Submitted:
23 May 2025
Posted:
26 May 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patients
2.2. Genomic Data Collection and Single Nucleotide Polymorphism Genotyping
2.3. Immunohistochemistry Analysis
2.4. Exploration of Gene Expression Profiles
2.5. Clinical and Histological Data Collection
2.6. Statistical Analysis
3. Results
3.1. Baseline Characteristics
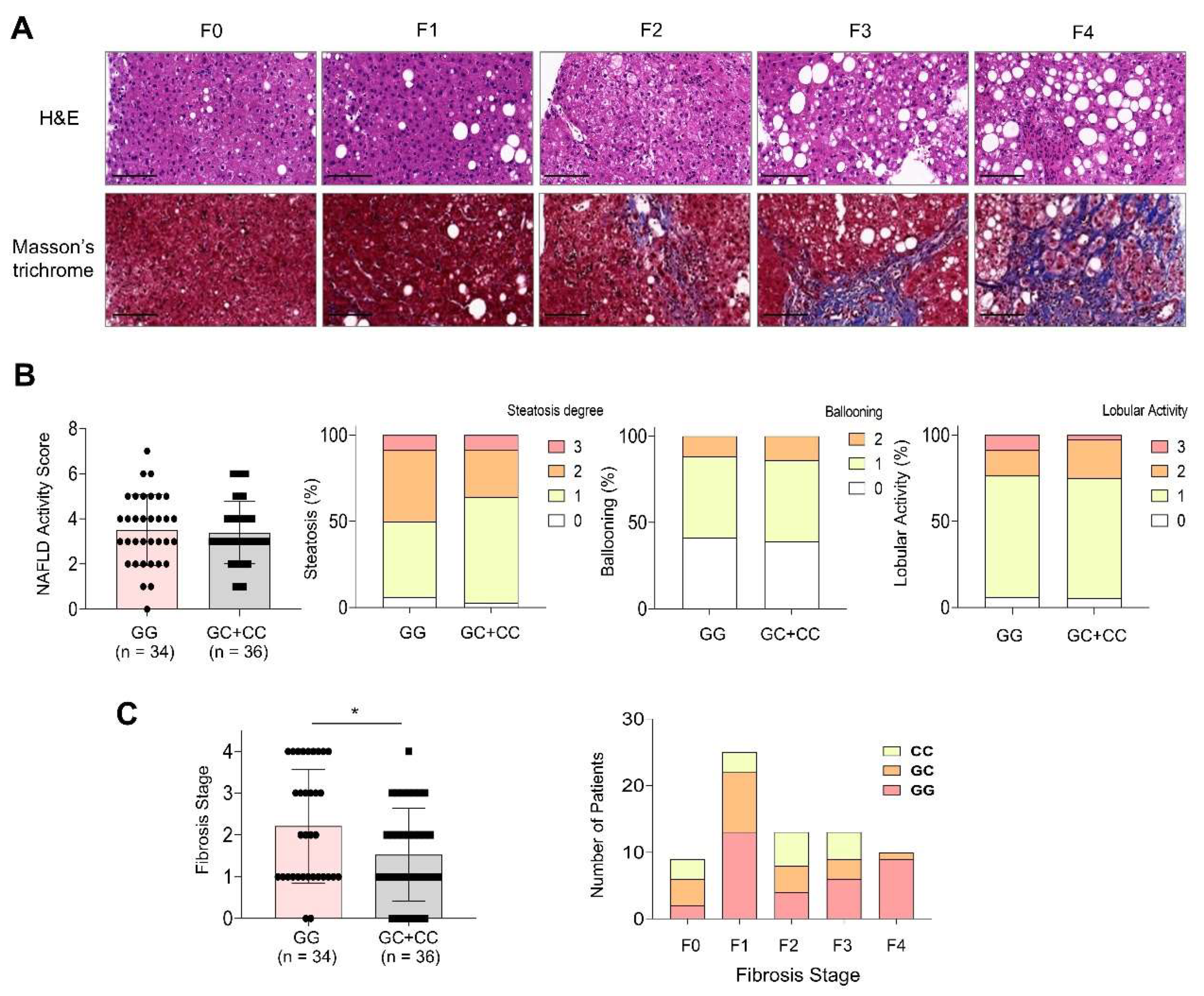
3.2. MASLD Severity According to the PNPLA3 I148M Genetic Variant
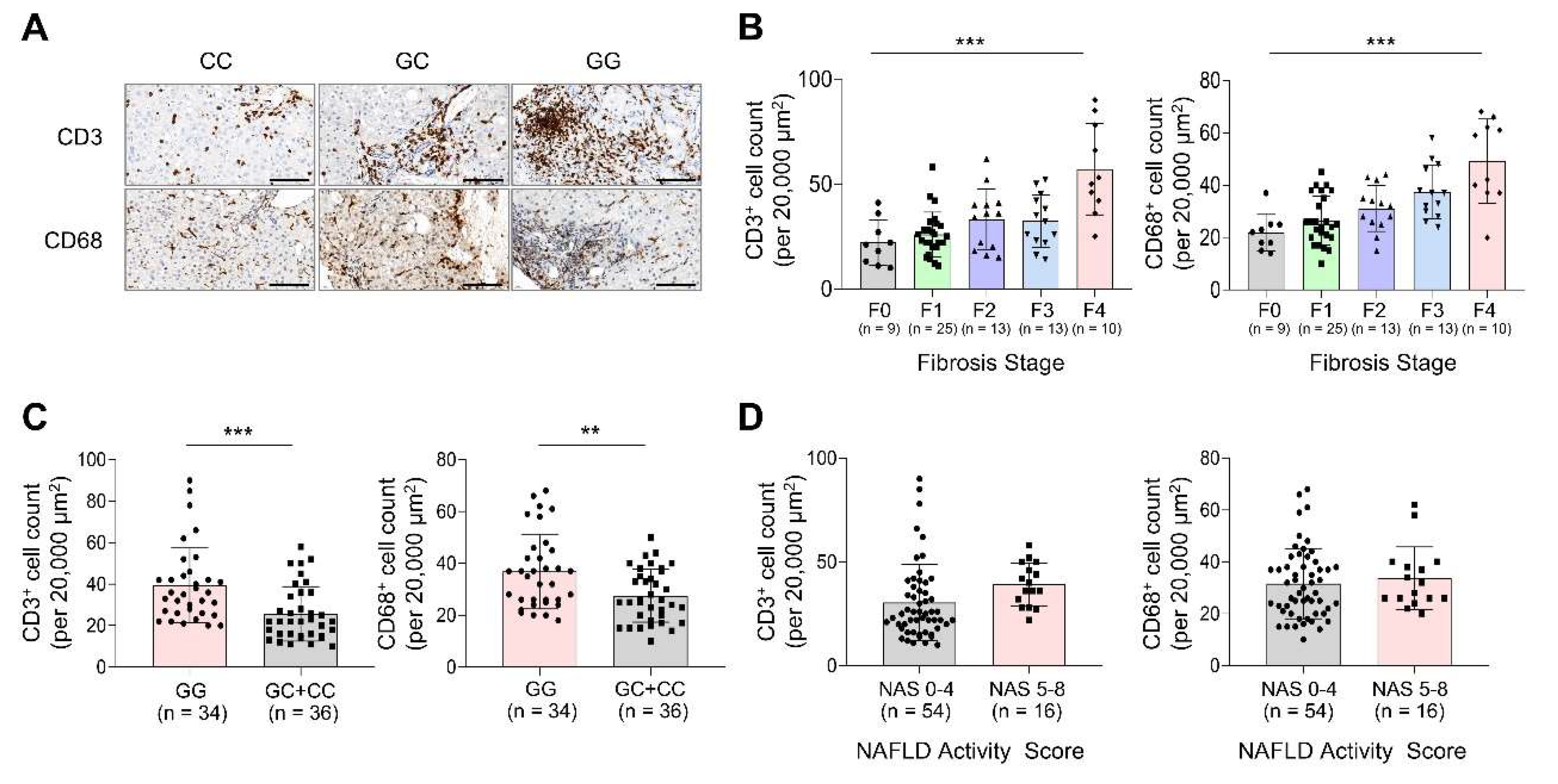
3.3. Differential Immune Cell Infiltration in MASLD Liver
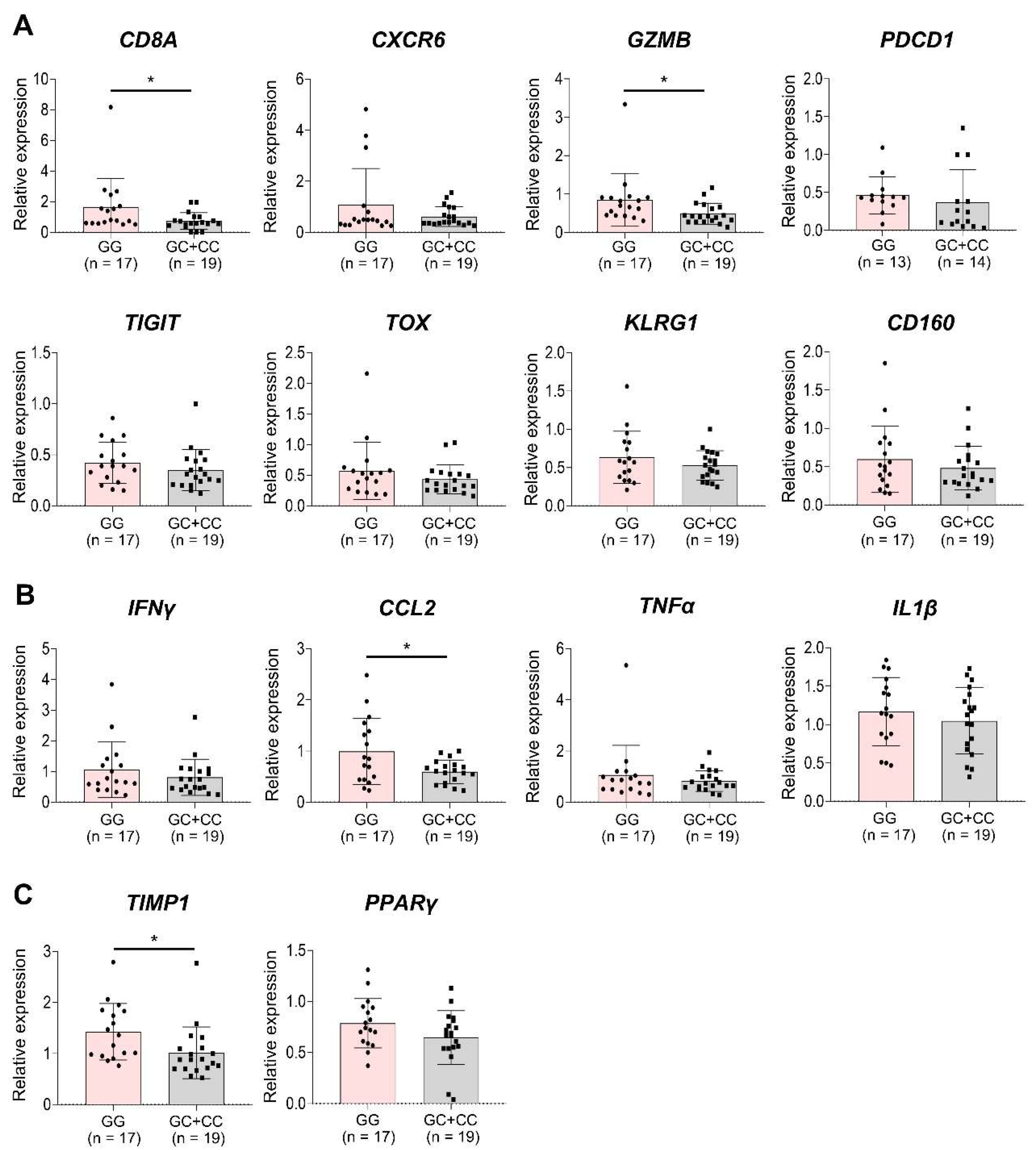
3.4. Association Between PNPLA3 I148M Genetic Variant and Immune-Related Gene Expression
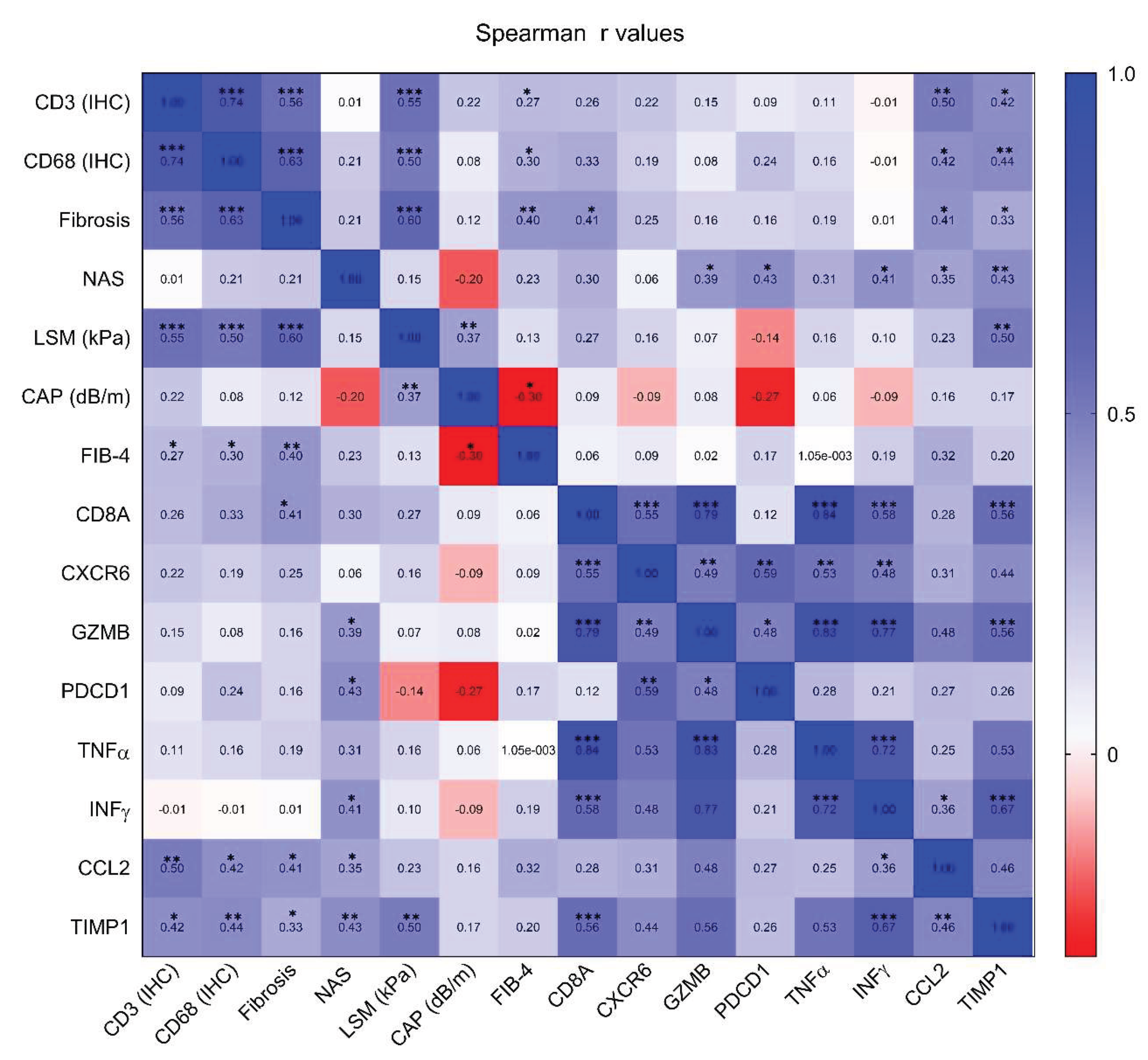
3.5. Correlation Between Immune-Related Markers and MASLD Severity
3.6. Sensitivity Analysis in MASLD with Early Fibrosis (F0–2)
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
References
- EASL-EASD-EASO Clinical Practice Guidelines on the Management of Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD). Obes Facts 2024, 17, 374–444. [CrossRef] [PubMed]
- Kim, D.Y. Changing etiology and epidemiology of hepatocellular carcinoma: Asia and worldwide. J Liver Cancer 2024, 24, 62–70. [Google Scholar] [CrossRef] [PubMed]
- Han, J.W.; Sohn, W.; Choi, G.H.; Jang, J.W.; Seo, G.H.; Kim, B.H.; Choi, J.Y. Evolving trends in treatment patterns for hepatocellular carcinoma in Korea from 2008 to 2022: a nationwide population-based study. J Liver Cancer 2024, 24, 274–285. [Google Scholar] [CrossRef] [PubMed]
- Esler, W.P.; Cohen, D.E. Pharmacologic inhibition of lipogenesis for the treatment of NAFLD. J Hepatol 2024, 80, 362–377. [Google Scholar] [CrossRef]
- Iwaki, M.; Fujii, H.; Hayashi, H.; Toyoda, H.; Oeda, S.; Hyogo, H.; Kawanaka, M.; Morishita, A.; Munekage, K.; Kawata, K.; et al. Prognosis of biopsy-confirmed metabolic dysfunction- associated steatotic liver disease: A sub-analysis of the CLIONE study. Clin Mol Hepatol 2024, 30, 225–234. [Google Scholar] [CrossRef]
- Harrison, S.A.; Bedossa, P.; Guy, C.D.; Schattenberg, J.M.; Loomba, R.; Taub, R.; Labriola, D.; Moussa, S.E.; Neff, G.W.; Rinella, M.E.; et al. A Phase 3, Randomized, Controlled Trial of Resmetirom in NASH with Liver Fibrosis. N Engl J Med 2024, 390, 497–509. [Google Scholar] [CrossRef]
- An, J.; Sohn, J.H. Pharmacological advances in the treatment of nonalcoholic fatty liver diseases : focused on global results of randomized controlled trials. Clin Mol Hepatol 2023, 29, S268–s275. [Google Scholar] [CrossRef]
- Puengel, T.; Tacke, F. Pharmacotherapeutic options for metabolic dysfunction-associated steatotic liver disease: where are we today? Expert Opin Pharmacother 2024, 25, 1249–1263. [Google Scholar] [CrossRef]
- Johnson, S.M.; Bao, H.; McMahon, C.E.; Chen, Y.; Burr, S.D.; Anderson, A.M.; Madeyski-Bengtson, K.; Lindén, D.; Han, X.; Liu, J. PNPLA3 is a triglyceride lipase that mobilizes polyunsaturated fatty acids to facilitate hepatic secretion of large-sized very low-density lipoprotein. Nat Commun 2024, 15, 4847–s41467. [Google Scholar] [CrossRef]
- Valenti, L.; Al-Serri, A.; Daly, A.K.; Galmozzi, E.; Rametta, R.; Dongiovanni, P.; Nobili, V.; Mozzi, E.; Roviaro, G.; Vanni, E.; et al. Homozygosity for the patatin-like phospholipase-3/adiponutrin I148M polymorphism influences liver fibrosis in patients with nonalcoholic fatty liver disease. Hepatology 2010, 51, 1209–1217. [Google Scholar] [CrossRef]
- Liu, Y.L.; Patman, G.L.; Leathart, J.B.; Piguet, A.C.; Burt, A.D.; Dufour, J.F.; Day, C.P.; Daly, A.K.; Reeves, H.L.; Anstee, Q.M. Carriage of the PNPLA3 rs738409 C >G polymorphism confers an increased risk of non-alcoholic fatty liver disease associated hepatocellular carcinoma. J Hepatol 2014, 61, 75–81. [Google Scholar] [CrossRef] [PubMed]
- Rosso, C.; Caviglia, G.P.; Birolo, G.; Armandi, A.; Pennisi, G.; Pelusi, S.; Younes, R.; Liguori, A.; Perez-Diaz-Del-Campo, N.; Nicolosi, A.; et al. Impact of PNPLA3 rs738409 Polymorphism on the Development of Liver-Related Events in Patients With Nonalcoholic Fatty Liver Disease. Clin Gastroenterol Hepatol 2023, 21, 3314–3321.e3313. [Google Scholar] [CrossRef] [PubMed]
- Petta, S.; Armandi, A.; Bugianesi, E. Impact of PNPLA3 I148M on Clinical Outcomes in Patients With MASLD. Liver Int 2024. [Google Scholar] [CrossRef]
- Sookoian, S.; Pirola, C.J. Genetics in non-alcoholic fatty liver disease: The role of risk alleles through the lens of immune response. Clin Mol Hepatol 2023, 29, S184–s195. [Google Scholar] [CrossRef] [PubMed]
- BasuRay, S.; Smagris, E.; Cohen, J.C.; Hobbs, H.H. The PNPLA3 variant associated with fatty liver disease (I148M) accumulates on lipid droplets by evading ubiquitylation. Hepatology 2017, 66, 1111–1124. [Google Scholar] [CrossRef]
- Vilar-Gomez, E.; Pirola, C.J.; Sookoian, S.; Wilson, L.A.; Liang, T.; Chalasani, N. PNPLA3 rs738409 and risk of fibrosis in NAFLD: Exploring mediation pathways through intermediate histological features. Hepatology 2022, 76, 1482–1494. [Google Scholar] [CrossRef]
- Li, Z.; Wang, S.; Xu, Q.; Su, X.; Wang, Y.; Wang, L.; Zhang, Y. The double roles of T cell-mediated immune response in the progression of MASLD. Biomed Pharmacother 2024, 173, 116333. [Google Scholar] [CrossRef]
- Huby, T.; Gautier, E.L. Immune cell-mediated features of non-alcoholic steatohepatitis. Nat Rev Immunol 2022, 22, 429–443. [Google Scholar] [CrossRef]
- Mao, T.; Yang, R.; Luo, Y.; He, K. Crucial role of T cells in NAFLD-related disease: A review and prospect. Front Endocrinol (Lausanne) 2022, 13, 1051076. [Google Scholar] [CrossRef]
- Seth, R.K.; Das, S.; Kumar, A.; Chanda, A.; Kadiiska, M.B.; Michelotti, G.; Manautou, J.; Diehl, A.M.; Chatterjee, S. CYP2E1-dependent and leptin-mediated hepatic CD57 expression on CD8+ T cells aid progression of environment-linked nonalcoholic steatohepatitis. Toxicol Appl Pharmacol 2014, 274, 42–54. [Google Scholar] [CrossRef]
- Kazankov, K.; Jørgensen, S.M.D.; Thomsen, K.L.; Møller, H.J.; Vilstrup, H.; George, J.; Schuppan, D.; Grønbæk, H. The role of macrophages in nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Nat Rev Gastroenterol Hepatol 2019, 16, 145–159. [Google Scholar] [CrossRef] [PubMed]
- Sung, P.S. Crosstalk between tumor-associated macrophages and neighboring cells in hepatocellular carcinoma. Clin Mol Hepatol 2022, 28, 333–350. [Google Scholar] [CrossRef]
- Sung, P.S.; Park, D.J.; Roh, P.R.; Mun, K.D.; Cho, S.W.; Lee, G.W.; Jung, E.S.; Lee, S.H.; Jang, J.W.; Bae, S.H.; et al. Intrahepatic inflammatory IgA(+)PD-L1(high) monocytes in hepatocellular carcinoma development and immunotherapy. J Immunother Cancer 2022, 10. [Google Scholar] [CrossRef] [PubMed]
- Carpino, G.; Del Ben, M.; Pastori, D.; Carnevale, R.; Baratta, F.; Overi, D.; Francis, H.; Cardinale, V.; Onori, P.; Safarikia, S.; et al. Increased Liver Localization of Lipopolysaccharides in Human and Experimental NAFLD. Hepatology 2020, 72, 470–485. [Google Scholar] [CrossRef]
- Krenkel, O.; Tacke, F. Liver macrophages in tissue homeostasis and disease. Nat Rev Immunol 2017, 17, 306–321. [Google Scholar] [CrossRef]
- EASL-EASD-EASO Clinical Practice Guidelines on the management of metabolic dysfunction-associated steatotic liver disease (MASLD). J Hepatol 2024, 81, 492–542. [CrossRef]
- Cha, J.H.; Park, N.R.; Cho, S.W.; Nam, H.; Yang, H.; Jung, E.S.; Jang, J.W.; Choi, J.Y.; Yoon, S.K.; Sung, P.S.; et al. Chitinase 1: a novel therapeutic target in metabolic dysfunction-associated steatohepatitis. Front Immunol 2024, 15, 1444100. [Google Scholar] [CrossRef] [PubMed]
- Sterling, R.K.; Lissen, E.; Clumeck, N.; Sola, R.; Correa, M.C.; Montaner, J.; M, S.S.; Torriani, F.J.; Dieterich, D.T.; Thomas, D.L.; et al. Development of a simple noninvasive index to predict significant fibrosis in patients with HIV/HCV coinfection. Hepatology 2006, 43, 1317–1325. [Google Scholar] [CrossRef]
- Brunt, E.M.; Kleiner, D.E.; Wilson, L.A.; Belt, P.; Neuschwander-Tetri, B.A. Nonalcoholic fatty liver disease (NAFLD) activity score and the histopathologic diagnosis in NAFLD: distinct clinicopathologic meanings. Hepatology 2011, 53, 810–820. [Google Scholar] [CrossRef]
- Kleiner, D.E.; Brunt, E.M.; Van Natta, M.; Behling, C.; Contos, M.J.; Cummings, O.W.; Ferrell, L.D.; Liu, Y.C.; Torbenson, M.S.; Unalp-Arida, A.; et al. Design and validation of a histological scoring system for nonalcoholic fatty liver disease. Hepatology 2005, 41, 1313–1321. [Google Scholar] [CrossRef]
- Xu, R.; Tao, A.; Zhang, S.; Deng, Y.; Chen, G. Association between patatin-like phospholipase domain containing 3 gene (PNPLA3) polymorphisms and nonalcoholic fatty liver disease: a HuGE review and meta-analysis. Sci Rep 2015, 5, 9284. [Google Scholar] [CrossRef] [PubMed]
- Armisen, J.; Rauschecker, M.; Sarv, J.; Liljeblad, M.; Wernevik, L.; Niazi, M.; Knöchel, J.; Eklund, O.; Sandell, T.; Sherwood, J.; et al. AZD2693, a PNPLA3 antisense oligonucleotide, for the treatment of MASH in 148M homozygous participants: two randomized phase I trials. J Hepatol 2025. [Google Scholar] [CrossRef]
- Boeckmans, J.; Gatzios, A.; Schattenberg, J.M.; Koek, G.H.; Rodrigues, R.M.; Vanhaecke, T. PNPLA3 I148M and response to treatment for hepatic steatosis: A systematic review. Liver Int 2023, 43, 975–988. [Google Scholar] [CrossRef] [PubMed]
- Park, J.; Zhao, Y.; Zhang, F.; Zhang, S.; Kwong, A.C.; Zhang, Y.; Hoffmann, H.H.; Bushweller, L.; Wu, X.; Ashbrook, A.W.; et al. IL-6/STAT3 axis dictates the PNPLA3-mediated susceptibility to non-alcoholic fatty liver disease. J Hepatol 2023, 78, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Sutti, S.; Albano, E. Adaptive immunity: an emerging player in the progression of NAFLD. Nat Rev Gastroenterol Hepatol 2020, 17, 81–92. [Google Scholar] [CrossRef]
- Yeh, M.M.; Brunt, E.M. Pathological features of fatty liver disease. Gastroenterology 2014, 147, 754–764. [Google Scholar] [CrossRef]
- Bruzzì, S.; Sutti, S.; Giudici, G.; Burlone, M.E.; Ramavath, N.N.; Toscani, A.; Bozzola, C.; Schneider, P.; Morello, E.; Parola, M.; et al. B2-Lymphocyte responses to oxidative stress-derived antigens contribute to the evolution of nonalcoholic fatty liver disease (NAFLD). Free Radic Biol Med 2018, 124, 249–259. [Google Scholar] [CrossRef]
- Grohmann, M.; Wiede, F.; Dodd, G.T.; Gurzov, E.N.; Ooi, G.J.; Butt, T.; Rasmiena, A.A.; Kaur, S.; Gulati, T.; Goh, P.K.; et al. Obesity Drives STAT-1-Dependent NASH and STAT-3-Dependent HCC. Cell 2018, 175, 1289–1306.e1220. [Google Scholar] [CrossRef]
- Ghazarian, M.; Revelo, X.S.; Nøhr, M.K.; Luck, H.; Zeng, K.; Lei, H.; Tsai, S.; Schroer, S.A.; Park, Y.J.; Chng, M.H.Y.; et al. Type I Interferon Responses Drive Intrahepatic T cells to Promote Metabolic Syndrome. Sci Immunol 2017, 2. [Google Scholar] [CrossRef]
- Sutti, S.; Jindal, A.; Locatelli, I.; Vacchiano, M.; Gigliotti, L.; Bozzola, C.; Albano, E. Adaptive immune responses triggered by oxidative stress contribute to hepatic inflammation in NASH. Hepatology 2014, 59, 886–897. [Google Scholar] [CrossRef]
- Breuer, D.A.; Pacheco, M.C.; Washington, M.K.; Montgomery, S.A.; Hasty, A.H.; Kennedy, A.J. CD8(+) T cells regulate liver injury in obesity-related nonalcoholic fatty liver disease. Am J Physiol Gastrointest Liver Physiol 2020, 318, G211–g224. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharjee, J.; Kirby, M.; Softic, S.; Miles, L.; Salazar-Gonzalez, R.M.; Shivakumar, P.; Kohli, R. Hepatic Natural Killer T-cell and CD8+ T-cell Signatures in Mice with Nonalcoholic Steatohepatitis. Hepatol Commun 2017, 1, 299–310. [Google Scholar] [CrossRef] [PubMed]
- Wolf, M.J.; Adili, A.; Piotrowitz, K.; Abdullah, Z.; Boege, Y.; Stemmer, K.; Ringelhan, M.; Simonavicius, N.; Egger, M.; Wohlleber, D.; et al. Metabolic activation of intrahepatic CD8+ T cells and NKT cells causes nonalcoholic steatohepatitis and liver cancer via cross-talk with hepatocytes. Cancer Cell 2014, 26, 549–564. [Google Scholar] [CrossRef] [PubMed]
- Eickmeier, I.; Seidel, D.; Grün, J.R.; Derkow, K.; Lehnardt, S.; Kühl, A.A.; Hamann, A.; Schott, E. Influence of CD8 T cell priming in liver and gut on the enterohepatic circulation. J Hepatol 2014, 60, 1143–1150. [Google Scholar] [CrossRef]
- Dudek, M.; Pfister, D.; Donakonda, S.; Filpe, P.; Schneider, A.; Laschinger, M.; Hartmann, D.; Hüser, N.; Meiser, P.; Bayerl, F.; et al. Auto-aggressive CXCR6(+) CD8 T cells cause liver immune pathology in NASH. Nature 2021, 592, 444–449. [Google Scholar] [CrossRef]
- Burtis, A.E.C.; DeNicola, D.M.C.; Ferguson, M.E.; Santos, R.G.; Pinilla, C.; Kriss, M.S.; Orlicky, D.J.; Tamburini, B.A.J.; Gillen, A.E.; Burchill, M.A. Ag-driven CD8 + T cell clonal expansion is a prominent feature of MASH in humans and mice. Hepatology 2025, 81, 591–608. [Google Scholar] [CrossRef]
- Sekine, T.; Perez-Potti, A.; Nguyen, S.; Gorin, J.B.; Wu, V.H.; Gostick, E.; Llewellyn-Lacey, S.; Hammer, Q.; Falck-Jones, S.; Vangeti, S.; et al. TOX is expressed by exhausted and polyfunctional human effector memory CD8(+) T cells. Sci Immunol 2020, 5. [Google Scholar] [CrossRef]
| Total (n = 70) |
PNPLA3 GG (n = 34) |
PNPLA3 GC+CC (n = 36) |
P | |
|---|---|---|---|---|
| Male sex | 36 (51.4) | 15 (44.1) | 21 (58.3) | 0.342 |
| Age | 51.3±17.4 | 53.4±15.1 | 49.3±19.2 | 0.323 |
| BMI (Kg/m2) | 28.3±6.0 | 28.0±6.5 | 28.5±5.5 | 0.743 |
| HTN | 28 (40.0) | 16 (47.1) | 12 (33.3) | 0.354 |
| DM | 31 (44.3) | 16 (47.1) | 15 (41.7) | 0.831 |
| WBC (109/L) | 7.3±2.4 | 6.6±2.2 | 7.9±2.4 | 0.025 |
| Hb (g/dL) | 14.2±1.9 | 13.9±1.9 | 14.4±2.0 | 0.372 |
| PLT (103/µL) | 236.2±80.7 | 219.4±80.7 | 252.1±78.6 | 0.090 |
| PT (INR) | 1.1±0.1 | 1.1±0.1 | 1.0±0.1 | 0.055 |
| Albumin (g/dL) | 4.4±0.4 | 4.3±0.4 | 4.5±0.4 | 0.020 |
| TB (mg/dL) | 0.8±0.5 | 0.9±0.6 | 0.8±0.4 | 0.310 |
| AST (IU/L) | 65.1±38.6 | 64.3±41.3 | 65.9±36.4 | 0.864 |
| ALT (IU/L) | 105.0±155.9 | 82.0±73.6 | 126.8±204.5 | 0.232 |
| GGT (U/L) | 93.4±93.2 | 85.4±69.3 | 101.0±111.8 | 0.486 |
| Triglycerides (mg/dL) | 161.3±117.8 | 140.0±71.3 | 182.0±148.2 | 0.140 |
| Total Cholesterol (mg/dL) | 186.7±43.3 | 178.4±38.7 | 194.8±46.5 | 0.116 |
| LDL-C (mg/dL) | 110.9±38.0 | 106.7±36.9 | 115.1±39.1 | 0.364 |
| HDL-C (mg/dL) | 47.1±11.4 | 47.2±11.6 | 47.0±11.3 | 0.932 |
| NAS ≥ 5 | 16 (22.9) | 9 (26.5) | 7 (19.4) | 0.678 |
| Fibrosis stage | 0.025 | |||
| F0 | 9 (12.9) | 2 (5.9) | 7 (19.4) | |
| F1 | 25 (35.7) | 13 (38.2) | 12 (33.3) | |
| F2 | 13 (18.6) | 4 (11.8) | 9 (25.0) | |
| F3 | 13 (18.6) | 6 (17.6) | 7 (19.4) | |
| F4 | 10 (14.3) | 9 (26.5) | 1 (2.8) | |
| CAP (dB/m) | 299.5±37.6 | 306.3±38.6 | 294.1±36.4 | 0.228 |
| LSM (kPa) | 11.2±6.4 | 12.7±7.8 | 10.0±4.9 | 0.115 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




