Submitted:
12 May 2025
Posted:
12 May 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Related Work
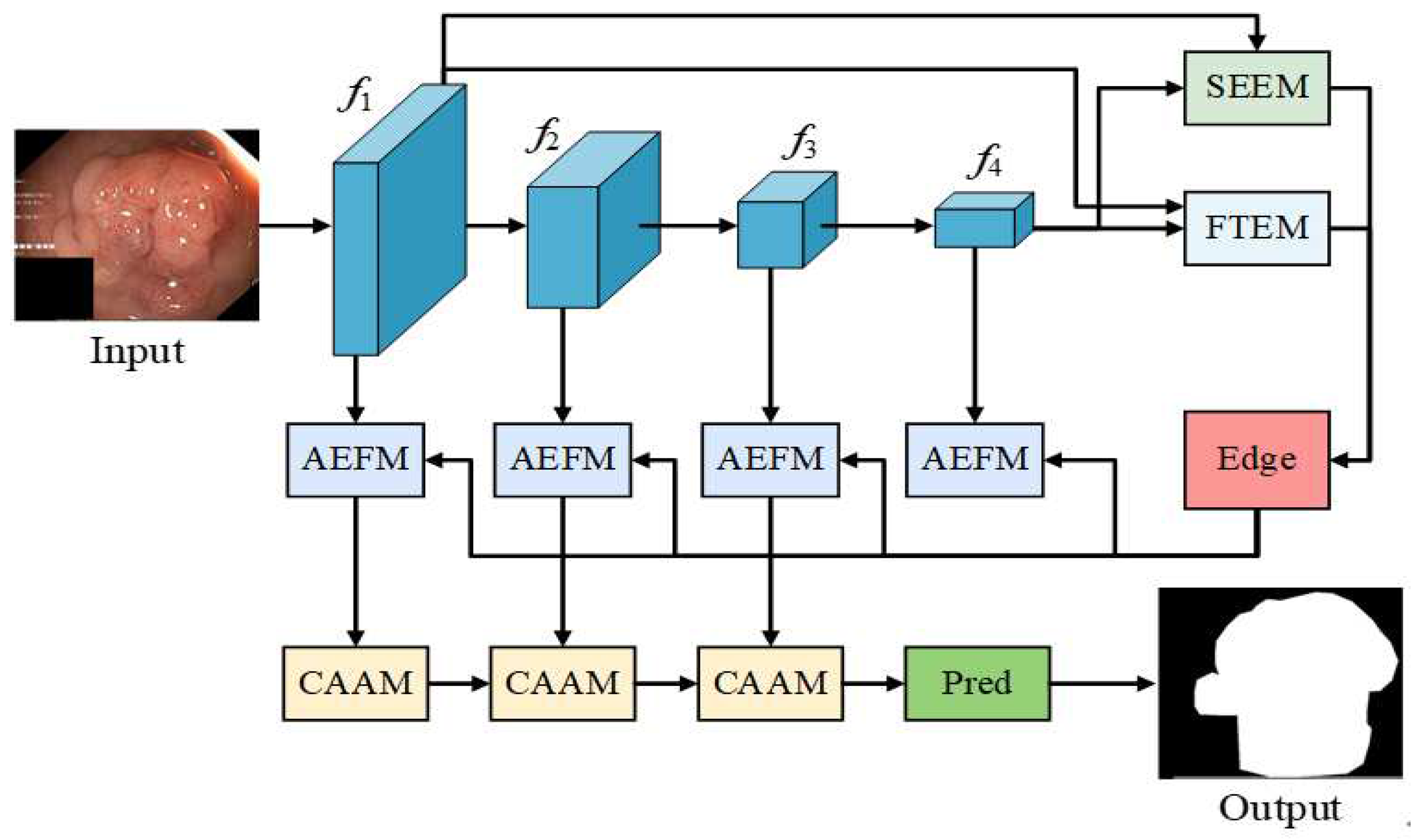
3. Methodologies
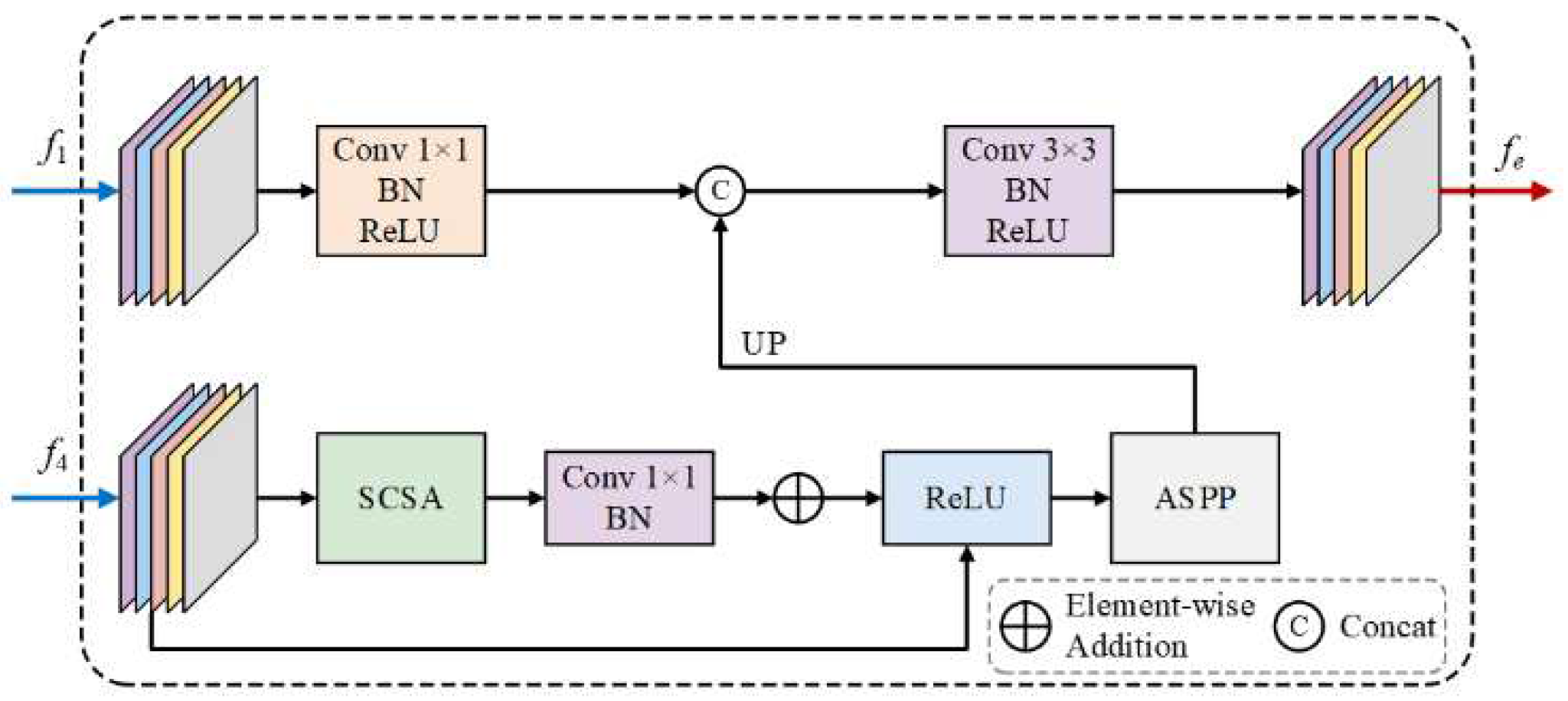
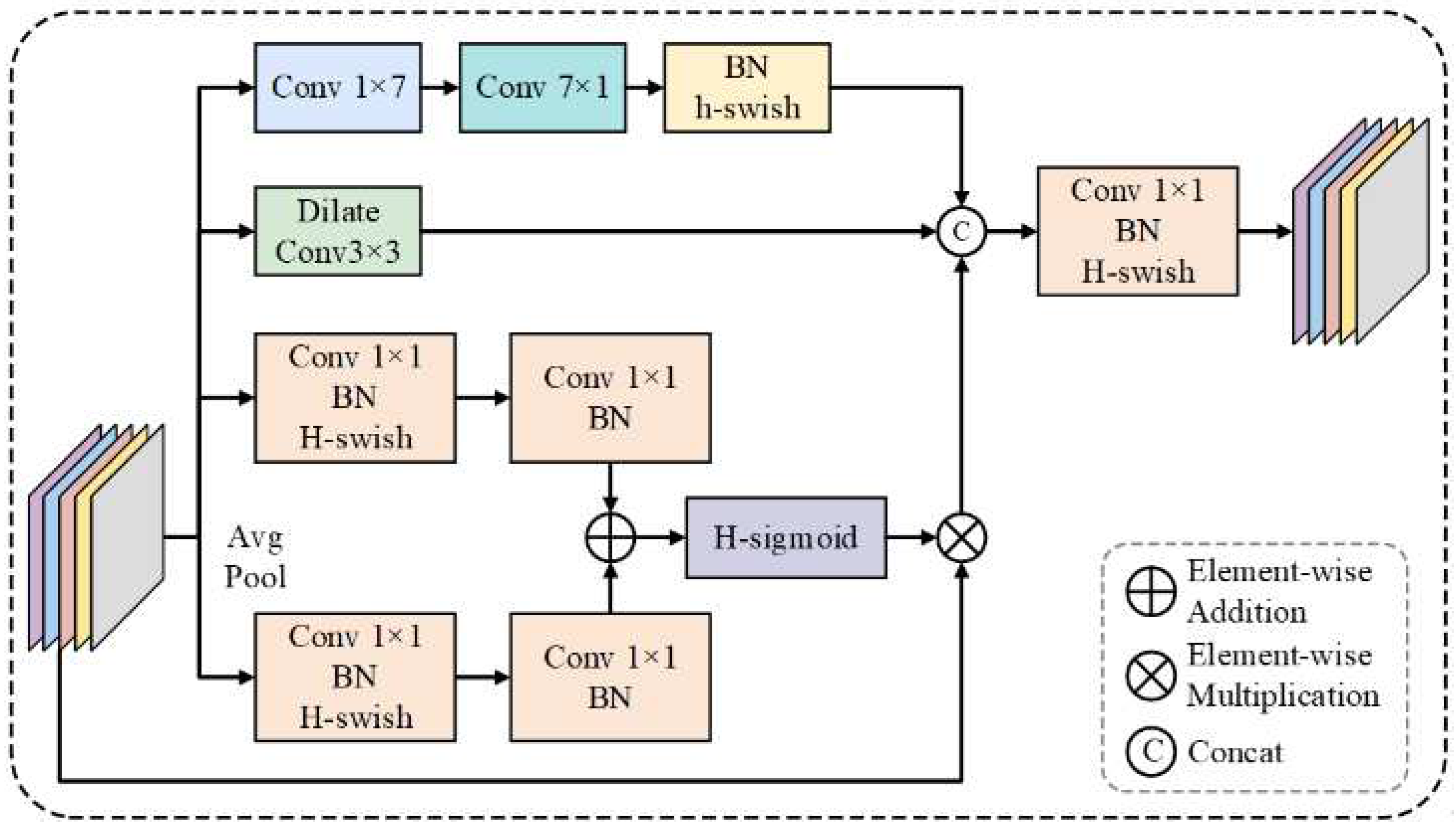
3.1. SEEM
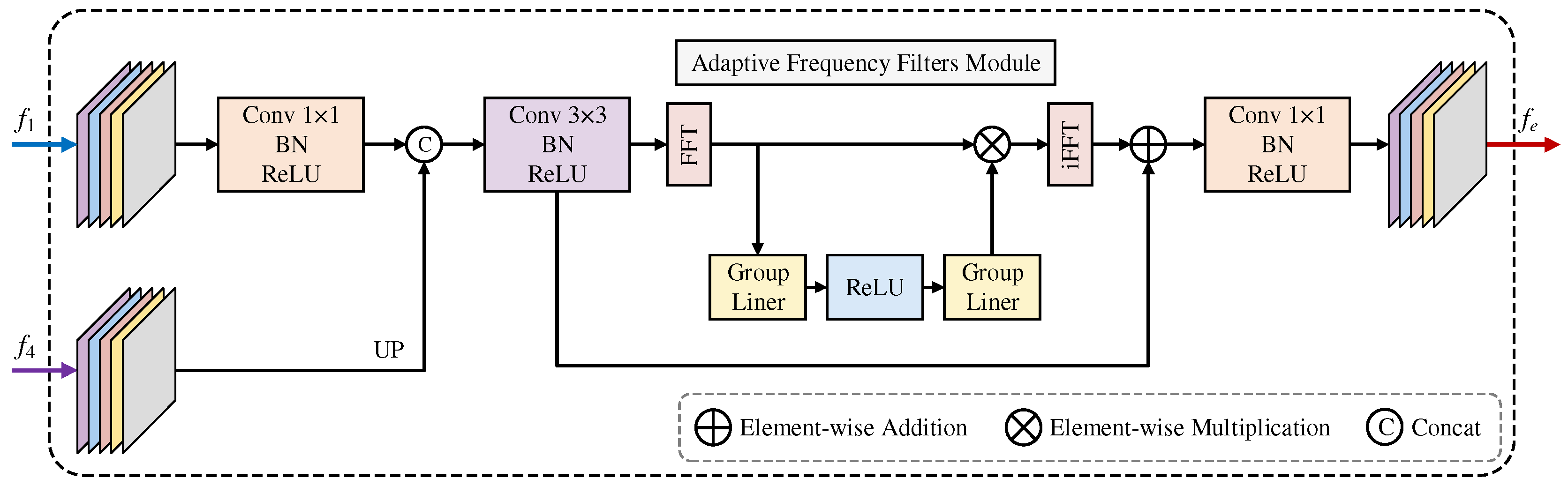
3.2. FTEM
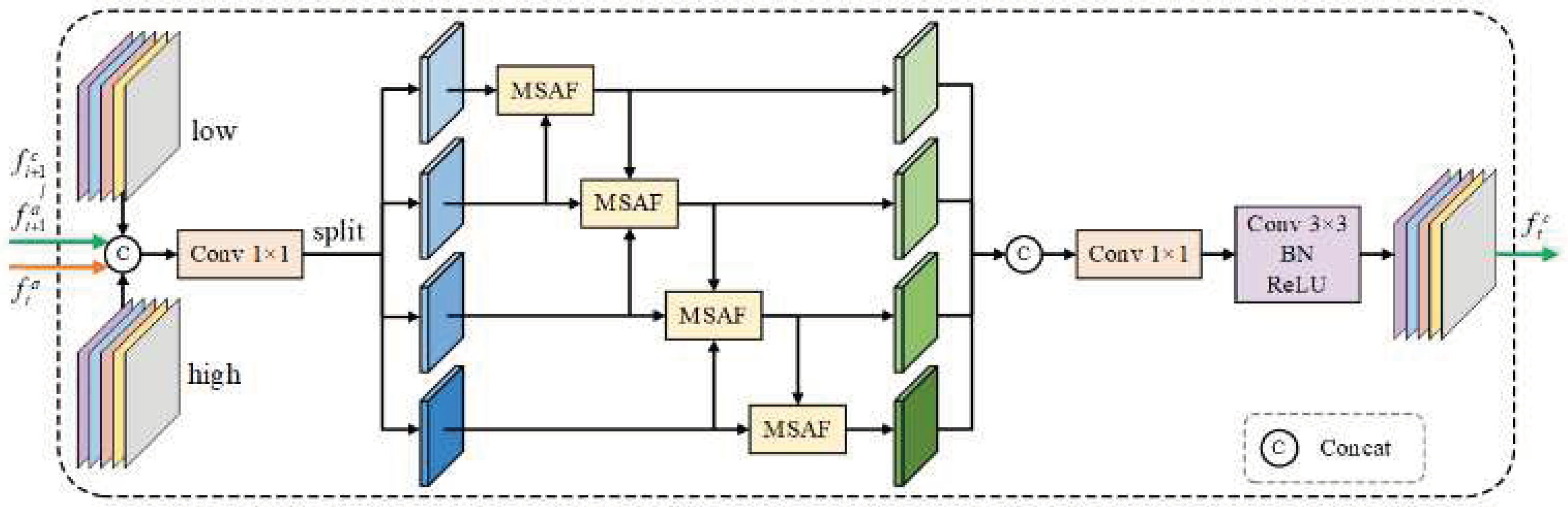
3.3. AEFM
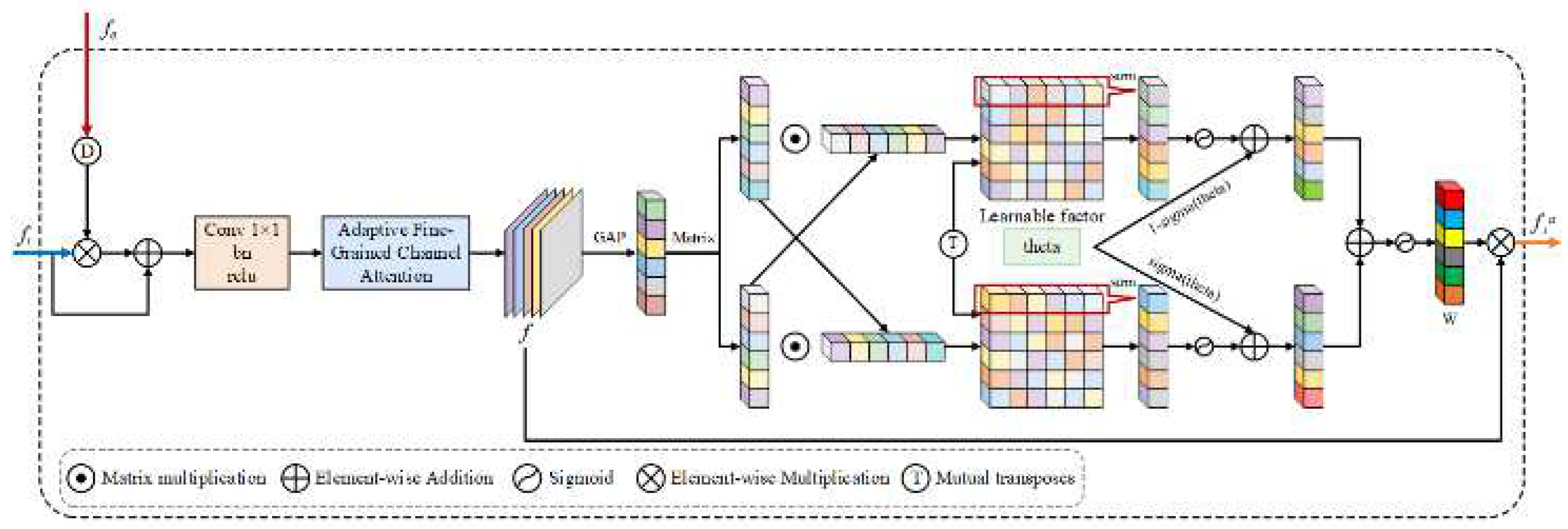
3.4. CAAM
4. Experiments Results
4.1. Implement Details
4.2. Implement Details
4.3. Evaluation Metrics
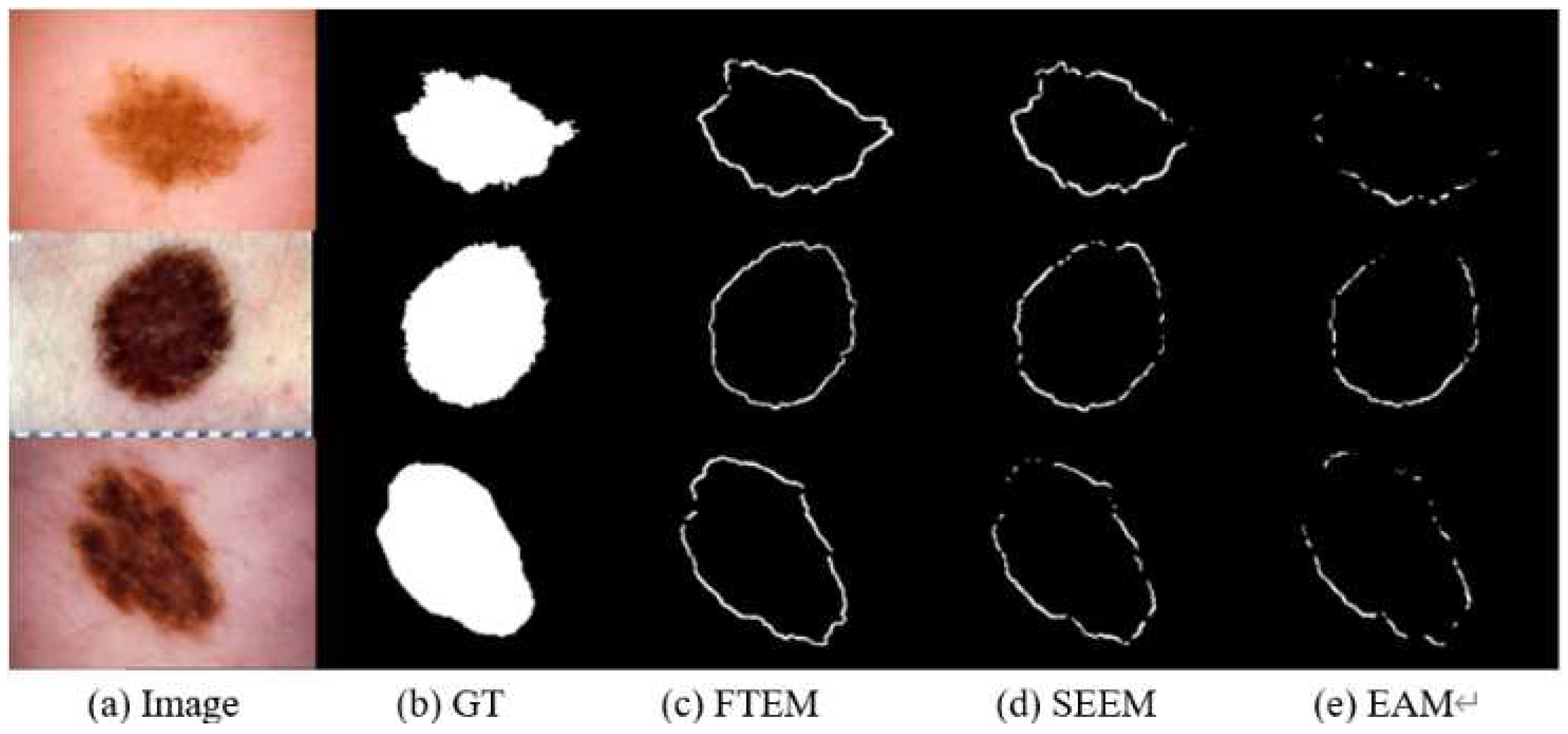
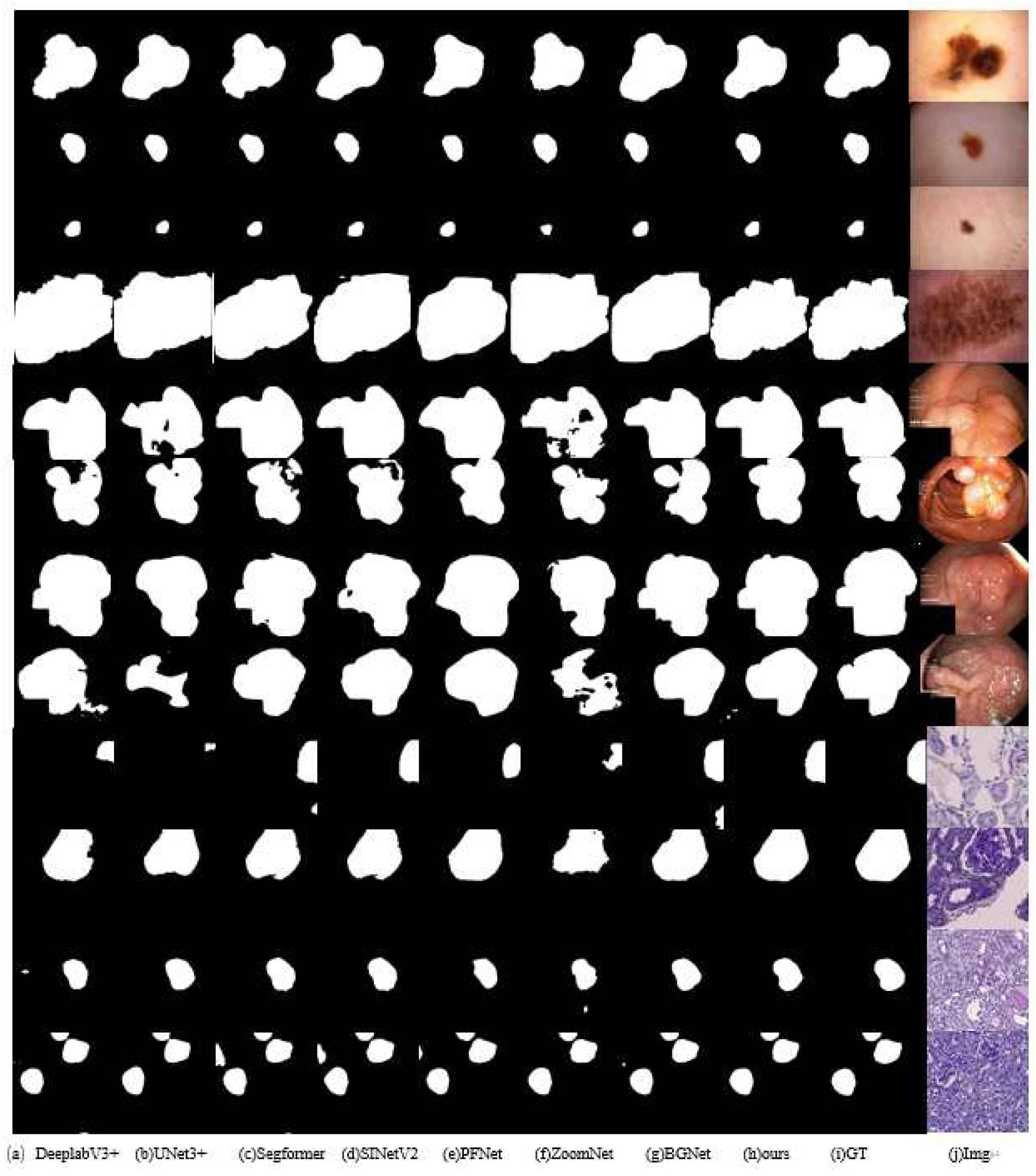
4.4. Comparative Experiments
4.5. Ablation Experiment
5. Discussion
6. Conclusions
References
- M. Kim and B. -D. Lee, “A Simple Generic Method for Effective Boundary Extraction in Medical Image Segmentation,” in IEEE Access, vol. 9, pp. 103875-103884, 2021. [CrossRef]
- J. Wang, X. Zhao, Q. Ning and D. Qian, “AEC-Net: Attention and Edge Constraint Network for Medical Image Segmentation,” 2020 42nd Annual International Conference of the IEEE Engineering in Medicine & Biology Society (EMBC), Montreal, QC, Canada, 2020, pp. 1616-1619. [CrossRef]
- Zhu, C., Zhang, X., Li, Y., Qiu, L., Han, K., & Han, X. (2022). SharpContour: A contour-based boundary refinement approach for efficient and accurate instance segmentation. In Proceedings of the IEEE/CVF Conference on computer vision and pattern recognition (pp. 4392-4401). [CrossRef]
- Fan, D. P., Ji, G. P., Cheng, M. M., & Shao, L. (2021). Concealed object detection. IEEE transactions on pattern analysis and machine intelligence, 44(10), 6024-6042. [CrossRef]
- Ji, G. P., Fan, D. P., Chou, Y. C., Dai, D., Liniger, A., Van Gool, L. (2023). Deep gradient learning for efficient camouflaged object detection. Machine Intelligence Research, 20(1), 92-108. [CrossRef]
- Ji, G. P., Zhu, L., Zhuge, M., & Fu, K. (2022). Fast camouflaged object detection via edge-based reversible re-calibration network. Pattern Recognition, 123, 108414. [CrossRef]
- H. Zhao, J. Shi, X. Qi, W. Wang, and J. Jia, Pyramid scene parsing network, in Proc. IEEE CVPR, pp. 6230-6239, (2017). [CrossRef]
- R. Azad, M. Asadi-Aghbolaghi, M. Fathy, and S. Escalera, Attention deeplabv3+: Multi-level context attention mechanism for skin lesion segmentation, in Proc. ECCV, LNCS 12535, (2020). [CrossRef]
- Z. Zhou, M. M. R. Siddiquee, N. Tajbakhsh, and J. Liang, UNet++: A Nested U-Net Architecture for Medical Image Segmentation, IEEE Trans. Med. Imaging, vol. 39, no. 6, pp. 1856-1867, (2020). [CrossRef]
- C. Qin, Y. Wu, W. Liao, J. Zeng, S. Liang, and X. Zhang, Improved U-Net3+ with stage residual for brain tumor segmentation, BMC Med. Imaging, vol. 22, no. 1, 14, (2022). [CrossRef]
- Oktay, O., Schlemper, J., Folgoc, L. L., Lee, M., Heinrich, M., Misawa, K., Rueckert, D. (2018). Attention u-net: Learning where to look for the pancreas. arXiv preprint. arXiv:1804.03999. [CrossRef]
- Chen, J., Mei, J., Li, X., Lu, Y., Yu, Q., Wei, Q., Zhou, Y. (2024). TransUNet: Rethinking the U-Net architecture design for medical image segmentation through the lens of transformers. Medical Image Analysis, 97, 103280. [CrossRef]
- H. Cao, Y. Wang, J. Chen, D. Jiang, X. Zhang, Q. Tian, and M. Wang, Swin-unet: Unet-like pure transformer for medical image segmentation, in Proc. CCF TCCV, pp. 205-218, (2022). [CrossRef]
- Xie, E., Wang, W., Yu, Z., Anandkumar, A., Alvarez, J. M., & Luo, P. (2021). SegFormer: Simple and efficient design for semantic segmentation with transformers. Advances in neural information processing systems, 34, 12077-12090. [CrossRef]
- Lee, H. J., Kim, J. U., Lee, S., Kim, H. G., & Ro, Y. M. (2020). Structure boundary preserving segmentation for medical image with ambiguous boundary. In Proceedings of the IEEE/CVF conference on computer vision and pattern recognition (pp. 4817-4826). [CrossRef]
- Wang, J., Chen, F., Ma, Y., Wang, L., Fei, Z., Shuai, J., Qin, J. (2023). Xbound-former: Toward cross-scale boundary modeling in transformers. IEEE Transactions on Medical Imaging, 42(6), 1735-1745. [CrossRef]
- Y. Pang, X. Zhao, T. Xiang, L. Zhang, and H. Lu, Zoom in and out: a mixed-scale triplet network for camouflaged object detection, in Proc. IEEE CVPR, pp. 2150-2160, (2022). [CrossRef]
- H. Mei, G.-P. Ji, Z. Wei, X. Yang, X. Wei, and D.-P. Fan, Camouflaged object segmentation with distraction mining, in Proc. IEEE CVPR, pp. 8768-8777, (2022). [CrossRef]
- Y. Sun, S. Wang, C. Chen, and T. Xiang, Boundary-guided camouflaged object detection, in Proc. IJCAI, pp. 1335-1341, (2022). [CrossRef]
- Zheng, P., Gao, D., Fan, D. P., Liu, L., Laaksonen, J., Ouyang, W., & Sebe, N. (2024). Bilateral reference for high-resolution dichotomous image segmentation. arXiv preprint. arXiv:2401.03407. [CrossRef]
- Si, Y., Xu, H., Zhu, X., Zhang, W., Dong, Y., Chen, Y., & Li, H. (2024). SCSA: Exploring the synergistic effects between spatial and channel attention. arXiv preprint. arXiv:2407.05128. [CrossRef]
- Sun, H., Wen, Y., Feng, H., Zheng, Y., Mei, Q., Ren, D., & Yu, M. (2024). Unsupervised bidirectional contrastive reconstruction and adaptive fine-grained channel attention networks for image dehazing. Neural Networks, 176, 106314. [CrossRef]
- Wang, G., Gan, X., Cao, Q., & Zhai, Q. (2023). MFANet: multi-scale feature fusion network with attention mechanism. The Visual Computer, 39(7), 2969-2980. [CrossRef]
- Wang, W., Xie, E., Li, X., Fan, D. P., Song, K., Liang, D., Shao, L. (2022). Pvt v2: Improved baselines with pyramid vision transformer. Computational visual media, 8(3), 415-424. [CrossRef]
- Y. Tang, Y. He, V. Nath, P. Guo, R. Deng, T. Yao, Q. Liu, C. Cui, M. Yin, Z. Xu, H. Roth, D. Xu, H. Yang, and Y. Huo, HoloHisto: End-to-end Gigapixel WSI Segmentation with 4K Resolution Sequential Tokenization, arXiv preprint, (2024). [CrossRef]
- Codella, N. C., Gutman, D., Celebi, M. E., Helba, B., Marchetti, M. A., Dusza, S. W., Halpern, A. (2018). Skin lesion analysis toward melanoma detection: A challenge at the 2017 international symposium on biomedical imaging (isbi), hosted by the international skin imaging collaboration (isic). In 2018 IEEE 15th international symposium on biomedical imaging (ISBI 2018) (pp. 168-172). IEEE. [CrossRef]
- Jha, D., Smedsrud, P. H., Riegler, M. A., Halvorsen, P., De Lange, T., Johansen, D., & Johansen, H. D. (2020). Kvasir-seg: A segmented polyp dataset. In MultiMedia modeling: 26th international conference, MMM 2020, Daejeon, South Korea, January 5–8, 2020, proceedings, part II 26 (pp. 451-462). Springer International Publishing. [CrossRef]
- Li, C., He, Y., Fu, Y., et al. (2023). GFUNet: A lightweight medical image segmentation network integrating global Fourier transform. Computers in Biology and Medicine, 165, 107255. [CrossRef]
- Ruan, Y., Han, J., Zhang, J., et al. (2024). MEW-UNet: Multi-axis external weights based UNet for medical image segmentation. arXiv preprint. arXiv:2312.17030. [CrossRef]
- Zhou, L., Xu, T., Yang, Z., et al. (2024). SF-UNet: A lightweight UNet framework based on spatial and frequency domain attention for medical image segmentation. arXiv preprint. arXiv:2406.07952. [CrossRef]

| Method | ISIC2018 | Kvasir-SEG | KPIs2024 | |||||||||
| mPA | mIoU | mDice | F1-score | mPA | mIoU | mDice | F1-score | mPA | mIoU | mDice | F1-score | |
| PSPNet | 0.950 | 0.881 | 0.899 | 0.906 | 0.970 | 0.890 | 0.899 | 0.898 | 0.979 | 0.842 | 0.850 | 0.860 |
| DeepLabV3plus | 0.951 | 0.886 | 0.907 | 0.911 | 0.974 | 0.904 | 0.911 | 0.912 | 0.977 | 0.832 | 0.840 | 0.845 |
| U-Net | 0.951 | 0.884 | 0.896 | 0.909 | 0.969 | 0.886 | 0.885 | 0.894 | 0.976 | 0.826 | 0.834 | 0.840 |
| Swin-UNet | 0.953 | 0.885 | 0.898 | 0.912 | 0.971 | 0.890 | 0.891 | 0.900 | 0.976 | 0.841 | 0.850 | 0.855 |
| U-Net3+ | 0.943 | 0.863 | 0.885 | 0.889 | 0.951 | 0.829 | 0.829 | 0.832 | 0.968 | 0.767 | 0.774 | 0.780 |
| Segformer | 0.950 | 0.883 | 0.900 | 0.909 | 0.977 | 0.914 | 0.918 | 0.924 | 0.987 | 0.907 | 0.920 | 0.925 |
| SINetV2 | 0.955 | 0.891 | 0.905 | 0.917 | 0.973 | 0.900 | 0.901 | 0.907 | 0.979 | 0.847 | 0.855 | 0.860 |
| PFNet | 0.946 | 0.873 | 0.885 | 0.899 | 0.965 | 0.877 | 0.879 | 0.884 | 0.976 | 0.823 | 0.830 | 0.835 |
| ZoomNet | 0.949 | 0.876 | 0.896 | 0.899 | 0.962 | 0.861 | 0.872 | 0.866 | 0.968 | 0.763 | 0.770 | 0.775 |
| BGNet | 0.951 | 0.885 | 0.902 | 0.910 | 0.975 | 0.908 | 0.914 | 0.915 | 0.983 | 0.877 | 0.882 | 0.885 |
| BiRefNet | 0.953 | 0.892 | 0.908 | 0.919 | 0.966 | 0.878 | 0.879 | 0.885 | 0.986 | 0.908 | 0.915 | 0.920 |
| FEA-Net(ours) | 0.968 | 0.912 | 0.918 | 0.924 | 0.981 | 0.923 | 0.923 | 0.929 | 0.988 | 0.915 | 0.923 | 0.931 |
| Method | ISIC2018 | Kvasir-SEG | KPIs2024 | |||||||||
| mPA | mIoU | mDice | F1-score | mPA | mIoU | mDice | F1-score | mPA | mIoU | mDice | F1-score | |
| SEEM+AEFM+CAAM | 0.959 | 0.903 | 0.910 | 0.916 | 0.977 | 0.915 | 0.918 | 0.921 | 0.985 | 0.901 | 0.908 | 0.911 |
| EAM+EFM+CAM (BGNet) | 0.951 | 0.885 | 0.902 | 0.910 | 0.975 | 0.908 | 0.914 | 0.915 | 0.983 | 0.877 | 0.882 | 0.885 |
| FTEM+ AEFM+CAAM | 0.965 | 0.907 | 0.913 | 0.919 | 0.979 | 0.918 | 0.920 | 0.923 | 0.986 | 0.911 | 0.915 | 0.923 |
| FTEM+EAM+ EFM+CAM | 0.960 | 0.901 | 0.909 | 0.916 | 0.977 | 0.914 | 0.917 | 0.920 | 0.985 | 0.907 | 0.909 | 0.914 |
| FTEM+SEEM+AEFM+CAAM (Ours) | 0.968 | 0.912 | 0.918 | 0.924 | 0.981 | 0.923 | 0.923 | 0.929 | 0.988 | 0.915 | 0.923 | 0.931 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




