Submitted:
10 June 2025
Posted:
12 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Works
2.1. The Deep Learning for Classification of MRI Brain Tumors
2.2. The Ensemble of Deep CNNs for Classification of MRI Brain Tumors
3. Materials and Methods
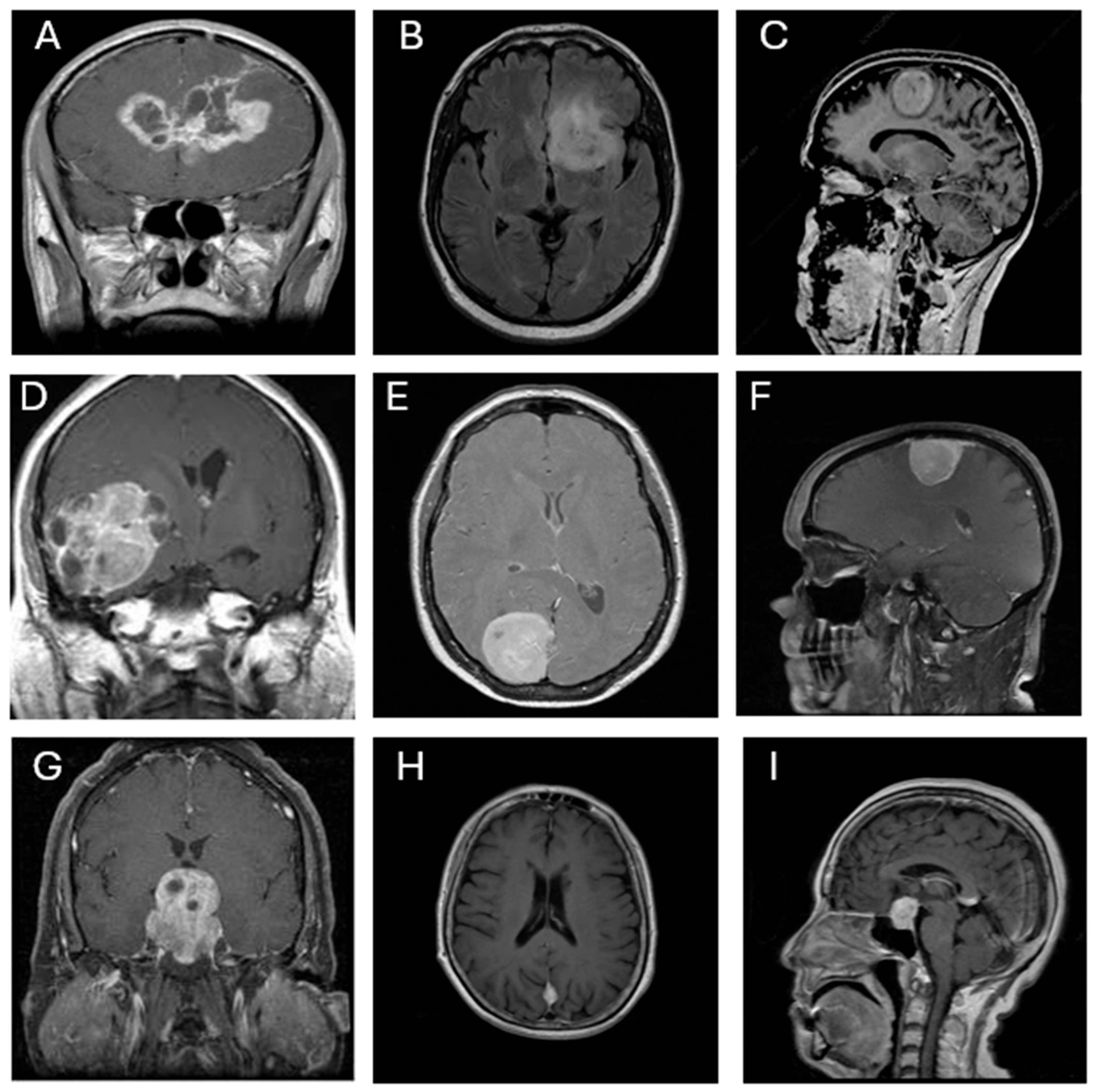
3.1. Dataset Description
3.2. Selected CNN Architectures and Voting Schema
3.3. Data Splitting and Training Configuration
- Training set: 70% of the data used for weight optimization.
- Validation set: 10% used to monitor generalization performance and apply early stopping.
- Testing set: 20% used exclusively for final model evaluation.
3.4. Evaluation Performance
- Accuracy (Acc): The proportion of correctly predicted instances among all predictions, reflecting the overall classification performance.
- Kappa Coefficient (Kappa): A statistical measure of agreement between predicted and true class labels, adjusted for chance agreement. Values closer to 1 indicate stronger consistency.
- True Positive Rate (TP): Also referred to as sensitivity or recall, this metric was computed for each class—TP1 to TP4—corresponding to glioma, meningioma, no tumor, and pituitary tumor, respectively. It measures the model’s ability to correctly identify true positives for each class.
- Precision (Pre): The ratio of true positive predictions to the total number of positive predictions, calculated as Pre1 to Pre4 for glioma, meningioma, no tumor, and pituitary tumor, respectively. This indicates the model’s reliability in its positive predictions.
- Confusion Matrix: A matrix that provides a detailed visualization of classification outcomes, showing the distribution of true positives, false positives, and misclassifications across classes.
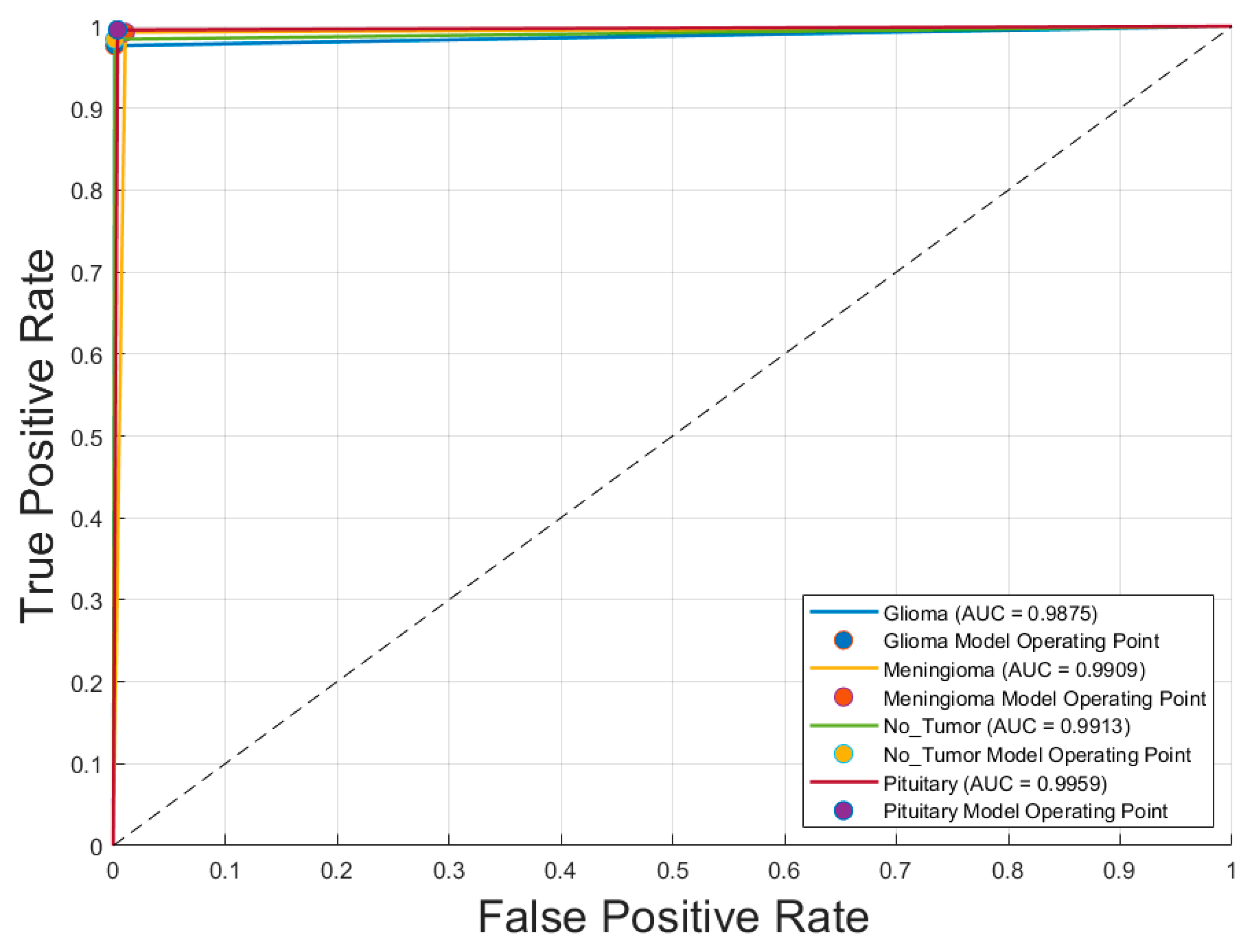
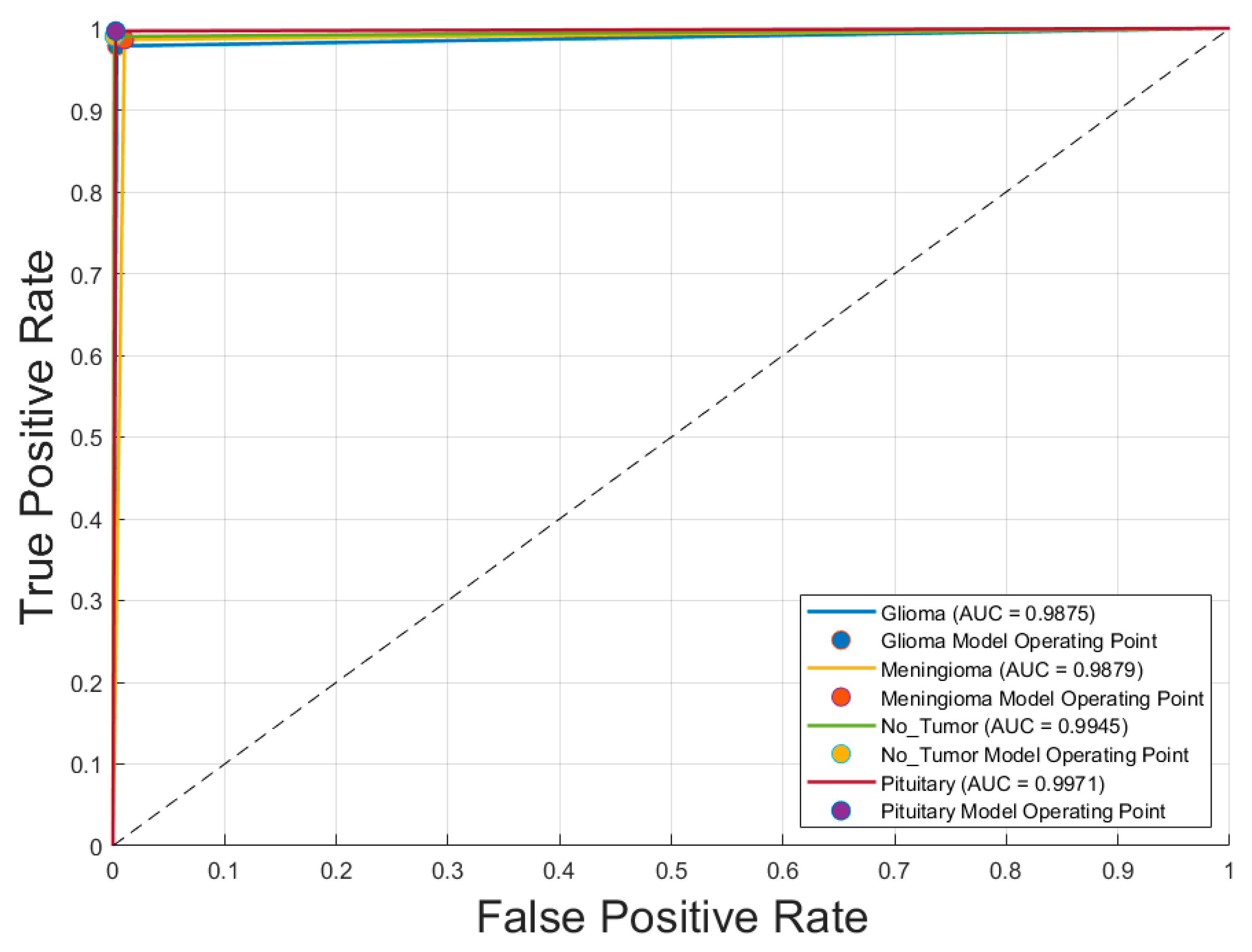
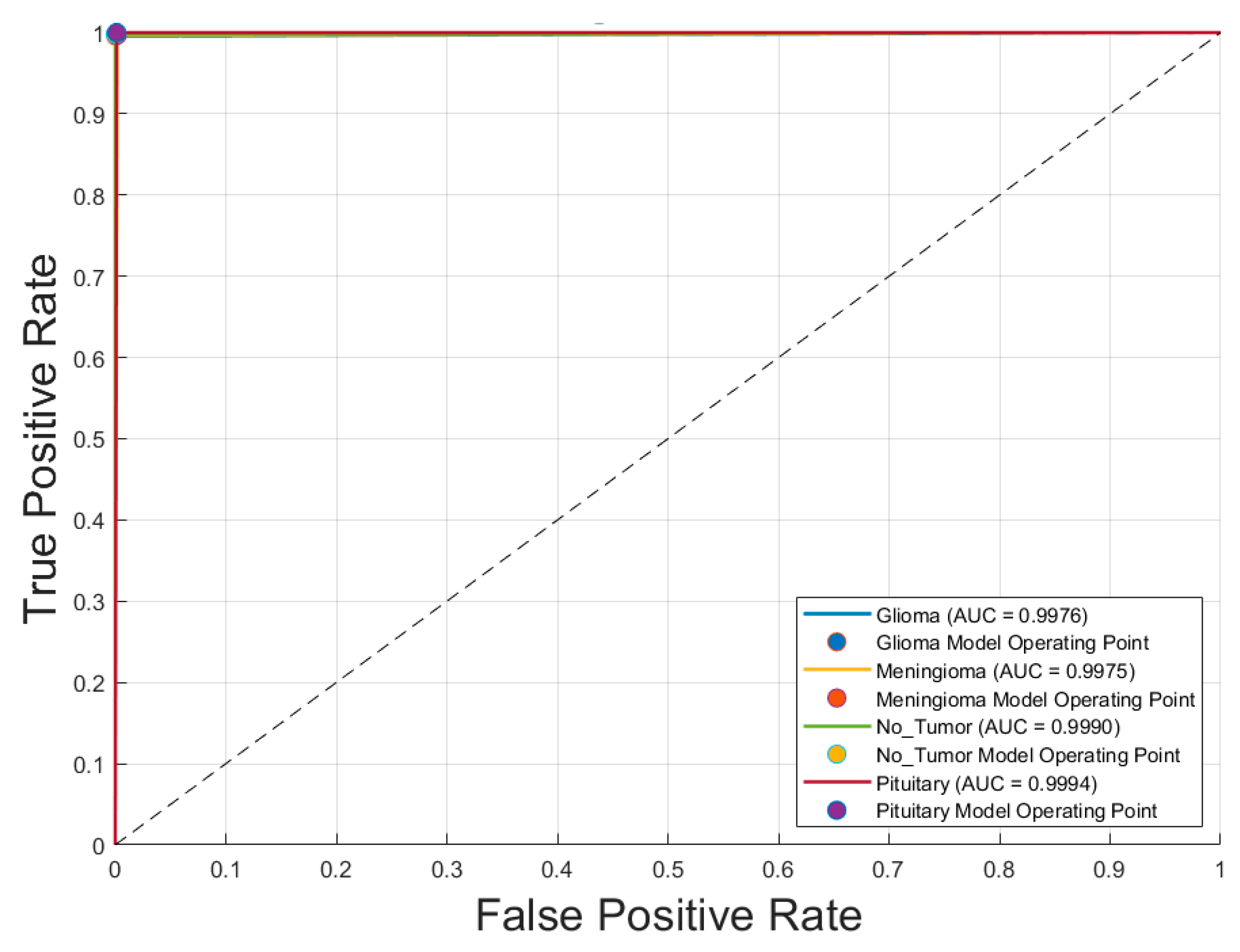
- Receiver Operating Characteristic (ROC) Curve: Plotted for each class to evaluate the model’s discriminative ability, specifically the trade-off between sensitivity (true positive rate) and specificity (1 − false positive rate).
4. Results
4.1. Comparative Classification Performance of CNN Models
- Optimizer choice significantly impacts model performance, with ADAM outperforming SGDM across all CNN architectures.
- Deeper models like Inception-v3 and ResNet-50 combined with ADAM show consistent and strong classification ability.
- Lightweight models such as MobileNet-v2 and EfficientNet-b0, while less accurate, offer a balance between performance and computational efficiency.
4.2. Confusion Matrix and ROC Analysis of the Optimal CNN Model
5. Discussion
5.1. Performance of Individual CNN Models
5.2. Advantage of the Voting Ensemble Schema
5.3. Evaluating the Proposed Method Against Related Works
6. Conclusions
6.1. Summary of Findings
6.2. Limitations and Future Work
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ait Amou M, Xia K, Kamhi S, Mouhafid M. A novel MRI diagnosis method for brain tumor classification based on CNN and Bayesian optimization. Healthcare (Basel). 2022 Mar 8;10(3):494. [CrossRef]
- Deepa S, Janet J, Sumathi S, Ananth JP. Hybrid optimization algorithm enabled deep learning approach brain tumor segmentation and classification using MRI. J Digit Imaging. 2023 Jun;36(3):847-868. [CrossRef]
- AlTahhan FE, Khouqeer GA, Saadi S, Elgarayhi A, Sallah M. Refined automatic brain tumor classification using hybrid convolutional neural networks for MRI scans. Diagnostics (Basel). 2023 Feb 23;13(5):864. [CrossRef]
- Gupta I, Singh S, Gupta S, Ranjan Nayak S. Classification of brain tumours in MRI images using a convolutional neural network. Curr Med Imaging. 2024;20:e270323214998. [CrossRef]
- Albalawi E, Thakur A, Dorai DR, Bhatia Khan S, Mahesh TR, Almusharraf A, Aurangzeb K, Anwar MS. Enhancing brain tumor classification in MRI scans with a multi-layer customized convolutional neural network approach. Front Comput Neurosci. 2024 Jun 12;18:1418546. [CrossRef]
- Nahiduzzaman M, Abdulrazak LF, Kibria HB, Khandakar A, Ayari MA, Ahamed MF, Ahsan M, Haider J, Moni MA, Kowalski M. A hybrid explainable model based on advanced machine learning and deep learning models for classifying brain tumors using MRI images. Sci Rep. 2025 Jan 10;15(1):1649. [CrossRef]
- Hassan E, Ghadiri H. Advancing brain tumor classification: A robust framework using EfficientNetV2 transfer learning and statistical analysis. Comput Biol Med. 2025 Feb;185:109542. [CrossRef]
- Iqbal S, Qureshi AN, Alhussein M, Aurangzeb K, Choudhry IA, Anwar MS. Hybrid deep spatial and statistical feature fusion for accurate MRI brain tumor classification. Front Comput Neurosci. 2024 Jun 24;18:1423051. [CrossRef]
- El Amoury S, Smili Y, Fakhri Y. Design of an optimal convolutional neural network architecture for MRI brain tumor classification by exploiting particle swarm optimization. J Imaging. 2025 Jan 24;11(2):31. [CrossRef]
- Kusuma PV, Reddy SCM. Brain tumor segmentation and classification using MRI: Modified segnet model and hybrid deep learning architecture with improved texture features. Comput Biol Chem. 2025 Aug;117:108381. [CrossRef]
- Huang KA, Alkadri A, Prakash N. Employing squeeze-and-excitation architecture in a fine-tuned convolutional neural network for magnetic resonance imaging tumor classification. Cureus. 2025 Mar 5;17(3):e80084. [CrossRef]
- Jarria SPA, Wesley AB. Hybrid fruit bee optimization algorithm-based deep convolution neural network for brain tumour classification using MRI images. Network. 2025 Mar 28:1-23. [CrossRef]
- da Costa Nascimento JJ, Marques AG, do Nascimento Souza L, de Mattos Dourado Junior CMJ, da Silva Barros AC, de Albuquerque VHC, de Freitas Sousa LF. A novel generative model for brain tumor detection using magnetic resonance imaging. Comput Med Imaging Graph. 2025 Apr;121:102498. [CrossRef]
- Chandraprabha K, Ganesan L, Baskaran K. A novel approach for the detection of brain tumor and its classification via end-to-end vision transformer - CNN architecture. Front Oncol. 2025 Mar 10;15:1508451. [CrossRef]
- Disci R, Gurcan F, Soylu A. Advanced brain tumor classification in MR images using transfer learning and pre-trained deep CNN models. Cancers (Basel). 2025 Jan 2;17(1):121. [CrossRef]
- Afzal S, Rauf M, Ashraf S, Bin Md Ayob S, Ahmad Arfeen Z. CART-ANOVA-based transfer learning approach for seven distinct tumor classification schemes with generalization capability. Diagnostics (Basel). 2025 Feb 5;15(3):378. [CrossRef]
- Elhadidy MS, Elgohr AT, El-Geneedy M, Akram S, Kasem HM. Comparative analysis for accurate multi-classification of brain tumor based on significant deep learning models. Comput Biol Med. 2025 Apr;188:109872. [CrossRef]
- Ali RR, Yaacob NM, Alqaryouti MH, Sadeq AE, Doheir M, Iqtait M, Rachmawanto EH, Sari CA, Yaacob SS. Learning architecture for brain tumor classification based on deep convolutional neural network: Classic and ResNet50. Diagnostics (Basel). 2025 Mar 5;15(5):624. [CrossRef]
- Hsu WW, Guo JM, Pei L, Chiang LA, Li YF, Hsiao JC, Colen R, Liu P. A weakly supervised deep learning-based method for glioma subtype classification using WSI and mpMRIs. Sci Rep. 2022 Apr 12;12(1):6111. [CrossRef]
- Özkaraca O, Bağrıaçık Oİ, Gürüler H, Khan F, Hussain J, Khan J, Laila UE. Multiple brain tumor classification with dense CNN architecture using brain MRI images. Life (Basel). 2023 Jan 28;13(2):349. [CrossRef]
- Abirami S, Ramesh K, Lalitha VaniSree K. Classification and pixel change detection of brain tumor using Adam Kooka-burra optimization-based Shepard convolutional neural network. NMR Biomed. 2025 Feb;38(2):e5307. [CrossRef]
- Mao Y, Kim J, Podina L, Kohandel M. Dilated SE-DenseNet for brain tumor MRI classification. Sci Rep. 2025 Jan 28;15(1):3596. [CrossRef]
- Aurna NF, Yousuf MA, Taher KA, Azad AKM, Moni MA. A classification of MRI brain tumor based on two stage feature level ensemble of deep CNN models. Comput Biol Med. 2022 Jul;146:105539. [CrossRef]
- Alsubai S, Khan HU, Alqahtani A, Sha M, Abbas S, Mohammad UG. Ensemble deep learning for brain tumor detection. Front Comput Neurosci. 2022 Sep 2;16:1005617. [CrossRef]
- Al-Azzwi ZHN, Nazarov AN. Brain tumor classification based on improved stacked ensemble deep learning methods. Asian Pac J Cancer Prev. 2023 Jun 1;24(6):2141-2148. [CrossRef]
- Tandel GS, Tiwari A, Kakde OG. Multi-class brain tumor grades classification using a deep learning-based majority voting algorithm and its validation using explainable-AI. J Imaging Inform Med. 2025 Jan 8. [CrossRef]
- He K, Zhang X, Ren S, Sun J. Deep residual learning for image recognition. In: 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2016 Jun 27–30; Las Vegas, NV, USA. IEEE; 2016. p. 770–8. [CrossRef]
- Szegedy C, Liu W, Jia Y, Sermanet P, Reed S, Anguelov D, Erhan D, Vanhoucke V, Rabinovich A. Going deeper with convolutions. In: 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2015 Jun 7–12; Boston, MA, USA. IEEE; 2015. p. 1–9. [CrossRef]
- Tan M, Le QV. EfficientNet: rethinking model scaling for convolutional neural networks. In: Proceedings of the 36th International Conference on Machine Learning (ICML); 2019 Jun 9–15; Long Beach, CA, USA. PMLR; 2019. arXiv:1905.11946 [cs.LG]. [CrossRef]
- Sandler M, Howard A, Zhu M, Zhmoginov A, Chen LC. MobileNetV2: inverted residuals and linear bottlenecks. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2018 Jun 18–23; Salt Lake City, UT, USA. IEEE; 2018. p. 4510–4520. arXiv:1801.04381 [cs.CV]. [CrossRef]
- Szegedy C, Vanhoucke V, Ioffe S, Shlens J, Wojna Z. Rethinking the Inception architecture for computer vision. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR); 2016 Jun 27–30; Las Vegas, NV, USA. IEEE; 2016. p. 2818–2826. arXiv:1512.00567 [cs.CV]. [CrossRef]
| CNN | No. of Layers | Parameters (MB) | Default Input Size | Key Advantages |
|---|---|---|---|---|
| ResNet-18 [27] | 18 | 11.7 | 224 × 224 | Lightweight, fast training, good for small datasets |
| GoogLeNet [28] | 22 | 7.0 | 224 × 224 | Inception modules for multi-scale feature extraction |
| EfficientNet-b0 [29] | 290 | 5.3 | 224 × 224 | Parameter-efficient, high accuracy with fewer resources |
| MobileNet-v2 [30] | 154 | 3.5 | 224 × 224 | Optimized for speed and mobile deployment |
| Inception-v3 [31] | 315 | 23.9 | 299 × 299 | High accuracy with reduced computation |
| ResNet-50 [27] | 177 | 25.6 | 224 × 224 | Deeper network with residual connections for feature reuse |
| ResNet-101 [27] | 347 | 44.6 | 224 × 224 | Strong performance on complex tasks due to depth |
| CNN | Optimizer | TP1 | TP2 | TP3 | TP4 | Pre1 | Pre2 | Pre3 | Pre4 | Accuracy | Kappa |
|---|---|---|---|---|---|---|---|---|---|---|---|
| ResNet-18 | SGDM | 0.986 | 0.964 | 0.970 | 0.976 | 0.960 | 0.966 | 0.980 | 0.995 | 0.974 | 0.965 |
| ResNet-18 | ADAM | 0.983 | 0.983 | 0.982 | 0.988 | 0.980 | 0.981 | 0.980 | 0.995 | 0.984 | 0.978 |
| GoogLeNet | SGDM | 0.997 | 0.974 | 0.992 | 0.990 | 0.976 | 0.993 | 0.984 | 0.996 | 0.987 | 0.983 |
| GoogLeNet | ADAM | 0.953 | 0.974 | 0.972 | 0.986 | 0.980 | 0.971 | 0.912 | 0.995 | 0.971 | 0.960 |
| EfficientNet-b0 | SGDM | 0.900 | 0.898 | 0.933 | 0.917 | 0.897 | 0.868 | 0.918 | 0.960 | 0.910 | 0.877 |
| EfficientNet-b0 | ADAM | 0.959 | 0.951 | 0.984 | 0.967 | 0.949 | 0.940 | 0.970 | 0.996 | 0.963 | 0.949 |
| MobileNet-v2 | SGDM | 0.910 | 0.905 | 0.924 | 0.925 | 0.905 | 0.859 | 0.920 | 0.981 | 0.915 | 0.885 |
| MobileNet-v2 | ADAM | 0.965 | 0.967 | 0.963 | 0.954 | 0.950 | 0.935 | 0.978 | 0.993 | 0.962 | 0.949 |
| Inception-v3 | SGDM | 0.967 | 0.931 | 0.926 | 0.962 | 0.934 | 0.923 | 0.950 | 0.990 | 0.949 | 0.931 |
| Inception-v3 | ADAM | 0.991 | 0.975 | 0.994 | 0.993 | 0.978 | 0.986 | 0.990 | 0.997 | 0.987 | 0.983 |
| ResNet-50 | SGDM | 0.939 | 0.893 | 0.929 | 0.888 | 0.887 | 0.851 | 0.912 | 0.991 | 0.909 | 0.877 |
| ResNet-50 | ADAM | 0.987 | 0.969 | 0.974 | 0.983 | 0.977 | 0.968 | 0.986 | 0.988 | 0.979 | 0.971 |
| ResNet-101 | SGDM | 0.932 | 0.897 | 0.877 | 0.897 | 0.879 | 0.818 | 0.954 | 0.988 | 0.903 | 0.869 |
| ResNet-101 | ADAM | 0.983 | 0.966 | 0.974 | 0.985 | 0.959 | 0.969 | 0.990 | 0.998 | 0.977 | 0.969 |
| Voting | 0.996 | 0.997 | 0.998 | 1.000 | 0.999 | 0.996 | 1.000 | 0.997 | 0.998 | 0.997 |
| Class | Glioma | Meningioma | No Tumor | Pituitary | TP | FP |
|---|---|---|---|---|---|---|
| Glioma | 904 | 18 | 3 | 1 | 0.976 | 0.024 |
| Meningioma | 2 | 927 | 0 | 5 | 0.993 | 0.007 |
| No Tumor | 1 | 4 | 492 | 3 | 0.984 | 0.016 |
| Pituitary | 0 | 3 | 1 | 897 | 0.996 | 0.004 |
| Precision | 0.953 | 0.974 | 0.972 | 0.986 | Accuracy | 0.987 |
| FP | 0.047 | 0.026 | 0.028 | 0.014 | Kappa | 0.983 |
| CNN | Glioma | Meningioma | No Tumor | Pituitary | TP | FP |
|---|---|---|---|---|---|---|
| Glioma | 906 | 19 | 1 | 0 | 0.978 | 0.022 |
| Meningioma | 6 | 921 | 2 | 5 | 0.986 | 0.014 |
| No Tumor | 1 | 3 | 495 | 1 | 0.990 | 0.010 |
| Pituitary | 1 | 2 | 0 | 898 | 0.997 | 0.003 |
| Precision | 0.953 | 0.974 | 0.0972 | 0.986 | Accuracy | 0.987 |
| FP | 0.047 | 0.026 | 0.028 | 0.014 | Kappa | 0.983 |
| CNN | Glioma | Meningioma | No Tumor | Pituitary | TP | FP |
|---|---|---|---|---|---|---|
| Glioma | 922 | 4 | 0 | 0 | 0.996 | 0.004 |
| Meningioma | 0 | 931 | 0 | 3 | 0.997 | 0.003 |
| No Tumor | 1 | 0 | 499 | 0 | 0.998 | 0.002 |
| Pituitary | 0 | 0 | 0 | 901 | 1.000 | 0.000 |
| Precision | 0.953 | 0.974 | 0.0972 | 0.986 | Accuracy | 0.998 |
| FP | 0.047 | 0.026 | 0.028 | 0.014 | Kappa | 0.997 |
| Authors | Year | Method | Task | Classes | Accuracy |
|---|---|---|---|---|---|
| Ait Amou et al. [1] | 2022 | CNN + Bayesian Optimization | Classification | 3 | 98.70% |
| Deepa et al. [2] | 2023 | Hybrid Optimization + DRN | Segmentation & Classification | 3 | 92.10% |
| AlTahhan et al. [3] | 2023 | Hybrid AlexNet-KNN | Classification | 4 | 98.60% |
| Gupta et al. [4] | 2024 | Custom CNN | Classification | 2 | 94.00% |
| Albalawi et al. [5] | 2024 | Multi-layer CNN | Classification | 4 | 99.00% |
| Nahiduzzaman et al. [6] | 2025 | PDSCNN + RRELM | Classification | 4 | 99.20% |
| Hassan & Ghadiri [7] | 2025 | EfficientNetV2 | Classification | 3 | 99.16% |
| Iqbal et al. [8] | 2024 | FusionNet (Statistical + CNN) | Classification | 2 | 97.53% |
| El Amoury et al. [9] | 2025 | PSO-Optimized CNN | Classification | 4 | 99.20% |
| Kusuma & Reddy [10] | 2025 | SegNet + Bi-LSTM | Segmentation & Classification | 4 | 98.00% |
| Huang et al. [11] | 2025 | ResNet50V2 + SE Blocks | Classification | 4 | 98.40% |
| Jarria & Wesley [12] | 2025 | Fruit Bee Optimized CNN | Classification | 3 | 92.60% |
| da Costa Nascimento et al. [13] | 2025 | YOLO + LLM | Detection & Classification | 2 | 98.00% |
| Chandraprabha et al. [14] | 2025 | ViT + CNN | Classification | 4 | 99.64% |
| Disci et al. [15] | 2025 | Transfer Learning (Xception, etc.) | Classification | 4 | 98.73% |
| Afzal et al. [16] | 2025 | ResNet18 + CART-ANOVA | Classification | 4 | 98.05% |
| Elhadidy et al. [17] | 2025 | Swin Transformer + EfficientNet | Classification | 4 | 98.72% |
| Ali et al. [18] | 2025 | Classic CNN + ResNet50 | Classification | 3 | 99.88% |
| Abirami et al. [21] | 2025 | AKO-Shepard CNN | Classification & Detection | 3 | 93.60% |
| Aurna et al. [23] | 2022 | Two-Stage Ensemble CNN | Classification | 4 | 99.13% |
| Alsubai et al. [24] | 2022 | CNN-LSTM | Classification | 3 | 99.10% |
| Al-Azzwi & Nazarov [25] | 2023 | Stacked Ensemble (VGG19, etc.) | Classification | 2 | 96.60% |
| Tandel et al. [26] | 2025 | Majority Voting + XAI | Classification | 4 | 98.47% |
| This Study | 2025 | Majority Voting Ensemble | Classification | 4 | 99.80% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




