Submitted:
24 June 2025
Posted:
30 June 2025
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Generalization Across MRI Datasets—Training AI models on scans from one or two hospital cohorts has the tendency to result in loss of accuracy when trained on another cohort because the type of scanner used, acquisition parameters, and patient population differ across sites, causing a domain shift.
- Class Imbalance in Brain Tumor Datasets—A majority of publicly available datasets contain imbalanced distributions of the tumor classes, causing biased predictions in favor of the majority class.
- Explainability & trust – Deep models are "black boxes" which makes it hard for clinicians to interpret AI-generated diagnoses and hinder real-world usage.
- Small and Early-Stage Tumor Detection: Because small tumors and early-stage tumor growth are usually hard to distinguish from normal tissues in MRI scans, most models are unable to detect them.
2. Literature Review
2.1. Pre-Trained Models for Brain Tumor Classification
2.2. Custom AI Models for Brain Tumor Classification
2.3. Comparative Performance Analysis
3. Methodology
3.1. Dataset Selection
-
Kaggle “Brain Tumor Classification (MRI)” Dataset [10] – Comprising 3,264 T1-weighted contrast-enhanced MRI images, categorized into four classes:
- (a)
- Glioma tumor
- (b)
- Meningioma tumor
- (c)
- Pituitary tumor
- (d)
- No tumor (healthy cases)
-
Figshare “Brain Tumor Dataset” [11] – Containing 7,000+ MRI images, categorized into three classes:
- (a)
- Glioma tumor
- (b)
- Meningioma tumor
- (c)
- Pituitary tumor
3.2. Data Preprocessing
- Image Resizing – All images were resized to 224 × 224 pixels, aligning with VGG16’s input size.
- Normalization – Pixel values were scaled to [0, 1] using min-max scaling to improve model convergence.
-
Image Augmentation – To increase dataset variability and reduce overfitting, we applied:
- (a)
- Rotation (±25°)
- (b)
- Horizontal & vertical flipping
- (c)
- Zoom (±20%)
- (d)
- Contrast adjustment
- Class Imbalance Handling – Class weights were computed and applied during training to mitigate bias toward majority tumor classes.
3.3. Data Augmentation
- Rotation: Random rotations () to simulate different viewing angles.
- Flipping: Horizontal and vertical flips to enhance spatial invariance.
- Zooming: Random zoom-in and zoom-out to introduce variations in tumor magnifications.
- Brightness Adjustment: Controlled intensity modifications to account for differences across MRI scanners.
- Shifting: Minor translations of the image to make the model robust to positional variations.
3.4. Class Imbalance Handling
- Class Weights: Adjusted loss function penalties to counterbalance the effect of dominant classes.
- Oversampling: Replicated minority class images to ensure a more balanced representation.
- Targeted Data Augmentation: Applied additional augmentations exclusively to underrepresented classes to synthetically increase their presence.
3.5. Transfer Learning Architecture
3.5.1. VGG16 Model Adaptation
- Pre-trained Base Model: The VGG16 model was loaded with ImageNet weights, excluding the fully connected layers (include_top=False).
- Frozen Layers: All convolutional layers in VGG16 were initially frozen, preventing their weights from being modified:

-
Custom Classification Head: The fully connected layers were replaced with a trainable classification head consisting of:
- −
- Global Average Pooling (GAP) – Reducing feature maps to a 512-dimensional vector.
- −
- Batch Normalization – Stabilizing activations for better convergence.
- −
-
Fully Connected Dense Layers:
- ∗
- 256 neurons (ReLU activation, dropout = 0.5)
- ∗
- 128 neurons (ReLU activation, dropout = 0.5)
- −
- Softmax Output Layer – Classifying MRI scans into 3 categories (glioma, meningioma, pituitary tumor).
| Layer (Type) | Output Shape | Params |
|---|---|---|
| VGG16 (Base Model) | (None, 7, 7, 512) | 14,714,688 |
| Global Average Pooling (GAP) | (None, 512) | 0 |
| Batch Normalization | (None, 512) | 2,048 |
| Dense (256 neurons, ReLU) | (None, 256) | 131,328 |
| Dropout (0.5) | (None, 256) | 0 |
| Dense (128 neurons, ReLU) | (None, 128) | 32,896 |
| Dropout (0.5) | (None, 128) | 0 |
| Dense (Softmax, 3 output classes) | (None, 3) | 387 |
3.5.2. Justification for VGG16 Selection
- Total Parameters: 14,881,347
- Trainable Parameters: 165,635
- Non-trainable Parameters: 14,715,712
3.6. Training Strategy
- Optimizer: Adam optimizer (learning_rate = 1e-4)
- Loss Function: Categorical Cross-Entropy
- Batch Size: 32
- Epochs: 100 (Early stopping after 10 epochs of no improvement)
-
Callbacks:
- −
- EarlyStopping – Monitors validation loss and stops training if no improvement is detected:

- −
- ModelCheckpoint – Saves the best model based on validation performance:

- Training Execution:

3.7. Evaluation Metrics & Comparative Analysis
- Accuracy – Overall classification correctness.
- Precision – Proportion of correctly classified tumors per class.
- Recall (Sensitivity) – True positive rate, measuring detection ability.
- F1-Score – Balancing precision and recall.
- Confusion Matrix – Visualizing misclassifications across tumor types.
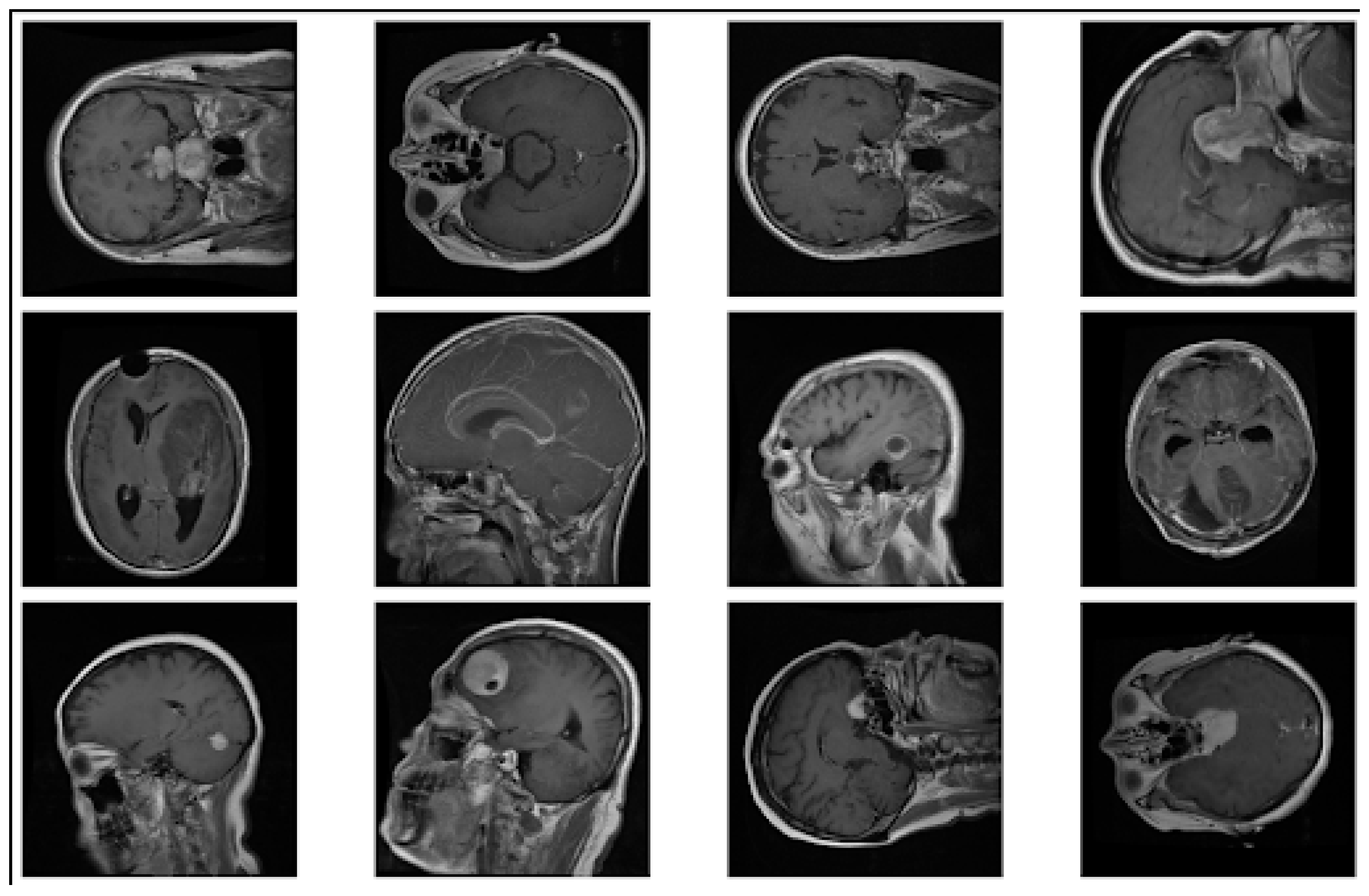
4. Visualization
4.1. Data Augmentation Previews
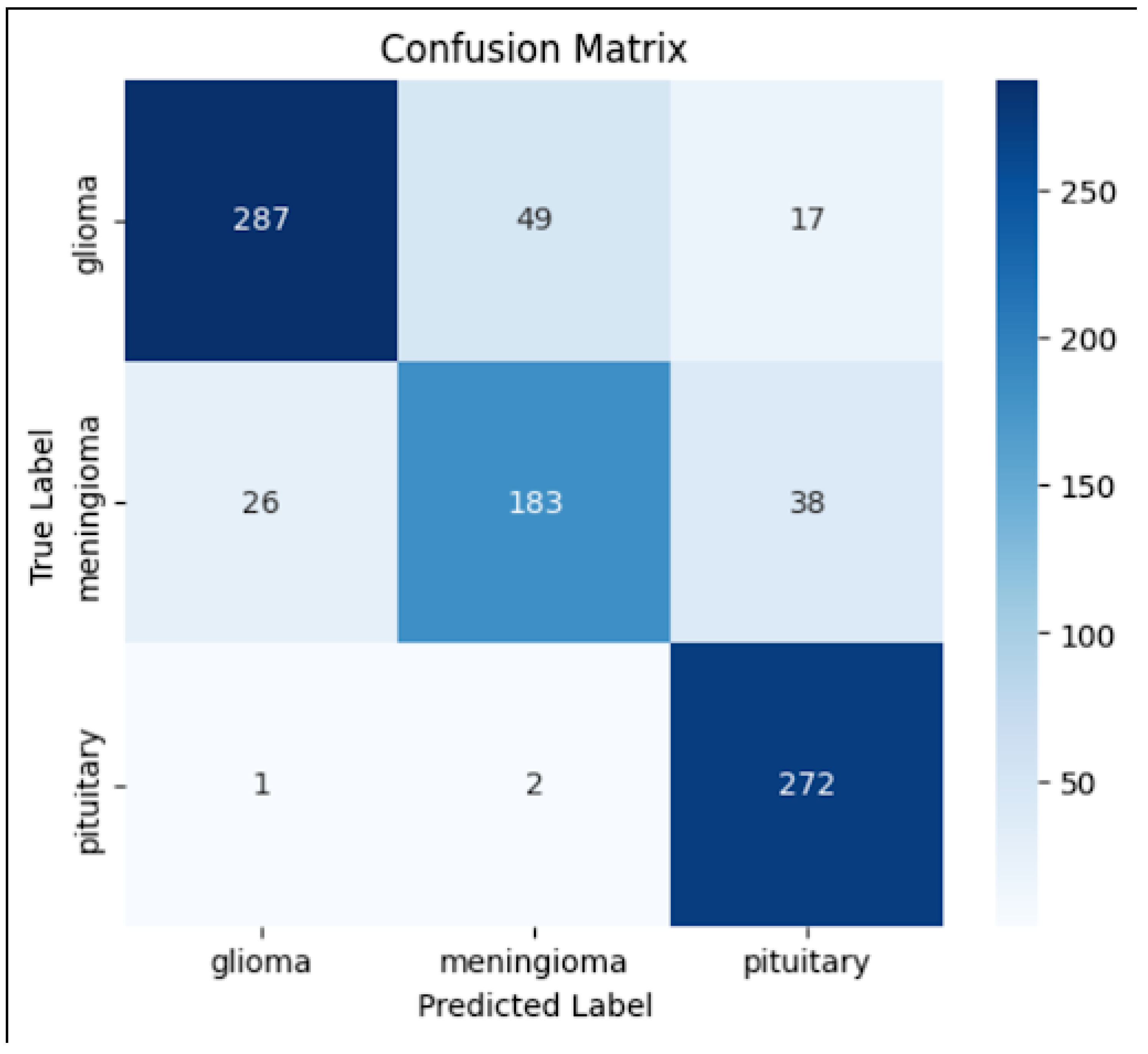
4.2. Confusion Matrix
- The diagonal values indicate correct classifications, whereas off-diagonal values highlight misclassifications.
- The model performed well in classifying pituitary tumors, but some misclassifications occurred between glioma and meningioma, which could be attributed to their structural similarities in MRI scans.
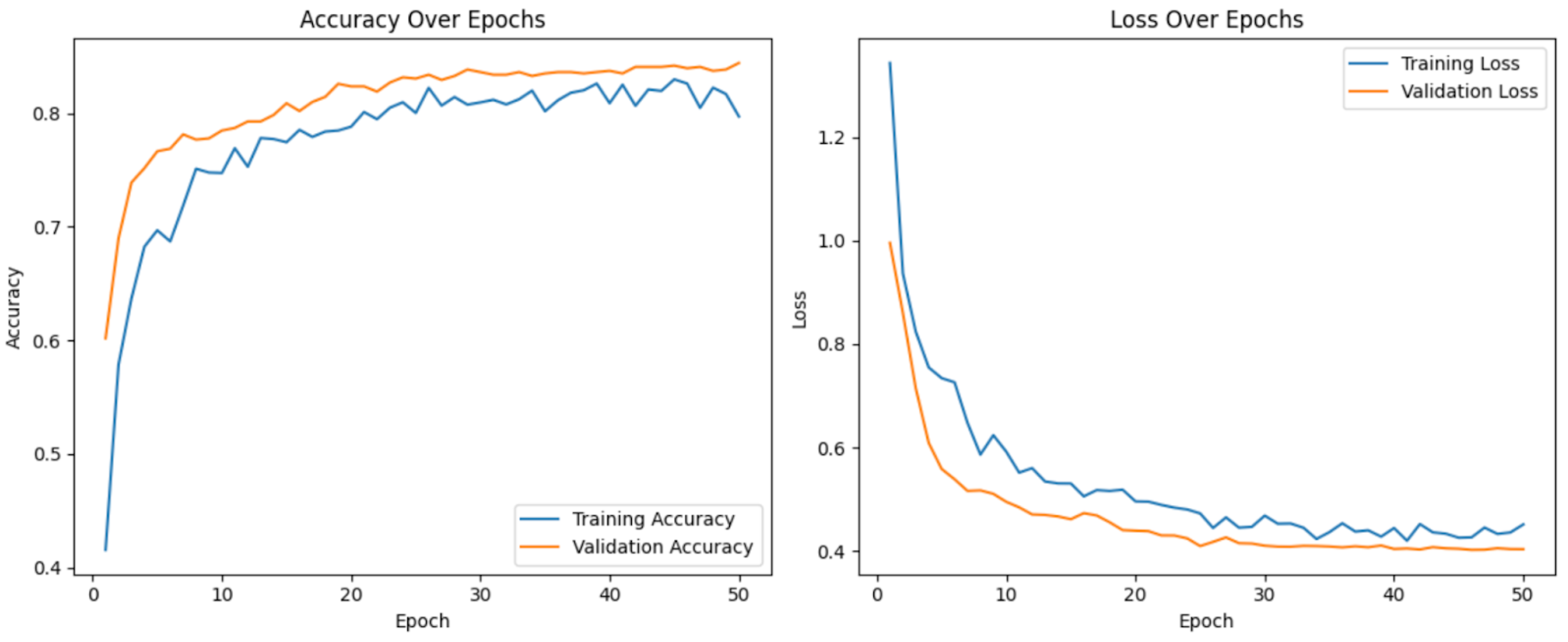
4.3. Training Curves
- The steady decrease in training loss indicates that the model is effectively learning from the dataset.
- The validation loss follows a similar trend, suggesting no significant overfitting. However, slight fluctuations in validation loss after epoch 10 suggest that further fine-tuning or regularization techniques could further improve generalization.
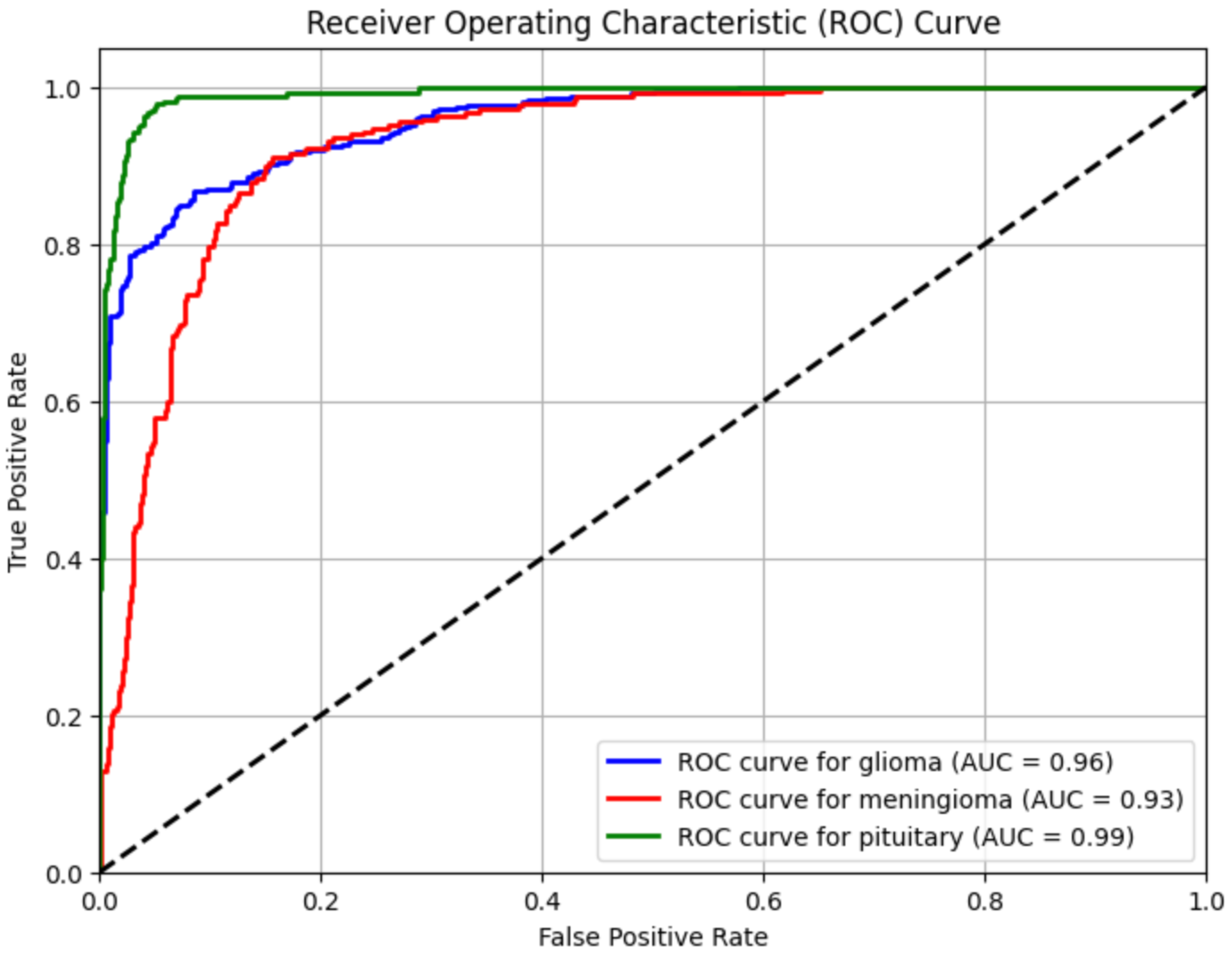
4.4. ROC Curve
- Glioma Tumor → AUC = 0.96
- Meningioma Tumor → AUC = 0.93
- Pituitary Tumor → AUC = 0.99
5. Evaluation Metrics
5.1. Classification Report
5.2. Confusion Matrix Analysis
5.3. ROC Curve Evaluation
5.4. Learning Curves
5.5. Model Explainability with Grad-CAM
5.6. Summary
6. Results and Discussion
6.1. Challenges Observed
- Class Imbalance: Despite class weighting and augmentation, minority classes (meningioma and pituitary) remained harder to classify in some experiments.
- Dataset Variability: Merging datasets introduced real-world diversity but also increased intra-class variability.
- Generalization: While validation metrics were strong, some alternate models (EfficientNet, MobileNet) underperformed on the test set, reinforcing the need for careful model selection and tuning.
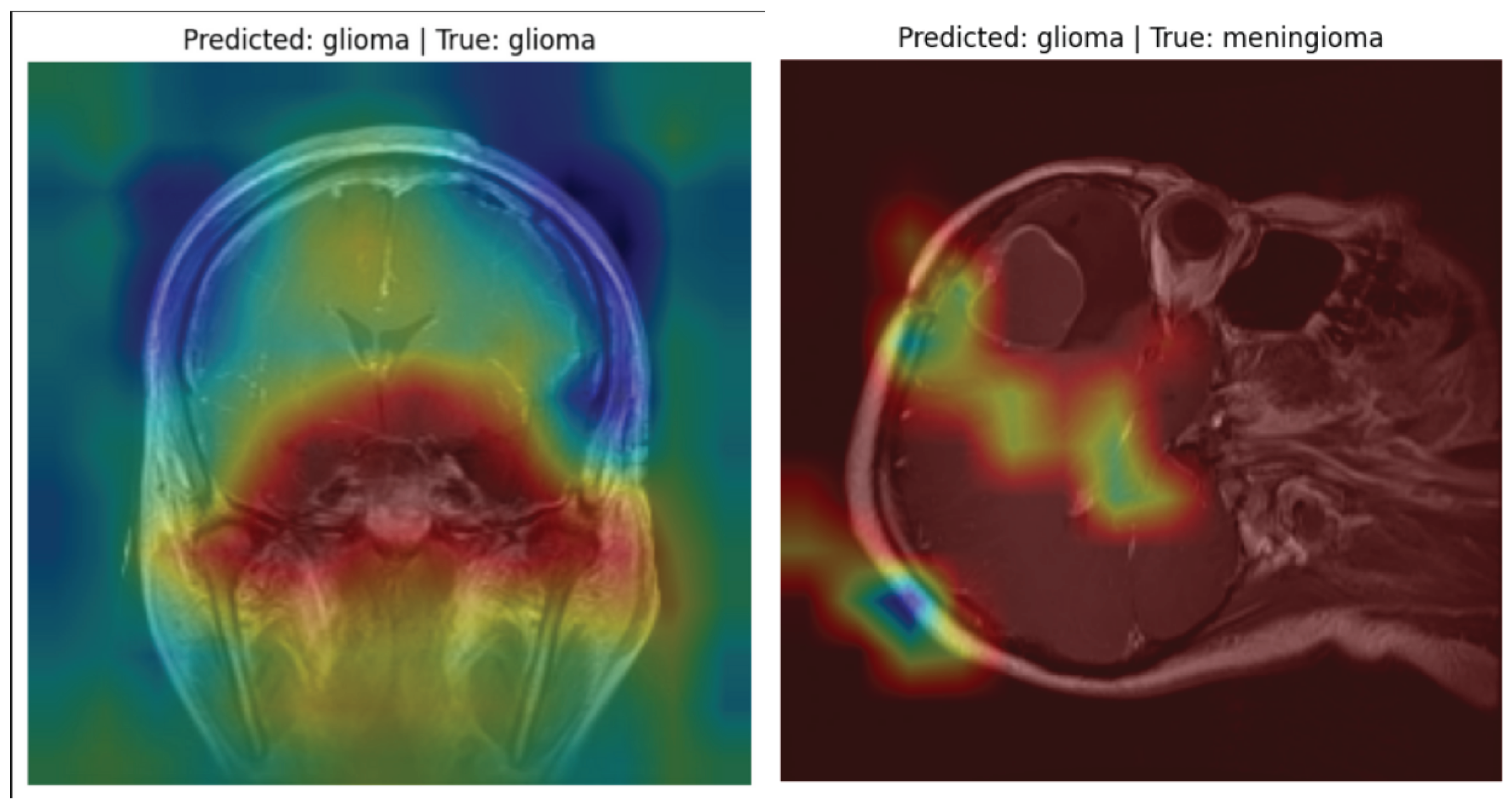
6.2. Model Explainability with Grad-CAM
- The heatmaps allow radiologists and researchers to verify whether the model is focusing on tumor regions.
- In correctly classified cases, Grad-CAM activation maps strongly corresponded with visible tumor boundaries.
- Even in misclassified samples, the model highlighted regions of abnormal tissue, showing it was identifying suspicious areas, though not always matching the ground-truth label.
7. Conclusions
References
- Abdusalomov, A.B.; Mukhiddinov, M.; Whangbo, T. Brain tumor detection based on deep learning approaches and magnetic resonance imaging. Cancers 2023, 15, 4172. [Google Scholar] [CrossRef] [PubMed]
- Amin, J.; Sharif, M.; Haldorai, A.; Yasmin, M.; Nayak, R.S. Brain tumor detection and classification using machine learning: A comprehensive survey. Complex & Intelligent Systems 2021, 8, 3161–3183. [Google Scholar] [CrossRef]
- Dulal, R.; Dulal, R. Brain tumor identification using improved YOLOv8, 2025. arXiv preprint.
- Khan, A.H.; Abbas, S.; Khan, M.A.; Farooq, U.; Khan, W.A.; Siddiqui, S.Y.; Ahmad, A. Intelligent model for brain tumor identification using deep learning. Applied Computational Intelligence and Soft Computing, 2022; 1–10. [Google Scholar] [CrossRef]
- Zahoor, A.; Malik, H.; Khan, S. Res-BRNet: A novel residual and boundary-aware network for brain tumor classification in MRI. Expert Systems with Applications 2024, 221, 119932. [Google Scholar] [CrossRef]
- Chen, R.; Zhang, X.; Li, P.; Wang, L. YOLO-NeuroBoost: Enhancing real-time object detection for brain tumor MRI scans. IEEE Transactions on Biomedical Engineering 2024, 71, 1125–1137. [Google Scholar] [CrossRef]
- Parida, A.; Capellán-Martín, D.; Jiang, Z.; Tapp, A.; Liu, X.; Anwar, S.M.; Ledesma-Carbayo, M.J.; Linguraru, M.G. Adult glioma segmentation in Sub-Saharan Africa using transfer learning on stratified fine-tuning data, 2024. arXiv preprint.
- Reddy, S.; Kumar, P.; Sharma, N. Fine-tuned vision transformers for multi-class brain tumor classification. Neural Computing and Applications 2024, 36, 517–531. [Google Scholar] [CrossRef]
- Krishnan, H.; Patel, S.; Gupta, R. RViT: A rotation-invariant vision transformer for brain tumor MRI classification. Medical Image Analysis 2024, 92, 102313. [Google Scholar] [CrossRef]
- Bhuvaji, S.; Kanchan, S.; Dedge, S.; Bhumkar, P.; Kadam, A. Brain tumor classification (MRI), 2020. Dataset on Kaggle.
- Cheng, J. Brain tumor dataset, 2017. Dataset on Figshare. [CrossRef]
| Study | Pre-Trained Model | Proposed Model | Dataset | Accuracy (%) |
|---|---|---|---|---|
| Khan et al. (2022) | ResNet-50 | N/A (Baseline Model) | Figshare Brain MRI Dataset | 96.50% |
| Zahoor et al. (2024) | ResNet-18, VGG16 | Res-BRNet (Boundary-Aware CNN) | Kaggle + Br35H (10k images) | 98.20% |
| Reddy et al. (2024) | ViT-B16, ViT-L32 | FTVT (Fine-Tuned ViTs) | Br35H (7,023 MRI images) | 98.70% |
| Krishnan et al. (2024) | ViT-B16 | RViT (Rotation-Invariant ViT) | Kaggle Brain MRI | 98.60% |
| Chen et al. (2024) | YOLOv8 | YOLO-NeuroBoost | Br35H, Roboflow MRI Dataset | 99.50% |
| Abdusalomov et al. (2023) | YOLOv7 | CBAM-YOLOv7 (Enhanced Detector) | Large MRI Dataset (10,288 images) | 99.40% |
| Current Study (2025) | VGG16 | VGG16 + Custom Dense Classifier (Fine-Tuned) | Kaggle + Figshare Brain MRI | 84.40% |
| Classification Report: | Precision | Recall | F1-Score | Support |
|---|---|---|---|---|
| glioma | 0.91 | 0.81 | 0.86 | 353 |
| meningioma | 0.78 | 0.74 | 0.76 | 247 |
| pituitary | 0.83 | 0.99 | 0.90 | 275 |
| accuracy | - | - | 0.85 | 875 |
| macro avg | 0.84 | 0.85 | 0.84 | 875 |
| weighted avg | 0.85 | 0.85 | 0.85 | 875 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




