Submitted:
02 May 2025
Posted:
07 May 2025
You are already at the latest version
Abstract
Keywords:
Article Highlights
- Single cell proteomics (SCP) is a promising technology defining the cellular proteomics at a single cell level.
- Absolute quantification of single-cell proteins is possible with the recent techniques.
- Innovative techniques such as, SCOPE, SCOPE2, nanoPOTS, CyTOF and IMC are used for SCP analysis.
- Multi-omics integration of SCP, SCG, scRNA-seq, SCM and SCEpi provide deep understanding of complex biological systems.
- SCP provides valuable insight of tumor heterogeneity, disease mechanisms, therapeutic development, neurobiology, and cellular and developmental biology.
Expert Opinion
Introduction
Methods of SCP
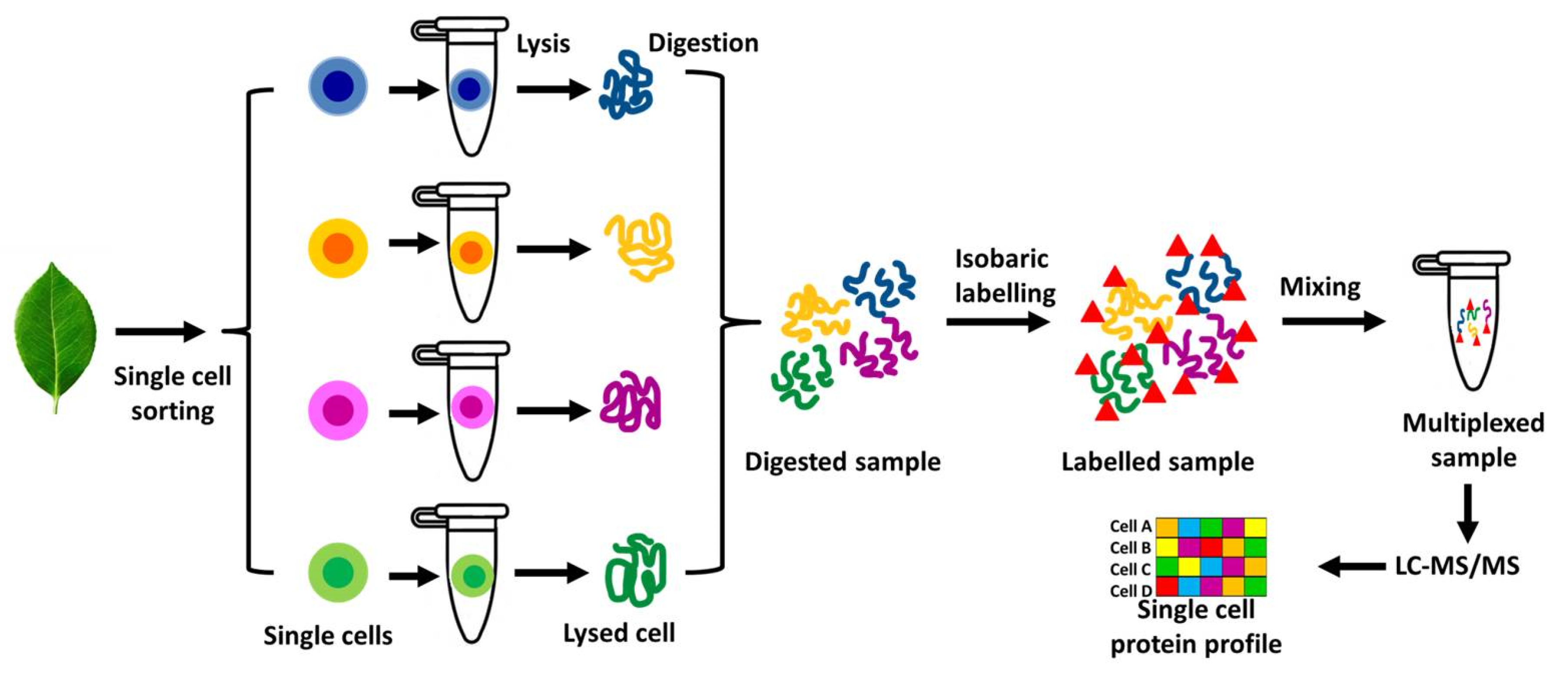
[1] Mass Spectrometry (MS)-Based Methods
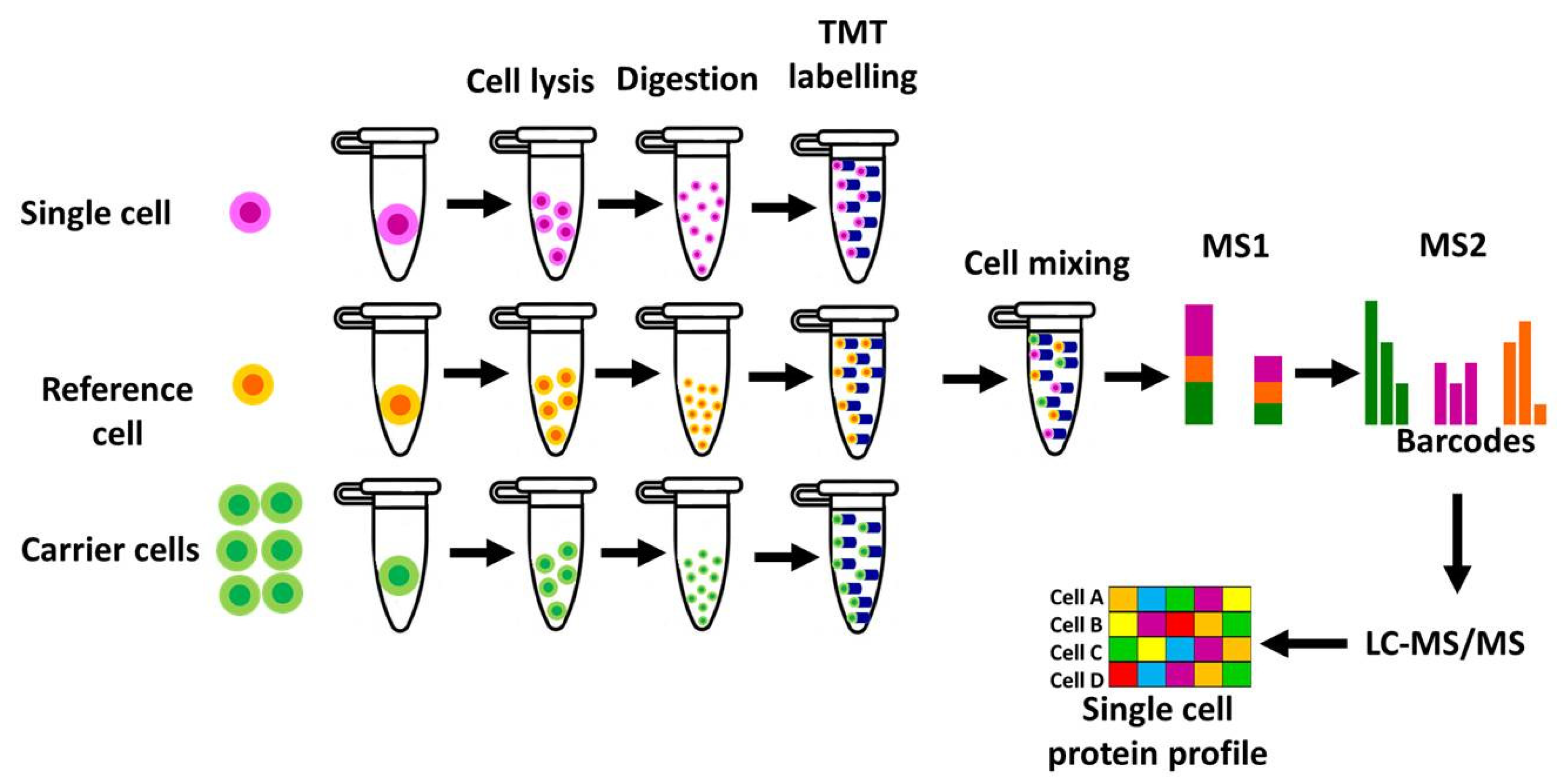
- I-Single Cell ProtEomics by Mass Spectrometry (SCoPE-MS)
- II-Single-Cell ProtEomics by Mass Spectrometry (SCoPE-MS) SCoPE2
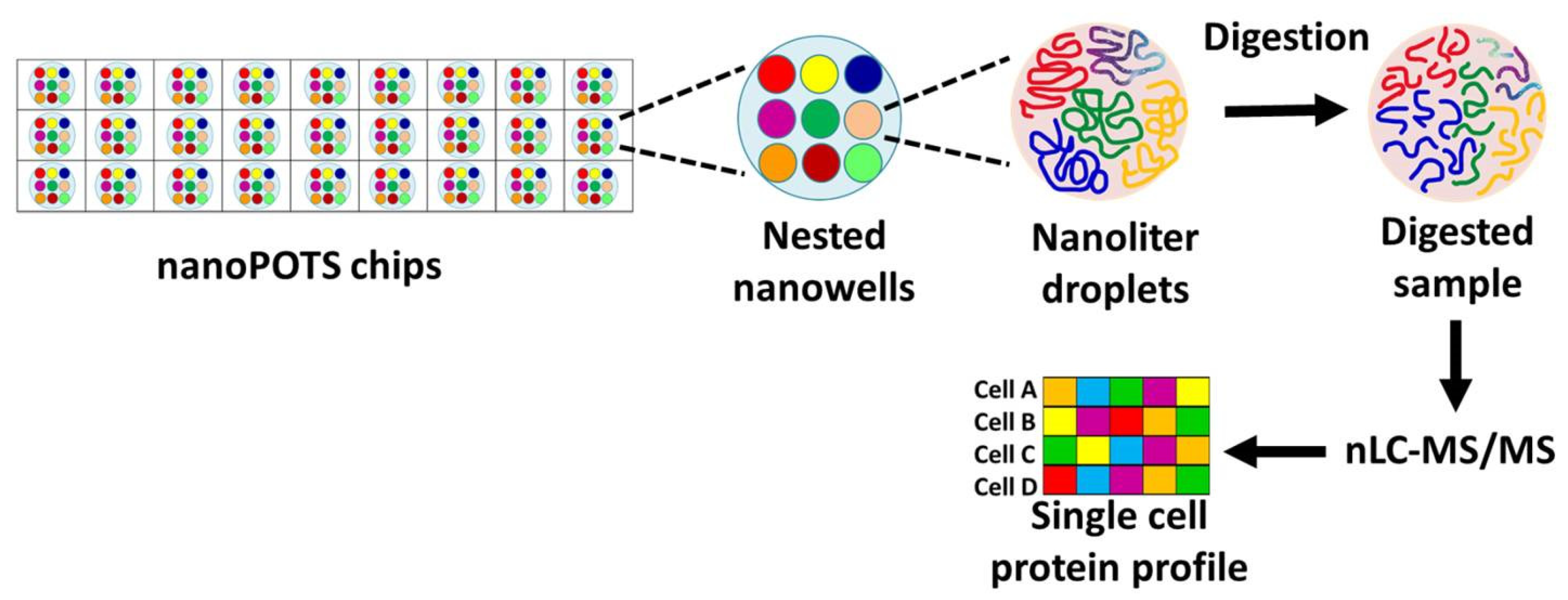
- III-Nanodroplet Processing in One Pot for Trace Samples (nanoPOTS)
[2] Antibody-Based Techniques
- I-Flow Cytometry (FCM)
- II-Mass Cytometry (CyTOF)
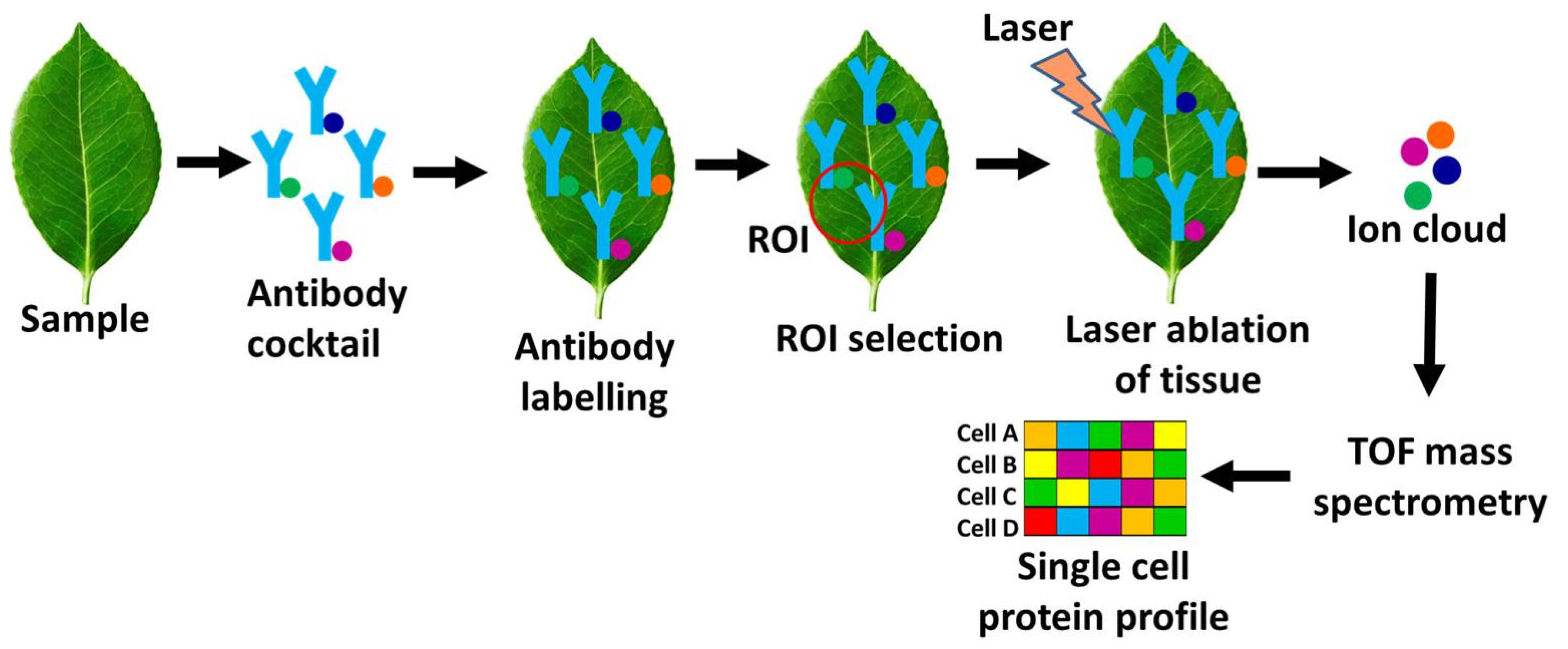
- [III] Imaging Mass Cytometry (IMC)
Integration with Other Omics Approaches
- Single-Cell Proteomics and single cell transcriptomics integration
- 2.
- Single-Cell Proteomics and single cell Epigenomics integration
- 3.
- Single-Cell Proteomics and single cell Metabolomics integration
- 4.
- Cellular Indexing of Transcriptomes and Epitopes by Sequencing (CITE-seq)
Technological Advancements and Future Directions
Applications
- ▪
- Cancer research: SCP holds immense promise for cancer research, particularly in understanding cancer progression, tumor heterogeneity, prediction of drug resistance and identification of new biomarkers and novel therapeutic targets.
- ▪
- Stem cell and developmental biology: SCP profiling of individual stem cells provides insights into cell differentiation and tissue regeneration for understanding developmental biology.
- ▪
- ▪ Neurobiology: SCP is important to study neurons or glial cells to explore proteome change during brain function, neurological disorders and neurodegenerative diseases.
- ▪
- Developmental biology: SCP explores cell differentiation and development to understand early embryonic development or organogenesis.
- ▪
- Cellular heterogeneity: SCP is important for understanding the cellular heterogeneity of complex tissues like tumors.
- ▪
- Biomarker discovery: SCP enables the discovery of biomarkers for specific cell types, states, or disease.
- ▪
- Drug response profiling: SCP is useful for monitoring the effects of drugs on individual cells, identifying drug-resistant cell populations, and discovering drug mechanisms in real time.
- ▪
- Immunology: SCP aid understanding the immune system at unprecedented resolution, identifying unique protein signatures in different immune cells, and monitoring immune responses during infection or vaccination.
- ▪
- Neurodegenerative diseases: SCP allows examination of protein misfolding, aggregation, and cellular stress responses in individual neurons in Alzheimer’s, Parkinson’s, and others neurological diseases.
Conclusions
Author Contributions
Disclosure Statement
Declaration of Interest
References
- Paolillo C, Londin E, Fortina P. Single-Cell Genomics. Clin Chem, 65(8), 972-985 (2019). [CrossRef]
- He S, Wang LH, Liu Y et al. Single-cell transcriptome profiling of an adult human cell atlas of 15 major organs. Genome Biol, 21(1), 294 (2020). [CrossRef]
- Emara S, Amer S, Ali A, Abouleila Y, Oga A, Masujima T. Single-Cell Metabolomics. Adv Exp Med Biol, 965, 323-343 (2017).
- Wen L, Tang F. Single cell epigenome sequencing technologies. Mol Aspects Med, 59, 62-69 (2018). [CrossRef]
- Vistain LF, Tay S. Single-Cell Proteomics. Trends Biochem Sci, 46(8), 661-672 (2021).
- Mund A, Coscia F, Kriston A et al. Deep Visual Proteomics defines single-cell identity and heterogeneity. Nat Biotechnol, 40(8), 1231-1240 (2022). [CrossRef]
- Wilson NK, Kent DG, Buettner F et al. Combined Single-Cell Functional and Gene Expression Analysis Resolves Heterogeneity within Stem Cell Populations. Cell Stem Cell, 16(6), 712-724 (2015). [CrossRef]
- Eze UC, Bhaduri A, Haeussler M, Nowakowski TJ, Kriegstein AR. Single-cell atlas of early human brain development highlights heterogeneity of human neuroepithelial cells and early radial glia. Nat Neurosci, 24(4), 584-594 (2021). [CrossRef]
- Vegliante R, Pastushenko I, Blanpain C. Deciphering functional tumor states at single-cell resolution. Embo j, 41(2), e109221 (2022). [CrossRef]
- Budnik B, Levy E, Harmange G, Slavov N. SCoPE-MS: mass spectrometry of single mammalian cells quantifies proteome heterogeneity during cell differentiation. Genome Biology, 19(1), 161 (2018). [CrossRef]
- Specht H, Emmott E, Petelski AA et al. Single-cell proteomic and transcriptomic analysis of macrophage heterogeneity using SCoPE2. Genome Biol, 22(1), 50 (2021). [CrossRef]
- Zhu Y, Piehowski PD, Zhao R et al. Nanodroplet processing platform for deep and quantitative proteome profiling of 10-100 mammalian cells. Nat Commun, 9(1), 882 (2018). [CrossRef]
- Bandura DR, Baranov VI, Ornatsky OI et al. Mass cytometry: technique for real time single cell multitarget immunoassay based on inductively coupled plasma time-of-flight mass spectrometry. Anal Chem, 81(16), 6813-6822 (2009). [CrossRef]
- Robinson JP, Roederer M. HISTORY OF SCIENCE. Flow cytometry strikes gold. Science, 350(6262), 739-740 (2015). [CrossRef]
- Ctortecka C, Hartlmayr D, Seth A et al. An Automated Nanowell-Array Workflow for Quantitative Multiplexed Single-Cell Proteomics Sample Preparation at High Sensitivity. Mol Cell Proteomics, 22(12), 100665 (2023). [CrossRef]
- Kim S, Kamarulzaman L, Taniguchi Y. Recent methodological advances towards single-cell proteomics. Proc Jpn Acad Ser B Phys Biol Sci, 99(8), 306-327 (2023). [CrossRef]
- Nalla LV, Kanukolanu A, Yeduvaka M, Gajula SNR. Advancements in Single-Cell Proteomics and Mass Spectrometry-Based Techniques for Unmasking Cellular Diversity in Triple Negative Breast Cancer. PROTEOMICS – Clinical Applications, 19(1), e202400101 (2025). [CrossRef]
- Petrosius V, Schoof EM. Recent advances in the field of single-cell proteomics. Transl Oncol, 27, 101556 (2023). [CrossRef]
- Huang B, Wu H, Bhaya D et al. Counting low-copy number proteins in a single cell. Science, 315(5808), 81-84 (2007). [CrossRef]
- Specht H, Harmange G, Perlman DH et al. Automated sample preparation for high-throughput single-cell proteomics. bioRxiv, 399774 (2018).
- Petelski AA, Emmott E, Leduc A et al. Multiplexed single-cell proteomics using SCoPE2. Nat Protoc, 16(12), 5398-5425 (2021). [CrossRef]
- Cong Y, Liang Y, Motamedchaboki K et al. Improved Single-Cell Proteome Coverage Using Narrow-Bore Packed NanoLC Columns and Ultrasensitive Mass Spectrometry. Anal Chem, 92(3), 2665-2671 (2020). [CrossRef]
- Woo J, Williams SM, Markillie LM et al. High-throughput and high-efficiency sample preparation for single-cell proteomics using a nested nanowell chip. Nat Commun, 12(1), 6246 (2021). [CrossRef]
- Lohani V, A.R A, Kundu S, Akhter MDQ, Bag S. Single-Cell Proteomics with Spatial Attributes: Tools and Techniques. ACS Omega, 8(20), 17499-17510 (2023). [CrossRef]
- Golden JP, Kim JS, Erickson JS et al. Multi-wavelength microflow cytometer using groove-generated sheath flow. Lab Chip, 9(13), 1942-1950 (2009). [CrossRef]
- Stavrakis S, Holzner G, Choo J, deMello A. High-throughput microfluidic imaging flow cytometry. Curr Opin Biotechnol, 55, 36-43 (2019). [CrossRef]
- Miura T, Mikami H, Isozaki A, Ito T, Ozeki Y, Goda K. On-chip light-sheet fluorescence imaging flow cytometry at a high flow speed of 1 m/s. Biomed Opt Express, 9(7), 3424-3433 (2018). [CrossRef]
- Holzner G, Mateescu B, van Leeuwen D et al. High-throughput multiparametric imaging flow cytometry: toward diffraction-limited sub-cellular detection and monitoring of sub-cellular processes. Cell Reports, 34(10), 108824 (2021). [CrossRef]
- Bendall SC, Davis KL, Amir el AD et al. Single-cell trajectory detection uncovers progression and regulatory coordination in human B cell development. Cell, 157(3), 714-725 (2014). [CrossRef]
- Porpiglia E, Samusik N, Ho ATV et al. High-resolution myogenic lineage mapping by single-cell mass cytometry. Nat Cell Biol, 19(5), 558-567 (2017). [CrossRef]
- Wroblewska A, Dhainaut M, Ben-Zvi B et al. Protein Barcodes Enable High-Dimensional Single-Cell CRISPR Screens. Cell, 175(4), 1141-1155.e1116 (2018). [CrossRef]
- Stern L, McGuire H, Avdic S et al. Mass Cytometry for the Assessment of Immune Reconstitution After Hematopoietic Stem Cell Transplantation. Front Immunol, 9, 1672 (2018). [CrossRef]
- Chang Q, Ornatsky O, Hedley D. Staining of Frozen and Formalin-Fixed, Paraffin-Embedded Tissues with Metal-Labeled Antibodies for Imaging Mass Cytometry Analysis. Curr Protoc Cytom, 82, 12.47.11-12.47.18 (2017). [CrossRef]
- Catena R, Montuenga LM, Bodenmiller B. Ruthenium counterstaining for imaging mass cytometry. J Pathol, 244(4), 479-484 (2018). [CrossRef]
- Jackson HW, Fischer JR, Zanotelli VRT et al. The single-cell pathology landscape of breast cancer. Nature, 578(7796), 615-620 (2020). [CrossRef]
- Kuett L, Catena R, Özcan A et al. Three-dimensional imaging mass cytometry for highly multiplexed molecular and cellular mapping of tissues and the tumor microenvironment. Nat Cancer, 3(1), 122-133 (2022). [CrossRef]
- Gerdtsson E, Pore M, Thiele JA et al. Multiplex protein detection on circulating tumor cells from liquid biopsies using imaging mass cytometry. Converg Sci Phys Oncol, 4(1) (2018). [CrossRef]
- Flynn E, Almonte-Loya A, Fragiadakis GK. Single-Cell Multiomics. Annu Rev Biomed Data Sci, 6, 313-337 (2023). [CrossRef]
- Lee J, Hyeon DY, Hwang D. Single-cell multiomics: technologies and data analysis methods. Experimental & Molecular Medicine, 52(9), 1428-1442 (2020). [CrossRef]
- Jha PK, Valekunja UK, Ray S, Nollet M, Reddy AB. Single-cell transcriptomics and cell-specific proteomics reveals molecular signatures of sleep. Commun Biol, 5(1), 846 (2022). [CrossRef]
- Ali A, Davidson S, Fraenkel E et al. Single cell metabolism: current and future trends. Metabolomics, 18(10), 77 (2022). [CrossRef]
- Vandereyken K, Sifrim A, Thienpont B, Voet T. Methods and applications for single-cell and spatial multi-omics. Nat Rev Genet, 24(8), 494-515 (2023). [CrossRef]
- Bi H, Weng X. Single-Cell Epigenomics and Proteomics Methods Integrated in Multiomics. Fundamental Research, (2024). [CrossRef]
- Orsburn BC. Metabolomic, Proteomic, and Single-Cell Proteomic Analysis of Cancer Cells Treated with the KRAS(G12D) Inhibitor MRTX1133. J Proteome Res, 22(12), 3703-3713 (2023).
- Stoeckius M, Hafemeister C, Stephenson W et al. Simultaneous epitope and transcriptome measurement in single cells. Nat Methods, 14(9), 865-868 (2017). [CrossRef]
- Kumar S, Orcales F, Shih BB et al. Cellular indexing of transcriptomes and epitopes (CITE-Seq) in hidradenitis suppurativa identifies dysregulated cell types in peripheral blood and facilitates diagnosis via machine learning. Res Sq, (2024). [CrossRef]
- Fulcher JM, Markillie LM, Mitchell HD et al. Parallel measurement of transcriptomes and proteomes from same single cells using nanodroplet splitting. Nat Commun, 15(1), 10614 (2024). [CrossRef]
- Zheng R, Matzinger M, Mayer RL, Valenta A, Sun X, Mechtler K. A High-Sensitivity Low-Nanoflow LC-MS Configuration for High-Throughput Sample-Limited Proteomics. Analytical Chemistry, 95(51), 18673-18678 (2023). [CrossRef]
- Hartlmayr D, Ctortecka C, Seth A, Mendjan S, Tourniaire G, Mechtler K. An automated workflow for label-free and multiplexed single cell proteomics sample preparation at unprecedented sensitivity. bioRxiv, 2021.2004.2014.439828 (2021). [CrossRef]
- Zhang T, Warden AR, Li Y, Ding X. Progress and applications of mass cytometry in sketching immune landscapes. Clin Transl Med, 10(6), e206 (2020). [CrossRef]
- Lun XK, Szklarczyk D, Gábor A et al. Analysis of the Human Kinome and Phosphatome by Mass Cytometry Reveals Overexpression-Induced Effects on Cancer-Related Signaling. Mol Cell, 74(5), 1086-1102.e1085 (2019). [CrossRef]
- Hiam-Galvez KJ, Allen BM, Spitzer MH. Systemic immunity in cancer. Nat Rev Cancer, 21(6), 345-359 (2021). [CrossRef]
- Neuperger P, Szalontai K, Gémes N et al. Single-cell mass cytometric analysis of peripheral immunity and multiplex plasma marker profiling of non-small cell lung cancer patients receiving PD-1 targeting immune checkpoint inhibitors in comparison with platinum-based chemotherapy. Front Immunol, 14, 1243233 (2023). [CrossRef]
- Kravchenko-Balasha N, Shin YS, Sutherland A, Levine RD, Heath JR. Intercellular signaling through secreted proteins induces free-energy gradient-directed cell movement. Proc Natl Acad Sci U S A, 113(20), 5520-5525 (2016). [CrossRef]
- Xue Q, Bettini E, Paczkowski P et al. Single-cell multiplexed cytokine profiling of CD19 CAR-T cells reveals a diverse landscape of polyfunctional antigen-specific response. J Immunother Cancer, 5(1), 85 (2017). [CrossRef]
- Su Y, Wei W, Robert L et al. Single-cell analysis resolves the cell state transition and signaling dynamics associated with melanoma drug-induced resistance. Proc Natl Acad Sci U S A, 114(52), 13679-13684 (2017). [CrossRef]
- Sinkala E, Sollier-Christen E, Renier C et al. Profiling protein expression in circulating tumour cells using microfluidic western blotting. Nat Commun, 8, 14622 (2017). [CrossRef]
- Hughes AJ, Spelke DP, Xu Z, Kang CC, Schaffer DV, Herr AE. Single-cell western blotting. Nat Methods, 11(7), 749-755 (2014). [CrossRef]
- Shahi P, Kim SC, Haliburton JR, Gartner ZJ, Abate AR. Abseq: Ultrahigh-throughput single cell protein profiling with droplet microfluidic barcoding. Sci Rep, 7, 44447 (2017). [CrossRef]
- Ramji R, Wang M, Bhagat AA et al. Single cell kinase signaling assay using pinched flow coupled droplet microfluidics. Biomicrofluidics, 8(3), 034104 (2014). [CrossRef]
- Reza KK, Dey S, Wuethrich A et al. In Situ Single Cell Proteomics Reveals Circulating Tumor Cell Heterogeneity during Treatment. ACS Nano, 15(7), 11231-11243 (2021). [CrossRef]
- Yang L, Ball A, Liu J et al. Cyclic microchip assay for measurement of hundreds of functional proteins in single neurons. Nat Commun, 13(1), 3548 (2022). [CrossRef]
- Piehowski PD, Zhu Y, Bramer LM et al. Automated mass spectrometry imaging of over 2000 proteins from tissue sections at 100-μm spatial resolution. Nat Commun, 11(1), 8 (2020). [CrossRef]
- Kwon Y, Piehowski PD, Zhao R et al. Hanging drop sample preparation improves sensitivity of spatial proteomics. Lab Chip, 22(15), 2869-2877 (2022). [CrossRef]
- Végvári Á, Rodriguez JE, Zubarev RA. Single-Cell Chemical Proteomics (SCCP) Interrogates the Timing and Heterogeneity of Cancer Cell Commitment to Death. Anal Chem, 94(26), 9261-9269 (2022). [CrossRef]
- Gebreyesus ST, Siyal AA, Kitata RB et al. Streamlined single-cell proteomics by an integrated microfluidic chip and data-independent acquisition mass spectrometry. Nat Commun, 13(1), 37 (2022). [CrossRef]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



