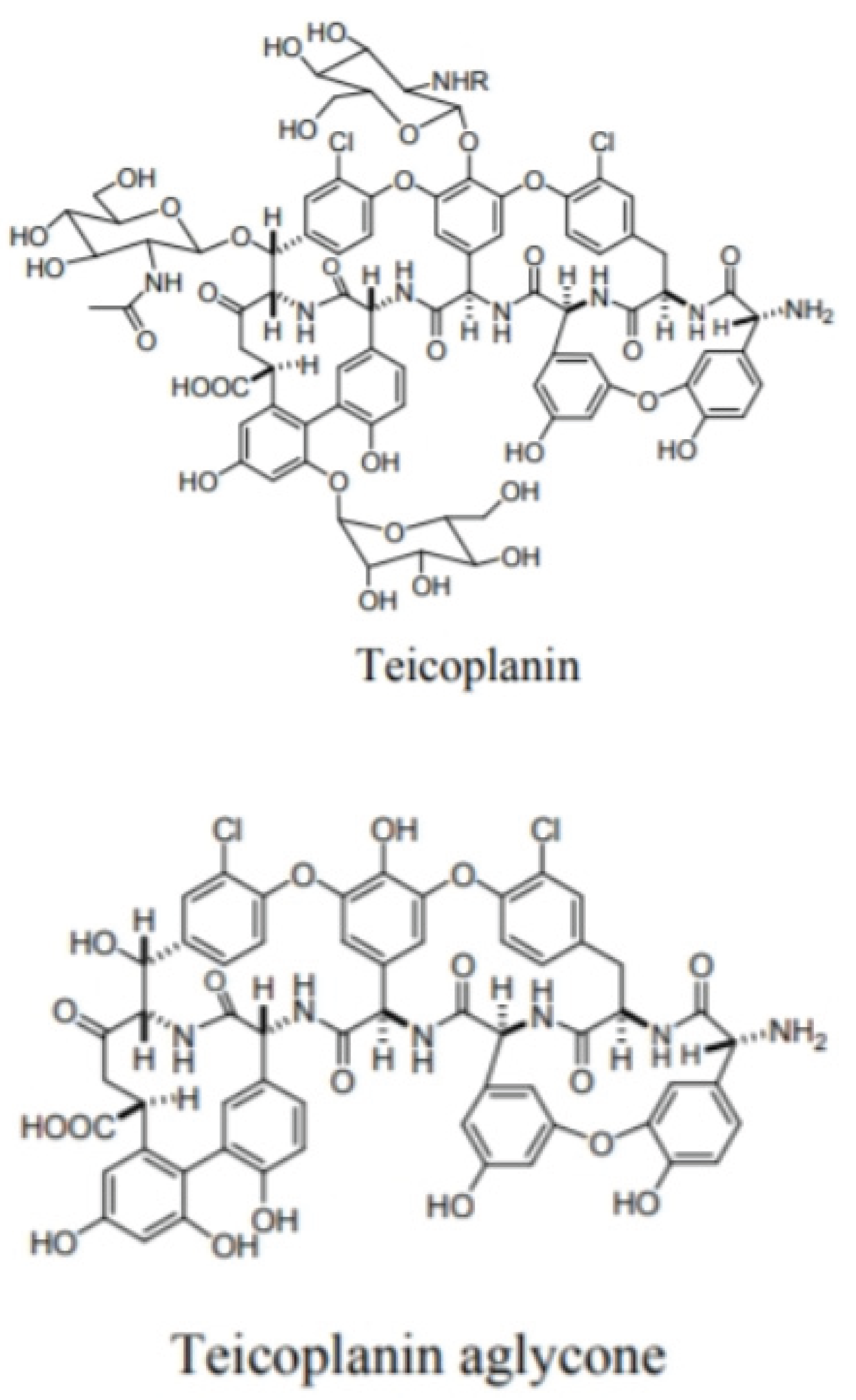
- Glycopeptide antibiotics like avoparcin, teicoplanin, ristocetin A, vancomycin, and their derivatives
- Polypeptide antibiotics, notably thiostrepton
1. Introduction
2. Earlier Models Used to Understand Chiral Discrimination
2.1. Three-Point Model
2.2. Molecular Docking of Enantiomers on a Fragment of the CSP in Vacuum
- Rigid docking setup – Both the enantiomer and a CSP fragment (e.g., a polysaccharide monomer or a short oligomer) are held fixed, typically without solvent.
- Conformational sampling – Multiple conformers of the enantiomers are generated and clustered.
- Energy estimation – Binding energies are averaged across clusters, or the energy of the most populated cluster is used as a predictive indicator.
- Elution inference – The enantiomer showing the weakest binding is predicted to elute first.
2.3. Molecular Docking Followed by Geometry Relaxation Using MD
- A single cellulose tris(3,5-dimethylphenyl carbamate) monomer may be used to dock enantiomers of three different pharmaceuticals. The most favorable pose is then refined through MD within a solvent mixture (e.g., 70/30 methanol, ethanol, or 2-propanol in n-hexane).
- In another study, a four-glucose segment of Chiralcel OJ (cellulose tris(4-methylbenzoate)) served as the docking target for pyrazole derivatives. Subsequent MD refinement—initiated from the top docking pose—was performed in ethanol, using an extended version of the polymer structure.
3. Molecular Dynamics (MD) Simulations as a Method of Describing the Dynamic Chromatographic Process
3.1. Factors That Need to Be Considered in Designing the Simulation System
-
Representing the stationary phase
- ○
- Option A: Include a fully atomistic, dynamic support—e.g., an amorphous silica slab with experimentally informed silanol distributions.
- ○
- Option B: Use a fixed silica layer where surface Si atoms serve as tethering sites for the CSP. Tether points may be arranged randomly, regularly, or based on known grafting densities. Example setups include trimethylsilyl end-capped selectors or chains grafted via amide bonds to siloxane-modified Si(111) wafers.
- ○
- Option C: Omit the solid support entirely and either restrain the polymer fragment or allow it to float freely as a simplified model.
- Choosing the CSP fragment size
-
o Large constructs such as quadruple 18-mer polysaccharide strands.
- ○
- A single 12-mer polymer chain.
- ○
- Smaller oligomers—hexamers, tetramers, or dimers—are often preferred. A monomer alone lacks the structural groove characteristic of helical polysaccharides.
- ○
- For brush-type CSPs, tether length to the silica substrate is an additional variable.
-
Modeling the solvent environment
- ○
- Explicit solvent: All molecules are represented atomistically, matching experimental composition.
- ○
- Continuum dielectric model: Solvent effect approximated via a variable dielectric medium.
- ○
- Vacuum: Used for simplified or initial exploratory simulations.
-
Applying rigid-body constraints
- ○
- To accelerate simulations, rigid-body treatments are used—especially for aromatic rings with stiff internal geometry. Forces may be applied to maintain translational and rotational dynamics, while internal atomic positions remain fixed (e.g., the dinitrophenyl and tetrahydrophenanthrene units in Whelk-O1).
- Choosing force fields [4]
- o Selections include AMBER with GAFF for organic molecules, CHARMM or OPLS for biomolecules, and specialized models such as Siepmann’s TraPPE-UA for solid phases or interfaces.
- Starting structure bias: Beginning from a docked complex or a known elution pose may precondition the results. Unbiased starting conditions are preferred for credibility.
- CSP truncation artifacts: Shortening polymer chains can introduce “end effects” that distort binding behavior.
- Omitting the substrate: Simulations lacking solid support may misrepresent interfacial recognition mechanisms.
- Finite-size effects: Box dimensions must be tested; results should stabilize as size increases.
3.2. It Is Important to Include Explicit Solvent [13] Molecules
- Electrostatic interactions grow stronger in low-dielectric (nonpolar) solvents and weaken in high-dielectric (polar) ones. In MD simulations, this is effectively modeled by the dielectric constant—ranging from ~1 in nonpolar environments up to ~80 in polar systems—being applied to the Coulomb term, which uniformly suppresses electrostatic interactions.
- As electrostatic contributions diminish in polar media, van der Waals forces take on a more prominent role, altering molecular dynamics and binding landscapes—especially at close range.
- Explicit solvent modeling is essential because it captures phenomena such as solvent-driven solvation/desolvation events and hydrogen-bond displacement when an enantiomer approaches the selector—dynamics that implicit or continuum models cannot accurately resolve.
- Experiments confirm that the mobile phase composition influences chiral recognition not just indirectly but by changing the structural behavior of the CSP and analyte. For example, the ADMPC polymer adopts different conformations when switching from methanol to hexane/isopropanol or acetonitrile. Solid-state NMR shows that hexane molecules can become embedded within CSP structures, while polar solvents can disrupt intra- and intermolecular hydrogen-bond networks.
4. MD Studies on Covalently Bonded Selectors
4.1. Preparing the Interface: Solvent Effects on Covalently-Bonded Selectors
- At typical chain densities of ~2.9 µmol/m² or higher and with tether lengths equivalent to eight carbon atoms or more, RPLC systems reproduce expected retention behaviors in silico.
- The retention mechanism at the molecular level is extraordinarily complex—dependent on subtle variations in chain alignment, solvent structuring near the surface, and grafting density—such that broad generalizations across systems are unreliable.
- PEPU (N-(1-phenylethyl)-N’-3-(triethoxysilyl)propyl-urea)
- R-DNB-leucine and R-DNB-phenylglycine
- Proline-based selectors with varying moieties
- Whelk-O1 (1-(3,5-dinitrobenzamido)-1,2,3,4-tetrahydrophenanthrene)
4.2. MD Simulations of Chiral Analytes on Whelk-O1 Selectors
- H-bonding via the amide hydrogen plus π–π stacking between the analyte’s aromatic ring and the selector’s dinitrophenyl group (the most common).
- H-bonds involving the amide oxygen.
- H-bonds with nitro oxygen atoms.
- π–π stacking with the tetrahydrophenanthrene moiety.
4.3. MD Simulations of Chiral Analytes on Proline-Type Selectors
5. MD Studies of Chiral Molecules Interacting with Cyclodextrins
- In trajectories lasting under 10 ns, water rarely penetrates the CD cavity.
- In longer runs (e.g., 50 ns), water ingress into the cavity occurs but only after tens of nanoseconds.
- Studies with β-CD and ibuprofen enantiomers in methanol show that methanol molecules preferentially occupy hydrogen-bonding sites, significantly altering binding compared to vacuum MD.
6. MD Simulations of Chiral Recognition Using Polysaccharide CSPs
6.1. The Structure of Polysaccharide-Based CSPs
6.2. MD of Chiral Interaction with Polysaccharide-Based CSP Without Explicit Solvent
6.3. MD Using a Single Polymer Strand in the Solvent System
- Equilibration of the ADMPC 12-mer in each solvent (heptane/IPA 90/10, methanol, acetonitrile) for 100 ns to obtain a representative backbone structure.
- Drug inclusion followed by additional 100 ns simulations, all initiated from the most populated backbone conformation (analyzed via φ/ψ dihedral maps akin to protein Ramachandran plots).
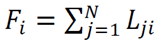
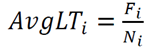
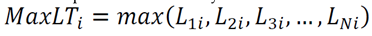
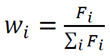
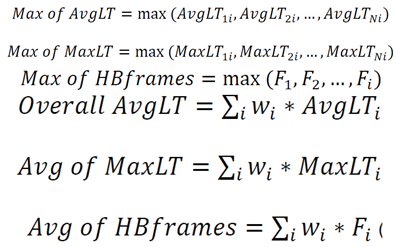
- For ambucetamide and etozoline, where experimental elution orders are not yet established, all MD metrics consistently predicted the same order. This consistency gives confidence in our assignments.
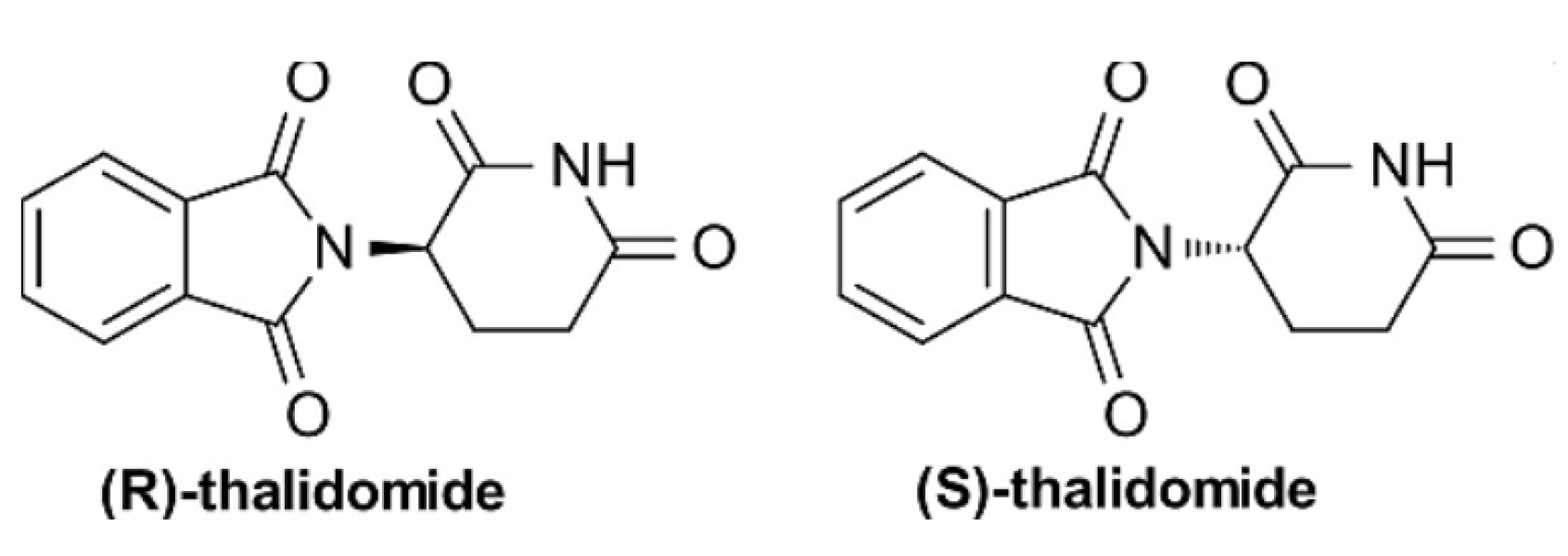
- For all analytes except thalidomide, the MD analysis correctly predicted elution order.
- The strongest statistical correlations emerged from metrics based on maximum hydrogen-bond lifetimes—either the average of individual maximum lifetimes for each donor–acceptor pair or the overall maximum lifetime.
- In contrast, the unrestrained (free-floating) 12-mer performed poorly: hydrogen-bonding metrics were inconsistent, and ring–ring interaction orientations failed to differentiate R and S for several analytes, indicating excessive polymer mobility compromise.
6.4. MD Using Multiple Polymer Strands Coated on Amorphous Silica
- Multiple 18-mer ADMPC polymer chains tethered onto an amorphous silica surface, replacing previous single-chain models.
- A fully dynamic, unrestrained system without artificial constraints on atomic movements.
- Consideration of chain–chain interactions, allowing enantiomers to simultaneously interact with adjacent chains.
- A realistic interfacial geometry, where analytes can only approach the exterior face of the brush layer, as on a real HPLC column.
- For benzoin and flavanone, all six MD metrics consistently matched experimental elution orders, with average hydrogen-bond lifetimes emerging as the most predictive.
- For thalidomide (in methanol), longer lifetimes for R-enantiomer H-bonds aligned with the experimentally observed elution of S first—unresolved in earlier single-chain models.
- For valsartan, R was correctly predicted to elute first. The multi-chain model found many long-lived interactions involving multiple donor–acceptor pairs, rendering separation factor predictions more realistic than before.
- Crucially, inter-chain interactions—instances where a single enantiomer engages with two adjacent polymer strands—were observed frequently (see simulations), especially for larger analytes like valsartan. These cooperative binding events were entirely missed in single-chain models.
7. MD Simulations in Terms of Quantitative Differential/ Discriminatory Aspects for One Enantiomer Relative to the Other
7.1. Counts of H-Bonds and Distribution of Hydrogen Bonding [7] Lifetimes
7.2. The role of Ring-Ring Interactions
7.3. Counts of Diastereomeric Complexes and Lifetimes of Diastereomer Types
7.4. Free Energy Profile
- CSP2: the normal setup with the wide opening exposed
- CSP1: the inverted orientation exposing the small opening
- Along the O1 path, energy quickly drops when the A-ring enters the cup, reaching a minimum at ξ≈−0.8ξ≈−0.8 Å for R and −0.2−0.2 Å for S—reflecting stabilization with the A-ring centered.
- A barrier is encountered as the molecule advances, followed by a second minimum at ξ=+4.1ξ=+4.1 Å, where the C-ring occupies the cavity and the B-ring rests near the narrow entrance.
- The O2 path’s energy profile mirrors the O1 pathway due to symmetry, reflecting similar energetics whether entering or exiting the wide rim.
- From these PMF profiles, Li calculated binding free energies ΔGbind:
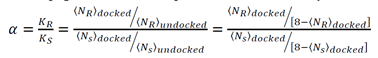
- O1: R is more strongly bound (−0.98−0.98 kcal/mol) than S (−0.61−0.61 kcal/mol), a difference of 0.37 kcal/mol.
- O2: S binds more tightly (−1.83−1.83 kcal/mol) compared to R (−0.27−0.27 kcal/mol), yielding a larger energy gap (1.56 kcal/mol).
- Consequently:
- With CSP1 (inverted cup), R is retained longer than S.
- With CSP2 (normal cup), S shows stronger retention, reversing elution order.
7.5. One-Body and Two-Body Distribution Functions
8. Future Work
- Predict the impact of functional-group modifications on the CSP’s selectivity
- Estimate how these changes would alter separation factors
- Assess elution preferences for enantiomers under refined conditions
Conclusions
- It supports virtual screening of large compound libraries.
- It predicts drug–target binding and unbinding kinetics, which increasingly inform drug efficacy and residence times.
- It deciphers allosteric mechanisms, offering insights into non-obvious regulatory sites.
- It probes water thermodynamics in binding pockets, guiding design of potent, tightly bound ligands.
- Identifying preferred analyte binding locations on a chiral stationary phase (CSP).
- Determining whether both enantiomers occupy the same binding site or different ones.
- Elucidating specific intermolecular forces responsible for binding or the formation of diastereomeric complexes.
- Uncovering which interactions contribute to enantiomeric discrimination.
- Pinpointing CSP fragments most critical to binding and whether they also drive selectivity.
- Characterizing the CSP conformations most effective for discrimination.
- Investigating cooperative effects or induced-fit mechanisms—i.e., conformational changes in the CSP, analyte, or both—that enhance binding.
- Assessing the role of solvent in binding strength and selectivity.
Acknowledgment:
Ethics Declaration:
References
- Kazakevich, Y. , & LoBrutto, R. (2007). *HPLC for Pharmaceutical Scientists*. Wiley-Interscience.
- Jameson, C. J. , Wang, X., & Murad, S. (2021). Molecular Dynamics Simulations of Enantiomeric Separations as an Interfacial Process in HPLC. *AIChE Journal*, 67(7), e17143. [CrossRef]
- Easson, L. H. , & Stedman, E. (1933). Studies on the relationship between chemical constitution and physiological action. *Biochemical Journal*, 27(5), 1257–1266. [CrossRef]
- Pirkle, W. H. , & Pochapsky, T. C. (1986). Considerations of chiral recognition relevant to the liquid chromatographic separation of enantiomers. *Chemical Reviews*, 86(3), 997–1013. [CrossRef]
- Schlick, T. (1996). Pursuing Laplace’s Vision on Modern Computers. *IMA Volumes in Mathematics and Its Applications*, 82, 219–247. [CrossRef]
- Frenkel, D. , & Smit, B. (2002). *Understanding Molecular Simulation: From Algorithms to Applications*. Academic Press.
- Cornell, W. D. , et al. (1995). A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. *Journal of the American Chemical Society*, 117(19), 5179–5197. [CrossRef]
- Brooks, B. R. , et al. (2009). CHARMM: The biomolecular simulation program. *Journal of Computational Chemistry*, 30(10), 1545–1614. [CrossRef]
- Pronk, S. , et al. (2013). GROMACS 4.5: A high-throughput and highly parallel open source molecular simulation toolkit. *Bioinformatics*, 29(7), 845–854. [CrossRef]
- Lipkowitz, K. B. (1995). Applications of computational chemistry to the study of chiral recognition in chromatography. *Chemical Reviews*, 95(6), 1625–1660. [CrossRef]
- Roux, B. (1995). The calculation of the potential of mean force using computer simulations. *Computer Physics Communications*, 91(1-3), 275–282. [CrossRef]
- Cann, N. M. , & Smith, D. A. (2019). Molecular modeling of chiral separations: from force fields to interaction lifetimes. *Journal of Chromatography A*, 1594, 158–168. [CrossRef]
- Wang, X. , & Murad, S. (2020). MD simulations of interfacial interactions in chiral chromatography. *Langmuir*, 36(12), 3156–3164. [CrossRef]
- Hunter, C. A. , & Sanders, J. K. M. (1990). The nature of π–π interactions. *Journal of the American Chemical Society*, 112(14), 5525–5534. [CrossRef]
- Meyer, E. A. , Castellano, R. K., & Diederich, F. (2003). Interactions with aromatic rings in chemical and biological recognition. *Angewandte Chemie International Edition*, 42(11), 1210–1250. [CrossRef]
- Morris, G. M. , et al. (1998). Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. *Journal of Computational Chemistry*, 19(14), 1639–1662. [CrossRef]
- Goodsell, D. S. , & Olson, A. J. (1990). Automated docking of substrates to proteins by simulated annealing. *Proteins: Structure, Function, and Bioinformatics*, 8(3), 195–202. [CrossRef]
- Wang, R. , et al. (2002). The PDBbind database: collection of binding affinities for protein–ligand complexes with known three-dimensional structures. *Journal of Medicinal Chemistry*, 47(12), 2977–2980. [CrossRef]
- Stojanov, A. , et al. (2022). Cyclodextrin-based selectors for enantioselective chromatography: Structure–function relationships. *Journal of Separation Science*, 45(2), 288–300. [CrossRef]
- Li, X. , et al. (2020). MD simulations and PMF analysis of flavanone enantiomers in β-cyclodextrin. *Journal of Chromatography A*, 1615, 460777. [CrossRef]
- Kirkwood, J. G. (1935). Statistical Mechanics of Fluid Mixtures. *Journal of Chemical Physics*, 3(5), 300–313. [CrossRef]
- Allen, M. P. , & Tildesley, D. J. (1987). *Computer Simulation of Liquids*. Oxford University Press.
- Berendsen, H. J. C. , et al. (1984). Molecular dynamics with coupling to an external bath. *Journal of Chemical Physics*, 81(8), 3684–3690. [CrossRef]
- Murad, S. , et al. (2022). Silica-supported polysaccharide CSPs and inter-chain interactions in HPLC: A MD study. *Journal of Molecular Liquids*, 360, 119505. [CrossRef]
- Zhang, H. , et al. (2021). Atomistic insights into hydrogen bonding in chiral stationary phases. *Analytica Chimica Acta*, 1154, 338320. [CrossRef]
- Bianchi, D. , & Mariella, R. (2021). Explicit-solvent simulations of cellulose-based CSPs: Elution order predictions. *Analytical Chemistry*, 93(7), 3429–3437. [CrossRef]
- Du, Q. , et al. (2019). A multi-strand simulation model of chiral recognition. *Chemical Physics Letters*, 721, 96–102. [CrossRef]
- Bereznitski, Y. , et al. (2000). Thermodynamics of chiral separation in ADMPC CSPs. *Journal of Chromatography A*, 875(1–2), 65–79. [CrossRef]
- Kasat, R. B. , et al. (2005). Structural aspects of polysaccharide CSPs from solid-state NMR. *Macromolecules*, 38(12), 4874–4881. [CrossRef]
- Roux, B. , & Berneche, S. (2002). Biological ion channels and free energy barriers. *Philosophical Transactions of the Royal Society B: Biological Sciences*, 357(1417), 119–128. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).